Abstract
The trillions of microorganisms inhabiting the human gut are intricately linked to human health. While specific microbes have been associated with diseases, microbial abundance alone cannot reveal the molecular mechanisms involved. One such important mechanism is the biosynthesis of functional metabolites. Here, we develop a biosynthetic enzyme-guided disease correlation approach to uncover microbial functional metabolites linked to disease. Applying this approach, we negatively correlate the expression of gut microbial sulfonolipid (SoL) biosynthetic enzymes to inflammatory bowel disease (IBD). Targeted chemoinformatics and metabolomics then confirm that SoL abundance is significantly decreased in IBD patient data and samples. In a mouse model of IBD, we further validate that SoL abundance is decreased while inflammation is increased in diseased mice. We show that SoLs consistently contribute to the immunoregulatory activity of different SoL-producing human microbes. We further reveal that sulfobacins A and B, representative SoLs, act on Toll-like receptor 4 (TLR4) and block lipopolysaccharide (LPS) binding, suppressing both LPS-induced inflammation and macrophage M1 polarization. Together, these results suggest that SoLs mediate a protective effect against IBD through TLR4 signaling and showcase a widely applicable biosynthetic enzyme-guided disease correlation approach to directly link the biosynthesis of gut microbial functional metabolites to human health.
Similar content being viewed by others
Introduction
The human gut microbiome, composed of trillions of microorganisms, is intricately linked to human health1. At the species abundance level, numerous human microbes have been rigorously correlated with disease phenotypes, however, the mechanisms by which these microbes influence human health remain largely unkown2,3,4. One important mechanism is through the biosynthesis of functional metabolites by microbes5,6,7,8,9. Microbial functional metabolites are in direct contact and constant exchange with human cells, granting them inherent biological activity in complex host-microbe interactions5,6,8. Accordingly, human microbiome research has begun moving towards revealing these microbial functional metabolites and their corresponding molecular mechanisms that drive specific disease phenotypes10,11.
Human microbiota-derived lipids are a prolific class of functional metabolites. While many studies have focused on common microbe-derived lipids, such as short-chain fatty acids (SCFAs) and phospholipids12,13, there are a significant number of underexplored lipids which may be equally capable of influencing human health14. One such class of microbe-derived lipids is sulfonolipids (SoLs), unique molecules that bear striking structural similarity to both bacterial and endogenous sphingolipids (SLs) which are known for their role in mediating immune signaling in humans15. SoL-producing bacteria do not biosynthesize SLs but instead produce SoLs in high abundance16,17, suggesting that SoLs may replace SLs as functional metabolites with similar but distinctly different functions. In fact, two SoL-producing genera, Alistipes and Odoribacter, have been negatively correlated with the two primary forms of IBD, ulcerative colitis (UC) and Crohn’s disease (CD)18,19,20,21, with some species shown to ameliorate IBD symptoms20,21. We have found that sulfobacin A (SoL A), a representative SoL produced by Chryseobacterium gleum F93 DSM 16776, exhibits unusual immunoregulatory activity in vitro by modulating inflammatory cytokine production, especially through suppression of the lipopolysaccharide (LPS)-induced inflammatory response22 which has been reported as a key contributor to the progression of IBD23,24,25,26. We have also elucidated the biosynthetic pathway of SoLs and shown that this pathway involves critical enzymes that are specifically involved in SoL biosynthesis, allowing it to be distinguished from the SL biosynthetic pathway22. Whether SoLs produced by Alistipes and Odoribacter, two genera negatively associated with IBD18,19, represent functional metabolites in this negative correlation is unknown. Furthermore, the molecular target(s) of SoLs as a whole class of unique and abundant lipids is also unknown.
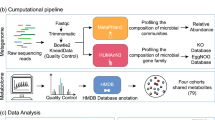
To probe the associations between microbial functional metabolites and disease, many studies have used untargeted metabolomics. However, this approach alone faces challenges such as the complexity of the metabolome, the lack of reference databases for identification, trace metabolite amounts, and a high degree of variability between different metabolomes27,28,29, all of which lead to difficulty in revealing specific gut microbial functional metabolites as drivers of molecular mechanisms in disease. In contrast, disease-related sequencing datasets such as the Human Microbiome Project30, DIABIMMUNE31, and numerous others hosted in the NCBI Sequence Read Archive32 are widely available, higher quality, less dimensional, and less variable. This presents an alternative approach to elucidating the potential connection between SoLs and disease. Thus, we developed an approach that leverages the critical SoL biosynthetic enzymes to examine their differential prevalence and expression in disease, allowing genomic and transcriptomic-level correlations between the biosynthesis of SoLs by human microbes and disease. Informed by this biosynthetic enzyme-guided disease correlation, we can further examine the abundance of SoLs in human disease-related metabolomics datasets to validate these genomic and transcriptomic-level associations at the metabolomic level.
In this work, we used this informatics-based, functional metabolites-focused approach to reveal a negative correlation between SoL biosynthesis and human IBD pathogenesis, followed by targeted chemoinformatic and metabolomic analysis. We then experimentally validated this multi-level correlation using a mouse model of IBD. Through bioactive molecular networking, we determined that SoLs consistently contribute to the immunoregulatory activity of SoL-producing human gut commensals. Using cell-based assays, we also revealed that SoLs primarily mediate their immunomodulatory activity through interaction with TLR4. Specifically, SoLs binds directly to TLR4 via the accessory protein myeloid differentiation factor 2 (MD-2) and displace LPS from MD-2 at higher concentrations, leading to suppression of TLR4 signaling pathways and macrophage M1 polarization. Together, these results demonstrate our biosynthetic enzyme-guided disease correlation approach to uncovering the chemical basis and molecular mechanisms of intricate host-microbe interactions and outline a potential mechanism by which gut microbial SoLs exert a protective effect against IBD progression through disruption of LPS-mediated TLR4 signaling.
Results
Biosynthetic enzyme-guided disease correlation analysis reveals a negative correlation between SoL biosynthesis and IBD
We began by systematically investigating the biosynthetic potential of SoLs from 285,835 human gut bacterial reference genomes including single amplified genomes (SAGs) and metagenome-assembled genomes (MAGs)33. Based on sequence homology with experimentally verified SoL biosynthetic enzymes22,34,35,36 (Supplementary Fig. 1, Supplementary Data 1), we identified a total of 562,214 homologous enzyme sequences, including 469,012 cysteate synthases (CYS), 33,486 cysteate fatty acyltransferases (CFAT), and 59,716 short-chain dehydrogenases/reductases (SDR) (Supplementary Fig. 2a). Uncovering phylogenetic trends, we found that these three enzymes were widely distributed in 255,572 genomes (Supplementary Fig. 2a) across 21 phyla, with the majority belonging to Bacteroidota and Firmicutes_A (Supplementary Fig. 2b). A subset of 6.21% (15,863/255,572) of the genomes was found to encode all three putative SoL biosynthetic enzymes (Supplementary Fig. 2a). To prioritize them for further analysis, we filtered the homologs on the basis of three rules: (1) the homology of both CFAT and CYS must equal or exceed 50% sequence similarity with experimentally validated CFATs and CYSs (Supplementary Data 1), as these enzymes are the first two specific enzymes in the biosynthetic pathway of SoLs that distinguish the biosynthesis between SoLs and SLs22,34,35,36; (2) the homologous regions of CYS, CFAT, and SDR should include protein domains with Pfam IDs PF00291, PF00155, and PF00106, respectively (hit score >50); (3) a set of homologous enzymes, especially SDR enzymes that show variable sequence similarities, should come from the same genome encoding all three enzymes as all three are required for SoL biosynthesis, thus ensuring co-occurrence. Applying these rules, we prioritized 9,731 CYS (1384 unique sequences), 9740 CFAT (917 unique sequences), and 10,319 SDR enzymes (1,076 unique sequences) (Fig. 1a, Supplementary Data 2) from 9,633 bacterial genomes. The prioritized enzymes were distributed among 42 species from Bacteroidota (99.99%, 9632/9633, 95% confidence interval: 99.94% ~ 100%) and one species from Firmicutes_A (0.01%, 1/9633, 95% confidence interval: 0.0018% ~ 0.059%) (Fig. 1b, Supplementary Data 3). Of note, among the 42 species from Bacteroidota, 71% (30/42) of them belong to bacterial families that have been previously reported to produce SoLs (Fig. 1b) including Rikenellaceae (containing genera Alistipes and Alistipes_A)16,37, Marinifilaceae (containing genus Odoribacter)16, and Weeksellaceae (containing genus Chryseobacterium B)22,38.
a Overview of SoL biosynthetic enzymes identified in human gut bacteria. 562,214 putative SoL biosynthetic enzymes were identified across 21 bacterial phyla. 6.21% of genomes encode 3 types of SoL biosynthetic enzymes (Pie chart, sections in red and purple). Bar chart shows the number of prioritized SoL biosynthetic enzymes encoded by 9,633 genomes (highlighted in red in the pie chart). b A circular phylogenetic tree shows the prioritized SoL biosynthetic enzymes found primarily in species from Bacteroidota (highlighted in green and orange). The tree is annotated with species names and colored by taxonomic families (Rikenellaceae: green; Marinifilaceae: orange; Weeksellaceae: pink; Lachnospiraceae: gray). c Principal Coordinate Analysis (PCoA) shows differences in the presence profile of overall SoL biosynthetic enzyme subfamilies between IBD and non-IBD groups based on Jaccard distance. Statistical significance was determined using PERMANOVA; p = 0.001. d 35 SoL biosynthetic enzyme subfamilies were significantly more prevalent in healthy individuals (red dots) than in IBD groups (blue dots) with a difference of prevalence >10%. All comparisons were significant by two-sided Fisher’s exact test with p < 0.05. e PCoA shows the differences in the expression profile of overall SoL biosynthetic enzyme subfamilies between IBD and non-IBD groups based on Bray-Curtis distances. Statistical significance was determined using PERMANOVA; p = 0.001. f Expression profiles of differential SoL biosynthetic enzyme subfamilies (n = 8; two-sided Mann-Whitney U test, adjusted p < 0.05). Upper panel: bar charts showing the prevalence of differential SoL biosynthetic enzyme subfamilies across non-IBD (red) and IBD individuals (dark blue). Statistical significance for prevalence was calculated using a two-sided Fisher’s exact test. Except CYS subfamily24 (CYS_24, no significance), all were significantly higher in prevalence in non-IBD than IBD groups (p < 0.05). Lower panel: box plots displaying the abundance profiles of differential SoL biosynthetic enzyme subfamilies in non-IBD (red) and IBD individuals (dark blue). All box plots include center lines representing the median, box limits representing upper and lower quartiles, whiskers representing the 1.5x interquartile range, and points representing outliers. Significance was further determined by one-sided Mann-Whitney U test, with adjusted p-value < 0.05.
