Abstract
Animal behavior is linked to the gene regulatory network (GRN) coordinating gene expression in the brain. Eusocial honeybees, with their natural behavioral plasticity, provide an excellent model for exploring the connection between brain activity and behavior. Using single-nucleus RNA sequencing and spatial transcriptomics, we analyze the expression patterns of brain cells associated with the behavioral maturation from nursing to foraging. Integrating spatial and cellular data uncovered cell-type and spatial heterogeneity in GRN organization. Interestingly, the stripe regulon is explicitly activated in foragers’ small Keyon cells, which are implicated in spatial learning and navigation. When worker age is controlled in artificial colonies, stripe and its key targets remained highly expressed in the KC regions of bees performing foraging tasks. These findings suggest that specific GRNs coordinate individual brain cell activity during behavioral transitions, shedding light on GRN-driven brain heterogeneity and its role in the division of labor of social life.
Similar content being viewed by others
Introduction
Animal behaviors are mainly coordinated by the nervous system transmitting signals across different body areas1,2. Brain, as a major part of the central nervous system, contains neurons that send electrochemical waves rapidly to modulate other cells, which drives specific behavioral responses3. In addition, dynamic genomic processes reflected by the transcriptional states within the brain cells are also implicated in behavioral changes4. Specific changes in brain gene expression have been linked to behavioral responses to environmental stimuli and even can accurately predict behavioral states5. While genes may not directly specify behaviors, they mediate the production of cell molecules, such as neurotransmitters6,7 and hormones8, that govern brain functions, and behavioral states are expressed. For the examination of the relationship between brain gene expression and behaviors, there is a particular focus on the transcriptional profile related to social behaviors4. Social behavior characterizes the complex interactions among individuals and is normally beneficial to the engaged individuals within the species9. Compared to other traits, social behaviors are highly plastic during the lifetime of the individuals10. Studies of the social phenotypes of multiple species have revealed that brain gene expression mediates plastic responses in the context of social organization11,12,13.
Insects provide ideal models for the study of social behavior as they exhibit extremes of social organization from solitary to complex societies where groups of individuals perform specific tasks14,15. The honeybee is one of the few models with a highly evolved social structure and a rich behavioral repertoire16, and the colony is organized by a complex and sophisticated division of labor17. In addition to the reproductive castes (queens and male drones), worker honeybees show distinct labor division patterns. The newly emerged adult bees spend the first ~2 weeks inside the hive and perform most nursing tasks, and then they transit to foraging foods outside the hive18. Naturally, nursing and foraging states are age-related, and many independent studies have documented the close relationship between behavioral maturation in honeybees and the bulk brain neuro-genomic state11,19,20,21. However, it still lacks the resolution to assign the cell heterogeneity and distinguish the expression differences across cell populations.
Although the honeybee possesses a relatively small brain consisting of ~1 million neurons, its neural organization is much more elaborate than Drosophila22. Moreover, the honeybee brain comprises distinct regions that perform various advanced functions, such as the mushroom bodies (MBs), optic lobes (OLs), and antennal lobes (ALs), coordinating the brain activity23,24,25 and, thus, resulting the behaviors. Recent studies using single-cell RNA sequencing technology find that the honeybee brain is composed of multiple cell types showing differential gene expression patterns, which are associated with the reproduction caste26 and the aggressive behavior in honeybees27. However, the comprehensive profiling of the entire brain regarding behavioral transition from within-hive tasks to foraging remains elusive. Particularly, behavior-specific neuro-genomic states were influenced by GRN which seek to describe the relationship between transcription factors (TFs) and their putative target genes19,28. Additionally, the behavior-associated GRNs in the brain are manifested across functional regions containing multiple cell types, which may show specific activities of GRNs with the behavior transitions29. Thus, uncovering both the cell and spatial heterogeneity in the GRN organization will provide insights into molecular characteristics of the brain associated with the behavioral states.
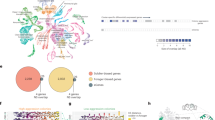
Here, we constructed a high-resolution molecular landscape of brain across natural behavior maturation of honeybee workers (10-day-old nurse and 22-day-old forager) using single-nucleus RNA sequencing (snRNA-seq) and spatial transcriptomics (Fig. 1a). Typical cell types, including Kenyon, glial, and optic lobe cells, with distinguished patterns of gene expression led by specific combinations of TFs, displayed different activity of GRN. Comparing the two classical state of workers, nurses and foragers, we uncovered the dynamic expression changes and deciphered the regulatory code for specific cell populations in spatial and cellular profiles. By integrating both of these profiles, we found brain regions, such as the mushroom body exhibiting distinct GRN activity during the behavioral transition.
a Schematic diagram of the experimental design. b Annotated tSNE visualization of the clustering of 121,247 single-nucleus transcriptomes obtained from nurse (70,060) and forager (51,187) bees. KC: Kenyon cells; Glia: glial cells; OLC: optic lobe cells; OPN: olfactory projection neurons; Hem: hemocytes; IPC: insulin-producing cells; Others: unknown cell type. Cluster identifiers from Seurat clustering at a resolution of 0.2 are indicated in parentheses. c Selected marker genes for the cell clusters annotated in nurse and forager bee brains. The expression levels of indicated genes were evaluated by the collapsed pseudo-bulk expression in each cluster as a percentage of unique molecular identifiers (UMIs) of the total cluster. Bars represent the means of 54 biological replicates + SEM. Cluster labels correspond to those shown in Supplementary Fig. S4. Source data are provided as a Source Data file. d Normalized UMIs per cell for the markers of Kenyon (dopR2, mAChR-DM1) and glial (repo, glnS) cells Heatmap plotted over global tSNE.
Results
Identification of discrete cell types in honeybee brain
We isolated nuclei for brains from 136 nurse and 80 forager honeybee individuals. All brain tissues were profiled by snRNA-seq using the DNBelab C4 droplet-based platform. We first included all samples, regardless of caste, to get a comprehensive description of cell types. We obtained transcriptomic data for 143,228 nuclei and filtered doublets and cells with a low number of genes. Finally, 121,247 high-quality cells were retained from 54 libraries (Supplementary Figs. S1–S3; Supplementary Dataset S1). An average sequencing depth of 88,867 reads per cell was achieved, and ~90% of reads could be mapped to the genome of A. mellifera, with ~50% of reads mapped to the exonic regions. The average number of detected genes per cell is 1213 in nurse and 1038 in forager bees.
Depending on the clustering resolution, we obtained 32–52 cell clusters (0.2–1.6; Supplementary Fig. S4). In global visualization using t-Distributed Stochastic Neighbor Embedding (tSNE), the cells from nurses and foragers did not cluster separately (Supplementary Fig. S5), indicating the consistency of cell types between the two castes. Cell types of 24 clusters (resolution = 0.2) were annotated using known markers from single-cell transcriptomes of Drosophila and Harpegnathos30,31,32,33,34. Generally, we observed characteristic cell types of the brain, including 29.0% Kenyon cells (KCs), 12.0% glia, 13.0% optic lobe cells (OLCs), 2.8% olfactory projection neurons (OPNs), 0.5% hemocytes, and 2.2% insulin-producing cells (Fig. 1b; Supplementary Figs. S6 and S7). The expression of classical markers documented in A. mellifera confirmed our annotation (Fig. 1c, d; Supplementary Dataset S2). Seven subclusters of the glia were characterized (Supplementary Fig. S8). Subpopulations of ensheathing (marked by CG7509), cortex (wrapper, CG6356, zyd), and surface (Glob1, Indy, Tret1) glia were identified. Three subclusters of astrocytes were distinguished by a set of markers (Eaat1, Gat, Rh50, CG1495, CG9657), and one cluster (G1) could not be assigned to a known transcriptional type. Eight distinct subclusters were identified from OLCs with different expression patterns. Cluster 8 was identified as OPNs by Oaz30 To further validate our annotation, the cell clusters from nurses and foragers were mapped to the whole-brain profiles of Drosophila melanogaster30 and Monomorium pharaonic ant35. The cross-species analysis based on AUROC scores revealed that the major cell types (KC, glia, OLC) were more conserved between honeybees and ants, but only glia showed relatively high similarity between honeybees and Drosophila. (Supplementary Fig. S9). In addition, the KCs and glial cells from nurses and foragers clustered according to cell types rather than worker castes (Supplementary Fig. S10). In addition, it revealed differentially expressed genes (DEGs) for distinct cell types (Supplementary Dataset S3). Gene ontology (GO) analysis on DEGs in each cell type revealed significant enrichment in different biological functions (Supplementary Fig. S11; Supplementary Dataset S4). For example, GO terms of “learning or memory” and “cell communication” were explicitly enriched in KCs, while OLCs and OPNs were enriched for “axon” and “neuron projection”. Taken together, our snRNA-seq atlas revealed the common cell types populating the nurse and forager honeybee brains.
