Abstract
This study aimed to highlight the molecular and biochemical changes induced by methylglyoxal (MGO) exposure in SH-SY5Y human neuroblastoma cells, and to explore how these changes contribute to its neurotoxicity, utilizing an integrated proteomics and metabolomics approach. Using label-free quantitative nanoLC-MS/MS proteomics and targeted LC-TQ-MS/MS-based metabolomics, the results revealed that MGO exposure, particularly at cytotoxic levels, significantly altered the proteome and metabolome of SH-SY5Y cells. Analysis of proteomics data showed significant alterations in cellular functions including protein synthesis, cellular structural integrity, mitochondrial function, and oxidative stress responses. Analysis of metabolomics and integration of metabolomics and proteomics data highlighted significant changes in key metabolic pathways including arginine biosynthesis, glutathione metabolism, cysteine and methionine metabolism, and the tricarboxylic acid cycle. These results suggest that MGO exposure induced both toxic effects and adaptive responses in cells. MGO exposure led to increased endoplasmic reticulum stress, disruptions in cellular adhesion and extracellular matrix integrity, mitochondrial dysfunction, and amino acid metabolism disruption, contributing to cellular toxicity. Conversely, cells exhibited adaptive responses by upregulating protein synthesis, activating the Nrf2 pathway, and reprogramming metabolism to counteract dicarbonyl stress and maintain energy levels. Furthermore, a set of key proteins and metabolites associated with these changes were shown to exhibit a significant concentration-dependent decrease or increase in their expression levels with increasing MGO concentrations, suggesting their potential as biomarkers for MGO exposure. Taken together, these findings provide insight into the molecular mechanisms underlying MGO-induced neurotoxicity and potential targets for therapeutic intervention.
Similar content being viewed by others
Introduction
Neurodegenerative disorders, such as Alzheimer’s disease (AD) and Parkinson’s disease (PD), are increasingly prevalent globally due to aging populations and the lack of effective therapies1. The pathogenic mechanisms underlying neurodegeneration are complex, involving various characteristics like oxidative stress, mitochondrial dysfunction, inflammation, and protein misfolding and aggregation2,3,4. The process of protein glycation, resulting in irreversible modifications in proteins, has been suggested to play important roles in these multifactorial diseases5,6. Protein glycation is a post-translational modification wherein reducing sugars or reactive dicarbonyl compounds, notably methylglyoxal (MGO) and glyoxal (GO), react non-enzymatically with amino acid residues on proteins, forming advanced glycation end products (AGEs)7. These modifications can alter protein structure and dynamics and induce the formation of protein cross-links, potentially promoting or stabilizing pathological protein aggregation, thereby contributing to neurodegeneration8,9. Furthermore, AGEs could induce cell dysfunction through interacting with the receptor for AGEs (RAGE), triggering oxidative stress and inflammatory responses10.
MGO, extensively found in various food products and endogenously formed as a by-product primarily from glycolysis, is the most reactive precursor for AGE formation both endogenously and exogenously11,12,13. MGO concentrations are typically low in healthy individuals due to the relatively low internal production and effective detoxification by the glyoxalase system14. Elevated in vivo MGO levels can occur under certain pathological conditions such as hyperglycemia, or exposure to high levels of exogenous glucose or MGO15,16. An increase in the steady-state level of reactive dicarbonyls, especially MGO, is the main cause of a dysfunctional metabolic state called dicarbonyl stress, resulting in increased modifications of cellular amino acids and biomacromolecules, including proteins and DNA, forming AGEs15.
Clinical research has revealed a correlation between elevated serum levels of MGO and accelerated cognitive decline in elderly individuals17 and found increased levels of MGO and the resulting AGEs in the cerebrospinal fluid of AD patients18,19. Similarly, increased levels of MGO-derived AGEs have been detected in the nigral neurons of PD patients20. In an animal study, aged mice orally treated with 100 mg/kg bw MGO for 4 weeks developed cognitive impairment, accompanied by increased expression of RAGE and Presenilin-1 (a critical component of the gamma-secretase complex involved in AD) in the hippocampus21. Additionally, in Caenorhabditis elegans, long-term dietary exposure to MGO led to AD-related toxicity including reduced locomotive behaviors and promoted amyloid β accumulation22. These clinical and animal findings underscore the pathological relevance of MGO in the context of neurodegeneration. Furthermore, in vitro studies using different neuronal cell models, such as the human SH-SY5Y and SK-N-MC neuroblastoma cell line have shown that exposure of cells to exogenous MGO reduced cell viability and mitochondrial redox activity, as well as increased the production of reactive oxygen species (ROS) and apoptosis23,24,25,26. Recent studies using physiologically relevant in vitro neuronal-like cells transdifferentiated from human mesenchymal stem cells and 3D human stem-cell-derived neuronal spheroids demonstrated that MGO at low μM concentrations—comparable to human sample levels measured in MGO-associated diseases—altered key neuronal markers, including MAP-2 and NSE, involved in biochemical pathways relevant to neurodegenerative diseases and aging27,28. However, the modes of action underlying these toxic effects induced by MGO, which may play roles in neurodegeneration, remain largely unclear.
Given that the neurotoxic effects of MGO may result from its complex interactions with free cellular amino acids and biomacromolecules, causing damage across the cellular metabolome, proteome, and genome29,30, traditional toxicological methods that focus on limited endpoints may not adequately capture the broader biological or toxicological impacts of MGO, especially the alterations in gene and protein expression in response to MGO exposure. Omics techniques, such as transcriptomics, proteomics, and metabolomics, offer a more comprehensive view, enabling the analysis of thousands of molecules in a single biological sample31, which could potentially enhance our understanding of the toxicological mechanisms and adverse outcome pathways associated with MGO exposure. A previous study using 2D fluorescence difference gel electrophoresis coupled with MALDI-TOF-MS identified 49 differentially expressed proteins (DEPs) in MGO-treated SH-SY5Y cells32. However, this gel-based proteomics approach is limited by its inability to detect low-abundance proteins and its ineffectiveness in resolving complex protein mixtures, which could lead to overlooking critical biomarkers33.
In contrast, in the present study, we utilized an untargeted, label-free quantitative proteomics approach with nanoLC-MS/MS to explore the cellular mechanisms of neurotoxicity induced by MGO in SH-SY5Y cells, a widely recognized neuronal cell model for investigating the pathological effects of neurotoxic compound34. This method substantially broadens the detectable proteome, providing a more detailed and comprehensive proteomic profile that could capture subtle changes, including those involving low-abundance proteins35. Additionally, a targeted metabolomics approach using LC-TQ-MS/MS was used to investigate the impact of MGO on the cellular metabolome, offering insights into metabolic alterations and the overall metabolic state of the cells. Integrating both proteomics and metabolomics could provide a holistic view of the phenotypic changes and the complex molecular interactions between proteins and metabolites triggered by MGO exposure, enabling a deeper exploration into the disrupted cellular pathways and their potential contributions to MGO neurotoxicity.
Materials and methods
Chemicals and reagents
MGO (40 wt% in water), acetic acid (purity ≥ 99%), tris base, and hydrochloric acid (37%) were purchased from Merck (Darmstadt, Germany). Methanol and acetonitrile (ACN) were obtained from Biosolve BV (Valkenswaard, The Netherlands). Formic acid (purity ≥ 99%) was purchased from VWR CHEMICA (Amsterdam, The Netherlands). Dulbecco’s Modified Eagle Medium/Nutrient Mixture F-12 (DMEM/F12) with GlutaMAX supplement cell culture medium with and without phenol red, trypsin-EDTA for cell dissociation, phosphate-buffered saline (PBS), penicillin/streptomycin, and nonessential amino acids (NEAA) were purchased from Gibco (Paisley, UK). Fetal calf serum (FCS) was purchased from Invitrogen (Breda, The Netherlands). BCA protein assay kit and protease inhibitor cocktail were obtained from Thermo Fisher Scientific (Bleiswijk, The Netherlands). Ultrapure water was prepared by a Milli-Q system (Millipore, MA, USA). All other reagents in this study were of analytical grade or purer.
