Abstract
In plants, aberrant RNAs produced by endogenous genes or transgenes are normally degraded by the nuclear and cytosolic RNA quality control (RQC) pathways. Under certain biotic or abiotic stresses, RQC is impaired, and aberrant RNAs are converted into siRNAs that initiate post-transcriptional gene silencing (PTGS) in the cytosol. How aberrant RNAs are selected and brought to the cytoplasm is not known. Here we show that the RNA-binding protein SUPPRESSOR OF GENE SILENCING (SGS)3 shuttles between the cytosol and the nucleus where it associates with the ISWI-like CHROMATIN REMODELER (CHR)11 and with RNAs transcribed from PTGS-sensitive transgene loci binding CHR11. Knocking down CHR11 and its paralog CHR17 strongly reduces transgene PTGS, suggesting that SGS3 recruitment by CHR11/17 facilitates PTGS initiation. CHR11 is also enriched at endogenous protein-coding genes (PCGs) producing nat-siRNAs and va-siRNAs under biotic or abiotic stresses, and this production is reduced in chr11 chr17 double mutants at genome-wide level. Moreover, impairing CHR11 and CHR17 rescues the lethal phenotype caused by the massive production of siRNAs from PCGs in RQC-deficient mutants. We propose that SGS3 recruitment by CHR11/17 allows exporting RNAs to the cytosol to initiate the production of siRNAs.
Similar content being viewed by others
Introduction
In plants, post-transcriptional gene silencing (PTGS) mediated by short interfering (si)RNAs is a defense mechanism that selectively eliminates RNAs in the cytosol. It can target RNAs of both exogenous or endogenous origins (for a recent review see ref. 1). In the case of viruses, double-stranded (ds)RNA intermediates of viral replication or viral single-stranded (ss)RNAs that are converted to dsRNA by cellular RNA-dependent RNA polymerases (RDRs) are processed by DICER-LIKE2/4 (DCL2/4) into 21-22-nt siRNAs that are loaded onto ARGONAUTE1/2 (AGO1/2)2. AGO/siRNA complexes cleave viral ssRNA, thus reducing the virus titer. Moreover, when DCL2-dependent 22-nt siRNAs are loaded onto AGO1, the RNA-binding protein SGS3 is supposed to interact with AGO13 and protect AGO1-associated RNA from degradation4. This step occurs in cytosolic siRNA-bodies where RDR6 is present in addition to AGO1 and SGS35,6,7,8. This suggests that viral RNAs bound to AGO1 and protected by SGS3 are transformed into dsRNA by RDR6, thus creating an amplification loop that increases the production of siRNAs and further contributes to decreasing virus titer.
PTGS also sometimes targets sense transgenes (a process referred to as S-PTGS or co-suppression when S-PTGS affects simultaneously sense transgenes and homologous endogenous genes9,10). Because sense transgenes are not designed to produce dsRNA, the way S-PTGS is initiated is not fully understood. It has originally been proposed that certain arrangements could be responsible for the production of dsRNA because transgenes insert randomly and sometimes in multiple copies in plant genomes, and thus could form inverted repeats, leading to IR-PTGS (also referred to as RNAi). Although this could happen in certain cases, the fact that, in most cases, S-PTGS is initiated locally and subsequently spreads to the rest of the plant rather suggests a localized initiation event. The current accepted hypothesis consists in a sudden/accidental burst of production of aberrant (ab)RNAs that saturate the capacity of the RNA Quality Control (RQC) pathway, allowing the excess of abRNAs to be transformed into dsRNA by RDR6 to initiate PTGS1. Supporting this hypothesis, mutations in RQC enhance S-PTGS but not IR-PTGS11. Moreover, an uncapped antisense abRNA was detected in transgenic Arabidopsis lines expressing a bacterial GUS reporter under the control of the viral p35S promoter and the plant tRbcS terminator12. Two representative lines were characterized. Line L1 spontaneously undergoes S-PTGS, i.e., it accumulates GUS siRNAs and lack GUS mRNAs in a wildtype background, whereas it lacks GUS siRNAs and accumulates GUS mRNA in ago1, dcl2dcl4, rdr6 or sgs3 mutant backgrounds. In contrast, line 6b4 does not initiate S-PTGS spontaneously but is capable of amplifying S-PTGS in RQC-deficient mutants or upon grafting onto the L1 line, indicating that line 6b4 is prone to undergo S-PTGS but is not capable of initiating the process spontaneously. An uncapped antisense RNA was detected in both lines, which accumulates at a high level in line L1 and at a low level in line 6b412. Nevertheless, it accumulated at a much higher level in line 6b4 when the nuclear and cytosolic exoribonucleases XRN3 and XRN4 were mutated, allowing 6b4 xrn3 xrn4 plants to trigger S-PTGS spontaneously. These results indicate that both lines are prone to initiate and amplify S-PTGS, but that they differ in their ability to naturally produce enough abRNAs to initiate S-PTGS12.
Endogenous protein-coding genes (PCGs) can also produce siRNAs, but this does not occur under regular conditions of growth. RDR6-SGS3-dependent PCG-derived siRNAs, referred to as rqc-siRNAs or va-siRNAs are only detected in RQC-deficient mutants11,13,14,15,16,17 or in virus-infected plants in which RQC is dysfunctional or saturated by viral RNAs18. Another category of RDR6-SGS3-dependent endogenous siRNAs, referred to as nat-siRNAs are produced from PCG/NAT pairs (i.e., PCGs that overlap other protein-coding or non-protein coding genes), which are induced under certain biotic and abiotic stresses19. Importantly, nat-siRNAs not only derive from the overlapping part of the PCG/NAT pairs but also from the adjacent segments of each RNA, suggesting that after cleavage of the dsRNA part of the duplex, each ssRNA overhang is considered as an abRNA and transformed into dsRNA by RDR620.
Together, these results indicate that many types of endogenous or exogenous genes can produce siRNAs after the transformation of abRNA into dsRNA by RDR6. Still, major questions remain unanswered regarding the way abRNAs are selectively exported to the cytosol for transformation into dsRNA by RDR6. Using biochemical, genetic, and genome-wide approaches, we show that transgenes and endogenous genes that produce siRNAs bind to the ISWI-like CHROMATIN REMODELER (CHR)11. We show that CHR11 and its paralog CHR17 interact with the RNA-binding protein SGS3 and that SGS3 shuttles between cytosol and the nucleus where it binds transgene RNAs. We propose a model where CHR11/CHR17 and SGS3 are major nuclear determinants promoting the initiation of siRNA production.
Results
SGS3 interacts with CHR11, an ISWI-like chromatin remodeling protein
Whereas AGO1, DCL2, DCL4, and RDR6 have clear functions in RNA cleavage or synthesis, the role of the RNA-binding protein SGS3 has so far remained elusive, except for a possible role in bridging AGO1 to RDR6 during the PTGS amplification step in the cytosol3,4. To gather further information, we searched for SGS3 partners. A yeast-two-hybrid screen was performed using an Arabidopsis cDNA library and SGS3 as bait (see “Methods” and Supplementary Fig. 1). Among the candidates identified in this screen, the one that showed the strongest interaction with SGS3 corresponded to CHR11 (At3g06400), which encodes one of the 42 chromatin remodeling proteins of Arabidopsis21. Deletion analyses revealed that, at least in yeast, the coiled-coil domain of SGS3 interacts with the HAND-SANT-SLIDE domains located at the C-terminus of CHR11 (Fig. 1a and Supplementary Fig. 1).
a SGS3 and CHR11 structures. CHR11 and SGS3 domains interacting in yeast two-hybrid are shown in gray. The sequence of the CHR11 clone retrieved in the in yeast two-hybrid screen is shown in Supplementary Fig. 1. b Localization of CHR11 full-length and truncated forms (stop1 and stop2) in N. benthamiana (the same results were obtained twice in N. benthamiana and more than twice in A. thaliana transgenics). Channels are indicated above each column. The overlay is shown for DAPI/YFP/chlorophyll fluorescences. c BiFC Interaction between SGS3 and CHR11 (full-length and truncated forms) in the nucleus of N. benthamiana (pointed by arrows). Subcellular localization of reconstructed YFP was determined in the leaf epidermis for SGS3 protein in fusion with the C- terminal part of YFP (YC) and CHR11 proteins in fusion with the N- terminal part of YFP (YN). Channels are indicated above each column. The overlay is shown for DAPI/YFP/chlorophyll fluorescences (the same results were obtained 4 times in N. benthamiana and once in A. thaliana). d SGS3 is associated with fractions enriched in microsomal and nuclear compartments in wild-type Arabidopsis, in the chr11 chr17 double mutant, and in the chr11 chr17 double mutant complemented with the p35S:FLAG-CHR11construct (the same results were obtained twice on two different subcellular fractionations). The sgs3-1 null allele is used as a negative control.
In planta confirmation of this interaction was obtained by BiFC experiments performed by agro-infiltration in N. benthamiana. Indeed, co-infiltration of YFPNter-CHR11 and SGS3-YFPCter generated a nuclear YFP signal similar to the GFP signal obtained when infiltrating GFP-CHR11 (Fig. 1b and c). Deletion of the HAND-SANT-SLIDE domains did not abolish the nuclear localization of CHR11 (GFP-CHR11stop1), but abolished the interaction with SGS3 (YFPNter-CHR11stop1 + SGS3-YFPCter), confirming the yeast-two hybrid results. Deletion of only the SLIDE domain did not abolish the nuclear localization of CHR11 (GFP-CHR11stop2), nor the interaction with SGS3 (YFPNter-CHR11stop2 + SGS3-YFPCter), indicating that SGS3 interacts with CHR11 through its HAND-SANT domains (Fig. 1b and c).
