Abstract
Rhabdomyosarcoma (RMS) is an aggressive cancer thought to arise from impaired myogenesis. This can be substantially overcome by increasing the levels of pannexin 1 (PANX1), a critical component of the myogenic program, but the mechanism involved is unknown. Using RNA-seq, we have previously found that overexpression of PANX1 dramatically reshapes the transcriptomic landscape of RMS including downregulation of a myogenic modulator, APOBEC2 (apolipoprotein B mRNA editing enzyme catalytic subunit 2). Following this clue, we investigated the role of APOBEC2 in the PANX1-mediated suppression of RMS malignancy. Here we show that, using a panel of patient-derived RMS cell lines and tumor specimens, APOBEC2 is expressed in RMS, but that its levels are lower than those in both differentiating myoblasts and skeletal muscle. In most RMS cell lines examined, APOBEC2 accumulates during proliferation and sustains their stem-like characteristics, as evidenced by its ability to promote the growth of spheroids upon increased expression. Yet, ectopic PANX1 expression led to a marked downregulation of APOBEC2 across a large proportion of RMS cell lines assessed. Strikingly, these were the same cells in which PANX1 triggers multinucleation. We further reveal that, like healthy myoblasts progressing through myogenesis, the multinucleation observed here in RMS cells results from cell fusion. Importantly, in RMS cells engineered to overexpress APOBEC2, PANX1 no longer enhances cell fusion, but its other anti-tumorigenic properties are still preserved. Collectively, our data indicate that PANX1 promotes RMS cell fusion by downregulating APOBEC2 expression, driving these tumor cells further into the myogenic program.
Similar content being viewed by others
Introduction
Rhabdomyosarcoma (RMS) is the most common soft tissue sarcoma in children [1, 2]. Pediatric RMS is comprised of two main histologic subtypes: embryonal (eRMS) and alveolar (aRMS) [2,3,4]. eRMS usually affects infants or children under ten years of age, whereas aRMS mostly occurs in adolescents and young adults and is associated with a worse prognosis [5]. Unfortunately, despite intense treatment regimens, patients with metastatic or relapsed RMS have a 30% and 17% 5-year survival rate, respectively [6,7,8]. RMS cells are positive for myogenic markers, but their ability to exit the cell cycle and complete differentiation and fusion is altered [9]. Myogenesis is a complex and tightly controlled multistep process in which precursor cells commit to myoblast lineage, differentiate, fuse into multinucleated myotubes, and mature to form myofibers [10]. Understanding the mechanism involved in the dysregulation of myogenic pathways in RMS may provide a new possibility to alleviate malignancy [11]. As terminal myogenesis switches off cell proliferation and migration, promoting RMS differentiation should antagonize tumor growth and metastasis [12].
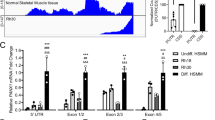
Pannexin 1 (PANX1 in humans; Panx1 in rodents) forms single membrane channels important in cellular communication and signaling [13]. Pannexin 1 is ubiquitously expressed in many organs and tissues including the skeletal muscle [14,15,16,17,18,19,20,21,22]. We have previously identified PANX1/Panx1 as a regulator of myogenesis [22]. Pannexin 1 expression is low in undifferentiated myoblasts and increases during myogenesis [22,23,24]. In vitro, increasing PANX1 levels promoted myoblast differentiation and fusion, while its inhibition reduced these processes [22]. Panx1 loss leads to reduced myoblast fusion ex vivo and smaller myofibers in male mice [25]. As opposed to differentiated skeletal muscle tissue and myoblasts, PANX1 expression is low in RMS tissue specimens and cell lines [26]. The canonical 5’ untranslated region (5’UTR) or 5’ leader of the transcript is completely lost in RMS cells and cannot be translated [27]. Notably, PANX1 overexpression inhibited RMS malignant properties in vitro and in vivo, independent of its canonical channel activity [26]. As this was the first time that a channel-independent function had been attributed to pannexins, the molecular changes brought on by increasing PANX1 expression in RMS were surveyed and characterized using a combination of unbiased genome-wide approaches. Using RNA-seq, we found that ectopic expression of PANX1 regulates the transcription level of various genes including APOBEC2 (apolipoprotein B mRNA editing enzyme catalytic subunit 2) [28].
APOBEC2 is highly expressed in skeletal muscle and affects muscle development in mice [29,30,31]. Notably, APOBEC2 knockout (KO) mice display early myoblast differentiation and fusion [32]. APOBEC2 may not have some of the functions shared by the other APOBEC family members, such as RNA editing, DNA methylation, and DNA mutation [33]. Instead, APOBEC2 may play a role in transcriptional regulation during myogenesis [33]. APOBEC2 occupies promoter regions during myoblast differentiation, modulating specific transcriptional programs [33]. Recent reports also suggest an important role of APOBEC2 in various cancers through the regulation of the tumor related genes PTEN, p53, and Eif4g2 [34,35,36]. In bladder cancer, APOBEC2 transcriptional level was significantly correlated with the tumor mutational burden [37]. In stomach adenocarcinoma, APOBEC2 was found to be downregulated compared to normal tissues [38].
Here, we showed that APOBEC2 is expressed in RMS cells and tumors but at a lower level than that of differentiating myoblasts and skeletal muscle tissue. APOBEC2 levels were found to increase as RMS cells proliferate, possibly to help sustain their stemness as APOBEC2 overexpression accelerates 3D spheroid growth. PANX1 overexpression led to the downregulation of APOBEC2 in half of the RMS cell lines assessed. Notably, these were the same cells in which we showed the ability of PANX1 to trigger RMS cell multinucleation. We demonstrated using a cell fusion assay that, like healthy myoblasts, the multinucleation observed in RMS results from cell fusion. As PANX1’s role in inhibiting RMS malignant properties is multifaceted, we have used various functional assays to identify the role of APOBEC2 downregulation in this context. In cells engineered to overexpress APOBEC2, PANX1 retained most of its anti-tumorigenic properties but could no longer enhance cell fusion. Collectively, our data indicate that PANX1 promotes RMS cell fusion by downregulating APOBEC2 expression.