To determine whether there is a link between gut microbial capacity to produce SoLs and IBD incidence, we conducted a comparative analysis of metagenomic and metatranscriptomic data obtained from the Inflammatory Bowel Disease Multi’omics Database (IBDMDB)19,39. We began by generating sequence similarity networks with a 90% sequence identity threshold to group enzymes with similar functions. Consequently, we categorized the prioritized biosynthetic enzymes into 214 subfamilies (79 CYS subfamilies; 25 CFAT subfamilies; and 110 SDR subfamilies) for the subsequent analyses (Supplementary Data 4). Looking for the presence of the prioritized 214 subfamilies in IBD cohorts, we identified 154 subfamilies in 667 metagenome samples (182 healthy samples and 485 IBD disease samples), of which 116 subfamilies were detected in ≥5% of samples (Supplementary Fig. 2c). Beta diversity of the presence of these 116 subfamily biosynthetic enzymes indicated that the overall composition of SoL biosynthetic enzyme subfamilies was significantly different between the healthy and IBD cohorts (Fig. 1c, Jaccard distance, PERMANOVA p = 0.001). Of note, 57 subfamilies had a significantly higher prevalence (two-sided Fisher’s exact test p < 0.05) in healthy individuals as compared to IBD cases (Supplementary Data 5), among which 35 subfamilies (18 CYS subfamilies, 2 CFAT subfamilies, and 15 SDR subfamilies) further show a difference of prevalence > 10% (Fig. 1d).
To further examine the difference between the expression profiles of SoL biosynthetic enzymes between the IBD and healthy groups, we extended our comparative analysis to the metatranscriptomic level. We found that 132 SoL biosynthetic enzyme subfamilies were expressed in 777 metatranscriptomic samples (193 healthy samples and 584 IBD disease samples), with about 42% (55/132) detected in at least 5% of samples (Supplementary Fig. 2d). Beta diversity of the expression profiles of SoL biosynthetic enzymes suggested that the overall expression of these enzyme subfamilies is significantly different between the healthy and IBD cohorts (Fig. 1e, Bray-Curtis distance, PERMANOVA p = 0.001). To capture more detail, we compared the prevalence and abundance differences of each enzyme subfamily in the metatranscriptomic samples. Nine subfamilies had higher prevalence (two-sided Fisher’s exact test p < 0.05, varying from 9% ~ 17%) in the non-IBD group than the IBD group (Supplementary Data 6). We further identified 8 subfamilies (6 CYS, 1 CFAT, and 1 SDR) as significantly different in abundance (expression) profiles between the healthy controls and IBD cases (Fig. 1f, two-sided Mann-Whitney U test, adjusted p < 0.05). Notably, 7 of the 8 subfamilies had a higher prevalence (Fig. 1f, upper panel, two-sided Fisher’s exact test p < 0.05) and a higher abundance (Fig. 1f, lower panel, one-sided Mann-Whitney U test, adjusted p < 0.05) in the non-IBD group than in the IBD group.
We finally looked for metabolomic evidence in the differences of detectable SoLs, the products of the biosynthetic enzymes mentioned above, among IBD and non-IBD groups from publicly accessible metabolomics datasets. We expected that the increased expression of SoL biosynthetic enzymes would correspond with an increased abundance of stool SoLs in non-IBD groups compared to IBD groups after possible uptake by the host. Using metabolomics data from two independent datasets (dataset 1: IBDMDB19,39, corresponding to the same dataset used for metagenomics and metatranscriptomics analysis; dataset 2: PRISM40), we identified metabolite features putatively corresponding to specific sulfonolipids16,41 (Supplementary Data 7), by exact mass comparison with mass error less than 5 ppm (Supplementary Data 8). Within each dataset individually, we indeed found that metabolomic features potentially corresponding to SoLs were decreased in stool samples of IBD groups compared to non-IBD groups. Since there were no MS/MS data available in either dataset, we utilized additional complementary approaches to confirm these features as SoLs including analysis of in-source fragmentation, correlation of co-eluting metabolomic features, and retention time matching between the dataset and data recreated using our own instrument, followed by experimental validation using targeted metabolomics with an additional set of samples from independent IBD disease case and control cohorts.
To validate the presence of SoLs in these datasets, we first tried to identify SoLs using in-source fragments (ISFs) of metabolites based on an established set of criteria42. We initially identified six groups of co-eluting metabolomic features as potential ISFs which showed peak-to-peak intensities highly correlated with putative SoL features (Supplementary Fig. 3, Pearson correlation coefficients ≥0.9). We then examined the reference MS/MS spectra of our isolated and literature-reported SoLs22,41 matching putative SoL masses, which contained limited m/z values corresponding to potential ISFs we identified. However, their relatively low intensity was not conclusive enough to classify them as high-confidence ISFs42. Thus, we proceeded with a complementary correlational approach to identify the putative SoL features. Among the originally identified six groups of co-eluting metabolomic features, five members were detected in both datasets mentioned above (Supplementary Fig. 3c–f, features highlighted in bold). Based on exact mass matching, these features corresponded to SoL analogs: SL 34:1;2 O, SL 17:0;O/16:1;O, SL 33:1;2 O | SL 17:0;O/16:1;O, SL 34:1;2 O | SL 17:0;O/17:1;O, and SL 32:0;O | SL 17:0;O/15:043. In addition, these features had higher peak area-peak area correlation with each other (Pearson correlation ≥0.9; Supplementary Fig. 3c–f). Notably, SoLs are often detected in metabolomics as a series of analogs with consecutive additions of CH2 and H2 moieties within the class and with different numbers of oxygens between classes16,43. Thus, these features likely represent a series of analogs chemically modified from a common parent metabolite, or co-produced by a specific microbe, which is consistent with SoL analogs. These metabolomic features were further positively correlated with species of the prolific SoL-producing genus Alistipes: A. putredinis, A. finegoldii, A. indistinctus, A. shahii, A. onderdonkii, and Bacteroidales bacterium ph8 (which belongs to A. obesi) with Spearman correlation coefficients ≥ 0.5 (Supplementary Figs. 4–6). This positive correlation indicated that the abundance of these metabolites increased with the increase in these species, supporting that these species likely produced these molecules. Furthermore, these metabolomic features had significantly higher abundance in non-IBD groups compared to IBD groups in both IBD datasets (Fig. 2a, Supplementary Fig. 7, Wilcoxon rank sum test, p < 0.05, one-sided), which was consistent with our exact mass matching analysis.
a Box plots showing the relative abundance of SoL candidates detected in dataset 1 (upper) from non-IBD (red boxes, n = 124 individuals) and IBD individuals (blue boxes, n = 348 individuals) and in dataset 2 (lower) from non-IBD (red boxes, n = 56 individuals) and IBD individuals (blue boxes, n = 164 individuals). For each feature, the corresponding SoL is noted in the bottom label. The prefixes of metabolomic feature names correspond to the detection method used: In dataset 1, HILp indicates the HILIC-positive method and HILn indicates the HILIC-negative method. In dataset 2, HILIC-pos indicates the HILIC-positive method and HILIC-neg indicates the HILIC-negative method. Details for the corresponding LCMS methods can be found in the original studies. Significance was determined using the one-sided Wilcoxon rank sum test with the hypothesis that the abundance of SoL was higher in the non-IBD than in IBD group. Exact p-values from left to right are: 1.2E-11, 3.9E-13, 2.3E-14, 1.1E-6, 1.1E-8 in dataset 1 (upper) and 3.3E-8, 6.2E-9, 1.2E-7, 4.5E-10, and 6.5E-11 in dataset 2 (lower). b Box plots showing the absolute abundance of SoLs B, C, and F measured by targeted metabolomics in an independent cohort of IBD patient stool samples. SoL B and F were found to be significantly decreased in IBD (blue boxes, n = 40) compared to non-IBD (red boxes, n = 20) samples. All box plots include center lines representing the median, box limits representing upper and lower quartiles, whiskers representing the 1.5x interquartile range, and points representing outliers. Source data are provided in the Source Data file. Significance was determined using two-sided Student’s t-test. The exact p-values for SoL B and SoL F were 0.0241 and 0.0482, respectively. For all p values: *0.01 <p < 0.05, **0.001 <p < 0.01, and ***p < 0.001.
To further validate our identification of SoLs in these datasets, we acquired one of the columns used to generate the original data19,40. We then selected several standard compounds used in dataset 2 and the candidate SoL B feature that we identified by exact mass matching, and subsequently analyzed the retention times of the standard compounds alongside our own standard SoL B using our in-house HPLC-MS instrument. Due to the inherent variability of retention time between instruments44,45, we calculated the relative retention time (RRT)46,47 using each of the standard’s retention time relative to that of our SoL B standard and compared these values to the RRTs calculated using the corresponding dataset standards and candidate SoL B. We found that the RRT values using our SoL B standard and the RRT values using the candidate SoL B shared a linear relationship (Supplementary Fig. 8, R2 = 0.9915), indicating that the shift in retention time was linear and thus suggesting that the candidate SoL B feature was SoL B.
Finally, to experimentally validate our informatic analysis, we obtained deidentified stool samples collected from an independent cohort of IBD patients (n = 40) and healthy controls (n = 40), and analyzed their SoL abundance by targeted HPLC-MS/MS. We detected SoLs B, C, and F as major SoLs and found that all their abundances were decreased in IBD samples compared to non-IBD samples (Fig. 2b), with SoL B and F being significantly decreased (Wilcoxon rank sum test, p < 0.05, one-sided). This independent validation was consistent with our bioinformatic analysis which also showed that major SoLs including SoL B were significantly decreased in IBD metabolomes and further supported our identification of SoLs in the metabolomics datasets as well as our chemoinformatic analysis showing decreased abundance of SoLs in stool samples of IBD.
Thus, our metagenomic analysis reflected that SoL biosynthetic enzymes were more prevalent in the non-IBD group than the IBD group, metatranscriptomics suggested that genes encoding these enzymes are more actively transcribed in the non-IBD group, and chemoinformatics and metabolomics indicated that representative SoLs are in higher abundance in stool samples from the non-IBD group. We further validated the metabolomics data in an independent cohort of IBD patient samples which showed that SoL abundance was indeed significantly decreased in IBD compared to non-IBD samples. Altogether, our findings establish a negative correlation directly between SoL biosynthesis and IBD, consistent with the previously reported negative association between SoL-producers, namely Alistipes and Odoribacter, and IBD22,23.