SGS3 shuttles between the nucleus and the cytosol
SGS3 was reported as localizing mostly in cytosolic foci that also contain AGO1, AGO7, and RDR6, which are referred to as siRNA bodies5,6,7,8. Nevertheless, one report also described SGS3 in the nucleoplasm after immuno-localization on isolated nuclei22. Moreover, two recent reports revealed that truncated forms of the SGS3 protein lacking its prion-like domain, which likely makes it non-functional and localized in the nucleus23,24, suggesting that SGS3 could have some function in the nucleus. To get further information on the localization of the native SGS3 protein, western blots were performed on fractionated cells (Fig. 1d). In addition to being found in the microsomal fraction7, SGS3 was associated with the fractions enriched in nuclear compartments (Fig. 1d), suggesting a dual localization of SGS3 in the nucleus and cytosol of Arabidopsis cells.
To further examine the partition of SGS3 between the nucleus and the cytosol, a tagged SGS3 protein was expressed using the SGS3 promoter to stay as close as possible to physiological levels, and introduced into sgs3 mutants carrying the PTGS transgene locus L1. Compared to L1/sgs3/pUBQ10:SGS3-GFP transformants, which all showed complementation of L1 PTGS, only 4% of the L1/sgs3/pSGS3:SGS3-Venus transformants showed complementation of L1 PTGS (Supplementary Table 1). Moreover, the few L1/sgs3/pSGS3:SGS3-Venus transformants complemented for L1 PTGS gave no detectable signal in confocal or spinning disk analysis, indicating that the SGS3 promoter is insufficient for imaging by conventional technics. Therefore, SGS3-GFP was subsequently expressed under the RDR6 promoter, which is expressed at a higher level than SGS3 according to the ePlant software tool (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi)25. Introduction of pRDR6:SGS3-GFP in L1/sgs3 plants restored L1 PTGS more efficiently than the pSGS3:SGS3-Venus construct (Supplementary Table 1), and GFP could be detected in confocal analysis of L1/sgs3/pRDR6:SGS3-GFP plants. The GFP signal was only cytosolic, similar to that observed using a p35S:SGS3-GFP construct (Fig. 2a), suggesting that the amount of SGS3 in the nucleus is very low, even when expressed under the control of the very strong p35S promoter, likely because it is actively exported to the cytosol.
a SGS3-GFP relocalizes to the nucleus after inhibition of nuclear export by leptomycin B. Two independent sgs3/pRDR6:SGS3-GFP complemented lines were analyzed (the same results were obtained three times on three independent treatments with leptomycin B). b CHR11 over-expression increases retention of SGS3 in the nucleus in different combinations of lines expressing SGS3-GFP under the 35S or RDR6 promoters (both complementing the sgs3-1 mutant) and FLAG-CHR11 under the 35S or UBQ10 promoters both complementing a chr11 chr17 double mutant (the same results were obtained 6 times, see also Supplementary Fig. 3). c Nuclear localization of a control line expressing GFP-CHR11 under the 35S promoter (the same results were obtained more than twice in A. thaliana transgenic plants).
To test this hypothesis, L1/sgs3/pRDR6:SGS3-GFP plants were treated with Leptomycin B (LMB), an inhibitor of nucleo-cytosolic export. After 28 hours, the GFP signal was mostly seen in the nucleus (Fig. 2a and Supplementary Fig. 2a), supporting the hypothesis that SGS3 shuttles between the nuclear and cytosolic compartments. To ensure that LMB treatment does not generally affect the localization of proteins that reside in cytosolic foci, pUBQ10:GFP-ATG8A, and p35S:RDR6-GFP plants were treated with LMB. No nuclear GFP signal was observed (Supplementary Fig. 2b), indicating that LMB treatment does not affect the localization of ATG8A, which remains in cytosolic autophagic bodies26, or the localization of RDR6, which remains in cytosolic siRNA bodies. Given that the two PTGS components, SGS3 and RDR6, resides in the same cytosolic siRNA bodies, these results clearly indicate that only SGS3 shuttles between the nuclear and cytosolic compartments.
Because SGS3 interacts with CHR11 in the nucleus, we asked whether increasing the CHR11 level in the nucleus could increase the retention of SGS3 in the nucleus. To test this, L1/sgs3/pRDR6:SGS3-GFP plants were transformed with a pUBQ10:FLAG-CHR11 construct. Whereas the GFP signal was only cytosolic in L1/sgs3/pRDR6:SGS3-GFP plants, a nuclear SGS3-GFP signal was detected in some L1/sgs3/pRDR6:SGS3-GFP/pUBQ10:FLAG-CHR11 transgenic plants (Fig. 2b and Supplementary Fig. 3), indicating that increasing CHR11 level increases the fraction of nuclear SGS3. This result was confirmed by crossing L1/sgs3/p35S:SGS3-GFP plants with p35S:FLAG-CHR11 plants. Despite the use of the strong p35S promoter, the GFP signal was only cytosolic in plants expressing only L1/sgs3/p35S:SGS3-GFP. In contrast, a GFP signal was detected in the nucleus of some L1/sgs3/p35S:SGS3-GFP x p35S:FLAG-CHR11 plants (Fig. 2b and Supplementary Fig. 3). Nevertheless, the nuclear SGS3-GFP signal appeared non-homogeneous in plants over-expressing CHR11, indicating that CHR11 contributes to retaining SGS3 in the nucleus but that it can never abolish its export to the cytosol. Of note, SGS3 import in the nucleus appears independent of CHR11 and CHR17. Indeed, western blot analysis showed that SGS3 is detected in the nucleus of chr11 chr17 plants (Fig. 1d). Contrasting SGS3, CHR11 is never detected in the cytosol, indicating that the CHR11-SGS3 complex is purely nuclear (Fig. 2c), and dissociates before SGS3 re-export to the cytosol.
The two ISWI-like chromatin remodeling factors CHR11 and CHR17 act redundantly in transgene S-PTGS
CHROMATIN REMODELING PROTEIN 11 (CHR11) is a chromatin remodeling factor that belongs to the ISWI subfamily, which also comprises CHR17. CHR11 and CHR17 proteins share 98.6 % identity and ensure redundant functions. chr11 or chr17 single mutants do not exhibit any obvious developmental defects, while the chr11 chr17 double mutant is dwarf and sterile27 (Supplementary Fig. 4). Previous analyses revealed that CHR11 binds to the body of 17393 genes in Arabidopsis and that 3222 genes are up-regulated and 4188 down-regulated in the chr11 chr17 double mutant, which likely explains its drastic phenotype28,29. It was also reported that the plant-specific ISWI complex containing CHR11 and CHR17 functions to partially derepress genes and transposable elements subjected to transcriptional gene silencing (TGS) through their function in regulating nucleosome distribution30. However, no report to date has established a link between the ISWI complex and PTGS.
Although CHR17 was not retrieved from our Y2H screen, which may be due to the limited size of the cDNA library used for the screen, we considered that CHR11 and CHR17 could play redundant roles in transgene S-PTGS by attracting SGS3 to transgene RNAs. To test this hypothesis, we examined the effect of impairing CHR11 and CHR17 on L1 PTGS. However, the only chr11 and chr17 mutants available belong to the SALK and GABI collections, which interferes with the expression of the p35S:GUS transgene carried by the L1 locus31,32, making the use of these mutants impossible. Therefore, we transformed L1 with a pUBQ10:CHR11 construct and screened for transformants exhibiting a phenotype resembling that of the chr11 chr17 double mutant due to co-suppression of the CHR11 and CHR17 endogenous copies by the pUBQ10:CHR11 transgene. Given the possible role of CHR11 and CHR17 in S-PTGS/co-suppression, we did not expect obtaining plants exhibiting a phenotype as strong as that of the chr11 chr17 double mutant. Indeed, among 50 primary transformants, 15 showed a phenotype resembling that of the chr11 chr17 double mutant, among which two had mild leaf defects and produced seeds (Supplementary Fig. 5). Analysis of the progeny of these two transformants showed that CHR11 and CHR17 mRNA levels were reduced (Supplementary Fig. 6) and that L1 S-PTGS was delayed compared to L1 controls, suggesting a role for CHR11/CHR17 in transgene S-PTGS (Fig. 3a and b).
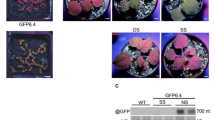
a GUS activity at 12 days in the progeny of line L1/ pUBQ10:CHR11 #46. Plants exhibiting a wild-type phenotype [wt] (#46.10.7 and #46.26.2) were harvested individually. Box plots show the five-number summary of each set of data: including the minimum score, first (lower) quartile, median, third (upper) quartile, and maximum score. Means are indicated by a cross, and the number of seedlings analyzed is indicated. Plants exhibiting a chr11 chr17 phenotype indicative of CHR11/CHR17 cosuppression (#46.10.7* and #46.26.2*) were pooled to form a unique sample. b GUS and CHR11 siRNA accumulation at 12 days in the progeny of line L1/ pUBQ10:CHR11 line #46. Plants were harvested in bulk. GUS activity is given below for each sample used for RNA extraction (the results are shown for the northern blot obtained from one unique extraction of each sample). c Kinetics of GUS activity in the progeny of lines L1/ pUBQ10:amirCHR11-17 #125 and #135, which exhibit strong CHR11/CHR17 cosuppression. At 7 and 12 days, plants were harvested in bulk (results at 17, 24, and 40 days are shown in Supplementary Fig. 8). d GUS mRNA, GUS siRNA and amiRCHR11-17 accumulation at 18 days in the progeny of lines L1/ pUBQ10:amirCHR11-17 lines #125 and #135 harvested in bulk. GUS activity is given below for each sample used for RNA extraction (the results are shown for the northern blot obtained from one unique extraction of each sample).