Results
APOBEC2 is expressed in RMS cells and tumor specimens, but at lower levels than in differentiating myoblasts and skeletal muscle
To assess the role of APOBEC2 in RMS, its expression was first examined in cell lines. As APOBEC2 levels increases during the differentiation of mouse myoblasts [29, 33], its levels in RMS were compared to that of undifferentiated and differentiating human skeletal muscle myoblasts (HSMM). We found that most cell lines express APOBEC2 transcript levels comparable to undifferentiated HSMM, except Rh41 and Rh36 which displayed higher expression (Fig. 1A). Nevertheless, the transcript level of APOBEC2 was significantly lower in undifferentiated HSMM and all RMS cell lines in comparison with differentiating HSMM (Fig. 1A). Similar results were found with APOBEC2 protein expression (Fig. 1B, C). This was further shown by immunolabeling analysis (Fig. 1D) in which APOBEC2 localization in myoblasts and RMS cells was mainly cytoplasmic. However, in the Rh36 and RD cell lines, based on the appearance of its staining pattern, endogenous APOBEC2 may be localized in other compartments such as the Golgi apparatus and the endoplasmic reticulum (Fig. 1D).
Endogenous APOBEC2 expression in six RMS cell lines, as well as in undifferentiated (Undiff. HSMM) and differentiating HSMM (Diff. HSMM), was quantified by (A) RT-qPCR (n = 3; ###p < 0.001 compared to Undiff. HSMM; ####p < 0.0001 compared to Undiff. HSMM; ****p < 0.0001 compared to Diff. HSMM (one-way ANOVA followed by Tukey’s post hoc tests)) and by (B) Western blots and their (C) quantification (n = 3 independent biological replicates; ***p < 0.001 compared to Diff. HSMM (one-way ANOVA followed by Tukey’s post hoc tests)). GAPDH was used as a loading control. D Representative pictures of APOBEC2 (green) immunolabeling in the RMS cells, undifferentiated and differentiating HSMM (blue = nuclei; bar = 30 μm). E Representative images of human RMS tumors and normal skeletal muscles immunolabeled for APOBEC2 (red), which was quantified in (F). The negative control without primary antibodies shows the labeling specificity. *p < 0.05 compared to skeletal muscle (unpaired two-tailed Student’s t test). Blue = nuclei, bar = 100 µm.
APOBEC2 was then immunolabeled in 9 pediatric RMS tumors and 7 skeletal muscle biopsies. We found that similar to murine muscles [29], APOBEC2 was present in human myofibers (Fig. 1E). APOBEC2 was also detected in RMS, but generally at a lower level than in skeletal muscle (Fig. 1F). Altogether these results show that APOBEC2 is expressed in RMS cell lines and tumor specimens, but at a lower level than in differentiating myoblasts and skeletal muscle, respectively.
PANX1 overexpression leads to APOBEC2 downregulation in Rh28, Rh30 and RD cell lines
Using RNA-seq analysis, we have found that the overexpression of PANX1 in the Rh30 cell line led to a decrease in APOBEC2 transcript levels [28]. Given this, we evaluated the regulation of APOBEC2 expression by PANX1 in RMS. APOBEC2 levels were analyzed over eight days in our stable RMS cell lines in which PANX1 was overexpressed under the control of a cumate switch system [26]. APOBEC2 levels increased in most control cells as they proliferate, which was significant after eight days in Rh28, Rh30, Rh41, and RD cells (Fig. 2A–L). Notably, PANX1 overexpression significantly decreased the expression of APOBEC2 in Rh28, Rh30, and RD cells compared to their respective controls but not in Rh36, Rh41 and Rh18 (Fig. 2A–L). This reduction was detected at around day four and was evident after eight days in culture.
APOBEC2 protein levels were examined following induction of PANX1 expression (PANX1) over a period of eight days in culture in Rh28 (A, B), Rh30 (C, D), Rh41 (E, F), Rh18 (G, H), Rh36 (I, J), and RD (K, L) cell lines (n = 3; **p < 0.01 and ***p < 0.001 compared to their respective control cells treated with cumate (GFP); ###p < 0.001 and ####p < 0.0001 compared to GFP at Day 0; ns non-significant (two-way ANOVA followed by Tukey’s post hoc tests)). GAPDH was used as a loading control. Relative APOBEC2 expression was examined following induction of PANX1 expression (PANX1) after eight days in M Rh28, N Rh30, O Rh41, P Rh18, Q Rh36, and R RD cell lines (n = 3; *p < 0.05 and **p < 0.01 compared to their respective control cells (GFP) treated with cumate; ns: non-significant (two-tailed Student’s t test)). S APOBEC2 levels were examined in Rh30 xenografts overexpressing PANX1 compared to their control counterparts (GFP), T and quantified (n = 3; mice randomly assigned; not blinded method; *p < 0.05; unpaired two-tailed Student’s t test).
The previous RNA-seq results were confirmed in Rh30 cells [28] by RT-qPCR while also examining APOBEC2 mRNA expression in the other five cell lines after eight days upon PANX1 upregulation (Fig. 2M–R). The decrease in APOBEC2 protein levels in Rh28, Rh30 and RD cells was similarly associated with a significant decrease in its mRNA expression (Fig. 2M, N, R). Accordingly, APOBEC2 transcript expression was not affected by the upregulation of PANX1 expression in Rh41, Rh18, and Rh36 cells (Fig. 2O, P, Q).
Western blot analysis showed a downregulation of APOBEC2 levels when PANX1 expression was increased in Rh30 xenografts (Fig. 2S, T), suggesting that this regulation also occurs in vivo. These results indicate that, in some RMS cell lines, increasing PANX1 levels leads to a downregulation of APOBEC2 expression at both the transcript and protein level.
The regulation of APOBEC2 levels by PANX1 does not involve a downregulation of Pax7
Intriguingly, the effect of PANX1 upregulation on APOBEC2 levels was found in three of the six cell lines tested. When examining the characteristics that these cells may share, no correlation was found with the patients’ sex, tumor subtype, known mutation or gene fusion. It has previously been shown that the downregulation of Pax7 (Paired Box 7) results in the subsequent reduction of APOBEC2 expression in differentiating pluripotent stem cells, affecting the balance between proliferation and differentiation [39]. Since RMS arises from a defect in myogenesis and Pax7, APOBEC2, and PANX1 play a role in this process [22, 39, 40], the expression of Pax7 was examined in the six RMS cell lines by immunofluorescence (Fig. 3A). We found that Rh41, Rh36, and RD had a higher proportion of cells expressing Pax7 than the other three cell lines (Fig. 3B).
Pax7 (green) was labeled in the RMS cell lines. Undifferentiated (Undiff.) mouse primary myoblasts were used as a positive control. Representative images are shown in A (blue = nuclei; bar = 30 μm). B The percentage of Pax7-positive cells were quantified (n = 3; ***p < 0.001 and ****p < 0.0001 compared to the other RMS cell lines (one-way ANOVA followed by Tukey’s post hoc tests)). The same was done following induction of PANX1 in culture using cumate. The percentage of Pax7-positive cells has been determined following two and eight days in culture in the C Rh28, D Rh30, E Rh41, F Rh18, G Rh36, and H RD cell lines (n = 3; ns non-significant (two-way ANOVA followed by Sidak’s post hoc tests) compared to their respective controls cells (GFP) treated with cumate)).