An experimental model of colitis demonstrates the negative correlation between SoL production and IBD progression
Encouraged by our informatically predicted negative correlation between SoL biosynthesis and IBD, we sought to further experimentally validate our prediction using a well-established mouse model of IBD. We used Il10-deficient (Il10–/–) mice that are genetically susceptible to developing intestinal inflammation and chronically treated them with the non-steroidal anti-inflammatory drug piroxicam, which induces the development of colitis through the disruption of the gut mucosal barrier in inflammation-susceptible hosts48,49. We selected this model due to its stability, as Il10–/– mice generally will not develop colitis when born and raised under specific pathogen-free conditions unless induced by external stimuli such as piroxicam treatment. This allowed us to more confidently ensure that the effects observed were dependent on the induction of colitis and not due to the Il10 deficiency. Stimulation of mucosal TLRs stemming from mucosal barrier breakdown was another factor in our selection of this model, as we have previously shown that SoL A suppresses LPS-induced inflammation and LPS is well-known to activate TLR signaling22,50. As has been previously reported48,49, we observed that the colonic tissues were inflamed in the piroxicam-treated (IBD) group of Il10–/– mice when compared to the control (pre-IBD) group as indicated by gross pathology and blinded histopathology analyses (Fig. 3a–c, Supplementary Data 9 and 10).
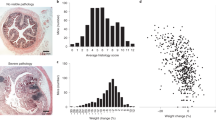
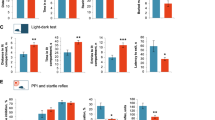
a Histological analysis of the mouse distal colon reveals that piroxicam treatment induced intestinal inflammation in Il10–/–- mice. b, c Histology and gross pathology scores indicate induction of colitis in Il10–/– mice treated with piroxicam (red bars, n = 7 female mice) compared to pre-IBD control Il10–/– mice (green bars, n = 4 female mice), confirming the successful establishment of the IBD model. The trends were consistent in male mice in another independent cohort using the same IBD model (Supplementary Fig. 10). Significance was determined using two-sided Mann-Whitney U test. Bars represent mean ± standard error. Exact p-values were 0.02037 and 0.01951 for Histology Score and Gross Pathology, respectively. d Total ion chromatograms (TICs) obtained from HPLC-HRMS analysis of fecal pellet extracts from control Il10–/– mice and Il10–/– + piroxicam mice reveal the presence of SoL A and SoL B. SoL abundances appear to be decreased in Il10–/– + piroxicam mice fecal pellets. e MS/MS spectra of SoLs A and B confirm their identities based on the presence of the 80 m/z fragment characteristic of sulfonate-containing compounds as well as other characteristic fragments (Supplementary Fig. 9a, b) and compared to literature fragmentation patterns16,22. f, g Peak areas were calculated using TICs obtained after MS/MS fragmentation and used to measure the abundance of SoLs A and B. Both SoLs A and B were significantly decreased in feces from inflamed mice (red bars, n = 7, female) compared to control mice (green bars, n = 4, female). Significance was determined using two-sided Student’s t-test. Bars represent mean ± standard error. Exact p values were 5.42E-5 and 0.01472 for SoL A and SoL B, respectively. h–k Gene expression of inflammatory markers TNFα, NOS2, IL-6, and IL-1β were significantly increased in the ceca of Il10–/– + piroxicam mice (red bars, n = 7, female) compared to control mice (green bars, n = 4, female). Significance was determined using two-sided Mann-Whitney U test. Bars represent mean ± standard error. Exact p values were 0.0061, 0.0061, 0.0061, and 0.0424 for TNFα, NOS2, IL-6, and IL-1β respectively. For all p values: *0.01 <p < 0.05, **0.001 <p < 0.01, and ****p < 0.0001. Source data are provided in the Source Data file.
To explore the link between SoL production and IBD, we collected fecal material from piroxicam-treated Il10–/– (IBD) mice (n = 7, female) and pre-IBD control Il10–/– mice (n = 4, female), extracted metabolites, and measured the abundance of SoLs by targeted metabolomics using high-performance liquid chromatography (HPLC)-high resolution mass spectrometry (HRMS) (Fig. 3d). We detected metabolites with m/z corresponding to major SoLs, specifically SoLs A (SL 34:0;2 O | SL 17:0;O/17:0;O) and B (SL 32:0;O | SL 17:0;O/15:0), in all fecal samples tested and unambiguously determined their identities by HPLC-MS/MS (Fig. 3e, Supplementary Fig. 9a, b). We then determined that the abundances of both SoLs A and B were significantly decreased in feces from piroxicam-treated mice compared to control (Fig. 3f, g). This result confirms our above-described informatic analysis and directly establishes a negative correlation between SoL production and colitis progression in the mouse model. In addition, we also observed significantly increased expression of the NF-κB-regulated inflammatory markers TNFα, NOS2, IL-6, and IL-1β in the IBD mouse group (Mann-Whitney U test, p ≤ 0.005; Fig. 3h–k), further indicating a negative correlation between SoL production and these inflammatory markers. Given our previously observed anti-inflammatory activity of SoL A against LPS22, a natural ligand of TLR4, this negative correlation suggests a potential role of SoLs in regulating IBD that may involve suppressing TLR4-mediated NF-κB activation. To exclude any differences caused by sex, we performed another independent study with male mice using the same model and observed the same negative correlation between SoL production and IBD progression (Supplementary Fig. 10).
Constant identification of SoLs’ contribution to immunomodulatory activity
We next examined the production of SoLs and their contribution to immunomodulatory activity in different human gut commensals. Unlike C. gleum F93 DSM 16776, which we experimentally investigated for its functional metabolites in relation to inflammatory activity22, the prolific SoL-producers Alistipes and Odoribacter had not yet been thoroughly chemically investigated to identify the biologically active components associated with remediation of IBD. In addition, Alistipes and Odoribacter produce a mixture of other SoLs16 and are likely to produce a multitude of other functional metabolites, both of which may complicate the potential immunomodulatory activity of these genera’s metabolites with respect to their bioinformatically predicted negative association with IBD. Thus, we conducted bioactive molecular networking of three Alistipes and two Odoribacter strains (Supplementary Data 11) to identify the constant contributor(s) to biological activity. We fractionated crude extracts of the Alistipes and Odoribacter strains and determined the biological activity of each fraction using a cell-based assay that measured the suppression of LPS-indued TNFα production (Fig. 4a). We simultaneously analyzed each fraction by untargeted HPLC-HRMS/MS to generate molecular networks using the Global Natural Products Social (GNPS) feature-based molecular networking (FBMN) pipeline51. We then correlated the relative expression of TNFα in each fraction with the relative peak area of molecular features across all fractions to generate a bioactivity score reflecting the contribution of specific features to the activity of the fractions. Bioactivity scores and relative peak areas were then mapped onto the molecular network to visualize these contributions. The resulting bioactive molecular network generated from Alistipes timonensis DSM 27924 is presented in Fig. 4b. The SoL-containing cluster contained the most abundant and most active molecular features, as indicated by the node size and color intensity compared to other clusters in the network. Additionally, this cluster contained several known SoLs but many more unannotated SoLs, suggesting that the family of biologically active SoLs is larger than what is currently known. In all other SoL-producers tested, we consistently identified SoLs as a major contributor in the active fractions of each strain (Supplementary Fig. 11). To exclude the possibility of observed SoL activity being influenced by LPS contamination, we confirmed the absence of leftover LPS in the SoL samples using a chromogenic LAL assay (Supplementary Fig. 12). Narrowing down the immunosuppressive activity of each of the strains to SoLs guided us to isolate pure SoLs A and B from A. timonensis DSM 27924 (structures confirmed by NMR spectroscopy; Supplementary Tables 1 and 2, Supplementary Figs. 13–24), as well as from each of the other Alistipes and Odoribacter strains tested. We thus reinforced the contribution of this class of lipids to the observed biological activity of Alistipes and Odoribacter.
a A crude extract of A. timonensis DSM 27924 was separated based on polarity into 5 fractions. Each fraction was used in an in vitro cell-based assay (n = 3 individual wells) to measure its respective capacity to suppress LPS-induced expression of TNFα. Fraction 2 (highlighted as a green bar) was found to have the most significant anti-inflammatory effect compared to LPS. All fractions were compared to LPS for statistical significance with only fractions 1 and 2 showing significant change. Fractions 3, 4, and 5 showed no significant change. Statistical significance was determined using two-sided Student’s t-test. Bars represent mean ± standard error. Exact p values were 0.02943 and 0.0133 for LPS against LPS + Fraction 1 and LPS against LPS + Fraction 2, respectively. For all p values: *0.01 <p < 0.05. Source data are provided in the Source Data file. b Untargeted HPLC-HRMS/MS was used to construct a molecular network for each fraction through GNPS FBMN. The relative peak area of each molecular feature in fraction 2 was mapped to the color of the nodes with more abundant features increasing from white to green. Bioactivity score was mapped to the node size with larger nodes indicating stronger negative correlations. Several known SoLs were annotated in this cluster and their structural variations are illustrated, further demonstrating that SoLs as a family of molecules contribute to the observed suppression of LPS-induced TNFα expression.
SoL A mediates dual immunomodulatory activity through TLR signaling
While the causes of IBD remain largely unknown, IBD progression has been linked to aberrant TLR signaling50. TLRs are pattern recognition receptors (PRRs) that initiate a variety of host processes, especially inflammatory responses, through the recognition of pathogen-associated molecular patterns (PAMPs) and other non-pathogenic microbial factors50,52,53. Specifically, TLR2 and TLR4 are well-known to recognize PAMPs in the gut microbiome50. In addition, their expression is significantly increased in IBD pathogenesis, reflecting a state of aberrant activation50,52. Thus, we expected that SoLs may interact with TLR4 or TLR2 to mediate their immunomodulatory activity. We treated primary mouse macrophages collected from wild-type C57BL/6 mice with SoL A (as a representative of SoLs) either alone or together with LPS (an agonist of TLR4) or Pam3CSK4 (an agonist of TLR2/1) and measured the expression of three inflammatory cytokines (IL-6, TNFα, and IL-1β). By itself, SoL A exhibited a mild to moderate effect on the expression of pro-inflammatory cytokines compared to control (Fig. 5), generally consistent with our previous finding22. As expected, the TLR ligands, LPS and Pam3CSK4, both showed significant induction of all three cytokines compared to control (Student’s t-test, p ≤ 0.0001) (Fig. 5). Notably, SoL A was found to significantly suppress the expression of all three cytokines induced by LPS (p ≤ 0.0001) (Fig. 5). Together, SoL A’s mild pro-inflammatory activity by itself and strong inhibition against LPS-induced inflammation constitute its dual immunomodulatory activity. SoL A also inhibited Pam3CSK4-induced IL-6 and TNFα to a smaller extent while increasing IL-1β expression induced by Pam3CSK4 (p ≤ 0.05) (Fig. 5). This result indicates that SoL A primarily affects LPS-induced inflammation and implies that interaction with TLR4 may be involved in SoL A’s mechanism of action. Interestingly, SoL A’s partial suppression of Pam3CSK4-induced inflammation suggests that SoL A-related anti-inflammatory activity may also extend to the TLR2/1 pathway and warrants further investigation. After identifying that SoL A’s primary effect is through TLR4, we further examined the biological activity of SoL B against LPS-induced inflammation. We found that SoL B also inhibited LPS-induced inflammation albeit to a lesser extent than SoL A (Supplementary Fig. 25). This activity is consistent with previous reports that SoL B exhibits anti-inflammatory activity both in vitro and in vivo in mouse models of acute inflammation54.