To test further this hypothesis, L1 was transformed with a pUBQ10:amiRCHR11-17 construct, which produces an artificial miRNA targeting both CHR11 and CHR17 mRNAs. Among 140 primary transformants, 30 showed mild to severe phenotypes resembling those of the chr11 chr17 double mutant. The two transformants exhibiting the most severe phenotypes showed impaired L1 S-PTGS, but these transformants were sterile (Supplementary Fig. 7). Analysis of the progeny of two fertile transformants exhibiting less severe phenotypes showed that CHR11 and CHR17 mRNA levels were reduced (Supplementary Fig. 6b) and that L1 S-PTGS was delayed compared to L1 controls (Fig. 3c, d and Supplementary Fig. 8), demonstrating a role of CHR11/17 in PTGS.
A simple explanation for S-PTGS impairment in chr11 chr17 double mutant could be that transgene transcription is reduced. To test this hypothesis, line 6b4 was transformed with the pUBQ10:amiRCHR11-17 construct. Line 6b4 carries the same p35S:GUS transgene as line L1; however, unlike line L1, which triggers PTGS spontaneously, line 6b4 does not trigger S-PTGS spontaneously. Rather, it stably expresses the p35S:GUS transgene, i.e., GUS activity levels are similar in 6b4 and 6b4 rdr6 or 6b4 sgs3 plants. This is likely because the 6b4 locus produces lower amounts of aberrant RNAs than the L1 locus. Nevertheless, the 6b4 locus triggers S-PTGS when RQC is impaired11,12,13,15,33,34,35,36, indicating that 6b4-derived aberrant RNAs can trigger S-PTGS when they are not degraded by the RQC machinery. Transformation of line 6b4 with the pUBQ10:amiRCHR11-17 generated transformants that showed GUS activity levels similar to 6b4 controls (Supplementary Fig. 9), even when CHR11 and CHR17 mRNA levels were reduced (Supplementary Fig. 6c), indicating that impairment of CHR11/17 does not modify the level of transcription of the p35S:GUS transgene. Together, these results suggest that the reason why CHR11/17 impairment affects S-PTGS is not due to changes in the level of transcription of transgene mRNA. Nevertheless, it remains possible that transgene transcription is qualitatively affected, modifying the ratio between functional mRNAs and aberrant RNAs.
CHR11 physically interacts with the p35S:GUS transgene
The way CHR11/17 promotes L1 S-PTGS could be by interacting with the p35S:GUS transgene locus to facilitate the addressing of GUS RNAs to SGS3 through the CHR11/SGS3 interaction. To test this hypothesis, L1/pUBQ10:GFP-CHR11 and L1/pUBQ10:Flag-CHR11 lines that complement the developmental defects of a chr11 chr17 double mutant were selected (Supplementary Fig. 10). ChIP-qPCR experiments performed on these plants revealed an enrichment of CHR11 on the p35S:GUS transgene at the L1 locus (Fig. 4).
ChIP-qPCR analyses were performed on 15-day-old seedlings of the indicated genotypes. a ChIP was performed on the L1/pUBQ10:GFP-CHR11 line using GFP antibodies for IP. The L1 line was used as a control, followed by normalization to GAPDH. Graphical representation shows the fold change of two biological repeats. b ChIP was performed on the L1/pUBQ10:FLAG-CHR11 using Flag antibodies for IP. Here, the L1/pUBQ10:GFP-CHR11, which expressed CHR11 at a similar level than in L1/pUBQ10:FLAG-CHR11, was used as a control, followed by normalization to GAPDH. Graphical representation shows the fold change as the mean of three biological repeats and displays every data point. Error bars represent the standard deviation.
Because the 6b4 locus was used as a control to rule out the hypothesis that S-PTGS impairment in chr11 chr17 double mutant is due to reduced transgene transcription (Supplementary Fig. 9), ChIP-qPCR experiments performed on 6b4/pUBQ10:GFP-CHR11 plants revealed that CHR11 also interacts with the p35S:GUS transgene at the 6b4 locus (Supplementary Fig. 11). In addition to validating the 6b4 locus as a control, this result supports the hypothesis that the 6b4 locus triggers S-PTGS in an RQC-deficient background through CHR11/SGS3 selection/recruitment, similar to the L1 locus. Altogether, these results suggest that CHR11/17 generally promotes transgene S-PTGS by binding to transgene loci, thus allowing the association of transgene RNAs with SGS3 in the nucleus and their export to the cytosol to initiate S-PTGS.
SGS3 associates with GUS RNAs in the nucleus
If SGS3 chaperones RNAs transcribed at loci binding CHR11 to promote their export to the cytosol, a direct interaction between SGS3 and GUS RNA should be detected in the nucleus. At first, RNA immunoprecipitation (RIP) was performed using the nuclei of pRDR6:SGS3-GFP plants. However, GUS RNAs were not detected after qRT-PCR, which could be due to the amount of SGS3 in the nucleus being insufficient to be immunoprecipitated. To circumvent this problem, SGS3-GFP was fused to an NLS signal and introduced into L1/sgs3 plants, hoping to increase the amount of nuclear SGS3. pRDR6:SGS3-NLS-GFP complemented L1 PTGS as efficiently as the pRDR6:SGS3-GFP construct (Supplementary Table 1). However, the GFP signal remained only cytosolic, confirming that the native SGS3 protein is actively re-exported to the cytosol (Supplementary Fig. 12). In a last attempt to increase the amount of nuclear SGS3, plants were submitted to heat stress. In pRDR6:SGS3-GFP plants, the SGS3-GFP signal accumulated exclusively in cytosolic foci which co-localized with stress granules5 (Supplementary Fig. 12). In contrast, pRDR6:SGS3-NLS-GFP plants showed a nuclear GFP signal, similar to the nuclear relocalization observed after LMB treatment (Supplementary Fig. 12), indicating that the added NLS is functional and that the export of the SGS3-NLS-GFP protein from the nucleus to the cytosol is inhibited or strongly diminished at 38 °C.
Nuclei of heat-stressed pRDR6:SGS3-NLS-GFP plants were isolated and used for RIP-qRT-PCR experiments. A significant enrichment of GUS RNA was observed in the fraction immunoprecipitated with SGS3 in the nucleus (Fig. 5), indicating that SGS3 is able of taking up its target RNAs in the nucleus and supporting the hypothesis that SGS3 chaperones these RNAs to cytosolic siRNA-bodies where they are transformed into dsRNA by RDR6. This is also in agreement with the fact that sgs3 is epistatic to rdr64 and that SGS3 is needed to stabilize mRNA before the synthesis of the complementary RNA strand by RDR6.
RIP analysis of GUS mRNA associated with SGS3 was performed using L1/sgs3 plants complemented by pRDR6:SGS3-NLS-GFP, and L1 line (without TAG) as control for normalization. RT-qPCR was performed to determine the accumulation levels of 5’ terminal part, central part of GUS transcript in the RIP. qPCR Fold enrichment of the immunoprecipitated (IP) was calculated as 2exp(−ΔΔCt [RIP in SGS3-NLS-GFP line/background in L1]). Fold enrichment of the qPCR internal control, PP2A was also calculated and arbitrarily set to 1. The mean values and standard deviation of three independent experiments are shown as every data point.
CHR11 promotes the production of siRNAs from endogenous protein-coding genes
The results presented above suggest a model in which SGS3 is recruited by CHR11 and presumably its paralog CHR17 to transgene DNA, allowing certain nascent transgene RNAs to bind SGS3, thus limiting their degradation by RQC. Moreover, SGS3 shuttling between nucleus and cytosol likely promotes the export of SGS3-bound transgene RNAs to the cytosol where they can be fully converted to dsRNA by RDR6 in siRNA bodies.
The next critical question was to assess how general is this mechanism for siRNA production from endogenous genes. To address this question, whole-genome data of CHR11 binding in wild-type plants29 were mined for various categories of genes producing different types of siRNAs. In addition, small RNA sequencing was performed to determine the accumulation of endogenous siRNAs in wildtype and chr11 chr17 plants. At first, TAS genes, which produce SGS3-dependent ta-siRNAs, were analyzed for CHR11 binding. Whereas 46% of the whole set of Arabidopsis genes bind CHR1129, none of the seven TAS genes showed binding (Fig. 6a and Supplementary Table 2a). Moreover, the accumulation of ta-siRNAs was not significantly affected in chr11 chr17 (Fig. 6c) as compared to wt. This result is coherent with the fact that TAS RNAs become substrates for SGS3 and RDR6 only after they are cleaved by miRNA-guided AGOs in the cytosol. Therefore, the binding of CHR11 to TAS genes and the association of SGS3 to TAS RNAs in the nucleus do not appear necessary to promote the export of TAS RNAs.