Using the same approach, the effect of PANX1 upregulation on Pax7 expression was examined. Interestingly, after either two or eight days, the proportion of Pax7-positive cells was not affected by PANX1 upregulation in any of the six cell lines (Fig. 3C–H). These data suggest that while the endogenous levels of APOBEC2 and Pax7 may be related to one another as they display a similar trend, modulation of Pax7 expression is likely not the mechanism by which PANX1 regulates APOBEC2 levels.
PANX1 overexpression induces multinucleation coinciding with cell lines where APOBEC2 is downregulated
As stated earlier, PANX1 overexpression increases multinucleation in Rh30 cells [26] and promotes the differentiation and fusion of human myoblasts [22, 40]. In vivo, we also found that Panx1 loss reduces myoblast fusion in mice [25]. On the other hand, APOBEC2 deficiency accelerates myogenic differentiation and fusion [32]. To assess whether there is a potential functional link between PANX1, APOBEC2, and multinucleation, the effect of PANX1 overexpression on multinucleation (Fig. 4A) was examined in the other RMS cell lines. Like in Rh30 cells, the number of multinucleated cells and the fusion index were significantly increased in Rh28 and RD cells when PANX1 was overexpressed (Fig. 4B–I). Comparable to our findings on multinucleation and fusion indexes in Rh41 and Rh36 cells (Fig. 4), our previous work showed that PANX1 overexpression did not increase Rh18 multinucleation [26]. Notably, these findings indicate a correlation in the ability of PANX1 to increase multinucleation and the downregulation of APOBEC2.
PANX1 was overexpressed and the number of nuclei was counted 48 h post-transfection compared to their control counterparts expressing the empty vector. Both vectors allow for endogenous GFP expression. PANX1 was labeled in red (blue = nuclei; bar = 30 μm). Representative images from the Rh28 cell line are shown in A with the cells expressing the empty vector shown on top panels and the ones expressing ectopic PANX1 shown in lower panels. Arrows indicate cells with more than one nucleus. The percentage of cells with two or more nuclei, as well as fusion indices, was calculated in B, C Rh28, D, E Rh41, F, G Rh36, and H, I RD transfected cells (n = 3; *p < 0.05 and **p < 0.01 compared to their respective control (GFP); non-significant (two-tailed unpaired Student’s t test)).
APOBEC2 upregulation does not alleviate the PANX1-mediated inhibition of RMS cell proliferation and viability
To delineate the role of APOBEC2 down-regulation in the PANX1-mediated inhibition of RMS progression, the Rh30 stable cell line that expresses PANX1 under the control of cumate was engineered to constitutively overexpress APOBEC2 (Fig. 5A). The APOBEC2 overexpression achieved was physiologically relevant with levels similar to differentiating HSMM.
A Representative Western blots showing the increase of APOBEC2 levels in the engineered stable Rh30 cell line as well as the induction of PANX1 with cumate treatment. Differentiating HSMM were used as a comparison with endogenous APOBEC2 levels. GAPDH was used as a loading control. Cell proliferation was assessed after two (B) and four (C) days in culture in which PANX1 levels were induced by cumate treatment (n = 3; *p < 0.05 and **p < 0.01 compared to their respective control (- Cumate) (two-way ANOVA followed by Tukey’s post hoc tests)). D Cell viability was quantified as percentage of viable cells after two and four days in culture where PANX1 expression was induced by cumate (+ Cumate) (n = 3; *p < 0.05 and **p < 0.01 compared to their respective control (- Cumate) (two-way ANOVA followed by Tukey’s post hoc tests)). Cells were grown in suspension to examine 3D spheroid formation and growth over 240 h. Representative images at 0 and 240 h are shown in (E). F Spheroid growth was quantified using the Incucyte live imaging system as mean image integrated intensity (n = 3; ****p < 0.0001 (two-way ANOVA followed by Tukey’s post hoc tests)). While not indicated here to prevent overcrowding the graphs presented in this figure, there was no significant effect of increasing APOBEC2 levels in PANX1 overexpressing cells (+ Cumate).
We have shown previously that the overexpression of PANX1 significantly reduced the proliferation and viability of Rh30 cells in vitro [26]. Here, using this newly generated cell line, we assessed whether the overexpression of APOBEC2 could reverse the PANX1-mediated inhibition of cell proliferation. In accordance with our previous work [26], PANX1 overexpression reduced the proliferation of Rh30 cells upon two (Fig. 5B) and four (Fig. 5C) days of induction. However, upregulation of APOBEC2 in these cells did not influence the PANX1-mediated reduction in cell proliferation (Fig. 5B, C). Similar to our previous work [26], PANX1 overexpression resulted in a significant reduction in cell viability after two and four days in culture (Fig. 5D), but this was also not affected by APOBEC2 upregulation (Fig. 5D).
As PANX1 overexpression prevents RMS spheroids growth, this assay was performed here [26]. As expected, PANX1 overexpression resulted in a significant reduction in Rh30 spheroid growth (Fig. 5E, F). However, APOBEC2 overexpression did not reverse the potent effect of PANX1 (Fig. 5E, F). Notably, APOBEC2 overexpression in absence of PANX1 upregulation significantly promoted Rh30 spheroid growth (Fig. 5E, F) compared to its empty vector counterpart. While PANX1 overexpression inhibits RMS cell viability and proliferation by a mechanism independent of APOBEC2 downregulation, our findings suggest that an increase in APOBEC2 expression in RMS may promote its malignancy.
APOBEC2 overexpression prevents the RMS cell fusion triggered by PANX1
As our findings suggest that the downregulation of APOBEC2 by PANX1 may be correlated with PANX1’s ability to trigger cell multinucleation, APOBEC2 was upregulated to examine its effect on this PANX1-mediated process. To this end, we used our cumate-inducible PANX1 overexpressing Rh30 stable cell line that constitutively overexpresses APOBEC2, together with their control cells, and counted the percentage of cells with two or more nuclei (Fig. 6A). As expected, the increase in PANX1 levels led to a significant elevation in multinucleation (Fig. 6B) and fusion (Fig. 6C). Notably, these effects were abolished following the overexpression of APOBEC2 (Fig. 6B, C).
