Abstract
The ADP-ribosyltransferase, PARP1 enzymatically generates and applies the post-translational modification, ADP-Ribose (ADPR). PARP1 roles in genome maintenance are well described, but recent work highlights roles in many fundamental processes including cellular identity and energy homeostasis. Herein, we show in both mouse and human skeletal muscle cells that PARP1-mediated PARylation is a regulator of the myogenic program and the muscle transcriptional response to steroid hormones. Chemical PARP1 modulation impacts the expression of major myocellular proteins, including troponins, key in dictating muscle contractile force. Whilst PARP1 in absence of DNA damage is often assumed to be basally inactive, we show PARylation to be acutely sensitive to extracellular glucose concentrations and the steroid hormone class, glucocorticoids which exert considerable authority over muscle tissue mass. Specifically, we find during myogenesis, a transient and significant rise in PAR. This early-stage differentiation event, if blocked with PARP1 inhibition, reduced the abundance of important muscle proteins in the fully differentiated myotubes. This suggests that PAR targets during early-stage differentiation are central to the proper development of the muscle contractile unit. We also show that reduced PARP1 in myoblasts impacts a variety of metabolic pathways in line with the recorded actions of glucocorticoids. Currently, as both regulators of myogenesis and muscle mass loss, glucocorticoids represent a clinical conundrum. Our work goes on to identify that PARP1 influences transcriptional activation by glucocorticoids of a subset of genes critical to human skeletal muscle pathology. These genes may therefore signify a regulatory battery of targets through which selective glucocorticoid modulation could be achieved. Collectively, our data provide clear links between PARP1-mediated PARylation and skeletal muscle homeostatic mechanisms crucial to tissue mass maintenance and endocrine response.
Similar content being viewed by others
Introduction
PARP1 irreversibly cleaves NAD+, producing nicotinamide for re-salvage and monomeric ADP-Ribose (ADPR) units. ADPR functions as a signaling moiety through site-specific attachment to target molecules, altering their biological activity. ADPR units can be mono- or poly-elongated: the latter produces poly-ADPR (PAR) chains, in a process termed PARylation [1]. To date, PARP1, alongside PARP2 and the tankyrases, are capable of performing PARylation. Recent studies of PARylation have revealed roles ancillary to the well-characterized genome repair. For example, PARP1 activity has been shown to govern fundamental processes including adipogenesis, RNA stability, and transdifferentiation [2,3,4,5,6].
The conserved and constitutive nature of PARP1 and PARylation underlies the variety of documented actions. Emerging studies implicate PARP1 within skeletal muscle metabolism and myogenesis [4, 7,8,9]. For example, in myogenic progression, downregulation of PARP1 in fully formed myotubes is required for oxidative stress resistance [9]. More recent work has demonstrated PARP1 binding to regulatory regions of the MYOD target muscle genes p57 and myogenin [4]. These works demonstrate PARP1 also exerts myogenic influence independent of its PARylating activity, however, the impacts of PARylation in skeletal muscle remain poorly characterized. Despite this incomplete understanding, PARP inhibitors have been suggested for alleviation of inflammation in non-communicable chronic diseases including myopathy.
Herein, we used transcriptomic and proteomic analyses to explore PARP1-mediated PARylation during myogenic progression. We show PARP1 and the PAR it applies during myogenesis are dynamic, and sensitive to both hormonal status and metabolic demand. Early-stage muscle cell differentiation sees a transient rise in PARylation that holds influence over the fully developed myotube. This indicates that the molecular targets of PARP1 during early-stage differentiation are pivotal in determining the functional quality of the muscle fiber. We also demonstrate that PARP1 holds impacts over the skeletal muscle transcriptional response to glucocorticoids, steroid hormones with governance over muscle protein synthesis and metabolic rate. We identify a subset of genes that are critical to muscle mass and contractile function that are co-regulated by PARP1. These results reveal whilst PARP1 inhibition mediates beneficial effects in fully developed muscle tissue, the potential for negative impacts exists for muscle differentiation and steroid hormone activation. Therefore, the use of clinically available PARP inhibition should be subject to greater consideration for impacts on the whole-body muscle mass.
Results
PARP1-mediated PARylation is dynamic during skeletal muscle differentiation
Molecular assessment of PARP1 and PAR during myogenesis remains limited. Given PARP1 non-enzymatically regulates the myogenic regulatory factor MYOD [4], we hypothesized that enzymatically driven PARylation also contributes to myogenic transition. To address this, C2C12 myoblasts were differentiated and interrogated for PAR and PARP1 levels on each day of differentiation. We report that levels of PAR were dynamic during myoblast alignment and fusion, reaching peak within day 1 and nadir by day 5 of differentiation (Fig. 1A, B). Identical dynamic regulation of PAR and PARP1 abundance was recorded in the human muscle cell line LHCN-M2 (Fig. 1C, D). Total levels of myogenin also increased, demonstrating the establishment of myogenic commitment in these myoblasts (Fig. 1A, C). Together, these data indicate, at least in mammals, a conserved process of PAR regulation driven by PARP1 that is associated with the initiation of the myogenic program.
A Western immunoblotting of differentiating C2C12 myoblasts probed for PARylation (PAR), PARP1, Myogenin, and Alpha-Tubulin (representative of n = 4). B Quantification of MYOG, PAR, and PARP1 present over differentiation. Each bar represents means ± S.D (n = 4) ***P < 0.001. C Western immunoblotting of differentiating LHCN-M2 human myoblasts probed for MYOG, PAR, PARP1, Myogenin, and Alpha-Tubulin (representative of n = 4). D Quantification of PAR and PARP1 present over differentiation. Each bar represents means ± S.D (n = 4) ***P < 0.001.
PARylation in skeletal muscle myogenesis is metabolically sensitive
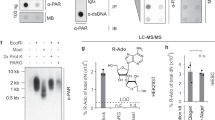
Conventionally, PARP1 is regarded as inactive in the absence of genotoxic stress. However, several studies have demonstrated that both PARP1 and PARylation are fundamental to a variety of biological processes [1]. As such, we explored the response of PARP1 and PAR to conditions of NAD+ excess or deficit during myogenesis. The NAD+ precursor nicotinamide riboside (NR) is bioavailable to human skeletal muscle and can directly generate NAD+ [10]. However, we find that NR supplementation of myoblasts during differentiation induction did not alter the levels of day 1 PAR accumulation (Fig. 2A, B). In contrast, we find NAD+ depletion through specific NAMPT inhibition with FK866 reduced day 1 PAR levels and myogenin expression, supporting an association of the enzymatic product of PARP1 with myogenic progression (Fig. 2C, D). We also found PAR during myogenesis to be regulated by the external cellular environment, with a dose-dependent increase in PAR in response to glucose deprivation (Fig. 2G, H) and is sensitive to steroid hormone levels following exposure to the glucocorticoid receptor (GR) agonist dexamethasone (Fig. 2E, F). Collectively, these data show that PARP1-mediated PARylation is active in differentiating myoblasts and exerts plasticity. Moreover, we find a synergistic interaction of both the inhibition of PARP1 with the inhibitor PJ34 and dexamethasone treatment during differentiation induction upon subsequent PAR abundance (Fig. 2I, J). Together, these data show PARP1 and PAR in skeletal muscle to be basally detected and responsive to the changing cellular environment. This implies PAR in skeletal muscle is a signaling motif with potential roles to play in the variety of pathways governing muscle homeostasis.
