Abstract
Postnatal respiration requires bulk formation of alveoli that produces extensive surface area for gas diffusion from epithelium to the circulatory system. Alveolar morphogenesis initiates at late gestation or postnatal stage during mammalian development and is mediated by coordination among multiple cell types. Here we show that fibroblast-derived Heparan Sulfate Glycosaminoglycan (HS-GAG) is essential for maintaining a niche that supports alveolar formation by modulating both biophysical and biochemical cues. Gli1-CreER mediated deletion of HS synthase gene Ext1 in lung fibroblasts results in enlarged and simplified alveolar structures. Ablation of HS results in loss of a subset of PDGFRαhi αSMA+ alveolar myofibroblasts residing in the distal alveolar region, which exhibit contractile properties and maintain WNT signaling activity to support normal proliferation and differentiation of alveolar epithelial cells. HS is essential for proliferation while preventing precocious apoptosis of alveolar myofibroblasts. We show that these processes are dependent upon FGF/MAPK signaling and forced activation of MAPK/ERK signaling partially corrected alveolar simplification and restored alveolar myofibroblast number and AT2 cell proliferation in HS deficient mice. These data reveal HS-dependent myofibroblast heterogeneity and function as an essential orchestrator for developing alveolar niche critical for the generation of gas exchange units.
Similar content being viewed by others
Introduction
Following the transition to air breathing, millions of alveoli are formed within the primary alveolar saccules to maximize the surface area for efficient gas exchange. Alveolar morphogenesis mainly depends on extensive structural remodeling coordinated by proliferation, differentiation, and cell-cell communications among multiple cell types, including mesenchymal, endothelial, and epithelial cells1,2. The arrest of alveolarization has been implicated in the pathogenesis of neonatal respiratory diseases such as bronchopulmonary dysplasia (BPD), a chronic lung disease of prematurity associated with impaired lung function and long-term morbidity3.
Formation and maturation of alveoli are directed and accompanied by dynamic changes in biochemical and biophysical cues2,4,5,6,7. During this process, alveolar type 1 (AT1) epithelial cells become thin and expansive, and migrate in proximity with endothelial cells, whereas alveolar type 2 (AT2) cells secrete surfactant proteins to reduce the surface tension of the alveoli. Recent advances in single-cell genomics and genetic mouse models have highlighted the developmental origin, cellular heterogeneity, and interactions among multiple lineages in alveolarization7,8,9. Despite the complexity of the contribution of each of the many cell types in alveologenesis, at least two major populations of fibroblasts were identified residing in the postnatal alveolar region in murine lung, including the Wnt2+ fibroblasts (also known as alveolar fibroblast 1) and the Tgfbi+ myofibroblasts8,10, with distinct cellular properties and lineage origins. Wnt2+ fibroblasts have characteristics of lipofibroblasts and express niche factors such as FGF7 and FGF10. In contrast, myofibroblasts are descendants of airway smooth muscle cells in late-stage lung development and serve as hallmarks of alveolar septation7,9,11. Myofibroblasts are contractile and produce elastin networks to establish alveolar scaffolds6. Recent studies have revealed two molecularly and morphologically distinct populations of myofibroblasts located in alveolar ducts and distal alveoli, respectively9,12. The latter, known as alveolar myofibroblasts or secondary crest myofibroblasts (SCMF), undergo developmental apoptosis and are cleared by immune cells at the end of alveolar septation12,13. Myofibroblasts can respond to SHH ligands sourced from AT1 epithelial cells7. A subset of hedgehog responsive PDGFRα+ myofibroblasts has also been proposed as critical niche cells in developing lungs to support the proliferative expansion of AT2 epithelial progenitors14. Our previous study and others have shown that myofibroblasts play a crucial role in producing extracellular matrix (ECMs) and mechanical forces that guide the formation of alveoli6,15. Ablation of PDGFRα+ fibroblasts or inhibition of growth factor signalings in myofibroblasts disrupts alveolar development5,15,16,17,18. While the cellular heterogeneity of lung mesenchymal cells is increasingly known, there is a paucity of information on signaling pathways that govern mesenchymal cell function and mediate interactions between cell types during alveolar morphogenesis.
While fibroblasts actively produce ECM components during lung development, cues from ECM can, in turn, impact cell identity or function. Heparan sulfate proteoglycans (HSPGs) are major components of the matrix proteins. Normal functions of HSPGs are largely dependent upon the heparan sulfate (HS) linear polysaccharide, which is attached to ECM and cell surface19. HS has been shown to exhibit dynamic affinities with signaling molecules such as FGF, WNT, SHH and BMPs to regulate their diffusion20. HS can also function as a coreceptor for FGF ligands and pathogens such as SARS-COV221,22. We have previously demonstrated that epithelial HS is required to produce active SHH forms during early lung development23. The role of mesenchymal HS in alveolar development remains unclear.
In the current study, we employ genetic mouse models and single-cell transcriptomics to interrogate the role of fibroblast-derived HS in pulmonary alveolarization. Postnatal ablation of the Ext1, which encodes for the HS synthase enzyme exostosin-1, resulted in alveolar simplification. We show that HS is required for maintaining the normal number and function of myofibroblasts for alveolar niche homeostasis. Upon loss of HS, a subset of PDGFRαhi αSMA+ alveolar myofibroblasts were selectively lost, and the remaining alveolar myofibroblasts exhibited impaired contractile gene expression and secreted less WNT11 that supports normal AT2 cell proliferation. We demonstrate that HS-dependent FGF/MAPK/ERK signaling is essential for the proliferation and function of alveolar myofibroblasts, which support AT2 cell growth, as well as for preventing their apoptosis. Restoration of MAPK signaling activity mitigated the loss of alveolar myofibroblasts and restored AT2 cell proliferation in HS-deficient lungs. Our study sheds light on how ECM molecules such as HS can regulate myofibroblast heterogeneity and cellular properties, thereby preserving the biophysical and biochemical cues of the alveolar niche essential for normal lung structure and function.
Results
HS level increases with the emergence of αSMA in PDGFRα+ fibroblasts during alveolar morphogenesis
Previous studies have identified two major fibroblast populations including alveolar fibroblasts marked by Wnt2 expression and myofibroblasts marked by Tgfbi expression8,10. Myofibroblasts play critical roles in secondary septation, an iterative process that generates numerous alveoli in terminal sacs. To determine the unique pathways that may be important in this process, we first compared the differentially expressed genes between Wnt2+ alveolar fibroblasts and myofibroblasts using previously published single-cell RNA-seq datasets from postnatal day 5 (P5) lungs (GSE160876 and GSE165063)8. We found that genes enriched in myofibroblasts are associated with pathways for ECM organization and smooth muscle contraction, which are consistent with the contractile nature of this population (Supplementary Fig. 1a). ECM-associated REACTOME terms related to HS-GAG functions were overrepresented in the functional enrichment analysis (Supplementary Fig. 1a). The HS polymerase complex is composed of proteins of the EXT family, which adds alternating units of glucuronic acid (GlcA) and N-acetylglucosamine (GlcNAc) to the nonreducing end of the growing chain24. To gain an overview of HS distribution in lung morphogenesis, we mapped the dynamic changes of Ext family genes involved in HS synthesis24. While the major components of the HS polymerase complex, Ext1, and Ext2, are broadly expressed in the developing lung, the expression of HS synthesis-associated genes is at a notably higher level in mesenchyme compared to the endothelium and epithelium compartments (Supplementary Fig. 1b–d). Time course analysis the mesenchyme population revealed that EXT family genes are highly expressed in the early postnatal period from P3 to P7, time points corresponding to active alveolar septation (Supplementary Fig. 1e). There is a slight decrease in expression of some critical genes such as Ext1 and Ext2 in mature lungs at P64 (Supplementary Fig. 1e).
The temporal increase in the expression of HS genes led us to test HS expression in PDGFRα+ fibroblast lineages, which are previously reported active and critical for alveolar septation16. While HS is highly expressed in alveolar epithelium marked by AT1 membrane marker AGER (Supplementary Fig. 2a), an increase of HS staining activity was notable at P4, P6 and P8 in mesenchyme, as demonstrated by colocalization of these HS signals with fibroblast markers, PDGFRα and COL1A1 (Supplementary Fig. 2b and c). Intriguingly, the increased intensity of HS staining in PDGFRα+ cells was correlated with the emergence of myofibroblast marker αSMA (Supplementary Fig. 2b). The staining of HS in PDGFRα+ cells decreased along with the disappearance of αSMA staining, detected at P15 and P21, time points corresponding to the end of the septation process (Supplementary Fig. 2b).
HS synthesis in fibroblasts is required for alveolar morphogenesis
To directly test the role of HS in lung alveologenesis, we generated a conditional knockout mice model using the previously defined Gli1CreERT2 and Ext1flox to specifically inactivate the Ext1 gene in fibroblast lineages. Newborn pups were administrated with tamoxifen at P0 and P1 to induce recombination (Fig.1a). The resulting conditional knockout mice (genotyped as Ext1flox/flox; Gli1CreERT2 with or without Rosa26tdTomato reporter) were referred to as Ext1Gli1∆/∆. We confirmed that two doses of tamoxifen treatment of Gli1CreERT2; Rosa26tdTomato mice at P0 and P1 induced widespread Cre activities in fibroblasts including the previously defined CDH4+ ductal myofibroblasts and PDGFRα+ lineages which includes alveolar fibroblasts and alveolar myofibroblasts (Supplementary Fig. 3a)12. Quantitative PCR (qPCR) analysis further confirmed loss of Ext1 expression in Magnetic Activated Cell Sorting (MACS) sorted PDGFRα+ fibroblasts (Supplementary Fig. 3b). Staining of HS with fibroblast markers confirmed ablation of HS in alveolar fibroblasts (PDGFRα+αSMA-), adventitial fibroblasts (PDGFRα+αSMA- cells surrounding airway, also known as alveolar fibroblast 2), alveolar myofibroblasts (PDGFRα+αSMA+), and ductal myofibroblasts (CDH4+αSMA+) from Ext1Gli1∆/∆ lungs (Supplementary Fig. 3c–e).
a Experimental design for tamoxifen-induced postnatal deletion of Ext1 using Gli1CreERT2. b Representative Hematoxylin & Eosin staining images of lung sections at multiple developmental time points, scale bars = 100 μm. c Morphometric quantification of mean linear intercept (MLI), n = 5 for control and Ext1Gli1∆/∆ at P6 and P8; n = 4 for control and 3 for Ext1Gli1∆/∆ at P15; n = 7 for control and 3 for Ext1Gli1∆/∆ at P21. d Lung volume measurement, n = 5 for control and Ext1Gli1∆/∆ at P6 and P8; n = 4 for control and 3 for Ext1Gli1∆/∆ at P15; n = 7 for control and 3 for Ext1Gli1∆/∆ at P21. e Maximum intensity projection of confocal z-stacks of AGER staining of P8 lungs, scale bars = 50 μm. f, g Quantification of relative alveolar airspace volume and surface area from z-stacks, n = 4 for control and 6 for Ext1Gli1∆/∆. (h–i) Lung compliance and FVC measured by PV test and FV test, respectively, in P21 mice, n = 10 for control and 8 for Ext1Gli1∆/∆. Data are presented as mean values ± SD. All p-values were determined by an unpaired 2-tailed t test. Source data are provided as a Source Data file.
