Abstract
Amyotrophic lateral sclerosis (ALS) is a swiftly progressive and fatal neurodegenerative ailment marked by the degenerative motor neurons (MNs). Why MNs are specifically susceptible in predominantly sporadic cases remains enigmatic. Here, we demonstrated N6-methyladenosine (m6A), an RNA modification catalyzed by the METTL3/METTL14 methyltransferase complex, as a pivotal contributor to ALS pathogenesis. By conditional knockout Mettl14 in murine MNs, we recapitulate almost the full spectrum of ALS disease characteristics. Mechanistically, pervasive m6A hypomethylation triggers dysregulated expression of high-risk genes associated with ALS and an unforeseen reduction of chromatin accessibility in MNs. Additionally, we observed diminished m6A levels in induced pluripotent stem cell derived MNs (iPSC~MNs) from familial and sporadic ALS patients. Restoring m6A equilibrium via a small molecule or gene therapy significantly preserves MNs from degeneration and mitigates motor impairments in ALS iPSC~MNs and murine models. Our study presents a substantial stride towards identifying pioneering efficacious ALS therapies via RNA modifications.
Similar content being viewed by others
Introduction
In recent years, the intricate world of RNA modifications, often referred to as the “epitranscriptome,” has emerged as a pivotal regulatory axis governing developmental processes1. Over 170 types of RNA modifications have been identified since the 1950s. High-throughput sequencing has revealed a diversity of messenger RNA (mRNA) modifications in various organisms2,3. Among these, m6A has received the most attention as it is the most abundant form of mRNA modification in mammals4. Various studies have shown that m6A levels can be dynamic and reversible. m6A is deposited by the m6A methyltransferase complex (termed the “writer”), with METTL3 (methyltransferase-like 3) acting as the catalytically active methyltransferase and METTL14 playing an essential structural role in facilitating catalysis. The larger methyltransferase holo complex contains accessory units, including WTAP (Wilms tumor 1-associated protein), VIRMA (vir-like m6A methyltransferase associated), RBM15 (RNA-binding motif protein 15) and its paralog (RBM15B), ZC3H13, and HAKAI5,6,7,8,9,10,11. Conversely, m6A marks are subject to removal by m6A demethylases, aptly termed “erasers”. Notable players in this dynamic process include FTO (fat mass and obesity-associated protein) and ALKBH5 (alkB homolog 5)12,13,14,15. This intricate interplay between writers and erasers crafts a finely tuned regulatory mechanism, orchestrating the reversible m6A modifications that play critical roles in shaping RNA function and developmental processes.
The m6A RNA modification acts on multiple molecular pathways, including in splicing, stability, nuclear export, localization, translational efficiency, and activation and decay of targeted mRNAs5,7,16,17. Recent studies have shown that constitutive knockout of Mettl14—a key facilitator of the m6A methyltransferase complex—is embryonically lethal in mice, whereas conditional knockout (cKO) of Mettl14 in neural progenitor cells disrupts cortical development and leads to premature death in mice18,19. Remarkably, levels of m6A are relatively low in mouse brain tissue during embryogenesis, but drastically increase by adulthood20, suggesting that m6A RNA modification plays a unique role in the adult central nervous system. That latter finding has also raised the possibility that m6A might play an important role in adult RNA homeostasis, with imbalances potentially leading to the onset or progression of neurodegeneration. This hypothesis is supported by studies demonstrating a positive correlation between m6A modification and gene expression homeostasis across tissues, as well as tissue-type-specific aging-associated m6A dynamics in primates21. In humans, functional impairment of m6A has also been shown to play a pivotal role in cancer22,23, cell fate transition and determination24,25, and disease26,27,28. Although a concordant decrease in m6A RNA methylation of brain tissue (specifically, the cingulate gyrus) from an Alzheimer’s disease (AD) mouse model and in human patient brain tissues has been reported29, whether m6A exerts a direct causative role in human neurodegeneration remains obscure.
To date, research efforts have focused almost exclusively on DNA sequencing (usually whole exome) to identify the genetic causes of neurodegenerative diseases. This is arguably the primary reason why the genetic and molecular bases for many neurodegenerative diseases remain unknown, as most neurodegenerative diseases are sporadic30,31. Accumulating evidence indicates that most aging-associated diseases, including amyotrophic lateral sclerosis (ALS), are linked to RNA metabolism, perhaps explaining why probing gene mutations by DNA sequencing fails to identify more ALS-causative genes32,33. Though dysregulated RNA processing has been identified in the majority of ALS patients, it remains unclear which aspects of RNA metabolism are critical and if they are directly causative of spinal motor neuron (MN) degeneration31,34. There are two major shortcomings of ALS-associated research efforts to date: (1) next-generation sequencing technologies are DNA-based and cannot directly sequence RNA or RNA isoforms with long reads and modifications, hindering analyses of RNA modifications from patient transcriptomes; and (2) only a small number of mouse models mimic to varying degrees the MN pathology of sporadic ALS (sALS), with most of them presenting relatively minor phenotypes when compared to familial ALS (fALS) models30. Accordingly, there is still no robust sALS animal model (>90% of ALS patients are sporadic) that fully recapitulates MN degeneration pathology. Although employing a gene mutation identified in ALS patients to generate an ALS murine model remains a robust methodology, it frequently only elicits some aspects of ALS pathology, occasionally resulting in excessively shortened lifespan30. Therefore, to advance research in this field necessitates: (1) discovering common disease-causing mechanisms present in both familial and sporadic ALS patients; and (2) establishing an ALS animal model based on these mechanisms. Such efforts would aim to replicate the primary hallmarks of familial and sporadic ALS at molecular (e.g., TARDBP/TDP43 aggregation), cellular (e.g., motor neuron degeneration), and physiological levels (manifesting muscle weakness and shortened lifespan).