Mouse peritoneal macrophages were treated with SoL A (10 μM), LPS (100 ng/mL), and Pam3CSK4 (500 ng/mL), either alone or in combination for 6 h. RT-qPCR analysis revealed that SoL A induces a mild pro-inflammatory effect compared to control but significantly suppresses LPS-induced cytokine expression levels and only partially suppresses Pam3CSK4-induced cytokine expression. Bars represent mean ± standard error. Experiments were independently repeated three times. For each treatment, n = 3 individual wells. Significance was determined using two-way ANOVA. For all p values: *p < 0.05, ****p < 0.0001. Source data are provided in the Source Data file.
Molecular docking and ELISA displacement assay suggest SoLs binding to TLR4/MD-2 complex
LPS stimulation of TLR4 occurs through a series of interactions ultimately resulting in LPS binding to MD-2, which forms a complex with TLR4 and induces dimerization to initiate signaling55,56,57. The TLR4/MD-2 heterodimer recognizes structurally diverse LPS molecules, giving it the flexibility to detect different LPS-related PAMPs in the human gut microbiome57. Interestingly, the TLR4/MD-2 complex was recently found to recognize human sulfatides, sphingolipid derivatives which bear a sulfated saccharide head group and dual acyl chains, presumably mimicking the disaccharide core and multiple acyl chains of LPS58. Comparing the chemical structure of SoL A to those of sulfatides and lipid A (the immunogenic portion of LPS) (Fig. 6a), we noted structural similarity in the negatively charged head groups and multiple acyl chains. We thus considered if multiple molecules of SoL A might bind to MD-2 in a similar configuration as sulfatides and lipid A. Inspired by sulfatides that bind in triplicate to MD-258, we used molecular docking to model the binding of three molecules of SoL A to MD-2. Our analysis predicted three molecules of SoL A indeed bind in the hydrophobic pocket of MD-2 (Fig. 6b), where lipid A is known to bind, with a docking score of -8.9 kcal/mol, better than that of lipid A which had a docking score of -6.2 kcal/mol. Additionally, SoL A was predicted to make hydrophobic contacts with several amino acids including I117, F119, I52, and F121 (Supplementary Data 12), all of which are also reported to contact the acyl chains of lipid A57. Notably, SoL A is also predicted to contact residues including R264 and R90 (Supplementary Data 12), consistent with contacts between these residues and the phosphate groups of lipid A57. This suggests that SoL A may bind directly to the TLR4/MD-2 complex and possibly compete for binding with LPS, allowing it to suppress LPS-induced activation of the TLR4 pathway. A critical aspect of lipid A binding to MD-2 is the exclusion of one acyl chain from the hydrophobic pocket of MD-2 which forms a bridge with TLR4 and is involved in inducing dimerization57. Likewise, we observed one acyl chain of SoL A excluded from the hydrophobic pocket in our docking analysis (Fig. 6b), further suggesting that SoL A mimics LPS as a ligand for TLR4. After successfully docking SoL A we then tested SoL B which lacks an extra hydroxy group, (Fig. 6a) potentially increasing its interactions with the hydrophobic binding pocket of MD-2. Our docking analysis indeed showed that SoL B also binds to MD-2 (Fig. 6b), with similar contacts as SoL A (Supplementary Data 12) but higher affinity (docking score of −9.6 kcal/mol) as predicted.
a Chemical structures of immunogenic lipid A (derived from LPS), sulfatide, SoL A, and SoL B illustrating structural similarity in multiple acyl chains and negatively charged head groups. b Molecular docking of lipid A (red), sulfatide (magenta), SoL A (blue), and SoL B (orange) into the hydrophobic pocket of MD-2 in complex with TLR4. Three molecules of SoLs A and B were used in molecular docking experiments to mimic the six acyl chains of lipid A as inspired by sulfatides58. c ELISA displacement assay used to measure the binding behavior of SoLs A and B in competition with 1 ng/mL LPS, a natural ligand of the TLR4/MD-2 complex. Compounds were added either simultaneously (blue), LPS first (green), or SoL first (red). Bars indicate mean ± standard deviation. Experiments were independently repeated three times. For each treatment, n = 3 individual wells. Source data are provided in the Source Data file.
To experimentally determine if SoLs A and B bind to MD-2 and to what extent the SoLs compete with LPS for binding to MD-2, we conducted an ELISA-based displacement assay. Taking advantage of biotinylated LPS, which retains the activity of unconjugated LPS59, we measured absorbance generated by an HRP-linked streptavidin probe to measure the relative amount of MD-2 which was bound with biotin-LPS as opposed to MD-2 bound with SoL A or B. We administered 0.1, 1.0, and 10 μM concentrations of SoL A or B and 1 ng/mL biotin-LPS to MD-2 in three sequences: 1) SoL first followed by LPS 1 h later, 2) LPS first followed by SoL 1 h later, and 3) both SoL A or B and LPS at the same time. After 1 h of incubation, we found that at all concentrations, when SoL A or B was added first, there was a marked decrease in percent absorbance as compared to when LPS was added first and when the two compounds were added together (Fig. 6c). This suggests that SoLs A and B both bind and occupy some sites of MD-2, preventing LPS from fully binding when it is added 1 h after SoL A or B. Furthermore, when moving from low to high concentrations of SoL A or B, we observed that the percent absorbance decreased dramatically. This indicates that with increasing concentration of SoL A or B, less LPS binds to MD-2, implying that SoLs indeed compete with LPS for binding to MD-2. Taken together, these results indicate that SoLs A and B can bind directly to MD-2 and more importantly compete with LPS for binding to this target, thus providing a potential molecular mechanism underlying SoL A’s pro-inflammatory activity by itself as well as its strong activity in suppressing LPS-induced inflammation which likely also expands to other members of the SoL family.
SoLs suppress LPS-induced TLR4 signaling to regulate macrophage polarization
Upon LPS binding, TLR4 initiates downstream signaling, such as through the NF-kB and MAPK pathways, resulting in the induction of inflammatory cytokine expression52. If a SoL binds to MD-2, it will block LPS-induced activation of the TLR4 pathway. Therefore, we investigated whether the addition of SoL A or B affected the phosphorylation of TLR4-downsteam signaling molecules, ERK1/2, and p38, or the degradation of IκBα, which are all critical for LPS-induced cytokine expression52 (Fig. 7a). We treated macrophages with LPS in the presence of increasing concentrations of SoL A or B (from 0 to 20 μM), then performed western blot analysis to examine the TLR4-downstream signaling pathways. We found that both SoLs reduced LPS-induced phosphorylation of p38 and ERK1/2 in a concentration-dependent manner. At the concentration of 20 μM, SoLs A and B almost completely blocked LPS-induced phosphorylation of p38 and ERK1/2 (Fig. 7b). Western blot also showed that SoLs concentration-dependently suppressed LPS-induced IκBα degradation (Fig. 7b). These results support that SoLs exert their anti-inflammatory effect by blocking LPS-mediated phosphorylation of downstream TLR4 proteins, effectively negating LPS activation of the TLR4 pathway. Also, SoL A or B alone at the concentration of 20 μM slightly enhanced the phosphorylation of certain signaling molecules (e.g., ERK1/2) compared to control, consistent with our observations that SoLs alone induced a mild pro-inflammatory effect on cytokine expression (Fig. 5) and supporting the dual immunomodulatory activity of SoLs which provides further opportunities to regulate homeostatic immune responses.
a Simplified pathway of TLR4 activation by LPS, highlighting proposed inhibition by SoLs competing for LPS binding. Proteins IκBα, ERK1/2, and p38 downstream of TLR4 which were selected for analysis are highlighted in red rectangles. Created in BioRender. Chen, H. (2024) BioRender.com/v71h443. b Western blot analysis of protein levels of IκBα as well as total and phosphorylated ERK1/2 and p38, after treatment with LPS (100 ng/mL) with or without various concentrations of SoL A or B. The housekeeping gene β-Actin was used as a loading control. The gel for IκBα was spliced to remove extra lanes (See Source Data for raw gel images). c, d THP-1-derived macrophages were treated with LPS + IFN-γ or IL-4 + IL-13 to polarize to M1 or M2 macrophages, respectively. Relative expression of markers IL-6, CXCL10, IL-12β, and TNFα (compared to M0) indicate that SoL A (10 μM) had a significant effect on suppressing M1 polarization (c); and relative expression of markers IL-10, CD206, and CD209 (compared to M0) indicate that SoL A had no significant effect on M2 polarization (d). Bars represent mean ± standard error. Experiments were independently repeated three times. For each treatment, n = 3 individual wells. Significance was determined using two-way ANOVA. For all p values: **0.001 <p < 0.01, ***0.0001 <p < 0.001, and ****p < 0.0001. Source data are provided in the Source Data file.
Because TLR4 signaling leads to macrophage polarization which has been shown to contribute to IBD60,61,62, we also examined the effects of SoL A on macrophage polarization. We treated THP-1 monocytes with IFN-γ and LPS to induce M1 polarization or IL-4 and IL-13 to induce M2 polarization. Successful induction of M1 and M2 polarization was confirmed by morphology changes and subsequent RT-qPCR quantification of cytokine profiles. When 10 μM of SoL A was added alongside the respective inducing agents, our relative cytokine expression results showed that SoL A significantly reduced the production of M1-polarized macrophage markers IL-6, CXCL10, IL-12β, and TNFα, compared to macrophages treated without SoL A (Fig. 7c) but had a mostly non-significant effect on M2 polarization (Fig. 7d). This suggested that SoL A suppresses macrophage M1 polarization, which supports our aforementioned result that SoLs interfere with TLR4 signaling potentially leading to inhibition of TLR4-mediated IBD.
Discussion
The results of this study have described two major points which are summarized here and discussed in detail below. First, we have leveraged our biosynthetic enzyme-guided disease correlation approach to directly connect the biosynthesis and production of a class of abundant yet underexplored human microbial metabolites, SoLs, to IBD, an existing human health condition with complex and poorly understood etiology, followed by an independent IBD patient cohort and mouse model of IBD to validate this informatically predicted negative correlation. Second, we have revealed that SoLs A and B, two representative gut microbial SoLs, modulate host immune responses through the TLR4/MD-2 complex and inhibit LPS-induced TLR4 activation and macrophage M1 polarization, which provides a mechanistic explanation for SoLs’ potential protective activity against IBD.