We monitored the alveologenesis process at multiple developmental stages from P6 to P21 in Ext1Gli1∆/∆ lungs. As shown in Fig. 1b, simplified alveolar structures and enlarged alveolar space were constantly presented in lungs from Ext1Gli1∆/∆ mice. Arrest of alveolar development was quantified by morphometric analysis for Mean Linear Intercept (MLI), which was significantly higher in Ext1Gli1∆/∆ lungs compared to control lungs at all stages examined without significant alternations of lung volume (Fig. 1c and d). To better visualize changes in the alveolar structure, we performed wholemount confocal imaging for AGER, a membrane marker for AT1 cells. 3D reconstruction of image stacks revealed larger alveoli with fewer grooves in Ext1Gli1∆/∆ lungs, evidence of defective secondary septation (Fig. 1e). Consistent with MLI measurement, mutant lungs exhibited increased airspace volume and decreased alveolar surface area (Fig. 1f and g). Alveolar simplification was associated with increased lung compliance and forced vital capacity (FVC) at P21 in Ext1Gli1∆/∆ mice compared to controls, as detected by pulmonary function test (Fig. 1h and i). These results indicate that fibroblast-derived HS is indispensable for the formation of alveolar architecture essential for normal lung function.
HS synthesis in fibroblasts is required for normal alveolar epithelial cell development
We next performed Fluorescence-Activated Cell Sorting (FACS) analysis to identify cell composition changes in mutant lungs, which revealed a significant decrease of epithelial cells in Ext1Gli1∆/∆ lungs (Supplementary Fig. 4a–g). Previous marker-based stereology identified that AT2 cells are a major epithelial type that undergoes proliferative expansion, bulk generation of alveoli results in a 6-fold increase of AT2 cell number postnatally25, suggesting that the decreased epithelial cells are most likely AT2 cells. To identify the cellular and molecular differences between control and mutant lungs, we performed single-cell RNA sequencing (scRNA-seq) on P8 lungs. Unsupervised clustering of cells from control and Ext1Gli1∆/∆ mutant lungs identified all major cell types including epithelial, mesenchymal, endothelial and immune cells (Fig. 2a). While all cell populations were presented in both control and mutant lungs, we discovered that the proportion of AT2 cells and a subpopulation of alveolar myofibroblasts (Alv-MyoFb) were greatly diminished in mutant lungs (Fig. 2a and b).
a UMAP plot of cell types of control and Ext1Gli1∆/∆ lungs colored by clusters. b Fraction of each identified cell type color codes correspond to Fig. 2a. c UMAP plot of alveolar epithelial cells. (d) Fraction of each alveolar epithelial cell type, color codes correspond to Fig. 2c. e Dot plot showing the expression of alveolar epithelial genes in AT1, AT2, and AT1/AT2 cells. f Representative IF images of Pro-SP-C and Ki67 in lungs collected at P6, scale bars = 50 μm. Yellow arrowheads indicate Ki67+ Pro-SP-C+ cells. (g) Quantification of the percentage of Ki67+ Pro-SP-C+ cells. n = 5 for control and Ext1Gli1∆/∆. h Representative IF images of Pro-SP-C, HOPX, and NKX2-1 in lungs collected at P8, scale bars = 25 μm. i Quantification of proportions of AT1 (HOPX+ NKX2-1+) and AT2 (Pro-SP-C+ NKX2-1+) cells in all epithelial cells (NKX2-1+) in alveolar regions of P8 mice, n = 8 for control and 6 for Ext1Gli1∆/∆. j Representative IF images of Pro-SP-C, HOPX, and NKX2-1 in lungs collected at P15, scale bars = 25 μm. k Quantification of proportions of AT1 (HOPX+ NKX2-1+) and AT2 (Pro-SP-C+ NKX2-1+) cells in all epithelial cells (NKX2-1+) in alveolar regions of P15 mice, n = 7 for control and 4 for Ext1Gli1∆/∆. Data are presented as mean values ± SD. All p-values were determined by an unpaired 2-tailed t test. Source data are provided as a Source Data file.
We first focused on AT2 cells since the loss of AT2 cells has been implicated in aberrant alveolar development and regeneration. Subsetting and reclustering of alveolar epithelial cells revealed a decreased proportion of AT2 cells and cells expressing both AT1 and AT2 markers, which were presumably identified as “bipotent progenitors”10, along with an increased proportion of the AT1 population (Fig. 2c–e). To examine whether the inactivation of HS disrupted the self-renewal capacity of AT2 cells, we co-stained lung sections with AT2 cell marker Pro-SP-C with Ki67 and discovered a significant decrease in the proliferation rate of AT2 cells in P6 Ext1Gli1∆/∆ lungs (Fig. 2f and g). Consistent with decreased AT2 cell proliferation, the percentage of Pro-SP-C marked AT2 cells decreased while that of HOPX marked AT1 cells increased in NKX2-1+ cells located in alveolar regions (Fig. 2h and i). The defects of AT2 cell number did not recover with the progression of alveolar development, as exemplified by the decrease in AT2 cell proportion observed at P15, a developmental stage that marks the end of alveolar septation in mice (Fig. 2j and k). These data suggest a cell non-autonomous role of mesenchymal HS in alveolar epithelial proliferation and differentiation during alveolar morphogenesis.
Loss of HS alters myofibroblast composition
Previous studies have shown that mesenchymal cells are essential for the development and repair of alveoli14,26,27,28. Since we have also observed a diminished number of a subset of myofibroblasts in mutant lungs, we then subsetted and reclustered the mesenchymal populations for a detailed scRNA-seq analysis. We prioritize our analysis on non-vascular mesenchymal cells because the Gli1CreERT2 line does not target PDGFRβ+ vascular mesenchymal cells15. Based on the recent axis-based classification of mesenchymal cells in the developing lung12, we identified 6 clusters including airway smooth muscle cells (ASMCs), ductal myofibroblasts (Ductal-MyoFb) and alveolar myofibroblasts (Alv-MyoFb) 1 and 2 in the epithelial axis and Wnt2+ alveolar fibroblasts (Alv-Fb) and adventitial fibroblasts (Adv-Fb) in the interstitial axis (Fig. 3a–c). Consistent with previous scRNA-seq analysis, we identified distinct marker genes for each of the clusters for ASMC, Ductal-MyoFb, and Alv-MyoFb (Fig. 3c). Within the two subpopulations of Alv-MyoFb, Alv-MyoFb-1 population expressed medium level of Alv-MyoFb marker Pdgfra along with low levels of ductal markers including Lgr6, Cdh4 and Hhip, whereas Alv-MyoFb-2 highly expressed Pdgfra (Fig. 3c). While cell number for Wnt2+ Alv-Fb and Adv-Fb in the interstitial axis were maintained, we identified a notable reduction of the Alv-MyoFb-2 population defined by Pdgfrahi Acta2+ in Ext1Gli1∆/∆ lungs. In contrast, the Ductal-MyoFb population, which is defined by Cdh4+ Acta2+Hhip+, and reportedly residing in the alveolar duct, was generally retained (Fig. 3b, c).
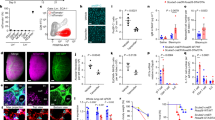
a UMAP plot of mesenchymal cell types of control and Ext1Gli1∆/∆ lungs colored by clusters. b Fraction of each mesenchymal cell type, color-coded as Fig. 3a. c Dot plot showing the expression of mesenchymal marker genes in fibroblasts. d Representative projection view of confocal z-stacks of αSMA, CDH4, and PDGFRα staining of P8 lungs. Z-plane projection is shown on the right side. P, proximal; D, distal. White arrowheads indicate PDGFRαhi αSMA+ cells (Alv-MyoFb-2), yellow arrowheads indicate PDGFRα+ αSMA+ CDH4+ (Alv-MyoFb-1) cells and red arrowheads indicate CDH4+αSMA+ (Ductal-MyoFb) cells. e, f Quantification of αSMA network volume and intensity in Fig. 3d, n = 3 for control and Ext1Gli1∆/∆. g Representative section IF of αSMA and PDGFRα staining of P8 lungs. Yellow arrowheads indicate PDGFRαhi αSMA+ cells. h Quantification of the percentage of PDGFRαhi αSMA+ cells in all αSMA+ cells, n = 3 for control and Ext1Gli1∆/∆. Scale bars = 25 μm for all images. Data are presented as mean values ± SD. All p-values were determined by an unpaired 2-tailed t test. Source data are provided as a Source Data file.
We employed 3D wholemount staining to label ductal (CDH4) and alveolar (PDGFRα) myofibroblast markers. Consistent with recent morphological characterization of myofibroblasts in developing lungs12, strong PDGFRα signals were always presented as the perinuclear pattern (Fig. 3d). We detected strong and perinuclear PDGFRαhi cells that correspond to the Pdgfrahi Acta2+ Alv-MyoFb-2 population in the distal alveolar region of the control lungs (Fig. 3d). In contrast, PDGFRαhi αSMA+ cells were fewer in alveolar region of Ext1Gli1∆/∆ lungs, and alveoli of Ext1Gli1∆/∆ lungs were populated with myofibroblasts with diffused PDGFRα staining and CDH4 expression, which were presumably Alv-MyoFb-1 (Fig. 3d). The Z-plane view of the confocal stack confirmed that PDGFRα signals were localized to the distal region, while CDH4 signals were found in the proximal region (Fig. 3d). In addition, there was a loss of the normal proximal-distal pattern of ductal and alveolar myofibroblast markers in Ext1Gli1∆/∆ lungs (Fig. 3d). The distribution of αSMA appeared more diffuse in the mesenchyme, but it remained primarily localized to the alveolar entrance rings and septa (Fig. 3d). While intensity for αSMA network did not change, the volume of αSMA network was significantly reduced in Ext1Gli1∆/∆ lungs (Fig. 3e and f). Quantification of αSMA+ and perinuclear PDGFRα+ cells on section stainings confirmed a 2-fold decrease in cell proportion of PDGFRαhi αSMA+ cells in P8 Ext1Gli1∆/∆ lungs (Fig. 3g and h).