Recently, two studies emphasized the roles of m6A in ALS. Barmada’s group reported that m6A hypermethylation modulates RNA binding by TDP43 and the disease pathogenesis of ALS and frontotemporal dementia (FTD)35, whereas Sun’s group indicated that globally reduced m6A levels in C9ORF72-associated ALS and FTD dysregulate RNA metabolism and contribute to neurodegeneration36. Although these two studies emphasize the importance of m6A homeostasis in ALS, their seemingly contradictory results necessitate further clarification. As both studies were performed primarily on cell models and postmortem spinal cord sections from patients, it is imperative to explore if manipulating m6A levels in animal models in vivo recapitulates ALS pathology.
Here, we used two different sets of MN Cre drivers to remove Mettl14, corroborating that m6A hypomethylation (hypo-m6A) elicits an ALS-like phenotype in vivo. Impairment of the m6A repertoire elicits dysregulation of many known ALS-related pathways, with a concomitant change in the chromatin landscape of spinal MNs. Additionally, iPSC~MNs from several familial and sporadic ALS patients exhibited hypo-m6A, and restoration of m6A homeostasis by means of a small molecule largely spared the MNs from degeneration in both familial and sporadic ALS contexts. Most importantly, intrathecal delivery of Fto-shRNA to knock down Fto, an m6A eraser enzyme, ameliorated the motor deficits of SOD1G93A mice (an ALS mouse model) and extended their lifespan. Accordingly, we speculate that m6A hypomethylation contributes to ALS and restoring the m6A reservoir can mitigate the symptoms of familial and sporadic ALS.
Results
m6A levels are reduced in human ALS iPSC~MNs and hypo-m6A leads to MN degeneration
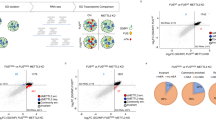
Given existing contradictory results35,36, we examined the extensive transcriptomic dataset derived from Answer ALS to establish if m6A hypermethylation (hyper-m6A) or hypo-m6A is associated with ALS. Our analysis focused on assessing expression levels of the methyltransferases METTL3/5/14/16, in which METTL3/14 are the ‘writer’ complex responsible for m6A for mRNAs, while METTL16 is largely for ncRNA and METTL5 is known for rRNA m6A modifications3 (Supplementary Fig. 1). Interestingly, only METTL3/14 exhibited a trend of down-regulation in numerous familial or sporadic human ALS iPSC~MNs (Supplementary Fig. 1a, b). In agreement with this outcome, we observed reduced METTL3 and METTL14 expression in the majority of postmortem cortex samples from ALS patients (Supplementary Fig. 1c), as well as reduced protein levels as examined by Li et al.36. To confirm this result, we assessed another independent study that conducted transcriptome analysis on a different set of human ALS iPSC~MNs and observed that most of those iPSC~MNs also exhibited consistent down-regulation of METTL3 and METTL14 (Supplementary Fig. 1d). Together, these findings indicate that compromised m6A pathways might be an overlooked aspect of ALS. To scrutinize if compromised m6A writer expression leads to hypo-m6A in a human context, we differentiated three familial ALS iPSC lines (SOD1+/L144F, C9ORF72exp~800 G4C2, and TDP43G298S), together with their corresponding isogenic rescue controls (Ctrl #1 and Ctrl #2) and healthy control (Ctrl #3), into spinal MNs, and then profiled their m6A dynamics along the differentiation process under stress-induced conditions (illustrated in Fig. 1a, results in Fig. 1b–g)37,38. To accelerate ALS disease progression, we applied cyclopiazonic acid (CPA), an endoplasmic reticulum stressor, as CPA has been shown previously to act as a selective stressor to accelerate the degeneration of human SOD1G93A iPSC~MNs but not wild-type controls37,38. Consistent with this scenario, we found that all of our Ctrl iPSC~MNs were relatively resistant to CPA stress, unlike the ALS iPSC~MNs that exhibited drastic loss after seven days of CPA treatment (Fig. 1b–d). No obvious degeneration was displayed by either Ctrl or ALS iPSC~MNs on day 4 (Fig. 1b–d). Thus, we could capture the progressive MN degeneration displayed by the familial ALS iPSC lines. Notably, even before the drastic MN loss following stress treatment on day 4, we consistently detected reduced m6A levels in the ALS iPSC~MNs (Fig. 1b–d). This trend was sustained at day 7 with a concomitant decrease in METTL3/14 expression, together with significantly increased expression of the demethylases FTO and ALKBH5 in most of the ALS iPSC~MNs (Supplementary Fig. 1e). In agreement with these data, we found that all three ALS iPSC lines displayed hypo-m6A before the MN loss caused by stressor treatment (Fig. 1e–g) revealed by m6A ELISA and dot blot (see methods for details).