Constructing spatial landscape of cell types in honeybee brain
Since we identified the major cell types in the bee brain, we next attempted to profile the spatial map with cellular resolution. We applied Stereo-seq to the brain sections of both nurses and foragers. The honeybee brain is oval in shape and contains several principal parts, including the MBs, optic lobes, central complex, and antennal lobes (Fig. 2a). The HE-stained section showed a relatively intact tissue in our sample, and the major structures could be distinguished (Fig. 2b). Two cryosection slices from nurse and forager were loaded on the chip, and the tissue regions were identified by ssDNA staining (Fig. 2c). We aggregated the datasets for each section into 50 × 50 DNB bins (bin 50) and obtained a total of 4741 bins. We retrieved ~18,000 unique transcripts of more than 1200 genes per bin from both samples (Supplementary Fig. S12). Unsupervised spatially constrained-clustering of the bins identified functional regions generally matching the localization of anatomically defined parts in the bee brain. Specifically, the paired cup-like calyx region of MBs was dominated by one specific cluster, and three contiguous clusters were distinguished within the outer layers of optic lobes, mainly resembling the regions of the retina, lamina, and part of the medulla (Fig. 2d, e). However, the anatomic regions of other brain parts (e.g., VL, AL, OC) could not be distinguished by clear boundaries of clusters, possibly because the slides are more posterior portions of the brains25. We next validated our spatial atlas by aligning the snRNA-seq data onto the brain sections. The projection by Tangram identified the localization of the KCs, OLCs, and glia across different brain regions (Fig. 2f, g). In both sections of nurse and forager bees, KCs and OLCs were condensed in the MB and optic lobe regions, respectively. In contrast, glial cells were spread on the whole central brain, which accords with the previous investigation by immunolabeling36. Thus, we generated a topographic transcriptomic atlas of the honeybee brain for the following investigations of spatial heterogeneity in cell activities.
a A schematic view of the honeybee brain with the main structures, including the mushroom body (MB), ocelli (OC), vertical lobe (VL), antennal lobe (AL), lobula (LO), medulla (ME), lamina (LA), and retina (RE). b Hematoxylin-eosin staining of one of the representative brain sections of the honeybee. c Nucleic-acid dye staining of the brain section shown in (b) attached on the Stereo-seq chip. Unsupervised clustering of the brain sections from nurse (d) and forager (e) bees based on the Stereo-seq data at bin-50 (50 × 50 DNB) resolution. Bins are colored based on the annotation of the clustering. Projection of our snRNA-seq data onto the brain sections of nurse (f) and forager (g) bees using Tangram. The color represents the relative probability of each cell type being mapped to the specific locations within the sections. Scale bars, 0.3 mm.
Spatial heterogeneity and evolution of the cell subtypes
Compared with other insects, honeybees’ MBs are more developed37. Morphological and molecular analysis has found four KC subtypes in the honeybee brain: Class-I large-type (lKCs), middle-type (mKCs), small-type (sKCs) KCs, and Class-II KCs. Preferentially expressed genes have been identified in KC subtypes38; we further reclustered all KC transcriptomes from our dataset. This identified five distinct populations within KCs (Fig. 3a), and the lKCs (marked by Mblk1), mKCs (mKAST), and sKCs (E74) could be distinguished by previously documented marker genes (Fig. 3b, c)38. We identified two subclusters for Class-II KCs: II-KC1 showed high relative expression of Trp and Fstl5, while II-KC2 was characterized by a set of markers (Apisα7-2, Mrjp3, PKC, sgg). Additionally, many other genes were expressed in a subtype-preferential manner (Supplementary Dataset S5), and the GO enrichment analysis suggested distinct functions of KC subtypes. DEGs in lKCs are involved in visual functions (e.g., ommatidial planar polarity), while those in Class-II KCs are associated with locomotor rhythm, associative learning, and memory (Fig. 3d).
a tSNE visualization for the subclustering of Kenyon cells. lKC: Class I large-type Kenyon cells; mKC: Class I middle-type Kenyon cells; sKC: Class I small-type Kenyon cells; II-KC: Class II Kenyon cells. Cluster identifiers from Seurat clustering at a resolution of 0.2 are indicated in parentheses. b Discernible marker genes for the subclusters annotated in Kenyon cells. Bars represent the means of 54 biological replicates +SEM. c Heatmap showing the average expression scale for marker genes in subclusters of Kenyon cells. d Representative GO terms enriched by the DEGs in different subclusters of KCs. Fold change is calculated as the proportion of detected genes within each GO term. Dot size represents the normalized fold change. Significance is assessed using the two-sided hypergeometric test. e Correspondence of KC clusters between Apis mellifera, Monomorium pharaonis35, and Drosophila melanogaster30 determined by MetaNeighbor. The lines with different thickness link KC clusters from A. mellifera with those from M. pharaonis and D. melanogaster according to the AUROC scores. Only transcription similarities with AUROC > 0.6 are shown (Supplementary Fig. S13). Source data are provided as a Source Data file.
Since different subtypes of KCs have also been characterized in other insect species, we then compared the transcriptional similarity of KCs between honeybee39 and other datasets from M. pharaonis35, H. saltator33, and D. melanogaster30,31,34 (Fig. 3e; Supplementary Fig. S13). Most KC clusters from honeybees showed high similarity to those from the two ant species, exhibiting advanced social organization. Specifically, the l- and mKCs of A. mellifera were similar to the Class-A KCs of ant species, while the Class-B KCs from ants were most similar to the Class-II KCs from honeybees. Comparatively, only the γ- and α’β’-type KCs of Drosophila showed relatively higher similarity to sKCs and II-KCs of the honeybee, respectively (AUROC > 0.9).
We also reclustered the eight subtypes belonging to the OLCs in our atlas (Fig. 4a). We compared the honeybee OLC clusters with those from Monomorium ants35 and two datasets of Drosophila optic lobe40,41. Although many honeybee OLC clusters showed high similarity to those from Monomorium ants35 (Fig. 4b; Supplementary Fig. S14c), some honeybee OLCs had strong similarity to OLC subclusters of Drosophila40,41 (Supplementary Fig. S14a, b). Interestingly, the OLC4 of A. mellifera showed a similar gene expression pattern to the c20 in M. pharaonic (AUROC = 0.98) and the lamina monopolar cells (L1-L5) in Drosophila (AUROC = 0.99) (Fig. 4b, Dataset S5). Specifically, OLC4 is highly similar to the L1–3 neurons of Drosophila expressing markers for chloride channel42. L1 and L2 depolarize in response to gamma-aminobutyric acid (GABA) and store vesicular glutamate43,44. Correspondingly, OLC4 of honeybee highly expressed marker genes related to chloride (ort, Lcch3), GABA (Grd, GABA-B-R3), and glutamate (GluClalpha, Ekar) channels (Supplementary Fig. S14d; Supplementary Dataset S6), suggesting relative conserved molecular functions of visual neurotics in the lamina45. In addition, the OLC1 and OLC8 were similar to the distal and proximal medulla neurons of Drosophila. Although the OLC6 of A. mellifera was not similar to the major OLC cell types from either Monomorium or Drosophila, it showed a high AUROC scores to several Drosophila subclusters (e.g., TmY3, Dm4 and Tm2; Supplementary Fig. S14a, b).
a tSNE visualization for the subclustering of OLCs from the worker brains, which are grouped into eight subtypes. Cluster identifiers from Seurat clustering at a resolution of 0.2 are indicated in parentheses. b Correspondence of OLC clusters between Apis mellifera, Monomorium pharaonis35, and Drosophila melanogaster40. The lines with different thickness link OLC clusters from A. mellifera with those from M. pharaonic and D. melanogaster according to the AUROC scores. Source data are provided as a Source Data file. c A schematic diagram of the structural characteristics of neurons from the optic ganglia of the honeybees (adapted from Ribi and Scheel23). svf, short visual fibers; lvf, long visual fibers; L, L-fibre; Tnf, tangential fiber with narrow-field ending; Twf, tangential fiber with wide-field ending; Tm, transmedullary neuron. Spatial visualization of the distribution of the OLC4 subcluster in the optic lobe regions of the brain sections of nurse (d) and forager (e) bees inferred by Tangram. The color of each merged bin is scaled according to the Tangram scores based on the projection of the snRNA-seq dataset. Scale bars, 0.3 mm.
The cell types and the topographical relationships of major neuronal elements in the optic lobes of worker honeybees have been well described by Ribi23,46 (Fig. 4c). Two types of fiber elements, the short (svf) and the long visual fibers (lvf) form the optic cartridges with the monopolar cells, L-fibers, within the lamina. In addition, two types of T-cells, the narrow (Tnf) and wide field (Twf) fibers and transmedullary cells (Tm), were also defined in bee brain. Interestingly, the Tangram projection of the snRNA-seq data revealed that the OLC4 subtype was specifically enriched in the lamina region of nurses and foragers (Fig. 4d, e). This observation suggests that the OLC4 cells exhibiting homology to the Drosophila laminar monopolar cells probably represent the L-fibers in the first optic ganglion of the honeybee46.
