Abstract
High crop yields require adequate nutrients, particularly nitrate (N) and phosphorus (P). Identifying regulators for efficient N-P utilization is critical in wheat. To explore N-P interactions, we analyze root transcriptomes under varying N-P supplies and identify TaTCP6 as a potential regulator. Nitrate-stimulated TaTCP6 directly triggers the expression of genes related to nitrogen utilization. TaTCP6 competes with TaSPX1/4 for the release of TaPHR2, and also interacts with TaPHR2 to enhance the transactivation capacity of downstream genes. Thus, through the dual roles of TaTCP6, the TCP6-SPX-PHR2 module activates the expression of phosphorus starvation response (PSR) genes. Inhibiting TaTCP6 reduces N and P absorption, negatively impacting yield, while overexpressing TaTCP6 increases grain yield. Notably, overexpression of TaSPX1 suppresses nitrogen utilization genes, especially under low phosphorus conditions. In conclusion, our findings highlight the role of TaTCP6 in coordinating N and P utilization and propose a strategy to reduce fertilizer inputs for sustainable agriculture.
Similar content being viewed by others
Introduction
Nutrients are critical for life through intertwined sensing, signaling, and metabolic processes. Plants have evolved sophisticated systems to respond and adapt to fluctuating nutrients availability from scarcity to abundant concentrations in the soil. Nitrate (N) and phosphorus (P) are the primary macronutrients, which can be metabolized into various organic compounds essential for numerous biochemical processes. Research in model plants has shown that nutrient acquisition is prioritized. For instance, phosphorus starvation responses (PSRs) are not activated until nitrate is available. Thus, it is crucial to investigate how crops efficiently coordinate N–P utilization in complex environments.
Previous researches have provided pieces of evidence that nitrate signaling is critical for PSRs. In Arabidopsis, the nitrate-NRT1.1-NLP7-NIGT1 cascade suppresses PHO2 expression to stabilize PHO1 and PHT1 proteins1,2,3. Recently, the nitrate-OsNRT1.1B-OsSPX4 cascade was proposed to integrate N and P signaling networks in rice4. AtNRT1.1/CHL1/AtNPF6.3 senses nitrate by the phosphorylation of Thr101, and its homolog in rice OsNRT1.1B shares similar competences. The NIN-like protein (NLP) AtNLP7 is regulated by nitrate concentration to enter the nucleus, a process that is subject to the calcium spiking and calcium-dependent protein kinases5. Recent studies have shown that AtNLP7 acts as an intracellular receptor and can directly bind nitrate ions, essential for its derepression6. AtPHR1 and OsPHR2 are the primary regulators of PSRs in Arabidopsis and rice, respectively7,8,9. Under P-replete conditions, the SPX protein inhibits the nuclear entry of PHR and its ability to bind DNA through interacting with PHR, ultimately preventing PHR from initiating PSRs10,11,12. Upon nitrate perception, OsNRT1.1B promotes the E3 ubiquitin ligase OsNBIP1 to degrade OsSPX4, thus releasing OsNLP3 and OsPHR2 to activate nitrate- and phosphate-responsive genes, respectively. AtNIGT1 and its analogs were strongly induced by nitrate or phosphorus starvation, but this stimulation was considerably attenuated in Atnlp7 and Atphr1 mutants. Conversely, AtNIGT1 facilitated phosphorus acquisition while impeding nitrate acquisition, thereby coordinating nutrient homeostasis in a complex N–P environment2,13,14,15,16,17,18,19. The GARP transcription factors AtGLK1/2 were reported to be induced by nitrate and could enhance the functionality of AtPHR1 in PSRs20. Among the members in the SPX family, only the SPX4 is not induced by phosphate deficiency. OsSPX4 primarily localizes to the cytoplasm and perimembrane and could be targeted and degraded by the NRT1.1B-NBIP1 complex. In contrast, OsSPX1 predominantly resides in the nucleus21,22. The distinct expression patterns and localization of SPX1 and SPX4 suggest the existence of additional and diverse mechanisms of nutrient regulation23,24. It is noteworthy that the bulk of these studies have been conducted in rice or Arabidopsis, with very limited investigation in wheat.
TCPs (TEOSINTE BRANCHED1/CINCINNATA/PROLIFERATING CELL FACTORs) are a group of ancient plant-specific transcription factors that can be divided into two subfamilies: Class I (PCF or TCP-P) and Class II (TCP-C). They play crucial roles in regulating diverse physiological processes across the life cycle, such as plant development and hormonal signaling pathways. In Arabidopsis, AtTCP20 and AtNLP6/7 form heterodimers to regulate the genes involved in nitrate assimilation and root meristem activity during N starvation25,26,27. In rice, OsTCP19 was repressed by nitrate treatment and negatively regulated tiller formation. The haplotype OsTCP19-H, with a 29-bp deletion in the upstream regulatory region, enhanced the response of tiller number to nitrate, that could be employed to produce more tillers in low or medium nitrate environments28.
Bread wheat (Triticum aestivum L.) is an allohexaploid (AABBDD, 2n = 6x = 42) and possesses a larger number of TCP genes compared to diploid plant species. In a recent study, over 66 TCPs were identified in the entire wheat genome29. In this study, we conducted transcriptome analysis using varying N–P supplies, revealing that nitrate promotes balanced utilization of N and P through the activation of nitrate and phosphate signaling pathways. Then, we identified a nitrate-induced transcription factor TaTCP6 as a regulator of nitrogen utilization. TaTCP6 competes with TaSPX1/4 for the release of TaPHR2 and also interacts with TaPHR2 to enhance its transactivation capacity. Thus, through the dual roles of TaTCP6, the TCP6-SPX-PHR2 module activates the expression of PSR genes. Importantly, overexpression of TaSPX1 suppressed the expression of nitrogen utilization genes, particularly in low-phosphorus conditions. These results demonstrate that the SPX family coordinates the utilization of N–P under low-phosphorus environments, reminiscent of the NIGT transcription factors. In addition, TaTCP19 might act as an antagonist to TaTCP6, negatively regulating N–P utilization. The functional and molecular characteristics of OsTCP6, the TaTCP6 homolog in rice, suggest that this mechanism might be conserved in other crops.
Results
Nitrate activates both N and P-use-related genes for a coordinated utilization of N and P
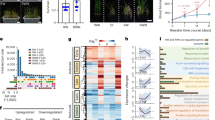
Plants frequently encounter multiple nutrient deficiencies in the soil. To explore the coordinated regulation of N and P in wheat, we cultivated seedlings in hydroponic conditions with different N–P supply combinations: high nitrate (HN, 2.5 mM KNO3); low nitrate (LN, 0.25 mM KNO3); high phosphate (HP, 0.3 mM KH2PO4); low phosphate (LP, 0.03 mM KH2PO4). Four weeks after treatment, we measured the length of the roots and shoots, as well as the biomass of the roots and shoots, and calculated the shoot/root ratio (Fig. 1a, b and Supplementary Fig. 1a). The significant and gradual increase in root biomass under LN and/or LP conditions indicates the efficacy of the treatment. Under LN conditions, the application of high phosphate had no significant effect on the total biomass of wheat; however, under HN conditions, it resulted in a remarkable increase in the total biomass (Fig. 1a, b). These findings demonstrate that the impact of increased phosphate supply on biomass is more pronounced when the nitrogen is sufficient.
a The phenotype of Fielder under different N and P conditions for 4 weeks. High nitrate (HN), 2.5 mM KNO3; low nitrate (LN), 0.25 mM KNO3; high phosphate (HP), 0.3 mM NaH2PO4; low phosphate (LP), 0.03 mM NaH2PO4. Scale bars, 10 cm. b Biomass analysis of wheat plants under different N and P conditions (5 wheat seedlings as a sample) (n = 12 biologically independent samples). Boxes represent the median values and the first and third quartiles; whiskers represent the minimum and maximum values. P values are from two-sided Student’s t-test. Exact P values and source data are provided as a Source Data file. c Venn diagram comparing genes differentially expressed in response to P treatment under LN and HN conditions. d Venn diagram comparing genes differentially expressed in response to N treatment under LP and HP conditions. Heatmap showing relative expression levels of NRG (Nitrate-related genes; e) and PRG (Phosphate-related genes; f), listed in Supplementary Data 1, in Fielder under different N and P conditions. (color codes in EB6223 to 2C7CC1). Heatmap showing relative expression levels of several nitrogen utilization genes (g) and PSR genes (h) in Fielder under different N and P conditions.
To further investigate the mechanism behind this phenomenon, we performed transcriptome sequencing of wheat roots cultured under different N–P conditions (Fig.1a, Supplementary Data 1). A total of 4710 and 149 differentially expressed genes (DEGs) induced by LP were identified under HN and LN conditions, respectively (Fig. 1c). The considerably higher count (>30-fold) suggests that a robust PSR necessitates a sufficient supply of nitrogen. We also observed a substantial decrease in the quantity of DEGs triggered by nitrate treatment under LP conditions (5251) in contrast to HP conditions (8864) (Fig. 1d). Only 46% (4077/8864) of the N-related DEGs under HP conditions were still stimulated under LP conditions. These results suggest that LP attenuates the nitrate signaling pathway. To obtain a comprehensive depiction of N–P interaction, we initially defined the genes related to N (n = 823) and P (n = 195) in wheat through analysis of the covariance of homologous genes and protein sequence comparison (Supplementary Data 2 and 3). The genes were categorized into four groups, including nitrate uptake, nitrate assimilation, phosphate uptake, and phosphate regulation. The expression patterns of the PSR and the nitrogen utilization genes illustrate a tight interdependency between N and P (Fig. 1e, f). Specifically, we observed a greater number and significantly higher expression of nitrogen utilization genes in HNHP compared to HNLP (Fig. 1g). The genes responsible for ammonium, amino acid, and small peptide transport are predominantly expressed under LN conditions (Supplementary Fig. 1b). Similarly, HN is the prerequisite for PSR genes in response to LP conditions, and TaSPX4 was the sole member in SPX family that was not influenced by phosphorus levels (Fig. 1h). The loss of responsiveness in LN conditions suggests that key regulators may be involved in both nitrate and phosphorus uptake. Interestingly, a group of genes related to potassium (K, Supplementary Data 4) and iron (Fe, Supplementary Data 5) uptake showed significant response to varying levels of nitrate and phosphate, indicating a broad interplay between different nutrients (Supplementary Fig. 1b). The majority of the genes involved in K uptake were downregulated by nitrate treatment, regardless of the phosphate levels in wheat. Previous studies in rice attempted to substitute long-term nitrate treatment with short-term nitrate induction4. The qPCR results for the sentinel genes indicated that nitrate stimulates the expression of nitrogen utilization genes predominantly after 6 h in wheat (Supplementary Fig. 1c). Thus, we also conducted transcriptome analysis after 1 and 6 h of nitrate induction, and identified 6523 genes as N-induced genes. Through comparison, we found that 1522 genes responded to both short- and long-term nitrate treatment (Supplementary Fig. 1d). Taken together, our results demonstrate that nitrate can directly activate both nitrate and phosphate signaling pathways to achieve a balanced N–P utilization.