Consistent with the above finding, isolated HS deficient PDGFRα+ fibroblasts showed fewer PDGFRα+ αSMA+ cells in cytospin analysis, and those cells were less efficient to contract the collagen gel in vitro (Supplementary Fig. 5a–f). During alveolarization, the deposition of elastin fibers and collagen fibers is largely dependent on myofibroblasts. 3D immunostaining showed that elastin fibers are presented at an equivalent level in the alveolar entrances (Supplementary Fig. 6a). In addition, staining for collagen fibers with Sirius Red did not reveal a detectable change between control and mutants in septa, suggesting a minimal contribution of HS for alveolar ECM remodeling and deposition (Supplementary Fig. 6b). These data demonstrate that HS is an essential regulator for myofibroblast composition, loss of which leads to loss of PDGFRαhi αSMA+ alveolar myofibroblasts within the developing alveolar niche.
HS maintains cell number and the contractile gene expression of alveolar myofibroblasts
To assess how HS affected alveolar myofibroblast function and composition, we next compared the differentially expressed genes in the remaining Alv-MyoFb from Ext1Gli1∆/∆ vs. those from control lungs. Gene Set Enrichment Analysis (GSEA) revealed increased gene signatures associated with hallmarks such as “Apoptosis” and the “P53 pathway”, and a decreased signature related to “E2F targets,” which are involved in cell proliferation, in HS-deficient Alv-MyoFb (Fig. 4a). Particularly, we found several pro-apoptotic genes, including the prominent P53 target, Cdkn1a, were among those upregulated in mutant Alv-MyoFb (Fig. 4b).
a GSEA analysis of differentially expressed genes in Alv-MyoFb in Ext1Gli1∆/∆ lungs compared to control lungs. b Dot plot showing the increased expression of apoptosis-related genes in Alv-MyoFb in Ext1Gli1∆/∆ lungs compared to control lungs. c Representative IF images of αSMA, PDGFRα, and cleaved-Caspase-3 (CASP3) in lungs collected at P6, P8, and P12, scale bars = 25 μm. Yellow arrowheads indicate CASP3+ PDGFRα+ αSMA+ cells. d Quantification of percentage of CASP3+ PDGFRα+ αSMA+ cells in total PDGFRα+ αSMA+ cells. n = 3 for control and Ext1Gli1∆/∆ at P6 and P8, n = 4 for control at P12. e Representative IF images of αSMA, PDGFRα, and Ki67 in lungs collected at P6 and P8, scale bars = 25 μm. Yellow arrowheads indicate Ki67+ PDGFRα+ αSMA+ cells. f Quantification of the percentage of Ki67+ PDGFRα+ αSMA+ cells in total PDGFRα+ αSMA+ cells. n = 4 for control and Ext1Gli1∆/∆ at P6, and n = 3 for control and Ext1Gli1∆/∆ at P8. Data are presented as mean values ± SD. All p-values were determined by an unpaired 2-tailed t test. Source data are provided as a Source Data file.
Alv-MyoFb are transiently presented during secondary septation and are cleared by a developmental apoptosis process starts approximately from P1212,13. To confirm whether HS prevents Alv-MyoFb apoptosis, we performed cleaved-caspase-3 staining. In control lungs, rare apoptotic PDGFRα+ αSMA+ cells were found at P6 and P8, and apoptosis was more frequently detected on P12; in contrast, deletion of Ext1 triggered an early emergence of apoptosis marker cleaved-caspase-3 in PDGFRα+ αSMA+ cells detected at P6 and P8 (Fig. 4c and d). Hardly any PDGFRα+ αSMA+ cells were presented in P12 Ext1Gli1∆/∆ lungs, which makes the apoptotic rate of this population non-quantifiable (Fig. 4c and d). Consistent with increased apoptosis, fewer HS deficient PDGFRα+ αSMA+ cells expressed proliferation marker Ki67 at P6 and P8 (Fig. 4e and f). The above data suggest that HS is required for the maintenance of the Alv-MyoFb population by preventing apoptosis and sustaining proliferation. Moreover, we noticed that increased apoptosis was associated with the loss of myogenic features. Genes related to myogenic property and actomyosin contractility, such as Tagln, Myl9, Myl6 and Mustn1 were downregulated in Alv-MyoFb from Ext1Gli1∆/∆ lungs (Supplementary Fig. 6c). Wholemount imaging confirmed less TAGLN activity in Ext1Gli1∆/∆ lungs (Supplementary Fig. 6d and e), suggesting that decreased contractile gene expression might also partially contributes to the reduced contraction of HS-deficient bulk PDGFRα+ fibroblasts in vitro (Supplementary Fig. 5e and f).
Since Gli1CreERT2 also targets Ductal-MyoFb, Wnt2+ Alv-Fb, and Adv-Fb, we compared gene expression profiles of control and Ext1Gli1∆/∆ lungs of each cell population using GSEA. While stress-related and hypoxia genes were activated in those cells from Ext1Gli1∆/∆ lungs, there was no significant enrichment for apoptosis-related pathways (Supplementary Data 1). These results are in accordance with the retained cell number of other populations (Fig. 3a and b). Together, these data demonstrate a selective requirement of HS for the maintenance of the Alv-MyoFb number and function by promoting proliferation, preventing precocious apoptosis, and preserving its myogenic gene expression for biophysical properties required for alveolar formation.
Disruption of myofibroblast heterogeneity is associated with loss of a WNT11 paracrine niche
To directly test the ability of HS-regulated myofibroblast heterogeneity to support alveolar progenitor proliferation and differentiation, we employed a 3D organoid system, in which we co-cultured wild-type AT2 cells (isolated from SftpcCreERT2, Rosa26tdTomato mice at P6) with MACS sorted PDGFRα+ fibroblasts from control and Ext1Gli1∆/∆ lungs (Fig. 5a). AT2 cells co-cultured with Ext1-deleted fibroblasts showed a decreased average organoid size compared to cells cultured with control fibroblasts (Fig. 5b and c). Immunofluorescence staining revealed that there was a decreased proportion of AT2 cells and an increased proportion of AT1 cells in alveolar organoids formed with Ext1-deleted fibroblasts (Fig. 5b and c). These data suggest that the loss of Alv-MyoFb-2 subpopulation accounted, at least partially, for the decreased AT2 cell proliferation.
a Schematics of workflow for AT2 cell-derived alveolar organoid assay with fibroblast co-culture. b Whole well scan for tdTomato+ alveolar organoids cultured for 21 days (upper panel) and IF for Pro-SP-C and HOPX on organoid sections (lower panel). Scale bar: 1 mm for tdTomato whole well scan, and 25 μm for section IF. c Quantification of average tdTomato+ organoid size (n = 7 for control and 10 for Ext1Gli1∆/∆), and percentages for Pro-SP-C+ HOPX+ and Pro-SP-C+ HOPX+ cells in sections from organoids (n = 8 for control and 7 for Ext1Gli1∆/∆). d Volcano plot displaying differentially expressed genes (DEG) in AT2 cells. e Violin plot showing the Wnt11 expression level in mesenchymal populations. f Violin plot showing the Wnt11 expression level in Alv-MyoFb of control and Ext1Gli1∆/∆ lungs. g Representative images of RNAscope assay for Axin2 and Wnt11 along with IF of Pro-SP-C and αSMA. h Quantification of the percentage of Axin2+ AT2 cells, n = 3 for control and Ext1Gli1∆/∆. Scale bars = 25 μm for all images. i Schematics and representative whole well scan images for AT2 cell-derived alveolar organoid cultured for 14 days with SFFF culture system. j Quantification of average tdTomato+ organoid size, n = 9 for -CHIR, 8 for -CHIR; +WNT11, 6 for + CHIR, and 5 for + CHIR + WNT11. Data are presented as mean values ± SD. p-values in c and h were determined by an unpaired 2-tailed t test. p-values in j were determined by the ANOVA test followed by the multiple comparisons test (Sidak). Source data are provided as a Source Data file.
To gain an insight into the molecular changes underlying alveolar epithelial proliferation and differentiation defects, we performed differential expression analysis on AT2 cells from the scRNA-seq data (Fig. 5d). Consistent with the previous study demonstrating that the activation of WNT signaling promotes AT2 self-renewal during alveolar formation4, WNT signaling/target genes associated with progenitor function, such as Fgfr2, Ctnnbip1, Clu and Znrf3, were downregulated in AT2 cells in KO lungs (Fig. 5d)29,30,31,32,33. Contrastly, genes associated with mature AT2 functions, such as Retnla, Chia1, and Lamp3 were increased (Fig. 5d)34. Consistent with increased AT1 cell proportion, there was increased expression of AT1 differentiation genes such as Hopx and Scnn1a in Ext1Gli1∆/∆ AT2 cells (Fig. 5d)25. These data demonstrate a potential loss of WNT activity, progenitor activity, and an increased potential of alveolar epithelial maturation in Ext1Gli1∆/∆ AT2 cells.
The defects in AT2 cell proliferation and differentiation observed in mutant lungs prompted us to investigate whether disruption of genes associated with WNT signaling in alveolar myofibroblasts impacts the self-renewal of alveolar epithelial cells. While the previous study identified mesenchymal Wnt5a as the main paracrine factor for regeneration-associated alveolar niche35, we found that Wnt5a and Wnt9a are mainly expressed by Alv-MyoFb1 and Ductal-MyoFb (Supplementary Fig. 6f). Intriguingly, we observed a predominant expression of Wnt11, a ligand that induces both canonical and non-canonical WNT activities36,37,38,39,40, in Alv-MyoFb population with a higher expression level in the Alv-MyoFb-2 cluster (Fig. 5e). Moreover, Wnt11 was remarkably decreased in the remaining Alv-MyoFb cells in mutant lungs (Fig. 5f). Single-molecule RNA in situ hybridization (smRNA-ISH) analysis confirmed a reduced proportion of Axin2 + AT2 cells and almost a complete loss of Wnt11 expression in remaining myofibroblasts (Fig. 5g and h).
