Abstract
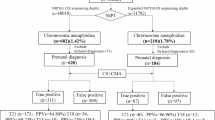
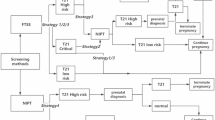
Non-invasive prenatal testing (NIPT) as a first-tier screening approach at population scale warrants close evaluation of its accessibility, affordability and extended application value. Here we present insights from a fully publicly funded NIPT programme for the pregnant population in Hebei province, China (July 2019–June 2022), with the latest NIPT coverage of 93.3%. Among 1,185,416 pregnant women, the incidences of fetal chromosome aneuploidies were 0.105% (trisomy 21), 0.026% (trisomy 18), 0.005% (trisomy 13), 0.056% (47,XXY), 0.043% (45,X), 0.039% (47,XYY) and 0.034% (47,XXX). Parental advanced age (≥35 years old) was related to a 4.31-fold risk for fetal trisomy 21 compared to parental non-advanced age. An elevated risk for small vulnerable newborns was observed in NIPT false-positive (relative risk: 1.49 (1.31, 1.68)) and NIPT high-risk without invasive prenatal diagnosis (relative risk: 1.85 (1.57, 2.18)) cases. NIPT reduced prenatal diagnosis by 97.4% compared to maternal serum screening, with better cost-effectiveness. Our findings indicate that first-tier NIPT for all pregnant women could be a feasible and cost-effective alternative to traditional maternal serum screening. NIPT positive results may be an early indicator of more adverse pregnancy outcomes beyond fetal aneuploidies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The primary data from this study are not publicly available owing to restrictions in the informed consent obtained from study participants, which do not allow for public data sharing. De-identified data may be made available from J.G. upon reasonable request and following approval by the relevant institutional and ethics committees. Requests will be considered, reviewed and actioned within 1 month of receipt. The data that support the findings of this study have been deposited in the CNGB Sequence Archive (CNSA) of the China National GeneBank DataBase (CNGBdb) under accession number CNP0006003.
Code availability
The code used for participant selection, data processing and analysis can be made available from J. Zhang upon reasonable request within 1 month of receipt.
References
Ravitsky, V. et al. The emergence and global spread of noninvasive prenatal testing. Annu. Rev. Genomics Hum. Genet. 22, 309–338 (2021).
Zhang, H. et al. Non-invasive prenatal testing for trisomies 21, 18 and 13: clinical experience from 146,958 pregnancies. Ultrasound Obstet. Gynecol. 45, 530–538 (2015).
Spencer, R. et al. Non-invasive prenatal testing for aneuploidy screening. BMJ 371, m3930 (2020).
Guy, G. P. et al. Secondary non-invasive prenatal screening for fetal trisomy: an effectiveness study in a public health setting. BJOG 128, 440–446 (2021).
Dar, P. et al. Cell-free DNA screening for trisomies 21, 18, and 13 in pregnancies at low and high risk for aneuploidy with genetic confirmation. Am. J. Obstet. Gynecol. 227, 251–259 (2022).
Battarbee, A. N. et al. Cost-effectiveness of ultrasound before non-invasive prenatal screening for fetal aneuploidy. Ultrasound Obstet. Gynecol. 61, 325–332 (2023).
Garshasbi, M. et al. Clinical application of cell-free DNA sequencing-based noninvasive prenatal testing for Trisomies 21, 18, 13 and sex chromosome aneuploidy in a mixed-risk population in Iran. Fetal Diagn. Ther. 47, 220–227 (2020).
Burgess, R. et al. Implementing publicly funded noninvasive prenatal testing for fetal aneuploidy in Ontario, Canada: clinician experiences with a disruptive technology. Qual. Health Res. 30, 1941–1952 (2020).
Dungan, J. S. et al. Noninvasive prenatal screening (NIPS) for fetal chromosome abnormalities in a general-risk population: an evidence-based clinical guideline of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 25, 100336 (2023).
Dugoff, L. et al. Cell-free DNA screening for trisomy 21 in twin pregnancy: a large multicenter cohort study. Am. J. Obstet. Gynecol. 229, 435.e1–435.e7 (2023).
Gadsbøll, K. et al. Current use of noninvasive prenatal testing in Europe, Australia and the USA: a graphical presentation. Acta Obstet. Gynecol. Scand. 99, 722–730 (2020).
Sium, A. F. et al. Indications, types, and diagnostic implications of prenatal genetic testing in Sub-Saharan Africa: a descriptive study. PLoS ONE 18, e0294409 (2023).
van der Meij, K. et al. TRIDENT-2: national implementation of genome-wide non-invasive prenatal testing as a first-tier screening test in the Netherlands. Am. J. Hum. Genet. 105, 1091–1101 (2019).