The nitrate-stimulated TaTCP6 positively regulates nitrate uptake and utilization
Among the structural genes for nitrogen uptake and utilization, TaNRT2.1-6B showed the most significant induction by nitrate (Supplementary Fig. 1c). In addition, the expression of TaNRT2.1 is notably influenced by P levels (Fig. 1g), indicating the existence of core cis-elements in the promoter of TaNRT2.1 for N and/or P regulations. To explore the regulatory factors involved in nitrate-enhanced utilization of N and P, the promoter of TaNRT2.1 was utilized as bait for Y1H screening, resulting in the identification of a group of TCP family transcription factors including TaTCP6 (TraesCS3B02G470700, with homoeologs TraesCS3A02G436100 and TraesCS3D02G428700) (Supplementary Data 6 and 7). The expression profile of the Class I TCP family in wheat revealed that TaTCP6 is the only gene displaying a similar expression pattern to TaNRT2.1, with the highest expression under HNHP conditions and a significant induction by nitrate (Fig. 2a). The direct interaction between TaTCP6 and the promoter of TaNRT2.1 was confirmed by Y1H and EMSA assays (Fig. 2b, c, k). Nitrate stimulated the transcription of TaTCP6, peaking at 4–6 h after treatment (Fig. 2d). OsTCP6 (Os01g0924400), the homologs of TaTCP6 in rice, was also transcriptionally induced by nitrate (Supplementary Fig. 2a, b). In consistence, the protein abundance of TaTCP6 significantly increased under HN conditions, as demonstrated by assays in wheat protoplasts (Fig. 2e). TaTCP6 is highly expressed in root cells (Supplementary Fig. 3a) and the protein was mainly localized in the nucleus (Supplementary Fig. 2c). To construct the networks of TaTCP6, we conducted Y2H library screening using TaTCP6 as bait. NLPs were identified as interactors of TaTCP6 (Supplementary Data 8). The interaction between TaTCP6 and TaNLP4 was validated through firefly luciferase complementation imaging (LCI) assays (Supplementary Fig. 5c).
a Expression heatmap of Class I TCP transcription factors under different N and P conditions and N induce treatment. b The schematic diagram of TaNRT2.1-6B4 (TraesCS6B02G044300) promoter. c Yeast one-hybrid analysis of TaTCP6-B binding to the TaNRT2.1-6B4 promoter. d RT-qPCR analysis of short-term induction of TaTCP6 in the roots of wheat seedlings under 0.2 mM and 2.5 mM nitrate solution, respectively. KCl was used as the negative control. Error bars represent the mean ± SD (n = 3). e Protein accumulation of TaTCP6 under the control of its native promotor following nitrate induction in wheat protoplasts. Scale bars, 10 μm. f Comparison of Seedling height among TaTCP6-SRDX-1, TaTCP6-SRDX-2, and Fielder control under HN or LN conditions. Data are mean ± SD (n = 3). g 15N uptake rate in seedlings of TaTCP6-SRDX-1, TaTCP6-SRDX-2, TaTCP6-CR-1 and Fielder. Data are mean ± SD (n = 3). h Transcription levels of TaNRT1.1B, TaNRT2.1-6B4, TaNIA1, and TaNIR in the roots of the Fielder, TaTCP6-OE-2 and TaTCP6-OE-4. Data are mean ± SD (n = 3). i Accumulation of TaTCP6-FLAG in the roots of TaTCP6 overexpressing transgenic wheat seedings (TaTCP6-OE-2) under different nitrate conditions. j Analysis of transcriptional activities of TaTCP6 on the promoters of TaNRT1.1B-1A2, TaNRT2.1-6B4, TaNIA1-6B1, and TaNIR-6B1 in wheat protoplasts. Data are mean ± SD (n = 6). k EMSA of TaTCP6 binding to the promoters of TaNRT1.1B-1A2, TaNRT2.1-6B4, TaNIA1-6B1, and TaNIR-6B1. These experiments in (e, i) were repeated three times independently with similar results. In (f–h, j), P values are from two-sided Student’s t-tests. Exact P values and source data are provided as a Source Data file.
To further investigate the function of TaTCP6, we generated overexpression lines (TaTCP6-OE; Supplementary Fig. 4c) as well as knockout lines (TaTCP6-CR) (Supplementary Fig. 4a). To avoid functional redundancy among TaTCPs, we also generated two transgenic lines pTaTCP6::TaTCP6-SRDX where TaTCP6 was fused with SRDX repression domain under the control of its native promoter. In addition, two knockout lines in rice, Ostcp6-1 and Ostcp6-2, were utilized in our study (Supplementary Fig. 4b). The TaTCP6-SRDX showed a more severe phenotype under HN conditions compared to LN conditions, suggesting that the function of TaTCP6 is dependent on N levels in wheat (Fig. 2f). Moreover, the protein abundance of TaTCP6 in Ubi::TaTCP6-Flag transgenic plants gradually increased with higher nitrate concentrations (Fig. 2i). This suggests that nitrate enhances the stability of TaTCP6 in a post-translational manner. To study the role of TaTCP6 in nitrate utilization, we first examined the expression levels of the genes related to nitrate uptake and assimilation. The results showed that the majority of these genes were significantly upregulated in TaTCP6 overexpression lines TaTCP6-OE (Fig. 2h). The nitrate uptake ability was significantly reduced in the seedlings of TaTCP6-SRDX (Fig. 2g). Transient luciferase activity assays indicated significant activation of LUC by TaTCP6 driven by the promoters of TaNRT1.1, TaNIA1 and TaNIR1 (Fig. 2j). EMSA assays revealed direct binding of TaTCP6 to the promoters of these genes (Fig. 2k; Supplementary Fig. 5a). Collectively, our data suggest that TaTCP6 is responsive to nitrate levels and positively regulates nitrate uptake and utilization.
To further clarify the effect of TaTCP6 on N uptake and utilization in wheat, we measured N use-related traits in TaTCP6 transgenic lines. The nitrate absorption in Mock and TaTCP6 overexpression lines was examined by using 15N-KNO3 as the nitrogen source at 0.25 and 2.5 mM. The accumulation of 15N in TaTCP6 overexpression lines was remarkably higher than that of Mock (Supplementary Fig. 16c, d). Moreover, we also analyzed the enzymatic activities of NR, the content of NO3- and Pi in shoots of Mock and TaTCP6 transgenic plants grown under HNHP, HNLP, LNHP, and LNLP conditions. Compared with Mock, TaTCP6 overexpression lines exhibited higher enzyme activities of NR under all four conditions, while the TaTCP6-SRDX lines displayed the opposite phenotypes (Supplementary Fig. 16e). The nitrate and Pi content in shoots showed that compared with Mock, TaTCP6 overexpression lines accumulated more nitrate and Pi, while the TaTCP6-SRDX lines accumulated less (Supplementary Fig. 16f, g). Moreover, these negative effects of TaTCP6-SRDX lines were weakened under the LN condition compared with the HN condition (Supplementary Fig. 16e–g). Interestingly, the nitrogen content of wheat plants under HN condition is significantly higher than that under low nitrogen conditions, but high and low nitrate treatments appeared to have subtle effect on the carbon content of wheat (Supplementary Fig. 16a, b). TaTCP6 overexpression lines significantly increased root nitrogen content but reduced shoot nitrogen content under high nitrogen conditions, whereas shoot nitrogen content was significantly increased under LN condition (Supplementary Fig. 16b). These results suggest that TaTCP6 may also be involved in nitrogen nutrient partitioning under different nitrogen conditions.
OsTCP19 has been well-studied as a negative regulator of tillering in response to external nitrate supply. However, its role in N–P uptake and utilization has received little attention, despite the significantly decreased expressions of OsNRT2.1, OsNRT2.2, OsNIA1, and OsPht1;2 in OsTCP19-OE lines28. Herein, we discovered that TaTCP19, the homolog of OsTCP19 in wheat (Supplementary Fig. 13c), exhibited a tissue-specific expression pattern similar to TaTCP6 (Supplementary Fig. 3a). TaTCP19 could remarkably inhibit the expression of LUC reporter genes driven by the promoters of TaNRT1.1B, TaNIA1 and TaNIR (Supplementary Fig. 13b). Further experiments showed that the expression of TaTCP19 was also regulated by nitrate concentration and reduced upon nitrate induction (Supplementary Fig. 2d). These results indicate that TaTCP19 might function antagonistically to TaTCP6 in the regulation of balanced N–P utilization in wheat.
TCP6 regulates key agronomic traits in both wheat and rice
The uptake and utilization of nutrients by crops are important determinants of crop yield. Field experiments were conducted to determine whether TaTCP6 and its homolog OsTCP6 in rice contribute to field yield (Beijing, 2023). The overexpression of TaTCP6 leads to a significant increase in plant height (PH) and thousand-grain weight (TGW), whereas these traits were significantly reduced in TaTCP6-SRDX lines (Fig. 3a–c). Though no statistical difference in PH was observed between WT and TaTCP6-CR-1, the TGW of TaTCP6-CR-1 was still significantly decreased (Fig. 3a–c). The TaTCP6-SRDX lines showed severely impaired spike development with reduced spike length (SL) and number of number of spikelets per spike per spike (Supplementary Fig. 6a, b). Tiller number (TN) was significantly increased in TaTCP6-OE but reduced in TaTCP6-SRDX lines (Supplementary Fig. 6b). TaTCP6 manipulation not only impacted grain development (Supplementary Fig. 7), but also influenced the nitrogen and phosphorus content in grains (Supplementary Fig. 6c). Similarly, rice mutants Ostcp6 exhibited significant decreases in PH, the number of primary branching peduncles, the number of grains per spike (GPS), grain width and TGW in comparison to the control ZH11 (Fig. 3d–g; Supplementary Fig. 6d, e). The growth and developmental abnormalities in mutants resulted in a substantial decrease in rice yield per panicle and per plant (Fig. 3d–g; Supplementary Fig. 6d, e). These findings indicate that TaTCP6 and its homolog in rice are essential for crop yield.
a Wheat plants at the heading stage with the indicated genotypes. Scale bars, 20 cm. b Grain shape of wheat seeds with the indicated genotypes. Scale bars, 1 cm. c Analysis of plant height, grain length, grain width, and 1000-grain weight with the indicated genotypes. Data are mean ± SD (n = 20 for plant height analysis and 10 for grain traits analysis). d Rice plants at the mature stage with the indicated genotypes. Scale bars, 40 cm. e Rice panicle at the mature stage with the indicated genotypes. Scale bars, 10 cm. f Grain shape of rice seeds with the indicated genotypes. Scale bars, 1 cm. g Analysis of plant height, primary branches per panicle, number of grains per panicle, grain width, 1000-grain weight, and grain yield per plant with the indicated genotypes. Data are mean ± SD (n = 12). In (c, g), P values are from two-sided Student’s t-tests. Exact P values and source data are provided as a Source Data file.