Cell type-specific gene expression changes associated with behavioral maturation
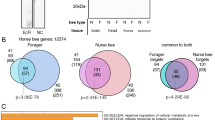
Since we have generated a cellular transcriptomic atlas for the honeybee brain, we examined the gene expression during the transition from in-hive tasks to foraging outside. We identified 1265 and 1016 genes upregulated in forager and nurse bees, respectively (Supplementary Fig. S15; Supplementary Dataset S7a). Among the top ten differentially expressed genes across four cell types, Mrjp1-5 were highly expressed in the KC, OLC, OPN, and Glia of nurses, which agrees with the previous finding that mrjp is involved in regulating phenotypic plasticity47. Whereas LOC724367 (l(2)efl) were generally upregulated in various cell types of foragers, which participate in eIF2 alpha phosphorylation, which may enhance the translation of specific mRNAs to promote memory consolidation48. However, some of the differentially expressed genes are restricted to one cell type, there are 48.4% and 63.5% of these genes was cell-type specific for forager and nurse, respectively (Fig. 5a, b). To examine the gene expression variations conditional on cell type, we first compared our snRNA-Seq data to three independent bulk assays by Whitfield et al.11, Alaux et al.49, and Khamis et al.19 (Fig. 5c, d). In foragers, the bulk data were dominated by expression changes identified in KCs. Conversely, in nurses, cellular changes in glia and OPNs could be captured only in the A-MEXP-755 dataset.
Bar plots display DEGs identified in different cell types of (a) foragers (n = 1265) and (b) nurses (n = 1016). Significance assessed by two-sided quasi-likelihood F-tests in edgeR, adjusted by Benjamini–Hochberg. DEGs were identified as genes with an absolute log2FC > 1 and an adjusted P value < 0.05. Bars indicate whether a gene is a DEG in a given cell type. Overlap of DEGs between single-cell clusters and bulk brain11,19,49 in forager (c) and nurse (d) bees. The color of each square represents the proportion of overlapping DEGs, normalized within each bulk dataset. Significance is assessed using the two-sided Fisher’s exact test. e Heatmap showing upregulated DEGs from different subclusters of cells in forager (upper) or nurse (down). Colors represent the normalized log2FC between nurses and foragers across different cell types. Source data are provided as a Source Data file.
Moreover, both DEGs in nurses and foragers exhibited subtype-specificity (Supplementary Fig. S16, Supplementary Dataset S7b). To identify subtypes with the most pronounced differential expression between nurse and forager bees, we normalized log2 fold change (log2FC) values of gene expression to a 0-1 scale across all subtypes (Fig. 5e). Among the genes analyzed, the Pban, which encodes the pheromone biosynthesis-activating neuropeptide50 and caveolin-3, known to enhance synaptic plasticity and learning ability51,52, displayed the most significant differential expression in sKCs. The astrocytes of nurses showed high expression of apidermin 2 (Apd-2), which is also upregulated in the heads of young bees53. Collectively, these results suggest that the gene expression shifts detected by bulk assays could be attributed to specific cell types, and single-cell resolution is implicated in finding cell heterogeneity in response to behavioral states.
Spatial heterogeneity of gene regulatory networks
In addition to the gene expression within the cell, the GRN coordinating the observed expression changes is an important driver of behavioral plasticity54. TFs are key proteins in linking signaling transduction networks to gene-specific transcriptional regulation55. In order to identify GRNs and the activities of TFs across individual cells, we built a SCENIC database for A. mellifera and inferred the co-expression networks. In total, 184 regulons (out of 602 initial co-expression modules) were identified with significantly enriched motifs (NES score >3) with their corresponding TFs (Fig. 6a; Supplementary Dataset S8). Notably, 19 TFs have been identified as potential key regulators of behavioral plasticity in the honeybee brain19,28,54. Cell clustering based on the AUCell scores resulted in 14 SCENIC clusters, showing a different t-SNE projection compared to that based on gene expression (Fig. 6b). Each SCENIC cluster was dominated by different cell types (Fig. 6c, d), and the SCENIC clusters were grouped according to the cell categories (Fig. 6e, f). Correspondingly, each cell type exhibited a specific pattern of regulon activities, indicating that the same type of cells possessed similar activities of regulatory networks. AUCell scoring revealed potential master regulators driving specific states for cell types (Supplementary Dataset S9a). KCs were mainly regulated by Vsx2 and hormone receptors (Hr51). Bsh was the marker regulon for both OPNs and OLCs, which corroborated with the optic lobe of Drosophila30. Glia was recognized by the activity of the Repo, Dll, and HGTX networks56.
a Normalized enrichment score (NES) and the size of all 184 regulons identified from honeybee brains. Nineteen regulons for transcription factors (TFs) previously associated with behavioral maturation of honeybees19,28,54 are indicated in red, with their highest-scoring motif shown. The inset shows the size distribution for target genes of each regulon. b SCENIC-based tSNE showing 14 clusters based on regulon activity (AUCell score) for each cell. Proportions of different (c) cell types and (d) subtypes in each SCENIC cluster. Cluster labels correspond to those shown in (b). SCENIC-based tSNEs showing the cluster of cell types (e) and subtypes (f) with similar regulatory states. Source data are provided as a Source Data file.
Based on the constructed SCENIC database, we compared the activity of the regulons from different cell populations in nurse versus forager bees (Fig. 7a; Supplementary Dataset S9b). The br and usp regulon showed significantly higher activity in most cell types of nurse bees. The Hr38 and stripe regulons were enriched in forager KCs, especially in sKCs. GRN is often combinatorial that TFs cooperate through regulating the same genes57. The regulatory network showed a substantial overlap among master regulons in KCs, including stripe, CrebB, Hr38, forming a complex network consisting of 525 genes (Fig. 7b; Supplementary Dataset S10). Moreover, the TF stripe and CrebB exhibited self-regulation by targeting each other or themselves. These indicate that honeybee brain cells show differential regulatory networks of TFs, which may be related to the behavioral patterns of honeybees.
a Differential regulon activity between forager and nurse bee brains across cell subtypes. Nineteen regulons previously associated with bee maturation are shown here19,28,54. The size of the square represents a -log10(adj. P) of the two-sided t-test, and P value is adjusted using the Benjamini–Hochberg method. The color indicates the normalized log2FC across different cell types. Log2FC > 0 indicates upregulation in foragers. b Regulatory network for the co-regulatory factors CrebB, stripe, Hr38. Regulons for each TF are represented by different edge colors. The target genes are shown in circles; the TFs and the corresponding motifs are shown in hexagon nodes. c Volcano plot showing the regulons with differential activity scores in functional subregions of the mushroom body and the optic lobe. The significance is determined by two-sided t-test, adjusted by Benjamini–Hochberg correction. d Spatial visualization of the normalized activity score of the regulon stripe at mushroom body regions of nurse and forager brains. Scale bars, 0.3 mm. Fluorescent in situ hybridization of stripe (e), Hr38 (f) and caveolin-3 (g) in the bee brains. Simultaneous detection of target gene mRNA (red) and (4’, 6-diamidino-2-phenylindole) DAPI-stained nuclei (white), using confocal microscopy. Brain sections from 22-day forager, 10-day forager and 10-day nurse bees, respectively. Scale bars = 20 um. Each experiment was repeated independently three times. h The bar plot displays the variation in fluorescence intensity within the mushroom body for stripe (top) and its target genes Hr38 (middle) and caveolin-3 (bottom). The comparative analysis was conducted using the two-sided LSD test, with a significance level of 0.05, adjusted by Benjamini–Hochberg correction. Each point represents a randomly selected fluorescence region (n = 5). Bars represent the average intensity + SEM. Source data are provided as a Source Data file.
We next explored the GRN activities and compared the activities of regulons across spatial locations (Fig. 7c). For nurse, the most outer layer of the optic lobe was distinguished by the highly active usp regulon. However, this is not consistent with the snRNA-seq analysis, which might be due to the complex structure of the bee optic lobes25. In comparison, both the snRNA- and Stero-seq analyses revealed that the stripe regulon was enriched in the MB regions of forager bees, and the projection-based analysis showed a higher activity score of the stripe regulon at MB regions of forager brains (Fig. 7d). To further confirm the expression patterns of the stripe regulon, we performed fluorescence in situ hybridization (FISH) assays to detect the activity of the TF stripe, and its target genes, Hr38 and caveolin-3. Here, we established single-cohort colonies and collected age-controlled forager and nurse bees (10- and 22-day old foragers, 10-day old nurse bees). Our FISH analyses showed that the fluorescence signal of stripe was concentrated in the small KCs localizing at the inner core of the MB calyces (Supplementary Fig. S17), which is consistent with the results based on the snRNA-seq data (Fig. 7a). Moreover, the stripe transcripts were highly expressed in both 10-day-old and 22-day-old foragers (Fig. 7e, h). Similarly, the target genes Hr38 (Fig. 7f, h) and caveolin-3, (Fig. 7g, h) also showed significantly higher activities in forager brains. Notably, cells expressing caveolin-3 exhibited widespread distribution in MB, consistent with the observed upregulation trend of caveolin-3 expression in the lKCs, mKCs, and sKCs of forager bees (Supplementary Fig. S18). Altogether, our data showed cell-type specific gene regulatory interactions and the GRNs operating in different spatial locations may coordinate the behavioral maturation of honeybees.