A, B Western immunoblotting of differentiating myoblasts differentiated in ±nicotinamide riboside (NR) (0.5 mM) (representative of n = 4) probed for PAR, PARP1, MYOD, and Alpha-Tubulin. C, D Western immunoblotting of differentiating myoblasts differentiated in ± NAMPT specific inhibitor FK866 (50 nM) (representative of n = 4) probed for PAR, PARP1, Myogenin, and Alpha-Tubulin, ***P < 0.001. E, F Western immunoblotting of differentiating myoblasts differentiated in ±dexamethasone (1 µM) probed for PAR, PARP1, MYOD and Alpha-Tubulin (representative of n = 4, **P < 0.01). G, H C2C12 myoblasts differentiated in differentiation medium containing different glucose concentrations before lysate harvest and immunoblotted for PAR, PARP1, Hexokinase II, and Alpha-Tubulin (representative of n = 3, *P < 0.05). I, J Western immunoblotting of differentiating myoblasts differentiated in ±dexamethasone (1 µM), PARP inhibitor PJ34 (10 µM) or both in combination (representative of n = 3, *P < 0.05, **P < 0.01).
Modulation of PARylation during early-stage myogenesis
To modulate PAR generation, we applied the broad-spectrum PARP inhibitor PJ34 during differentiation induction and demonstrate significant reduction of early-stage PAR accumulation in both murine and human skeletal muscle cells (Fig. 3A–D). PJ34 treatment also significantly increased cellular NAD+ (Supplementary Fig. 1A). To further investigate if the transient day 1 PAR accumulation is a determinant of developed myotube functional quality, we also assessed levels of Troponin T1 (TNNT1), a subunit of the sarcomere. This was reduced in myotubes exposed to PJ34 during early-stage differentiation and suggests PARylation is required for proper thin filament assembly (Fig. 3A, C). These PAR trends were also replicated using Rucaparib (Supplementary Fig. 2A, B). Inhibition of PARP1 with PJ34 is also shown by immunofluorescence, with the day 1 PAR accumulation impaired in the PJ34-treated differentiating myoblasts (Fig. 3E). To identify the major transcriptional processes governed by PARP1 inhibition, we conducted RNAseq on PJ34 treated early-stage differentiating C2C12 myoblasts compared to vehicle controls (Fig. 3F). Gene Ontology analysis revealed suppression of pathways regulating assembly of the skeletal muscle myofibrils and detection of muscle stretch, a table of differentially expressed genes is provided (Supplementary Table S1). Additionally, in line with documented actions of PARP1 in transcriptional regulation [11] suppression of the phosphorylation of RNA polymerase II suggests global shifts in transcriptional rate as a result of reduced PARP1 activity. Collectively, these data confirm the dynamic nature of PAR in skeletal muscle and a background of transcriptional coordination during early-stage differentiation.
A Western immunoblotting of differentiating C2C12 myoblasts differentiated in ±PARP inhibitor PJ34 (10 µM) probed for PAR, PARP1, Troponin 1 (TNNT1), and Alpha-Tubulin (representative of n = 4). B Quantification of PAR and PARP1 during differentiation. Each bar represents means ± S.D (representative of n = 4). C Western immunoblotting of differentiating LHCN-M2 myoblasts differentiated in ±PARP inhibitor PJ34 (10 µM) probed for PAR, PARP1, TNNT1, and Alpha-Tubulin (n = 4) *P < 0.05, **P < 0.01, and ***P < 0.001. D Quantification of PAR and PARP1 in human myoblasts during differentiation. Each bar represents means ± S.D (representative of n = 4). E Immunostaining of PAR (red), DAPI (purple), and MYOD (green) in differentiating C2C12 myoblasts (n = 4). F Dot plot of gene ontology (GO) overrepresentation analysis of C2C12 myoblasts differentiated in ±PARP inhibitor PJ34 (10 µM) (n = 3) on day 1 of differentiation. The x axis shows the gene ratio which represents the percentage of genes enriched in a term. The y axis represents the enriched pathways: size of the node represents the number of enriched genes in the term.
Differentiating skeletal muscle and impacts of PARP1 upon the proteome
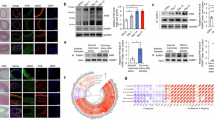
To gain insight into proteomic changes during differentiation regulated by PARP1-generated PAR, we subjected lysates of differentiating myoblasts treated with PJ34 (Fig. 3A) to unbiased SWATH-MS analysis. These recovered peptides aligned with 2911 identified proteins expressed from day 0 to day 6 of differentiation ±PJ34 (Fig. 4A). Examination of PJ34 treated day 1 differentiating lysates revealed differentially abundant proteins associated with documented roles of PARP1 in chromosome biology, and downregulated proteins (>1.5 fold change) included CEBPZ (−2.70 fold change, Pvalue 0.04), CBX6 (−1.59 fold change Pvalue 0.05), STIM2 (−2.37 fold change, Pvalue 0.004) and CDN1B (−2.09 fold change Pvalue 0.03) (Fig. 4B). These data describe muscle PARP1 as dictating chromatin features, consistent with regulation of transcriptional programs across tissues [12]. Subsequently, downregulated proteins from PJ34 treated day 3 differentiating lysates shifted towards association with skeletal muscle functions including MYL4 (−1.68 fold change Pvalue 0.10), and TNNI1 (−1.60 fold change Pvalue 0.11) (Fig. 4B). Proteins fundamental to skeletal muscle phenotype and contractility also shift in PJ34 treated day 5 differentiating lysates, including TNNT1 (−3.35 fold change, Pvalue 0.05), INSR (-4.36 fold change P value 0.04). A tabular list of differentially abundant proteins is provided (Supplementary Table S2).
A Heatmap showing 2911 detected proteins in differentiating C2C12 myoblasts differentiated in ± PARP inhibitor PJ34 (10 µM) over days 0, 1, 2, 3, 4, 5, and 6 of differentiation (n = 3 per condition). B Volcano plots showing differentially abundant proteins in presence of PJ34 on each day of the 6 days of differentiation. Downregulated proteins are green (>−1.5 fold change), upregulated are red (>1.5 fold change). Arrows denote named proteins.
Proper myoblast differentiation requires PARP1 activity
PJ34 is a broad PARP inhibitor that can target other members of the PARP family of enzymes, including the ADP-ribosylating tankyrases [13]. Given this, we sought to explore the observed proteomic changes and challenge the extent of day 1 PARP1 mediated PARylation over myogenesis. To do this, we also treated C2C12 myoblasts with the highly specific PARP1 inhibitor BYK204165 [14] during differentiation induction. We assessed levels of PAR as well as subsequent impacts on differentiation trajectory and observed ablation in total PAR protein levels in BYK204165 treated groups (Supplementary Fig. S1B). Myoblasts were treated on day 1 with BYK204165 for 24 h before washout and subsequently left to differentiate for 6 days with media changed every 48 h (Fig. 5A). Lysates were harvested and subjected to unbiased SWATH-MS analysis. This detected 2921 proteins, of which 180 were significantly differentially expressed in BYK204165-treated differentiating lysates (Fig. 5B and Supplementary Table S3). Of the downregulated proteins, these again included muscle contractile proteins including TNNT1 (−1.31 fold change), MYL4 (−1.86 fold change), TNNT3 (−1.53 fold change), MYL1 (−1.81 fold change) and MYH3 (−1.83 fold change) (Fig. 5C). Pathway overrepresentation analysis of these samples reveals these proteins associated to biological processes governing actin binding, cytoskeletal protein binding, cytoskeletal motor binding, troponin binding, actin filament binding and structural constituent of muscle (Supplementary Fig. S1C). These observations reflect our SWATH-MS analysis of PJ34-treated differentiating lysates (Fig. 4B) and highlight the importance of PARP1-mediated PARylation events that take place on day 1 of differentiation. To better understand how PARylation during early-stage differentiation influences the fully differentiated myotube, we employed Giemsa-Jenner staining of C2C12 myoblasts treated with either PJ34 or BYK204165 during differentiation induction. Cells were fixed on day 6 of differentiation and subjected to an unbiased method of quantification of myogenic differentiation [15]. This analysis revealed subsequently reduced fusion index of myotubes treated with PJ34 (20.15 ± 6.41 S.D) and BYK204165 (24.94 ± 3.21 S.D) (Fig. 5D). These suggest PAR elevation at early-stage differentiation impacts myocyte fusion and PARylation targets during early-stage differentiation are critical to the proper formation of the muscle sarcomere, impacting the overall number of contractile units per fiber.