To test the function of WNT11 for AT2 self-renew, we isolated WT AT2 cells from SftpcCreERT2, Rosa26tdTomato mice at P6 and subjected them to a serum-free feeder-free (SFFF) organoid culture system. We found that the addition of recombinant WNT11 increased the average size of organoids cultured without canonical WNT signaling activator CHIR, and WNT11 did not further increase organoid size upon activation of WNT by CHIR, suggesting that WNT11 promotes AT2 cell self-renew, at least partially, through canonical WNT pathway (Fig. 5i and j).
We also tested whether the expression level of growth factors in other mesenchymal cell types was reduced since Wnt2+ Alv-Fb was identified as niche cells for alveolar regeneration41. However, we found that Fgf7, Fgf10 and Wnt2, which support AT2 self-renew42,43, were highly expressed in Wnt2+ Alv-Fb and Adv-Fb of Ext1Gli1∆/∆ lungs; whereas Bmp4, which inhibits AT2 proliferation, was expressed at a lower level (Supplementary Fig. 6g)44, suggesting that reduced AT2 cell proliferation in Ext1Gli1∆/∆ lungs is not likely caused by decreased production of niche factors from Wnt2+ Alv-Fb. Taken together, these data demonstrate that HS-dependent myofibroblast heterogeneity and function are essential for maintaining a specialized alveolar niche promoting AT2 self-renew associated with WNT11 paracrine function during alveolar formation.
HS is required for FGF/MAPK signaling in alveolar myofibroblasts
We then asked what molecular programs mediate the function of HS to maintain homeostasis of the alveolar niche. Pathway activity inference from scRNA-seq data suggests that MAPK signaling was mostly active in Alv-MyoFb population, and Alv-MyoFb-2 cluster has a higher level of MAPK signaling activity than that of Alv-MyoFb-1 (Fig. 6a and b). HS has been shown to function as co-receptor of FGFR, our data demonstrate that Alv-MyoFb-2 expressed higher levels of FGF target genes such as Etv1, Etv4 and Anxa2 (Supplementary Fig. 7a)45,46. Further analysis of FGF target genes in a previously published mouse lung developmental atlas dataset revealed that in the Alv-MyoFb-2 population, Etv4 and Anxa2 expression emerged readily after birth and diminshed at P14, suggesting a putative role of FGF/MAPK signaling for Alv-MyoFb development (Supplementary Fig. 7b and c). Consistent with the emergence of FGF targets, Wnt11 expression also emerged while the expression of secreted WNT inhibitor Notum was inhibited after birth (Supplementary Fig. 7c), suggesting a possible role of alveolar myofibroblast specific activation of local WNT signaling by FGF/MAPK.
a Heatmap of pathway activity of each cell cluster inferred from scRNA-seq data. b Feature plot of MAPK activity distribution in fibroblasts. c Homer motif analysis of less accessible genomic regions in Ext1Gli1∆/∆ compared to control lungs, as identified in Rosa26-tdTomato+ cells with ATAC-seq. d Violin plot showing the expression level of FGF target genes (Etv4 and Anxa2) and antagonists (Spry1 and Dusp1). e Representative IF images of αSMA, PDGFRα, and p-ERK staining in P6 lungs, scale bars = 25 μm. White arrowheads indicate cells with no nuclear p-ERK staining and yellow arrowheads indicate cells with nuclear p-ERK staining. f Quantification of p-ERK intensity of αSMA+ PDGFRα+ cells from control and Ext1Gli1∆/∆ lungs. Each dot represents a cell, data were obtained from 3 controls and 3 Ext1Gli1∆/∆ mutants, and at least 300 cells were measured for each group. g Representative Hematoxylin & Eosin staining (upper panel) and corresponding maximum intensity projection view of confocal z-stacks of AGER staining (lower panel) of P8 lungs, scale bars = 100 μm for Hematoxylin & Eosin staining and 50 μm for AGER staining. h MLI quantification of each group. n = 5 for control, 5 for Ext1Gli1∆/∆ and 8 for Ext1Gli1∆/∆; KrasLSL-G12D. i, j Quantification of relative alveolar airspace volume and surface area from z-stacks, n = 4 for each group. Data are presented as mean ± SD, except for the scatter dot plot in (f), which does not include error bars. p-value in (f) was determined by an unpaired 2-tailed t test. p-values in (h–j) were determined by the ANOVA test followed by the multiple comparisons test (Tukey). Source data are provided as a Source Data file.
We speculate that HS-dependent extracellular signaling may induce profound changes in the cell-autonomous transcriptional program, which could help identify the signaling pathways that play a dominant role in modulating myofibroblast fate. Thus, we performed bulk ATAC-seq from the Rosa26-tdTomato sorted cells from control and Ext1Gli1∆/∆ lungs, which demonstrates a widespread decrease of chromatin accessibility in Ext1Gli1∆/∆ fibroblasts (Supplementary Fig. 8a). In support of the decrease of myogenic signature genes, sites with less chromatin accessibility annotated genes that are critical for GO terms such as “actin filament organization” and “muscle tissue development” (Supplementary Fig. 8b). Decreased chromatin accessibilities were detected in genes including Myl9, Tagln and Wnt11 (Supplementary Fig. 8c–e). Importantly, Homer analysis revealed that these sites contain multiple similar motifs for ETS and ETV family members (Fig. 6c). Consistent with this finding, FGF target genes Etv4 and Anxa2 were decreased, whereas expression of antagonists for FGF signaling such as Spry1 and Dusp1 were increased in the Alv-MyoFb population of Ext1Gli1∆/∆ lungs (Fig. 6d)47,48. These data indicate a suppressed FGF transcriptional program in mutant Alv-MyoFb. In support of the scRNA-seq data, immunostaining for p-ERK suggests that PDGFRα+ αSMA+ cells in Ext1Gli1∆/∆ lungs have less MAPK signaling as evidenced by reduced nuclear p-ERK staining, which was confirmed by quantification of p-ERK intensity within these cells (Fig. 6e and f). These results demonstrate that HS is essential for the emergence of FGF/MAPK signaling in Alv-MyoFb during alveolar morphogenesis.
Activation of MAPK signaling alleviates alveolarization defects
MAPK signaling is closely related to cell mitotic activity, to test whether manipulation of MAPK signaling can influence the proliferation of alveolar myofibroblasts and AT2 cells, we crossed Gli1CreERT2 and KrasLSL-G12D mice and administered tamoxifen at P0 and P1 to the newborn pups. Gli1CreERT2-mediated excision of the stop elements enabled the expression of a constitutively active mutant of KRAS in fibroblasts. Immunofluorescence staining confirmed that p-ERK intensity was significantly increased in PDGFRα+ αSMA+ cells with KRAS activation (Supplementary Fig. 9a and b). Expression of KrasG12D did not alter PI3K activity in PDGFRα+ αSMA+ cells, as demonstrated by the unchanged p-AKT level (Supplementary Fig. 9c and d). There was a subtle increase of cellularity in Gli1CreERT2; KrasLSL-G12D mice at P8, morphometric analysis revealed a non-significant decrease of MLI (mean value of 41.35 for Gli1CreERT2 vs. 40.79 for Gli1CreERT2; KrasLSL-G12D, p = 0.0817). Gli1CreERT2 and Gli1CreERT2; KrasLSL-G12D mice have comparable lung architecture, as demonstrated by MLI at P15 and P21. (Supplementary Fig. 9e and f). A similarly rare occurrence of apoptosis was observed in PDGFRα+ αSMA+ cells at P8 (Supplementary Fig. 9g and h). While a subtle decrease in apoptosis of PDGFRα+ αSMA+ cells was noted at P15 (p = 0.0740) in Gli1CreERT2; KrasLSL-G12D mice, no PDGFRα+ αSMA+ cells were detected in both Gli1CreERT2 and KrasLSL-G12D mice at P21, indicating that KRAS activation did not disrupt clearance of these cells (Supplementary Fig. 9g and h). Increased proliferation of PDGFRα+ αSMA+ cells, as well as a higher percentage of PDGFRαhi αSMA+ cells were observed in Gli1CreERT2; KrasLSL-G12D lungs at P8 (Supplementary Fig. 9i–k). Concurrently, the proliferation of AT2 cells was also increased (Supplementary Fig. 9l and m). These data suggest that MAPK activation is sufficient to drive alveolar myofibroblast proliferation.
To test whether mesenchymal FGF/MAPK signaling is functionally relevant to HS-mediated alveolar morphogenesis, we introduced the KrasLSL-G12D allele into the mutant mice line. KRAS activation in Ext1Gli1∆/∆ mice activated MAPK/ERK signaling regardless of the blockade of HS synthesis (Supplementary Fig. 10a–c). Similarly, PI3K signaling, as demonstrated by p-AKT staining, was not altered in compound mutants (Supplementary Fig. 10d and e). Expression of KRASG12D in Ext1-deleted fibroblasts significantly attenuated the alveolar simplification phenotype compared to Ext1Gli1∆/∆ mice alone (Fig. 6g–j). Quantification of immunofluorescence stainings demonstrated that Ext1Gli1∆/∆; KrasLSL-G12D compound mutants had an overall higher proliferation rate of AT2 cells compared to Ext1Gli1∆/∆ mice (Fig. 7a and b), and introducing the KrasLSL-G12D allele into Ext1Gli1∆/∆ mutants also partially corrected alveolar AT1-AT2 epithelial cell differentiation (Fig. 7c and d).