Discussion
The MBs, a higher-order center for sensory integration and memory, are conserved in insect brains58. Compared with other insects, A. mellifera possessed a more elaborate MB structure59. Consistently, we find a large proportion of KCs (29.0%) in A. mellifera, which is comparable to that in the social ants (36.0%) but much larger than that in Drosophila (5%)30,33,35, indicating that the large volume of MB is implicated in the development of sociality60. Previous studies suggest that KCs were more prevalent in the brains of social insects35,61. Consistently, the cross-species analysis revealed that both Class-I and -II types of KCs are conserved between the honeybee and the two species of ants. In A. mellifera, morphological analysis documents that four subtypes of KCs localize at specific brain regions, including Class-I large-, small-, middle-type KC and IIKC38. Our findings reveal distinct transcriptomic and functional profiles among the four types of Kenyon cells, underscoring their specialized roles in neural processing and behavior. The spatial transcriptomic data confirmed the location of KCs in the MBs, while we could not distinguish the distribution of specific subtypes within the MB calyces58. Among them, the mKC is a recently defined subtype from Class-I KCs62, and our atlas found that the mKCs showed a high expression level of mKast together with a set of marker genes, which are specifically enriched in the pathway of cellular response to ecdysone (Fig. 3d). Ecdysone is a major regulator of behaviors in bees, and the ecdysone-regulated genes are selectively expressed in the MBs63. The transition of the ecdysone signaling mode in the MBs is involved in the worker age-polyethism, and the mKCs are active in the brains of foragers64. Thus, these suggest that the sensory information processing in the mKC is significant during behavior maturation62.
With an extraordinary ability to recognize colors and navigate routes accurately, the OLs are a vital sensory organ for honeybees to receive information from the outside environment16. Previous studies have demonstrated that the morphological features and neural structures of the visual systems in honeybees exhibit caste-specific adaptations65.The nurse bees spend most of their day inside the colony, while the foragers need to detect flowers and navigate during their foraging flights outside the hive66. Specifically, the OLC4 localizing in the lamina layer exhibits a highly similar gene expression pattern to the monopolar cells in Drosophila. It probably represents the L-fiber in the honeybee OLs, transmitting signals from the retina to the medulla67. The monopolar cells in Drosophila express GABA receptors and accept GABAergic inputs from the centrifugal interneurons68. Correspondingly, the genes related to GABA receptors and channels are upregulated in OLC4. It has been found that GABAergic neuron activity was significantly higher in a part of the OL regions of the forager bees69. Taken together, the region-specific GABA-mediated neuronal networks within the brain may be implicated in processing complex information during foraging tasks in the field.
Behaviors stem from the coordinated activities of cells and the genes operating within the cells of the brain. It has been long recognized that significant changes in brain gene expression profiles are related to the behavioral transition by worker bees from hive work to foraging21. The altered brain profiles are mainly driven by behavior rather than age and can accurately predict the behavioral state of individual workers11. Moreover, the shift in brain gene expression accompanying behavioral maturation is quite reproducible, validated by independent studies28,70. This implies that many cells experience similar state transitions that the aggregate expression profiles of the whole brain still reflect the changes. However, we found that the altered gene expression profiles are highly cell-type specific, while the identified DEGs largely overlapped between the bulk studies and our cellular measurements. Thus, the construction of a higher-resolution brain gene expression map resolves the cellular heterogeneity in transcription and suggests that individual cells switch states in a coordinated manner during the behavioral transition.
The gene expression changes associated with behaviors are coordinated by the GRN, controlled by the active TFs and their interactions with the cis-regulatory regions. We found that the GRNs underlying the transcriptional states of brain cells differed significantly between the two worker castes. In the forager’s brain, the expression level of the TF stripe is upregulated in the MBs. The stripe (EGR1), as an immediate early gene, is activated and transcribed in response to neurological signals, which contribute to the learning and memory system in both vertebrates71,72 and insects73. In honeybees, the stripe is rapidly activated during the orientation flight, which may contribute to spatial learning and navigation behavior74,75. Correspondingly, our spatial transcriptomic map showed that the stripe regulon is more active in the MB calyces of foragers. Furthermore, there is heterogeneity in cell subtypes of the stripe regulon activity, which is specifically active in the forager’s sKCs related to sensory information processing during the foraging flights62. Previous studies have identified stripe as an immediate early gene highly expressed in sKCs76,77. Our findings reveal that stripe functions as a TF together with a group of target genes shows elevated regulon activity in sKCs, highlighting its broader regulatory influence on honeybee behaviors. Conversely, the stripe regulon is less active in the glia. During brain development, stripe is involved in brain epigenetic programming, with its binding sites becoming hypomethylated in mature neurons but remaining methylated in the glia78. Thus, the upregulated stripe in response to orientation flights may lead to the demethylation of its binding sites of the mediated genes shifting to a more active transcriptional state.
The division of labor in honey bee colonies is based on the age, with worker bees performing discrete sets of behaviors throughout their lifespan18. These behavioral states are associated with distinct brain transcriptomic states driven by GRN19. Thus, the brains gene expressions will be affected by both age and natural behavioral maturation49. In our snRNA analysis, the ages for nurse and forager bees were not controlled, we found that many genes showed obvious changes. Specifically, the TF stripe and its target genes are largely affected in the KCs. Considering the potential age effect, we built the artificial colony and controlled for the age of workers. The FISH analysis revealed that stripe and its key target genes, such as Hr38 and caveolin-3, remained highly expressed in specific KCs of foragers (Fig. 7). However, the current study may not fully detangle the effects of age and caste. Previous studies have found that age-related differences in brain gene expression are most apparent in early adult life, typically much earlier than they start nursing and foraging79. In future studies, by comparing the cellular GRN between natural and age-controlled foragers and nurses, it will distinguish the key features for the division of labor.
In this study, our spatial transcriptomic analysis could not delineate the anatomic regions of certain parts of the honeybee brain, such as the VL, AL and OC. This limitation may arise from the bee brain’s intricate spatial organization, where the capture of different cell types can vary depending on the slice position25. Additionally, snRNA-seq captures cells from the entire brain, but the spatial transcriptomic analysis was based on a single-layer sampling. Such difference in cellular sampling can amplify discrepancies in regulon activity between the two datasets, particularly in the OL regions, which consists of the lamina, medulla, and lobula arranged sequentially from distal to proximal along the body axis80. Future studies can construct a 3D expression atlas of the entire brain through serial sectioning, providing more comprehensive insights. On the other hand, while our study focused on gene expression changes within individual cells, dynamic interplay between neuronal networks and GRNs may contribute to the generation of social behavior29. The electrochemical signals, such as hormones, transmitted through the neuronal networks may affect the activity of GRN, and in turn, GRN can also control neural activity by regulating the response elements81. Here, our spatial transcriptomic analysis identified distinct GRN patterns between nurses and foragers across different brain regions. Future integration of both neuronal networks and GRNs will provide insight into the evolution from solitary to social life.
Methods
Honeybee sample collection
The fieldwork took place in August 2020 at the apiary of China Agricultural University, Beijing, China. To control the age of nurse bees, single-cohort colonies were set up82. Briefly, brood frames were collected from a single hive, and adult bees were brushed off. The frames were then incubated in the laboratory at 35 °C and 50% relative humidity. In two days, ~1000 bees emerged from each frame in the incubator. Newly emerged bees were individually marked on their thoraces with a spot of paint (Uni-Paint PX-20). Then, marked bees and the frames were immediately returned to their original hives. Ten-day-old marked bees placing their heads into honeycomb cells containing an egg or larva were collected as nurse bees. Age-matched 21- to 23-day-old bees returning to the colony with loads of pollen on their legs were captured as foragers. There is no current requirement regarding insect care and use in research. Honeybees were cared for daily with adequate food during the experimental period. For tissue collection, bees were collected gently and immediately euthanized by CO2 anesthesia before dissection to reduce any unnecessary duress.
All bees were collected between 9:00 and 11:00 a.m. on the same day (2020/8/20). Honeybee brains were dissected immediately using a dissecting microscope (Canon). Bees were placed on beeswax and fixed using two insect needles through the thorax. The head cuticle was removed, and the whole brain was placed on a glass slide. We then carefully removed the hypopharyngeal glands, salivary glands, three simple eyes, and two compound eyes. Dissected brains were kept frozen at −80 °C until analysis.