A Representative images showing the nuclei (blue) in cells overexpressing APOBEC2 (red) in the absence (- Cumate) or presence of ectopic PANX1 (+ Cumate) in the Rh30 cell line. These cells constitutively express GFP. Arrows indicate cells with more than one nucleus (bar = 30 μm). B The percentage of cells with 2 or more nuclei was counted (n = 3; ****p < 0.0001 compared to its respective control; ns non-significant (one-way ANOVA followed by Tukey’s post hoc tests)). C The fusion index was calculated for each condition (n = 3; ***p < 0.001 compared to its respective control; ns non-significant (one-way ANOVA followed by Tukey’s post hoc tests)). The Rh30 cell line was engineered to stably express mKate2 (Nuclight red (D)) or GFP (Nuclight green (E)) in their nuclei to then be used in the cell fusion assay (blue = nuclei; bar = 200 μm). An equal number of these cell lines were seeded together and transfected with PANX1. F Representative images of a PANX1 (blue) expressing cell containing red (arrow heads) and green (arrow) nuclei are shown (bar = 100 μm).
While it is well established that multinucleation results from cell fusion during myogenesis in the context of health, we wanted to confirm that this is also the case here in RMS cells. To this end, we generated two stable Rh30 cell lines, one with red nuclei (Fig. 6D) and the other one with green nuclei (Fig. 6E). Equal numbers of Rh30 cells with red and green nuclei were seeded and then transfected to overexpress PANX1. We found several multinucleated cells overexpressing PANX1 (blue) containing both red and green-labeled nuclei (Fig. 6F), suggesting that the multinucleation observed with increased PANX1 expression in RMS cells was due to cell fusion. Collectively, our results suggest that the downregulation of APOBEC2 plays an important role in the mechanism by which PANX1 promotes RMS cell fusion.
Discussion
This is the first study to our knowledge that has examined the role of APOBEC2 in RMS, whereby we have found that APOBEC2 is expressed in RMS cells and tumor specimens, but its levels are lower than that of differentiating myoblasts and myofibers. This lower expression of APOBEC2 may reflect the undifferentiated state of RMS similar to what has been previously observed in undifferentiated mouse myoblasts [29, 32, 33, 41, 42] and in undifferentiated HSMM in this current study. In multinucleated differentiated mouse C2C12 cells, APOBEC2 was detected in both the cytoplasm and nuclei [33]. However, in differentiating human myoblasts, APOBEC2 was mainly cytoplasmic. Rh30 and Rh41 showed a similar subcellular localization, while other RMS cells displayed different localization patterns reminiscent of compartments such as the Golgi apparatus. To our knowledge, such subcellular localization has not been previously reported for APOBEC2 but is worth noting as it may suggest an alteration of its function in cancer cells. This alternate localization may also reflect distinct functions stemming from the access of APOBEC2 to different pools of interactors and downstream players [43].
APOBEC2 levels increase during myogenesis in vitro [29, 33] and seem to closely reflect the fusion status [29]. In RMS, APOBEC2 expression is also dynamic as it increased in most RMS cell lines over the course of eight days. As APOBEC2 overexpression promoted the growth of Rh30 spheroids in vitro, it may be therapeutically beneficial to prevent the increase in APOBEC2 levels in proliferating RMS tumor cells. This increase in expression may help sustain stem-like characteristics, which are more evident in 3D tumor models than in 2D culture. Accordingly, single-myofiber cultures revealed that muscle stem cells from APOBEC2 KO mice display lower self-renewal properties [44]. Notably, our findings indicate that PANX1 overexpression reduced the increase in APOBEC2 levels in three (Rh28, Rh30 and RD) of the six cell lines assessed. Little is known regarding the regulation of APOBEC2 levels but it has been reported that in Xenopus, APOBEC2 expression is regulated by TGF-β (transforming growth factor β) [43, 45, 46]. In human hepatocytes, APOBEC2 expression is significantly increased in response to both TNF-α (tumor necrosis factor α) and IL-1β (interleukin-1β) through NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells) activation [47]. Interestingly, we found that NF-κB2 protein levels are elevated when PANX1 levels are increased in Rh30 cells. NF-κB2 encodes the precursor protein p100 that, when cleaved, produces the functional protein p52 [48]. p52 protein levels were very low in all conditions, but p100 was significantly increased after eight days of PANX1 induction (Supplemental Fig. 1). As p100 forms complexes (kappaBsomes) that interact with and inhibits other NF-κB proteins [48], its increase in PANX1 overexpressing cells could potentially lead to NF-κB inhibition and subsequent downregulation of APOBEC2 expression.
Although the reason why increasing PANX1 levels does not reduce APOBEC2 expression in some RMS cells remains unclear, it may be due to mutations or inactivation of the relevant downstream pathways at play. Future exploration of transcriptomic, epigenetic mutational profiles may provide an explanation. Nevertheless, the use of several patient-derived cell lines in this current study was beneficial in identifying the functional relevance of this regulation. Indeed, the RMS cells in which PANX1 promoted cell multinucleation are coinciding with cell lines where APOBEC2 was downregulated. Further, APOBEC2 overexpression inhibited the PANX1-mediated RMS cell multinucleation, which was found to be due to cell fusion – a key process of myogenesis. While Panx1 regulates myoblast fusion only in male mice [25], the effect seen here in RMS is not sex dependent. Indeed, PANX1 overexpression promoted APOBEC2 downregulation and multinucleation in cell lines from male (Rh30 and Rh28) and female (RD) individuals. Our finding that the downregulation of APOBEC2 levels promotes cell fusion of RMS cells is in accordance with those obtained using healthy skeletal muscle cells. Ohtsubo et al. demonstrated that myoblasts from APOBEC2 KO mice displayed earlier upregulation of myogenin levels together with enhanced fusion following induction of differentiation [32]. Myogenin promotes myocyte fusion by binding to the myomaker promoter, being required for the expression of myomaker and other genes essential for myocyte fusion [49]. We have previously reported that PANX1 overexpression led to an increase in the proportion of Rh30 cells that express myogenin [26]. However, we have not found a PANX1-mediated regulation of myogenin expression in the other cell lines (RD and Rh28) in which PANX1 triggers APOBEC2 downregulation and cell fusion (data not shown). This suggests that RMS cell fusion induced by PANX1-mediated downregulation of APOBEC2 involves a novel mechanism that differs from normal skeletal muscle cells.
While still demonstrating a role of APOBEC2 in myoblast differentiation and fusion, results from a recent study using C2C12 myoblasts targeting APOBEC2 using shRNA were contradictory [33]. This may be due to the experimental set up and myogenic status of the cells as the knockdown was done prior to inducing differentiation. Nevertheless, it was demonstrated that APOBEC2 binds chromatin and regulates transcription of non-muscle genes during myoblast differentiation, playing a role in cell fate specification [33].