a Representative IF images of Pro-SP-C and Ki67 in Ext1Gli1∆/∆ and Ext1Gli1∆/∆; KrasLSL-G12D lungs collected at P8. Yellow arrowheads indicate Ki67+ AT2 cells. b Quantification of the percentage of Ki67+ Pro-SP-C+ cells, n = 8 for Ext1Gli1∆/∆ and 5 for Ext1Gli1∆/∆; KrasLSL-G12D. c Representative IF images of Pro-SP-C, HOPX, and NKX2-1 in lungs collected at P8, scale bars = 25 μm. d Quantification of proportions of AT1 (HOPX+ NKX2-1+) and AT2 (Pro-SP-C+ NKX2-1+) cells in all epithelial cells (NKX2-1+) in alveolar regions, n = 4 for Ext1Gli1∆/∆ and 7 for Ext1Gli1∆/∆; KrasLSL-G12D. e Representative IF images of αSMA, PDGFRα, and cleaved-Caspase-3 (CASP3) on Ext1Gli1∆/∆ and Ext1Gli1∆/∆; KrasLSL-G12D lungs collected at P8. f Quantification of the percentage of PDGFRαhi αSMA+ cells in total αSMA+ cells and (g) apoptotic PDGFRα + αSMA+ cells. n = 7 for Ext1Gli1∆/∆ and 6 for Ext1Gli1∆/∆; KrasLSL-G12D. h Representative IF images of αSMA, PDGFRα, and Ki67 in lungs collected at P8, scale bars = 25 μm. Yellow arrowheads indicate Ki67+ PDGFRα+ αSMA+ cells. i Quantification of the percentage of Ki67+ PDGFRα+ αSMA+ cells in total PDGFRα+ αSMA+ cells. n = 5 for Ext1Gli1∆/∆ and Ext1Gli1∆/∆; KrasLSL-G12D. j Representative images of RNAscope assay for Axin2 and Wnt11 along with co-staining of Pro-SP-C and αSMA in P8 lungs from Ext1Gli1∆/∆ and Ext1Gli1∆/∆; KrasLSL-G12D animals. k Quantification of the percentage of Axin2+ AT2 cells, n = 3 for Ext1Gli1∆/∆ and 4 for Ext1Gli1∆/∆; KrasLSL-G12D. Scale bars = 25 μm for all images. Data are presented as mean values ± SD. All p-values were determined by an unpaired 2-tailed t test. Source data are provided as a Source Data file.
To determine whether the decreased FGF/MAPK signaling is responsible for Alv-MyoFb loss, we quantified PDGFRαhi αSMA+ cells and observed a partial recovery of this population in Ext1Gli1∆/∆; KrasLSL-G12D lungs (Fig. 7e and f). In addition, PDGFRα+ αSMA+ cells of Ext1Gli1∆/∆; KrasLSL-G12D mice were less apoptotic and more proliferative than those in Ext1Gli1∆/∆ animals (Fig. 7e, g–i), confirming a cell-autonomous requirement of FGF/MAPK signaling to promote proliferation and prevent precocious apoptosis for alveolar development. smRNA-ISH assay demonstrated that there were more Axin2 + AT2 cells in Ext1Gli1∆/∆; KrasLSL-G12D lungs compared to those in Ext1Gli1∆/∆ lungs (Fig. 7j and k). Furthermore, the reoccurrence of Wnt11 expression in myofibroblasts was detected in lungs from Ext1Gli1∆/∆; KrasLSL-G12D animals (Fig. 7j), supporting that FGF/MAPK signaling is required for Alv-MyoFb to produce WNT11. Taken together, these findings demonstrate that HS mediates alveolar morphogenesis by maintaining the number and function of alveolar myofibroblasts in a process mediated, at least partially by FGF/MAPK signaling.
Discussion
The generation and normal function of alveolar structures depends upon coordinated actions among various cell types. Stromal cells produce ECM molecules to shape the alveolar scaffold and secret niche factors to support the proliferation and differentiation of epithelial and endothelial cells for gas exchange and homeostasis14,15,17,27,28,44,49,50. Our data supports that fibroblast-derived Heparan Sulfate Glycosaminoglycans (HS-GAGs) are essential for alveolar morphogenesis by maintaining alveolar myofibroblast number and function. We show that HS-dependent FGF/MAPK signaling is critical for alveolar myofibroblast proliferation, survival and maintaining the developing alveolar niche related to its function to produce WNT11 (Supplementary Fig. 11). Thus, the present study identifies HS as the core ECM molecules that maintain myofibroblasts heterogeneity, which confers the biophysical and biochemical cues required for structural remodeling and alveolar epithelial cell proliferation and fate selection.
ECM remodeling and mechano-transduction are closely associated, and accompany and regulate the generation and regeneration processes of various organs51,52. Alveologenesis is such a process that depends on biophysical forces mediated by the contractile myofibroblasts, a subset of which emerge postnatally and disappear when the lungs reach maturation. Myofibroblasts form “fishnet” structures and are embedded in the alveolar epithelial surface to divide the primary alveolar sacs into alveoli11. We and others have shown that growth factor signalings are part of the ECM remodeling program15,17,49. However, the role of myofibroblast heterogeneity and its regulatory machinery in alveolar morphogenesis are less well-known. HS is a major component of ECM and has been broadly implicated in developmental signaling pathways53. Our data establish HS as an essential factor in maintaining the alveolar myofibroblast population residing in the distal alveolar region, defined by Pdgfrahi Acta2+ in single-cell transcriptomics and immunofluorescence analysis. The Alv-MyoFb-2 population seen lost in Ext1Gli1∆/∆ lungs likely represents the same cell population termed “partitioning myofibroblasts” in another study as defined by Fgf18+ Lgr6-, which subdivide alveolar complexes after the formation of alveolar entrance9.
Recent studies have shown that impaired myofibroblast proliferation is a hallmark of alveolar simplification and that alveolar myofibroblasts undergo developmental apoptosis at the terminal stage of septation12,54. However, the factors regulating the proliferation and survival of alveolar myofibroblasts remain less well-known. Decreased number of alveolar myofibroblasts in HS-deficient lungs aligns with reduced proliferation and increased apoptotic rate of this population in Ext1Gli1∆/∆ lungs in the early alveolarization phase, suggesting an essential role of HS in the maintenance of niche homeostasis in alveolar complexes. Intriguingly, scRNA-seq analysis suggested a decreased transcriptional program for genes associated with contractility of myofibers in the remaining Alv-MyoFbs along with an enhanced activity P53-associated apoptosis pathway. It is unclear whether the loss of the myogenic feature is due to increased apoptotic stress or caused by other HS-regulated signaling pathways. Nonetheless, our data support the central role of HS in biophysical force generation, one of the key aspects of the myofibroblasts-directed septation process, by maintaining the cell number and myogenic gene expression of alveolar myofibroblasts.
Current data support a cell non-autonomous function of HS in regulating AT2 proliferation and differentiation. The defects of alveolar epithelial cells observed in Ext1Gli1∆/∆ animals are consistent with the recent study demonstrating myofibroblasts as niche cells supporting AT2 cell growth during development14. While the factors secreted by Wnt2+ Alv-Fb such as WNT2, FGF7, and FGF10 have been implicated for alveolar generation and regeneration27,43,55, their expression was not reduced in Wnt2+ Alv-Fb cells from HS deficient cells. Moreover, Ext1Gli1∆/∆ derived PDGFRα+ fibroblasts were less effective in expanding AT2 populations in the organoid assay, demonstrating the requirement of additional factors for AT2 proliferation and fate selection during alveolar formation. Given that Wnt2+ Alv-Fb are presented both in developing and adult lungs, selective expression of Wnt11 in the transient Alv-MyoFbs perhaps is associated with the highly proliferative nature of postnatal AT2 cells, which exhibit a 6-fold increase in cell number during the critical stage of alveolar morphogenesis25. Consistent with this notion, the addition of WNT11 in the AT2 SFFF organoid system augmented organoid growth, demonstrating a direct signaling communication between the alveolar myofibroblast population and AT2 cells. A recent human organoid study demonstrates that NOTUM+ myofibroblast inhibits the AT2 cell fate of distal tip progenitors during embryonic development55. Consistently, the decrease of the WNT inhibitor Notum, along with the emergence of Wnt11 in the Alv-MyoFb population after birth suggests a possible shift of myofibroblast identity/function after birth to facilitate WNT activation for bulk alveoli generation, a process likely controlled by HS-dependent FGF/MAPK signaling. While our data argue that HS-dependent Wnt11 expression in Alv-MyoFb modulates WNT responsiveness of AT2 cells, identifying additional factors that mediate the temporal-spatial cell-cell communication between myofibroblast populations and AT2 cells during alveolar formation will be of high priority for future study.
Lung mesenchymal cells are highly heterogeneous in transcriptional signatures, morphologies, and temporal and spatial locations. The heterogeneity of postnatal myofibroblast populations has been less well-known. Our single-cell transcriptomics data is consistent with the previous three-axis classification of mesenchymal cells in developing lungs12. In the epithelial axis, parallel to its accompanying epithelial cells, ASMC, Ductal-MyoFb, and Alv-MyoFb form a proximal-distal pattern. The specific requirement of HS-dependent FGF/MAPK signaling for the maintenance of distally located Alv-MyoFb population coincides with the role of FGF signaling on proximal-distal patterning seen in epithelial cells, where FGF activity is mostly restricted to distal tips23,42,46. Changes in mesenchymal proximal vs. distal identity have been implicated in the pathogenesis of emphysema, an adult lung disease with similar histopathological changes in BPD50. Our mouse model phenocopies multiple aspects of human BPD, suggesting a potential role for disrupted intercellular communication driven by aberrant HS-GAG synthesis and/or the loss of alveolar myofibroblast identity, in its pathogenesis.
The rescuing effects of KRAS activation on alveolar simplification phenotype provide proof of concept that increasing the cell number of Alv-MyoFb can help mitigate BPD. KRAS has been shown to activate two major signaling pathways, MAPK and PI3K, which drive cell proliferation in various cell types56. Our data demonstrate that KRAS activates MAPK/ERK but not PI3K/AKT in PDGFRα+αSMA+ cells, indicating that MAPK/ERK signaling is one of the main contributors to the proliferation and survival of alveolar myofibroblasts. However, additional signaling pathways may also play a role in mediating the proliferative effects of KRAS, which warrants further investigation. Since sustained activation of KRAS is oncogenic, the development of therapeutic strategies that promote a more physiologically relevant and controllable MAPK activation in alveolar myofibroblasts could offer novel therapeutic avenues for the treatment of BPD.