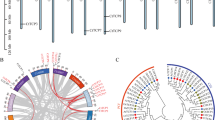
a Number and proportion of whole Arabidopsis genes (whole-genome or sub-categories producing siRNAs) binding to CHR11 or not according to ref. 22. ta-siRNA: trans-acting siRNAs, va-siRNA: virus-activated siRNAs, nat-siRNA: natural antisense transcripts siRNAs, rqc-siRNA: endogenous siRNAs produced when RQC is impaired, ct-siRNA: endogenous siRNAs produced by coding transcripts. b Number and proportion of Arabidopsis PCGs (whole-genome or sub-categories producing siRNAs) binding CHR11 or not according to22. c Variation of 21/22nt ta-siRNA accumulation between chr11 chr17 and Col-0. Values are log2 fold changes according to DESeq2 differential analysis (n = 3 biologically independent samples per genotype and 7 ta-siRNA genes). The median is represented by the horizontal line, the box indicates the interquartile range (IQR, 25th-75th percentiles), whiskers correspond to 1.5 IQR, and black points indicate points outside this range. d Accumulation of va-siRNAs upon TCV infection of Col-0 and chr11 chr17. Values are log2 fold changes according to DESeq2 differential analysis compared to Mock treatment (n = 3 biologically independent samples per genotype and condition, and 324 TCV va-siRNA genes). The number shown above indicates the p-values of the difference of log2 fold change determined by a Wilcoxon test (the value displayed “< 2.2e-16” is the smallest p-value R will return for a Wilcoxon test). The median is represented by the horizontal line, the box indicates the interquartile range (IQR, 25th-75th percentiles), whiskers correspond to 1.5 IQR, and black points indicate points outside this range. e Variation of va-siRNAs upon TCV infection of Col-0 and chr11 chr17. Values of the heat map are log2 fold changes according to DESeq2 differential analysis compared to Mock treatment.
In a second time, protein-coding genes (PCGs) that produce siRNAs without the requirement of AGO/miRNA-guided RNA cleavage were examined because these PCGs could require CHR11 binding and SGS3 association in the nucleus to allow RNA export to the cytosol. Generally, PCGs do not produce siRNAs, except when RQC is impaired11,13,14,15,16,17. Compared to the 50% of Arabidopsis PCGs that bind CHR1129, a significant enrichment was found for PCGs producing siRNAs in dcp2 (74%), vcs (73%), ski2 xrn4 (72%), urt1 xrn4 (69%) and cstf64 (79%) (Fig. 6b and Supplementary Table 2B). In addition to RQC deficiency, specific conditions, including biotic and abiotic stresses, allow PCGs to produce siRNAs. Remarkably, a significant enrichment (76%) was also observed for PCG/NAT pairs (i.e., PCGs that overlap other protein-coding or non-protein coding genes), which, under certain biotic and abiotic stresses, produce endogenous siRNAs referred to as nat-siRNAs19. Lastly, an enrichment in CHR11 binding was observed for PCGs producing endogenous siRNAs, referred to as va-siRNAs, which are produced after infection with viruses such as CMV (85%) or TuMV (71%) (Fig. 6b and Supplementary Table 2B). Together, these results support the relevance of CHR11 in the production of siRNAs from PCGs.
To further test whether CHR11 could promote siRNA production from PCGs, the effect of CHR11/17 impairment on endogenous siRNAs synthesis was investigated. It is likely that chr11 chr17 mutants would not tolerate all biotic or abiotic stresses required to induce the production of nat-siRNAs. In contrast, it appeared possible to infect chr11 chr17 double mutants with viruses. Infection with CMV was not efficient enough in our hands, but infection with TCV was very successful on the chr11 chr17 double mutant. Small RNAseq analysis in TCV-infected wildtype plants revealed that TCV infection results in the production of va-siRNAs from PCGs that also produce va-siRNAs in CMV-and TuMV-infected plants (Supplementary Fig. 13). Moreover, enrichment of CHR11 binding was also found for PCGs producing va-siRNAs under TCV infection (83%) (Fig. 6b and Supplementary Table 2b). Comparing TCV-infected wildtype plants and TCV-infected chr11 chr17 mutants revealed that the induction of va-siRNAs was significantly reduced in chr11 chr17 plants (Fig. 6d and e), further supporting the link between CHR11/17 and the production of endogenous vasiRNAs. To determine if this reduced induction of va-siRNAs was not simply due to reduced transcription of these PCGs in chr11 chr17 double mutants, mRNAseq data of wildtype plants and chr11 chr17 double mutants29 were compared to our data. No relation or bias could be found between the variation of va-siRNAs accumulation between chr11 chr17 and Col plants with the differential accumulation of their mRNA transcripts between the two genotypes for the full set of genes (Supplementary Fig. 14a) or for the TCV va-siRNA-producing genes (Supplementary Fig. 14b). These results rule out the possibility that the reduced induction of va-siRNAs in chr11 chr17 double mutants is due to reduced transcription of these PCGs, similar to what was observed for transgenes (Supplementary Fig. 9).
At last, we investigated whether impairing CHR11 and CHR17 activity could rescue the growth of RQC-deficient mutants. Indeed, RQC-deficient mutants such as the vcs and dcp2 single mutants15 or the ski2 xrn4 double mutant17 die at an early stage of development, likely because of the massive production of siRNAs from PCGs. This lethal phenotype can be rescued by impairing RDR6 or SGS3, which prevents the production of most of these siRNAs15,17. The chr11 and chr17 mutations could not be combined with dcp2 or vcs because DCP2 and CHR17 are genetically linked, and so are VCS and CHR11. Therefore, chr11 and chr17 were combined with ski3 and xrn4. The ski3 xrn4 double mutant die at an early stage of development, whereas the ski3 xrn4 rdr6 triple mutant is viable (supplementary Fig. 15). The chr17 ski3 xrn4 triple mutant die at the two-cotyledons stage, whereas the chr11 chr17 ski3 xrn4 quadruple mutant grew as well as chr11 chr17 (Fig. 7), indicating that impairing CHR11 and CHR17 rescues the lethal phenotype of RQC-deficient mutants. This result, therefore, suggests that CHR11 and CHR17 participate to the production of rogue siRNAs from endogenous gene when RQC is deficient.
Pictures of representative plants of the indicated genotypes grown for 27 days in short days. All plants were grown at the same time. All pictures at the same scale. a ski3 xrn4 chr17. b ski3 xrn4 chr11 chr17. c chr11 chr17.
Discussion
Although sense transgene PTGS (S-PTGS) was first described 30 years ago9,10, the way it is initiated is still not fully understood. It is assumed that transgene aberrant RNAs are transformed into dsRNA by the RNA-dependent RNA polymerase RDR6, but how these aberrant RNAs are selected and brought to RDR6 and how they escape degradation by RNA quality control (RQC) pathways was not resolved. Here, we propose a model in which the binding of CHR11 (and presumably CHR17) to transgene DNA allows the recruitment of SGS3 to nascent transgene RNAs, protecting them from a full degradation by RQC and allowing their export to the cytosol to initiate S-PTGS (Fig. 8). CHR11 and CHR17 domains are shared by the ISWI and SWR1 complexes involved in nucleosome sliding and H2AZ deposition at a large number of genomic loci29,30,37. CHR11 and CHR17 interact with actors of the ISWI and SWR1 complexes by their SLIDE domain, which is dispensable for interaction with SGS3. Therefore, by interacting with CHR11 HAND and/or SANT domains, SGS3 appears, to form a complex that has not been described before. Of note, Luo et al.29 did not recover SGS3 by IP-MS using a pCHR11:CHR11-FLAG transgenic line. This could be explained by the fact that the interaction between SGS3 and CHR11 was only detected by Bi-FC when a TAG epitope was fused to the N terminal part of CHR11 (this work).
In the nucleus, SGS3 is recruited at CHR11-binding loci. When the genomic arrangement allows the production of sense and antisense RNAs forming dsRNA with a 5’ overhang, SGS3 binds to the dsRNA structure, which probably releases the CHR11-SGS3 interaction, allowing its export to the cytosol. In the cytosol, RNAs bound to SGS3 are converted to dsRNA by RDR6, followed by processing by DCL4 and DCL2 into 21-nt siRNAs that can execute PTGS and 22-nt siRNAs that allow PTGS amplification. The model described for the p35S-GUS transgene locus likely applies for endogenous PCG/NAT pairs. (This figure was entirely created by authors using ppt).
SGS3 recruitment by CHR11/17 is reminiscent of the recruitment of the SGS3-like protein IDN2 by SWI3B, a sub-unit of the SWI/SNF complex that promotes RNA-directed DNA methylation (RdDM) and transcriptional gene silencing (TGS) mediated by 24-nt siRNAs derived from transposons and repeats transcribed by the plant-specific polymerases Pol IV and Pol V38. Similar to SGS3, which interacts with CHR11 through its coiled-coil domain, IDN2 interacts with SWI3B through a similar domain. Whether IDN2 protects nascent Pol V-dependent RNAs against degradation by nuclear RQC is not known, but it is a reasonable hypothesis explaining the role of IDN2 in RdDM and TGS. If SGS3 plays a similar role in PTGS, it should protect aberrant RNAs in both nuclear and cytosolic compartments and presumably travel from nucleus to cytosol with its RNA partners. SGS3 was reproducibly found in the cytosol6,7,8 but was described only once in the nucleoplasm of isolated nuclei22. However, truncated forms of the SGS3 protein lacking its prion-like domain, which likely makes it non-functional, were found in the nucleus23,24; suggesting that SGS3 could have some function in the nucleus. Here, we show that, in fact, the native SGS3 shuttles between cytosol and nucleus. When nucleo-cytosolic export is not inhibited, SGS3 is hardly detectable in the nucleus, except when CHR11 level is increased in the nucleus, or when SGS3 fused to a canonical NLS is forced to remain in the nucleus after heat stress, suggesting that SGS3 is actively exported from the nucleus to the cytosol. This efficient export, together with the RNA binding and protecting effect of SGS3 against degradation by RQC, certainly contributes to addressing transgene RNAs to the cytosol to initiate PTGS.