A Schematic representation of the PARP inhibitor treatment protocol during C2C12 myoblast differentiation. B Heatmap representing differential abundance of proteins within samples differentiated in ±PARP1 specific inhibitor BYK204165 (10 µM) (Vehicle n = 7, BYK204165 n = 6). C Volcano plot of SWATH-LCMS lysates recovered from day 6 differentiated C2C12 myoblasts differentiated in ± PARP inhibitor PJ34 (10 µM). Differential protein abundance shown with downregulated proteins marked green and upregulated proteins marked red. D Myotube fusion index of differentiating myoblasts differentiated in ±PARP inhibitor PJ34 (10 µM) or ±PARP1 specific inhibitor BYK204165 (10 µM) (n = 3). Cells were fixed on days 1, 3, and 6 of differentiation. Upper panel shows days 1, 3, and 6 fusion index, lower panel shows day 6 fusion index. Each bar represents means ± S.D (n = 3) ***P < 0.001, **P < 0.01. E Representative photographs of Jenner-Giemsa stained differentiating myoblasts over days 1, 3, and 6 of differentiation (n = 3).
Reduction of PARP1 impacts the myoblast transcriptome
The muscle differentiation program is transcriptionally regulated by myogenic regulatory factors, some of which have been shown to be influenced by PARP1 [4]. However, the broader elements of PARP1 roles in the muscle transcriptome remain elusive. To provide insight, we conducted RNAseq on undifferentiated C2C12 myoblasts transfected with siRNA-mediated knockdown of PARP1 (siPARP1) (Fig. 6A–C). As PARP1 is a major NAD+ consuming enzyme, accounting for up to 90% of cellular PARylation [16], we examined by qPCR, impacts of siPARP1 on the expression of PARP2, PARG, NAMPT, and SIRT1, genes involved in NAD+ dependent homeostasis (Fig. 6D). No significant changes were observed in the expression of these genes. Additionally, RNAseq of siPARP1 myoblasts showed the glucocorticoid binding partner NR3C2 gene which codes for the mineralocorticoid receptor (MR), as being significantly downregulated (Fig. 6E and Supplementary Table S4). Gene Set Enrichment Analysis of the siPARP1 differentially expressed genes revealed that consistent with PARP1 established DNA damage repair response functions, the double-strand break repair process was the major deregulated process (Fig. 6E, F). Also suppressed, were pathways and metabolic processes regulating fundamental metabolism including pantothenate metabolism, palmitoylation, and mitochondrial import. Gene Set Enrichment Analysis also reported shifts in major pathways of hypoxia regulation (Normalized Enrichment Score (NES) 1.5, Pvalue 0.001), myogenesis (NES 1.3, P value 0.052), and TNF-α via NF-κB (NES −1.3, P value 0.039). PARP1 regulation of hypoxia, as is interaction with the NF-κB immune signaling cascades, has been documented [17, 18]. Moreover, these features are reported as being regulated by glucocorticoid signaling [19].
A qPCR of PARP1 in scrambled sequence controls (n = 5) and siRNA PARP1 transfected (siPARP1) C2C12 myoblasts (n = 4) ***P < 0.001. B Western immunoblotting of protein lysates collected from scrambled and siPARP1 C2C12 myoblasts (n = 5). C Quantification of scrambled and siPARP1 transfections by western blotting (n = 5). D qPCR of Scrambled control and siPARP1 cDNA for PARP1, PARP2/PARP2, PARG, NAMPT, and SIRT1 transcripts. Scrambled (n = 5) and siPARP1 (n = 4). E Representative volcano plot of differential gene expression following RNAseq of scrambled and siPARP1 C2C12 myoblasts. F Gene set enrichment analysis (GSEA) of pathways over and under-represented in RNAseq data of siPARP1 C2C12 myoblasts. G Enrichment plots of GSEA in siPARP1 C2C12 myoblasts for hypoxia, H myogenesis and, I TNFα via NF-κB.
Reduced PARP1 impacts the glucocorticoid transcriptional response
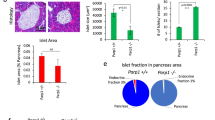
Finally, as we show PARylation as being a glucocorticoid-sensitive post-translational modification (Fig. 2D, E), as well as NR3C2 (MR) being downregulated in siPARP1 C2C12 myoblasts (Fig. 6E), and glucocorticoid signaling being a determinant of muscle turnover [19,20,21,22], we set out to test if PARP1 influences the glucocorticoid activated transcriptional program. To do this, we treated siPARP1 and scrambled transfected C2C12 myoblasts with dexamethasone for 24 hours. RNA was collected and sequenced for differential gene expression. We found that siPARP1 myoblasts retained the ability to upregulate genes in response to dexamethasone, sharing activation of typical glucocorticoid-regulated genes including SerpinA3N (9.50 fold change; 5.22 × 10−08 False Discovery Rate (FDR), HIF3α (7.40 fold change; 4.29 × 10−10 FDR) and Mt2 (3.97 fold change; 9.07 × 10−24 FDR). However, we observed that the expression of 86 glucocorticoid-induced genes was lost in dexamethasone-treated siPARP1 myoblasts (Fig. 7A, B and Supplementary Table S5). A tabular selection of these is presented for scrambled and siPARP1 myoblasts alongside classically glucocorticoid-activated genes (Fig. 7B), and the full list is provided (Supplementary Table S5). This cohort of genes, differentially regulated by dexamethasone in scrambled controls but not siPARP1 myoblasts, were subjected to functional profiling using g:Profiler [23]. This revealed enrichment for gene ontological processes and functions of significance to skeletal muscle physiology including calcium release (Padj. 1.49 × 10−2), as well as biological processes including straited muscle differentiation (Padj. 1.03 × 10−8) and cellular development (Padj. 3.34 × 10−6) (Fig. 7C). Furthermore, enrichment of the cellular compartment forming the thin filament of the myofibril was recorded (I band (FDR 5.06 × 10−6); Z disc (FDR 5.59 × 10−5)). Mapping of these 86 genes for transcription factor motifs generated enrichment for key regulatory factors of muscle including Pax4, a regulator of muscle protein turnover [24] (Padj. 4.67 × 10−3), LKLF (Padj. 5.72 × 10−3), PTF1 (Padj. 8.84 × 10−3) and myogenin (Padj. 8.98 × 10−3). Finally, human phenotype mapping identifies this gene cohort as being characteristic of skeletal muscle pathology including proximal muscle weakness, muscular dystrophy and Gower’s sign [25]. These data indicate there are a subset of genes regulated by glucocorticoids in skeletal muscle which are also dependent on PARP1.
A Venn diagram of genes positively regulated by glucocorticoids in Scrambled and siPARP1 C2C12 myoblasts (>1.5 Fold Change, <0.05 FDR) (n = 5 per treatment). B Selected list of genes responsive to dexamethasone and gene expression lost in siPARP1 C2C12 myoblasts. C Manhattan plot produced following g:GOSt analysis using the 86 genes whose response to dexamethasone is lost in siPARP1 C2C12 myoblasts. The plot shows overrepresented processes by Molecular function (red), Biological process (orange), Cellular compartment (green), Transcription factors (blue), and Human phenotype (purple).