Our current study has several limitations. Like the most frequently used Pdgfra-driven Cre lines, the Gli1CreERT2 used by our study exhibited activity in all fibroblasts and thus may cause non-specific effects. We found that loss of HS only affected the cell number of the PDGFRαhi αSMA+ population, demonstrating a relatively specific role of HS for alveolar myofibroblast proliferation and function. Although the cell number and niche factor expression in Wnt2+ Alv-Fbs were unaffected, these cells exhibited elevated expression of stress-related genes following HS ablation, potentially due to disruption of the secretome caused by alterations in ECM composition. The identification of alveolar myofibroblast-specific genes within emerging scRNA-seq datasets is expected to facilitate the development of mouse models enabling more precise targeting of myofibroblast subpopulations. In addition, a significant portion of the HS present in the ECM is secreted by mesenchymal cells. However, we currently lack the ability to determine the extent to which the secreted HS is required for signal transduction in other cell types, such as AT2 cells, potentially influencing their proliferation. Unlike other ECM molecules, HS is often bound to core proteins and exhibits distinct glycosylation modifications. While we discovered striking alveolarization defects after ablating the HS side chain in the mesenchymal cells using genetic approaches, it is unknown which HSPGs play a central role in mediating myofibroblast FGF/MAPK signaling during alveolar morphogenesis. Earlier studies have suggested glypicans are required for FGF signaling transduction57. Identifying the specific roles of individual HSPGs or HS modifications in pulmonary alveologenesis should provide additional insights into the pathogenesis of BPD, which may allow the development of novel therapeutics to enhance the generation of alveoli in preterm infants.
Methods
Animals
All animal experiments were performed under the IACUC protocol 2021-054 approved by West China Second University Hospital of Sichuan University. Animals were housed under 12/12 h light/dark cycle with humidity levels between 40–60%. The Gli1CreERT2 (Strain No. 007913), and Rosa26tdTomato (Strain No. 007914) mouse lines were acquired from Jackson Laboratories (Bar Harbor, ME). SftpcCreERT2 mice were acquired from Dr. Hal Chapman58. Ext1flox mice were described previously23,59. KrasLSL-G12D (Strain No. T004551) mice were purchased from GemPharmatech (Nanjing, China). Mice were maintained on a mixed genetic background. To induce Cre activity, 100 μg tamoxifen (Aladdin Scientific, T137975) dissolved in corn oil was administrated to each pup via intraperitoneal injection on P0 and P1. Both male and female mice were used in all experiments and littermates of Ext1flox/flox and Ext1flox/+; Gli1CreERT2 genotypes were used as controls. Littermates carrying Gli1CreERT2 were used as control when Cre activity was needed to induce tdTomato.
Pulmonary function test
Mice aged at P21 were anesthetized with sodium pentobarbital before subjecting to the test. A pulmonary function test was conducted on a DSI Buxco PFT system, and compliance and FVC were measured with the Quasistatic Pressure Volume test and the Fast Flow Volume test, respectively.
Histology and MLI analysis
The lungs were harvested at indicated time points and perfused with cold PBS from the right heart ventricle to remove the residual blood, followed by gravity inflation of 2% PFA in PBS from the trachea at a constant pressure of 25 cm H2O. Tissues were fixed overnight at 4 °C. Lung volume was determined using Archimedes’ principle60. The fixed lungs were fully immersed in PBS, and the volume of displaced PBS was measured to represent the lung volume. Fixed lungs were washed with PBS and were then subjected to dehydration through 30, 50, 70, 90%, and 100% gradient ethanol. Tissues were cleared by xylene and embedded as paraffin blocks. Tissue sections were prepared at 5 μm in thickness for subsequent experiments. Standard hematoxylin and eosin (H&E) staining was performed, and images were taken on an Olympus APX-100 imager. Morphometric analysis for MLI was conducted by manual counting in FIJI software15. Hematoxylin and eosin (H&E) staining images were overlaid with a 15 × 11 grid using FIJI software to determine the number of intercepts between alveolar walls and the grid lines, both vertically and horizontally. Lines crossing large blood vessels and airways were excluded from the count. The Mean Linear Intercept (MLI) was calculated using the following equation: MLI = total line length / total number of intercepts.
Immunofluorescence
For section immunofluorescence staining, paraffin sections were rehydrated, and immersed in Tris-EDTA (pH 9.0) buffer for heat-mediated antigen retrieval using a microwave. Tissue sections were then blocked with 5% donkey serum in PBS for 1 h at room temperature, followed by incubation with primary antibody at 4 °C overnight. On the next day, samples were washed with PBS-Tween20 (0.05%) and re-probed with fluorophore-conjugated secondary antibodies. Antibody against Pro-SP-C was generated by immunizing guinea pigs with a peptide (MDMSSKEVLMESPPDYSAGPRSQFRIPCCPVHLKR) corresponding to the 1–35 aa of mouse Pro-SP-C protein. For targets that require signal amplification, biotinylated secondary antibodies were used, then HRP was introduced with streptavidin-HRP reagent (ApexBio, K1229), and sections were incubated with biotinylated Tyramide (Aladdin scientific, B288371) followed by streptavidin-fluorophores. All sections were counterstained with DAPI (Thermo Fisher, D3571) and sealed with Prolong-Gold antifade mounting medium (Thermo Fisher, P36934). Detailed information for all antibodies and conjugated fluorephores was included in Supplementary Table 1. Images were acquired under Leica Stellaris 5 or Olympus FV3000 confocal microscopes using identical laser settings across each group. Cell counting was performed manually using the “Cell Counter” plugin in FIJI. Cells with DAPI surrounded by αSMA or PDGFRα signals were identified as αSMA-positive or PDGFRα-positive, respectively. To quantify PDGFRαhi cells, the PDGFRα channel was auto-thresholded using the “Intermodes” method in FIJI, with signals exceeding the threshold classified as bright PDGFRα staining. PDGFRαhi cells were manually identified based on their perinuclear pattern, using the thresholding mask for guidance. P-ERK or p-AKT intensity for each PDGFRα+ αSMA+ cell was automatically measured using the CellProfiler software equipped with a plugin to run a deep learning-based segmentation algorithm, Cellpose261,62. Briefly, PDGFRα+ and αSMA+ cells were identified using the “RunCellpose” module. PDGFRα+ αSMA+ cells were then identified by the “RelateObjects” module that performs a colocalization test of each identified object. The mean fluorescence intensity of p-ERK or p-AKT within the PDGFRα-marked regions of the identified double-positive cells was then measured. The pipeline has been deposited at GitHub: https://github.com/hehua860/pipelines.
Wholemount immunofluorescence for AGER PDGFRα, αSMA, CDH4, and TAGLN was performed on lung strips collected from the left lobe25. Tissues were washed with PBS-Triton (0.3%) for 2 h, followed by blocking with 5% BSA in PBS-Triton (0.3%) for 6 h at room temperature. After blocking, tissues were incubated with primary antibodies diluted in the blocking solution for 48 h at 4 °C. Secondary antibody incubation was carried out for an additional 48 hours at 4 °C. Stained tissues were then cleared in 75% glycerol before imaging. Antibody dilutions were listed in Supplementary Table 1. For the AGER membrane staining, 34 μm-thick z-stacks were collected using confocal microscopes. 3D Images were reconstructed using the maximum intensity projection method within FIJI software, and quantification of airspace volumes and surface areas was done by using IMARIS software with default settings for surface rendering15. Elastins were stained with Cy5-Hydrazide (Aladdin, C171353, 1:1000 in PBS) and images were acquired using the same settings as in AGER staining. Co-staining of PDGFRα and αSMA with CDH4 or TAGLN was imaged at a depth of 44 μm, αSMA volume was quantified using FIJI. αSMA or TAGLN mean intensity was quantified using the maximum projection image from z-stacks.
Single-molecule RNA in situ hybridization using RNAscope
RNAscope Multiplex Fluorescent Reagent Kit v2 (ACD #323110) was performed according to the manufacturer’s instructions. Paraffin sections were baked for 1 h at 60 °C, and were then dewaxed in xylene and anhydrous ethanol. Slides were incubated with Hydrogen Peroxide followed by 1X target retrieval reagents in a microwave. Slides were then treated with Protease Plus reagents in the kit and hybridized with specific probes for Wnt11 (ACD, 405021) and Axin2 (ACD, 400331-C2) at 40 °C for 2 h, followed by incubation of amplifier for each probe including Multiplex FLv2 Amp1 and Multiplex FLv2 Amp2 according to the kit specifications. RNA fluorescent signals were then incorporated into the samples using two rounds of HRP-TSA reactions by series incubation with Multiplex FLv2 HPR-C1/C2 and TSA-vivid 570/650 dyes (TOCRIS, 7526 and 7527). Slides were counterstained with anti-αSMA and Pro-SP-C antibodies and DAPI. Axin2 positive cells were counted manually using the “Cell Counter” plugin in FIJI software.
Lung digestion
Mouse lungs were collected in ice-cold PBS, following removal of non-pulmonary tissues, lungs were minced into 1 mm3 blocks. Tissues were digested with 5 µ/mL Dispase (Corning, 354235), 0.025 mg/mL DNase I (Roche, 10104159001) and 480 µ/mL Collagenase type I (Thermo Fisher, 17100017) in PBS solution at 37 °C for 1 h. The tissues were subsequently transferred to C-Tubes (Miltenyi, 130-093-237) and mechanically dissociated using a gentleMACS Dissociator (Miltenyi). The samples were filtered through a 70 μm strainer and then centrifuged at 300 g for 5 min. Samples were treated with 1X RBC lysis buffer (Biolegend, 420301) for 5 min on ice. Cells were then collected in FACS buffer (PBS containing 1 mM EDTA, 25 mM HEPES, and 0.5% BSA) for downstream experiments.
FACS analysis and sorting
Single cells were spun down and resuspended in 100 μl FACS buffer, cells were incubated with anti-mouse CD16/32 (FcR) antibody (Biolegend, 101301) at 4 °C for 1 h to block non-specific antibody bindings. Cells were then stained with FACS antibodies listed in Supplementary Table 1. Before analysis or sorting, DAPI (5 mg/ml, Thermo Fisher, D3571) was added at the concentration of 1:10000. FACS analysis was performed using the BD FACS Fortessa II and Beckman CytoFLEX S systems. FACS sorting was performed using a BD FACSAria Fusion system. Data analysis was performed using FlowJo software.