a Timeline of MN differentiation in control (Ctrl) and ALS (SOD1+/L144F, C9ORF72exp~800 G4C2, TDP43G298S) iPSC lines. CPA (cyclopiazonic acid) was applied to accelerate MN degeneration (annotated as basal time point day 0). NF neurotrophic factors. b–d MN degeneration index and m6A RNA methylation levels of Ctrl and ALS iPSC~MNs. m6A percentages of Ctrl/ALS types at indicated time points were normalized to those at day 0. ALS iPSC~MNs undergo dramatic degeneration from day 7 to day 21 post-CPA treatment, while a significant reduction in m6A level was already exhibited at day 4. The degeneration index measures the neurite fragmentation. e–g Quantification of the m6A ratio in mRNAs from day 4 post-CPA treatment of Ctrl and ALS (SOD1+/L144F, C9ORF72exp~800 G4C2, TDP43G298S) iPSC lines. The mRNAs were extracted by poly(A) purification. The panels at the right show representative images of an m6A dot blot and methylene blue staining (for loading controls) (b–g: n = 3 independent experiments). h Timeline of STM2457 (20 μM)-mediated inhibition of m6A modification during MN differentiation of wild-type iPSC lines. STM2457 was applied for 6 days to accelerate MN degeneration. i and j Inhibition of METTL3-mediated m6A modification in human wild-type iPSC~MNs results in a sharp decline in m6A levels after four days of treatment for STM2457, as assayed by m6A ELISA in mRNA (i) and mRNA m6A dot blot (j) (i, j: n = 3 independent experiments). m6A methylation was normalized to the vehicle (DMSO), which served as a non-stressed control. Note that dramatic neurite degeneration was observed on day 6 in k upon STM2457 treatment. Scale bar, 200 µm. l Quantification of the degeneration index at an indicated time point normalized to the vehicle control (n = 6 independent experiments). Illustrations in a and h created in BioRender. Chen, J. (2025) https://BioRender.com/3nizfu7. All Data are presented as mean ± SD, significant P values from two-tailed t-tests. N.S. non-significant. Source data are provided as a Source data file.
To confirm if hypo-m6A leads to MN degeneration, we adopted two approaches. First, we used a specific METTL3 inhibitor (METTL3i), STM245722, to impair m6A production during human MN differentiation (Fig. 1h), which revealed that METTL3i reduces the m6A-mRNA repertoire assayed by m6A ELISA and dot blot (Fig. 1i, j), with concomitantly drastic neurite degeneration and a reduced MN population (Fig. 1k, l). Secondly, we transfected the HEK293T with METTL3 or METTL14-shRNA (Supplementary Fig. 2a) and revealed a reduction in m6A-mRNA levels (Supplementary Fig. 2b). Furthermore, we infected the Lentivirus (LV)-shMETTL3 and shMETTL14 observed a concomitant neurite degeneration (Supplementary Fig. 2c–e). Thus, our results together with the large set of available ALS patient data support the notion that METTL3/METTL14 might be a critical regulatory complex linked to both familial and sporadic ALS disease, and that MNs appear to be more sensitive to METTL3/METTL14 down-regulation. Most notably, global down-regulation of the m6A methylation repertoire elicited obvious MN degeneration.
Impairment of the m6A production enzyme Mettl14 in spinal MNs elicits MN degeneration
To determine if hypo-m6A promotes MN degeneration in vivo, we conditionally deleted Mettl14 specifically in MNs either at the developmental stage by using Olig2-Cre39 or at the postmitotic and postnatal stage by adopting ChAT-Cre (see Methods for details)18. First, we verified that both conditional mouse lines displayed a significant reduction in the population of Mettl14on cells in the ventral horn of the spinal cord (Supplementary Fig. 3a, b). Although Olig2-Cre; Mettl14floxed mice mostly exhibited early postnatal lethality (~P24 to P28, Supplementary Fig. 3c) with a shivering phenotype (Supplementary Movie 1), all MN subtypes appeared normal based on immunostaining (Supplementary Fig. 3d–g). As Olig2 is known to be expressed at a later stage in oligodendrocytes, we consider that the shivering phenotype might be a reflection of compromised oligodendrocyte precursors (Olig2on and Sox9on double-positive cells) (Supplementary Fig. 3h, i), consistent with a previous study40. Conversely, the ChAT-Cre; Mettl14floxed mice displayed normal MN development and an ordinary appearance at the postnatal and juvenile stages (Fig. 2a). Nevertheless, two months later, we observed a gradual decline in body weight from P70 (Fig. 2a) and a kyphosis phenotype from P100 for all of the ChAT-Cre; Mettl14floxed mice (Supplementary Fig. 3j). Strikingly, all of the ChAT-Cre; Mettl14floxed mice (both male and female, n > 90) exhibited premature death at P160~P300 (Fig. 2b).