Through both bioinformatic and chemoinformatic analyses, we revealed that the expression of SoL biosynthetic enzymes and abundance of SoLs in the gut metabolome are negatively correlated with IBD status in humans. Our IBD model with male and female piroxicam-treated Il10–/– mice supported this sex-independent negative correlation with a concurrent negative association between SoL production and TLR-4-related inflammatory markers TNFα, NOS2, IL-6, and IL-1β. Our findings were consistent with literature reports which have shown that the bacterial genera of abundant SoL production, Alistipes and Odoribacter, are associated with the remediation of IBD symptoms20,21. Considering that these SoL-producers are commensal members of the human gut microbiome16,63, their constant production of SoLs in the human gut may help to maintain intestinal immune homeostasis, thereby preventing IBD. Both Alistipes and Odoribacter are also known to produce SCFAs, which have been implicated in reducing intestinal inflammation12,21,63,64,65. Besides SCFAs, there are other gut microbial functional metabolites such as secondary bile acids and indole derivatives which have also been linked to modulating host inflammation and immunity66,67,68,69,70. Thus, the role SoLs play individually and/or synergistically with other factors in IBD pathogenesis remains interesting and awaits further investigation.
Toward understanding the mechanism of the immunomodulatory activity of SoLs, we showed that the representative SoLs A and B primarily target the TLR4 pathway and that they both block LPS binding to MD-2 to suppress TLR4 signaling. Our analysis indicated that SoL A preferentially suppresses LPS-induced inflammation as compared to Pam3CSK4-induced inflammation, suggesting that TLR4 activation is more strongly affected by SoL A than TLR2/1 activation. The selectivity for TLR4 may stem from SoL A’s structural similarity to LPS, a hypothesis supported by the recently reported human TLR4 ligands, sulfatides, which share highly similar structural features to SoLs58. As microbial functional metabolites, it is reasonable that SoLs are directly recognized by TLR4, which has evolved specifically to recognize PAMPs71. Our molecular docking also suggested SoLs A and B’s recognition by TLR4, indicating that three molecules of either SoL indeed bind with stronger predicted binding score compared to lipid A and make contacts with important amino acids in the pocket of MD-2, consistent with lipid A binding as well as structurally-related sulfatides57,58. Our ELISA-based displacement analysis then confirmed that both SoLs A and B directly bind to MD-2, block LPS binding when added prior to LPS, and could displace bound LPS from MD-2 at higher concentrations. We further demonstrated that increasing concentration of SoLs A and B suppressed LPS-induced TLR4 signaling pathways in a dose-dependent manner and SoL A significantly suppressed M1 polarization of macrophages, further indicating their capacity to reduce downstream TLR4 signaling responses. Notably, increased numbers of M1 macrophages is a characteristic feature of IBD62 and this suppressive effect may represent one explanation of how SoL-producing bacteria are able to remediate symptoms of IBD. Taken together, our results represent the first report of SoLs A and B’s binding to MD-2 and establish that SoLs are likely to mediate their dual immunomodulatory activity by occupying the hydrophobic binding pocket of MD-2 (pro-inflammatory by SoLs alone) but primarily by blocking LPS binding to the TLR4/MD-2 complex (anti-inflammatory against LPS-induced inflammation). This discovery suggests SoLs’ mechanistic role in regulating a multitude of TLR4-related inflammatory conditions, most notably IBD which is associated with dysregulation of TLRs, especially TLR423,24,25,26, leading to aberrant macrophage activation62,72.
While many studies have focused on the association of certain microbes with specific diseases by analyzing the abundance and distribution of microbial strains18,19,73,74,75,76, we describe here an approach which leverages disease-related sequencing data to connect the biosynthesis of functional metabolites directly to human health conditions. Our approach takes advantage of critical biosynthetic enzymes specifically required for the biosynthesis of their corresponding functional metabolites. Increased or decreased expression of these enzymes, reflecting the differential production of a specific functional metabolite, can be correlated with different disease states to reveal positive or negative associations. Through these correlations, our biosynthetic enzyme-guided disease correlation approach can reveal trends in functional metabolite biosynthesis in the context of human health conditions, enabling a more focused, targeted chemoinformatic analysis to rapidly fish out metabolites of interest and further confirm their association with disease. By applying this approach, we have identified a negative relationship between SoL biosynthesis and IBD directly from patients’ data followed by verification using an IBD mouse model, which has guided us to further reveal a molecular target and a potential mechanistic explanation for the protective effect that SoLs and SoL-producing bacteria exert against IBD. We are now further characterizing the biological role and therapeutic potential of SoLs through elucidating the mechanism of SoL delivery and uptake in the host and probing their effects on IBD pathogenesis. Our future studies include colonization of SoL-producers in the Il10–/– mouse model, determining whether SoLs are gut-restricted microbial metabolites or not, and exploring the role of microbial delivery systems such as outer membrane vesicles. Furthermore, with the increasing availability of disease-related sequencing data32,77 and the rapidly advancing investigation of microbial biosynthetic pathways78,79,80, we envision that our approach can be broadly applied to uncover previously unknown human microbial metabolites with potentially important human health implications and facilitate the investigation of their complex effects on human health.
Methods
Identification of SoL biosynthetic enzymes from human gut bacterial reference genomes
We collected experimentally validated enzymes involved in SoL biosynthesis as reference amino acid sequences of CYS, CFAT, and SDR (Supplementary Data 1). Reference amino acid sequences were used as seed sequences to search for homologs in the human gut bacterial reference genomes using the DIAMOND blastp model81 with an e-value threshold of 10−5. We then investigated the taxonomic distribution of the resulting homolog sets based on the taxonomy annotation for each genome33. To prioritize SoL biosynthetic enzymes, the homologs of CFATs and CYSs from genomes which encode copies of CFAT, CYS, and SDR were first used to generate sequence similarity networks with experimentally validated CFAT and CYS at a threshold of 50% similarity using MMseqs282. The prioritized CFATs and CYSs meet co-occurrence with SDRs in the same genome. The filtered enzymes were further subjected to Pfam domain analysis by hmmsearch (HMMER v3.3) with default parameters against the Pfam-A database (v33.1). Enzymes containing corresponding Pfam domains with hit score >50 were selected as prioritized SoL biosynthetic enzymes. Prioritized enzymes were used to generate CYS, CFAT, and SDR subfamilies using sequence similarity networks with at least 90% similarity by MMseqs2 clustering82. Maximum-likelihood trees were generated using the representative genome (Supplementary Data 3) of each species by GTDB-Tk (v2)83 with the following parameters (refer to the GTDB-Tk user guide: https://ecogenomics.github.io/GTDBTk/commands/index.html): 1. GTDB-Tk reference data release 207 was used. 2. Parameters of classify: gtdbtk classify_wf --genome_dir --out_dir --force. 3. Parameters of infer tree from multiple sequence alignment: gtdbtk infer --msa_file MSA_FILE --out_dir OUT_DIR. 4. Using the convert_to_itol command to make the tree suitable for visualization in iTOL: gtdbtk convert_to_itol --input_tree --output_tree. Finally, we annotate their phylogenetic trends using iTOL84.
Quantification of enzyme abundances in metagenomic and metatranscriptomic samples
Metagenomic and metatranscriptomic whole-genome sequencing datasets of human gut microbiomes related to IBD were downloaded from the Inflammatory Bowel Disease Multi’omics Database19,39 (IBDMDB, https://ibdmdb.org/tunnel/public/summary.html) For both metagenomic and metatranscriptomic samples, reads were quality filtered and adapter removed using bbduk.sh with the following parameters: qtrim=rl ktrim=r mink=11 trimq=10 minlen=40 (read quality cutoff is 10, read length cutoff is 40). High-quality reads were mapped to the nucleotide sequences of corresponding contigs containing the SoL biosynthetic genes using the BWA mem algorithm85 with default parameters. The reads mapped to genes were counted by featurecounts86 with the following parameters: -f -t CDS -M -O -g transcript_id -F GTF -s 0 -p --fracOverlap 0.25 -Q 10 –primary. Enzymes encoded or expressed in at least 5% of samples were considered as common distribution in humans and were included in the comparative analysis. Transcripts per million (TPM) were calculated for each SoL biosynthetic enzyme-encoding gene. The abundance of each subfamily was calculated by the sum of the relative abundances of all genes in the subfamily. Beta diversity was performed to quantify the prevalence and relative abundance differences in the overall composition of SoL biosynthetic enzymes between the IBD and the control groups. PERMANOVA was performed to show the encoding and expression profile differences of SoL biosynthetic enzymes between IBD and control groups. Both beta diversity and PERMANOVA were performed using the R package vegan87,88. To explore the differences between IBD and control groups of single SoL biosynthetic enzyme subfamilies, we used the Shapiro–Wilk test to evaluate the normality of a specific gene family’s relative abundance. We then calculated the significance of relative abundance between healthy and IBD individuals using either two-sample Student’s t-test (for normally distributed data) or a two-sample Wilcoxon rank sum test (for not normally distributed data). Significance tests were performed in Python using packages Pandas and SciPy89,90,91. The Benjamini-Hochberg method was used to adjust p-values to correct for multiple testing92. SoL biosynthetic enzyme subfamilies were considered differential if the adjusted p-value was less than 0.05. For differential Sol biosynthetic enzymes, we performed a two-sided Fisher’s exact test to explore their difference in prevalence across IBD and non-IBD groups.
Analysis of publicly available metabolomics datasets
Processed per-subject metabolomic feature tables and microbial species relative abundance profiles were collected from two publicly available IBD datasets: IBDMDB19,39 (https://ibdmdb.org/downloads/html/rawfiles_MBX_2017-03-01.html) and PRISM40 (https://www.nature.com/articles/s41564-018-0306-4#Sec31). Known SoL-related metabolites were collected from MS-DIAL441 and reference16 (Supplementary Data 7). Search parameters were set to the exact mass of SoLs using a 5-ppm match tolerance for parent ions. Identified metabolomic features were considered SoL candidates. Next, features co-eluting with SoL candidates (within ± 0.05 min retention time window) were selected to build a feature similarity network (Pearson correlation). Relative abundance was normalized using the equation below following the calculation described in dataset 2:
Here, Ni represents the absolute intensity of a metabolomic feature, Sum(Ni) represents the total absolute intensity of all features in the sample. In this step, all the metabolomic samples were included (dataset 1: 546 samples; dataset 2: 220 samples). The correlation coefficients ≥0.9 between SoL candidates and other metabolomic features were subjected to further analysis hereafter. If the co-eluting features could be identified in SoL candidates’ MS/MS spectra in MS-DIAL441 and a previous in-depth study16, these features were considered SoL candidates’ in-source fragments. If not, these metabolites might be co-produced by a specific microbe, or chemically modified from a common parent metabolite, and these metabolites were considered as SoL analog candidates. Of note, each data type, detected using the same LCMS method19,39,40, was analyzed separately in each cohort.
To further examine whether SoL candidates correlated with the abundance of any taxa, the Spearman correlation between species-level relative abundances and SoL candidates’ relative abundance was calculated. To do so, a series of SoL analog candidates (SoL candidates correlated with other SoL candidates) were selected for further analysis. In total, 472 samples collected from dataset 1 and 220 samples collected from dataset 2 were included. Of note, corresponding microbial species’ relative abundance profiles and metabolomic features data were paired in these samples. Species that were present in less than 5% of samples were excluded. Finally, the relative abundance of SoL analogs was used to calculate differential abundances between non-IBD and IBD samples. The one-sided Wilcoxon rank sum test was used to measure statistical significance with the hypothesis that the abundance of SoL was higher in the non-IBD group than in UC or CD.