MACS sorting
Single cells with FcR blocked as described above were incubated with biotin-conjugated anti-mouse CD140a antibody at 4 °C for 30 min. Cells were resuspended in 80 μl FACS buffer and incubated with 20 μl anti-biotin microbeads (Miltenyi, 130-090-485) at 4 °C for 30 min. Cells were then passed through LS columns (Miltenyi, 130-122-729) to enrich CD140a (PDGFRα) positive cells. For subsequent staining, cells were cytospun and fixed with 4% PFA at room temperature for 5 min.
scRNA-seq library preparation and analysis
For each sample, approximately 15000 to 20000 cells were loaded into the 10X Genomics Chromium system, The libraries were constructed using the Chromium Next GEM Single Cell 3’ Reagent Kits v3.1 according to the manufacturer’s protocol. Libraries were sequenced on the Illumina Nova-seq 6000 platform. FASTQ files were processed within the Cell Ranger using the count command with default settings. The output matrices were then analyzed in the Seurat R package63, cells with more than 6000 or less than 300 genes or a percentage of mitochondrial genes greater than 20% were excluded from analysis. Cells from Control and Ext1Gli1∆/∆ samples were integrated using the canonical correlation analysis (‘CCA’) method. Cells were clustered with the Louvain algorithm with a resolution of 0.4. Major cell types were annotated according to markers identified from LungMAP CellCards and CellRef64,65. Differentially expressed genes were determined by the FindMarkers function using the MAST test method66. scRNA-seq data were visualized by the scCustomize package (https://samuel-marsh.github.io/scCustomize/). For pathway activity inference, the decoupleR package67 was used with the default parameters. Functional enrichment analysis of myofibroblast enriched genes for REACTOME terms was conducted using ToppFun (https://toppgene.cchmc.org/enrichment.jsp). GSEA analysis on genes ranked by log2foldchange was performed using GSEA software and visualized by the GseaVis package. Adjusted p-value (FDR) < 0.25 was considered significant, significant terms were highlighted in Supplementary Data 1.
qPCR
RNA was isolated using RNA Easy Fast Tissue/Cell Kit (TIANGEN, DP451) and was then reverse transcribed using HiScript III All-in-one RT SuperMix (Vazyme, R333-01) to obtain the first cDNA strand. Quantitative PCR was conducted on a LightCycler® 96 system (Roche) using ChamQ Universal SYBR qPCR Master Mix (Vazyme, Q711-02) following the manufacturer’s specification. qPCR primers are as follows: Ext1-F TGGAGGCGTGCAGTTTAGG, Ext1-R GAAGCGGGGCCAGAAATGA; Actb-F GGCTGTATTCCCCTCCATCG, Actb-R CCAGTTGGTAACAATGCCATGT.
Alveolar organoid
A total of 5000 live FACS-purified AT2 cells (DAPI- tdTomato +) from P6 SftpcCreER; Rosa26tdTomato mice were mixed with 50000 MACS-enriched PDGFRα+ cells from P6 lungs in 45 μl organoid culture medium (MTEC/Plus medium with 25 ng/mL EGF (Peprotech, AF-100-15), 10 μM SB431542 (SelleckChem, S1067)), followed by mixing with 45 μl Corning Matrigel GFR Membrane Matrix (Corning, 356231). The resulting cell mixture was then carefully added to the center of a transwell insert and incubated at 37 °C for 20 min to solidify. Then medium was added into the bottom chamber of the transwell. Culturing of SFFF organoid was performed as described previously68, 5000 FACS purified AT2 cells were embedded in 50 μl Matrigel (Corning, 356231) and solidified on a 6-well plate. Cells were cultured in DMEM/F12 medium (Corning, 10-092-CV) containing 10 μM SB431542 (Selleckchem, S1067), 1 μM BIRB796 (Selleckchem,S1574), 1X ITS (Beyotime, C0343), 50 ng/mL hEGF (Peprotech, AF-100-15), 5 μg/mL Heparin (Sigma, H3149), 1X B27 supplements (Thermofisher, 17504044), 1X N2 supplements (Thermofisher, 17502048), 1.25 mM N-Acetyl Cysteine (Sigma-Aldrich, A9165) and 10 ng/mL mFGF10 (Peprotech, 450-61-25UG). SFFF organoids were cultured with or without 3 μM CHIR (Selleckchem, S1263), and rhWNT11 (R&D systems, 6179-WN) was used at 25 ng/ml. For both systems, cells were cultured with medium containing 10 μM ROCK inhibitor (Y-27632.2HCL, SelleckChem, S1049) at the first 48 h, the medium was replaced every other day with no ROCK inhibitor added. 10 ng mIL-1β (211-11B-10UG, Peprotech) was added to the SFFF medium within the first 2 days of culture. Organoid cultures were imaged with an Olympus APX-100 system and were subsequently fixed with 4% PFA overnight at 4 °C. Fixed organoids were then embedded in 3% agarose gel, and the gel blocks were dehydrated and embedded in paraffin. Samples were processed as described above for immunofluorescence staining. Mean organoid size was quantified using the “Analyze Particles” function in FIJI software.
Collagen gel contraction assay
Purified PDGFRα+ cells from P6 mice were loaded into 500 μl (1 mg/ml) rat collagen I gel (Corning, 356236) solution at 105 per well on a 24-well plate. The gel solutions were neutralized with 1 μl 1 M NaOH solution. Gels were let solidify at 37 °C for 20 min and transferred to a 12-well plate. The floating gels containing fibroblasts were cultured with DMEM supplemented with 10% FBS for 5 days, the medium was changed every other day. Images were taken every 24 hours using the Olympus APX-100 system. FIJI software was used to measure the gel size.
ATAC-seq library preparation and analysis
Live tdTomato+ mesenchymal cells from P8 Control (Ext1flox/+; Gli1CreERT2; Rosa26tdTomato) and KO (Ext1flox/flox; Gli1CreERT2; Rosa26tdTomato) mice were isolated using FACS sorting. 50000 cells were subjected to ATAC-seq library preparation using TruePrep DNA Library Prep Kit V2 for Illumina (Vazyme, TD502) and TruePrep® Index Kit V2 for Illumina (Vazyme, TD202). Libraries were sequenced on the Illumina Nova-seq 6000 platform. FASTQ files were trimmed with Trimgalore (https://github.com/FelixKrueger/TrimGalore) to remove adapters and aligned to the mm10 reference genome using bowtie269 with a parameter of --very-sensitive -X 2000. MACS2 was used to call peaks with parameters of --nomodel --shift -100 --extsize 20070. The csaw R package was used to determine differentially accessible regions with a parameter of |log2foldchange | > 0.58, p.adjust < 0.0571. Peak annotation and enrichment was carried out using ChIPseeker72. HOMER motif analysis was performed to determine the motifs enriched in the differential peaks73.
Statistical analysis
Graph plotting and data analysis were performed in GraphPad Prism software. Data are presented as mean and standard deviation. Unpaired t-test was used to determine significance for two-group comparison, and one-way ANOVA followed by multiple comparisons (Sidak or Tukey test) was used to determine significance where three or more groups were included. p-values less than 0.05 were considered significant, and specific n-values are listed in figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The single-cell RNA sequencing and ATAC-seq data have been deposited in the NCBI Gene Expression Omnibus with accession number GSE264372. Publicly available datasets GSE160876 and GSE165063 were also used. Source data are provided in this paper.
Code availability
CellProfiler pipelines for automatic quantification of pERK or pAKT fluorescent intensity can be found in the following GitHub repository: https://github.com/hehua860/pipelines.
References
Vila Ellis, L. & Chen, J. A cell‐centric view of lung alveologenesis. Dev. Dynam 250, 482–496 (2021).
Zepp, J. A. & Morrisey, E. E. Cellular crosstalk in the development and regeneration of the respiratory system. Nat. Rev. Mol. Cell Biol. 20, 551–566 (2019).
Thébaud, B. et al. Bronchopulmonary dysplasia. Nat. Rev. Dis. Prim. 5, 78 (2019).
Frank, D. B. et al. Emergence of a wave of Wnt signaling that regulates lung alveologenesis by controlling epithelial self-renewal and differentiation. Cell Rep. 17, 2312–2325 (2016).
Zhang, K. et al. A mammalian Wnt5a–Ror2–Vangl2 axis controls the cytoskeleton and confers cellular properties required for alveologenesis. Elife 9, https://doi.org/10.7554/elife.53688 (2020).
Li, R., Li, X., Hagood, J., Zhu, M.-S. & Sun, X. Myofibroblast contraction is essential for generating and regenerating the gas-exchange surface. J. Clin. Investig. 130, 2859–2871 (2020).
Zepp, J. A. et al. Genomic, epigenomic, and biophysical cues controlling the emergence of the lung alveolus. Science 371, https://doi.org/10.1126/science.abc3172 (2021).
Negretti, N. M. et al. A single-cell atlas of mouse lung development. Development 148, https://doi.org/10.1242/dev.199512 (2021).
Gillich, A. et al. Alveoli form directly by budding led by a single epithelial cell. Preprint at https://doi.org/10.1101/2021.12.25.474174 (2021).
Guo, M. et al. Single cell RNA analysis identifies cellular heterogeneity and adaptive responses of the lung at birth. Nat. Commun. 10, 37 (2019).
Branchfield, K. et al. A three-dimensional study of alveologenesis in mouse lung. Dev. Biol. 409, 429–441 (2016).
del Pilar, O. N., Garay, M. J. G. & Chen, J. Three-axis classification of mouse lung mesenchymal cells reveals two populations of myofibroblasts. Development 149, https://doi.org/10.1242/dev.200081 (2022).
Hagan, A. S., Zhang, B. & Ornitz, D. M. Identification of an FGF18-expressing alveolar myofibroblast that is developmentally cleared during alveologenesis. Development 147, https://doi.org/10.1242/dev.181032 (2019).
Gao, F. et al. Hedgehog-responsive PDGFRa(+) fibroblasts maintain a unique pool of alveolar epithelial progenitor cells during alveologenesis. Cell Rep. 39, 110608 (2022).
He, H., Snowball, J., Sun, F., Na, C. L. & Whitsett, J. A. IGF1R controls mechanosignaling in myofibroblasts required for pulmonary alveologenesis. JCI Insight 6, https://doi.org/10.1172/jci.insight.144863 (2021).
Li, R. et al. Pdgfra marks a cellular lineage with distinct contributions to myofibroblasts in lung maturation and injury response. Elife 7, https://doi.org/10.7554/elife.36865 (2018).