Identification of the target genes of TaTCP6
To better understand the mechanism by which TaTCP6 promotes nutrition utilization, transcriptome analysis was conducted on five lines overexpressing TaTCP6 with varying levels (Supplementary Fig. 4c): low (TaTCP6-OE-2), medium (TaTCP6-OE-3 and TaTCP6-OE-4), and high (TaTCP6-OE-5 and TaTCP6-OE-6). In comparison to the WT, different numbers of DEGs were detected in the five lines (Supplementary Data 9). Although the minimum number (883) of DEGs were identified in TaTCP6-OE-2, Gene Ontology (GO) analysis showed that these genes were significantly enriched in nitrogen and phosphorus utilization pathways (Supplementary Fig. 5d). Further analysis was conducted on TaTCP6-OE-3/4/5/6 with a higher number of DEGs. A total of 11,818 DEGs were identified in at least three lines, with 8421 DEGs overlapping in these four lines (Fig. 4a). Among them, 554 DEGs were nitrogen-regulated genes (Fig. 4b). Next, CUT&Tag assays were conducted, followed by high-throughput sequencing, to identify the TaTCP6 DNA-binding sites on the wheat genome. A total of 85,072 peaks were identified, which were mapped to 11,813 annotated genes (Supplementary Data 10). Density plots of TaTCP6 DNA-binding sites show a wide range of binding signals on gene body regions (Fig. 4c). The motif analysis of TaTCP6 CUT&Tag peaks revealed a marked enrichment of the GTGGGNCCCACC motif which is highly consistent with previously reported TBS (Fig. 4d). A genome distribution analysis revealed that these peaks were highly enriched in intergenic regions, accounting for 78.98% of all peaks. Notably, 11.97% of peaks (7764 genes) were located in putative promoter regions 2 kb upstream of transcription start sites (Fig. 4e). Among the genes directly bound by TaTCP6, 1,889 genes (16.0%) were differentially expressed between Fielder and TaTCP6-OE, of which 942 and 947 genes were upregulated and downregulated, respectively (Fig. 4f). GO analysis indicated that these TaTCP6 target genes were enriched in biological processes related to iron transport, response to hormone, seedling development, response to glucose and response to nitrate (Fig. 4g). Specifically, Integrative Genomics Viewer showed the specific TaTCP6-binding peaks on the promoter regions of genes related to nitrogen utilization such as TaNRT2.1-6B4, TaNRT1.1B, TaNIA1-6B1, TaNIR-6B1, TaGS-4A1, and TaGOGAT-3A1 (Fig. 4h). It is noteworthy that several key regulatory genes for nitrogen utilization and plant growth like TaNLP3-3D, TaNLP4-5D, TaNAC2-3B30, TaZnF-4B31, TaSPX4-5D, and TaBZR2-3A, also exhibited specific TaTCP6-binding peaks in their promoters (Fig. 4i). Electrophoretic mobility shift assays (EMSA) were performed to confirm the direct binding of TaTCP6 to the promoters of TaNAC2-3B, TaNLP4-5D, and TaSPX4-5D in vitro (Fig. 4j). Importantly, these genes were all upregulated in TaTCP6 overexpression lines (Fig. 4k). These results suggest that TaTCP6 controlled nitrate uptake, assimilation and wheat development by directly regulating the related essential genes.
a Venn diagram showing the numbers of DEGs in the roots of 2-week-old wheat seedlings between four TaTCP6-OE lines and Mock. b Venn diagram showing the overlap of DEGs of N induces in the whole plants of 2-week-old wheat seedlings, HN/LN HP, and TaTCP6-OE/Mock in the roots of 2-week-old wheat seedlings. c A density plot of the putative TaTCP6-binding peaks located within 2 kb upstream of the TSS (Transcription start sites) and 2 kb downstream of the TES (Transcription end sites). d A putative TaTCP6-binding motif was identified using only the promoter regions of the binding peaks from the CUT&Tag data identified by DREME. e Pie chart showing the distribution of annotated TaTCP6-binding peaks in wheat genome relative to nearby genomic features including TSS, exons, introns, and untranslated regions. f Venn diagram showing the overlap of genes associated with TaTCP6-binding sites and the DEGs identified by RNA-seq in TaTCP6-OE-3/4/5/6. g Gene Ontology (GO) terms enriched in the genes associated with TaTCP6-binding sites. h Chromatin binding peaks of TaTCP6 at the genomic regions of TaNRT2.1-6B4, TaNRT1.1B-1D, TaNIA-6B1, TaNIR-6D1, TaGS-4A1, and TaGOGAT-3A1 genes. CUT&Tag reads were visualized using the Integrative Genomics Viewer. The control is shown in green and TaTCP6 in blue. i Chromatin binding peaks of TaTCP6 at the genomic regions of TaNLP3-3D, TaNLP4-5D, TaNAC2-3B, TaZnF-4B, TaSPX4-5D, and TaBZR2-3A genes. CUT&Tag reads were visualized using the Integrative Genomics Viewer. The control is shown in green and TaTCP6 in blue. j EMSA of TaTCP6 binding to the promoters of TaNAC2-3B, TaNLP4-5D, and TaSPX4-5D. k Heatmaps showing the relative expression changes (z-normalized) of nitrogen-related plant development genes determined by RNA-seq in TaTCP6-OE. The values used to generate the heatmaps are the average FPKMs of two independent RNA-seq replicates.
TaTCP6 plays a pivotal role in phosphate uptake and utilization in wheat
Based on the repressed expression of OsPht1;2 in OsTCP19-OE rice plants28 and the effect of TaTCP6 on phosphorus content in the grains of TaTCP6 transgenic plants (Supplementary Fig. 6c), we assumed that TaTCP6 may be also involved in phosphorus uptake and utilization. The enhanced expression of genes involved in phosphorus uptake and utilization, such as TaPAP10c-5A, TaPHF-5A, and TaSPX1 (Fig. 5a), in addition to the increased Pi content of TaTCP6 overexpression lines (Fig. 5b), further suggested that TaTCP6 plays important roles in phosphorus usage. Knocking out OsTCP6 also weakened the phosphate uptake rate of rice (Supplementary Fig. 5e). Furthermore, our Y2H screening using TaTCP6 as the bait identified multiple GARP family transcription factors (TFs), including TaPHR2 (Supplementary Data 8), a key transcriptional regulator of phosphate starvation responses. GARP TFs contain a unique MYB-related domain, with a consensus sequence (SHLQ(K/M)(Y/F)), and play a crucial role in nutritional and hormonal responses32. Protein interaction between TaTCP6 and truncated variants of several typical GARP TFs, including TaKAN1, TaMYR1 TaGLK1, and TaNIGT133, proved that TaTCP6 interacts with the MYB domains of these TFs (Fig. 5c–e). It was further verified by bimolecular fluorescence complementation (BiFC), co-immunoprecipitation (Co-IP), pull-down, and LCI that TaTCP6 interacts with TaRLI1 and TaPHR2 in the nucleus (Fig. 5f–i).
a Heatmaps showing the relative expression changes (z-normalized) of genes mediating phosphorus uptake and utilization determined by RNA-seq in TaTCP6-OE. The values used to generate the heatmaps are the average FPKMs of two independent RNA-seq replicates. b Pi contents of Mock and TaTCP6-OE wheat plants. wheat seedlings grown for 2 weeks were sampled for analysis Pi contents. Data are mean ± SD (n = 6); P values are from two-sided Student’s t-tests. Exact P values and source data are provided as a Source Data file. c Yeast two-hybrid (Y2H) analysis of protein interaction between TaKAN1, TaMYR1, TaCLK1, TaNIGT1, and TaTCP6. TaTCP6TCP only contains the TCP domain (Pfam: PF03634). d A schematic diagram of the protein domain structure of several GARP family transcription factors. e–i Yeast two-hybrid analysis of protein interaction domain between TaRLI1, TaPHR2, and TaTCP6. Interaction between TaRLI1, TaPHR2, and TaTCP6 confirmed by Y2H (e), bimolecular fluorescence complementation (BiFC; f), firefly luciferase complementation imaging (LCI; g), pull-down (h), and co-immunoprecipitation (Co-IP; i) assays. Scale bars, 10 μm. These experiments in (h, i) were repeated three times independently with similar results.
TaTCP6 competes with TaSPXs to interrupt TaPHR2-TaSPXs interaction
Since SPX domain-containing proteins play crucial roles in the repression of PSR and were identified through Y2H screening, we examined the interactions between TaTCP6 and TaSPXs. TaSPX1, TaSPX3, TaSPX4, and TaSPX6 were isolated based on the phylogenetic analysis of the SPX family in wheat, rice and Arabidopsis (Supplementary Fig. 9a). The protein stability of TaSPXs is negatively correlated with nitrate status, with the exception of TaSPX1 (Supplementary Fig. 9b). We also found that the interaction between SPX domain and RLI1 is conserved in wheat and rice (Supplementary Fig. 9c–e). The subcellular localization analysis indicated that TaSPX1 is exclusively located in the nucleus, TaSPX4 in the cytoplasm and around membranes and other SPXs distributed in both the nucleus and cytoplasm (Supplementary Fig. 10a, b). We discovered that TaTCP6 could interact with all of the TaSPX proteins, with TaPHR2 and TaRLI1 serving as positive controls in BiFC assays (Fig. 6a). Interestingly, the interaction between TaTCP6 with TaSPXs was predominately confined to the nucleus, while the interactions between TaSPXs with TaPHR2 and TaRLI1 were influenced by the subcellular localization of TaSPXs (Fig. 6a; Supplementary Fig. 10a–c). Co-IP, pull-down, and LCI further confirmed that TaTCP6 was able to interact with TaSPX1 and TaSPX4 (Fig. 6b–d). The interaction between OsSPXs-OsPHR2 has been proven to be dependent on phosphate levels10,21,22,34. Here, we found that the interactions between TaTCP6 and TaSPX1 as well as TaSPX4 are phosphate-independent, as demonstrated by the in vitro pull-down assay (Fig. 6c). Given that both TaTCP6 and TaSPXs interact with TaPHR2, we postulated that TaTCP6 competes with TaSPXs to free TaPHR2 and enable it to perform its function. To test this hypothesis, we conducted experiments to examine the impact of TaTCP6 on TaPHR2 in the presence of SPXs. The transient expression assays in wheat protoplasts showed that TaSPX4 could sequester TaPHR2 in the cytoplasm. However, the presence of TaTCP6 leads to the release of TaPHR2 into the nucleus (Fig. 6e). In N. benthamiana, the fluorescence signals were strongest in the presence of TaPHR2-nLUC and cLUC-TaSPX1 alone, and diminished significantly as the amount of TaTCP6 increased (Fig. 6f). In addition, the Co-IP assays showed that the addition of either TaTCP6 or TaPHR2 diminished the interaction of TaSPX1 with the other one (Supplementary Fig. 11a). These results indicated that TaTCP6 disrupts the interaction between TaPHR2 and TaSPX1, promoting the shuttling of TaPHR2 to the nucleus. Consistently, the in situ hybridization results show a similar mRNA expression pattern of TaTCP6, TaTCP19, TaPHR2, and TaSPX1 in the wheat root (Supplementary Fig. 3b). The interaction of TaTCP19 with TaPHR2 and TaSPX1/4 were also verified by LCI, which showed a strong interaction between TaTCP19 with TaPHR2, but a very weak or no interaction with TaSPX1/4 (Supplementary Fig. 11b–d).