Animals
B6.129P2-Il10tm1Cgn/J (Il-10 KO; Il10–/–) mice from the C57BL/6 background were originally purchased from The Jackson Laboratory. Mice were bred and maintained in specific pathogen-free conditions at the University of South Carolina and were maintained on a 12-hour light/dark cycle with unlimited access to water and food (Inotiv Teklad, 8604, https://insights.envigo.com/hubfs/resources/data-sheets/8604-datasheet-0915.pdf). All animal protocols were approved by the University of South Carolina Institutional Animal Care and Use Committee.
Piroxicam-accelerated Il10 –/– mouse IBD model and analysis
At 8-12 weeks of age, male (n = 5 mice distributed in 2 cages) and female (n = 7 mice distributed in 2 cages) Il10–/– mice were switched to a normal chow diet (Inotiv Teklad, 8604) supplemented with 100 ppm of piroxicam (Cayman Chemical, 13368; Inotiv Teklad, TD.210442) to induce colitis development as previously described49. In parallel to these mice, male (n = 3 in 1 cage) and female (n = 4 in 1 cage) Il10–/– mice were maintained on the control diet for the duration of the experiment to serve as pre-colitic controls. Mice were euthanized at 18 days to collect tissues for inflammation assessment and intestinal contents for quantification of SoLs. At necropsy, colitis severity was first grossly assessed, which included qualitative evaluations of cecal atrophy (0–5), thickening of cecal (0–5) and colon tissues (0–5), the extent of content loss in the cecum (0–4) and stool consistency/diarrhea (0–3). For histopathology, segments of the colon were first washed in PBS and then fixed in 10% neutral buffered formalin. The tissues were embedded in paraffin, cut into 5-mm sections, and stained with hematoxylin and eosin (H&E) at the Instrumentation Resource Facility at the University of South Carolina School of Medicine. Inflammation scores of colon sections were blindly assessed as previously described93 using an Echo Revolve light microscope and accompanying software. Briefly, colitis severity was assessed based on the following histopathological features: length measurements in microns of crypt hyperplasia converted to a score from 0-4, qualitative assessment of goblet cell loss (0–5), crypt abscesses per 10X field counts converted to a score from 0-4, and qualitative assessment of submucosal edema (0-3). RNA isolations and RT-qPCR were performed as previously described94. Briefly, RNA was isolated from snap-frozen cecal tissues using the TriZol method (Thermo Fisher Scientific). cDNA was synthesized using SuperScript III reverse transcriptase (ThermoFisher Scientific). qRT-PCR was performed at the Functional Genomics Core at the University of South Carolina. The relative abundance of mammalian mRNA transcripts was calculated using the ΔΔCT method and normalized to Eef2 levels. The oligonucleotides used for qRT-PCR were: Eef2 forward: TGTCAGTCATCGCCCATGTG, reverse: CATCCTTGCGAGTGTCAGTGA; Tnfa forward: AGCCAGGAGGGAGAACAGAAAC, reverse: CCAGTGAGTGAAAGGGACAGAACC; Nos2 forward TTGGGTCTTGTTCACTCCACGG, reverse: CCTCTTTCAGGTCACTTTGGTAGG; Il6 forward: GAAATGATGGATGCTACCAAACTG, reverse: CTCTCTGAAGGACTCTGGCTTTG; Il1b forward: CTCAATGGACAGAATATCAACCAAC, reverse: GGCTGTGCCGTCTTTCATTAC. Fecal samples were collected and immediately flash-frozen and stored at –80° C.
SoL extraction from mouse and patient fecal material
For SoL extraction and quantification, frozen mouse or human fecal samples were lyophilized to remove remaining water and resuspended in methanol (Thermo Fisher Scientific). Fecal sample suspensions were vortexed for 1 min prior to sonication for 10 min. The methanol extract was collected by centrifugation at 20,000 x g for 10 min and dried under a gentle stream of nitrogen. The resulting residue was then redissolved in methanol + 0.1% ammonium hydroxide (ThermoFisher Scientific) and filtered through a 0.22 μm filter prior to analysis.
Liquid chromatography and mass spectrometry analysis
High-resolution mass spectra were collected using a ThermoFisher Scientific Q-Exactive HF-X hybrid Quadrupole-Orbitrap mass spectrometer using electrospray ionization in negative mode. Liquid chromatography (LC) used a ThermoFisher Scientific Vanquish HPLC coupled to the aforementioned mass spectrometer. LC was performed using a Waters Xbridge BEH C18 XP column (2.1 x 100 mm) with alkaline mobile phase (pH 10.7) consisting of solvent A (water + 0.1% ammonium hydroxide) and solvent B (acetonitrile + 0.1% ammonium hydroxide) in a gradient starting from 35% B and increasing to 100% B over 10 min, hold at 100% B for 5 min, then re-equilibration at 35% B for 5 min at a flow rate of 0.4 mL/min. MS scans were obtained in the orbitrap analyzer which was scanned from 500 to 2000 m/z at a resolution of 60,000 (at 200 m/z). Targeted MS/MS fragmentation was conducted for SoLs in a ± 0.5 Da window around their expected m/z. MS data were analyzed by Xcalibur (ThermoFisher Scientific 4.2.47). Relative retention time was calculated by dividing RT of the analyte of interest by the RT of a standard compound.
Anaerobic culture and bioactive molecular networking
Three Alistipes (A. putredinis DSM 17216, A. timonensis DSM 25383, and A. timonensis DSM 27924) and two Odoribacter strains (O. laneus DSM 22474 and O. splanchnicus DSM 20712) were cultured in Reinforced Clostridial Medium (RCM, BD Biosciences) under anaerobic conditions at 37 °C. After three days of growth, cultures were harvested by centrifugation at 12,000 x g for 30 min. The resulting cell-free supernatant was extracted with an equal volume of methyl ethyl ketone and the cell pellets were extracted by resuspension in methanol and sonication before both extracts were combined and concentrated in vacuo. The combined crude extract was then fractionated on a silica gel column using a stepwise gradient of dichloromethane and methanol (DCM:MeOH; 15:1, 7:1, 5:1, 3:1, 1:1). Each fraction was then used in an in vitro cell-based assay measuring the suppression of LPS-induced TNFα expression. Simultaneously, samples of the fractions were subjected to untargeted HPLC-HRMS/MS as described above. MS/MS was conducted using data-dependent acquisition with a resolution of 30,000, isolation window of 2.0 m/z, and dynamic exclusion time of 15 s. HPLC-HRMS/MS data was processed using MZmine3 following the GNPS FBMN workflow with minimal changes95. Molecular networks were constructed using the quickstart GNPS FBMN setting with no changes51. Raw data used for this analysis was deposited in the University of California, San Diego Center for Computational Mass Spectrometry MassIVE database (https://doi.org/doi:10.25345/C5028PP9T; ftp://massive.ucsd.edu/MSV000091884/). Bioactivity scores were assigned using a custom R script which calculated Pearson correlation coefficients between each molecular feature and the activity of each fraction96. Finally, bioactive molecular networks were visualized in Cytoscape v3.9.197.
Purification of SoLs A and B
Fractions containing SoLs were further purified by Sephadex LH-20 run in 1:1 DCM:MeOH. Finally, pure SoLs A and B were isolated by semi-preparative scale HPLC running an isocratic solvent composition of 47% water + 0.1% ammonium hydroxide and 53% acetonitrile + 0.1% ammonium hydroxideon a ThermoFisher Scientific Ultimate 3000 semi-preparative scale HPLC equipped with a Waters Xbridge Prep C18 5 μm OBD column (19 x 100 mm) with a flow rate of 5.0 mL/min. 1H, 13C, 1H-13C HSQC, 1H-13C HMBC, and 1H-1H COSY NMR spectra for SoLs A and B were acquired in methanol-d4 on a Bruker Avance III HD 400 MHz spectrometer with a 5 mm BBO 1H/19F-BB-Z-Gradient prodigy cryoprobe. Data were collected and reported as follows: chemical shift, integration multiplicity (s, singlet; d, doublet; t, triplet; m, multiplet), coupling constant. Chemical shifts were reported using the methanol-d4 resonance as the internal standard for 1H-NMR methanol-d4: δ = 3.31 ppm and 13C-NMR methanol-d4: δ = 49.0 ppm. Pure SoLs A and B were confirmed to be free of LPS using a Chromogenic Endotoxin Quant Kit (Pierce).
Preparation and treatment of macrophages
Primary mouse macrophages were prepared by first introducing 3 mL of 3% thioglycolate to mice via intraperitoneal injection. After 3 days, 10 mL of chilled PBS was introduced intraperitoneally to flush out macrophages. The cell suspension was then separated by centrifugation at 300 x g for 5 min. Cells were seeded in culture dishes containing DMEM with 10% FBS for 1 h before being washed with serum-free DMEM two times to remove unattached cells. The cells were incubated in serum-free DMEM for 16 h before treatment. To treat the macrophages, cells were incubated for 6 to 24 h in DMEM without FBS with addition of LPS (Sigma-Aldrich), Pam3CSK4 (Invivogen), or SoL A. Cells were finally washed twice with Dulbecco’s phosphate-buffered saline before being lysed for total RNA or protein extraction.
mRNA extraction and RT-qPCR in macrophage-based assays
Treated mouse macrophage cells were lysed with TriZol (Invitrogen) and total RNA was extracted from the cell lysate using a Direct-zol RNA miniprep kit (Zymo Research) according to the manufacturer’s protocol. The quality and quantity of RNA was then determined using a nanodrop and 1000 ng of mRNA from each sample was used for cDNA synthesis using a First-strand cDNA Synthesis System (Marligen Bioscience). qPCRs reactions were prepared in a 20 uL final volume containing Fast Start Universal SYBR Green Master (Rox) (Roche Applied Science), cDNA template, deionized water, and primers and probes for IL-1β, TNFα, IL-6, and the 18S rRNA which was used as a housekeeping gene Cycling conditions were 95 °C for 10 min followed by 40 cycles of 95 °C for 10 s, 60 °C for 15 s, and 68 °C for 20 s, then a melting curve analysis from 60 °C to 95 °C every 0.2 °C was obtained. Amplifications were performed on an Eppendorf Realplex Mastercycler (Eppendorf). Relative gene expression levels were calculated using the ΔΔCT method and expression levels of 18S were used to normalize the results.
Molecular docking
The crystal structure of the TLR4/MD-2 comlpex was retrieved from the Protein Data Bank (PDB ID: 3FXI)57 and prepared using AutoDock Tools98. Molecular structures of SoL A, SoL B, sulfatide, and lipid A were constructed, and energy minimized using Marvin version 21.17.0, ChemAxon (https://www.chemaxon.com). Models of SoL A, SoL B, sulfatide, and lipid A were also prepared using AutoDock Tools and docked against the TLR4/MD-2 complex using AutoDock Vina99,100 in a 32x32x32 angstrom box surrounding the MD-2 monomer. Docking results were visualized using PyMol101.