Cell culture and MGO treatment
The human neuroblastoma cell line SH-SY5Y (ATCC CRL-2266) was obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). Cells were maintained in a humidified incubator at 37 °C and 5% CO2. The culture medium consisted of DMEM/F12 GlutaMAX supplemented with 10% FCS, 1% NEAA, and 1% penicillin/streptomycin. Medium was refreshed every 4–7 days, and cells were subcultured at a 1:20 ratio upon reaching confluency using a 0.05% trypsin-EDTA solution.
For the proteomics and metabolomics studies, three concentrations of MGO—500, 750, and 1000 μM—were used to treat the cells. The selection of the three MGO exposure concentrations, including both toxic and non-toxic levels, was made to ensure the elicitation of discernible cellular responses and to allow for the delineation of the concentration-dependent cellular impacts of MGO. These concentrations were chosen based on our previous MTT cell viability assay36, which showed that 500 and 750 μM MGO did not significantly affect cell viability, while at 1000 μM, there was a significant reduction, with 68.0% of viability remaining. The cell seeding density for the proteomics and metabolomics studies was the same as that used in the MTT assay.
Label-free quantitative proteomics analysis
Proteomic sample preparation
SH-SY5Y cells were seeded in T175 flasks (Greiner Bio-one, Alphen aan den Rijn, The Netherlands) at a density of 4.69 × 104 cells/cm2 and incubated for 24 h. After incubation, the cells were treated with an assay medium (DMEM/F12 without phenol red and FCS supplement) that contained MGO at three concentrations (500, 750, and 1000 μM, final concentration, added from 200 times concentrated stock solutions in nano pure water) or with a solvent control (0.5% nano pure water in DMEM/F12 without phenol red and FCS supplement) for 24 h. After exposure, the cells were washed twice with PBS. Subsequently, 1.8 mL of ice-cold PBS was added to each flask, and the cells were gently scraped off. The cell suspension was then transferred into a 2-mL protein low-binding tube (Eppendorf, Nijmegen, The Netherlands) and centrifuged at 500 × g for 2 min at 4 °C to collect the cell pellets. These pellets were washed twice using 1.5 mL ice-cold 100 mM Tris-hydrochloride (Tris-HCl, pH 8) under identical centrifugation conditions. The washed pellets were resuspended in 90 μL 100 mM Tris-HCl pH 8 with 1% protease inhibitor cocktail (Thermo Fisher Scientific, Bleiswijk, The Netherlands), followed by sonication in an ice-water bath (Qsonica sonicator, Qsonica, Newtown, USA) for 30 min using a pulse mode (30 s on and 30 s off) at 100% amplitude. Protein concentrations were measured using a BCA protein assay kit according to the manufacturer’s instructions (Thermo Fisher Scientific, Bleiswijk, The Netherlands). The Protein Aggregation Capture method was utilized to prepare peptide samples for nanoLC-MS/MS analysis37. These peptide samples were immediately stored at −20 °C until further analysis. All treatments were conducted in four independent biological replicates.
NanoLC-MS/MS-based label-free quantitative proteomics analysis
NanoLC-MS/MS analysis was conducted using a Thermo Vanquish Neo nanoLC system connected to a Thermo Orbitrap Exploris 480 mass spectrometer (Thermo Fisher Scientific, Bremen, Germany). An in-house prepared 0.10 mm × 250 mm analytical column filled with 1.9 μm ReproSil-Pur 120 C18-AQ beads was used. The mobile phase consisted of 0.1 v/v% formic acid in water (eluent A) and 0.1% formic acid in ACN (eluent B). The elution was carried out using a three-step linear gradient, starting from 0 to 9% B over 1 min, increasing from 9% to 34% B over 50 min, and then from 34% to 50% B over 4 min. This was followed by maintaining the gradient at 50% B for 1 min, after which it was returned to the initial conditions and held there for a duration equivalent to 2.5 column volumes at 925 bar to prepare for the next injection. The injection volumes were adjusted to correct for variations in protein concentrations among the samples, based on the BCA assay results, with injection volumes ranging from 1 to 2 µL. The elution flow rate was maintained at 500 nL/min. An electrospray potential of 3.5 kV was applied to the eluent prior to entering the column, and automatic gain control targets for MS and MS/MS were set to 300% and 100%, respectively, with maximum ion injection times capped at 50 ms for MS and 30 ms for MS/MS. Higher energy collisional dissociation was utilized to fragment the most abundant 2–5+ charged peaks identified in the MS scan, employing an isolation width of 1.2 m/z and 28% normalized collision energy, with MS/MS scans conducted in a data-dependent mode at a resolution of 15,000, applying a threshold of 5e4 and a 15-s exclusion duration for previously selected m/z ± 10 ppm.
Protein identification and quantification
The peptides were identified and quantified using the MaxQuant software package (version 2.0.3.0). A human database (UP000005640) retrieved from UniProt along with a contaminants database that contains sequences of common contaminants like Trypsins (P00760, bovine and P00761, porcin) were utilized for analysis38. Data were filtered to include only the proteins that were reliably identified by at least two peptides, at least one of which was unique, and one was unmodified and had a false discovery rate (FDR) below 1% on both the protein and peptide level. Reversed hits were removed from the MaxQuant result table.
Targeted metabolomics analysis
Metabolite extraction
SH-SY5Y cells were seeded in 6-well plates (Greiner Bio-One) at a density of 4.69 × 104 cells/cm2 and incubated for 24 h. For each sample for the metabolomics analysis, a parallel setup was implemented: one well designated for metabolite extraction and another for cell counting, with both wells undergoing identical exposure conditions as also used for the proteomics study and specified in the section proteomic sample preparation above. In the wells designated for metabolite extraction, metabolic activity was quenched by adding 500 µL 80% methanol (methanol, 80:20 v/v) at −70 °C. The cells were then carefully scraped while maintained on dry ice and transferred to a 2-mL tube (Eppendorf). Concurrently, cells in the parallel counting wells were detached using trypsin-EDTA, resuspended in 500 µL PBS (2% FCS), and cell numbers and concentrations were determined from a 20 µL aliquot using a Cellometer (Nexcelom Bioscience, Lawrence, USA). The suspension collected in the 2-mL tube from the wells designated for metabolite extraction was centrifuged at 6000 × g for 5 min, and the supernatants were collected in a new 2 mL Eppendorf tube. The cell pellets underwent two additional extractions with 500 µL 80% methanol each, with all extracts pooled directly into the same tube. These combined extracts were dried under nitrogen gas, reconstituted in 100 µL 50% methanol with 1 µL acetic acid, and centrifuged at 13,000 × g for 5 min to remove debris. The prepared samples were then immediately stored at −80 °C until LC-MS analysis. For all treatments four independent biological replicates were conducted.
LC-TQ-MS/MS-based quantification of metabolites
The metabolomics study for the cell metabolite extracts was performed using a Shimadzu Nexera XR LC-20AD XR UHPLC system coupled with a Shimadzu 8050 triple quadrupole mass spectrometer with an electrospray ionization interface (Shimadzu Corporation, Kyoto, Japan), accompanied by the LC/MS/MS Method Package for Primary Metabolites ver. 2 (Shimadzu Corporation). This method package offers optimized analytical conditions including chromatogram acquisition, detection of mass spectral peaks via an incorporated mass spectral library, and method files that specify the analytical conditions, which enables simultaneous quantification of 97 metabolites including amino acids, organic acids, nucleosides, and nucleotides. These metabolites are important in central cellular pathways such as the glycolytic system, tricarboxylic acid (TCA) cycle, and amino acid metabolism.