Single-nucleus suspension preparation and snRNA-seq with DNBelab C4 system
For the construction of each library, four frozen brain tissues were transferred into a 2-ml Dounce homogenizer (Sigma, D8938) with 2 ml ice-cold 1×Homogenization Buffer containing 0.1 mM Tris-HCl (pH = 7.5, Thermo Fisher, 15567027)), 0.1 mM NaCl (Invitrogen, AM9760G), 0.03 mM MgCl2 (Thermo Fisher, AM9530G), 0.1 mM DTT (Sigma, 43816), 1 × Protease inhibitor cocktail (Roche, 4693116001), 0.4 U/μl RNase inhibitor (BGI, 01E019MM), 0.1% NondietP-40(Roche, 11332473001). After transferring, the homogenizer was put into ice for 20 min, and then tissues were homogenized by 25 strokes of the tight pestle B. Then, the mixture was filtered through a 30-μm strainer (MILTENYI, 130-110-915) into a 15-ml tube and centrifuged at 700 × g for 10 min at 4 °C to pellet the nuclei. The pellet was resuspended in 1 ml of ice-cold PBS containing 1.5% bovine serum albumin (SIGMA, V900933) and 0.2U/μl RNase inhibitor. The suspension was centrifuged at 700 × g for 5 min at 4 °C to pellet the nuclei. Nuclei were resuspended in PBS containing 0.04% bovine serum albumin and 0.2U/μl RNase inhibitor at a final concentration of 1000 nuclei per μl for library preparation.
snRNA-Seq libraries were prepared by the DNBelab C Series Single-Cell Library Prep Set (MGI, 1000021082)83. First, single-nucleus suspensions were converted to barcoded snRNA-seq libraries by droplet encapsulation, emulsion breakage, mRNA-captured bead collection, reverse transcription, cDNA amplification, and purification. Then, indexed sequencing libraries were constructed and quantified by Qubit ssDNA Assay Kit (Thermo Fisher Scientific, Q10212). Then snRNA libraries were sequenced by the DIPSEQ T1 platform at the China National GeneBank (Shenzhen, China) with the following sequencing strategy: 41-bp read length for Read 1 and 100-bp read length for Read 2.
snRNA-seq data processing
Raw sequencing reads from DNBSEQ-T1 were filtered and demultiplexed using PISA (v0.2; https://github.com/shiquan/PISA). Reads were aligned to the genome of A. mellifera (GCF_003254395.2_Amel_HAv3.1_genomic) using STAR (v2.5.1b)84, and the bam files were sorted by sambamba for downstream calculation (v0.7.0)85. The reads were annotated against the GTF file of the A. mellifera genome with ‘PISA anno’. The barcode rank plots were generated, and the ambient RNA noise was reduced based on the cutoff point between empty and nonempty barcodes in SoupX (v1.4.8)86. The nucleus versus gene UMI count matrix was generated using PISA.
Cell quality control and integration
The Seurat (v4.0.5)87 pipeline was executed on each library. Cells from the nurse and forager bees sorted by the nFeature_RNA (number of expressed genes in a cell) and 5th to 95th percentile were retained, with each gene detected in at least three cells. Cells with high mitochondrial content were discarded (the percentage of UMIs classified as mitochondrial genes <0.03). The UMI count matrix was normalized via the ‘NormalizeData’ function, and the mean-variance relationship was controlled to select the most variable genes by ‘FindVariableFeatures’. Principle components (PCs) were calculated using the ‘RunPCA’ function of Seurat.
For each library, we used DoubletFinder (v2.0)88 to remove potentially doublet cells. The expected doublet rate was estimated as 5%, and the pseudo-doublets were generated with pN (the number of artificial doublets) set to 0.25 and pK (the neighborhood size) of 0.01. Cells were identified as doublets based on their rank order in the distribution of the proportion of artificial nearest neighbors (pANN), and cells with the top pANN values (> pK) were predicted as doublets.
Using ‘FindIntegrationAnchors’ and ‘IntegrateData’ functions, all libraries were integrated into a single matrix with the first 30 principal components. The following parameters were used to correct for batch effects of sequencing: reduction = “rpca”, normalization.method = “LogNormalize”. Integrated expression values were then scaled using the ‘ScaleData’ function, and principal components were calculated by ‘RunPCA’ based on the top 2000 variable genes. Graph-based unsupervised clustering was performed by ‘FindNeighbors’ and ‘FindClusters’ with different resolution parameters (0.2, 0.5, 0.8, 1.2, 1.6), followed by ‘clustering trees’ using the R package clustree (v0.4.3)89 to visualize the relationships between multiple resolutions. The embedding plot TSNE (‘RunTSNE’) was used for visualization.
Cell type annotation
We annotated cell type by searching for specific expression patterns for marker genes that have been defined in other insects, including D. melanogaster30,31,32, Harpegnathos saltator33, and Monomorium pharaonic35. The gene sets of A. mellifera (GCF_003254395.2_Amel_HAv3.1), D. melanogaster (GCF_000001215.4_Release_6_plus_ISO1_MT), and H. saltator (GCA_003227715.2_Hsal_v8.6) were retrieved from National Center for Biotechnology Information (NCBI). The reference genome of M. pharaonic was retrieved from Gao et al.90. To define homologous genes between A. mellifera and other species (D. melanogaster, H. saltatory, M. pharaonic), we referred to the orthologs provided by OrthDB (v10.1)91 and also included the protein sequences with two-way reciprocal best hits identified by DIAMOND (v0.8.23) (e-value < 1e−5, percentage of identical matches >30%)92. Different numbers of homologous genes were defined between A. mellifera and D. melanogaster (7955 genes), H. saltator (9464 genes), and M. pharaonic (8545 genes).
According to the established neuronal and glial marker genes, clusters were assigned to different cell types according to aggregated cluster-level expression profiles. Moreover, we also detected specifically overexpressed genes for each cluster by a differential expression analysis of the cells with the ‘FindAllMarkers’ function of Seurat. Significance was defined based on the Wilcoxon rank-sum test (P value < 0.01, log2FC > 0.1). Only genes detected in at least 25% of the cells within the sub-cluster were considered. Then, we performed Gene Ontology enrichment analyses on these overexpressed genes for different clusters using the R package clusterProfiler (v3.18.1)93. The GO terms of genes from A. mellifera were retrieved from Elsik et al.94. The Hypergeometric test (two-sided) was used to test the frequency of terms associated with a different set of genes relative to the overall 12,374 protein-coding genes of A. mellifera as background. The significance was adjusted using the Benjamini–Holm correction.
Transcriptional similarity of cell clusters between species
We assessed the pairwise transcriptional similarity of cell clusters between different species as described by Li et al.35. In brief, MetaNeighbor (v1.10.0)95 was used to calculate the AUROC score for identifying clusters with high similarities between species, including A. mellifera, D. melanogaster, M. pharaonic, and H. saltator. Orthologous genes between species identified as markers of different cell clusters (log2FC > 1.25, FDR < 0.05) were used for the MetaNeighbor analysis. The UMI counts from a pseudo-cell generated by combining ten cells randomly selected from a cell cluster without replacement were summed by genes. The expression level of the marker genes was calculated as CP10K (UMI counts per 10,000) and was standardized by z-transformation. The correlations between the cell clusters from different species were compared based on Pearson’s coefficients.
Differentially expressed gene analysis
We performed DEG analysis across different cell clusters and subclusters. The number of UMI for each gene from all cells in the cluster from each library was summed together and normalized by Trimmed Mean of M-values (TMM) method. Common, trended, and tagwise negative binomial dispersions were calculated by ‘estimateDisp’ function in edgeR (v3.32.1)96. We then performed Quasi-likelihood F-tests using ‘glmQLFit’ and ‘glmQLFTest’ to identify differentially expressed genes between nurse and forager bees. DEGs were defined as genes with an absolute log2FC > 1 and adjusted P value < 0.05 (Benjamini–Hochberg correction). In addition, we compared our DEGs with three DEG sets between nurse and forager bees obtained from the previous bulk analysis, including A-MEXP-36 (529 upregulated genes in nurses; 317 in foragers)11, A-MEXP-755 (467 in nurses; 439 in foragers)49 and CAGEscan (457 in nurses; 416 in foragers)19. Enrichment over gene sets from previous bulk data and DEGs in each cluster from our snRNA data was statistically assessed using the hypergeometric distribution (Fisher’s exact test). For all cases, we used all 12,374 protein-coding genes in Apis mellifera as background.
Stereo-seq library preparation and sequencing
Single fresh brain tissue of nurse or forager bee was collected within 2 min after dissection. The hypopharyngeal glands and salivary glands were removed, while three simple eyes and two compound eyes were preserved to ensure brain tissue integrity. Brain tissues were then frozen in an OCT embedding medium (Sakura Finetek USA, Inc) bathed in chilled isopentane. After embedding, the tissue orientation was marked on the disposable base molds (25 mm × 25 mm × 5 mm). OCT-embedded bee brains were cryosectioned to a thickness of 10 μm using a Leika CM1950 cryostat. Stereo-seq library preparation and sequencing were performed as previously described with minor modification97. In brief, the selected intact brain tissue sections of nurse and forager bees were immediately placed on the Stereo-seq chip and were fixed in 100% methanol at −20 °C for 30 min. The chips were then checked by nucleic acid dye staining (Thermo fisher, Q10212) and light microscopy (Ti-7 Nikon Eclipse Microscope). Meanwhile, tissue sections adjacent to those used for Stereo-seq were stained by hematoxylin and eosin for histological examination. For RNA release, tissue sections on the Stereo-seq chip were permeabilized with 0.1% pepsin (Sigma, P7000) in 0.01 M HCl buffer (pH = 2), incubated at 37 °C for 6 minutes and then washed with 0.1x SSC buffer (Thermo, AM9770) containing 0.05 U/μLRNase inhibitor (NEB, M0314L). Then, mRNAs were in situ captured by the DNBs and reverse transcribed overnight at 42 °C using SuperScript II (Invitrogen, 18064-014, 10 U/μL reverse transcriptase, 1 mM dNTPs, 1 M betaine solution PCR reagent, 7.5 mM MgCl2, 5 mM DTT, 2 U/μLRNase inhibitor, 2.5 μM Stereo-TSO, and 1x First-Strand buffer). Subsequently, the tissues were removed by incubating at 37 °C for 30 min with Tissue Removal buffer (10 mM Tris-HCl, 25 mM EDTA, 100 mM NaCl, 0.5% SDS). The resulting cDNAs were subjected to Exonuclease I (NEB, M0293L) treatment for 3 h at 37 °C. They were finally amplified with PCR mix containing KAPA HiFi Hotstart Ready Mix (Roche, KK2602) and 0.8 μM cDNA-PCR primer. The PCR reactions were incubated at 95 °C for 5 min, followed by 15 cycles of 98 °C for 20 s, 58 °C for 20 s and 72 °C for 3 min, and finally 72 °C for 5 min. A total of 20 ng of the resulting PCR products were subject to the following steps for DNA nanoballs (DNBs) generation: fragmentation (in-house Tn5 transposase), amplification (KAPA HiFi Hotstart Ready Mix), and purification (VAHTS DNA clean beads). In the end, DNBs were loaded into the patterned Nano arrays and sequenced on an MGI DNBSEQ-T10 sequencer (50 bp for read 1, 100 bp for read 2).