a BiFC assays of TaTCP6, TaPHR2, and TaRLI1 with TaSPX1, TaSPX3, TaSPX4, and TaSPX6 in wheat protoplasts. Scale bars, 10 μm. b Co-IP assays of TaTCP6 with TaPHR2, TaRLI1, TaSPX1, and TaSPX4. c The in vitro pull-down assays of TaSPX1/4 with TaTCP6, TaPHR2, and TaRLI1 without additional phosphate or InsP6/8 treatments. d LCI assays of TaTCP6 with TaSPX1 and TaSPX4 in N. benthamiana leaves. e The effect of TaSPX4 and TaTCP6 on the fluorescence intensity of TaPHR2-GFP in wheat protoplasts. TaPHR2-GFP was co-transformed with EV (mCherry), EV, and TaSPX4-mCherry or TaTCP6-FLAG and TaSPX4. Error bars represent the mean ± SD (n = 16). Scale bars, 20 μm. f The effect of TaTCP6 on the interaction between TaPHR2 and TaSPX1 in LCI assays. The ratios of different Agrobacterium strains containing the indicated constructs are shown in brackets. The pseudocolor bar indicates the range of luminescence intensity. Error bars represent the mean ± SD (n = 5). These experiments in (b, c) were repeated three times independently with similar results. In (e, f), different letters above the bars indicate statistically significant differences (adjusted P < 0.05, one-way ANOVA with Tukey’s test). Exact P values and source data are provided as a Source Data file.
TaTCP6 interacts with and enhances TaPHR2 activity to initiate PSR
To determine if the interaction with TaTCP6 impacts the transcriptional activation ability of TaPHR2, transient expression assays were performed in wheat protoplasts. It turned out that TaPHR2 exhibited high transactivation activity towards the promoters of PSR genes (Fig. 7a and Supplementary Fig. 12a). Importantly, the co-expression with TaTCP6 significantly enhanced this transactivation activity (Fig. 7a). The EMSA experiments indicate that TaTCP6 did not affect the DNA-binding ability of TaPHR2, nor did TaPHR2 affect the binding of TaTCP6 to the TBS motif. (Supplementary Fig. 12b, c). In addition, TaTCP6 significantly restored the compromised transcriptional activation activity of TaPHR2 caused by TaSPX1 (Fig. 7b, c). We further demonstrated that TaTCP19, TaPCF3, and TaTCP9 have varying effects on the transcriptional activation ability of TaPHR2. In particular, TaTCP19 exhibited a pronounced inhibitory effect, as opposed to TaTCP6 (Fig. 7b, c). These results suggest that nitrate supply promotes the transcriptional activation capability of TaPHR2 by upregulating TaTCP6 and downregulating TaTCP19 expressions, thereby enhancing phosphorus uptake and utilization. Then, wheat TaPHR2 was isolated based on its homology to OsPHR2 (Supplementary Fig. 8), and three homozygous transgenic lines that constitutively overexpress TaPHR2 (TaPHR2-OE-2/4/5) were generated (Fig. 7d). Consistently, overexpression of TaPHR2 promotes the expression of PSR genes and increased Pi content (Fig. 7e, f). Further quantitative analysis revealed that the Pi contents in the TaPHR2-OE × TaTCP6-SRDX T1 plants were significantly lower than those in TaPHR2-OE wheat plants (Fig. 7g), which may be majorly attributed to the suppressed activity of OsPHR2 by TaTCP6-SRDX. In summary, these results provide compelling evidence that TaTCP6 can promote Pi accumulation in wheat by increasing the activity of TaPHR2.
a Analysis of transcriptional activation ability of TaPHR2, TaTCP6 and TaPHR2-TaTCP6 complex on the promoters of TaIPS1.1, TaPht1;2, TaPht1;6, TaIPS1.2, TaPht1;11, and TaRLI1 in wheat protoplasts. Error bars represent the mean ± SD (n = 6). b GAL4-dependent chimeric transactivation assay of TaPHR2 and TaPHR2 co-expression with TaTCP6, TaTCP19, TaPCF3, TaTCP9, TaSPX1, and TaSPX1-TaTCP6 complex in wheat protoplasts. The upper part is a schematic illustration of the effector and reporter constructs. M1: EV; M2: TaPHR2; M3: TaPHR2-TaTCP6; M4: TaPHR2-TaTCP19; M5: TaPHR2-TaPCF3; M6: TaPHR2-TaTCP9; N1: EV; N2: TaTCP6; N3: TaPHR2; N4: TaPHR2-TaTCP6; N5: TaPHR2-TaSPX1; N6: TaPHR2-TaSPX1-TaTCP6. TaPHR2 with BD (GAL4-DNA-Binding Domain). Error bars represent the mean ± SD (n = 6). c Analysis of the transcriptional activation properties of TaPHR2 in wheat protoplasts. An LUC reporter driven by the artificial promoters 4×P1BS-min35S was used to test the transcriptional activity of TaPHR2. The different combinations were co-transformed in wheat protoplasts, and the combination is the same as shown in (b). Empty vectors with 4×P1BS-min35S were used as the control. Error bars represent the mean ± SD (n = 6). d Transcription levels of TaPHR2 in three transgenic lines (TaPHR2-OE-2, TaPHR2-OE-4, and TaPHR2-OE-5) independently overexpressing TaPHR2. Data are mean ± SD (n = 6). e Pi contents of Mock and TaPHR2-OE wheat plants. Wheat seedlings grown for 2 weeks were sampled for analysis Pi contents. Data are mean ± SD (n = 9). f Transcription levels of TaIPS1.1, TaPht1;2, and TaPht1;6 in the roots of TaPHR2-OE-2, TaPHR2-OE-5, and Fielder wheat plants. Data are mean ± SD (n = 3). g Pi contents of Mock, TaPHR2-OE × Mock and TaPHR2-OE × TaTCP6-SRDX plants. The flag leaves of wheat plants grown in greenhouse before heading were sampled for measurement. Data are mean ± SD (n = 9). In (a-c), different letters above the bars indicate statistically significant differences (adjusted P < 0.05, one-way ANOVA with Tukey’s test). Exact P values and source data are provided as a Source Data file. In (d–g), P values are from two-sided Student’s t-tests. Exact P values and source data are provided as a Source Data file.
TaSPX1 is involved in the repression of the nitrogen utilization genes under low phosphate condition
As mentioned above, phosphate deficiency significantly reduced the expression levels of nitrogen utilization genes under HN conditions (Fig. 1f, h). However, the underlying mechanism remains largely elusive. Since the interaction of TaTCP6-SPXs could affect PSRs, we hypothesized that SPX might regulate nitrogen utilization through TaTCP6 as well. RT-qPCR showed that the transcription of nitrogen utilization genes was significantly inhibited in the TaSPX1-OE lines, and this inhibition is more significant under LP conditions (Fig. 8a–c). The LCI assay in N. benthamiana and BiFC assay in wheat protoplast showed that low phosphate significantly weakened TaSPX1-TaPHR2 interaction, but had no effect on the conjugation of TaSPX1 with TaTCP6 (Fig. 8d, e). It has been reported that the OsSPXs-OsPHR2 interaction occurs in a phosphate-dependent manner. Thus, we propose that phosphate deficiency dissociates the TaSPX1-TaPHR2 complex, allowing more TaSPX1 proteins to interact with the positive regulatory factors in the nitrate signaling pathway, such as TaTCP6 and TaNLP3/4. The expression of TaNAC2, TaNLP4, and TaSPX4 in TaTCP6-OE × TaSPX1 T1 plants was significantly reduced compared to TaTCP6-OE lines, indicating that TaSPX1 suppressed the activity of TaTCP6, thereby impairing its function of activating nitrogen utilization genes (Fig. 8f). Moreover, the transcription factor TaNIGT1, induced by low phosphorus, markedly suppressed the expression of the nitrogen utilization genes, TaNRT1.1 and TaNRT2.1 (Supplementary Fig. 13a). Overall, our study has unveiled mechanisms underlying the low phosphate-mediated inhibitory of the expression of nitrogen utilization genes, as well as nitrogen uptake and utilization.
a Transcription levels of TaSPX1 in two transgenic lines (TaSPX1-OE-1, TaSPX1-OE-2) independently overexpressing TaSPX1. Data are mean ± SD (n = 3). Transcription levels of TaNRT1.1B, TaNRT2.1, TaNIA1, and TaNIR in the roots of TaSPX1-OE-1, TaSPX1-OE-2, and Fielder wheat plants under HP (b) and LP (c) conditions. Data are mean ± SD (n = 3). d The interaction assays of SPX1-PHR2 and SPX1-TCP6 with InsP6 treatments in N. benthamiana leaves by LCI assays. Incubated with 300 µM InsP6 for 1 h Data are mean ± SD (n = 4). e The interaction assays of SPX1-PHR2 and SPX1-TCP6 with NaH2PO4 treatments (LP: 0 mM; HP: 0.18 mM) in wheat protoplasts by BiFC assays. wheat protoplasts isolated from 10-day-old Pi-starved seedlings (Endosperm was removed 3 days after seedling emergence and transplanted to phosphorus-deficient Hoagland’s culture solution) were co-transfected with the indicated vectors. Photos were taken after 18 h of incubation with indicated treatments. Scale bars, 50 μm. f Transcription levels of TaNLP4, TaNAC2, and TaSPX4 in Mock, TaTCP6-OE-2, TaTCP6-OE-2 × TaSPX1-OE-1, and TaTCP6-OE-2 × TaSPX1-OE-2. Data are mean ± SD (n = 3). In (a–d), P values are from two-sided Student’s t-tests. Exact P values and source data are provided as a Source Data file. In (f), different letters above the bars indicate statistically significant differences (adjusted P < 0.05, one-way ANOVA with Tukey’s test). Exact P values and source data are provided as a Source Data file.