Ökem, Z. G. et al. Economic analysis of prenatal screening strategies for Down syndrome in singleton pregnancies in Turkey. Eur. J. Obstet. Gynecol. Reprod. Biol. 219, 40–44 (2017).
Wilmot, H. C. et al. Down syndrome screening and diagnosis practices in Europe, United States, Australia, and New Zealand from 1990–2021. Eur. J. Hum. Genet. 31, 497–503 (2023).
Chitty, L. et al. Uptake, outcomes, and costs of implementing non-invasive prenatal testing for Down’s syndrome into NHS maternity care: prospective cohort study in eight diverse maternity units. BMJ 354, i3426 (2016).
Xu, Y. et al. Cost-effectiveness analysis of non-invasive prenatal testing for Down syndrome in China. Int. J. Technol. Assess. Health Care 35, 237–242 (2019).
van der Meij, K. et al. Non-invasive prenatal test uptake in socioeconomically disadvantaged neighborhoods. Prenat. Diagn. 41, 1395–1400 (2021).
Wang, S. et al. A cost-effectiveness analysis of screening strategies involving non-invasive prenatal testing for trisomy 21. Front. Public Health 10, 870543 (2022).
Yang, L. & Tan, W. Prenatal screening in the era of non-invasive prenatal testing: a Nationwide cross-sectional survey of obstetrician knowledge, attitudes and clinical practice. BMC Pregnancy Childbirth 20, 579 (2020).
Kim, S. et al. Prospective observations study protocol to investigate cost-effectiveness of various prenatal test strategies after the introduction of noninvasive prenatal testing. BMC Pregnancy Childbirth 18, 307 (2018).
Liu, C. et al. Application of ultrasound combined with noninvasive prenatal testing in prenatal testing. Transl. Pediatr. 11, 85–98 (2022).
Van Den Bogaert, K. et al. Outcome of publicly funded nationwide first-tier noninvasive prenatal screening. Genet. Med. 23, 1137–1142 (2021).
Cuckle, H. & Benn, P. Review of epidemiological factors (other than maternal age) that determine the prevalence of common autosomal trisomies. Prenat. Diagn. 41, 536–544 (2021).
Bu, X., Ye, W. & Zhou, J. Paternal age, risk of congenital anomalies, and birth outcomes: a population-based cohort study. Eur. J. Pediatr. 182, 3519–3526 (2023).
Donate, A. et al. Paternal age and numerical chromosome abnormalities in human spermatozoa. Cytogenet. Genome Res. 148, 241–248 (2016).
Loughry, L., Pynaker, C., White, M., Halliday, J. & Hui, L. State-wide increase in prenatal diagnosis of Klinefelter syndrome on amniocentesis and chorionic villus sampling: impact of non-invasive prenatal testing for sex chromosome conditions. Prenat. Diagn. 43, 156–161 (2023).
Elmerdahl Frederiksen, L. et al. Maternal age and the risk of fetal aneuploidy: a nationwide cohort study of more than 500,000 singleton pregnancies in Denmark from 2008 to 2017. Acta Obstet. Gynecol. Scand. 103, 351–359 (2024).
Chen, X. et al. Trends in the prevalence, prenatal diagnosis, and outcomes of births with chromosomal abnormalities: a hospital-based study in Zhejiang Province, China during 2014–2020. Orphanet J. Rare Dis. 17, 446 (2022).
Franasiak, J. M. et al. The nature of aneuploidy with increasing age of the female partner: a review of 15,169 consecutive trophectoderm biopsies evaluated with comprehensive chromosomal screening. Fertil. Steril. 101, 656–663 (2014).
Sánchez, X. C. et al. Associations of psychiatric disorders with sex chromosome aneuploidies in the Danish iPSYCH2015 dataset: a case-cohort study. Lancet Psychiatry 10, 129–138 (2023).
van Rijn, S. A review of neurocognitive functioning and risk for psychopathology in sex chromosome trisomy (47,XXY, 47,XXX, 47,XYY). Curr. Opin. Psychiatry 32, 79–84 (2019).
Dowlut-McElroy, T. et al. Cell-free DNA screening positive for monosomy X: clinical evaluation and management of suspected maternal or fetal Turner syndrome. Am. J. Obstet. Gynecol. 227, 862–870 (2022).
Butler, G. et al. Klinefelter syndrome: going beyond the diagnosis. Arch. Dis. Child 108, 166–171 (2022).
Samango-Sprouse, C. et al. Incidence of sex chromosome aneuploidy in a prenatal population: 27-year longitudinal study in Northern Italy. Ultrasound Obstet. Gynecol. 62, 266–272 (2023).