Stereo-seq raw data processing
Stereo-seq raw data processing and unsupervised clustering were performed97. In brief, sequences were generated by an MGI DNBSEQ-Tx sequencer with CID and MID appended on each sequencing read. First, CID sequences on Read 1 were mapped to the designed coordinates of the in situ captured chip with 1-base mismatch tolerance, and reads with MID containing either N or >2 bases with a low-quality score (<10) were filtered out. STAR was used to map reads to the reference genome of A. mellifera. Only UMI sequences with a MAPQ > 10 were annotated to their corresponding genes and included in the subsequent calculation. Finally, the CID-containing expression profile matrix was generated by a publicly available pipeline SAW.
Binning and unsupervised clustering of Stereo-seq data
Transcripts captured by 50 × 50 DNBs were merged as one bin 50, the fundamental unit for the downstream analysis. The “SCTransform” function of Seurat was applied to normalize and identify highly variable genes (HVGs). All HVGs were included in the principal-component analysis. Clusters were identified using the ‘FindNeighbors’ and ‘FindClusters’ functions.
Mapping snRNA data onto Stereo-seq data
Tangram (v0.21.1)98 was used for mapping single-nucleus gene expression data onto our spatial gene expression data to project annotation in the snRNA-seq on space. The top 100 differentially expressed genes in each cell type were identified by the ‘sc.tl.rank_genes_groups’ function. The DEGs were then used as training data for the Tangram projection with the ‘cluster’ mode.
SCENIC analysis
We first created the motif databases for the cisTarget and SCENIC analysis on A. mellifera. We identified TFs with Position Weight Matrix (PWM) models in D. melanogaster. We collected 850 TFs with 16,882 motifs in D. melanogaster from the SCENIC database, and ortholog analysis (described above) identified 602 orthologous TFs associated with 9737 PWM models in A. mellifera. A custom cisTarget database was created on A. mellifera using ‘create_cisTarget_databases’ (https://github.com/aertslab/create_cisTarget_databases). We scored the 9738 PWMs on the regulatory regions up to 5 kb upstream and 2 kb downstream of each gene in A. mellifera. The gene-motif rankings were then built, taking the best-scoring region for each motif and gene.
The SCENIC analysis was conducted as described in the workflow99 using pySCENIC (v0.6.6)100. Candidate regulatory modules were inferred from co-expression patterns between TFs and genes by GRNBoost2. Co-expression modules were refined by eliminating indirect targets based on TF motif information using cisTarget to identify putative direct-binding targets. We only retained the motifs that showed NES > 3.0 with their corresponding TF. These processed modules with significant motif enrichment of the correct regulator were retained. We used the AUCell algorithm to assess the activity of each regulon in every single cell. This calculated the enrichment of the regulon as an area under the recovery curve across the ranking of all genes in a particular cell, whereby genes were ranked by their expression value. Based on gene regulatory networks described by the AUC matrix, we ran the cell clustering with a resolution of 0.2 in Seurat. The marker regulons across cell types were defined by the Wilcox test with log2FC > 0.1 and P value < 0.01.
For each regulon, the log2FC between nurse and forager bees was calculated within each subtype at single cell level. To assess significance, differential regulon activity was determined using the t-test. The resulting p-values were then corrected using the Benjamini–Hochberg false discovery rate method101. To investigate regulon activity in space, we utilized AUCell to assess their activity within bin 50 of the spatial transcriptomic sections. Using the same approach, we then examined the differences in regulon activity between nurse and forager in the corresponding brain regions.
Fluorescence in situ hybridization
To verify the expression of genes regulated by the stripe regulon in KCs, the field experiment was performed again in May 2024 at the apiary of China Agricultural University, Beijing, China. Three hives were selected to set up single-cohort colonies to control the age of adult bees. Ten-day-old marked bees placing their heads into honeycomb cells containing an egg or larva were collected as nurse bees. 10-day-old and 22-day-old bees returning to the colony with loads of pollen on their legs were captured as young and old foragers.
We performed SweAMI-FISH (GF002, Servicebio) and high-throughput microscopy scans102. RNA probes targeting the stripe (LOC726302), Hr38 (LOC551232) and caveolin-3 (LOC408310) incorporating the fluorescent dye Cy3 at the 5′ ends were designed. Detailed information of the probe sequences can be found in Supplementary Dataset S11. After fixation in 4% paraformaldehyde in PBS for 24 h, the brains were dehydrated, embedded in paraffin, and sectioned at a thickness of 4 μm. Sections with intact tissue underwent dewaxing and antigen retrieval using citrate buffer (pH 6.0). Proteolytic digestion was performed using proteinase K (20 μg/ml) for 5 min. Tissue samples were washed three times in PBS, each for 5 min. A pre-hybridization solution was applied at 37 °C for 1 h, followed by overnight hybridization in a 40 °C incubator using a probe- containing hybridization solution. Post-hybridization washes were conducted using 2× Saline Sodium Citrate (SSC) at 37 °C for 10 min, 1× SSC at 37 °C twice for 5 min each, and 0.5× SSC at room temperature for 10 min. For branched DNA probe hybridization, slides were gently dried and covered with 60 μl of pre-warmed hybridization solution, followed by incubation in a humidified chamber at 40 °C for 45 min. Post-hybridization washes included 2× SSC, 1× SSC, 0.5× SSC, and 0.1× SSC at 40 °C for 5 min each. Cy3-labeled signal probes were applied at a dilution 1:200 and incubated at 42 °C for 3 h. Further washes were performed with 2× SSC at 37 °C for 10 min, 1× SSC at 37 °C twice for 5 min each, and 0.5× SSC at 37 °C for 10 min. Sections were stained with DAPI in the dark for 8 min, rinsed, and mounted with an anti-fade mounting medium. Hybridized brain tissues were imaged using a Zeiss 810 Laser Scanning Confocal microscope with a 20× objective (Carl Zeiss Microscopy GmbH, Jena, Germany).
Fluorescence intensity was analyzed using ImageJ software (v1.8.0)103. In the fluorescence region, five distinct areas of equal size were randomly chosen. To detect differences of intensity, an analysis of variance (ANOVA) model was fitted using the ‘aov’ function in the ‘stats’ package in R. Multiple comparisons were performed after the one-way ANOVA using the ‘LSD.test’ function from the ‘agricolae’ package (v1.3.5).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data of the snRNA-seq used in this study are available in the CNGB Sequence Archive104 of China National GeneBank DataBase105 under accession code CNP0003087. The raw data of the spatial transcriptomics used in this study are available in the CNGB Sequence Archive of China National GeneBank DataBase under accession code CNP0003197. The reference genomes used in this study were retrieved from NCBI: Apis mellifera GCF_003254395.2 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_003254395.2/], Drosophila melanogaster GCF_000001215.4 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001215.4/], and Harpegnathos saltator GCF_003227715.2 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_003227715.2/]. The reference genome of Monomorium pharaonic was retrieved from the CNGB Sequence Archive CNP0001417. The public datasets used in this study include scRNA-seq data of the adult Drosophila brain from Davie et al.30, available at GEO GSE107451, and from Li et al., 2022, available at ArrayExpress E-MTAB-10519. The scRNA-seq data of the adult Drosophila midbrain from Croset et al.31, have been deposited in NCBI SRP128516. The snRNA-seq data of the whole brain of Monomorium pharaonic are accessible at NCBI PRJNA833256, while the RNA-seq data generated for the Harpegnathos saltator midbrain have been deposited in NCBI GSE149668. The scRNA-seq data of the Apis mellifera brain are available at NCBI GSE142044. The scRNA-seq data of the Drosophila visual system from Kurmangaliyev et al. have been deposited in NCBI GSE156455, while those from Özel et al. are available at NCBI GSE142789. The bulk brain gene expression data of nurse and forager honeybees based on microarray A-MEXP-755 have been deposited at ArrayExpress E-TABM-512, while those based on microarray A-MEXP-36 are available at ArrayExpress E-MEXP-24. Additionally, the CAGEscan sequences for nurse and forager samples have been deposited in NCBI GSE64315. Source data are provided with this paper.