TaTCP6 contributes to the nitrogen and phosphorus starvation adaptation and significantly increases the yield in the field
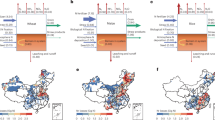
We next determined the effects of TaTCP6 on agronomic traits in field trials. The performance of the Mock and transgenic lines TaTCP6-OE-2/5, and TaTCP6-SRDX-1/2, were evaluated in field experiments under normal (NN&NP) and low-N and low-P (LN&LP) conditions in Shijiazhuang (Fig. 9a) and normal conditions in Beijing. Various agronomic traits, such as plant height, tiller number, and grain number per spike showed significant deterioration in LN&LP conditions, while 1000-grain weight (TGW) showed no significant difference and even increased compared with NN&NP condition (Fig. 9g, h). TaTCP6-OE-2/5 significantly increased the number of productive tillers under both NN&NP and LN&LP conditions (Fig. 9b, e–g). TaTCP-OE-2 with medium TaTCP6 expression also showed higher plant height and TGW under the NN&NP condition, while it showed higher grain number per spike and relatively lower TGW under LN&LP (Fig. 9b, e–g). TaTCP6-OE-5 with high TaTCP6 expression showed no significant difference in these agronomic traits compared with control (Fig. 9b, e–g). Under both conditions, TaTCP6-OE-2 and TaTCP6-OE-5 were significantly higher single plant yield (Fig. 9b, c, e–g). It is noteworthy that under NN&NP condition, TaTCP6-OE-2 obtained a higher yield due to its better spike and grain performance compared with TaTCP6-OE-5, although its tiller number is less than that of TaTCP6-OE-5 (Fig. 9b, c, e–g). The overexpression of TaTCP6-SRDX severely impaired the growth of wheat under NN&NP condition, with a greater impact as expression levels increased. However, under the LN&LP condition, this effect was weakened (Fig. 9d, e, f, h). The field performance in Beijing is comparable to that in Shijiazhuang. However, the increase in single plant yield by overexpression of TaTCP6 is less pronounced than in Shijiazhuang. This difference can be attributed to the decrease in the TGW of TaTCP6 overexpression lines in Beijing compared to the control group (Fig. 9d, i). These results suggested that elevating TaTCP6 to an optimal level could significantly boost grain yield under both NN&NP and LN&LP conditions in the field.
a Total nitrogen and phosphorus content and wheat growth performance in two plots we used for field experiments. Shijiazhuang, Hebei (2024). NN&NP (Normal nitrogen and normal phosphorus), LN&LP (Low nitrogen and low phosphorus). Data are mean ± SD. Phenotype of Mock and TaTCP6-OE lines under HN&HP (b) and LN&LP (c) conditions at a mature stage in Shijiazhuang. d Phenotype of Mock and TaTCP6-SRDX lines at the mature stage under normal field conditions in Beijing, 2024. Phenotype of TaTCP6-SRDX lines with Mock (e) and TaTCP6-OE with Mock (f) at the 10 d postanthesis (DPA) under HN&HP and LN&LP conditions in Shijiazhuang. Scale bars, 20 cm. g Plant height, number of productive tillers, spikelet number per spike, grain number per spike 1000-grain weight, and yield per plant of Mock and TaTCP6 -OE lines under HN&HP and LN&LP conditions in Shijiazhuang. Data are mean ± SD. h Plant height, number of productive tillers, spikelet number per spike, grain number per spike 1000-grain weight, and yield per plant of Mock and TaTCP6-SRDX lines under HN&HP and LN&LP conditions in Shijiazhuang. Data are mean ± SD. i Plant height, number of productive tillers, spikelet number per spike, grain number per spike 1000-grain weight, and yield per plant of TaTCP6-SRDX lines with Mock and TaTCP6-OE lines with Mock under normal conditions in Beijing, 2024. Data are mean ± SD. In (a, g–i), Boxes represent the median values and the first and third quartiles; whiskers represent the minimum and maximum values. P values are from two-sided Student’s t-test. Exact P values and source data are provided as a Source Data file.
Discussion
In this study, we collected a series of transcriptome data for wheat seedlings under different nitrogen and phosphorus conditions. From these data, we identified a Class I TCP transcription factor, TaTCP6, that responds to external nitrate and directly regulates nitrate uptake and assimilation. Additionally, TaTCP6 interacts with TaSPXs and TaPHR2, releasing TaPHR2 from the TaSPXs-TaPHR2 complex and increasing its transcriptional activation capacity, thus significantly improving P utilization efficiency in wheat (Fig. 10). Overexpression of TaTCP6 significantly increased yield, while knockout lines exhibited substantial yield loss. Importantly, we proved that low phosphorus inhibited nitrogen utilization through enhancing TaSPXs-TaNLP3/TaTCP6 interactions. All these findings demonstrate that TaTCP6 acts as a central regulator of coordinated N–P uptake and assimilation.
In the presence of elevated nitrogen levels, the expression of TaTCP6 was observed to undergo a substantial increase. This protein has been shown to bind directly with and promote the expression of genes involved in NUG (Nitrogen utilization genes). Furthermore, the increased expression of TCP6 has been found to interact with TaPHR2 and TaSPXs, thereby promoting the expression of genes associated with PSR (Phosphorus starvation response). Conversely, in a low-phosphorus condition. TaSPX1 interactions with TaPHR2 were weakened, thereby promoting the expression of PSR genes (including TaSPXs). The abundantly expressed TaSPXs were forced to interact with TaTCP6 and TaNLPs to repress the expression of NUG.
Nitrate serves as both the primary nitrogen source and a crucial signal that governs the balanced utilization of N–P, C–N, and diverse agronomic traits such as tiller number in plants. Plants fix CO2 through photosynthesis during carbon assimilation, while C–N balance encompasses both nutrient and energy equilibrium. The two widely recognized transcription factors in rice, OsGRF4 and OsDREB1C, are adapted to low nitrogen conditions and play positive roles in carbon assimilation and nitrate uptake35,36. In this study, the anatomical observations and measurements of nitrate content in wheat kernels indicated that TaTCP6 appeared to contribute more to carbon assimilation than to nitrate assimilation, as evidenced by a greater net increase in carbohydrates compared to proteins in the grains. Thus, our research uncovers an aspect of the mechanism through which the efficient supply of N improves crop photosynthetic efficiency.
Under HP conditions, SPX and PHR will form a complex and exert opposed roles for PSR, while they tend to act in concert to inhibit plant development under LP conditions. OsPHR2 has received much more attention due to the ease of transcription factor research, and equal attention should be paid to SPX. The multifunctionality of SPX proteins in plants has been increasingly revealed. For instance, the interaction between OsSPX and OsBZR1 is implicated in BR response and plant defense against disease in LP environments37. In this study, we highlighted the significance of the interactions between SPXs and TCPs in wheat. Interestingly, SPXs showed much weaker interactions with TaTCP19 compared to TaTCP6, unlike the robust interactions observed between TaTCP6/TaTCP19 and TaPHR2. Resolving this incident necessitates additional efforts using methods such as structural biology.
A recent discovery has revealed that OsRLI1 in rice exists in two isoforms. The truncated version RLI1a lacks the coiled-coil (CC) domain, resulting in the inability to form dimers. The absence of a CC domain in RLI1a enables it to activate a wider range of target genes compared to RLI1b, the full-length isoform. RLI1a, but not RLI1b, is stimulated by HP and has the ability to regulate both the BR pathway and leaf inclination38. In this study, we searched the reference genome and designed primers to explore the presence of alternative splicing (AS) of TaRLI1. Although TaRLI1 is induced by HP, no AS events were detected in wheat. Consequently, substantial efforts were needed to investigate the reasons for the loss of AS for TaRLI1 and its role in regulating leaf angle under HP conditions in wheat.
In addition, an in-depth study is required to explore the involvement of epigenetic modifications in response to N and P signalings39,40. The stability of OsNGR5 is enhanced by nitrogen, and it aids in the recruitment of PRC2 to suppress genes that inhibit branching through H3K27me3 modification39. Under LP conditions, AtPHR1/AtPHL1 is responsible for the PSR throughout the entire genome, with the rapid gene response being influenced by alterations in genome accessibility41. It has been reported that OsHMGB1, as a chromatin structural protein, binds to the promoters and regulates a variety of genes responsive to LP by enhancing chromatin accessibility42. Previous studies suggested that TCP proteins could directly or indirectly regulate the level of epigenetic modifications of downstream genes. Here, we found the interaction between TaTCP6 and acetyltransferase TaHAG1 from our Y2H screening. The Tahag1 mutants, particularly Tahag1-Aabbdd, showed no significant developmental defects in normal environments but manifested striking phenotypic traits under stress. Similarly, the impairments in nitrate and phosphorus uptake appear to occur only in nutrients-deprived environments (Supplementary Fig. 14). This prompted our curiosity as to how the TaTCP6-TaHAG1 module responds to nutrient deficiency.
Excessive nitrogen promotes the formation of nonproductive tillers in rice, which decreases nitrogen use efficiency (NUE). OsGATA8-H was identified as a high-NUE haplotype with enhanced nitrogen uptake and a higher proportion of productive tillers43. Compared with TaTCP6-OE-2, TaTCP6-OE-5 with higher expression levels of TaTCP6 has relatively more tillers but lower final yield which may be due to an increased incidence of nonproductive tillering in TaTCP6-OE-5 (Fig. 9b, e–g). Nonproductive tillers compete with productive tillers for nutrients, soil and sunlight, which deteriorates population quality and ultimately affects wheat yield. This view is supported by the better spike and grain performance of TaTCP6-OE-2 (Fig. 9b, e–g; Supplementary Fig. 15 and Supplementary Data 11). Furthermore, we observed that the yield-increasing effect of overexpression of TaTCP6 is not as good in Beijing as in Shijiazhuang (Fig. 9), which may be attributed to the narrow plant spacing in Beijing (15 plants/2 m in Shijiazhuang and 15 plants/m in Beijing). Overexpression of TaTCP6 in Beijing promoted tillering, but it reduced the TGW of the whole plant spike, although the TGW of the main stem spike in the overexpression lines was still better than that of mock (Fig. 9i; Supplementary Fig. 15 and Supplementary Data 11). Resource constraints may affect the potential of tillering to promote yield. It is noteworthy that increasing nitrogen application often contributed to plant height and tiller numbers, but the formation of spike traits is related to both early vegetative growth and nutritional conditions. If there are too many tillers, it may result in fewer GPS and TGW. If there are few tillers and not that many nutrients, it may gain a higher TGW. In addition, if nutrition is limited, wheat may still be preferentially supplied to the main stem spike, which may lead to inconsistent performance between the main stem spike and the whole plant (Fig. 9i; Supplementary Fig. 15 and Supplementary Data 11). For example, from the perspective of the main stem spike, the performance has improved, but overall, the performance has deteriorated (Fig. 9; Supplementary Fig. 15 and Supplementary Data 11). This highlights the necessity of using more indicators to objectively describe the adaptation of crops to soil nutrients and yield formation.
Nitrate supply appears to dominate the PSR, a phenomenon that still cannot be fully explained. The comprehensive mechanism and to what extent nitrate controls the PSR still need more effort. These researches will contribute to the rational application of various nutrient elements and the reduction of fertilizer inputs for sustainable agriculture.