A Supelco Discovery HS F5-3 column (2.1 mm × 150 mm, 3 μm) was employed during the experiment with a flow rate of 0.25 mL/min. Ultrapure water containing 0.1% formic acid (A) and ACN with 0.1% formic acid (B) were used as mobile phase. The following gradient was used: 0–2 min, 100% A; 2–5 min, 100–75% A; 5–11 min, 75–65% A; 11–15 min, 65–5% A; 15–20 min, 5% A; 20–20.1 min, 5–100% A; 20.1–28 min, 100% A. The injection volume was 1 μL. The analysis settings for MS were determined according to the manufacturer’s instructions included in the method package for primary metabolites ver. 2 (Shimadzu Corporation).
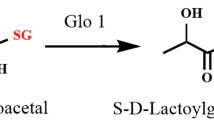
In addition to the original Shimadzu method which quantifies 97 primary metabolites, we have also incorporated the quantification of methylglyoxal-derived hydroimidazolone 1 (MG-H1). MG-H1 is the most abundant free-form AGE in vivo, formed from MGO and arginine39. It was monitored at the [M + H]+ of precursor to product ion transitions of m/z 229.0 > 114.0 (collision energy = −10 eV).
Statistical analysis
Data pre-processing
For proteomics data, zero label-free quantitation (LFQ) intensity values were replaced with a value of 107 (slightly lower than the lowest measured value) to allow ratio calculations, and LFQ intensities were log10 scaled for further statistical analysis.
For metabolomics data, peak identification and integration were carried out using LabSolutions software 5.114 (Shimadzu Corporation) to obtain intensity values for detected metabolites. Intensity values of metabolites in each sample were divided by the corresponding cell count to correct for variations in cell numbers, ensuring comparable metabolite measurements across samples. Data were normalized using a pooled sample from the control group, then log-transformed and mean-centered using MetaboAnalyst 6.0 (https://www.metaboanalyst.ca/)40.
Multivariate analysis
Principal component analysis (PCA) was conducted on the log-transformed LFQ intensities for proteomics data using the prcomp function from the stats package in R software 4.3.141, with data scaled to normalize variance across different measurements. For metabolomics data, PCA was applied on the pre-processed data using the same function in R without scaling to unit variance.
Univariate analysis
The fold change (FC) of proteins and metabolites was calculated as the ratio of the average intensities of each MGO-treated group to that of the control. Student’s t-test was performed on the respective datasets to compare differences between the control and different concentrations of MGO-treated groups, with p values below 0.05 considered statistically significant. Proteomics data were analyzed using Perseus version 1.6.2.142, while metabolomics data were analyzed using MetaboAnalyst.
To determine if there was a concentration-dependent increase or decrease in the expression levels of proteins and metabolites, Jonckheere–Terpstra test, a non-parametric method used for identifying ordered differences among groups43, was performed. Results with p values below 0.05 were considered to be statistically significant.
Selection of differentially expressed proteins (DEPs) and differential metabolites (DMs)
DEPs and DMs across varying concentrations of MGO-treated groups were identified using the same selection criteria. Both DEPs and DMs were chosen based on FCs >1.2 or <0.83, accompanied by p values <0.05 relative to the control group.
Pathway and network analysis
For proteomics data, enrichment analysis for DEPs was conducted using DAVID v2023q4 (https://david.ncifcrf.gov/, accessed April 2024), including GO terms, KEGG pathways, and WikiPathways44. A protein–protein interaction (PPI) network was constructed for DEPs with a high confidence level set at 0.7 utilizing the STRING database 12.0 (https://string-db.org/, accessed April 2024)45. Molecular Complex Detection (MCODE) (version 2.0.3)46 in Cytoscape (version 3.10.2)47 was used to identify key functional modules in PPI network with parameters set as follows: degree cutoff = 2, node score cutoff = 0.2, K-core = 2, and max depth = 100. Hub proteins were identified using the CytoHubba plugin with the maximal clique centrality (MCC) method48 in Cytoscape.
For metabolomics data, pathway enrichment analysis of DMs identified from the high-concentration MGO-treated group (1000 μM MGO) was performed using MetaboAnalyst according to the KEGG database. Pathways with pathway impact greater than 0.20 and p values < 0.05 were considered significant.
Integrated analyses of metabolomics and proteomics
To better understand the interactions between proteins and metabolites, an integrated analysis of metabolomics and proteomics data was conducted, including correlation analysis and joint pathway analysis. Given the large number of DEPs, the correlation analysis focused on all DMs and only the proteins identified in the top 5 key functional modules in PPI network, both derived from the high-concentration MGO-treated group. Correlation between proteomics and metabolomics data was quantified using Spearman’s correlation (cor.test() function in R) using only proteins identified in the top 5 key functional modules in PPI network and all DMs. The matrices of median concentrations of each treatment group were merged column-wise before taking pair-wise correlation among proteins and metabolites. Only correlation >0.75 (in absolute value) and with p values < 0.01 were retained for analysis and visualized using Cytoscape.
Subsequently, DEPs and DMs were subjected to joint pathway analysis using MetaboAnalyst based on KEGG database. Pathways with an impact >0.20 and p values < 0.05 were deemed significant.
Data visualization
The bar charts, enrichment results, heatmaps depicting the intensity of DEPs and DMs, and volcano plots illustrating these entities were visualized through the SRplot webserver (http://www.bioinformatics.com.cn/srplot)49, accessed on April 2024.
Results
Effects of MGO exposure on the proteome of SH-SY5Y cells
To assess the effects of MGO exposure on the proteomic profile of SH-SY5Y cells and to provide insight into the pathways disturbed by the exposure, quantitative proteomics was performed using MGO concentrations of 500, 750, and 1000 µM, with the highest concentration being cytotoxic as determined by our previous MTT viability assay36. In total, 3160 proteins were identified and quantified using a 1% FDR filter. PCA was performed on these proteins. The 3D PCA plot, as shown in Fig. 1a, revealed a distinct separation between the MGO-treated groups and the control group. The PCA accounted for 55.8% of the variation in the protein profiles with PC1, PC2, and PC3. The variation observed suggested that MGO exposure, particularly at the highest MGO concentration, substantially altered the proteome in SH-SY5Y cells compared to the untreated control cells.
a 3D PCA plot based on the expression levels of 3160 proteins. b The number of upregulated and downregulated differentially expressed proteins (DEPs) in different MGO-treated groups. c Volcano plot of proteins quantified in different MGO-treated groups (red dots: upregulated DEPs, blue dots: downregulated DEPS, gray dots: proteins with no significant change). d Heatmap of the 453 DEPs identified in the high-concentration MGO-treated group presented for all treatment groups. Columns represent groups and rows represent proteins. The color gradient from blue to red indicates the protein expression level from low to high. Treatment groups are labeled as follows: control (C-x) for 0 μM MGO, low (L-x) for 500 μM MGO, medium (M-x) for 750 μM MGO, and high (H-x) for 1000 μM MGO. The “x” in each label represents the replicate number, ranging from 1 to 4.