a Body weight of male and female ChAT-Cre; Mettl14floxed and littermate control mice. Data are presented as mean ± SD. b Kaplan–Meier survival curves reflect that both male and female ChAT-Cre; Mettl14floxed mice die prematurely compared to littermate controls. Immunostaining (c) and quantification (d) of lumbar ChATon MN numbers reveal a gradual reduction starting after P100 and a significant loss of ChAT-Cre; Mettl14floxed MNs at P160 (Data are presented as mean ± SD, with significant P values from two-tailed t-tests. NS non-significant. n = 3 in ChAT-Cre; Mettl14floxed and control mice at P30, P70, P160, and P250, respectively; n = 5 in ChAT-Cre; Mettl14floxed mice and n = 4 in control mice at P100, Scale bar, 100 µm). Preferential loss of the cholinergic C-bouton nerve terminals of MNs in the ChAT-Cre; Mettl14floxed mice from P70 (e), with respective quantification in (f). Scale bar, 50 µm. Data are presented as mean ± SD, n = 3 mice in ChAT-Cre; Mettl14floxed and control at P30, P100, P160, and P250, respectively; n = 5 in ChAT-Cre; Mettl14floxed mice and n = 3 in control mice at P70, with significant P values from two-tailed t-tests. N.S. non-significant. Source data are provided as a Source data file.
Apart from their kyphotic appearance and movement defects, we further investigated a series of molecular and cellular ALS disease features in the ChAT-Cre; Mettl14floxed mice. At the molecular level, we observed (1) that the numbers of ChATon MNs in the lumbar region of spinal cords were comparable before P70 but gradually declined after P100 (Fig. 2c, d). However, C-boutons, a source of cholinergic input to MNs, already showed a prominent decrease from P70 (Fig. 2e, f); (2) prominent neuroinflammation upon microglia (Iba1on) activation in the spinal cords of the ChAT-Cre; Mettl14floxed mice relative to controls at P160 (Fig. 3a, b), but not before P120 (Supplementary Fig. 4a, b); and (3) significant cytoplasmic aggregation of Tdp43 in the ChAT-Cre; Mettl14floxed mice, whereas control littermates mainly presented nuclear localizations for that protein after P120 (Fig. 3c, d, and Supplementary Fig. 4c, d). Notably, another RNA-binding protein, Fus, which is often shown as mislocalization in ALS patients, also exhibited cytoplasmic aggregation in the ChAT-Cre; Mettl14floxed mice (Supplementary Fig. 4e, f). Furthermore, the ChAT-Cre; Mettl14floxed mice exhibited drastically reduced endplate area and muscle denervation (Fig. 3e–h). Overall, our findings indicate that the ChAT-Cre; Mettl14floxed mouse model demonstrates progressive MN degeneration, mirroring several key molecular pathological features observed in human ALS patients.
Images illustrate microglial activation, as determined by immunostaining for Iba1 (a), and quantification (b) of lumbar Iba1on numbers at the ventral region, revealing significant microglial activation in ChAT-Cre; Mettl14floxed mice compared to littermate controls. Staining was repeated on n = 5 mice. c Tdp43 (green) is localized in the nucleus of the MNs of control mice. In the ChAT-Cre; Mettl14floxed mutant mice, numerous Tdp43 inclusions exist in the cytoplasm (arrow). High magnifications of the highlighted Tdp43 aggregates in MNs are shown in the rightmost panels. Respective quantification is presented in d, (n = 3 mice). e, g Representative z-stack confocal images of neuromuscular junctions (NMJs) in gastrocnemius (GA) muscles dissected from P150 ChAT-Cre; Mettl14floxed and littermate control mice. Motor nerves were visualized using a combination of SV2/NF(2H3) (green) and post-synaptic AChRs with α-BTX (magenta). Arrowheads identify denervated synapses, abnormal axonal swellings (e), and smaller endplates (g). f, h Quantification of the denervation ratio of NMJs and endplate area from (e, g). (n = 3 mice, quantification for all NMJs from all views of captured images). Scale bar, 100 µm. Data are presented as mean ± SD with significant P values from two-tailed t-tests. Source data are provided as a Source data file.
Mice with m6A hypomethylation recapitulate ALS-associated behavioral phenotypes
Although some previous ALS mouse models exhibit molecular hallmarks of ALS pathology, their MN physiology or gross behaviors appeared relatively normal30,34. To examine if the ChAT-Cre; Mettl14floxed mice represent an improved potential ALS mouse model, we conducted a series of behavioral assays on the ChAT-Cre; Mettl14floxed mice to further characterize the phenotypes observed from P40 to P210. First, the results of a rotarod test and forelimb grip strength assay corroborated that motor ability gradually declined, together with concomitant muscle weakness, recapitulating two major pathological manifestations observed for ALS patients (Fig. 4a–d, Supplementary Movie 2). Next, through an open field test, we noted that general activity levels of the ChAT-Cre; Mettl14floxed mice gradually became compromised and their exploratory behavior in a novel environment was reduced, reflecting a motor deficit and a frontotemporal dementia (FTD)-like phenotype (Fig. 4e, f, and Supplementary Fig. 5). Finally, we performed a kinematic analysis, which revealed that whereas the spinal interneuron (IN) circuit remained largely intact, motor outputs were interrupted in the ChAT-Cre; Mettl14floxed mice (Fig. 4g-I, Supplementary Fig. 5a, and Supplementary Movie 3). This detailed scrutiny of the ChAT-Cre; Mettl14floxed mice at the molecular, cellular, physiological, and behavioral levels indicates that the m6A reservoir is a critical factor in maintaining adult MN function, and that compromising m6A levels prompts a MN degeneration process that recapitulates ALS disease progression.