Here we employed two distinct expression systems to induce PANX1 expression that included appropriate negative controls, either an empty plasmid or a GFP-alone vector. Our experiments demonstrated that PANX1 overexpression induced cellular fusion in three of the six RMS cell lines assessed. This strongly suggests that the observed phenotype is not a generalized artifact of overexpression but rather a function dependent on the biological context. These findings support the specificity of PANX1 in driving the observed effect. However, overexpression systems may alter downstream pathways or binding partner availability in ways that are not easily predicted. Unbiased RNA-seq or proteomics may be beneficial to comprehensively exclude such possibilities in future studies.
While we demonstrated that PANX1 is involved in RMS cell fusion, the inhibition of cell proliferation and viability by PANX1 overexpression seen here and in our previous studies [26, 28] seems to occur independently of the downregulation of APOBEC2 levels. APOBEC2 was overexpressed in Rh30 cells at a level comparable to that of its endogenous expression in differentiating HSMM for physiological relevance. However, an inducible system may have been beneficial to observe subtle effects. Nevertheless, our findings suggest that the potent inhibitory effect of PANX1 on RMS malignancy occurs via several downstream pathways regulating different functions and phenotypes. It is interesting to note that increasing PANX1 levels alone can drive RMS cells further into the myogenic program whereas when examined using myoblasts, differentiation needed to be triggered by serum deprivation to facilitate extensive fusion. Despite this, no significant reversal of PANX1-mediated inhibition of RMS spheroid growth was identified by increasing APOBEC2 levels. This could suggest that the induction of cell fusion mediated by APOBEC2 downregulation is not sufficient by itself to alter spheroid growth. However, the rapid apoptosis triggered by PANX1 overexpression in this assay [26] likely hinders the ability to examine the effect of cell fusion in this context. While the role of APOBEC2 in the PANX1-mediated inhibition of tumor progression or promotion of differentiation in vivo was not examined here, it is valuable to note that Rh30 xenografts overexpressing PANX1, which grew significantly slower than their control counterparts [26], also displayed a reduction in APOBEC2 levels.
Approaches that induce RMS differentiation have increasingly gained significant interest as they may overcome the myogenic blockade and/or make tumor cells more sensitive to chemotherapy [50]. Based on our work, elevating PANX1 levels or reducing APOBEC2 expression in RMS would be of therapeutic benefit. While investigating the mechanisms regulating PANX1 expression, we discovered that quercetin can significantly increase PANX1 levels in RMS [27]. Notably, quercetin treatment promoted the differentiation of RMS cells in a PANX1-dependent manner [27]. The therapeutic potential of this approach was shown in vitro as quercetin treatment prevented 3D RMS spheroid growth and induced complete regression of established spheroids [27]. Future studies will investigate the benefit of quercetin, as well as strategies targeting APOBEC2, on tumor growth and differentiation in vivo, when used alone or in combination with chemotherapeutic drugs. Indeed, we have previously demonstrated that increasing PANX1 levels in RMS cells inhibits their malignant properties through a novel mechanism independent of its canonical channel activity that we are further delineating. Here, we have used a panel of RMS cell lines to show that PANX1 triggers RMS cell fusion by downregulating APOBEC2 expression, forcing these undifferentiated malignant cells further into the myogenic pathway.
Materials and methods
Tissue specimens, cell lines, and xenografts
Seven paraffin-embedded pediatric skeletal muscle and nine RMS samples were used. All human RMS cell lines were from Dr. P. Houghton (St. Jude Children’s Hospital, Memphis, TN, USA), except the RD (eRMS; female) cell line (American Type Culture Collection): Rh28 (aRMS; male), Rh30 (aRMS; male), Rh41 (aRMS; female), Rh36 (eRMS; male) [51, 52]. Rh18 was generated from a patient (female) tumor xenografted in immunodeficient mice. The tumor was predominately eRMS with some aRMS histologic elements, but the resulting xenograft from which the Rh18 cell line was derived was entirely embryonal [52]. Immortalized HSMM were from Dr. B. Chazaud (University Claude Bernard Lyon, Lyon, France) [40, 53]. Mouse myoblasts were isolated as previously described [25]. RMS cells and HSMM were cultured as previously described [28, 40]. Cells were mycoplasma-negative.
Xenografts were from cumate-treated Rh30 cells injected into the gastrocnemius muscle of 4- to 6-week-old female SCID mice (3 mice: PANX1-expressing cells into the right leg; empty vector cells into the left leg) [26]. Mice were injected with cumate daily [26] and sacrificed when tumors reached 2000 mm3. Experiments were approved by the Animal Care Committee at the University of Ottawa and conducted per the Canadian Council of Animal Car guidelines.
Stable cell lines generation and transfections
Stable Rh18 and Rh30 cell lines in which PANX1 expression is controlled by a cumate (30 µg/ml) switch system were previously described [26]. The other cumate-inducible RMS cell lines were generated using the same methodology. These cells express GFP constitutively.
APOBEC2 overexpression in the cumate-inducible Rh30 cell line and its control counterpart were generated using an APOBEC2 expression vector (pLV[Exp]-Bsd-TRE>hAPOBEC2[NM_006789.4], ID: VB210721-1157dvf) or its control vector (pLV[Exp]-Bsd-TRE > ORF_Stuffer, ID: VB210809-1167qsh) obtained from Vector Builder. Vectors were packaged into lentiviruses (SBI, #LV100A-1) and tittered using the Global UltraRapid Lentiviral Titer Kit (SBI, #LV961A-1). Selection was performed with blasticidin. An MOI of 100 was used as the level of APOBEC2 achieved was similar to that of differentiating HSMM.
Stable Rh30 cell lines with red or green nuclei were generated by infection using Incucyte Nuclight red (nuclear-restricted mKate2, Sartorius, #4625)) or Nuclight green (nuclear-restricted green fluorescent protein, Sartorius, #4624)) lentiviruses, followed by selection using puromycin.
The untagged PANX1 overexpression vector, pRP[Exp]-Puro-CAG>hPANX1 (ID: VB90091-9898zkn), was from Vector Builder. Transfections were performed using Lipofectamine 2000 reagent (ThermoFisher Scientific).