Xiang, J. et al. Clinical impacts of genome-wide noninvasive prenatal testing for rare autosomal trisomy. Am. J. Obstet. Gynecol. MFM 5, 100790 (2023).
van Prooyen Schuurman, L. et al. Clinical impact of additional findings detected by genome-wide non-invasive prenatal testing: follow-up results of the TRIDENT-2 study. Am. J. Hum. Genet. 109, 1140–1152 (2022).
Becking, E. C. et al. Fetal fraction of cell-free DNA in noninvasive prenatal testing and adverse pregnancy outcomes: a nationwide retrospective cohort study of 56,110 pregnant women. Am. J. Obstet. Gynecol. 231, 241–244 (2023).
United Nations, Department of Economic and Social Affairs, Population Division. World Population Prospects 2022. https://population.un.org/wpp/Download/Standard/Fertility/ (2022).
de Graaf, G. et al. Estimation of the number of people with Down syndrome living in Canada. Genet. Med. 27, 101422 (2025).
Stanley, K. E., Thienpont, B. & Vermeesch, J. R. Expanding the scope of non-invasive prenatal screening. Nat. Genet. 57, 2644–2654 (2025).
Zhang, J. et al. Prospective prenatal cell-free DNA screening for genetic conditions of heterogenous etiologies. Nat. Med. 30, 470–479 (2024).
Heesterbeek, C. J. et al. Prenatal cell-free DNA testing of women with pregnancy-associated cancer: a retrospective cross-sectional study. Lancet Reg. Health Eur. 45, 101024 (2024).
Turriff, A. E. et al. Prenatal cfDNA sequencing and incidental detection of maternal cancer. N. Engl. J. Med. 391, 2123–2132 (2024).
Adil, M. et al. Preeclampsia risk prediction from prenatal cell-free DNA screening. Nat. Med. 31, 1312–1318 (2025).
Guo, Z. et al. Genome-wide nucleosome footprints of plasma cfDNA predict preterm birth: a case-control study. PLoS Med. 22, e1004571 (2025).
Ashorn, P. et al. Small vulnerable newborns-big potential for impact. Lancet 401, 1692–1706 (2023).
Acknowledgements
We thank all the pregnant women who participated in the project. We thank the related staff of the 12 specified PD institutions and 288 specified prenatal testing institutions. We thank the leading group and technical expert group of Hebei NIPT project. This study was supported by the S&T Program of Hebei (grant nos. 215A9907D to S.H. and 21377722D to Y. Zhou) and the S&T Program of Shijiazhuang (grant no. 235790429H to D.T.). The funders had no role in the study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J. Zhang, L.Z., J.G., B.Z. and J. Zhu are joint senior authors and designed the study. J.L., M.W., S.H., B.Z. and J. Zhang drafted the paper. M.W., L.Y., L.T., G.H., H.J. and B.Z. contributed to the data analysis. C.Z., Yunfang Wang, Y. Zhang, Yuzhen Wang, L.C., C.G., H.L., H.X. and B.Y. contributed to the implementation of the project. S.H., Y.L., Y. Zhou, Q.H., R.D., D.T. and S.G. contributed to the data collection. J.L., J. Zhao, Y.C., J. Zhang, L.Z., J.G., B.Z. and J. Zhu contributed to the organization and supervision of the project. All authors were responsible for reviewing and revising the article. The corresponding author attests that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Health thanks David Barad and Ming Chen for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Manonmani Soundararajan, in collaboration with the Nature Health team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 First-tier NIPT coverage for detecting fetal common trisomies.
PD: prenatal diagnosis. Regions were categorized by socioeconomic statuses based on GDP per capita from nationwide and regional public data in 2019 (cities of top3 GDP were defined as advantaged, cities of last 3 GDP were defined as disadvantaged, and the remaining cities were defined as intermediate). The sample size for estimating NIPT coverage (left figure) was 1,529,416 (all registered pregnancies). The sample size for PD rate (right figure) was 2,309 (NIPT high-risk cases on T21, T18, T13).
Extended Data Fig. 2 Distribution of NIPT high-risk for rare autosomal aneuploidies.
n = 953. The number of NIPT high-risk cases on each autosomal chromosome (except chromosome 13, 18, 21) was labeled above the bar. The black bar presented the prenatal diagnosis confirmed fetal RAA, and the gray bar was women without prenatal diagnosis.
Supplementary information
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Table 2 (download XLSX )
Statistical source data.
Extended Data Figs. 1 and 2 and Tables 1 and 2 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Wu, M., He, S. et al. Large-scale implementation of non-invasive prenatal testing for fetal aneuploidies. Nat. Health (2026). https://doi.org/10.1038/s44360-026-00104-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44360-026-00104-2