Subsequently, DEPs in each MGO treatment group were identified by comparison to the control group (see section “Selection of differentially expressed proteins (DEPs) and differential metabolites (DMs)” for selection criteria). As shown in Fig. 1b, there were 93 DEPs (59 upregulated and 34 downregulated) in the low-concentration MGO-treated group (500 µM), 204 DEPs (108 upregulated and 96 downregulated) in the medium-concentration MGO-treated group (750 µM), and 453 DEPs (223 upregulated and 230 downregulated) in the high-concentration MGO-treated group (1000 µM). The volcano plots in Fig. 1c illustrate the distribution of DEPs in each MGO-treated group compared to the control group, and the expression levels of the 453 DEPs identified in the high-concentration MGO-treated group were visualized across all four treatment groups in a clustering heatmap (Fig. 1b). These results indicate that low-concentration MGO exposure (500 µM) had a lesser impact on the protein profile of SH-SY5Y cells, whereas exposure to higher concentrations, especially the highest tested concentration of 1000 µM, which is cytotoxic, substantially altered the protein expression profile. Considering the substantial changes in protein profile and the significant reduction in cell viability in the high-concentration MGO-treated group (1000 µM), subsequent analyses focused on this treatment group to further explore the biological functions and pathways perturbed by MGO, relevant to its neurotoxicity. The p values and FCs of the DEPs identified in the high-concentration MGO-treated group, relative to control, can be found in Supplementary Table 3. GO, KEGG, and WikiPathways analyses were conducted on DEPs identified in SH-SY5Y cells treated with the high concentration of MGO. The GO enrichment analysis (Fig. 2a) revealed significant impacts on ribosomal RNA processing and ribosome biogenesis, highlighting potential disruptions in protein synthesis. Alterations were also observed in cellular components such as the nucleoplasm and extracellular exosomes, suggesting changes in both intracellular architecture and extracellular communication. Molecular function changes, including those affecting RNA, protein, and nucleosomal DNA binding activities, underscored potential disruptions in molecular interactions. These findings emphasize the extensive cellular and molecular disturbances induced by MGO.
a GO enrichment analysis for the categories of biological process (BP), cellular component (CC), and molecular function (MF); the top six enriched terms for each category are shown, selected based on the p values. b KEGG pathway enrichment analysis. c WikiPathways enrichment analysis (top 20 enriched terms are shown according to the p values).
KEGG pathway analysis (Fig. 2b) further demonstrated the impact on ribosome biogenesis and protein processing in the endoplasmic reticulum (ER), essential for proper protein folding and synthesis. Additionally, pathways related to focal adhesion and the regulation of the actin cytoskeleton were also significantly affected, indicating changes crucial for cell morphology and structural integrity. MGO’s impact on pathways associated with diabetic cardiomyopathy underscored its potential role in metabolic dysregulation, a common feature in diabetes characterized by elevated MGO levels50. This was further evidenced by significant enrichments in metabolic pathways, biosynthesis of amino acids, and oxidative phosphorylation, all pointing to MGO’s extensive impact on cellular metabolism. WikiPathways analysis (Fig. 2c) revealed significant effects of MGO on the cellular oxidative stress response, particularly through the enrichment of the Nrf2 signaling pathway.
PPI analysis was conducted on DEPs identified from the high-concentration MGO-treated group to delve deeper into the essential proteins and their association with pathways significantly enriched, as analyzed above. The PPI network constructed with DEPs, consisting of 451 nodes and 1681 edges (Supplementary Fig. 1), was further analyzed using the MCODE plugin in Cytoscape to identify key functional modules. Based on their network scores, the top five densely connected modules were extracted and analyzed for their specific protein compositions (Fig. 3). All selected modules had MCODE scores >3.5. Enrichment analysis was subsequently performed for each module to further elucidate their associated biological functions (Table 1). Notably, Modules 1 and 2 were both linked to protein synthesis: Module 1 primarily involved proteins related to ribosome biogenesis, such as RBM28, TBL3, UTP4, and WDR3; while Module 2 focused on proteins involved in translation initiation, including EIF4A1, EIF1AX, and EIF3M. Module 3 was crucial for maintaining cell structure and mediating cell–matrix interactions, featuring proteins like ITGB1, COL1A1, and LAMB1. Module 4 played a key role in energy metabolism and mitochondrial function through oxidative phosphorylation, with proteins such as NDUFA7, UQCRB, and COX5B. Finally, Module 5 was linked to oxidative stress responses through the Nrf2 pathway, with critical proteins like NQO1, HMOX1, and GCLM.
Enrichment results for each module are presented in Table 1.
Using the CytoHubba plugin in Cytoscape, the top 10 hub proteins (Supplementary Fig. 2)—PES1, NOC2L, GTPBP4, PDCD11, NOP2, MRTO4, GNL3, GRWD1, RPF2, and TSR1—were identified based on their MCC scores and were all found to be localized in Module 1, indicating the importance of this module.
Overall, the enrichment and PPI network analysis results suggest that exposure of cells to MGO leads to notable changes in several key cellular processes; these changes include abnormal protein synthesis, compromised cellular structure integrity, metabolic dysfunction, and enhanced responses to oxidative stress.
Effects of MGO exposure on the metabolome of SH-SY5Y cells
Considering the effect of MGO on cellular metabolism as revealed in the proteomics study, a targeted metabolomic study using the same exposure conditions as those used in the proteomics study was conducted to further explore the cellular metabolic response upon MGO exposure. In total, 75 out of the 98 compounds were detected and quantified in the samples. The PCA plot showed a distinct separation between the MGO-treated groups and the control (Fig. 4a), indicating MGO exposure, especially at the highest MGO concentration, greatly altered the metabolic profiles in cells. Subsequently, DMs in each MGO treatment group were identified by comparison to the control group (see section “Selection of differentially expressed proteins (DEPs) and differential metabolites (DMs)” for selection criteria). As shown in Fig. 4b, there were 12 DMs (5 upregulated and 7 downregulated) in the low-concentration MGO-treated group (500 µM), 29 DMs (18 upregulated and 11 downregulated) in the medium-concentration MGO-treated group (750 µM), and 31 DMs (17 upregulated and 14 downregulated) in the high-concentration MGO-treated group (1000 µM). The volcano plots in Fig. 4c further illustrate the distribution of DMs in each MGO-treated group compared to the control, and the levels of the 31 DMs identified in the high-concentration MGO-treated group were visualized across all four treatment groups in a clustering heatmap (Fig. 4d). Among all the DMs, MG-H1 that is reported to be the most abundant in vivo free-form AGE39, appeared to be the most significantly changed compound in all the three MGO-treated groups compared to control (Fig. 4c). There was an increase of up to 64-fold in the intensity of MG-H1 in the high-concentration MGO-treated group compared to control (Supplementary Table 6). The p values and FCs for other DMs identified in the high-concentration MGO-treated group can also be found in Supplementary Table 6.
a 3D PCA plot based on the levels of 75 metabolites. b The number of upregulated and downregulated differential metabolites (DMs) in different MGO-treated groups. c Volcano plot of metabolites quantified in different MGO-treated groups (red dots: upregulated DMs, blue dots: downregulated DMs, gray dots: metabolites with no significant change). d Heatmap of the 31 DMs identified in the high-concentration MGO-treated group presented for all treatment groups. The color gradient from blue to red indicates the metabolite level from low to high. Treatment groups are labeled as follows: control (C-x) for 0 μM MGO, low (L-x) for 500 μM MGO, medium (M-x) for 750 μM MGO, and high (H-x) for 1000 μM MGO. The “x” in each label represents the replicate number, ranging from 1 to 4.