Discussion
Herein, we demonstrate PARP1 and PARylation as being dynamically regulated during murine and human skeletal muscle differentiation. Specifically, we detail a conserved early-stage PAR accumulation that occurs within 24 h of myogenic induction that is both active and metabolically sensitive. We also show PARP1 may have roles in the cellular response to glucocorticoids, via regulation of a cohort of genes required for functional skeletal muscle.
PARP1 and PARylation are dynamic during skeletal muscle myogenesis
Suppression of PARP1 elicits beneficial metabolic effects which include enhanced exercise performance, increased energy expenditure, enhanced mitochondrial function, and resistance to oxidative stress [7,8,9, 26]. Additionally, PARP1 and PAR levels are dynamically and differentially regulated in response to exercise between aged untrained and aged trained muscle [27]. Our results establish PAR deposition by PARP1 as well as its auto-PARylation occurring within the first 24 h of myogenesis, supporting recent work showing PARP1-mediated PAR deposition following MYOD-driven transdifferentiation of fibroblasts into myoblasts [28]. This increased PAR accumulation could be ascribed to metabolic pathway shifts from glycolysis to oxidative phosphorylation as the major energy source during myogenesis, which has been reported in embryonic stem cells and induced pluripotent stem cells [29, 30]. Initiation of myogenesis upregulates the muscle-restricted gene NMRK2, and this switch increases NAD+ biosynthesis to drive PARP1 activity [31]. While the molecular recipients of the differentiating day 1 PAR accumulation are yet to be identified, existing evidence indicates MYOD is a likely target. Our data also show that this PAR accumulation is sensitive to changes in metabolism. Notably, we observed higher PAR levels in glucose-deprived conditions during myogenesis. However, PAR levels are induced in conditions of high glucose levels and fed states [7, 32]. These energetic requirements of day 1 differentiating myoblasts are likely specific to this point of myogenesis. This similarly has been found with autophagy and mitochondrial biogenesis which also upregulate PAR on day 1 [33, 34]. We also demonstrate that within myogenesis, that the troponins are also governed by PARP1 inhibition. It is therefore probable that PARP1, either through the direct or enzymatic application of PAR chains, coregulates their mRNA transcription. Incidentally, impacts over troponin expression by PARP1 hold relevance for cardiac muscle [35]. Given the nature of PARP1 PARylating activity, we also showed NAD+ precursor supplementation did not significantly impact PAR dynamics. This is possibly due to buffering mechanisms in response to NAD+ availability [36], the specific inhibition of PARP1 was successful in reducing PAR supported via observed increases in NAD+ (Supplementary Fig. 1A).
PARP1-mediated PARylation contributes to myogenesis
We postulated that PARP1-mediated PAR accumulation occurring on day 1 of myogenesis is to an extent, not dependent on myotube formation, but rather exerts consequences for the developed myotube. Using the broad-spectrum PARP inhibitor PJ34 and the PARP1-specific inhibitor BYK204165, we show that the expression profile changes are indeed related to PARP1-mediated PARylation on the first day of myogenesis. We also found downregulation of PARP1 alone in undifferentiated C2C12 myoblasts was sufficient to cause shifts in the muscle transcriptome, and subsequent biological processes comparable to those of early-stage PARP1 inhibition during myogenesis. Further insight into which substrates are PARylated within the first day of myogenesis where PAR disposition is at its highest will be useful in defining downstream signaling. In this regard, nuclear PARP1-mediated PARylating activities are increased in the myoblast following successful transdifferentiation from fibroblasts, providing merit for this [28]. It is, however, likely that site-specific PARylation of key myogenic regulators occurs, although this remains to be tested, once performed, it would reveal specific amino acid sequences serving to impact the myogenic protein activity and the continual process of muscle turnover. With regards to skeletal muscle, the myogenic transcriptional regulator Yin Yang1 (YY1) has been directly shown to be both a recipient of PARylation by PARP1 [37], and regulated by the immunomodulatory NF-kB pathway during myogenesis [38]. Crosstalk between PARP1 and NF-kB has been reported in several cell and tissue types [17, 39, 40]. Therefore, it is probable that a greater level of coordination between the NF-kB pathway and PARylation exists and remains to be explored in skeletal muscle. Another direct recipient of PARylation and regulator of myogenic progression is the transcription factor CCAAT/enhancer binding protein beta (C/EBPb), which has been demonstrated to be PARylated during adipogenesis [2, 3]. Given C/EBPb roles in satellite cells, the myocyte, and its interactions with MYOD [41,42,43], there is potential for C/EBPb PARylation in muscle tissue. Additional mechanisms of both PARP1 and catalytic PAR control are also not discounted and exert influence over 3D chromatin organization [44, 45]. However, it should be noted that PARP2 which also contributes to PAR accumulation, has demonstrated roles in myogenesis and skeletal muscle structure [46]. Collectively, our presented results suggest that both PARP1 and its PARylating activity hold influence over the skeletal muscle phenotype, and further underscores PARP1’s role as a multifaceted protein being able to bind to nucleic acids as well catalyzing the PARylation of target substrates [1, 47, 48].
PARP1 exerts influence over skeletal muscle glucocorticoid transcriptional response
Evidence for PARP1 in regulation of GR-mediated transcriptional response has been presented in other cells [49]. Furthermore, PARP1 null mice have increased cortisol levels [50]. Our transcriptomic analysis of siPARP1 C2C12 myoblasts reveals differentially expressed changes in the N3RC2 gene coding for MR, and while not as ubiquitous as its GR counterpart, MR can also bind glucocorticoids with higher affinities [19]. Moreover, we demonstrate pathways commonly classified as glucocorticoid controlled, including hypoxia, myogenesis, and TNF-α response through NF-κB, being shifted in siPARP1 myoblasts (Fig. 6g, h, i). Because PARP1 has documented roles in the NF-κB pathway for control of inflammatory response [17] and the mechanism of anti-inflammatory effects of glucocorticoids involves the repression of NF-κB, it is plausible that this glucocorticoid-mediated effect occurs via PARP1. Furthermore, NF-κB has also been implicated in differentiation programs of other tissues, including myogenesis [38, 39, 51]. In this regard, PARP1 and its PARylating activity exert influence over osteoclast differentiation and bone remodeling via NF-κB dependent transcription of IL-1β [39], and glucocorticoid-induced osteoporosis is a consequence of the inhibition of IL-1 production [52]. Furthermore, as with PARP1, the suppression of NF-κB is fundamental for driving myogenesis [38, 51], while dexamethasone treatment at the myoblast stage enhances myogenesis [20,21,22]. Concurrently, we demonstrate that application of dexamethasone during differentiation induction of myoblasts reduces PAR levels (Fig. 2D) indicating glucocorticoids have impacts on PARP1-mediated PARylating activity. Recent studies have demonstrated that the tumor susceptibility gene TSG101 binds to and enhances GR transcriptional activity [53] and similarly, PARP1 drives enzymatic activity for DNA damage-induced IKK-NF-κB activation [54]. Therefore, we speculate a similar paradigm in which glucocorticoids can regulate PARP1-mediated PARylation within the skeletal muscle via PARP1 interacting partners. Together with our findings, there is suggested the potential for PARP1 and GR interactions for influencing glucocorticoid-mediated outcomes and muscle turnover.