a Schematic illustration of the behavioral tests conducted to assess motor functions. Created in BioRender. Chen, J. (2025) https://BioRender.com/mrtnwlk. Locomotor coordination on an accelerating rotarod is displayed as the rotation speed at which mice fell off (b) and the latency to fall (c). (Ctrl: n = 3/3, 8/9, 5/10, 11/10, and 10/10 mice; ChAT-Cre; Mettl14floxed: n = 3/3, 7/11, 7/15, 9/12, and 8/10 mice at P40, P70, P100, P130, and P160 from males/females, respectively). There was a significant decrease in locomotor activity at P70 and thereafter for males and at P100 and thereafter for females. d Forelimb grip strengths for ChAT-Cre; Mettl14floxed male and female mutant mice. (Ctrl: n = 3/6, 12/12, 12/14, 12/10, and 6/10 mice and ChAT-Cre; Mettl14floxed: n = 3/6, 10/14, 10/17, 8/12, and 7/9 mice at P40, P70, P100, P130, and P160 from males/females, respectively). e Travel pathways (red) of representative trajectory diagrams filmed for 10 and 60 minutes in the open field test arena (square perimeter) for the early-onset (P70) and disease progression (P160) stages of ChAT-Cre; Mettl14floxed and littermate control mice. f Total distance traveled in the open field test. (Ctrl: n = 3/3, 7/9, 6/8, 8/9, and 8/9 mice; ChAT-Cre; Mettl14floxed: n = 3/3, 6/11, 6/11, 7/12, and 7/9 mice at P40, P70, P100, P130, and P160 from males/females, respectively). g Schematic illustration of the behavioral tests from the treadmill conducted to assess motor function. Created in BioRender. Chen, J. (2025) https://BioRender.com/mrtnwlk. h, i Stride width (usually mediated by INs) is not compromised, whereas stride length (mediated by MNs) is drastically reduced in the ChAT-Cre; Mettl14floxed mice (Speed = 15 cm/s). n = 6 mice (3 from P180 and 3 from > P210 mice). Data are presented as mean ± SD, significant P values from two-tailed t-tests. N.S. non-significant. Source data are provided as a Source data file.
Uncovering the dysregulated m6A-modified genes leading to MN degeneration
To gain insights into the potential mechanisms underlying how hypo-m6A promotes MN degeneration, we aimed to systematically identify the dysregulated genes possessing m6A modifications in our ChAT-Cre; Mettl14floxed mice (Figs. 5, 6). To identify m6A-modified transcripts, we adopted a direct RNA sequencing platform, which enables the identification of the MN m6A epitranscriptome at single-nucleotide resolution (Fig. 5). Since it is technically challenging to obtain sufficient adult MNs from the mouse spinal cord for direct RNA sequencing, we employed an enhanced method using mouse ESC-derived MNs (mESC~MNs), matured with a conditioned medium (Fig. 5a–c)38,41. First, we confirmed that this approach successfully generated MNs expressing mature neuronal markers, with longer and more mature neurite structures revealed by discrete Syn1-positive puncta (Fig. 5b, c). We then subjected these mature MNs to the Oxford Nanopore Technologies (ONT) platform, which provides a powerful framework for detecting RNA modifications through advanced machine-learning algorithms applied to sequencing metrics (Fig. 5d, details in Methods). As expected, several adult mature MN genes are abundantly expressed in the Nanopore direct RNA-seq results (Fig. 5e). We subsequently employed two supervised machine learning tools, namely EpiNano and m6Anet42,43, which set a stringent criterion that the predicted m6A sites need to occur in at least two samples for either one of the algorithms (see Methods for details). We identified 30,340 high-confidence m6A modification sites (probability > 0.5) corresponding to 7,921 genes (Fig. 5d, genes of interest are illustrated in Supplementary Data 1; refer to the Methods for details). Interestingly, a deeper estimate of the predicted m6A stoichiometry indicated that the high variation levels are displayed across different m6A sites in the same gene (Supplementary Data 3). Consistent with previous findings29, our analysis also revealed enriched distributions of m6A sites in coding sequences (CDSs) and 3′ UTRs, especially near the stop codons (Fig. 5f and Supplementary Fig. 6a). By analyzing the enrichment of Gene Ontology (GO) and KEGG pathways for the m6A-modified transcripts, we noticed a striking enrichment for ALS-related genes (Fig. 5g). In addition, although only 35.56% of transcripts (7921 out of 22,280) contained m6A in our dataset, among the 81 identified ALS risk genes to date, our Nanopore direct RNA-seq demonstrated that 41.98% of ALS risk genes (34 out of 81) are m6A-modified (P = 2.06e-6, one-tailed hypergeometric test; Fig. 5h and Supplementary Data 2).