To further explore the metabolic pathways influenced by MGO, a pathway analysis based on the KEGG database was conducted on the DMs identified in the high-concentration MGO-treated group. Figure 5a shows that nine metabolic pathways were significantly enriched, especially arginine and proline metabolism, cysteine and methionine metabolism, glutathione (GSH) metabolism, arginine biosynthesis, alanine, aspartate and glutamate metabolism and TCA cycle. These findings suggest that MGO, particularly at cytotoxic levels, induced metabolic dysfunction in SH-SY5Y cells.
a Metabolic pathway enrichment analysis of identified DMs in the high-concentration MGO-treated group. Integrated analyses of proteomics and metabolomics, including: b Correlation network of DMs and proteins identified in the top 5 key functional modules, both derived from the high-concentration MGO-treated group. The network was generated based on Spearman’s correlation analysis using a significance threshold of p < 0.01 and an absolute correlation coefficient >0.75. Blue rectangle nodes represent proteins; green diamond nodes represent metabolites. Red edges indicate negative correlations, and blue edges indicate positive correlations; c joint pathway enrichment analysis of DEPs and DMs in the high-concentration MGO-treated group. Pathways with a pathway impact greater than 0.20 and a p value < 0.05 were considered significant.
Integrated analyses of proteomics and metabolomics in SH-SY5Y cells after MGO exposure
Integrated analyses of proteomics and metabolomics were conducted to further explore the interactions between DEPs and DMs. Initially, we assessed the correlation between proteins identified in the top 5 key functional modules (Fig. 3) and DMs, both derived from the high-concentration MGO-treated group. The resulting correlation network, depicted in Fig. 5b, revealed a dense web of interactions among the proteins and DMs. Importantly, the network underscored the significant involvement of key metabolic pathways such as GSH metabolism, cysteine and methionine metabolism, and the TCA cycle. Key metabolites highlighted in these pathways include GSH, cystathionine, and various TCA cycle intermediates.
Subsequently, a joint metabolic pathway enrichment analysis was performed to examine the effects of MGO exposure on DEPs and DMs further. This analysis revealed that arginine biosynthesis was the most significantly affected pathway, showing the highest pathway impact and the most notable p value (Fig. 5c). Moreover, the analysis confirmed significant impacts on the TCA cycle, GSH metabolism, and cysteine and methionine metabolism pathways, corroborating the initial findings suggested by the correlation network. These pathways, affected by MGO exposure in SH-SY5Y cells, are extensively discussed in the discussion section to elucidate their roles in cellular responses.
A further screening of potential protein and metabolite biomarkers for MGO exposure
For the proteins enriched in the top five densely connected modules (Table 1), as well as for DMs, both sourced from the high-concentration MGO-treated group, a Jonckheere–Terpstra test was performed on the pre-processed intensity data of these proteins and DMs across all treatment groups. This analysis was conducted to determine if the respective expression levels exhibited a concentration-dependent increase or decrease with increasing concentrations of MGO, enabling the identification of potential sensitive and reliable biomarkers for MGO exposure.
The results in Table 2 show that 31 proteins exhibited a significant concentration-dependent increase in expression levels with increasing MGO concentrations (p < 0.05), and 11 proteins showed a significant concentration-dependent decrease (p < 0.05). Proteins with a concentration-dependent increase in their levels with increasing MGO concentrations were mainly linked to terms including “ribosome biogenesis,” “rRNA processing,” “translational initiation,” and the “Nrf2 pathway,” suggesting an enhanced protein synthesis response and activation of oxidative stress defenses. In contrast, proteins with a concentration-dependent decrease in their levels with MGO concentrations were mainly associated with “focal adhesion” and “ECM–receptor interaction,” indicating potential disruptions in cell adhesion and extracellular matrix (ECM) integrity.
Similarly, 16 metabolites were found to exhibit a significant concentration-dependent increase of their levels with increasing MGO concentrations (p < 0.05), while 4 metabolites showed a significant concentration-dependent decrease (p < 0.05) (Table 3). The metabolites showing concentration-dependent increases are primarily related to the TCA cycle (including lactic acid, citric acid, isocitric acid, malic acid, fumaric acid, and 2-ketoglutaric acid) and GSH synthesis (5-glutamylcysteine and GSH).
Taken together, these results reveal that the majority of the proteins (enriched from the top 5 densely connected modules in Table 1) and DMs, both identified from the high-concentration MGO-treated group, exhibited concentration-dependent increases or decreases in response to MGO exposure. Proteins and metabolites that exhibited concentration-dependent changes in response to MGO exposure could serve as biomarkers, indicative of MGO-induced cellular responses.
Discussion
The objective of this study was to investigate the molecular and biochemical changes induced by MGO exposure in SH-SY5Y human neuroblastoma cells and to elucidate how these changes contribute to neurotoxicity, through integrated proteomics and metabolomics analyses. Our comprehensive analyses revealed that exposure to MGO, especially at cytotoxic level, induced extensive alterations in both the proteome and metabolome of SH-SY5Y cells. Analysis of proteomics data identified a substantial number of DEPs involved in critical cellular functions, including protein synthesis, cellular structural integrity, mitochondrial function and oxidative stress responses. Analysis of metabolomics and subsequent integration of metabolomics and proteomics data underscored significant changes in key metabolic pathways such as arginine biosynthesis, GSH metabolism, cysteine and methionine metabolism, and the TCA cycle. These changes may represent cellular adaptations or protective responses to dicarbonyl stress, or they could directly lead to cellular dysfunction and subsequent toxicity.
First, the enrichment and the PPI network analysis indicated significant alterations in protein synthesis-related pathways, such as ribosome biogenesis and translation initiation, upon exposure of cells to MGO. The expression levels of the majority of proteins involved in these pathways were significantly upregulated, indicative of heightened protein synthesis activity (Table 1). This upregulation is likely a cellular adaptation to meet the increased demand for new proteins and to manage extensive dicarbonyl stress-induced protein damage, crucial for replacing damaged or dysfunctional proteins. However, this enhanced protein synthesis could lead to an increased load of nascent proteins entering the ER, elevating the risk of protein misfolding and aggregation—primary indicators of ER stress51. The “protein processing in the endoplasmic reticulum” pathway, significantly enriched in our KEGG enrichment analysis, further corroborated the cellular response to this stress. It highlighted the upregulation of chaperones such as heat shock proteins HSPA8 and HSPH1, which assist in managing misfolded proteins52. The downregulation of protein disulfide isomerases such as PDIA3, PDIA4, and PDIA6 in the same pathway suggested a compromised ability to maintain protein structural integrity and an enhanced risk of further ER stress53. In response to ER stress, the unfolded protein response (UPR) may be activated54. Previous studies indicated that dicarbonyl stress from increased levels of MGO can trigger the UPR, characterized by an increased abundance of not only heat shock proteins but also components like ubiquitin ligases that encapsulate and remove proteins exhibiting misfolding due to dicarbonyl modifications54,55. While initially protective, prolonged activation of the UPR can lead to cellular dysfunction, as persistent ER stress challenges the cell’s capacity to restore proteostasis and can drive pathways leading to apoptosis and chronic inflammation55. Moreover, chronic ER stress and prolonged activation of UPR have been reported to be strongly associated with the pathology of neurodegenerative diseases, exacerbating neuronal loss and synaptic dysfunction which are critical factors in the progression of these disorders56.
Furthermore, our proteomic analysis revealed significant alterations in cellular adhesion mechanisms, particularly within the focal adhesion and ECM–receptor interaction pathways (Table 1). There was downregulation of integrins such as ITGB1, ITGAV, ITGA3, and ITGA6, along with crucial ECM components like COL1A1 and LAMB1. These changes suggest substantial alterations in the mechanisms that maintain cellular architecture and mediate signal transduction between the ECM and the cytoskeleton57, potentially underlying the cytotoxic effects induced by MGO. Integrins, serving as critical transmembrane receptors, are pivotal in mediating both physical and biochemical linkages between the extracellular matrix and the cytoskeleton58. Their downregulation can impair cell adhesion, impacting neuronal connectivity and synaptic stability, essential for neural development and function59. Similarly, alterations in COL1A1 and LAMB1 can disrupt the structural integrity and signaling capabilities of cells, potentially contributing to the pathophysiology of neurodegeneration60. Additionally, it is noteworthy that in our previous study36 we have observed that cytoskeletal proteins, crucial for the structural integrity and functionality of focal adhesion complexes61, seem to be particularly susceptible to modifications by MGO in SH-SY5Y cells. This observation leads us to hypothesize that MGO-induced modifications of these cytoskeletal proteins could play a role in causing instability and subsequent downregulation of integrins and other adhesion-related proteins.