Methods
Plant materials
For generating overexpression lines of TaTCP6 (TaTCP6-OE) and TaSPX1 (TaSPX1-OE), the full-length coding sequence (CDS) of TaTCP6-B and TaSPX1-D fused with 3×Flag tag sequence was cloned into the PLC41-Z2 vector respectively to generate Ubipro::TaTCP6-B-Flag and Ubipro::TaSPX1-D-Flag. To generate the material of TaTCP6-SRDX, the full-length coding sequence (CDS) of TaTCP6-B and its 3K promoter fused with SRDX tag sequence was cloned into the PLC41-Z2 vector to generate TaTCP6-Bpro::TaTCP6-B-SRDX. For generating overexpression lines of TaPHR2 (TaPHR2-OE), the full-length coding sequence (CDS) of TaPHR2-B fused with GFP tag sequence was cloned into the PLC41-Z2 vector respectively to generate Ubipro::TaPHR2-B-GFP. These vectors were further transformed into wheat cultivar “Fielder” respectively via Agrobacterium tumefaciens-mediated gene transformation. The Tatcp6 mutant plants were generated by CRISPR/Cas9-based gene editing. In brief, the sgRNA target sequences for TaTCP6-A/B/D gene editing were first designed using the web-based CRISPR-Cereal (http://crispr.hzau.edu.cn/CRISPR-Cereal/). Then, the sgRNA sequences were introduced into the pBUE414 vector via T4 DNA ligase (M0202, NEB). The resulting constructs were transferred into the A. tumefaciens strain EHA105 and transformed into wheat cultivar “Fielder” by A. tumefaciens-mediated gene transformation. All these wheat materials were validated by PCR and sequencing. The primers used in this study are listed in Supplementary Data 12.
Growth conditions and methods of field experiment
The field experiments for wheat growing were conducted in the Shangzhuang Experimental Station which belongs to the College of Agronomy and Biotechnology, China Agricultural University in Beijing, and Zhaoxian Experimental Station which belongs to the Institute of Grain and Oil Crops, Hebei Academy of Agriculture and Forestry Sciences in Shijiazhuang, Hebei Province. Two treatments (NN&NP and LN&LP) were carried out at Dishang Experimental Station (2024). The NN&NP treatment had 18 g N/m2 and 8 g P/m2 applied prior to sowing and 9.0 g N/m2 applied at the stem elongation stage. The LN&LP treatment had no N and P application. Twenty-five seeds of each line were sown in 2 m rows, spaced 20 cm apart. Two consecutive field experiments were also conducted in Shangzhuang Experimental Station (2023–2024) with 15 seeds of each line sown in 1 m rows, spaced 20 cm apart. All rice plants were grown under natural paddy conditions at the Shangzhuang Experimental Station (2023). Seeds were firstly sown in the seedbed for germination. After 21 days, seedlings grown in the seedbed were transplanted into the paddy field with plant spacing: 15 cm, and row spacing: 25 cm.
Growth conditions and methods of hydroponics experiment
The wheat seeds (Fielder genetic background) were initially germinated on wet filter papers for about 2 days. Then the seedlings were cultivated in the plant hydroponic box (127 × 87 × 114 mm) for 4 days with 1 L distilled water, changing the water every 2 days. After removal of residual endosperm, the seedlings were transferred to plastic containers (30 × 15 × 15 cm) that contained 6 L of indicated nutrient solution: HNHP (2.5 mM KNO3, 0.3 mM NaH2PO4), HNLP (2.5 mM KNO3, 0.03 mM NaH2PO4), LNHP (0.25 mM KNO3, 0.3 mM NaH2PO4), LNLP (0.25 mM KNO3, 0.03 mM NaH2PO4). The nutrient solution was adjusted based on basal nutrient solution consists of the following macronutrients: 2.5 mM KNO3, 0.3 mM NaH2PO4·2H2O, 1 mM CaCl2·2H2O, 1 mM MgSO4·7H2O, 0.77 µM ZnSO4·7H2O, 0.32 µM CuSO4·5H2O, 40 µM EDTA–FeNa·3H2O, 9 µM MnSO4·4H2O, 0.05 µM (NH4)6·Mo7O24·4H2O and 20 µM H3BO3, with the pH adjusted to 5.8. KNO3 and NaH2PO4·2H2O was added to the basal nutrient solution at indicated concentrations for different treatments. The nutrient solution was renewed every 3 days. Five weeks after cultivation, wheat plants were harvested for phenotypic evaluation. All plants were grown in plant growth chambers (BPC500HL, Fujian Jiupo Biotechnology Co., Ltd) under 16 h of light at 24 °C and 8 h of dark at 20 °C with about 70% relative humidity.
Assay for short-term nitrate-induced wheat transcriptome sequencing
For all the short-term nitrate induction assays in this study, germinated seedlings (Chinese spring) were first grown in distilled water for 2 days. After removal of residual endosperm, the seedlings were then subjected to NO3− deficiency nutrient solution (0.25 mM (NH4)2SO4 and 2.5 mM KCl) for 12 days. Subsequently, the seedlings were exposed to a nutrient solution with 2.5 mM NO3− was added (0.25 mM (NH4)2SO4 and 2.5 mM KNO3−) for different durations. Total RNA from whole plants was extracted using the TRIzol reagent (Invitrogen), following the manufacturer’s protocol.
Total RNA extraction and real time-PCR (RT-PCR)
Under normal conditions and treatment, the roots of 14-day-old seedlings were collected for RNA extraction. Total RNAs were extracted using TRIzolTM Reagent (15596018, Thermo Fisher Scientific). The reverse transcription of cDNAs was performed by using HiScript II Q Select RT SuperMix for qPCR (+gDNA wiper) (R223-01, Vazyme Biotech) according to the manufacturer’s recommendations. Taq Pro Universal SYBR qPCR Master Mix (Q712-02, Vazyme Biotech) was used for RT-PCR analysis. All primers were shown in Supplementary Data 12 and β-ACTIN was used as an internal control.
RNA-seq analysis
Raw sequencing reads were processed and filtered using the software fastp with the default parameters. The remaining high-quality reads were mapped to the wheat IWGSC RefSeq v1.1 reference genome using STAR with the default parameters. Feature Counts were then used to summarize the reads mapped to each gene. To avoid confounding effects from low-quality gene models, only high-confidence gene models with an FPKM of more than 1 in at least one sample were considered expressed. DESeq2 was used for differential expression analysis. The genes with an absolute fold change value greater than 1 and an FDR less than 0.05 were considered DEGs. The GO enrichment analysis was performed using GO Analysis Toolkit and Datadase for Agriculture Community (agriGO), and terms with an FDR less than 0.05 were considered significant.
CUT&Tag assay
The CUT&Tag assays were conducted by Igenebook Biotechnology Co., Ltd according to the previously described44. Wheat seedlings were cleaved into nucleus suspension, and 10 w nuclei were harvested and incubated with concanavalin A coated magnetic beads for 15 min at RT. Then, bead-bound cells were resuspended and incubated with the appropriate primary antibody (1:50; F1804, Sigma) and IGG(1:50; 2729S, CST) overnight at 4 °C. DNA from CUT&Tag was used to construct sequencing libraries following the protocol provided with Hyperactive Universal CUT&Tag Assay Kit for Illumina Pro (TD904, Vazyme Biotech Co., Ltd).
Analysis of CUT&Tag data
Trimmomatic (version 0.36) was used to filter out low-quality reads. Clean reads were mapped to the Chinese Spring reference genome (IWGSC, RefSeq v1.1) by Bwa (version 0.7.15). Samtools (version 1.3.1) was used to remove potential PCR duplicates.MACS2 software (version 2.1.1.20160309) was used to call peaks by default parameters (bandwidth, 300 bp; model fold, 5, 50; p value, 0.00001). If the midpoint of a peak is located closest to the TSS of one gene, the peak will be assigned to that gene. HOMER (version 3) was used to predict motif occurrence within peaks with default settings for a maximum motif length of 12 base pairs. ClusterProfiler (http://www.bioconductor.org/packages/release/ bioc/html/clusterProfiler.html) in the R package was employed to perform GO (http://geneontology.org/). The GO and KEGG enrichment analysis were calculated using a hypergeometric distribution with a q value cutoff of 0.05.
Yeast one-hybrid (Y1H) assay
The partial TaNRT2.1-6B promoter sequence was amplified, recombined with the pAbAi vector, and then integrated with the genome of the Y1HGold yeast strain. CDS of the TaTCP6 was amplified, cloned into the pGADT7 vector, and transferred into the bait yeast strain. The detailed procedure was according to the Matchmaker Gold Y1H Library Screening System (Clontech). Different concentrations of the aureobasidin A (AbA) antibiotic were used to select the positive bait yeast strain on a medium lacking Ura and Leu (-L/-U).
Yeast two-hybrid (Y2H) assay
The coding sequence for the truncated TaTCP6 was amplified and cloned into the pGBKT7 vector to generate pGBKT7-TaTCP6tcp, which was used as a bait to screen a yeast cDNA library derived from Fielder wheat seedlings to identify the candidate TaTCP6-interacting proteins. The Y2H screening was performed according to the user’s manual (PT3024-1, Clontech Laboratories, Inc.) The coding sequences of genes of interest were cloned into the pGADT7 vectors (Clontech), and various combinations of plasmids were co-transformed into the yeast AH109 strain. Positive transformants were selected on synthetic drop-out (SD) medium lacking Trp and Leu (-L/-T), while the screening of interactions was performed on SD medium lacking Trp, Leu, His, and Ade (-L/-T/-H/-A).
Split-luciferase complementation imaging (LCI) assay
The CDSs of indicated genes were separately cloned into the pCambia1300-nLUC (nLUC) and pCambia1300-cLUC (cLUC) vectors. The resulting constructs were transformed into Agrobacterium tumefaciens strain GV3101, which were further co-infiltrated in N. benthamiana leaves. About 36 h after infiltration, the LUC signal within the infiltration region was analyzed using NightSHADE LB 985 (Berthold Technologies).
Bimolecular fluorescent complimentary (BiFC) assay
The CDSs of indicated genes were separately cloned into the pUC57-cYFP and pUC57-nYFP vectors. The indicated constructs were transiently expressed in wheat protoplasts using a PEG-calcium mediated method as described previously45. The fluorescence signals were detected at 488 nm for YFP using the confocal 507 microscope (LSM900; Carl Zeiss, Heidenheim, Germany).