RT-qPCR analysis
Total RNA was extracted using RNeasy Mini Kit (Quigen, MD, USA) followed by reverse transcription by High-Capacity cDNA Reverse Transcription Kit (ThermoFisher Scientific). The synthesized cDNA was used as a template with the SsoAdvanced™ Universal SYBR® Green Supermix (Bio-Rad, California, USA) for quantitative PCR on Mastercycler ep-realplex (Eppendorf, Hamburg, Germany) real-time PCR system with validated primers for APOBEC2 (forward 5’- AGAACCTGGACGACCCTGAGAA -3’; reverse 5’- CAACCACATAGCAGAGGAAGGTC -3’) and GAPDH (forward 5’- CAAGACCTTGGGCTGGGAC -3’; reverse 5’- AGGCTGCGGGCTCAATTTAT -3’) from Origene Technologies (Rockville, Maryland, USA). Relative expression and fold change were then determined using a comparative Ct method.
Western blotting
Cell lysis was performed as previously desbribed [40]. After blocking, membranes were incubated with anti-APOBEC2 (1:1000, Sigma-Aldrich, #HPA017957) or anti-PANX1 (1:2000, Sigma-Aldrich, #HPA016930), and then with Alexa 680- (1:5000, ThermoFisher Scientific, #A21009) or infrared fluorescent-labeled secondary antibodies IRDye800 (Li-COR Biosciences, Nebraska, USA, #925-32210) used at 1:5000. Anti-GAPDH (Advanced ImmunoChemistry, California, USA, Mab 6C5) was used for normalization. Western blots were processed using the Odyssey CLx Imaging System (Li-COR).
Immunofluorescence microscopy
Specimen sections were deparaffinized and rehydrated. Antigen retrieval was performed using 10 mM citrate buffer (pH 6.0) in a pressure cooker. Following permeabilization and blocking, samples were labeled with anti-APOBEC2 (1:100, Sigma-Aldrich, #HPA017957) and then with anti-rabbit secondary antibodies conjugated to Alexa Fluor 594 (ThermoFisher Scientific, #A11012). A negative control was done without primary antibody. Sections were mounted with DAPI Fluoromount-G (ThermoFisher Scientific, #00-4959-52) and visualized with the Olympus Fluoview FV1000 confocal microscope. Quantification was done using the ImageJ software and averaged from four random fields of the same area for each sample. Positive labeling in normal skeletal muscle and negative labeling in the absence of primary antibody were used to optimize the identification of cut-offs for positive labeling in the absence of image saturation as well as background. Once set, these parameters where then consistently applied to all samples sequentially without adjustment. Results are presented as the average of integrated fluorescence arbitrary units.
Cells were fixed and permeabilized. Non-specific labeling was blocked, and cells were incubated with anti-APOBEC2 (1:200), anti-PANX1 (1:200), or anti-Pax7 (0.3 μg/mL, DHSB, Iowa City, Iowa, USA). After washing, cells were incubated with secondary antibodies (anti-rabbit Alexa Fluor 488 (ThermoFisher Scientific, #A-11008), anti-rabbit Alexa Fluor 594 (ThermoFisher Scientific, A-11012), or anti-mouse Alexa Fluor 488 (ThermoFisher Scientific, A-11001)), and mounted with DAPI Fluoromount-G. Images were collected sequentially maintaining constant settings using the EVOS FL Auto (ThermoFisher Scientific) or the Olympus Fluoview FV1000 confocal microscope.
The fusion index, which represents the percentage of fusion events in a cell population, is defined as [(N-S)/T] × 100, where N is the number of nuclei in the syncytia, S is the number of syncytia (cells with at least three nuclei), and T is the number of nuclei counted [54, 55].
Proliferation assay
Cells were treated with cumate to induce PANX1 expression. After two or four days, cells were incubated with 10 µM BrdU incorporating reagent (BrdU Cell Proliferation ELISA Kit (Abcam, Cambridge, UK, #ab126556)) for two hours. Results were analyzed using a single read at 450 ηm on a Synergy HTX plate reader (BioTek, Vermont, USA).
Viability assay
Cells were treated with or without cumate for two or four days. Cells and supernatants were combined with 0.4% Trypan Blue solution. Viability was determined using the Countess automated cell counter (ThermoFisher Scientific).
Spheroid formation assay
Spheroid were analyzed using the IncuCyte Zoom system (Essen Bioscience, Michigan, USA), as previously described [26].
Cell fusion assay
Stable Rh30 cell lines with red and green nuclei were seeded together (1:1) and transfected to overexpress PANX1. Following PANX1 labeling and mounting with DAPI Fluoromount G, images were collected with the Olympus Fluoview FV1000 confocal microscope.
Statistics
Statistical significance was analyzed using unpaired two-tailed Student’s t tests, and analysis of variance (ANOVA) followed by Tukey’s or Sidak’s post hoc tests. Data are presented as mean ± standard deviation (SD).
Data availability
All data generated or analysed during this study are included in this published article (and its supplementary files).
References
Chen C, Dorado Garcia H, Scheer M, Henssen AG. Current and future treatment strategies for rhabdomyosarcoma. Front Oncol. 2019;9:1458.
Paulino AC, Okcu MF. Rhabdomyosarcoma. Curr Probl Cancer. 2008;32:7–34.
Dobson CC, Langlois S, Grynspan D, Cowan KN, Holcik M. Engaging cell death pathways for the treatment of rhabdomyosarcoma. Crit Rev Oncog. 2016;21:221–39.
Malempati S, Hawkins DS. Rhabdomyosarcoma: review of the Children’s Oncology Group (COG) Soft-Tissue Sarcoma Committee experience and rationale for current COG studies. Pediatr Blood Cancer. 2012;59:5–10.
Parham DM, Barr FG. Classification of rhabdomyosarcoma and its molecular basis. Adv Anat Pathol. 2013;20:387–97.
Punyko JA, Mertens AC, Gurney JG, Yasui Y, Donaldson SS, Rodeberg DA, et al. Long-term medical effects of childhood and adolescent rhabdomyosarcoma: a report from the childhood cancer survivor study. Pediatr Blood Cancer. 2005;44:643–53.
Zarrabi A, Perrin D, Kavoosi M, Sommer M, Sezen S, Mehrbod P, et al. Rhabdomyosarcoma: current therapy, challenges, and future approaches to treatment strategies. Cancers. 2023;15:5269.
Chisholm JC, Marandet J, Rey A, Scopinaro M, de Toledo JS, Merks JHM, et al. Prognostic factors after relapse in nonmetastatic rhabdomyosarcoma: a nomogram to better define patients who can be salvaged with further therapy. J Clin Oncol. 2011;29:1319–25.
Dasgupta R, Fuchs J, Rodeberg D. Rhabdomyosarcoma. Semin Pediatr Surg. 2016;25:276–83.
Biressi S, Molinaro M, Cossu G. Cellular heterogeneity during vertebrate skeletal muscle development. Dev Biol. 2007;308:281–93.