a Timeline of in vitro differentiation and maturation from mESC~MNs. Created in BioRender. Chen, J. (2025) https://BioRender.com/xap28jf. b The heatmap from qPCR verification shows that postnatal upregulation of functionally relevant genes. Slc5a7 serves as a positive control as it is constantly expressed after the postmitotic stage. c Upper panel: Immunostaining of Syn1 in Day 7 and Day 12 mESC~MNs. Smi32 labels the MNs and neurites. Scale bar, 100 µm. n = 3 independent experiments. All quantification information is provided in the Methods and Source data file. Lower panel: Quantifications reveal a significant increase in neurite thickness (left), neurite complexity (middle), and mature structure revealed by the puncta number (right) in the Day 12 mature mESC~MNs. d Overview of the experimental and analysis workflow for conducting Nanopore direct RNA-seq on mature mESC~MNs. Created in BioRender. Chen, J. (2025) https://BioRender.com/cq89p4h. e The heatmap from Nanopore direct RNA-seqshows that several feature adult genes (ChAT, Spp1, Bag, Slc5a7, Fos, and Syn1) are more enriched in the Day12 mature mESC~MNs compared to embryonic genes (Isl1, Olig2, and Mnx1). f Motif preference of m6A peaks identifies the DRACH consensus motif (D = A, T, or G, R = A or G, and H = A, T, or C). Metagene profile of enrichment of m6A-modified sites across the mRNA transcriptome. 5’UTR, 5’ untranslated region; CDS, coding sequence; 3′UTR, 3′ untranslated region. Replicates 1, 2, and 3 (R1, R2, and R3) represent the triplicate biological repeats. g KEGG pathway analysis of the m6A-modified MN epitranscriptome reveals distinct biological pathways related to neurodegenerative diseases. Terms of interest in this study are highlighted in bold purple. h Schematic for analyzing the m6A-modified MN epitranscriptome, showing that 41.98% of m6A-modified genes are ALS risk genes. i, j The verification of the predicted m6A-modified sites in Tardbp and Atp13a2 by m6A pull-down qPCR of selected high m6A-modified sites and low m6A-modified sites. Points represent individual biological experiments. All data are presented as mean ± SD, n = 3, with significant P values from two-tailed t-tests. N.S. non-significant. Source data are provided as a Source data file.
a Overview of the experimental workflow for single nucleus multiomics. Created in BioRender. Chen, J. (2025) https://BioRender.com/esjho7r. b Immunostaining showing localization of Sun1-sfGFP-myc in MNs that carry R26-CAG-LSL-Sun1-sfGFP-myc together with a Cre driver. Scale bar, 200 µm. The staining experiment was independently repeated. (n = 3 mice). c Uniform manifold approximation and projection (UMAP) representation of all nuclei that passed quality filtering. Dimensionality reduction and clustering were performed based on gene expression (RNA, left), chromatin accessibility (ATAC, middle), and weighted nearest neighbor (WNN) integration of RNA and ATAC data (right). Clusters are color-coded and annotated using label transfer prediction, referencing Blum et al., 2021. d Major cell type proportions are unaffected at P100~P120 in Sun1sfGFP; ChAT-Cre; Mettl14floxed mice, a stage before MN degeneration. e Schematic for cross-referencing DEGs, particularly those down-regulated in Sun1sfGFP; ChAT-Cre; Mettl14floxed MNs, and the m6A-modified MN epitranscriptome. The resulting data reveals distinct biological pathways (Gene Ontology, right) and KEGG (f) that might cause MN degeneration in the Sun1sfGFP; ChAT-Cre; Mettl14floxed mice. Terms of interest in this study are highlighted in bold and purple. Schematic for analyzing the dot-plot data (g), with the outcome (h) showing ALS disease risk genes displaying significant changes in expression in Sun1sfGFP; ChAT-Cre; Mettl14floxed mice in each cholinergic neuronal subtype.
To validate our computationally predicted m6A sites, we used Tardbp (Tdp43) as a benchmark to validate our methodology. Consistent with a previous report35, we confirmed the existence of a previously identified m6A site in Tardbp with a high probability rate with stoichiometry ranging from 57.38 to 92.53% (Supplementary Data 3) and verified via m6A antibody pull-down assay (Fig. 5i, site 1). Additionally, we uncovered and validated an additional high m6A-modified site within the Tardbp transcript from Nanopore direct RNA-seq (Fig. 5i, site 2). Moreover, we substantiated the existence of predicted m6A sites in Atp13a2 (Park9) (Fig. 5j)44, an ALS risk gene not previously shown to have m6A modifications in MNs. Finally, we further verified several newly identified m6A-modified sites in ALS risk genes, including Dctn1, Epha4, C9orf72, Glt8d1, Cacna1h, Chrna3, Bscl2, Fig4, Hnrnpa2b1, Ubqln2, Hnrnpa1, Tuba4a, Sod1, Chmp2b, and PIKfyve (Supplementary Fig. 6b). Thus, these observations confirm the sensitivity, accuracy, and reliability of Nanopore technology to identify m6A-modified sites in MNs.