ELISA displacement assay
Solid-phase sandwich ELISA kits were purchased from Invitrogen. The ELISA experiments were performed according to the kit instructions, using 50 nM hMD-2 (Novus Biologicals), 1 ng/mL LPS-EB Biotin (Invivogen, Cat. No.: tlrl-lpsbiot, derived from E. coli 0111:B4), and 0.1, 1.0, and 10 μM purified SoL A. SoL A was added to the assay 1 h before, 1 h after, or simultaneously with LPS-EB Biotin. Absorbance was measured at 450 nm using a BioTek microplate reader.
Western blot
Macrophages were treated with 100 ng/mL of LPS and 5 or 20 μM SoL A for 30 min. Following treatment, all cellular protein was extracted using MPER lysis buffer (Thermo Scientific). Protein samples were loaded onto SDS-PAGE gels for separation, then transferred to nitrocellulose membranes (Amersham Biosciences). Primary antibodies and HRP-conjugated secondary antibodies (Cell Signaling Technology) were used to detect target proteins. Antibodies used included p38 (Cell Signaling Technology, Cat. No. 8690, 1:1000 dilution), phospho-p38 (Cell Signaling Technology, Cat. No. 9211, 1:1000 dilution), ERK1/2 (Cell Signaling Technology, Cat. No. 4695, 1:1000 dilution), phospho-ERK1/2 (Cell Signaling Technology, Cat. No. 9101, 1:1000 dilution), IκBα (Cell Signaling Technology, Cat. No. 4814, 1:1000 dilution), and β-actin (Cell Signaling Technology, Cat. No. 3700, 1:1000 dilution). Signal was detected using an ECL kit (Thermo Scientific).
Macrophage polarization
THP-1 monocytes were maintained in RPMI1640 with 10% heat inactivated FBS, 1% Penicillin-Streptomycin-Amphotericin B, and 50 μM 2-mercaptoethanol prior to differentiation. The cells were differentiated into macrophages with 150 nM PMA for 48 h. M1 polarization was induced by adding 20 ng/mL IFN-γ and 100 pg/mL LPS for 24 h. M2 polarization was induced by adding 20 ng/mL IL-4 and 20 ng/mL IL-13 for 24 h. In all tests, 10 μM SoL A was added at the same time as M1 and M2 differentiation agents. After 24 h of treatment, total RNA was collected, and RT-qPCR was performed as described above.
Statistics and reproducibility
Statistical tests in the biosynthetic enzyme-disease correlation portion of this study used publicly available data, thus aspects including sample size, data exclusion, randomization, and blinding were not applicable to our re-analysis. For all other experiments, sample sizes were chosen to achieve statistical significance, and data from outliers were excluded based on Grubbs’ test for outliers. For animal experiments, randomization was applied to assign animals to groups and cages and blinding was applied in the analysis of histopathology data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
285,385 human microbial reference genomes were obtained from a previously published collection31. Whole-genome sequencing datasets of human gut microbiomes related to IBD used for biosynthetic enzyme-guided disease correlation were downloaded from the NCBI SRA (SRA Accessions: PRJNA398089 and PRJNA389280). Targeted metabolomic analysis performed in this work was based on the data obtained from two publicly available metabolomics datasets downloaded from IBDMDB and Metabolomics Workbench (Accession: PR000677 [https://doi.org/10.21228/M85H44]). Tables for the processed biosynthetic enzymes and metabolite abundance are available in Supplementary Data 2, 3, and 8. The mass spectrometry data generated in this study for bioactive molecular networking have been deposited in the UCSD CCMS MassIVE database under accession MSV000091884 [https://doi.org/10.25345/C5028PP9T]102. Source data are provided with this paper.
Code availability
The code used for analysis in support of our findings are available in the GitHub repository: https://github.com/ZHANGJianArya/SoL. The code can also be accessed in Zenodo with the https://doi.org/10.5281/zenodo.13896673103.
References
Donia, M. S. & Fischbach, M. A. Small molecules from the human microbiota. Science 349, 1254766 (2015).
Haiser, H. J. & Turnbaugh, P. J. Developing a metagenomic view of xenobiotic metabolism. Pharmacol. Res. 69, 21–31 (2013).
Flint, H. J., Scott, K. P., Duncan, S. H., Louis, P. & Forano, E. Microbial degradation of complex carbohydrates in the gut. Gut. Microbes 3, 289–306 (2012).
Dai, H. et al. Recent advances in gut microbiota-associated natural products: structures, bioactivities, and mechanisms. Nat. Prod. Rep. 40, 1078–1093 (2023).
Lavelle, A. & Sokol, H. Gut microbiota-derived metabolites as key actors in inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 17, 223–237 (2020).
Skelly, A. N., Sato, Y., Kearney, S. & Honda, K. Mining the microbiota for microbial and metabolite-based immunotherapies. Nat. Rev. Immunol. 19, 305–323 (2019).
Spanogiannopoulos, P., Bess, E. N., Carmody, R. N. & Turnbaugh, P. J. The microbial pharmacists within us: a metagenomic view of xenobiotic metabolism. Nat. Rev. Microbiol. 14, 273–287 (2016).
Cao, Y. et al. Commensal microbiota from patients with inflammatory bowel disease produce genotoxic metabolites. Science 378, eabm3233 (2022).
Yao, L. et al. A biosynthetic pathway for the selective sulfonation of steroidal metabolites by human gut bacteria. Nat. Microbiol. 7, 1404–1418 (2022).
Fischbach, M. A. Microbiome: Focus on causation and mechanism. Cell 174, 785–790 (2018).
Chaudhari, S. N., McCurry, M. D. & Devlin, A. S. Chains of evidence from correlations to causal molecules in microbiome-linked diseases. Nat. Chem. Biol. 17, 1046–1056 (2021).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Bäckhed, F. From dietary fiber to host physiology: Short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Chiurchiù, V., Leuti, A. & Maccarrone, M. Bioactive lipids and chronic inflammation: Managing the fire within. Front. Immunol. 9, 38 (2018).
Bae, M. et al. Akkermansia muciniphila phospholipid induces homeostatic immune responses. Nature 608, 168–173 (2022).
MacEyka, M. & Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nature 510, 58–67 (2014).
Walker, A. et al. Sulfonolipids as novel metabolite markers of Alistipes and Odoribacter affected by high-fat diets. Sci. Rep. 7, 11047 (2017).
Pitta, T. P., Leadbetter, E. R. & Godchaux, W. Increase of ornithine amino lipid content in a sulfonolipid-deficient mutant of Cytophaga johnsonae. J. Bacteriol. 171, 952–957 (1989).
Durack, J. & Lynch, S. V. The gut microbiome: relationships with disease and opportunities for therapy. J. Exp. Med. 216, 20–40 (2018).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Dziarski, R., Park, S. Y., Kashyap, D. R., Dowd, S. E. & Gupta, D. Pglyrp-regulated gut microflora Prevotella falsenii, Parabacteroides distasonis and Bacteroides eggerthii enhance and Alistipes finegoldii attenuates colitis in mice. PLOS ONE 11, e0146162 (2016).
Lima, S. F. et al. Transferable immunoglobulin A–coated Odoribacter splanchnicus in responders to fecal microbiota transplantation for ulcerative colitis limits colonic inflammation. Gastroenterology 162, 166–178 (2022).
Hou, L. et al. Identification and biosynthesis of pro-inflammatory sulfonolipids from an opportunistic pathogen Chryseobacterium gleum. ACS Chem. Biol. 17, 1197–1206 (2022).
Pasternak, B. A. et al. Lipopolysaccharide exposure is linked to activation of the acute phase response and growth failure in pediatric Crohnʼs disease and murine colitis. Inflamm. Bowel Dis. 16, 856–869 (2010).
Pastor Rojo, O. et al. Serum lipopolysaccharide-binding protein in endotoxemic patients with inflammatory bowel disease. Inflamm. Bowel Dis. 13, 269–277 (2007).
Im, E., Riegler, F. M., Pothoulakis, C. & Rhee, S. H. Elevated lipopolysaccharide in the colon evokes intestinal inflammation, aggravated in immune modulator-impaired mice. Am. J. Physiol.-Gastrointest. Liver Physiol. 303, G490–G497 (2012).
Stephens, M. & von der Weid, P.-Y. Lipopolysaccharides modulate intestinal epithelial permeability and inflammation in a species-specific manner. Gut. Microbes 11, 421–432 (2020).
Chaleckis, R., Meister, I., Zhang, P. & Wheelock, C. E. Challenges, progress and promises of metabolite annotation for LC–MS-based metabolomics. Curr. Opin. Biotechnol. 55, 44–50 (2019).
Cui, L., Lu, H. & Lee, Y. H. Challenges and emergent solutions for LC-MS/MS based untargeted metabolomics in diseases. Mass Spec. Rev. 37, 772–792 (2018).
Schrimpe-Rutledge, A. C., Codreanu, S. G., Sherrod, S. D. & McLean, J. A. Untargeted metabolomics strategies—challenges and emerging directions. J. Am. Soc. Mass Spectrom. 27, 1897–1905 (2016).
Proctor, L. M. et al. The integrative human microbiome project. Nature 569, 641–648 (2019).
Vatanen, T. et al. Genomic variation and strain-specific functional adaptation in the human gut microbiome during early life. Nat. Microbiol. 4, 470–479 (2019).
Katz, K. et al. The sequence read archive: a decade more of explosive growth. Nucleic Acids Res. 50, D387–D390 (2022).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39, 105–114 (2021).
Liu, Y. et al. Identification and characterization of the biosynthetic pathway of the sulfonolipid capnine. Biochemistry 61, 2861–2869 (2022).
Vences‐Guzmán, M. Á. et al. Identification of the Flavobacterium johnsoniae cysteate‐fatty acyl transferase required for capnine synthesis and for efficient gliding motility. Environ. Microbiol. 23, 2448–2460 (2021).
Radka, C. D., Miller, D. J., Frank, M. W. & Rock, C. O. Biochemical characterization of the first step in sulfonolipid biosynthesis in Alistipes finegoldii. J. Biol. Chem. 298, 1–13 (2022).
Radka, C. D., Frank, M. W., Rock, C. O. & Yao, J. Fatty acid activation and utilization by Alistipes finegoldii, a representative Bacteroidetes resident of the human gut microbiome. Mol. Microbiol. 113, 807–825 (2020).
Kamiyama, T. et al. Sulfobacins A and B, novel von Willebrand factor receptor antagonists: I. Production, isolation, characterization and biological activities. J. Antibiotics 48, 924–928 (1995).
Schirmer, M. et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat. Microbiol. 3, 337–346 (2018).
Franzosa, E. A. et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat. Microbiol. 4, 293–305 (2019).