By examining the transcriptomic changes in siPARP1 C2C12 myoblasts treated with dexamethasone, we observe that the expression of glucocorticoid canonical target genes such as FKBP5 remained intact (Fig. 7B). We observed differential expression in a cohort of genes whose response following dexamethasone treatment was lost in siPARP1 myoblasts. These imply that PARP1 exerts influence over glucocorticoid transcriptional response in skeletal muscle. Furthermore, PARP1 has demonstrated involvement in myopathy [55], as does chronic glucocorticoid exposure [21, 56]. These indicate potential for PARP1 roles in the transcriptional pathways manifesting glucocorticoid-induced myopathy, necessitating broader studies. PARP1 mediates the glucocorticoid responsiveness of Gonadotropin-Releasing Hormone receptors [57], and here our findings demonstrate aspects of glucocorticoid-mediated impacts on skeletal muscle may occur through PARP1. However, it should also be appreciated that differential expression of glucocorticoid response genes could potentially be regulated by downstream events who themselves are impacted by siPARP1. Further investigation into direct GR and PARP1 interactions within the skeletal muscle would therefore establish clearer links into the extent PARP1 holds over overall glucocorticoid response.
Collectively, our studies demonstrate further potential for PARylation actions in skeletal muscle physiology and response to glucocorticoids (Fig. 8). While this might provide novel avenues for PARP inhibition in amelioration of chronic glucocorticoid-induced side effects, potential negative impacts over skeletal muscle turnover and repair should be considered.
PARP1 in myoblasts also influences transcriptional regulation of glucocorticoids. These actions highlight the importance of NAD+ availability and PARylation to skeletal muscle.
Materials and methods
Cell culture
The murine muscle myoblast cell line C2C12 was purchased (ATCC, VA, USA) and maintained in proliferation media, composed of Dulbecco’s Modified Eagle’s Medium (DMEM) 25 mM glucose (Lonza, UK) supplemented with 10% (v/v) fetal bovine serum (FBS) (Thermo, UK) and 1% Penicillin/Streptomycin (P/S) (Thermo, UK). Upon cells reaching 70–80% confluence, the differentiation medium, composed of DMEM 25 mM glucose supplemented with 2% horse serum (HS) (Thermo, UK) and 1% P/S, was added to induce differentiation. The myoblasts were allowed to differentiate for 6 days with fresh differentiation medium added every other day, sufficient for the successful development of mature myotubes. The human muscle myoblast cell line LHCN-M2 (Evercyte, Germany) was grown in proliferation media as described [58], composed of a 4:1 ratio of DMEM and Medium 199 (Sigma-Aldrich, UK) supplemented with 15% FBS, 200 mM HEPES (Thermo, UK), 0.03 µg/ml zinc sulfate (Sigma-Aldrich, UK), 1.4 µg/ml vitamin B12 (Sigma-Aldrich, UK), 0.055 µg/ml dexamethasone (Sigma-Aldrich, UK), 2.5 ng/ml hepatocyte growth factor (Proteintech, UK), 10 ng/ml basic fibroblast growth factor (Sigma-Aldrich, UK) and 1% P/S. The differentiation media of LHCN-M2 was composed of a 4:1 ratio of DMEM and Medium 199 supplemented with 2% HS and 1% P/S. LHCN-M2 differentiation was performed similarly to C2C12 myoblasts.
Cell treatments
Myoblasts were treated with either vehicle controls or the PARP inhibitors BYK204165 (Tocris, UK) and PJ34 (MedChemExpress, NJ, USA) at working concentrations of 10 µM, as well as Rucaparib (Selleck Chemical) treatment at a working concentration of 1 µM. Treatments with nicotinamide riboside (NR) (Chromadex, CA, USA) were performed at 0.5 mM, FK866 (Sigma-Aldrich, UK) at 50 nM, and dexamethasone at 1 µM. All treatments were diluted in the differentiation medium and performed during differentiation induction of myoblasts, after which treatments were washed out after 1 day (24 h) of differentiation and replaced with fresh differentiation medium, and myoblasts were left to differentiate as described.
Transfection of silencing RNA
Cells were seeded at an approximate density of 100,000 cells/well in six-well cell culture plates and incubated for at least 3 h to allow adherence. The siRNA transfection mix was composed of Lipofectamine 2000 (Thermo, UK) and 0.5 µg/µl of siRNA PARP1 predesigned from three pooled siRNA PARP1 sequences (Merck, UK) or scrambled siRNA control sequence (Merck, UK). Transfection was conducted in Opti-Mem serum and antibiotic-free media (Thermo, UK) and incubated for 24 h, after which the transfection mix was replaced with fresh medium.
RNA extraction and quantitative PCR
Cells were washed with 1x PBS and harvested in TRIzol Reagent (Life Technologies, UK). Total RNA was isolated using chloroform extraction and isopropanol precipitation. RNA clean-up and purification were conducted using a column purification kit (Zymo, Germany) according to instructions and quantity was measured by Nanodrop. Samples designated for RNAseq were further assessed for quality using the 2100 Bioanalyzer Instrument (Agilent Technologies, UK) with all samples having at least an RNA integrity number of >9.5.
For quantitative PCR (qPCR), reverse transcription was performed on extracted RNA using a High-Capacity cDNA Reverse Transcription Kit (Thermo, UK) according to the manufacturer’s instructions. cDNA was added to TaqMan™ Universal PCR Master Mix (Thermo, UK) with the appropriate primer pairs as follows: PARP1 forward: CCTGAACAACGCAGACAGC, PARP1 reverse: CGTTGTGCGTGGTAGCATGA, PARP2 forward: GGAAGGCGAGTGCTAAATGAA, PARP2 reverse: GGAAGGCGAGTGCTAAATGAA, NAMPT forward: CGCCATCTCCTTGAATGA, NAMPT reverse: GCACCACTAATCATCAGACC, PARG forward: GTGCCAGTTTCGATCCGTAGA, PARG reverse GGCCAGCATCGTGTAGATGA, SIRT1 forward: GGCTACCGAGACAACCTCCTG, SIRT1 reverse: AGTCCAGTCACTAGAGCTGGCG. Reactions were performed in 384 well plates and conducted using QuantStudio™ 7 Flex Real-Time PCR System (Applied Biosystems, UK) in single-plex format. All reactions were normalized to 18S rRNA (VICTM) (Applied Biosystems, UK). Data were collected as Ct values and used to obtain ΔΔCt values, and subsequently expressed as fold change ± standard error of the mean (S.D).
RNAseq
Library preparation and sequencing were conducted by Novogene Inc., UK, original fastq files can be found on EBI Array Express E-MTAB-12343. Reads were mapped using the kallisto RNAseq quantification program [59]. Analyses were carried out using R in RStudio and Bioconductor. Briefly, transcript quantification data were summarized to genes by tximport and normalized using edgeR. Normalized and filtered data were variance stabilized with voom function from limma and differentially expressed genes were identified with limma. Functional enrichment of differential expressed genes was conducted using g:Profiler [60].
Cell lysis and western blotting
For whole-cell lysis, cells were first washed in ice-cold 1x PBS and subsequently scraped in RIPA buffer (50 mM Tris pH 8.0, 150 mM NaCl, 0.5% (w/v) sodium deoxycholate, 0.1% (w/v) SDS, 1% NP-40) supplemented with 1× Pierce™ EDTA-free protease inhibitor cocktail (Thermo, UK) and incubated on ice for 30 min. Lysates were clarified by centrifugation at 12,000 rpm for 15 min at 4oC. The supernatant was recovered and stored at -80oC until use. Total protein concentration was determined by detergent-compatible protein assay (Bio-Rad, UK) according to the kit’s instructions.