Moreover, analysis of proteomic data revealed significant alterations in proteins involved in oxidative phosphorylation, indicating changes in mitochondrial function and energy metabolism (Table 1). Metabolomics and integrated omics analyses confirmed that these changes extend to the TCA cycle, suggesting a comprehensive disruption in energy metabolism (Fig. 5). Figure 6a specifically depicts the effects of MGO exposure on the energy metabolism in SH-SY5Y cells, beginning with a notable downregulation of glycolytic enzymes such as PFKL and ENO2. This reduction likely compelled the cells to depend more on the TCA cycle, as evidenced by increased intermediates like citric acid and isocitric acid, and enzymes such as IDH3B and IDH3G, responding to higher energy needs or managing oxidative stress. Support from gluconeogenesis and other pathways likely provided necessary substrates. Furthermore, the electron transport chain showed a compromised functionality with significant downregulation of components such as NDUFA7, COQ6, and COX5B, indicating a reduced efficiency. Conversely, the upregulation of components like ATP5F1D suggests compensatory efforts to maintain ATP production despite these inefficiencies. Previous in vitro studies have confirmed that mitochondrial impairments, including reduced mitochondrial membrane potential and the overproduction of ROS mediated by mitochondria, are key contributors to the cytotoxic effects induced by MGO in SH-SY5Y cells25,62. Our findings provide further evidence at the molecular level of mitochondrial dysfunction’s role in these effects within SH-SY5Y cells. It is worth noting that contrasting findings have been reported in different cell types and experimental conditions. For example, exposure to 200 µM MGO (cytotoxic level) in C2C12 myoblast cells showed a decrease in TCA cycle intermediates without significant glycolysis changes63, while non-toxic MGO concentrations in adipocytes enhanced aerobic glycolysis64. This indicates that metabolic responses may vary with cell type and MGO toxicity levels.
a Effects on energy metabolism, showing disruptions in glycolysis, TCA cycle, and oxidative phosphorylation. b Effects on Nrf2/GSH axis, depicting activation of the Nrf2 pathway and upregulation of GSH levels. c Effects on amino acid catabolism, detailing disruptions in arginine biosynthesis and related impacts on the urea cycle induced by MGO. Red arrows indicate upregulation, blue arrows indicate downregulation, based on integrated omics analyses. Figure created with BioRender.com.
Additionally, the proteomic analysis highlighted significant engagement of the Nrf2 pathway (Table 1) and alterations in GSH synthesis-related pathways, including cysteine and methionine metabolism and GSH metabolism, as prominently featured in our metabolomics and integrated omics analyses (Fig. 5). Figure 6b illustrates the impact of MGO exposure on these pathways. MGO has been reported to activate the Nrf2 pathway directly by modifying Keap1 and potentially indirectly through ROS production induced by MGO65,66. Consistent with these findings, we observed the upregulation of several Nrf2-regulated proteins, including HMOX1 and GCLM, with the latter being critical for GSH synthesis67. The metabolites detected in the cysteine and methionine metabolism and GSH metabolism pathways indicated an increased cellular effort to synthesize GSH. For instance, the observed downregulation of glutamate and cystathionine suggested their increased consumption in GSH synthesis. A 2.1-fold increase in GSH levels in the high-concentration MGO-treated cells was observed reflecting the cells’ attempt to enhance their detoxification capacity via the GSH-dependent glyoxalase system54. Together, the activation of Nrf2 and the increase in GSH levels can be seen as cellular protective mechanisms against dicarbonyl stress, working to mitigate the cytotoxic effects induced by MGO exposure.
Finally, the metabolomic analysis and integrated omics studies revealed significant alterations in several amino acid metabolism pathways, particularly in the arginine biosynthesis pathway, which plays a critical role in feeding into the urea cycle. Significant downregulation was observed in key metabolites including ornithine, citrulline, arginosuccinate, and arginine—which are all fundamental components of the urea cycle (Fig. 6c). Additionally, glutamate levels were reduced, likely reallocating resources to GSH synthesis to combat dicarbonyl stress. This reduction in glutamate, a key precursor in both the arginine biosynthesis and the urea cycle, likely contributed to decreased substrate availability for these cycles. The depletion of urea cycle metabolites may also stem from their consumption in the reaction of arginine with MGO to form MG-H1, which was substantially increased. The disruption of this pathway can compromise nitrogen waste processing and amino acid catabolism68, thereby potentially influencing broader cellular functions and contributing to the neurotoxicity observed in SH-SY5Y cells.
It is important to note that the interpretation of the proteomics and metabolomics data discussed above was primarily based on the DEPs and DMs identified in cells from the high-concentration MGO-treated group, where the most substantial changes were evaluated. Enrichment analyses were also conducted for DEPs identified at two lower concentrations, with results presented in Supplementary Figs. 3 and 4. These analyses show that fewer terms were enriched at lower concentrations compared to the highest concentration MGO-treated group. Terms related to cellular adhesion and signaling were enriched across all three treatment groups, whereas terms associated with protein synthesis and responses to oxidative stress were only enriched in the medium- and high-concentration MGO-treated groups. Furthermore, the concentration-dependent analysis, taking into account all treatment groups, shows that the majority of proteins (enriched from the top five densely connected modules (Table 1)) and DMs, identified from the high-concentration MGO-treated group, exhibited either a positive or negative correlation with MGO exposure concentrations (Tables 2 and 3). These results further confirm that the response to MGO is more pronounced at the highest MGO concentration used in this study, supporting the reliability of our enrichment analysis and the interpretation of data based on these analyses at the highest MGO concentration. Consequently, this validates that the patterns or changes observed under the highest MGO concentration conditions are robust indicators of MGO-induced effects.
Moreover, it should also be noted that the use of Student’s t-test alone in the current study, without corrections for multiple testing, may introduce false positives. Our current strategy is based on the exploratory aspect of omics studies in toxicology and pharmacology, aimed at capturing a broad spectrum of potential targets. Consequently, it would be of interest for future studies to further validate the DEPs and the pathways of interest identified in this study. Despite this limitation, several findings from the current study, such as increased ER stress and the activation of the Nrf2-mediated pathway upon MGO exposure, are supported by existing literature66,69. Different or additional methods, such as PCR or western blot, could be employed to quantify the differential expression of proteins of interest for further validation. Additionally, we have deposited the proteins and metabolites identified across all three concentration groups, along with their corresponding raw p values, in Supplementary Tables 7 and 8 and on the open-source platform Figshare (https://doi.org/10.6084/m9.figshare.27268275). This will allow future researchers to apply statistical corrections and explore the data from different perspectives.