Code availability
The scripts used in this study have been made available on GitHub repository (https://github.com/mumu8910/snRNA_NF.git) with the identifier (https://doi.org/10.5281/zenodo.14909482)106.
References
Chiel, H. J. & Beer, R. D. The brain has a body: adaptive behavior emerges from interactions of nervous system, body and environment. Trends Neurosci. 20, 553–557 (1997).
Kohl, J. et al. Functional circuit architecture underlying parental behaviour. Nature 556, 326–331 (2018).
Chen, P. & Hong, W. Neural circuit mechanisms of social behavior. Neuron 98, 16–30 (2018).
Robinson, G. E., Fernald, R. D. & Clayton, D. F. Genes and social behavior. Science 322, 896–900 (2008).
Robinson, G. E. Genomics. Beyond nature and nurture. Science 304, 397–399 (2004).
Nassel, D. R. Functional roles of neuropeptides in the insect central nervous system. Naturwissenschaften 87, 439–449 (2000).
Nassel, D. R. & Homberg, U. Neuropeptides in interneurons of the insect brain. Cell Tissue Res. 326, 1–24 (2006).
Tibbetts, E. A., Laub, E. C., Mathiron, A. G. E. & Goubault, M. The challenge hypothesis in insects. Horm. Behav. 123, 104533 (2020).
Taborsky, B. & Oliveira, R. F. Social competence: an evolutionary approach. Trends Ecol. Evol. 27, 679–688 (2012).
Blomberg, S. P., Garland, T. Jr. & Ives, A. R. Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57, 717–745 (2003).
Whitfield, C. W., Cziko, A. M. & Robinson, G. E. Gene expression profiles in the brain predict behavior in individual honey bees. Science 302, 296–299 (2003).
Rittschof, C. C. et al. Neuromolecular responses to social challenge: common mechanisms across mouse, stickleback fish, and honey bee. Proc. Natl Acad. Sci. USA 111, 17929–17934 (2014).
Mukherjee, D. et al. Salient experiences are represented by unique transcriptional signatures in the mouse brain. Elife 7, e31220 (2018).
Sokolowski, M. B. Social interactions in “simple” model systems. Neuron 65, 780–794 (2010).
Szathmary, E. & Smith, J. M. The major evolutionary transitions. Nature 374, 227–232 (1995).
Srinivasan, M. V. Honey bees as a model for vision, perception, and cognition. Annu Rev. Entomol. 55, 267–284 (2010).
Ament, S. A., Wang, Y. & Robinson, G. E. Nutritional regulation of division of labor in honey bees: toward a systems biology perspective. Wiley Interdiscip. Rev. Syst. Biol. Med 2, 566–576 (2010).
Robinson, G. E. Regulation of division of labor in insect societies. Annu. Rev. Entomol. 37, 637–665 (1992).
Khamis, A. M. et al. Insights into the transcriptional architecture of behavioral plasticity in the honey bee Apis mellifera. Sci. Rep. 5, 11136 (2015).
Scheiner, R. et al. Learning, gustatory responsiveness and tyramine differences across nurse and forager honeybees. J. Exp. Biol. 220, 1443–1450 (2017).
Zayed, A. & Robinson, G. E. Understanding the relationship between brain gene expression and social behavior: lessons from the honey bee. Annu. Rev. Genet. 46, 591–615 (2012).
Menzel, R. The honeybee as a model for understanding the basis of cognition. Nat. Rev. Neurosci. 13, 758–768 (2012).
Ribi, W. A. & Scheel, M. The second and third optic ganglia of the worker bee: Golgi studies of the neuronal elements in the medulla and lobula. Cell Tissue Res. 221, 17–43 (1981).
Bicker, G. Histochemistry of classical neurotransmitters in antennal lobes and mushroom bodies of the honeybee. Microsc Res. Tech. 45, 174–183 (1999).
Rybak, J. et al. The Digital Bee Brain: Integrating and managing neurons in a common 3D reference system. Front. Syst. Neurosci. 4, 30 (2010).
Zhang, W. et al. Single-cell transcriptomic analysis of honeybee brains identifies vitellogenin as caste differentiation-related factor. iScience 25, 104643 (2022).
Traniello, I. M. et al. Single-cell dissection of aggression in honeybee colonies. Nat. Ecol. Evol. 7, 1232–1244 (2023).
Chandrasekaran, S. et al. Behavior-specific changes in transcriptional modules lead to distinct and predictable neurogenomic states. Proc. Natl Acad. Sci. USA 108, 18020–18025 (2011).
Sinha, S. et al. Behavior-related gene regulatory networks: A new level of organization in the brain. Proc. Natl Acad. Sci. USA 117, 23270–23279 (2020).
Davie, K. et al. A single-cell transcriptome atlas of the aging Drosophila brain. Cell 174, 982–998 e20 (2018).
Croset, V., Treiber, C. D. & Waddell, S. Cellular diversity in the Drosophila midbrain revealed by single-cell transcriptomics. Elife 7, e34550 (2018).
Konstantinides, N. et al. Phenotypic convergence: distinct transcription factors regulate common terminal features. Cell 174, 622–635 e13 (2018).
Sheng, L. et al. Social reprogramming in ants induces longevity-associated glia remodeling. Sci. Adv. 6, eaba9869 (2020).
Li, H. et al. Fly Cell Atlas: A single-nucleus transcriptomic atlas of the adult fruit fly. Science 375, eabk2432 (2022).
Li, Q. et al. A single-cell transcriptomic atlas tracking the neural basis of division of labour in an ant superorganism. Nat. Ecol. Evol. 6, 1191–1204 (2022).
Shah, A. K., Kreibich, C. D., Amdam, G. V. & Munch, D. Metabolic enzymes in glial cells of the honeybee brain and their associations with aging, starvation and food response. PLoS One 13, e0198322 (2018).
Kubo, T. Neuroanatomical dissection of the honey bee brain based on temporal and regional gene expression patterns. In Honeybee Neurobiology and Behavior (eds Galizia, C., Eisenhardt, D. & Giurfa, M.) 341–357 (Springer, Dordrecht, 2012).
Kaneko, K., Suenami, S. & Kubo, T. Gene expression profiles and neural activities of Kenyon cell subtypes in the honeybee brain: identification of novel ‘middle-type’ Kenyon cells. Zool. Lett. 2, 14 (2016).
Traniello, I. M. et al. Meta-analysis of honey bee neurogenomic response links Deformed wing virus type A to precocious behavioral maturation. Sci. Rep. 10, 3101 (2020).
Kurmangaliyev, Y. Z., Yoo, J., Valdes-Aleman, J., Sanfilippo, P. & Zipursky, S. L. Transcriptional programs of circuit assembly in the Drosophila visual system. Neuron 108, 1045–1057 e6 (2020).
Ozel, M. N. et al. Neuronal diversity and convergence in a visual system developmental atlas. Nature 589, 88–95 (2021).
Gengs, C. et al. The target of Drosophila photoreceptor synaptic transmission is a histamine-gated chloride channel encoded by ort (hclA). J. Biol. Chem. 277, 42113–42120 (2002).
Kolodziejczyk, A., Sun, X., Meinertzhagen, I. A. & Nassel, D. R. Glutamate, GABA and acetylcholine signaling components in the lamina of the Drosophila visual system. PLoS One 3, e2110 (2008).
Reiff, D. F., Plett, J., Mank, M., Griesbeck, O. & Borst, A. Visualizing retinotopic half-wave rectified input to the motion detection circuitry of Drosophila. Nat. Neurosci. 13, 973–978 (2010).
Strausfeld, N. J. Brain organization and the origin of insects: an assessment. Proc. Biol. Sci. 276, 1929–1937 (2009).
Ribi, W. A. The first optic ganglion of the bee. III. Regional comparison of the morphology of photoreceptor-cell axons. Cell Tissue Res 200, 345–357 (1979).
Dobritzsch, D., Aumer, D., Fuszard, M., Erler, S. & Buttstedt, A. The rise and fall of major royal jelly proteins during a honeybee (Apis mellifera) workers’ life. Ecol. Evol. 9, 8771–8782 (2019).
Sharma, V. et al. eIF2α controls memory consolidation via excitatory and somatostatin neurons. Nature 586, 412–416 (2020).
Alaux, C. et al. Regulation of brain gene expression in honey bees by brood pheromone. Genes Brain Behav. 8, 309–319 (2009).
Hummon, A. B. et al. From the genome to the proteome: uncovering peptides in the Apis brain. Science 314, 647–649 (2006).
Mandyam, C. D. et al. Neuron-targeted caveolin-1 improves molecular signaling, plasticity, and behavior dependent on the hippocampus in adult and aged Mice. Biol. Psychiatry 81, 101–110 (2017).
Egawa, J. et al. Neuron-targeted caveolin-1 promotes ultrastructural and functional hippocampal synaptic plasticity. Cereb. Cortex 28, 3255–3266 (2018).