Total proteins were loaded and resolved on fixed percentage acrylamide SDS-PAGE gels and subsequently transferred onto PVDF membranes using Trans-Blot Turbo Transfer System (Bio-Rad, UK). Membranes were blocked with 5% bovine serum albumin (BSA) diluted in 1x TBS-T and incubated with primary antibodies overnight at 4oC: PARP1 (39559, Active-Motif, Belgium), PAR (MABE1031, Merck Millipore, UK), MYOD (sc-377460, Santa Cruz Biotechnology, Inc., USA), Hexokinase II (ab227198, Abcam, UK), Myogenin (ab1835, Abcam, UK), NF-kB (8242, Cell Signalling Technology, UK), α-Tubulin (sc-5286, Santa Cruz Biotechnology, Inc., USA), and subsequently with HRP-conjugated anti-mouse or anti-rabbit secondary antibodies (Dako, Denmark) for 1 h at room temperature. Blots were developed with Pierce™ ECL Western Blotting Substrate (Thermo, UK) and visualized using the G:BOX Chemi XX6 system (Syngene, UK). Bands were measured using Image J densitometry and normalized to those of loading controls.
Immunofluorescence
Cells grown on coverslips were rinsed with 1× PBS and fixed with 4% PFA (Thermo, UK). Permeabilization was performed with 0.1% Triton X-100 (Sigma-Aldrich, UK) and blocked with 10% goat serum (Life Technologies, UK). Coverslips were incubated with primary antibodies overnight at 4 oC, before incubation with secondary antibodies and DAPI nuclear dye (Thermo, UK) for 1 h at room temperature. Coverslips were mounted on slides and allowed to set prior to imaging.
Unbiased proteomics
Unbiased mass spectrometry was carried out on a SCIEX TripleTOF 6600 instrument with samples analyzed in both SWATH (Data Independent Acquisition) and IDA (Information Dependent Acquisition) modes for quantitation and spectral library generation respectively. IDA data were searched together using ProteinPilot 5.0.2 to generate a spectral library and SWATH data was analyzed using Sciex OneOmics software extracted against the locally generated library. Log fold change and statistical analysis were calculated as described [60].
NAD+ measurement
NAD+ was measured in lysates extracted in 0.6 N perchloric acid as described [61]. Briefly, standards or samples in phosphate buffer were combined with the cycling mixture composed of 2% ethanol, 100 µg/ml alcohol dehydrogenase, 10 µg/ml diaphorase, 20 µM resazurin, 10 µM flavin mononucleotide, 10 mM nicotinamide and 0.1% BSA, in 100 mM phosphate buffer pH 8.0. Resorufin accumulation was evaluated by reading excitation at 544 nm and emission at 590 nm.
Giemsa–Jenner staining of myotubes
Cells stained during differentiation time points were performed as described [15]. Briefly, cells were fixed in methanol and stained with Jenner (Alfa Aesar, UK) diluted in 1 mM sodium phosphate pH 5.6, followed by Giemsa (Alfa Aesar, UK) diluted in 1 mM sodium phosphate pH 5.6. Cells were imaged using a camera attached to an inverted microscope. Protein-rich myotube fibers presented themselves as dark/deep purple while nuclei-stained shallow purple/pink. Total number of nuclei per image was manually scored and the number of nuclei inside myotubes was expressed as a percentage of total nuclei to obtain the fusion index.
Statistical analysis
Students t test or ANOVA statistical comparisons were used with the Graphpad Software Inc. Prism version 9. Western blots were quantified by densitometry using imageJ. Data are presented as mean ± S.D with statistical significance determined as *P < 0.05, **P < 0.01, ***P < 0.001. Differences between two groups were determined using unpaired t-test compared treatments or genotypes. Statistical analysis derived from qPCR data was determined using ΔΔCt values throughout.
Data availability
The datasets generated for this study are available on request to the corresponding author. RNAseq files are available at Array Express.
Code availability
Code generated using RStudio version 4.2 and is available upon request.
References
Gibson BA, Kraus WL. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol. 2012;13:411–24.
Luo X, Ryu KW, Kim DS, Nandu T, Medina CJ, Gupte R, et al. PARP-1 controls the adipogenic transcriptional program by PARylating C/EBPβ and modulating its transcriptional activity. Mol Cell [Internet] 2017;65:260–71. https://pubmed.ncbi.nlm.nih.gov/28107648/. Accessed 1 Jul 2022.
Ryu KW, Nandu T, Kim J, Challa S, DeBerardinis RJ, Lee, et al. Metabolic regulation of transcription through compartmentalized NAD+ biosynthesis. Science 2018;360:eaan5780.
Matteini F, Andresini O, Petrai S, Battistelli C, Rossi MN, Maione R. Poly(ADP-ribose) Polymerase 1 (PARP1) restrains MyoD-dependent gene expression during muscle differentiation. Sci Rep. [Internet]. 2020;10:15086. Accessed 29 Jun 2022.
Wang Y, Zhang Y, Zhang S, Kim B, Hull VL, Xu J, et al. PARP1-mediated PARylation activity is essential for oligodendroglial differentiation and CNS myelination. Cell Rep. 2021;37:109695.
Jones A, Kraus WL. Multiomics analysis of the NAD +-PARP1 axis reveals a role for site-specific ADP-ribosylation in splicing in embryonic stem cells. Genes Dev. 2022;36:601–17.
Bai P, Cantó C, Oudart H, Brunyánszki A, Cen Y, Thomas C, et al. PARP-1 inhibition increases mitochondrial metabolism through SIRT1 activation. Cell Metab. 2011;13:461–8.
Pirinen E, Cantó C, Jo YS, Morato L, Zhang H, Menzies KJ, et al. Pharmacological Inhibition of poly(ADP-ribose) polymerases improves fitness and mitochondrial function in skeletal muscle. Cell Metab. 2014;19:1034–41.
Oláh G, Szczesny B, Brunyánszki A, Lopez-García IA, Gerö D, Radák Z, et al. Differentiation-associated downregulation of poly(ADP-Ribose) polymerase-1 expression in myoblasts serves to increase their resistance to oxidative stress. PLoS One [Internet] 2015;10:e0134227. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0134227. Accessed 29 Jun 2022.
Elhassan YS, Kluckova K, Fletcher RS, Schmidt MS, Garten A, Doig CL, et al. Nicotinamide riboside augments the aged human skeletal muscle NAD+ metabolome and induces transcriptomic and anti-inflammatory signatures. Cell Rep. 2019;28:1717–28.e6.
Eleazer R, Fondufe-Mittendorf YN. The multifaceted role of PARP1 in RNA biogenesis. Wiley Interdiscip Rev RNA [Internet]. 2021;12:e1617. https://pubmed.ncbi.nlm.nih.gov/32656996/. Accessed 11 Nov 2022.
Jubin T, Kadam A, Gani AR, Singh M, Dwivedi M, Begum R. Poly ADP-ribose polymerase-1: beyond transcription and towards differentiation. Semin Cell Dev Biol [Internet]. 2017;63:167–79. https://pubmed.ncbi.nlm.nih.gov/27476447/. Accessed 30 Sep 2022.
Wahlberg E, Karlberg T, Kouznetsova E, Markova N, Macchiarulo A, Thorsell AG, et al. Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat Biotechnol [Internet]. 2012;30:283–8. https://pubmed.ncbi.nlm.nih.gov/22343925/. Accessed 29 Jun 2022.
Eltze T, Boer R, Wagner T, Weinbrenner S, McDonald MC, Thiemermann C, et al. Imidazoquinolinone, imidazopyridine, and isoquinolindione derivatives as novel and potent inhibitors of the poly(ADP-ribose) polymerase (PARP): a comparison with standard PARP inhibitors. Mol Pharm [Internet]. 2008;74:1587–98. https://pubmed.ncbi.nlm.nih.gov/18809672/. Accessed 29 Jun 2022.