In conclusion, our integrated proteomics and metabolomics analyses provide a comprehensive view of the cellular alterations induced by MGO in SH-SY5Y cells. It was observed that MGO exposure, particularly at cytotoxic levels, significantly impacts both the proteome and metabolome, inducing a range of both toxic effects and adaptive responses. Contributing to toxicity are increased ER stress, disruptions in cellular adhesion and ECM integrity, mitochondrial dysfunction, and disrupted amino acid metabolism, notably in pathways such as arginine biosynthesis which directly impacts the urea cycle. On the adaptive side, cells upregulate protein synthesis, activate the Nrf2 pathway, and reprogram metabolism to counteract dicarbonyl stress and maintain energy levels, consistent with increasing evidence that cells are not merely passive targets of electrophilic damage but can actively adapt and protect themselves70,71,72. Furthermore, the majority of key proteins and metabolites associated with these changes, initially identified under cytotoxic conditions, were found to show a concentration-dependent decrease or increase in their expression levels with increasing MGO concentrations, suggesting their potential as biomarkers for MGO exposure. These findings enhance our understanding of MGO-induced neurotoxicity and identify potential therapeutic targets for mitigating its harmful effects.
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary Information file and on the open-source platform Figshare, which can be accessed at https://doi.org/10.6084/m9.figshare.27268275.
References
Rehman, M. U. et al. Mitochondrial dysfunctions, oxidative stress and neuroinflammation as therapeutic targets for neurodegenerative diseases: an update on current advances and impediments. Neurosci. Biobehav. Rev. 144, 104961 (2023).
Jurcau, A. Insights into the pathogenesis of neurodegenerative diseases: focus on mitochondrial dysfunction and oxidative stress. Int. J. Mol. Sci. 22, 11847 (2021).
Gandhi, J. et al. Protein misfolding and aggregation in neurodegenerative diseases: a review of pathogeneses, novel detection strategies, and potential therapeutics. Rev. Neurosci. 30, 339–358 (2019).
Niranjan, R. Recent advances in the mechanisms of neuroinflammation and their roles in neurodegeneration. Neurochem. Int. 120, 13–20 (2018).
Li, J., Liu, D., Sun, L., Lu, Y. & Zhang, Z. Advanced glycation end products and neurodegenerative diseases: mechanisms and perspective. J. Neurol. Sci. 317, 1–5 (2012).
Ramasamy, R. et al. Advanced glycation end products and RAGE: a common thread in aging, diabetes, neurodegeneration, and inflammation. Glycobiology 15, 16R–28R (2005).
Chellan, P. & Nagaraj, R. H. Protein crosslinking by the Maillard reaction: dicarbonyl-derived imidazolium crosslinks in aging and diabetes. Arch. Biochem. Biophys. 368, 98–104 (1999).
Vicente Miranda, H. & Outeiro, T. F. The sour side of neurodegenerative disorders: the effects of protein glycation. J. Pathol. 221, 13–25 (2010).
Sirangelo, I. & Iannuzzi, C. Understanding the role of protein glycation in the amyloid aggregation process. Int. J. Mol. Sci. 22, 6609 (2021).
Srikanth, V. et al. Advanced glycation endproducts and their receptor RAGE in Alzheimer’s disease. Neurobiol. Aging 32, 763–777 (2011).
Bellier, J. et al. Methylglyoxal, a potent inducer of AGEs, connects between diabetes and cancer. Diabetes Res. Clin. Pract. 148, 200–211 (2019).
Hellwig, M., Gensberger-Reigl, S., Henle, T. & Pischetsrieder, M. Food-derived 1,2-dicarbonyl compounds and their role in diseases. Semin. Cancer Biol. 49, 1–8 (2018).
Thornalley, P. J. Dicarbonyl intermediates in the Maillard reaction. Ann. N. Y. Acad. Sci. 1043, 111–117 (2005).
Rabbani, N. & Thornalley, P. J. Methylglyoxal, glyoxalase 1 and the dicarbonyl proteome. Amino Acids 42, 1133–1142 (2012).
Rabbani, N. & Thornalley, P. J. Dicarbonyl stress in cell and tissue dysfunction contributing to ageing and disease. Biochem. Biophys. Res. Commun. 458, 221–226 (2015).
Zhang, X., Scheijen, J. L., Stehouwer, C. D., Wouters, K. & Schalkwijk, C. G. Increased methylglyoxal formation in plasma and tissues during a glucose tolerance test is derived from exogenous glucose. Clin. Sci. 137, 697 (2023).
Beeri, M. S. et al. Serum concentration of an inflammatory glycotoxin, methylglyoxal, is associated with increased cognitive decline in elderly individuals. Mech. Ageing Dev. 132, 583–587 (2011).
Ahmed, N. et al. Protein glycation, oxidation and nitration adduct residues and free adducts of cerebrospinal fluid in Alzheimer’s disease and link to cognitive impairment. J. Neurochem. 92, 255–263 (2005).
Kuhla, B. et al. Methylglyoxal, glyoxal, and their detoxification in Alzheimer’s disease. Ann. N. Y. Acad. Sci. 1043, 211–216 (2005).
Fan, X. et al. Vitamin C is a source of oxoaldehyde and glycative stress in age‐related cataract and neurodegenerative diseases. Aging Cell 19, e13176 (2020).
Pucci, M. et al. Methylglyoxal affects cognitive behaviour and modulates RAGE and Presenilin-1 expression in hippocampus of aged mice. Food Chem. Toxicol. 158, 112608 (2021).
Wei, C.-C., Li, S.-W., Wu, C.-T., How, C. M. & Pan, M.-H. Dietary methylglyoxal exposure induces Alzheimer’s disease by promoting amyloid β accumulation and disrupting autophagy in Caenorhabditis elegans. J. Agric Food Chem. 70, 10011–10021 (2022).
Nan, F. et al. Ginsenoside Rb1 mitigates oxidative stress and apoptosis induced by methylglyoxal in SH-SY5Y cells via the PI3K/Akt pathway. Mol. Cell. Probes 48, 101469 (2019).
Kikuchi, S. et al. Neurotoxicity of methylglyoxal and 3‐deoxyglucosone on cultured cortical neurons: synergism between glycation and oxidative stress, possibly involved in neurodegenerative diseases. J. Neurosci. Res. 57, 280–289 (1999).
Wang, Y.-H., Yu, H.-T., Pu, X.-P. & Du, G.-H. Myricitrin alleviates methylglyoxal-induced mitochondrial dysfunction and AGEs/RAGE/NF-κB pathway activation in SH-SY5Y cells. J. Mol. Neurosci. 53, 562–570 (2014).
Suh, K. S., Chon, S., Jung, W. W. & Choi, E. M. Protective effects of sciadopitysin against methylglyoxal‐induced degeneration in neuronal SK‐N‐MC cells. J. Appl. Toxicol. 42, 274–284 (2022).
Coccini, T. et al. Methylglyoxal-induced neurotoxic effects in primary neuronal-like cells transdifferentiated from human mesenchymal stem cells: impact of low concentrations. J. Appl. Toxicol. https://doi.org/10.1002/jat.4515 (2023).
Teresa, C. et al. 3D human stem-cell-derived neuronal spheroids for in vitro neurotoxicity testing of methylglyoxal, highly reactive glycolysis byproduct and potent glycating agent. Curr. Res. Toxicol. 7, 100176 (2024).
Lo, T. W., Westwood, M. E., McLellan, A. C., Selwood, T. & Thornalley, P. J. Binding and modification of proteins by methylglyoxal under physiological conditions. A kinetic and mechanistic study with N alpha-acetylarginine, N alpha-acetylcysteine, and N alpha-acetyllysine, and bovine serum albumin. J. Biol. Chem. 269, 32299–32305 (1994).
Suantawee, T., Cheng, H. & Adisakwattana, S. Protective effect of cyanidin against glucose- and methylglyoxal-induced protein glycation and oxidative DNA damage. Int. J. Biol. Macromol. 93, 814–821 (2016).