Li, C. et al. The secondary crest myofibroblast PDGFRα controls elastogenesis pathway via a secondary tier of signaling networks during alveologenesis. Development 146, https://doi.org/10.1242/dev.176354 (2019).
Gao, F. et al. Decoding the IGF1 signaling gene regulatory network behind alveologenesis from a mouse model of bronchopulmonary dysplasia. Elife 11, https://doi.org/10.7554/elife.77522 (2022).
Sarrazin, S., Lamanna, W. C. & Esko, J. D. Heparan sulfate proteoglycans. Cold Spring Harb. Perspect. Biol. 3, a004952–a004952 (2011).
Yan, D. & Lin, X. Shaping morphogen gradients by proteoglycans. Cold Spring Harb. Perspect. Biol. 1, a002493–a002493 (2009).
Clausen, T. M. et al. SARS-CoV-2 Infection depends on cellular heparan sulfate and ACE2. Cell 183, 1043–1057.e15 (2020).
Chen, L. et al. Structural basis for FGF hormone signalling. Nature 618, 862–870 (2023).
He, H., Huang, M., Sun, S., Wu, Y. & Lin, X. Epithelial heparan sulfate regulates Sonic Hedgehog signaling in lung development. PLoS Genet .13, e1006992 (2017).
Kreuger, J. & Kjellén, L. Heparan sulfate biosynthesis. J. Histochem. Cytochem. 60, 898–907 (2012).
Yang, J. et al. Development and plasticity of alveolar type 1 cells. Development https://doi.org/10.1242/dev.130005 (2015).
Barkauskas, C. E. et al. Type 2 alveolar cells are stem cells in adult lung. J. Clin. Investig. 123, 3025–3036 (2013).
Zepp, J. A. et al. Distinct mesenchymal lineages and niches promote epithelial self-renewal and myofibrogenesis in the lung. Cell 170, 1134–1148 (2017).
Kato, K. et al. Pulmonary pericytes regulate lung morphogenesis. Nat. Commun. 9, 2448 (2018).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Schepeler, T., Mansilla, F., Christensen, L. L., Orntoft, T. F. & Andersen, C. L. Clusterin expression can be modulated by changes in TCF1-mediated Wnt signaling. J. Mol. Signal 2, 6 (2007).
Hassan, D. & Chen, J. CEBPA restricts alveolar type 2 cell plasticity during development and injury-repair. Nat. Commun. 15, 4148 (2024).
Toth, A. et al. Alveolar epithelial progenitor cells require Nkx2-1 to maintain progenitor-specific epigenomic state during lung homeostasis and regeneration. Nat. Commun. 14, 8452 (2023).
Hao, H. X. et al. ZNRF3 promotes Wnt receptor turnover in an R-spondin-sensitive manner. Nature 485, 195–202 (2012).
He, H. et al. PRDM3/16 regulate chromatin accessibility required for NKX2-1 mediated alveolar epithelial differentiation and function. Nat. Commun. 15, 8112 (2024).
Nabhan, A. N., Brownfield, D. G., Harbury, P. B., Krasnow, M. A. & Desai, T. J. Single-cell Wnt signaling niches maintain stemness of alveolar type 2 cells. Science 359, 1118–1123 (2018).
Lako, M. et al. Isolation, characterisation and embryonic expression of WNT11, a gene which maps to 11q13.5 and has possible roles in the development of skeleton, kidney and lung. Gene 219, 101–110 (1998).
Tao, Q. et al. Maternal Wnt11 activates the canonical Wnt signaling pathway required for axis formation in Xenopus embryos. Cell 120, 857–871 (2005).
Ouko, L., Ziegler, T. R., Gu, L. H., Eisenberg, L. M. & Yang, V. W. Wnt11 Signaling promotes proliferation, transformation, and migration of IEC6 intestinal epithelial cells. J. Biol. Chem. 279, 26707–26715 (2004).
Pandur, P., Läsche, M., Eisenberg, L. M. & Kühl, M. Wnt-11 activation of a non-canonical Wnt signalling pathway is required for cardiogenesis. Nature 418, 636–641 (2002).
Cha, S.-W., Tadjuidje, E., Tao, Q., Wylie, C. & Heasman, J. Wnt5a and Wnt11 interact in a maternal Dkk1-regulated fashion to activate both canonical and non-canonical signaling in Xenopus axis formation. Development 135, 3719–3729 (2008).
Tsukui, T., Wolters, P. J. & Sheppard, D. Alveolar fibroblast lineage orchestrates lung inflammation and fibrosis. Nature 631, 627–634 (2024).
Brownfield, D. G. et al. Alveolar cell fate selection and lifelong maintenance of AT2 cells by FGF signaling. Nat. Commun. 13, 7137 (2022).
Zacharias, W. J. et al. Regeneration of the lung alveolus by an evolutionarily conserved epithelial progenitor. Nature 555, 251–255 (2018).
Chung, M.-I., Bujnis, M., Barkauskas, C. E., Kobayashi, Y. & Hogan, B. L. M. Niche-mediated BMP/SMAD signaling regulates lung alveolar stem cell proliferation and differentiation. Development 145, https://doi.org/10.1242/dev.163014 (2018).
Lü, J., Izvolsky, K. I., Qian, J. & Cardoso, W. V. Identification of FGF10 targets in the embryonic lung epithelium during bud morphogenesis. J. Biol. Chem. 280, 4834–4841 (2005).
Herriges, J. C. et al. FGF-Regulated ETV transcription factors control FGF-SHH feedback loop in lung branching. Dev. Cell 35, 322–332 (2015).
Hacohen, N., Kramer, S., Sutherland, D., Hiromi, Y. & Krasnow, M. A. Sprouty encodes a novel antagonist of FGF signaling that patterns apical branching of the drosophila airways. Cell 92, 253–263 (1998).
Bermudez, O., Pagès, G. & Gimond, C. The dual-specificity MAP kinase phosphatases: critical roles in development and cancer. Am. J. Physiol. Cell Physiol. 299, C189–C202 (2010).
Li, R., Herriges, J. C., Chen, L., Mecham, R. P. & Sun, X. FGF receptors control alveolar elastogenesis. Development 144, 4563–4572 (2017).
Wang, C. et al. Expansion of hedgehog disrupts mesenchymal identity and induces emphysema phenotype. J. Clin. Investig. 128, 4343–4358 (2018).
Piccolo, S., Sladitschek-Martens, H. L. & Cordenonsi, M. Mechanosignaling in vertebrate development. Dev. Biol. 488, 54–67 (2022).
Di, X. et al. Cellular mechanotransduction in health and diseases: from molecular mechanism to therapeutic targets. Signal Transduct. Target Ther. 8, 282 (2023).
Häcker, U., Nybakken, K. & Perrimon, N. Heparan sulphate proteoglycans: the sweet side of development. Nat. Rev. Mol. Cell Biol. 6, 530–541 (2005).
Khan, I. S. et al. Impaired myofibroblast proliferation is a central feature of pathologic post-natal alveolar simplification. Elife 13, RP94425 (2024).
Lim, K. et al. Organoid modeling of human fetal lung alveolar development reveals mechanisms of cell fate patterning and neonatal respiratory disease. Cell Stem Cell 30, 20–37 (2023).
Huang, L., Guo, Z., Wang, F. & Fu, L. KRAS mutation: from undruggable to druggable in cancer. Signal Transduct. Targeted Ther. 6, 386 (2021).
Yan, D. & Lin, X. Drosophila glypican Dally-like acts in FGF-receiving cells to modulate FGF signaling during tracheal morphogenesis. Dev. Biol. 312, 203–216 (2007).
Chapman, H. A. et al. Integrin α6β4 identifies an adult distal lung epithelial population with regenerative potential in mice. J. Clin. Investig. 121, 2855–2862 (2011).
Zhang, R. et al. Heparan sulfate biosynthesis enzyme, Ext1, contributes to outflow tract development of mouse heart via modulation of FGF signaling. PLoS ONE 10, e0136518 (2015).
Hsia, C. C. W., Hyde, D. M., Ochs, M. & Weibel, E. R. An official research policy statement of the american thoracic society/European respiratory society: Standards forquantitative assessment of lung structure. Am. J. Respir. Crit. Care Med. 181, 394–418 (2010).
Pachitariu, M. & Stringer, C. Cellpose 2.0: how to train your own model. Nat. Methods 19, 1634–1641 (2022).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinform. 22, 433 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Sun, X. et al. A census of the lung: CellCards from LungMAP. Dev. Cell 57, 112–145 (2022).
Guo, M. et al. Guided construction of single cell reference for human and mouse lung. Nat. Commun. 14, 4566 (2023).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Badia-i-Mompel, P. et al. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinform. Adv. 2, https://doi.org/10.1093/bioadv/vbac016 (2022).
Konishi, S., Tata, A. & Tata, P. R. Defined conditions for long-term expansion of murine and human alveolar epithelial stem cells in three-dimensional cultures. STAR Protoc. 3, https://doi.org/10.1016/j.xpro.2022.101447 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Lun, A. T. L. & Smyth, G. K. csaw: a Bioconductor package for differential binding analysis of ChIP-seq data using sliding windows. Nucleic Acids Res. 44, e45–e45 (2016).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
This research was supported by grants from NSFC (32192400 to X.L., 82200098 to H.H., 82271748 to F.S., U21A20333 to HM. L. and 32350710191 to X.L.), the National Key R&D Program of China (2022YFA0806200) to H.H., F.S. and X.L., the Fundamental Research Funds for the Central Universities (SCU2022D006) to H.H., State Key R&D Project of the Ministry of Science and Technology of Sichuan Province (2024YFFK0071) to HM. L. and the Science and Technology Commission of Shanghai Municipality (20DZ2261200) to X.L.
Author information
Authors and Affiliations
Contributions
H.H., F.S., HM.L., and X.L. conceived experiments and prepared the manuscript. H.H. and C.M. conducted experiments and performed data analyses. W.W., W.H., Y.L., S.S., Q.M., J.L., and HX. L. conducted experiments. H.H. and M.L. performed bioinformatic analyses.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Pao-Tien Chuang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, H., Ma, C., Wei, W. et al. Heparan sulfate regulates myofibroblast heterogeneity and function to mediate niche homeostasis during alveolar morphogenesis. Nat Commun 16, 1834 (2025). https://doi.org/10.1038/s41467-025-57163-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57163-4