Hettmer S, Li Z, Billin AN, Barr FG, Cornelison DDW, Ehrlich AR, et al. Rhabdomyosarcoma: current challenges and their implications for developing therapies. Cold Spring Harb Perspect Med. 2014;4:a025650.
Nanni P, Nicoletti G, Palladini A, Astolfi A, Rinella P, Croci S, et al. Opposing control of rhabdomyosarcoma growth and differentiation by myogenin and interleukin 4. Mol Cancer Ther. 2009;8:754–61.
Wu Y-L, Yang A-H, Chiu Y-H. Recent advances in the structure and activation mechanisms of metabolite-releasing Pannexin 1 channels. Biochem Soc Trans. 2023;51:1687–99.
Zoidl G, Petrasch-Parwez E, Ray A, Meier C, Bunse S, Habbes H-W, et al. Localization of the pannexin1 protein at postsynaptic sites in the cerebral cortex and hippocampus. Neuroscience. 2007;146:9–16.
Langlois S, Cowan KN. Regulation of skeletal muscle myoblast differentiation and proliferation by Pannexins. Adv Exp Med Biol. 2017;925:57–73.
Alhouayek M, Sorti R, Gilthorpe JD, Fowler CJ. Role of pannexin-1 in the cellular uptake, release and hydrolysis of anandamide by T84 colon cancer cells. Sci Rep. 2019;9:7622.
Penuela S, Bhalla R, Gong X-Q, Cowan KN, Celetti SJ, Cowan BJ, et al. Pannexin 1 and pannexin 3 are glycoproteins that exhibit many distinct characteristics from the connexin family of gap junction proteins. J Cell Sci. 2007;120:3772–83.
Cowan KN, Langlois S, Penuela S, Cowan BJ, Laird DW. Pannexin1 and Pannexin3 exhibit distinct localization patterns in human skin appendages and are regulated during keratinocyte differentiation and carcinogenesis. Cell Commun Adhes. 2012;19:45–53.
Pelegrin P, Surprenant A. Pannexin-1 mediates large pore formation and interleukin-1beta release by the ATP-gated P2X7 receptor. EMBO J. 2006;25:5071–82.
Bruzzone R, Hormuzdi SG, Barbe MT, Herb A, Monyer H. Pannexins, a family of gap junction proteins expressed in brain. Proc Natl Acad Sci USA. 2003;100:13644–9.
Celetti SJ, Cowan KN, Penuela S, Shao Q, Churko J, Laird DW. Implications of pannexin 1 and pannexin 3 for keratinocyte differentiation. J Cell Sci. 2010;123:1363–72.
Langlois S, Xiang X, Young K, Cowan BJ, Penuela S, Cowan KN. Pannexin 1 and pannexin 3 channels regulate skeletal muscle myoblast proliferation and differentiation. J Biol Chem. 2014;289:30717–31.
Pham TL, St-Pierre M-E, Ravel-Chapuis A, Parks TEC, Langlois S, Penuela S, et al. Expression of Pannexin 1 and Pannexin 3 during skeletal muscle development, regeneration, and Duchenne muscular dystrophy. J Cell Physiol. 2018;233:7057–70.
Suarez-Berumen K, Collins-Hooper H, Gromova A, Meech R, Sacco A, Dash PR, et al. Pannexin 1 regulates skeletal muscle regeneration by promoting bleb-based myoblast migration and fusion through a novel lipid based signaling mechanism. Front Cell Dev Biol. 2021;9:736813.
Freeman E, Langlois S, Scott K, Ravel-Chapuis A, Jasmin BJ, Cowan KN. Sex-dependent role of Pannexin 1 in regulating skeletal muscle and satellite cell function. J Cell Physiol. 2022;237:3944–59.
Xiang X, Langlois S, St-Pierre M-E, Barré JF, Grynspan D, Purgina B, et al. Pannexin 1 inhibits rhabdomyosarcoma progression through a mechanism independent of its canonical channel function. Oncogenesis. 2018;7:89.
Xiang X, Hoang H-D, Gilchrist VH, Langlois S, Alain T, Cowan KN. Quercetin induces pannexin 1 expression via an alternative transcript with a translationally active 5’ leader in rhabdomyosarcoma. Oncogenesis. 2022;11:9.
Xiang X, Langlois S, St-Pierre M-E, Blinder A, Charron P, Graber TE, et al. Identification of pannexin 1-regulated genes, interactome, and pathways in rhabdomyosarcoma and its tumor inhibitory interaction with AHNAK. Oncogene. 2021;40:1868–83.
Sato Y, Probst HC, Tatsumi R, Ikeuchi Y, Neuberger MS, Rada C. Deficiency in APOBEC2 leads to a shift in muscle fiber type, diminished body mass, and myopathy. J Biol Chem. 2010;285:7111–8.
Liao W, Hong SH, Chan BH, Rudolph FB, Clark SC, Chan L. APOBEC-2, a cardiac- and skeletal muscle-specific member of the cytidine deaminase supergene family. Biochem Biophys Res Commun. 1999;260:398–404.
Conticello SG. The AID/APOBEC family of nucleic acid mutators. Genome Biol. 2008;9:229.
Ohtsubo H, Sato Y, Suzuki T, Mizunoya W, Nakamura M, Tatsumi R, et al. APOBEC2 negatively regulates myoblast differentiation in muscle regeneration. Int J Biochem Cell Biol. 2017;85:91–101.
Lorenzo JP, Molla L, Amro EM, Ibarra IL, Ruf S, Neber C, et al. APOBEC2 safeguards skeletal muscle cell fate through binding chromatin and regulating transcription of non-muscle genes during myoblast differentiation. Proc Natl Acad Sci USA. 2024;121:e2312330121.
Kostic C, Shaw PH. Isolation and characterization of sixteen novel p53 response genes. Oncogene. 2000;19:3978–87.
Okuyama S, Marusawa H, Matsumoto T, Ueda Y, Matsumoto Y, Endo Y, et al. Excessive activity of apolipoprotein B mRNA editing enzyme catalytic polypeptide 2 (APOBEC2) contributes to liver and lung tumorigenesis. Int J Cancer. 2012;130:1294–1301.
Li A, Wu J, Zhai A, Qian J, Wang X, Qaria MA, et al. HBV triggers APOBEC2 expression through miR‑122 regulation and affects the proliferation of liver cancer cells. Int J Oncol. 2019;55:1137–48.
Shi R, Wang X, Wu Y, Xu B, Zhao T, Trapp C, et al. APOBEC-mediated mutagenesis is a favorable predictor of prognosis and immunotherapy for bladder cancer patients: evidence from pan-cancer analysis and multiple databases. Theranostics. 2022;12:4181–99.