Vailati-Riboni, M., Palombo, V. & Loor, J. J. What are omics sciences? in Periparturient Diseases of Dairy Cows (ed. Ametaj, B.) Ch. 1 (Springer, 2017) https://doi.org/10.1007/978-3-319-43033-1_1.
Li, G. et al. Proteomics analysis of methylglyoxal-induced neurotoxic effects in SH-SY5Y cells. Cell Biochem. Funct. 29, 30–35 (2011).
Rabilloud, T., Chevallet, M., Luche, S. & Lelong, C. Two-dimensional gel electrophoresis in proteomics: past, present and future. J. Proteom. 73, 2064–2077 (2010).
Ioghen, O. C., Ceafalan, L. C. & Popescu, B. O. SH-SY5Y cell line in vitro models for Parkinson disease research—old practice for new trends. J. Integr. Neurosci. 22, 20 (2023).
Neilson, K. A. et al. Less label, more free: approaches in label‐free quantitative mass spectrometry. Proteomics 11, 535–553 (2011).
Zheng, L. et al. Proteomics-based identification of biomarkers reflecting endogenous and exogenous exposure to the advanced glycation end product precursor methylglyoxal in SH-SY5Y human neuroblastoma cells. Int. J. Biol. Macromol. 132859 https://doi.org/10.1016/j.ijbiomac.2024.132859 (2024).
Batth, T. S. et al. Protein aggregation capture on microparticles enables multipurpose proteomics sample preparation. Mol. Cell. Proteom. 18, 1027a (2019).
The UniProt Consortium. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Arakawa, S. et al. Mass spectrometric quantitation of AGEs and enzymatic crosslinks in human cancellous bone. Sci. Rep. 10, 18774 (2020).
Pang, Z. et al. MetaboAnalyst 6.0: towards a unified platform for metabolomics data processing, analysis and interpretation. Nucleic Acids Res. https://doi.org/10.1093/nar/gkae253 (2024).
Team, R. C. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2013).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote) omics data. Nat. Methods 13, 731–740 (2016).
Jonckheere, A. R. A distribution-free k-sample test against ordered alternatives. Biometrika 41, 133–145 (1954).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Szklarczyk, D. et al. The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 4, 1–27 (2003).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Chin, C.-H. et al. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8, 1–7 (2014).
Tang, D. et al. SRplot: A free online platform for data visualization and graphing. PLoS ONE 18, e0294236 (2023).
Vulesevic, B. et al. Methylglyoxal-induced endothelial cell loss and inflammation contribute to the development of diabetic cardiomyopathy. Diabetes 65, 1699–1713 (2016).
Ghemrawi, R. & Khair, M. Endoplasmic reticulum stress and unfolded protein response in neurodegenerative diseases. Int. J. Mol. Sci. 21, 6127 (2020).
Duncan, E. J., Cheetham, M. E., Chapple, J. P. & van der Spuy, J. The role of HSP70 and its co-chaperones in protein misfolding, aggregation and disease. Subcell. Biochem. 243–273 https://doi.org/10.1007/978-3-319-11731-7_12 (2015).
Victor, P., Sarada, D. & Ramkumar, K. M. Crosstalk between endoplasmic reticulum stress and oxidative stress: focus on protein disulfide isomerase and endoplasmic reticulum oxidase 1. Eur. J. Pharmacol. 892, 173749 (2021).
Rabbani, N., Xue, M. & Thornalley, P. J. Dicarbonyl stress, protein glycation and the unfolded protein response. Glycoconj. J. 38, 331–340 (2021).
Irshad, Z. et al. Activation of the unfolded protein response in high glucose treated endothelial cells is mediated by methylglyoxal. Sci. Rep. 9, 7889 (2019).
Hetz, C. & Saxena, S. ER stress and the unfolded protein response in neurodegeneration. Nat. Rev. Neurol. 13, 477–491 (2017).
Bachir, A. I., Horwitz, A. R., Nelson, W. J. & Bianchini, J. M. Actin-based adhesion modules mediate cell interactions with the extracellular matrix and neighboring cells. Cold Spring Harb. Perspect. Biol. 9, a023234 (2017).
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Lilja, J. & Ivaska, J. Integrin activity in neuronal connectivity. J. Cell Sci. 131, jcs212803 (2018).
Soles, A. et al. Extracellular matrix regulation in physiology and in brain disease. Int. J. Mol. Sci. 24, https://doi.org/10.3390/ijms24087049 (2023).
Cronin, N. M. & DeMali, K. A. Dynamics of the actin cytoskeleton at adhesion complexes. Biology 11, 52 (2021).
Furstenau, C. R., de Souza, I. C. C. & de Oliveira, M. R. Tanshinone I induces mitochondrial protection by a mechanism involving the Nrf2/GSH axis in the human neuroblastoma SH-SY5Y Cells exposed to methylglyoxal. Neurotox. Res. 36, 491–502 (2019).
Todoriki, S. et al. Methylglyoxal induces inflammation, metabolic modulation and oxidative stress in myoblast cells. Toxins 14, https://doi.org/10.3390/toxins14040263 (2022).
Komanetsky, S. M. et al. Proteomic identification of aerobic glycolysis as a potential metabolic target for methylglyoxal in adipocytes. Nutr. Res. 80, 66–77 (2020).
Zheng, L., van Dongen, K. C. W., Bakker, W., Miro Estruch, I. & Rietjens, I. The influence of intracellular glutathione levels on the induction of Nrf2-mediated gene expression by alpha-dicarbonyl precursors of advanced glycation end products. Nutrients 14, https://doi.org/10.3390/nu14071364 (2022).
Bollong, M. J. et al. A metabolite-derived protein modification integrates glycolysis with KEAP1-NRF2 signalling. Nature 562, 600–604 (2018).
Lu, S. C. Glutathione synthesis. Biochim. Biophys. Acta 1830, 3143–3153 (2013).
Dimski, D. S. Ammonia metabolism and the urea cycle: function and clinical implications. J. Vet. Intern. Med. 8, 73–78 (1994).
Chan, C. M. et al. Methylglyoxal induces cell death through endoplasmic reticulum stress‐associated ROS production and mitochondrial dysfunction. J. Cell. Mol. Med. 20, 1749–1760 (2016).
Jacobs, A. T. & Marnett, L. J. Systems analysis of protein modification and cellular responses induced by electrophile stress. Acc. Chem. Res. 43, 673–683 (2010).
Ken, I., Kit, I. T. & Masayuki, Y. Molecular mechanism activating nrf2–keap1 pathway in regulation of adaptive response to electrophiles. Free Radic. Biol. Med. 36, 1208–1213 (2004).
Kumagai, Y., Akiyama, M. & Unoki, T. Adaptive responses to electrophilic stress and reactive sulfur species as their regulator molecules. Toxicol. Res. 35, 303–310 (2019).
Acknowledgements
H.W. is thankful for the financial support provided by the China Scholarship Council, under Grant Number: CSC202307720061.
Author information
Authors and Affiliations
Contributions
Conceptualization: I.M.C.M.R., E.S., and L.Z.; methodology: L.Z., H.W., W.B., S.B., E.S., and I.M.C.M.R.; investigation and validation: L.Z. and H.W.; formal analysis: L.Z. and H.W.; writing—original draft preparation: L.Z. and H.W.; writing—review and editing: L.Z., E.S., S.B., W.B., and I.M.C.M.R.; supervision: I.M.C.M.R., E.S., and L.Z.; resources and project administration: I.M.C.M.R. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, H., Boeren, S., Bakker, W. et al. An integrated proteomics and metabolomics analysis of methylglyoxal-induced neurotoxicity in a human neuroblastoma cell line. npj Sci Food 8, 84 (2024). https://doi.org/10.1038/s41538-024-00328-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-024-00328-0