Veliça P, Bunce CM. A quick, simple and unbiased method to quantify C2C12 myogenic differentiation. Muscle Nerve [Internet]. 2011;44:366–70. https://onlinelibrary.wiley.com/doi/full/10.1002/mus.22056. Accessed 25 Oct 2022.
Kamaletdinova T, Fanaei-Kahrani Z, Wang ZQ. The enigmatic function of PARP1: from PARylation activity to PAR readers. Cells [Internet]. 2019;8:1625. Accessed 5 Oct 2022.
Oliver FJ, Ménissier-de Murcia J, Nacci C, Decker P, Andriantsitohaina R, Muller S, et al. Resistance to endotoxic shock as a consequence of defective NF-κB activation in poly (ADP-ribose) polymerase-1 deficient mice. EMBO J. 1999;18:4446–54.
Martí JM, Garcia-Diaz A, Delgado-Bellido D, O’Valle F, González-Flores A, Carlevaris O, et al. Selective modulation by PARP-1 of HIF-1α-recruitment to chromatin during hypoxia is required for tumor adaptation to hypoxic conditions. Redox Biol. 2021;41:101885.
Vettorazzi S, Nalbantoglu D, Gebhardt JCM, Tuckermann J. A guide to changing paradigms of glucocorticoid receptor function—a model system for genome regulation and physiology. FEBS J [Internet]. 2022;289:5718–43. https://onlinelibrary.wiley.com/doi/full/10.1111/febs.16100. Accessed 30 Sept 2022.
Guerriero V, Florini JR. Dexamethasone effects on myoblast proliferation and differentiation. Endocrinol [Internet]. 1980;106:1198–202. https://pubmed.ncbi.nlm.nih.gov/7188899/. Accessed 29 Jun 2022.
Han DS, Yang WS, Kao TW. Dexamethasone treatment at the myoblast stage enhanced C2C12 myocyte differentiation. Int J Med Sci [Internet]. 2017;14:434–43. https://pubmed.ncbi.nlm.nih.gov/28539819/. Accessed 29 Jun 2022.
Lin JW, Huang YM, Chen YQ, Chuang TY, Lan TY, Liu YW. et al. Dexamethasone accelerates muscle regeneration by modulating kinesin-1-mediated focal adhesion signals. Cell Death Discov. 2021;7:1–16. Accessed 29 Jun 2022.
Raudvere U, Kolberg L, Kuzmin I, Arak T, Adler P, Peterson H, et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res [Internet]. 2019;47:W191–8. https://academic.oup.com/nar/article/47/W1/W191/5486750. Accessed 28 Oct 2022.
Volodin A, Kosti I, Goldberg AL, Cohen S. Myofibril breakdown during atrophy is a delayed response requiring the transcription factor PAX4 and desmin depolymerization. Proc Natl Acad Sci USA [Internet]. 2017;114:E1375–84. https://www.pnas.org/doi/abs/10.1073/pnas.1612988114. Accessed 7 Oct 2022.
Caria F, Cescon M, Gualandi F, Pichiecchio A, Rossi R, Rimessi P, et al. Autosomal recessive Bethlem myopathy: a clinical, genetic and functional study. Neuromuscul Disord [Internet]. 2019;29:657–63. https://pubmed.ncbi.nlm.nih.gov/31471117/. Accessed 7 Oct 2022.
Mohamed JS, Wilson JC, Myers MJ, Sisson KJ, Alway SE. Dysregulation of SIRT-1 in aging mice increases skeletal muscle fatigue by a PARP-1-dependent mechanism. Aging [Internet]. 2014;6:820–34. https://pubmed.ncbi.nlm.nih.gov/25361036/. Accessed 17 Sept 2021.
Cobley JN, Sakellariou GK, Murray S, Waldron S, Gregson W, Burniston JG, et al. Lifelong endurance training attenuates age-related genotoxic stress in human skeletal muscle. Longev Healthspan [Internet]. 2013;2:11 https://pubmed.ncbi.nlm.nih.gov/24472304/. Accessed 17 Sept 2021.
Bisceglie L, Hopp AK, Gunasekera K, Wright RH, le Dily F, Vidal E, et al. MyoD induces ARTD1 and nucleoplasmic poly-ADP-ribosylation during fibroblast to myoblast transdifferentiation. iScience. 2021;24:102432.
Chung S, Dzeja PP, Faustino RS, Perez-Terzic C, Behfar A, Terzic A. Mitochondrial oxidative metabolism is required for the cardiac differentiation of stem cells. Nat Clin Pract Cardiovasc Med [Internet]. 2007;4:S60–7. https://pubmed.ncbi.nlm.nih.gov/17230217/. Accessed 6 Jan 2023.
Folmes CDL, Nelson TJ, Martinez-Fernandez A, Arrell DK, Lindor JZ, Dzeja PP, et al. Somatic oxidative bioenergetics transitions into pluripotency-dependent glycolysis to facilitate nuclear reprogramming. Cell Metab [Internet]. 2011;14:264–71. https://pubmed.ncbi.nlm.nih.gov/21803296/. Accessed 6 Jan 2023.
Fletcher RS, Ratajczak J, Doig CL, Oakey LA, Callingham R, da Silva Xavier G, et al. Nicotinamide riboside kinases display redundancy in mediating nicotinamide mononucleotide and nicotinamide riboside metabolism in skeletal muscle cells. Mol Metab. 2017;6:819–32.
Garcia Soriano F, Virág L, Jagtap P, Szabó É, Mabley JG, Liaudet L, et al. Diabetic endothelial dysfunction: the role of poly(ADP-ribose) polymerase activation. Nat Med [Internet]. 2001;7:108–13. https://pubmed.ncbi.nlm.nih.gov/11135624/. Accessed 5 Jan 2023.
Rodríguez-Vargas JM, Oliver-Pozo FJ, Dantzer F. PARP1 and poly(ADP-ribosyl)ation signaling during autophagy in response to nutrient deprivation. Oxid Med Cell Longev [Internet]. 2019;2019:2641712. https://pubmed.ncbi.nlm.nih.gov/31281570/. Accessed 15 Feb 2023.
Sin J, Andres AM, Taylo RDJR, Weston T, Hiraumi Y, Stotland A, et al. Mitophagy is required for mitochondrial biogenesis and myogenic differentiation of C2C12 myoblasts. Autophagy [Internet]. 2016;12:369–80. https://pubmed.ncbi.nlm.nih.gov/26566717/. Accessed 15 Feb 2023.
Zhang D, Hu X, Li J, Liu J, Baks-te Bulte L, Wiersma M, et al. DNA damage-induced PARP1 activation confers cardiomyocyte dysfunction through NAD+ depletion in experimental atrial fibrillation. Nat Commun [Internet]. 2019;10. https://pubmed.ncbi.nlm.nih.gov/30898999/. Accessed 7 Oct 2022.
Oakey LA, Fletcher RS, Elhassan YS, Cartwright DM, Doig CL, Garten A, et al. Metabolic tracing reveals novel adaptations to skeletal muscle cell energy production pathways in response to NAD + depletion. Wellcome Open Res. 2019;3:147.
Oei SL, Shi Y. Poly(ADP-Ribosyl)ation of transcription factor Yin Yang 1 under conditions of DNA damage. Biochem Biophys Res Commun. 2001;285:27–31.