Wei L, Wu X, Wang L, Chen L, Wu X, Song T, et al. Expression and prognostic value of APOBEC2 in gastric adenocarcinoma and its association with tumor-infiltrating immune cells. BMC Cancer. 2024;24:15.
Florkowska A, Meszka I, Nowacka J, Granica M, Jablonska Z, Zawada M, et al. PAX7 balances the cell cycle progression via regulating expression of Dnmt3b and Apobec2 in differentiating PSCs. Cells. 2021;10:2205.
Freeman E, Langlois S, Leyba MF, Ammar T, Léger Z, McMillan HJ, et al. Pannexin 1 dysregulation in Duchenne muscular dystrophy and its exacerbation of dystrophic features in mdx mice. Skelet Muscle. 2024;14:8.
Taulli R, Bersani F, Foglizzo V, Linari A, Vigna E, Ladanyi M, et al. The muscle-specific microRNA miR-206 blocks human rhabdomyosarcoma growth in xenotransplanted mice by promoting myogenic differentiation. J Clin Investig. 2009;119:2366–78.
Keller C, Guttridge DC. Mechanisms of impaired differentiation in rhabdomyosarcoma. FEBS J. 2013;280:4323–34.
Powell C, Cornblath E, Goldman D. Zinc-binding domain-dependent, deaminase-independent actions of apolipoprotein B mRNA-editing enzyme, catalytic polypeptide 2 (Apobec2), mediate its effect on zebrafish retina regeneration. J Biol Chem. 2014;289:28924–41.
Ohtsubo H, Sato Y, Suzuki T, Mizunoya W, Nakamura M, Tatsumi R, et al. Data supporting possible implication of APOBEC2 in self-renewal functions of myogenic stem satellite cells: Toward understanding the negative regulation of myoblast differentiation. Data Brief. 2017;12:269–73.
Etard C, Roostalu U, Strähle U. Lack of Apobec2-related proteins causes a dystrophic muscle phenotype in zebrafish embryos. J Cell Biol. 2010;189:527–39.
Vonica A, Rosa A, Arduini BL, Brivanlou AH. APOBEC2, a selective inhibitor of TGFβ signaling, regulates left-right axis specification during early embryogenesis. Dev Biol. 2011;350:13–23.
Matsumoto T, Marusawa H, Endo Y, Ueda Y, Matsumoto Y, Chiba T. Expression of APOBEC2 is transcriptionally regulated by NF-kappaB in human hepatocytes. FEBS Lett. 2006;580:731–5.
Ghosh G, Wang VY-F. Origin of the Functional Distinctiveness of NF-κB/p52. Front Cell Dev Biol. 2021;9:764164.
Ganassi M, Badodi S, Ortuste Quiroga HP, Zammit PS, Hinits Y, Hughes SM. Myogenin promotes myocyte fusion to balance fibre number and size. Nat Commun. 2018;9:4232.
Pomella S, Danielli SG, Alaggio R, Breunis WB, Hamed E, Selfe J, et al. Genomic and epigenetic changes drive aberrant skeletal muscle differentiation in rhabdomyosarcoma. Cancers. 2023;15:2823.
Hazelton BJ, Houghton JA, Parham DM, Douglass EC, Torrance PM, Holt H, et al. Characterization of cell lines derived from xenografts of childhood rhabdomyosarcoma. Cancer Res. 1987;47:4501–7.
Hinson ARP, Jones R, Crose LES, Belyea BC, Barr FG, Linardic CM. Human rhabdomyosarcoma cell lines for rhabdomyosarcoma research: utility and pitfalls. Front Oncol. 2013;3:183. https://doi.org/10.3389/fonc.2013.00183.
Massenet J, Gitiaux C, Magnan M, Cuvellier S, Hubas A, Nusbaum P, et al. Derivation and characterization of immortalized human muscle satellite cell clones from muscular dystrophy patients and healthy individuals. Cells. 2020;9:1780.
Sugimoto J, Schust DJ, Yamazaki T, Kudo Y. Involvement of the HERV-derived cell-fusion inhibitor, suppressyn, in the fusion defects characteristic of the trisomy 21 placenta. Sci Rep. 2022;12:10552.
Karakis V, Britt JW, Jabeen M, San Miguel A, Rao BM. Derivation of human trophoblast stem cells from placentas at birth. J Biol Chem. 2025;301:108505.
Acknowledgements
We gratefully acknowledge Dr. David Grynspan for his help in retrieving the patient samples. We are also thankful for the staff for the sectioning services provided by the Louise Pelletier Histology Core Facility at the University of Ottawa.
Funding
Canadian Institutes of Health Research-Institute of Cancer Research (CIHR-ICR)/Cancer Research Society (Operating Grant 943994, KNC), Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grant (RGPIN-2021-03801), Ontario Graduate Scholarship (OGS, XX), NSERC Canada Graduate Scholarship (AW), Queen Elizabeth II Graduate Scholarship in Science and Technology (QEII-GSST, AW, XX, and EF), University of Ottawa Destination 2020 (XX), Dr. Alex Mackenzie Trainee Grant (EF).
Author information
Authors and Affiliations
Contributions
Conceptualization and design: AW, SL, and KNC Development of methodology: AW, AB, SL, XX, and KNC Acquisition of data: AW, AB, XX, KG, and EF Analysis and interpretation of data: AW, AB, XX, SL, and KNC Writing of manuscript: SL, AB, and KNC Review of Manuscript: AW, AB, SL, XX, KG, EF, and KNC Project administration: KNC Study Supervision: SL, and KNC
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
After institutional ethics board approval (CHEO Research Ethics Board, protocol CHEOREB #11/148X), archived paraffin-embedded pediatric skeletal muscle and RMS samples were obtained, as secondary use of clinical samples, from the CHEO Department of Pathology and Laboratory Medicine (Ottawa, Ontario, Canada). Experiments were carried out in accordance with the CHEO Ethics Board guidelines and regulations. The study was conducted in accordance with the principles of the Declaration of Helsinki. Data were anonymized and all identifying information was removed prior to analysis. The secondary use of samples is consistent with the original consent obtained from all participants. Animal experiments were approved by the Animal Care Committee at the University of Ottawa and conducted per the Canadian Council of Animal Care guidelines (CHEO-3685 and CHEO-2123).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Welten, A., Bera, A., Langlois, S. et al. Pannexin 1 induces Rhabdomyosarcoma cell fusion by downregulating APOBEC2. Oncogenesis 14, 43 (2025). https://doi.org/10.1038/s41389-025-00586-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41389-025-00586-x