Next, we reasoned that most m6A-modified ALS risk genes could potentially contribute to the observed ALS-like pathologies upon m6A impairment in the ChAT-Cre; Mettl14floxed mice, so we probed the consequence of hypo-m6A for MNs by performing 10x Genomics single-nuclei multimodal profiling of ATAC/RNA (snATAC/RNAseq) on Ctrl (ChAT-Cre; Mettl14f/+) and ChAT-Cre; Mettl14floxed mice, allowing us to assess chromatin accessibility and gene dysregulation in a range of MN subtypes simultaneously (Fig. 6a, b). To selectively enrich for nuclei from spinal cholinergic neurons, we bred ChAT-Cre; Mettl14floxed mice expressing the nuclear envelope reporter CAG-Sun1/sfGFP45 (Fig. 6b). We harvested lumbar spinal cords at P100~120, a stage when the MN population is not greatly diminished, and collected GFPon cells by fluorescence-activated cell sorting (FACS) for single-nuclei multimodal profiling (Fig. 6c and Supplementary Fig. 7a). Each sample underwent rigorous quality control (QC) measures and was subsequently filtered to retain only cells that only met our QC criteria (Supplementary Fig. 7a) (see Methods for details). We then integrated three replicates from the Ctrl and Sun1sfGFP; ChAT-Cre; Mettl14floxed (KO) samples (Supplementary Fig. 7b, c). Principal component analysis (PCA) revealed negligible sequencing and batch confounding variables among our sample preparations (Supplementary Fig. 7d). PC1 and PC2 largely separated the major cell types in the population, and PC3 segregated the total population based on whether the cells were from the Ctrl or Sun1sfGFP; ChAT-Cre; Mettl14floxed cohorts. Using recognized markers for spinal cord cell type annotation45,46, we detected the three major cholinergic cell populations, i.e., skeletal MNs (Tns1on/Bcl6on), visceral MNs (Nos1on), and cholinergic INs (Pax2on) (Fig. 6d and Supplementary Fig. 7e). We did not observe any changes in the proportions of these major cell types, consistent with our in vivo characterization of the pre-onset stage of ChAT-Cre; Mettl14floxed mice (Fig. 6c, d). When we further analyzed subtypes within the skeletal MNs, we identified α (Htr1dlow, Rbfox3high, Vipr2on), γ (Htr1dhigh, Rbfox3low, Spp1low, Creb5on, Pard3bon) and γ* (Htr1dhigh, Rbfox3low, Spp1low, Stxbp6on, Plch1on) MNs (Supplementary Fig. 8a-d). Among α MNs, we could further distinguish fast-fatigue-resistant (Chodlon, Kcnq5on), slow-firing (Sv2aon), and fast-fatigable (Chodlon, Kcnq5off) cell types (Supplementary Fig. 8e, f). Therefore, our snATAC/RNAseq dataset encompasses all major adult MN subtypes identified from other studies45,46.
To discern the molecular alterations underlying the MN degeneration observed in Sun1sfGFP; ChAT-Cre; Mettl14floxed mice, we concentrated on differentially expressed genes (DEGs) within distinct types of cholinergic neurons. By performing a differential expression analysis between ChAT-Cre; Mettl14floxed and Ctrl samples in individual cell types, we identified 652, 500, and 604 DEGs (with p-adjusted <0.05) for skeletal MNs (down-regulated = 291 and up-regulated = 361), visceral MNs (down-regulated = 190 and up-regulated = 310), and cholinergic INs (down-regulated = 165 and up-regulated = 439), respectively (Supplementary Data 4). To identify specific candidates presenting direct dysregulation due to m6A modifications and thus potentially contributing to neuronal degeneration, we conducted GO and KEGG pathway analyses focusing on DEGs exclusively recognized as m6A-modified based on Nanopore direct RNA-seq across all cholinergic neurons from ChAT-Cre; Mettl14floxed mice (Fig. 6e). In alignment with our findings of MN denervation and compromised neuromuscular junction (NMJ) size in the ChAT-Cre; Mettl14floxed mice (Fig. 3e-h), together with neurite degeneration in METTL3i-treated and METTL3/METTL14 knockdown iPSC~MNs (Fig. 1h-l and Supplementary Fig. 2c-e), GO analysis of m6A-modified and down-regulated genes revealed enrichment for pathways related to axonogenesis, synapse organization, cytoskeleton, and tubulin-related gene terms (Fig. 6e and Supplementary Fig. 9a). Notably, we observed the down-regulation of RNA splicing-related genes (e.g., Malat1, Srsf2, Fus, Srek1, Hnrnph1, Tra2a) in the ChAT-Cre; Mettl14floxed samples (Supplementary Fig. 9b), corroborating previous studies showing that m6A-mediated splicing induces changes in gene expression3. Among this group of analyzed genes, we noted that ALS-associated genes were prominent in our KEGG pathway analysis (Fig. 6f). Specifically, in MNs, these down-regulated genes are linked to neurofilament (Nefm, Nefl), tubulin (Kif5c, Tubb3), gene encoding DNA/RNA binding protein (Fus), and nucleoporin (Nup93) (Supplementary Fig. 9c). Mutation or aberrant expression of these genes has been shown to contribute to ALS pathologies, such as cytoskeletal defects and nucleocytoplasmic transport31. Moreover, the down-regulation of calcium signaling, which is often linked to neurodegenerative diseases47, appeared as the top enriched pathway for m6A-modified and down-regulated genes in cholinergic neurons (Fig. 6f). Among the m6A-modified ALS risk genes (34 out of 81 genes in Supplementary Data 2), expression of two genes—Fus and Bscl2—was significantly reduced, whereas that of PIKfyve was increased in the skeletal MNs of ChAT-Cre; Mettl14floxed mice (Fig. 6g, h), in accordance with dysregulated ALS risk genes identified from patients48. These alterations are likely major contributors to MN degeneration and the subsequent motor behavior deficits observed in our ChAT-Cre; Mettl14floxed mice. Taken together, these findings strongly imply a direct association between decreased m6A levels, neurodegeneration, and ALS pathology.