Immunoprecipitation and co-immunoprecipitation assay
Total proteins of wheat seedlings were extracted with SDS-loading buffer (200 mM Tris pH 6.8, 40% glycerol, 8% SDS, and 20% β-mercaptoethanol) at 100 °C for 5 min. The Extracted proteins were then separated on SDS-PAGE and transferred onto the PVDF membrane. The membrane-carrying proteins were blocked with 5% milk in TBST containing 0.1% Tween-20 at room temperature for 1 h and incubated with the corresponding antibodies, such as anti-rabbit IgG-HRP and anti-mouse IgG-HRP (Sigma-Aldrich). The proteins were detected by IMMOBILON WESTERN CHEMILUM HRP SUBSTRATE (WBKLS, Millipore) under a fluorescence chemiluminescence imaging system (ChemiScope 6100). For Co-IP assays in N. benthamiana leaf tissues, the full-length CDS of indicated genes were cloned into pCAMBIA1300-221-Flag, pSuper1300-MYC, pSuper1300-mCherry, and pSuper1300-GFP vector and then they were transformed into Agrobacterium strains and co-infiltrated into N. benthamiana leaves. About 36 h after infiltration, the total proteins of infiltrated N. benthamiana leaves were extracted with the lysis buffer as described above. For Co-IP assays in wheat protoplast, the full-length CDS of indicated genes were cloned into pUC57-GFP, pUC57-MYC, and pUC57-Flag. About 16 h after infiltration, the total proteins of the infiltrated wheat protoplast were extracted with the lysis buffer as described above. Anti-GFP Magnetic Beads (M8823, Millipore) were used for TaTCP6-GFP protein immunoprecipitation. The immunoprecipitated proteins were detected by anti-Flag (1:5000; F3165, Sigma-Aldrich), anti-MYC (1:5000; ab9106, Abcam), anti-mCherry (1:2000; AE002, Abconal), anti-Actin (1:2000; AC009, Abconal) and anti-GFP antibodies (1:5000; ab32146, Abcam).
Subcellular localization
The CDS of TaTCP6, TaPHR2, TaRLI1, TaSPX1, TaSPX3, TaSPX4, and TaSPX6 was cloned into the pSuper1300-GFP vector and TaSPX4 was also cloned into the pSuper1300-mCherry vector. The indicated constructs were transiently expressed in wheat protoplasts using a PEG-calcium mediated method as described previously45. The fluorescence signals were detected at 488 nm for GFP and 546 nm for mCherry using the confocal microscope (LSM900; Carl Zeiss, Heidenheim, Germany). The resulting vectors of TaSPX1-GFP and TaSPX4-GFP were also introduced into Agrobacterium tumefaciens strain GV3101 and the bacterial suspensions infiltrated into about 4-week-old N. benthamiana leaves. After 2 days, the infiltrated leaves were collected and imaged on an LSM900 confocal microscope (Zeiss, Heidenheim, Germany). All primers used are listed in Supplementary Data 12.
Dual-luciferase transcriptional activity assay
The transcriptional activities of TaTCP6 on its downstream genes were carried out in Nicotiana benthamiana. TaTCP6 were cloned into the pSuper1300 vector as effectors. The promoter of TaNRT1.1B, TaNRT2.1, TaNIA1, TaNIR, TaIPS1.1, TaPht1;2, TaPht1;6, TaIPS1;2, TaPht1;11, and TaRLI1 was cloned into the dual-luciferase vector pGreenII 0800-LUC, respectively, to drive the firefly luciferase reporter gene (LUC), and luciferase from Renilla reniformis (REN) as an internal control was driven by 35S promoter. All plasmids were transformed into Agrobacterium GV3101 and expressed in N. benthamiana leaves by Agrobacterium-mediated transient transformation. Infected tissues were analyzed 48 h after infiltration. Activities of LUC and REN were examined using Dual-Luciferase Reporter Assay System kit (E1960; Promega). The expression level of a tested promoter is normalized by calculating the LUC:REN ratio.
Transient dual-luciferase assays were also carried out to detect the effect of TaTCPs, TaSPX1, and TaTCPs-TaSPX1 complex on the transcriptional activation activity of TaPHR2. The TaPHR2 CDS were cloned into a vector to fuse TaPHR2 with the GAL4-BD domain and generate the effector construct BD-TaPHR2. TaTCPs and TaSPX1 were also cloned into the pCAMBIA1300 vector respectively as effectors. To create the reporter construct, a promoter with a 6× GAL4 upstream activating sequence and a TATA box was introduced into the pGreenII 0800-LUC vector. The reporter and effector constructs were co-transformed N. benthamiana leaves as indicated by Agrobacterium-mediated transient expression, using the empty effector construct GAL4-BD as a control. Infected tissues were analyzed 48 h after infiltration. Activities of LUC and REN were examined using Dual-Luciferase Reporter Assay System kit (E1960; Promega). The expression level of a tested promoter is normalized by calculating the LUC:REN ratio.
Electrophoretic mobility shift assay (EMSA)
The full-length CDS of TaTCP6 were cloned into pMAL-c2x vector to generate pMAL-c2x-TaTCP6. pMAL-c2x, and pMAL-c2x-TaTCP6 were expressed in the E. coli Rosetta (DE3) strain (Weidi, Shanghai, China) and purified using amylose resin (New England Biolabs, Beijing, China) respectively. Oligonucleotide probes were synthesized and their 5′ ends were labeled with biotin. EMSA was performed using the Light Shift Chemiluminescent EMSA Kit (20148; Thermo Fisher Scientific), following the manufacturer’s instructions. The binding reactions were performed according to the corresponding manufacturer’s protocol. For the binding system of MBP-TaTCP6 fusion protein, about 2 μg protein, 0.02 μM biotin-labeled probe and appropriate amount of competitive probe were incubated in 20 μL reaction mixtures containing 15 mM Tris-HCl, pH 7.5, 50 mM KCl, 2 mM EDTA, 0.05% NP-40, 2 μg/μL BSA, 200 μM ZnCl2, 50 ng/μL poly (dI-dC), 5% glycerol for 20 min on 25 °C and then employed for electrophoretic analysis.
Pull-down assay
MBP, MBP-TaTCP6, MBP-TaPHR2, and MBP-TaRLI1 fused proteins were expressed in the E. coli Rosetta (DE3) strain (Weidi, Shanghai, China) and purified using amylose resin (New England Biolabs, Beijing, China) respectively. GST-TaSPX1 fused proteins were expressed in the E. coli Rosetta (DE3) strain (Weidi, Shanghai, China) and purified using ProteinIso® GST Resin (Transgen Biotech, Beijing, China). HIS-TF-TaTCP6 For HIS fused protein was expressed in the E. coli Rosetta (DE3) strain (Weidi, Shanghai, China) and purified using ProteinIso® HIS Resin (Transgen Biotech, Beijing, China). pull-down assays, the GST-TaSPX1 protein was co-incubated separately with MBP, MBP-TaTCP6, MBP-TaPHR2, and MBP-TaRLI1 in lysis buffer (25 mM Tris (PH 7.5), 50 mM NaCl, 1 mM DTT) with amylose resin at 4 °C for 2 h. Afterward, the beads were washed thoroughly, and proteins were analyzed by western blot using anti-GST (1:2000; AE001, ABclonal), anti-HIS (1:2000; AE003, ABclonal) and anti-MBP (1:2000; AE016, ABclonal) antibodies.
Phylogenetic analysis
To perform phylogenetic analysis, protein sequences of TCP gene family, GARP gene family, and SPX gene family from different plant species were downloaded on Ensembl Plants (http://plants.ensembl.org/index.html). Mega X46 was employed to construct the phylogenetic tree with the programs as follows: first, all protein sequences were aligned in Clustal W with default parameters; second, the Maximum Likelihood method was used to build phylogenetic tree with the Poisson model and 1000 replicates bootstrap; next, the phylogenetic tree was colored and beautified on the website Interactive Tree of Life.
Histological analysis
The collected samples of mature wheat grains were fixed in an FAA solution [10% (v/v) formaldehyde, 50% (v/v) alcohol, 5% (v/v) acetic acid, and 35% (v/v) water overnight at 4 °C, and then were embedded in paraffin, dehydrated and decolorized as described previously47. The samples were then cut into 4-µm thick cross sections using a Leica Ultracut rotary microtome (Leica Biosystems, Wetzlar, Germany), and stained with Periodic acid Schiff (PAS) or 1% sarranine and 0.5% fast green (G1031, Servicebio, Wuhan). Photographs were taken with a microscope imaging system (DS-U3, Nikon, Japan).
15N uptake rate analysis
For 15N uptake rate analysis, wheat seedlings were cultured in the hydroponic culture (supplemented with 2.5 mM KNO3, pH 5.8) for 2 weeks. The seedlings were then transferred into modified hydroponic culture containing 2.5 mM 15N-KNO3 for 3 h. The roots of the treated seedlings were washed in 0.1 mM CaSO4 for 2 min to remove the unabsorbed 15N-KNO3 on the surface of the roots. Shoots and roots of the treated seedlings were collected separately and dried at 65 °C. The samples were ground to a fine powder and used for 15N content measurement using an isotope ratio mass spectrometer equipped with an elemental analyzer (Thermo Finnigan Delta Plus XP; Flash EA 1112).
Measurements of Pi content in wheat plants
A frozen sample after fresh weight measurement was homogenized in 0.1 mL Pi extraction buffer (0.1 M NaCl, 1 mM EDTA (pH 8.0, 10 mM Tris-HCl (pH 8.0), 0.07‰ (w/v) 2-hydroxy-1-ethanethiol, and 1 mM PMSF), using an ice-cold mortar and pestle. The homogenate was then diluted 10 times with 0.9 mL 1% (w/v) acetic acid and placed on ice for 30 min. After centrifugation at 10,000 × g for 10 min at 4 °C, The supernatant was collected, diluted to an appropriate concentration, then mixed with a molybdenum blue reagent (0.4% (w/v) ammonium molybdate melted in 0.5 m H2SO4 (solution A) was mixed with 10% ascorbic acid (solution B; A:B = 6:1) in a 7:3 ratio and incubated for 30 min at 42 °C. The absorption values of the solution at 820 nm were determined using a microplate reader (Tecan Spark). A standard curve was calculated from varying concentrations of KH2PO4, then the Pi content was calculated according to the standard curve and dilution ratio.
Pi-depletion assay
For the Pi-depletion experiment, a group of three seedlings was transferred into a flask with 500 mL of depletion solution (a modified nutrient salt depletion solution containing 100 μM NaH2PO4, Kimura B). All flasks were kept on a shaking table at 130 rpm. A 500 mL volume of depletion solution was withdrawn at the indicated time, and the Pi concentration was measured.
Measurement of nitrate reductase activity
Leaves of wheat seedlings growing under four different conditions were used to measure NR activity, following the instruction manual of the NR Kit (Solarbio LIFE SCIENCES, Beijing, China; BC0080). 0.1 g leaf samples were extracted in 1 mL extraction solution and the mixture was centrifuged at 4000 × g for 10 min. The resulting supernatant was collected for further analysis. The absorbance at 340 nm was used for the calculation of NR activity, one unit (U) of NR activity was defined as the amount of 1 μmol NADH consumed per hour per gram of sample48.