Kucharski, R., Maleszka, J. & Maleszka, R. Novel cuticular proteins revealed by the honey bee genome. Insect. Biochem. Mol. Biol. 37, 128–134 (2007).
Jones, B. M. et al. Individual differences in honey bee behavior enabled by plasticity in brain gene regulatory networks. Elife 9, e62850 (2020).
Garcia-Alonso, L., Holland, C. H., Ibrahim, M. M., Turei, D. & Saez-Rodriguez, J. Benchmark and integration of resources for the estimation of human transcription factor activities. Genome Res. 29, 1363–1375 (2019).
Xiong, W. C., Okano, H., Patel, N. H., Blendy, J. A. & Montell, C. repo encodes a glial-specific homeo domain protein required in the Drosophila nervous system. Genes Dev. 8, 981–994 (1994).
Wasserman, W. W. & Sandelin, A. Applied bioinformatics for the identification of regulatory elements. Nat. Rev. Genet. 5, 276–287 (2004).
Suenami, S., Oya, S., Kohno, H. & Kubo, T. Kenyon cell subtypes/populations in the honeybee mushroom bodies: possible function based on their gene expression profiles, differentiation, possible evolution, and application of genome editing. Front. Psychol. 9, 1717 (2018).
Oya, S., Kohno, H., Kainoh, Y., Ono, M. & Kubo, T. Increased complexity of mushroom body Kenyon cell subtypes in the brain is associated with behavioral evolution in hymenopteran insects. Sci. Rep. 7, 13785 (2017).
Fahrbach, S. E. Structure of the mushroom bodies of the insect brain. Annu. Rev. Entomol. 51, 209–232 (2006).
Kuwabara, T., Kohno, H., Hatakeyama, M. & Kubo, T. Evolutionary dynamics of mushroom body Kenyon cell types in hymenopteran brains from multifunctional type to functionally specialized types. Sci. Adv. 9, eadd4201 (2023).
Kaneko, K. et al. Novel middle-type Kenyon cells in the honeybee brain revealed by area-preferential gene expression analysis. PLoS One 8, e71732 (2013).
Yamazaki, Y., Kiuchi, M., Takeuchi, H. & Kubo, T. Ecdysteroid biosynthesis in workers of the European honeybee Apis mellifera L. Insect. Biochem. Mol. Biol. 41, 283–293 (2011).
Yamazaki, Y. et al. Differential expression of HR38 in the mushroom bodies of the honeybee brain depends on the caste and division of labor. FEBS Lett. 580, 2667–2670 (2006).
Streinzer, M., Brockmann, A., Nagaraja, N. & Spaethe, J. Sex and caste-specific variation in compound eye morphology of five honeybee species. PLoS One 8, e57702 (2013).
Withers, G. S., Fahrbach, S. E. & Robinson, G. E. Selective neuroanatomical plasticity and division of labour in the honeybee. Nature 364, 238–240 (1993).
Dyer, A. G., Paulk, A. C. & Reser, D. H. Colour processing in complex environments: insights from the visual system of bees. Proc. Biol. Sci. 278, 952–959 (2011).
Davis, F. P. et al. A genetic, genomic, and computational resource for exploring neural circuit function. Elife 9, e50901 (2020).
Kiya, T. & Kubo, T. Analysis of GABAergic and non-GABAergic neuron activity in the optic lobes of the forager and re-orienting worker honeybee (Apis mellifera L.). PLoS One 5, e8833 (2010).
Zayed, A., Naeger, N. L., Rodriguez-Zas, S. L. & Robinson, G. E. Common and novel transcriptional routes to behavioral maturation in worker and male honey bees. Genes Brain Behav. 11, 253–261 (2012).
Veyrac, A., Besnard, A., Caboche, J., Davis, S. & Laroche, S. The transcription factor Zif268/Egr1, brain plasticity, and memory. Prog. Mol. Biol. Transl. Sci. 122, 89–129 (2014).
Brito, V. et al. Hippocampal Egr1-dependent neuronal ensembles negatively regulate motor learning. J. Neurosci. 42, 5346–5360 (2022).
Liu, T., Wang, L. & Li, Q. Drosophila ortholog of mammalian immediate-early gene Npas4 is specifically responsive to reversal learning. Neurosci. Bull. 37, 99–102 (2021).
Lutz, C. C. & Robinson, G. E. Activity-dependent gene expression in honey bee mushroom bodies in response to orientation flight. J. Exp. Biol. 216, 2031–2038 (2013).
Geng, H. et al. Visual learning in a virtual reality environment upregulates immediate early gene expression in the mushroom bodies of honey bees. Commun. Biol. 5, 130 (2022).
Shah, A., Jain, R. & Brockmann, A. Egr-1: a candidate transcription factor involved in molecular processes underlying time-memory. Front. Psychol. 9, 865 (2018).
Ugajin, A., Kunieda, T. & Kubo, T. Identification and characterization of an Egr ortholog as a neural immediate early gene in the European honeybee (Apis mellifera L). FEBS Lett. 587, 3224–3230 (2013).
Sun, Z. et al. EGR1 recruits TET1 to shape the brain methylome during development and upon neuronal activity. Nat. Commun. 10, 3892 (2019).
Whitfield, C. W. et al. Genomic dissection of behavioral maturation in the honey bee. Proc. Natl Acad. Sci. USA 103, 16068–16075 (2006).
Strausfeld, N. J. The lobula plate is exclusive to insects. Arthropod Struct. Dev. 61, 101031 (2021).
Hobert, O., Carrera, I. & Stefanakis, N. The molecular and gene regulatory signature of a neuron. Trends Neurosci. 33, 435–445 (2010).
Huang, Z. Y. & Robinson, G. E. Honeybee colony integration: worker-worker interactions mediate hormonally regulated plasticity in division of labor. Proc. Natl Acad. Sci. USA 89, 11726–11729 (1992).
Liu, C. Y. et al. A portable and cost-effective microfluidic system for massively parallel single-cell transcriptome profiling. bioRxiv, https://www.biorxiv.org/content/10.1101/818450v1 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. GigaScience 9, giaa151 (2020).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e21 (2019).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 e4 (2019).
Zappia, L. & Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. GigaScience 7, giy083 (2018).
Gao, Q. et al. High-quality chromosome-level genome assembly and full-length transcriptome analysis of the pharaoh ant Monomorium pharaonis. GigaScience 9, giaa143 (2020).
Kriventseva, E. V. et al. OrthoDB v10: sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Res. 47, D807–D811 (2019).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Elsik, C. G. et al. Hymenoptera Genome Database: integrating genome annotations in HymenopteraMine. Nucleic Acids Res. 44, D793–D800 (2016).
Crow, M., Paul, A., Ballouz, S., Huang, Z. J. & Gillis, J. Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor. Nat. Commun. 9, 884 (2018).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 e21 (2022).
Biancalani, T. et al. Deep learning and alignment of spatially resolved single-cell transcriptomes with Tangram. Nat. Methods 18, 1352–1362 (2021).
Van de Sande, B. et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 15, 2247–2276 (2020).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Benjamini, Y., Drai, D., Elmer, G., Kafkafi, N. & Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res. 125, 279–284 (2001).
Jia, Y. et al. Long non-coding RNA NORAD/miR-224-3p/MTDH axis contributes to CDDP resistance of esophageal squamous cell carcinoma by promoting nuclear accumulation of β-catenin. Mol. Cancer 20, 162 (2021).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Guo, X. et al. CNSA: a data repository for archiving omics data. Database 2020, baaa055 (2020).
Chen, F. Z. et al. CNGBdb: China National GeneBank database. Yi Chuan 42, 799–809 (2020).
Mu, X. H. et al. Single-nucleus and spatial transcriptome identify brain landscape of gene regulatory networks associated with behavioral maturation in honeybees. Github:snRNA_NF. https://doi.org/10.5281/zenodo.14909482 (2025).
Acknowledgements
The work was funded by the National Natural Science Foundation of China Project 32170495, the National Science Fund for Distinguished Young Scholars of China (grant 32425012) to Hao. Z., and the National Natural Science Foundation of China Project 32300385 to Z.Z.
Author information
Authors and Affiliations
Contributions
Hao Z. supervised the study. Hao Z., S.L., and X.X. designed the study; Z.Z., X.M., Y.L., H.C., L.Z., Yifan Z., and X.H. collected samples; X.M., Z.Z., J.M., Y.Q., Q.G., P.Z., Y.W., Haorong L., He Z., Qiye L., H.Y., and J.W. performed the single-nucleus RNA sequencing and data analysis; Z.Z., X.M., Qun L., Yinging Z., N.Z., D.L., R.Z., Q.J., A.J., S.P., Xiawei L., Xuemei L., J.S., L.M., and J.L. conducted the collection and analysis of spatial transcriptomics data; Z.Z., X.M., and Haoyu L. performed fluorescence in situ hybridization; Hao. Z., X.M., and Z.Z. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mu, X., Zhang, Z., Liu, Q. et al. Single-nucleus and spatial transcriptomics identify brain landscape of gene regulatory networks associated with behavioral maturation in honeybees. Nat Commun 16, 3343 (2025). https://doi.org/10.1038/s41467-025-58614-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58614-8