Tsugawa, H. et al. A lipidome atlas in MS-DIAL 4. Nat. Biotechnol. 38, 1159–1163 (2020).
Guo, J., Shen, S., Xing, S., Yu, H. & Huan, T. ISFrag: De novo recognition of in-source fragments for liquid chromatography–mass spectrometry data. Anal. Chem. 93, 10243–10250 (2021).
Folz, J. et al. Human metabolome variation along the upper intestinal tract. Nat. Metab. 5, 777–788 (2023).
Witting, M. & Böcker, S. Current status of retention time prediction in metabolite identification. J. Sep. Sci. 43, 1746–1754 (2020).
García, C. A., Gil-de-la-Fuente, A., Barbas, C. & Otero, A. Probabilistic metabolite annotation using retention time prediction and meta-learned projections. J. Cheminformatics 14, 33 (2022).
Sun, L. et al. A simple method for HPLC retention time prediction: linear calibration using two reference substances. Chin. Med. 12, 16 (2017).
Wang, Y. et al. A simple method for peak alignment using relative retention time related to an inherent peak in liquid chromatography-mass spectrometry-based metabolomics. J. Chromatographic Sci. 57, 9–16 (2019).
Hale, L. P., Gottfried, M. R. & Swidsinski, A. Piroxicam treatment of IL-10-deficient mice enhances colonic epithelial apoptosis and mucosal exposure to intestinal bacteria. Inflamm. Bowel Dis. 11, 1060–1069 (2005).
Berg, D. J. et al. Rapid development of colitis in NSAID-treated IL-10-deficient mice. Gastroenterology 123, 1527–1542 (2002).
Cario, E. Toll-like receptors in inflammatory bowel diseases: a decade later. Inflamm. Bowel Dis. 16, 1583–1597 (2010).
Nothias, L.-F. et al. Feature-based molecular networking in the GNPS analysis environment. Nat. Methods 17, 905–908 (2020).
Kawasaki, T. & Kawai, T. Toll-like receptor signaling pathways. Front. Immunol. 5, 1–8 (2014).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Maeda, J. et al. Inhibitory effects of sulfobacin B on DNA polymerase and inflammation. Int. J. Mol. Med. 26, 521–527 (2010).
Shimazu, R. et al. MD-2, a molecule that confers lipopolysaccharide responsiveness on Toll-like receptor 4. J. Exp. Med. 189, 1777–1782 (1999).
Miyake, K. Roles for accessory molecules in microbial recognition by Toll-like receptors. J. Endotoxin Res. 12, 195–204 (2006).
Park, B. S. et al. The structural basis of lipopolysaccharide recognition by the TLR4–MD-2 complex. Nature 458, 1191–1195 (2009).
Su, L. et al. Sulfatides are endogenous ligands for the TLR4–MD-2 complex. Proc. Natl Acad. Sci. 118, 1–12 (2021).
Luk, J. M., Kumar, A., Tsang, R. & Staunton, D. Biotinylated lipopolysaccharide binds to endotoxin receptor in endothelial and monocytic cells. Anal. Biochem. 232, 217–224 (1995).
Mosser, D. M. & Edwards, J. P. Exploring the full spectrum of macrophage activation. Nat. Rev. Immunol. 8, 958–969 (2008).
Zhang, Y. et al. ECM1 is an essential factor for the determination of M1 macrophage polarization in IBD in response to LPS stimulation. Proc. Natl Acad. Sci. USA 117, 3083–3092 (2020).
Zhang, X. & Mosser, D. Macrophage activation by endogenous danger signals. J. Pathol. 214, 161–178 (2008).
Parker, B. J., Wearsch, P. A., Veloo, A. C. M. & Rodriguez-Palacios, A. The genus Alistipes: Gut bacteria with emerging implications to inflammation, cancer, and mental health. Front. Immunol. 11, 906 (2020).
Chang, P. V., Hao, L., Offermanns, S. & Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl Acad. Sci. USA 111, 2247–2252 (2014).
Trapecar, M. et al. Gut-liver physiomimetics reveal paradoxical modulation of IBD-related inflammation by short-chain fatty acids. Cell Syst. 10, 223–239.e9 (2020).
Bhattarai, Y. et al. Bacterially derived tryptamine increases mucus release by activating a host receptor in a mouse model of inflammatory bowel disease. iScience 23, 101798 (2020).
Dodd, D. et al. A gut bacterial pathway metabolizes aromatic amino acids into nine circulating metabolites. Nature 551, 648–652 (2017).
Paik, D. et al. Human gut bacteria produce ΤΗ17-modulating bile acid metabolites. Nature 603, 907–912 (2022).
Sato, Y. et al. Novel bile acid biosynthetic pathways are enriched in the microbiome of centenarians. Nature 599, 458–464 (2021).
Li, W. et al. A bacterial bile acid metabolite modulates Treg activity through the nuclear hormone receptor NR4A1. Cell Host Microbe 29, 1366–1377.e9 (2021).
Rakoff-Nahoum, S. & Medzhitov, R. Toll-like receptors and cancer. Nat. Rev. Cancer 9, 57–63 (2009).
Moreira Lopes, T. C., Mosser, D. M. & Gonçalves, R. Macrophage polarization in intestinal inflammation and gut homeostasis. Inflamm. Res. 69, 1163–1172 (2020).
Bhattarai, Y., Muniz Pedrogo, D. A. & Kashyap, P. C. Irritable bowel syndrome: a gut microbiota-related disorder? Am. J. Physiol.-Gastrointest. Liver Physiol. 312, G52–G62 (2016).
Wang, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012).
Thomann, A. K. et al. Depression and fatigue in active IBD from a microbiome perspective-a Bayesian approach to faecal metagenomics. BMC Med. 20, 366 (2022).
Mars, R. A. T. et al. Longitudinal multi-omics reveals subset-specific mechanisms underlying irritable bowel syndrome. Cell 182, 1460–1473.e17 (2020).
Mukherjee, S. et al. Twenty-five years of Genomes OnLine Database (GOLD): data updates and new features in v.9. Nucleic Acids Res. 51, D957–D963 (2023).
Blin, K. et al. antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res. 49, W29–W35 (2021).
Blin, K. et al. The antiSMASH database version 2: a comprehensive resource on secondary metabolite biosynthetic gene clusters. Nucleic Acids Res. 47, D625–D630 (2019).
Terlouw, B. R. et al. MIBiG 3.0: a community-driven effort to annotate experimentally validated biosynthetic gene clusters. Nucleic Acids Res. 51, D603–D610 (2023).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Mirdita, M., Steinegger, M., Breitwieser, F., Söding, J. & Levy Karin, E. Fast and sensitive taxonomic assignment to metagenomic contigs. Bioinformatics 37, 3029–3031 (2021).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
R. Core Team. R: A Language and Environment for Statistical Computing. https://www.R-project.org/ (2021).
Oksanen, J. et al. Vegan: Community Ecology Package. https://CRAN.R-project.org/package=vegan (2020).
Van Rossum, G. & Drake, F. L. Python 3 Reference Manual. (CreateSpace, Scotts Valley, CA, 2009).
McKinney, W. Data structures for statistical computing in python. in Proceedings of the 9th Python in Science Conference 51–56 https://doi.org/10.25080/Majora-92bf1922-00a (2010).
Virtanen, P. et al. SciPy 1.0: Fundamental algorithms for scientific computing in python. Nat. Methods 17, 261–272 (2020).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc.: Ser. B (Methodol.) 57, 289–300 (1995).
Erben, U. et al. A guide to histomorphological evaluation of intestinal inflammation in mouse models. Int J. Clin. Exp. Pathol. 7, 4557–4576 (2014).
Ellermann, M. et al. Endocannabinoids Inhibit the Induction of Virulence in Enteric Pathogens. Cell 183, 650–665.e15 (2020).
Schmid, R. et al. Integrative analysis of multimodal mass spectrometry data in MZmine 3. Nat. Biotechnol. 41, 447–449 (2023).
Nothias, L.-F. et al. Bioactivity-based molecular networking for the discovery of drug leads in natural product bioassay-guided fractionation. J. Nat. Prod. 81, 758–767 (2018).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput Chem. 30, 2785–2791 (2009).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Computational Chem. 31, 455–461 (2010).
Eberhardt, J., Santos-Martins, D., Tillack, A. F. & Forli, S. AutoDock Vina 1.2.0: New docking methods, expanded force field, and python bindings. J. Chem. Inf. Model. 61, 3891–3898 (2021).
Schrödinger, L. L. C. The PyMOL molecular graphics system, version 2.0. (2015).
Li, J. MassIVE MSV000091884 - GNPS - Bioactive molecular networking for sulfonolipids - MZmine Processing for FBMN: This dataset contains the raw data used for MZmine processing and Feature-based molecular networking of semi-purified fractions obtained from A. timonensis used for bioactive molecular networking to reveal contribution of sulfonolipids to observed biological activity. MassIVE https://doi.org/10.25345/C5028PP9T (2023).
Zhang, J. ZHANGJianArya/SoL: initial release for citation. Zenodo https://doi.org/10.5281/ZENODO.13896673 (2024).
Acknowledgements
This work is partially funded by a National Institutes of Health (NIH) grant P20GM103641 awarded to M.N. and J.L., an NIH grant 1R35GM150565 awarded to J.L., a National Science Foundation grant 2239561 awarded to J.L., a Hong Kong Research Grants Council ECS grant HKU27107320 awarded to Y.X.-L., and the Hong Kong Branch of Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou) grant (SMSEGL20SC02) awarded to Y.X.-L. We acknowledge Dr. Kristen Hogan and Dr. Maria Marjorette Pena of the University of South Carolina Center for Colon Cancer Research Mouse Core Facility for assistance with mouse microbiome studies, as well as Dr. Michael Walla of the University of South Carolina Mass Spectrometry Facility for assistance in acquiring HRMS and HRMS/MS data. We also acknowledge Emily Quinn, Dorathea Lee, and Andrew Campbell for assistance with SoL purification.
Author information
Authors and Affiliations
Contributions
E.A.O., J.Z., Y.-X.L., and J.L. designed the study; E.A.O., J.Z., Z.E.F., D.X., Z.Z., M.K.M, M.M., and Y.W. collected the data; E.A.O., J.Z., H.C., D.F., M.E., Y.-X.L., and J.L. analyzed the data; P.N., M.N., Y.-X.L., and J.L. funded the study; E.A.O., J.Z., Y.-X.L., and J.L. prepared the manuscript; All authors participated in revising the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Alexander Rodriguez-Palacios, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Older, E.A., Zhang, J., Ferris, Z.E. et al. Biosynthetic enzyme analysis identifies a protective role for TLR4-acting gut microbial sulfonolipids in inflammatory bowel disease. Nat Commun 15, 9371 (2024). https://doi.org/10.1038/s41467-024-53670-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53670-y
This article is cited by
-
Genetics-mediated regulation of intestinal gene expression on microbiome contributes to human disease heritability
Molecular Systems Biology (2025)