Nitrate concentration measurements
The nitrate concentrations in leaves of wheat seedlings were measured by the salicylic acid (SA) concentrated sulfuric acid colorimetry method. Samples of 1 g fresh wheat leaves or roots were homogenized, and 10 mL sterile water was added, and then nitrate was extracted for 30 min in a boiling water bath. 0.1 mL extracted solution, 0.4 mL 5% SA-concentrated sulfuric acid (w/v) solution, and 9.5 mL 8% NaOH (w/v) solution were well mixed and OD410 was measured when the solution was cooled to 25 °C. The nitrate concentration of the extracted solution was calculated using a standard curve.
Statistics and reproducibility
Quantitative data, including three biological and at least three technical replicates, are presented in the form of mean ± SD. Means of two samples were compared using Student’s two-tailed t-tests. Analysis of variance (one-way ANOVA) was conducted using GraphPad Prism 9.0.0 software with default parameters. Significant differences were determined by one-way ANOVA or the Student’s t-test: *P < 0.05 and **P < 0.01. Tukey’s multiple comparison test was used to determine exactly which group means were different.
Three biological replicates were performed, with similar results; the results from one replicate are shown in the figures.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw data generated by this research have been deposited in the National Center for Biotechnology Information (NCBI) under accession number PRJNA1082516, PRJNA1178185 for RNA-seq and CUT&Tag. Source data are provided with this paper.
References
Park, B. S., Seo, J. S. & Chua, N. H. NITROGEN LIMITATION ADAPTATION recruits PHOSPHATE2 to target the phosphate transporter PT2 for degradation during the regulation of Arabidopsis phosphate homeostasis. Plant Cell 26, 454–464 (2014).
Medici, A. et al. AtNIGT1/HRS1 integrates nitrate and phosphate signals at the Arabidopsis root tip. Nat. Commun. 6, 6274 (2015).
Li, Q., Zhou, L., Li, Y., Zhang, D. & Gao, Y. Plant NIGT1/HRS1/HHO transcription factors: key regulators with multiple roles in plant growth, development, and stress responses. Int. J. Mol. Sci. 22, 8685 (2021).
Hu, B. et al. Nitrate-NRT1.1B-SPX4 cascade integrates nitrate and phosphorus signalling networks in plants. Nat. Plants 5, 401–413 (2019).
Liu, K. H. et al. Discovery of nitrate-CPK-NLP signalling in central nutrient-growth networks. Nature 545, 311–316 (2017).
Liu, K. H. et al. NIN-like protein 7 transcription factor is a plant nitrate sensor. Science 377, 1419–1425 (2022).
Wang, P. et al. Medicago SPX1 and SPX3 regulate phosphate homeostasis, mycorrhizal colonization, and arbuscule degradation. Plant Cell 33, 3470–3486 (2021).
Shi, J. et al. A phosphate starvation response-centered network regulates mycorrhizal symbiosis. Cell 184, 5527–5540 (2021).
Das, D. et al. PHOSPHATE STARVATION RESPONSE transcription factors enable arbuscular mycorrhiza symbiosis. Nat. Commun. 13, 477 (2022).
Wild, R. et al. Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 352, 986–990 (2016).
Zhong, Y. et al. Rice SPX6 negatively regulates the phosphate starvation response through suppression of the transcription factor PHR2. New Phytol. 219, 135–148 (2018).
Jia, X., Wang, L., Nussaume, L. & Yi, K. Cracking the code of plant central phosphate signaling. Trends Plant Sci. 28, 267–270 (2023).
Liu, H. et al. Overexpressing HRS1 confers hypersensitivity to low phosphate-elicited inhibition of primary root growth in Arabidopsis thaliana. J. Integr. Plant Biol. 51, 382–392 (2009).
Nakano, Y., Naito, Y., Nakano, T., Ohtsuki, N. & Suzuki, K. NSR1/MYR2 is a negative regulator of ASN1 expression and its possible involvement in regulation of nitrate reutilization in Arabidopsis. Plant Sci. 263, 219–225 (2017).
Maeda, Y. et al. A NIGT1-centred transcriptional cascade regulates nitrate signalling and incorporates phosphorus starvation signals in Arabidopsis. Nat. Commun. 9, 1376 (2018).
Ueda, Y. et al. Gene regulatory network and its constituent transcription factors that control nitrogen-deficiency responses in rice. New Phytol. 227, 1434–1452 (2020).
Zhang, Z. et al. Modulation of nitrate-induced phosphate response by the MYB transcription factor RLI1/HINGE1 in the nucleus. Mol. Plant 14, 17–529 (2021).
Medici, A. et al. Identification of molecular integrators shows that nitrogen actively controls the phosphate starvation response in plants. Plant Cell 31, 1171–1184 (2019).
Wang, X. et al. The transcription factor NIGT1.2 modulates both phosphate uptake and nitrate influx during phosphate starvation in Arabidopsis and maize. Plant Cell 32, 3519–3534 (2020).
Li, Y. et al. Nitrate-inducible GLK1 modulates phosphate starvation response via the PHR1-dependent pathway. New Phytol. 236, 1871–1887 (2022).
Puga, M. I. et al. SPX1 is a phosphate-dependent inhibitor of Phosphate Starvation Response 1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 111, 14947–14952 (2014).
Wang, Z. et al. Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc. Natl. Acad. Sci. USA 111, 14953–14958 (2014).
Hu, B. & Chu, C. Nitrate-phosphorus interplay: old story with molecular tale. New Phytol. 225, 1455–1460 (2020).
Paz-Ares, J. et al. Plant adaptation to low phosphorus availability: core signaling, crosstalks, and applied implications. Mol. Plant 15, 104–124 (2022).
Li, C., Potuschak, T., Colón-Carmona, A., Gutiérrez, R. A. & Doerner, P. Arabidopsis TCP20 links regulation of growth and cell division control pathways. Proc. Natl. Acad. Sci. USA 102, 12978–12983 (2005).
Guan, P. et al. Nitrate foraging by Arabidopsis roots is mediated by the transcription factor TCP20 through the systemic signaling pathway. Proc. Natl. Acad. Sci. USA 111, 15267–15272 (2014).
Guan, P. et al. Interacting TCP and NLP transcription factors control plant responses to nitrate availability. Proc. Natl. Acad. Sci. USA 114, 2419–2424 (2017).
Liu, Y. et al. Genomic basis of geographical adaptation to soil nitrate in rice. Nature 590, 600–605 (2021).
Zhao, J. et al. Genome-wide identification and expression profiling of the TCP family genes in spike and grain development of wheat (Triticum aestivum L.). Front. Plant Sci. 9, 1282 (2018).
He, X. et al. The nitrate-inducible NAC transcription factor TaNAC2-5A controls nitrate response and increases wheat yield. Plant Physiol. 169, 1991–2005 (2015).
Song, L. et al. Reducing brassinosteroid signalling enhances grain yield in semi-dwarf wheat. Nature 617, 118–124 (2023).
Safi, A. et al. The world according to GARP transcription factors. Curr. Opin. Plant Biol. 39, 159–167 (2017).
Ueda, Y. et al. NIGT1 family proteins exhibit dual mode DNA recognition to regulate nutrient response-associated genes in Arabidopsis. PLoS Genet. 16, e1009197 (2020).
Dong, J. et al. Inositol pyrophosphate InsP8 acts as an intracellular phosphate signal in Arabidopsis. Mol. Plant 12, 1463–1473 (2019).
Li, S. et al. Modulating plant growth-metabolism coordination for sustainable agriculture. Nature 560, 595–600 (2018).
Wei, S. et al. A transcriptional regulator that boosts grain yields and shortens the growth duration of rice. Science 377, eabi8455 (2022).
He, Y. et al. The OsBZR1–OsSPX1/2 module fine-tunes the growth–immunity trade-off in adaptation to phosphate availability in rice. Mol. Plant 17, 258–276 (2024).
Guo, M. et al. Alternative splicing of REGULATOR OF LEAF INCLINATION 1 modulates phosphate starvation signaling and growth in plants. Plant Cell 34, 3319–3338 (2022).
Wu, K. et al. Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice. Science 367, eaaz2046 (2020).
Zhang, J. et al. Sugar transporter modulates nitrogen-determined tillering and yield formation in rice. Nat. Commun. 15, 9233 (2024).
Barragán-Rosillo, A. C. et al. Genome accessibility dynamics in response to phosphate limitation is controlled by the PHR1 family of transcription factors in Arabidopsis. Proc. Natl. Acad. Sci. USA 118, e2107558118 (2021).
Wang, Y. et al. Rice chromatin protein OsHMGB1 is involved in phosphate homeostasis and plant growth by affecting chromatin accessibility. New Phytol. 240, 727–743 (2023).
Wu, W. et al. The elite haplotype OsGATA8-H coordinates nitrogen uptake and productive tiller formation in rice. Nat. Genet. 56, 1516–1526 (2024).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Ni, P. et al. Efficient and versatile multiplex prime editing in hexaploid wheat. Genome Biol. 24, 156 (2023).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Feng, Z. et al. The decreased expression of GW2 homologous genes contributed to the increased grain width and thousand‑grain weight in wheat-Dasypyrum villosum 6VS·6DL translocation lines. Theor. Appl. Genet. 134, 3873–3894 (2021).
Jing, Y. et al. TaLBD41 interacts with TaNAC2 to regulate nitrogen uptake and metabolism in response to nitrate availability. New Phytol. 242, 641–657 (2024).
Acknowledgements
This study was supported by the National Key Research and Development Program of China (No. 2023YFF1000601), the National Natural Science Foundation of China (No. U22A6009, No. 32201824), Chinese Universities Scientific Fund (No. 2024TC161), and Pinduoduo-China Agricultural University Research Fund (No. PC2023A01003). Our confocal microscopy and isotope mass spectrometry (IMS) work was performed at the CAB Public Instrument Platform of China Agricultural University.
Author information
Authors and Affiliations
Contributions
J.X., Q.S., and Z.N. conceived, conceptualized, and designed the study. B.L. and W.X. carried out most of the experiments. Y.N., Q.L., B.C., J.Q., Y. Zhao, Y. Zhou, L.S., and D.C. contributed to the molecular cloning, phenotyping work, and experimental data analysis. Q.S., Z.N., M.Y., Y.Y., M.X., and Z.L. contributed to the discussion and revised the manuscript. J.X., B.L., and W.X. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hatem Rouached, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, B., Xu, W., Niu, Y. et al. TaTCP6 is required for efficient and balanced utilization of nitrate and phosphorus in wheat. Nat Commun 16, 1683 (2025). https://doi.org/10.1038/s41467-025-57008-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57008-0
This article is cited by
-
Genome-wide identification of the GLK gene family in wheat (Triticum aestivum L.) and analysis of expression responses in different environments
BMC Genomics (2026)
-
Expression profiling of TaNF-YA transcription factor genes under abiotic stress conditions in wheat (Triticum aestivum L.) seedlings
Plant Biotechnology Reports (2026)
-
Wheat genomics frontiers for gene discovery and breeding applications
WheatOmics (2025)