Whole-genome analysis revealed that CHR11 bind to 50% of the protein-coding genes (PCGs) of the Arabidopsis genome (Supplementary Table 2B), suggesting that half of the endogenous PCGs could have the potential to produce siRNAs through the SGS3 pathway. So far, only 5000 endogenous PCGs were shown to produce siRNAs, either when RQC is impaired by mutations15,16,17,39, or when biotic or abiotic stress somehow challenge the functioning of RQC18,20. However, not all cell types could be analyzed because of the dramatic developmental defects of RQC-deficient mutants and infected plants, so the number of endogenous PCGs producing siRNAs is certainly underestimated. Nevertheless, given that the vast majority (70-85%) of the known siRNA-producing PCGs binds CHR11 (Fig. 6b and Supplementary Table 2B), it is likely that CHR11 favor the attraction of SGS3 to the RNAs transcribed by these PCGs. Although the genetic requirement for the production of these PCG-derived siRNAs has not always been determined, most of them were shown to depend on RDR1 or RDR6, which generally act in concert with SGS3 to produce 21- and 22-nt siRNAs40.
Given that CHR11 binds to p35S:GUS loci (Fig. 4), we propose a revised model in which S-PTGS is initiated in the nucleus owing to the interaction between CHR11 and SGS3 (Fig. 8). Consistent with the strong affinity of SGS3 to dsRNA exhibiting a 5’ overhang41, we propose that SGS3 binds to dsRNA formed by the annealing of the full-length GUS mRNA to the uncapped antisense RNA that is complementary to the 3’ end of the GUS mRNA. We previously suggested that this uncapped RNA, referred to as SUG, results from the processing of a read-through transcript that originates in the adjacent and convergent pNOS-NPT-tNOS transgene because the NOS terminator does not efficiently terminate transcription12. Following cleavage at the polyadenylation site of the NPT transcript, the downstream part of the read-through transcript, i.e., the uncapped SUG RNA, could anneal with the GUS mRNA. SGS3 could then bind to the GUS/SUG duplex in the nuclei because it has a 5’ overhang.
To explain how SGS3 and CHR11 separate to allow exporting the SGS3-dsRNA complex to the cytosol, we propose a model similar to that described for the shuttling of AGO142. In the AGO1 model, the native AGO1 protein produced in the cytosol has an exposed NLS and a buried NES, allowing its import in the nucleus. When it binds miRNAs in the nucleus, AGO1 changes conformation, which exposes its NES, allowing the AGO1/miRNA complexes to be exported to the cytosol. We propose, that when it binds to dsRNA, SGS3 changes conformation, which disrupts the CHR11-SGS3 association. The binding of SGS3 to dsRNA and/or the dissociation of SGS3 from CHR11 may also expose its NES, allowing the SGS3-dsRNA complex to be exported to the cytosol. Following its export to the cytosol, the SGS3-bound GUS/SUG duplex could be processed into 21- and 22-nt siRNAs by DCL4 and DCL2 to initiate PTGS, which is subsequently amplified by RDR6. GUS and/or SUG RNAs could also be separately transformed into dsRNA by RDR6 prior to DCL2/DCL4-mediated processing into 21- and 22-nt siRNAs.
This model could also explain the formation of nat-siRNAs that originate from overlapping transcription units producing natural antisense pairs of RNA (NAT transcripts). Indeed, nat-siRNAs derive from the overlapping region of an mRNA and its NAT, but also from the rest of the transcripts19. This suggests that the two annealed transcripts are protected from degradation by SGS3 and exported to the cytosol, where each are transformed into dsRNA by RDR6 to produce siRNAs. Therefore, the high affinity of NAT pairs for CHR11/17 likely explains how CHR11-mediated SGS3 recruitment to NAT transcripts may lead to producing nat-siRNAs, similar to what happens at transgene loci producing unintended antisense RNAs. Given the large overlap between the set of PCGs producing nat-siRNAs and the sets of PCGs producing siRNAs when RQC is impaired or during virus infection (Supplementary Fig. 16), nat-siRNAs, rqc-siRNAs and va-siRNAs could represent a unique class of siRNAs whose common determinant is their arrangement as PCG/NAT pairs. The complete list of bona fide PCG/NAT pairs is not known because the NAT partner could produce a non-conventional RNA that cannot be easily identified. Indeed, it could produce a non-coding RNA that cannot be predicted based on the genomic sequence and/or that is expressed only under particular conditions. Alternatively, it could produce uncapped or non-polyadenylated RNAs that cannot be cloned in regular RNAseq experiments. Although it is not possible to prove that all rqc-siRNAs and va-siRNAs originate from bona fide PCG/NAT pairs, the fact that impairing CHR11 and CHR17 rescues the lethal phenotype caused by the massive production of siRNAs from PCGs in the RQC-deficient ski3 xrn4 double mutant (Fig. 7) strongly suggests that our model for transgene S-PTGS applies to a majority of PCG-derived siRNAs.
Methods
Plant material, growth conditions, transformation, and virus inoculation
p35S:GUS lines 6b4 and L1 and mutants sgs3-1, ski3-3, xrn4-5, chr11-1 and -2, chr17-1 and -2 have been described previously6,28,36,43,44,45,46,47, chr17-3 (SALK 629656) and chr17-4 (SALK 585156) derive from the SALK T-DNA collection. The three double mutants chr11-1 chr17-1, chr11-2 chr17-1, and chr11-2 chr17-4 were generated and used indifferently for analyses since they share the same developmental defects. Arabidopsis seeds were sown in vitro on a nutritive medium (1.3% S-medium Duchefa, 1% Phytoblend agar) and vernalized at 4 °C for at least 2 days before being transferred to soil in culture chambers. Plants were grown at 23 °C, 70% humidity, 120 μE.m−2 lighting, and 16 h light/8 h dark (long-days) or 8 h light/16 h dark (short-days) photoperiod. To generate transgene/mutant combination lines, 6b4 and L1 were transformed by agrobacterium or crossed to the corresponding mutants/transgenic lines, and F2 progenies were genotyped to identified plants homozygous for both the transgene and the mutation(s). Arabidopsis plants were transformed by the floral dip method6. For virus infection, 20-day-old plants grown under short-day conditions were infected with TCV by mechanical inoculation43. Plants were grown for 3 weeks, after which the total aerial parts of 4 to 12 plants were harvested.
Cloning and constructs
pSGS3:SGS3-GFP was first obtained using a recombineering-based gene tagging system as described in ref. 25. To ensure that all cis-regulatory sequences are included, we have used a bacterial homologous recombination system to insert Venus fluorescent gene into SGS3 genomic sequence harbored by transformation-competent bacterial artificial chromosomes (TACs) before transfer to Agrobacterium and plant transformation. cDNAs or coding sequence plus introns from ATG to stop of SGS3 and CHR11 were cloned in the GATEWAY™ compatible vector pDONR207 (Invitrogen) using the following primers: attB2SGS3f/ attB2SGS3R and attB1CHR11Fbis/ attB1CHR11Rbis (Supplementary Table 3). SGS3 was transferred to the binary vectors pB7FWG2 to make the p35S:SGS3-GFP and in pRDR6-pGWB4 to make the pRDR6:SGS3-GFP. pRDR6-pGWB4 was obtained by cloning RDR6 promoter in pGWB448 digested by HindIII after amplification by the primers pairs proSGS2F-hIII / proSGS2R-HIII. pRDR6:SGS3-NLS-GFP was obtained by cloning canonical NLS sequences in speI of SGS3-pDONR207 prior to recombination in the compatible destination vector pRDR6-PGWB4.
CHR11 was transferred to the binary vectors pB7WGF2 to make the p35S:GFP-CHR11, to pGWB12 to make p35S:Flag-CHR11, pUB-Dest49 to make pUBQ10:CHR11, and pUBC‐GFP-DEST to make pUBQ10-GFP-CHR11. pUBQ10:Flag-CHR11 was obtained by cloning in pDONR207 CHR11 cDNA amplified by attB1FlagCHR11F and attB2CHR11Rbis primers prior to recombination in the compatible destination vector pUB-DEST. QuikChange™ (Stratagene, Arcueil, France) PCR‐based mutagenesis was used to introduce premature stop1 and stop2 in CHR11 cloned in pDONR207 using CHR11stop1Fbis /CHR11stop1Rbis and CHR11stop2F/CHR11stop2R pairs of primers to obtained p35S:GFP-CHR11stop1 and stop2.
We used the MIR319a precursor to engineer artificial micro RNA (amiRNA) for targeting both CHR11 and 1750. We replace the original miR319a with the artificial sequence amiRCHR11-17, 5’-TATCATGAAACGGTCGCACTA-3’, using WMD3-Web microRNA designer (http://wmd3.weigelworld.org/) to choose the best sequence for targeting both CHR11 and CHR17. AmiRCHR11-17 was synthesized and cloned in pUC57 by GenScript Corporation (New Jersey 08854-3900). Primers amiRf and amiRr were used to amplified amiRCHR11-17 in its precursor backbone for cloning in the pDONR207 vector prior to recombination in the compatible destination vector pUB-Dest to obtained pUBQ10:amirCHR11-17.
CDS of RDR6 without its stop codon was cloned in SalI and NotI of pENTR1A vector after amplification by the primers pairs RDR6f/RDR6r (Supplementary Table 3) prior recombination in the compatible destination vector pH7FWG2 to obtained p35S:RDR6-GFP.
The genomic sequence of H2B (At3g45980) under the control of 1 kb of it’s own promoter have been cloned in the Gateway pGWB553 vector48 with mRFP C-terminus tag, using At3g45980-F1 and At3g45980-R1 primers (Supplementary Table 3). The genomic sequence of NUP54 (At1g24310) under the control of 1 kb of it’s own promoter have been cloned in the Gateway pGWB543 vector48, with CFP C-terminus tag, using At1g24310-F1 and At1g24310-R1 primers (Supplementary Table 3).
GUS activity and molecular analyses
GUS protein was extracted, and GUS activity was quantified by monitoring the quantity of 4-methylumbelliferone products generated from the substrate 4-methylumbelliferyl-b-D-glucuronide (Duchefa) on a fluorometer (Thermo Scientific fluoroskan ascent)33.