Wang H, Hertlein E, Bakkar N, Sun H, Acharyya S, Wang J, et al. NF-kappaB regulation of YY1 inhibits skeletal myogenesis through transcriptional silencing of myofibrillar genes. Mol Cell Biol [Internet] 2007;27:4374–87. https://pubmed.ncbi.nlm.nih.gov/17438126/. Accessed 7 Oct 2022.
Robaszkiewicz A, Qu C, Wisnik E, Ploszaj T, Mirsaidi A, Kunze FA, et al. ARTD1 regulates osteoclastogenesis and bone homeostasis by dampening NF-κB-dependent transcription of IL-1β. Sci Rep [Internet]. 2016;6. Accessed 29 Jun 2022.
Ke B, Li A, Fu H, Kong C, Liu T, Zhu Q, et al. PARP-1 inhibitors enhance the chemosensitivity of leukemia cells by attenuating NF-кB pathway activity and DNA damage response induced by Idarubicin. Acta Biochim Biophys Sin [Internet]. 2022;54:91–8. https://pubmed.ncbi.nlm.nih.gov/35130631/. Accessed 7 Oct 2022.
Marchildon F, Fu D, Lala-Tabbert N, Wiper-Bergeron N. CCAAT/enhancer binding protein beta protects muscle satellite cells from apoptosis after injury and in cancer cachexia. Cell Death Dis 2016 7:2 [Internet]. 2016;7:e2109. https://www.nature.com/articles/cddis20164. Accessed 7 Oct 2022.
Marchildon F, Lala N, Li G, St.-Louis C, Lamothe D, Keller C, et al. CCAAT/enhancer binding protein beta is expressed in satellite cells and controls myogenesis. Stem Cells [Internet]. 2012;30:2619–30. https://academic.oup.com/stmcls/article/30/12/2619/6404629. Accessed 7 Oct 2022.
Lala-Tabbert N, AlSudais H, Marchildon F, Fu D, Wiper-Bergeron N. CCAAT/enhancer binding protein β is required for satellite cell self-renewal. Skelet Muscle [Internet]. 2016;6:1–11. https://skeletalmusclejournal.biomedcentral.com/articles/10.1186/s13395-016-0112-8. Accessed 7 Oct 2022.
Muthurajan UM, Hepler MRD, Hieb AR, Clark NJ, Kramer M, Yao T, et al. Automodification switches PARP-1 function from chromatin architectural protein to histone chaperone. Proc Natl Acad Sci USA [Internet]. 2014;111:12752–7. Accessed 7 Oct 2022.
Posavec Marjanović M, Crawford K, Ahel I. PARP, transcription and chromatin modeling. Semin Cell Dev Biol. 2017;63:102–13.
Jankó L, Sári Z, Kovács T, Kis G, Szántó M, Antal M, et al. Silencing of PARP2 blocks autophagic degradation. cells [Internet]. 2020;9. https://pubmed.ncbi.nlm.nih.gov/32046043/. Accessed 5 Jan 2023.
Gupte R, Liu Z, Kraus WL. PARPs and ADP-ribosylation: recent advances linking molecular functions to biological outcomes. Genes Dev. 2017;31:101–26.
Kim DS, Challa S, Jones A, Kraus WL. PARPs and ADP-ribosylation in RNA biology: from RNA expression and processing to protein translation and proteostasis. Genes Dev. 2020;34:302–20.
Trotter KW, King HA, Archer TK. Glucocorticoid receptor transcriptional activation via the BRG1-dependent recruitment of TOP2β and Ku70/86. Mol Cell Biol. 2015;35:2799–817.
Drazen DL, Bilu D, Edwards N, Nelson RJ. Disruption of poly(ADP-ribose) polymerase (PARP) protects against stress-evoked immunocompromise. Mol Med. 2001;7:761–6.
Guttridge DC, Albanese C, Reuther JY, Pestell RG, Baldwin AS Jr. NF-κB controls cell growth and differentiation through transcriptional regulation of cyclin D1. Mol Cell Biol [Internet]. 1999;19:5785. Accessed 7 Oct 2022.
Ilias I, Milionis C, Zoumakis E. An overview of glucocorticoid-induced osteoporosis. J Clin Rheumatol [Internet]. 2022;5. https://www.ncbi.nlm.nih.gov/books/NBK278968/. Accessed 7 Oct 2022.
White JT, Rives J, Tharp ME, Wrabl JO, Thompson EB, Hilser VJ. Tumor susceptibility gene 101 regulates the glucocorticoid receptor through disorder-mediated allostery. Biochem [Internet]. 2021;60:1647–57. https://pubmed.ncbi.nlm.nih.gov/34009973/. Accessed 6 Jan 2023.
Tufan AB, Lazarow K, Kolesnichenko M, Sporbert A, von Kries JP, Scheidereit C. TSG101 associates with PARP1 and is essential for PARylation and DNA damage-induced NF-κB activation. EMBO J [Internet]. 2022;41:e110372. https://onlinelibrary.wiley.com/doi/full/10.15252/embj.2021110372. Accessed 6 Jan 2023.
Sharma V, Nand Pandey S, Khawaja H, Brown JK, Hathout Y, Chen YW. PARP1 differentially interacts with promoter region of DUX4 Gene in FSHD myoblasts. J Genet Syndr Gene Ther. 2016;7:303.
Schakman O, Kalista S, Barbé C, Loumaye A, Thissen JP. Glucocorticoid-induced skeletal muscle atrophy. Int J Biochem Cell Biol [Internet]. 2013;45:2163–72. https://pubmed.ncbi.nlm.nih.gov/23806868/. Accessed 29 Jun 2022.
Lee C, Cederberg RA, White BR. Poly [ADP-Ribose] polymerase-1 (PARP-1) confers glucocorticoid responsiveness of the porcine GnRH receptor (GnRHR) Gene. Biol Reprod. 2011;85Suppl_1:2–4.
Zhu CH, Mouly V, Cooper RN, Mamchaoui K, Bigot A, Shay JW, et al. Cellular senescence in human myoblasts is overcome by human telomerase reverse transcriptase and cyclin-dependent kinase 4: consequences in aging muscle and therapeutic strategies for muscular dystrophies. Aging Cell. 2007;6:515–23.
Bray NL, Pimentel H, Melsted P, Pachter L. Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol. 2016;34:525–7. https://www.nature.com/articles/nbt.3519. Accessed 5 Oct 2022.
Aguilan JT, Kulej K, Sidoli S. Guide for protein fold change and p-value calculation for non-experts in proteomics. Mol Omics. 2020;16:573–82.
Luongo TS, Eller JM, Lu MJ, Niere M, Raith F, Perry C, et al. SLC25A51 is a mammalian mitochondrial NAD+ transporter. Nat [Internet]. 2020;588:174. Accessed 18 Oct 2022.
Acknowledgements
We would like to thank Prof. Mark Christian of Nottingham Trent University for helpful insight. We would also like to thank Chromadex for the supply of nicotinamide riboside.
Funding
This work was part funded by The Physiological Society Research grant and Bioscientifica COVID-19 relief fund awarded to CLD. CLD is also funded by a Quality research award from Nottingham Trent University.
Author information
Authors and Affiliations
Contributions
AT and CLD performed study concept and design; AT, AE, JC, CLD, AC, DB, and CC performed experimental work. AT and CLD performed development of methodology and writing, edits and revision of the paper; CLD, DB, CC and AT provided acquisition, analysis and interpretation of data, and statistical analysis; GGL and CS provided technical and material support. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tan, A., Younis, A.Z., Evans, A. et al. PARP1 mediated PARylation contributes to myogenic progression and glucocorticoid transcriptional response. Cell Death Discov. 9, 133 (2023). https://doi.org/10.1038/s41420-023-01420-2
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41420-023-01420-2