All RNA gel blot analyses were performed using 10 μg of total RNA. UidA, 25S DNA probes, and U6 oligo probes have been described before33. Oligo probe amiRCHR11 was 32 P‐end‐labeled to detect amiRCHR11-17, and CHR11 PCR fragment amplified by primers attB1CHR11Fbis/ attB2CHR11Rbis was used as a probe to detect CHR11 siRNA accumulated in co-suppressed L1/pUBQ10:CHR11 lines (Supplementary Table 3).
For the reverse transcription, RNA used for northern blot analysis were treated with DNaseI (Invitrogen) and 1 μg of DNA‐free RNA was reverse transcribed with an oligo d(T)18NN using the RevertAid H Minus Reverse Transcriptase (ThermoFisher, http://www.thermofisher.com/). Specific amplification of CHR11 and CHR17 were performed by using the CHR11f1 and r1, CHR17f and r primers (Supplementary Table 3), and qPCR results were normalized to GAPDH51.
Yeast two‐hybrid and BiFC cloning assays
SGS3 cDNA and the C‐terminal three coiled‐coil domains of SGS3 (corresponding to the amino acids 463–626) were cloned into the pLex10 vector to produce fusion proteins with the LexA DNA‐binding domain6. SGS3 cDNA was used as bait in yeast two‐hybrid screen of the 3-days-old etiolated Arabidopsis cDNA library CD4-2252. L40 strain containing LexA/SGS3 was transformed with the cDNA library fused to the activation domain GAL4. A total of 4.3 106 transformants were plated on a selective medium without Tryptophan, Leucine, and Histidine, after 2H, 12H or without expression on a permissive medium without Tryptophan (which allowed the weak to strong interactants to be screened). 264 His + colonies (2 from the screen without expression, 242 after 2H of expression, and 20 after 12H of expression) were tested for beta-galactosidase activity53. Out of the 264 His+ clones, 64 were LacZ + (2 clones issued from screen without expression, 42 obtained after 2H of expression, and 20 after 12H of expression). A fraction of the yeast plasmid minipreparation from each His + /LacZ + clone was used to transform HB101 E. coli. Plasmids extracted from the 64 E. coli clones was transformed again into either the L40 strain carrying LexA/SGS3 to confirm interaction or an L40 carrying an unrelated hybrid protein, LexA/Lamin, to test for false positive clones. Only 37 His + /LacZ+ clones were specific for LexA/SGS3 and in frame with the GAL4 sequence. Among these 37 clones, 28 interact with the C‐terminal three coiled‐coil domains of SGS3, which included the two clones recovered on a selective medium without expression (suggesting a strong interaction between the two partners). One of these two clones corresponded to the C-terminal part of CHR11.
SGS3 and CHR11 cDNAs without their stop codon were transferred into the BiFC GATEWAY™‐modified vector prior to being recombined with the N‐ and C‐terminal parts of YFP (YN and YC) to produce p35S:YN‐CHR11, p35S:CHR11-YN, p35S:YC‐CHR11, p35S:CHR11-YC, p35S:YN-SGS3, p35S:SGS3‐YN, p35S:YC-SGS3 and p35S:SGS3‐YC destination vectors6. The eight possible combinations were tested in BiFC, but only the combinations YN‐CHR11 / SGS3‐YC and YN-CHR11 / YC-SGS3 gave a YFP signal. Results are shown for the YN‐CHR11 / SGS3‐YC combination, which gave the best signal. Nicotiana benthamiana plants grown in the glasshouse under 13 h light, 25 °C day temperature, and 17 °C night temperature were used for all the agro-infiltration experiments. The leaves were infiltrated with an overnight culture of Agrobacterium tumefaciens strain C58C1 that was re-suspended to an absorbance at 600 nm of 0.1 with the infiltration medium (10 mM MES, pH5.6, 10 mM MgCl2, 200 μM acetosyringone). For co‐infiltration, equal volumes of both Agrobacterium cultures were mixed before infiltration. Observations were performed 72 h after infiltration. Confocal microscopy was performed on an inverted TCS‐SP2‐AOBS spectral confocal laser‐scanning microscope (Leica Microsystèmes SAS, Rueil‐Malmaison, France)6. Samples were excited with a 514 nm argon laser (50%) with an emission band of 520–550 nm for YFP detection and 640–700 nm for chlorophyll autofluorescence.
Imaging
For confocal imaging, Arabidopsis roots were directly imaged on a Leica TCS-SP5 (Leica Microsystems) equipped with a photomultiplier tube and hybrid detectors with an HCX PL APO CS 20.0 × 0.70 IMM objective. GFP was imaged with 488 nm excitation using an Argon laser.
Leptomycin B treatment
For the nuclear export inhibition experiment, five days-old seedlings were transferred to 100ul liquid MS-medium with Leptomycin B (Sigma LMB, at 5 uM final concentration corresponding to methanol at 4.7% final) and incubated 28 h before imaging. Controls were incubated 28H in 100 ul liquid MS-medium with methanol added at 4.7% final.
Subcellular fractionations and western blot
Two grams of rosette leaves of 31 days-old plants were ground in 16 ml buffer supplemented with 1.14 M sucrose (10 mM Tris–HCl pH 7.5, 5 mM MgCl2, 4 mM spermidine, 1 mM spermine, 1 mM DTT and protease inhibitor cocktail sigma p9599). The sample was filtered through Miracloth and centrifuged at 1000 × g for 10 min at 4 °C. Supernatants recovered after the first centrifugation was supplemented with 0.15% Triton X100 and was centrifuged at 100,000 × g 45 min at 4 °C to obtain the microsomal fractions (pellet). The first pellet was washed twice with 10 ml buffer (10 mM Tris–HCl pH 7.5, 5 mM MgCl2, 4 mM spermidine, 1 mM spermine, 1 mM DTT, and protease inhibitor cocktail sigma p9599) supplemented with 0.15% Triton X100. The last pellet corresponding to the nuclear fraction was resuspend in lysis buffer (10 mM Tris–HCl pH 7, 2% SDS, 10% glycerol, and protease inhibitor cocktail sigma p9599) and sonicate on bioruptor. Protein concentration was quantified using a detergent-compatible BCA kit (Bio-Rad), and 50 μg of protein were loaded on 8% SDS PAGE. Proteins were electroblotted onto nitrocellulose membranes (Amersham Hybond ECL). The membrane was blocked in 5% non-fat dry milk in 1 × TBSTT (0.25% Tween-20, 0.1% Triton X100, NaCl 150 mM) for 1 h at room temperature, rinsed for 5 min in 1 × TBST, and incubated with primary antibody in 5% non-fat dry milk and 1 × TBST for 1 h at room temperature. The membrane was then rinsed in 1 × TBST for 20 min before incubation with HRP-coupled secondary IgGs. Antigens were detected using chemiluminescence for HRP immunoblot (Amersham ECL Plus). The antibodies used were anti-SGS3 (ref B152 from Santacruz, 1/200e dilution) anti-H3 (ab1791 from Abcam, 1/3000e dilution).
ChIP
ChIP was performed on chromatin from 2 g crosslinked in vitro plantlets 15 days after germination mainly54. After 2 × 5 cycles of sonication (30 s “ON,” 30 s “OFF,” High intensity) with a Bioruptor UCD200 (Diagenode), the chromatin solution was diluted 10-fold to a final volume of 3 mL with ChIP dilution buffer before IP. To stabilize protein complexes, double crosslink was also tested using first disuccinimidyl glutarate 2 mM for 45 min at RT, then formaldehyde 1% final concentration for 7 minutes55, followed by sonication using Covaris S220 ultrasonicator 12 min with 5% duty cycle, 105 W peak power and 200 cycles per burst.
For ChIP on L1/pUBQ10:GFP-CHR11, 30 μL of GFP-trap-M beads (gfm-20/500 ul chromotech) was washed twice and resuspended in 60 μL of ChIP dilution buffer.
For ChIP on L1and 6b4/pUBQ10:Flag-CHR11, 60 μL of magnetic beads G was washed twice and resuspended in 60 μL of ChIP dilution buffer. 5 μL of anti-FLAG M2 clone SIGMA(F3165) were added to the beads G and incubated for at least 3 h at 4 °C with gentle rotation on a wheel. After three washes the beads plus antibodies were resuspended in 100 μL of ChIP dilution buffer.
1 mL of the chromatin solution was added to the antibodies plus beads (GFP-trap-M or G beads plus anti-Flag) and was incubated overnight at 4 °C with gentle rotation for GFP or Flag capture.
The washing of beads was performed at 4 °C twice (5mn) in low salt buffer (150 mM NaCl, 0.1% SDS, 1% TritonX-100, 2 mM EDTA, 20 mM Tris-HCl pH8.1), twice in high salt buffer (500 mM NaCl, 0.1% SDS, 1% TritonX-100, 2 mM EDTA, 20 mM Tris-HCl pH8.1), twice in LiCl buffer (0.25 M LiCl, 1% NP40, 1% sodium deoxycholate, 1 mM EDTA, 10 mM Tris-HCl pH 8.1) and twice in TE Buffer (10 mM Tris-HCl pH 8, 1 mM EDTA)56. After the last TE wash, the reverse crosslinking (5 h at 65 °C) and elution were performed using an IPure kit (Diagenode). The final elution was performed in 60 μL, and the chromatin was stored at − 20 °C until analysis.
Using Biorad-CFX-Maestro Software, the ChIP was analyzed by qPCR on 2 μL of the chromatin. Each primer pair are listed in Supplementary Table 3. The mean of the three qPCRs results (with SD < 0.4 cycle threshold) was used for each point. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH)) was used as an internal ref.57. Results are represented as fold change: normalized expression (delta delta Cq) given by the ratio of Relative Quantity of the sample (2(Cq control-Cq sample) for each identical primer with 100% of efficiency) divided by the Relative Quantity of internal ref.58. Here control is L1 for ChIP on L1/pUBQ10:GFP-CHR11 and L1/ pUBQ10:GFP-CHR11 for ChIP on L1/pUBQ10:Flag-CHR11, sample is our target and internal reference is GAPDH. Two to four biological replicates were analyzed each time. Results show the mean and SD of the independent biological replicates.
RIP RT-qPCR
10 g of 14‐day‐old L1/sgs3/pRDR6:SGS3-NLS-GFP and L1 control plants grown in liquid MS medium were directly ground in liquid nitrogen after treatment 1h30 at 38 °C. Cross-linking was performed by adding the equivalent of 20 ml of the powder obtained after grinding to 100 ml of fixation buffer (10 mM Tris-HCl pH8, 0,4 M sucrose BM grade, 10 mM MgCl2, 5 mM DTT, 0.6% Triton X-100, 4 tablets COMPLETE™, PROTEASE INHIBITOR COCKTAIL Roche, 100 ul for 100 ml RNAse Inhibitor SIGMA R7397 and 1% formaldehyde) and incubation at room temperature for 7 min with gentle agitation. To stop the crosslinking reaction, Glycine was added to the solution (0.125 M final). The cross-linked material was transferred on ice, and nuclear isolation was performed using a Dounce tissue grinder set. The lysate was filtered through a miracloth mesh and centrifuged at 2000g for 20 min at 4°C. The pellet was dissolved gently in 20 ml of extraction buffer II (10 mM Tris-HCl pH8, 0,25 M sucrose, 10 mM MgCl2, 5 mM DTT, 1% TRITON X100, protease-inhibitor Roche cOmplete tablets) and centrifuged at 2000 × g for 20 min at 4 °C, dissolved again in 20 ml of extraction buffer II and centrifuged at 2000g for 10 min at 4 °C. The pellet was dissolved in 0.6 ml of extraction buffer III (10 mM Tris-HCl pH8, 1.7 M sucrose, 2 mM MgCl2, 5 mM DTT, 0.15% TRITON X100) and overlayed onto 0.6 ml of the same buffer in a 2 ml tube. After centrifugation at 10000 g for 10 min at 4°C, the pellet was dissolved to a final volume of 1.2 ml in nuclei lysis buffer (50 mM Tris-HCl, pH8, 0.1% SDS, 10 mM EDTA, protease-inhibitor Roche, cOmplete tablets, and 40U of Recombinant Ribonuclease Inhibitor Invitrogen™ RNaseOUT™ 10777019). Sonication was carried out using a cooled PicoRuptor, 5 cycles 30” ON - 30” OFF (low intensity), in Diagenode TPX microtubes. After centrifugation for 5 min at 10000 × g, the supernatant was transferred to a new tube and diluted twice in ChIP dilution buffer (16.7 mM Tris-HCl, pH7.5, 167 mM NaCl, 1.1% Triton-X-100, 1.2 mM EDTA). 5% of the final volume was reserved at − 20 °C as input fraction (total RNA). The remaining nuclei solution was divided in 2 tubes, and 30ul of the antibodies plus beads GFP-trap (chromtek M270) were added to each tube and incubated ON at 4 °C with gentle rotation. The next day the beads were washed on a turning wheel 5mn at 4 °C in 1 ml of the following washing buffers: three times in wash buffer I (Tris-HCl pH8 50 mM, NaCl 150 mM, Triton 1%, SDS 0,1%, Na deoxicholate 0,5%), three times in wash buffer II (10 mM Tris-HCl pH8, 1% Na Deoxycholate, 1% IPEGAL, 1 mM EDTA) and once in Tris-HCl pH8 20 mM. Reverse cross-link and elution of the immune complex from the beads was performed by incubating the beads and the input fraction 2hrs at 55 °C in PK buffer (100 mM Tris-HCl pH8, 50 mM NaCl, 10 mM EDTA, 40U of Recombinant Ribonuclease Inhibitor Invitrogen™ RNaseOUT™ 10777019 and 4 mg/ml proteinase K ROCHE). For RNA extracted from IP eluate and input fraction, TRIzol reagent (Invitrogen) was used according to the manufacturer’s protocol. Extracted RNA was further concentrated by precipitation in 20 μg glycogen, 300 mM sodium acetate pH 5.2, and 1 volume of ispopropanol, followed by mixing, incubation at − 80 °C for 1 h, and centrifugation for 30 min at 13,000 × g at 4 °C. RNA pellets were solubilized in 20ul RNase-free water. For RT-qPCR analysis, RNA was reverse transcribed just after DNAse treatment by SuperScript II reverse transcriptase (Invitrogen) using random Hexamer. cDNA from input and IP were amplified with primers listed in Supplementary Table S3.
Fold enrichment in the immunoprecipitated fraction (IP) compared to input fraction was calculated as 2exp(−ΔΔCt [RIP in SGS3-NLS-GFP line/RIP background in L1 line]), which correspond to ratio between percentage of input calculated for SGS3-NLS-GFP line / L1 control line. Fold enrichment was calculated for the GUS mRNA target, and for a negative control, PP2A transcript arbitrarily set to 1. Mean values and standard errors of three independent experiments are shown.
Small RNAseq library construction, sequencing and analysis
Small RNA (sRNA) libraries were constructed from 1 µg of total RNA treated with DNaseI using the NEBNext® Multiplex Small RNA Library Prep Set for Illumina® kit (New England Biolabs) according to the manufacturer’s instructions. Libraries were sequenced on a NextSeq 500 Sequencing System (Illumina) using 75-nt single-end reads. The sequencing adapters were removed using cutadapt (v2.10) and sequence-matching Arabidopsis ribosomal or transfer RNA were discarded using bowtie (v1.3.1). The reads of length 21nt, 22nt, and 24nt were then mapped on the TAIR10 genome with the help of ShortStack (v3.8.5) without mismatch (--mismatches 0), keeping all primary multi-mapping (--bowtie_m all) and correcting for multi-mapped reads according to the uniquely mapped reads (--mmap u). For each gene annotation in Araport11 (coding genes, non-coding RNA, and miRNA precursor), the accumulation of 21/22nt was counted with ShortStack. The number of sequences corresponding to each mature miRNA of Arabidopsis in miRBase (v22) was used to measure miRNA abundance. 21/22nt sRNA accumulation in each library was normalized using the median ratio of the corresponding miRNA counts inside DESeq2 (v1.34.0). Differential 21/22nt sRNA accumulation and log2 fold changes between each genotype/condition couple were computed using DESeq2. FDR correction of the p-value was used. See Supplementary Data 1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this work are available within the paper and its Supplementary Information files. A reporting summary for this Article is available as a Supplementary Information file. The datasets and plant materials generated and analyzed during the current study are available from the corresponding author upon request. Small RNAseq data are accessible through NCBI’s Gene Expression Omnibus accession code GSE278529. All relevant data are available from the authors. Source data are provided in this paper.
References
Vaucheret, H. & Voinnet, O. The plant siRNA landscape. Plant Cell 36, 246–275 (2024).
Bologna, N. G. & Voinnet, O. The diversity, biogenesis, and activities of endogenous silencing small RNAs in Arabidopsis. Annu. Rev. Plant Biol. 65, 473–503 (2014).
Yoshikawa, M. et al. 3’ fragment of miR173-programmed RISC-cleaved RNA is protected from degradation in a complex with RISC and SGS3. Proc. Natl. Acad. Sci. USA 110, 4117–4122 (2013).
Yoshikawa, M., Peragine, A., Park, M. Y. & Poethig, R. S. A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes Dev. 19, 2164–2175 (2005).
Blagojevic, A. et al. Heat stress promotes Arabidopsis AGO1 phase separation and association with stress granule components. iScience 27, 109151 (2024).
Elmayan, T. et al. A neomorphic sgs3 allele stabilizing miRNA cleavage products reveals that SGS3 acts as a homodimer. FEBS J. 276, 835–844 (2009).
Jouannet, V. et al. Cytoplasmic Arabidopsis AGO7 accumulates in membrane-associated siRNA bodies and is required for ta-siRNA biogenesis. EMBO J. 31, 1704–1713 (2012).
Kumakura, N. et al. SGS3 and RDR6 interact and colocalize in cytoplasmic SGS3/RDR6-bodies. FEBS Lett. 583, 1261–1266 (2009).
Napoli, C., Lemieux, C. & Jorgensen, R. Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2, 279–289 (1990).
van der Krol, A. R., Mur, L. A., Beld, M., Mol, J. N. & Stuitje, A. R. Flavonoid genes in petunia: addition of a limited number of gene copies may lead to a suppression of gene expression. Plant Cell 2, 291–299 (1990).
Moreno, A. B. et al. Cytoplasmic and nuclear quality control and turnover of single-stranded RNA modulate post-transcriptional gene silencing in plants. Nucleic Acids Res. 41, 4699–4708 (2013).
Parent, J. S. et al. Post-transcriptional gene silencing triggered by sense transgenes involves uncapped antisense RNA and differs from silencing intentionally triggered by antisense transgenes. Nucleic Acids Res. 43, 8464–8475 (2015).
Lange, H. et al. RST1 and RIPR connect the cytosolic RNA exosome to the Ski complex in Arabidopsis. Nat. Commun. 10, 3871 (2019).
Li, T. et al. A genetics screen highlights emerging roles for CPL3, RST1 and URT1 in RNA metabolism and silencing. Nat. Plants 5, 539–550 (2019).
Martinez de Alba, A. E. et al. In plants, decapping prevents RDR6-dependent production of small interfering RNAs from endogenous mRNAs. Nucleic Acids Res. 43, 2902–2913 (2015).
Scheer, H. et al. The TUTase URT1 connects decapping activators and prevents the accumulation of excessively deadenylated mRNAs to avoid siRNA biogenesis. Nat. Commun. 12, 1298 (2021).
Zhang, X. et al. Plant biology. Suppression of endogenous gene silencing by bidirectional cytoplasmic RNA decay in Arabidopsis. Science 348, 120–123 (2015).
Cao, M. et al. Virus infection triggers widespread silencing of host genes by a distinct class of endogenous siRNAs in <em>Arabidopsis</em>. Proc. Natl. Acad. Sci. USA 111, 14613–14618 (2014).
Borsani, O., Zhu, J., Verslues, P. E., Sunkar, R. & Zhu, J. K. Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 123, 1279–1291 (2005).
Zhang, X. et al. Genome-wide analysis of plant nat-siRNAs reveals insights into their distribution, biogenesis and function. Genome Biol. 13, R20 (2012).
Knizewski, L., Ginalski, K. & Jerzmanowski, A. Snf2 proteins in plants: gene silencing and beyond. Trends Plant Sci. 13, 557–565 (2008).
Pontes, O. et al. Intersection of small RNA pathways in Arabidopsis thaliana sub-nuclear domains. PLoS ONE 8, e65652 (2013).
Kim, E. Y. et al. Ribosome stalling and SGS3 phase separation prime the epigenetic silencing of transposons. Nat. Plants 7, 303–309 (2021).
Tan, H. et al. Phase separation of SGS3 drives siRNA body formation and promotes endogenous gene silencing. Cell Rep. 42, 111985 (2023).
Waese, J. et al. ePlant: Visualizing and exploring multiple levels of data for hypothesis generation in plant biology. Plant Cell 29, 1806–1821 (2017).
Chen, Q. et al. Overexpression of ATG8 in arabidopsis stimulates autophagic activity and increases nitrogen remobilization efficiency and grain filling. Plant Cell Physiol. 60, 343–352 (2019).
Huanca-Mamani, W., Garcia-Aguilar, M., Leon-Martinez, G., Grossniklaus, U. & Vielle-Calzada, J. P. CHR11, a chromatin-remodeling factor essential for nuclear proliferation during female gametogenesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 102, 17231–17236 (2005).
Li, G. et al. Imitation Switch chromatin remodeling factors and their interacting RINGLET proteins act together in controlling the plant vegetative phase in Arabidopsis. Plant J. 72, 261–270 (2012).
Luo, Y. X. et al. A plant-specific SWR1 chromatin-remodeling complex couples histone H2A.Z deposition with nucleosome sliding. EMBO J. 39, e102008 (2020).
Zhang, Q. et al. DDT-RELATED PROTEIN4-IMITATION SWITCH alters nucleosome distribution to relieve transcriptional silencing in Arabidopsis. Plant Cell 35, 3109–3126 (2023).
Daxinger, L. et al. Unexpected silencing effects from T-DNA tags in Arabidopsis. Trends Plant Sci. 13, 4–6 (2008).
Mlotshwa, S. et al. Transcriptional silencing induced by Arabidopsis T-DNA mutants is associated with 35S promoter siRNAs and requires genes involved in siRNA-mediated chromatin silencing. Plant J. 64, 699–704 (2010).
Gy, I. et al. Arabidopsis FIERY1, XRN2, and XRN3 are endogenous RNA silencing suppressors. Plant Cell 19, 3451–3461 (2007).
Hematy, K. et al. The Zinc-Finger Protein SOP1 Is Required for a Subset of the Nuclear Exosome Functions in Arabidopsis. PLoS Genet. 12, e1005817 (2016).
Lange, H. et al. The RNA helicases AtMTR4 and HEN2 target specific subsets of nuclear transcripts for degradation by the nuclear exosome in Arabidopsis thaliana. PLoS Genet. 10, e1004564 (2014).
Yu, A. et al. Second-site mutagenesis of a hypomorphic argonaute1 allele identifies SUPERKILLER3 as an endogenous suppressor of transgene posttranscriptional gene silencing. Plant Physiol. 169, 1266–1274 (2015).
Li, G. et al. ISWI proteins participate in the genome-wide nucleosome distribution in Arabidopsis. Plant J. 78, 706–714 (2014).
Zhu, Y., Rowley, M. J., Bohmdorfer, G. & Wierzbicki, A. T. A SWI/SNF chromatin-remodeling complex acts in noncoding RNA-mediated transcriptional silencing. Mol. Cell 49, 298–309 (2013).
Krzyszton, M. & Kufel, J. Analysis of mRNA-derived siRNAs in mutants of mRNA maturation and surveillance pathways in Arabidopsis thaliana. Sci. Rep. 12, 1474 (2022).
Hua, X. et al. Global analysis of RNA-dependent RNA polymerase-dependent small RNAs reveals new substrates and functions for these proteins and SGS3 in arabidopsis. Noncoding RNA 7, https://doi.org/10.3390/ncrna7020028 (2021).
Fukunaga, R. & Doudna, J. A. dsRNA with 5’ overhangs contributes to endogenous and antiviral RNA silencing pathways in plants. EMBO J. 28, 545–555 (2009).
Bologna, N. G. et al. Nucleo-cytosolic shuttling of ARGONAUTE1 prompts a revised model of the plant microRNA pathway. Mol. Cell 69, 709–719 (2018).
Beclin, C., Boutet, S., Waterhouse, P. & Vaucheret, H. A branched pathway for transgene-induced RNA silencing in plants. Curr. Biol. 12, 684–688 (2002).
Elmayan, T. et al. Arabidopsis mutants impaired in cosuppression. Plant Cell 10, 1747–1758 (1998).
Jauvion, V., Elmayan, T. & Vaucheret, H. The conserved RNA trafficking proteins HPR1 and TEX1 are involved in the production of endogenous and exogenous small interfering RNA in Arabidopsis. Plant Cell 22, 2697–2709 (2010).
Mourrain, P. et al. Arabidopsis SGS2 and SGS3 genes are required for posttranscriptional gene silencing and natural virus resistance. Cell 101, 533–542 (2000).
Souret, F. F., Kastenmayer, J. P. & Green, P. J. AtXRN4 degrades mRNA in Arabidopsis and its substrates include selected miRNA targets. Mol. Cell 15, 173–183 (2004).
Nakagawa, T. et al. Improved Gateway binary vectors: high-performance vectors for creation of fusion constructs in transgenic analysis of plants. Biosci. Biotechnol. Biochem. 71, 2095–2100 (2007).
Grefen, C. et al. A ubiquitin-10 promoter-based vector set for fluorescent protein tagging facilitates temporal stability and native protein distribution in transient and stable expression studies. Plant J. 64, 355–365 (2010).
Schwab, R., Ossowski, S., Riester, M., Warthmann, N. & Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18, 1121–1133 (2006).
Butel, N., Le Masson, I., Bouteiller, N., Vaucheret, H. & Elmayan, T. sgs1: a neomorphic nac52 allele impairing post-transcriptional gene silencing through SGS3 downregulation. Plant J. 90, 505–519 (2017).
Kim, J., Harter, K. & Theologis, A. Protein-protein interactions among the Aux/IAA proteins. Proc. Natl. Acad. Sci. USA 94, 11786–11791 (1997).
Kieffer, F. et al. Cloning of Rac and Rho-GDI from tobacco using an heterologous two-hybrid screen. Biochimie 82, 1099–1105 (2000).
Le Masson, I. et al. Mutations in the Arabidopsis H3K4me2/3 demethylase JMJ14 suppress posttranscriptional gene silencing by decreasing transgene transcription. Plant Cell 24, 3603–3612 (2012).
Bouyer, D. et al. Genome-wide identification of RETINOBLASTOMA RELATED 1 binding sites in Arabidopsis reveals novel DNA damage regulators. PLOS Genet. 14, e1007797 (2018).
Gendrel, A. V., Lippman, Z., Martienssen, R. & Colot, V. Profiling histone modification patterns in plants using genomic tiling microarrays. Nat. Methods 2, 213–218 (2005).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W. R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17 (2005).
Butel, N. et al. Contrasting epigenetic control of transgenes and endogenous genes promotes post-transcriptional transgene silencing in Arabidopsis. Nat. Commun. 12, 2787 (2021).
Acknowledgements
We thank Hervé Ferry and Philippe Maréchal for taking care of the plants. We thank François Roudier and Jerémie Bazin for the discussion and advices respectively, on the ChIP protocol adapted for chromatin remodeler proteins and on the RIP protocol. We also thank Martin Lacroix for fruitful discussions. Research in the Vaucheret laboratory is supported by grants from the French Agence Nationale pour la Recherche (ANR-16-CE12-0032 and ANR-20-CE12-0009). The IJPB benefits from the support of the LabEx Saclay Plant Sciences-SPS (ANR-17-EUR-007). This work has benefited from the support of IJPB’s Plant Observatory technological platforms.
Author information
Authors and Affiliations
Contributions
T.E. and H.V. designed the experiments. T.E., T.B., E.E.M., I.L.M., A.C., and N.B. performed the experiments. T.E, T.B., M.D.C., and H.V. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yuichiro Watanabe, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Elmayan, T., Blein, T., Elvira-Matelot, E. et al. Arabidopsis SGS3 is recruited to chromatin by CHR11 to select RNA that initiate siRNA production. Nat Commun 16, 2978 (2025). https://doi.org/10.1038/s41467-025-57394-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57394-5