Hypo-m6A MNs exhibit an increase of closed chromatin regions
In contrast to the down-regulated DEGs, the up-regulated DEGs with m6A modifications exhibited conspicuous enrichment in genes responsible for regulating chromatin and histone modification (Fig. 7a and Supplementary Fig. 9d). Notably, several of these genes up-regulated in response to hypo-m6A—including members of the ATP-dependent Chd family, Bcl7c, Ncoa6, and Ube2b—have been implicated in the DNA damage response and apoptosis. These pathways are commonly implicated in diverse neurodegenerative diseases49 (Supplementary Fig. 9e). Accordingly, we noticed that the MNs of ChAT-Cre; Mettl14floxed mice displayed a drastic increase of γH2AX signals, together with a striking increase in repressive histone modification marks (i.e., H3K9me3) (Fig. 7b-e, Supplementary Fig. 10a-c). Our discovery of up-regulatedgenes linked to chromatin/histone modification prompted us to examine if changes in chromatin are associated with the aforementioned identified DEGs. To do so, we probed changes in chromatin accessibility in our snATAC/RNAseq dataset. Among the cholinergic cell types, ChAT-Cre; Mettl14floxed skeletal MNs displayed the most drastic peak changes (open peaks = 3278; closed peaks = 10,744), followed by visceral MNs (open peaks = 2495; closed peaks = 920), with cholinergic INs exhibiting the fewest peak changes (open peaks = 171; closed peaks = 602), most of these peaks are in intergenic or intronic regions (Fig. 7f). Subsequently, we integrated and scrutinized the snRNAseq and snATACseq data, revealing that only a modest subset of DEGs align with alterations in chromatin accessibility (Supplementary Fig. 9f and Supplementary Data 5). To further investigate the relationship between changes in chromatin accessibility and dysregulated gene expression, we first identified linked peak-to-gene associations and performed a correlation analysis between gene expression and changes in chromatin accessibility following Mettl14 ablation (Supplementary Fig. 10d and Supplementary Data 5). Our analysis revealed a low correlation between these two factors. Notably, only 17%, 11%, and 5% of the genes associated with regions of differential chromatin accessibility for skeletal MNs, visceral MNs, and cholinergic neurons, respectively, were both m6A-modified and differentially expressed upon Mettl14 ablation. These discoveries underscore the critical importance of preserving a nuanced equilibrium in the m6A transcriptome within adult MNs to maintain neuronal homeostasis. Diminished m6A levels may lead to compromised expression of pivotal neuronal and disease-associated genes governed by versatile regulatory mechanisms, i.e., either through direct modification of m6A-affected transcripts or by reshaping the chromatin landscape within MNs.
a Schematic for cross-referencing of DEGs, particularly those up-regulated in Sun1sfGFP; ChAT-Cre; Mettl14floxed MNs, and the m6A-modified MN epitranscriptome, with the outcome revealing distinct biological pathways (Gene Ontology, right) that might increase repressive histone modification and the DNA damage response (highlighted in bold purple) in the Sun1sfGFP; ChAT-Cre; Mettl14floxed mice. b–e Representative images illustrate a dramatic increase in repressive H3K9me3 mark (b) and DNA damage γH2AX (d) signals. Quantifications of lumbar H3K9me3on (c) and lumbar γH2AXon (e) signal intensities in the ventral regions from the spinal cord of P120 ChAT-Cre; Mettl14floxed mice compared to littermate controls (Ctrl: n = 5 mice, ChAT-Cre; Mettl14floxed: n = 6 mice, quantification for all MN nuclei from all views of captured images; Data are presented as mean ± SD with significant P values from two-tailed t-tests; Scale bars, 20 µm). f The bar plot shows changes in the number of peaks and distribution of their annotated genomic locations in cholinergic neuronal subtypes derived from Sun1sfGFP; ChAT-Cre; Mettl14floxed mice and control (Ctrl) snATAC-seq data. Source data are provided as a Source data file.
Restoring m6A homeostasis rescues MN degeneration in both familial and sporadic ALS models
The consistent manifestation of hypo-m6A in human ALS iPSC~MNs, together with our ChAT-Cre; Mettl14floxed mice recapitulating ALS pathology, prompted us to explore if bolstering the m6A reservoir could represent a therapeutic strategy. Thus, we deployed several familial ALS patient (C9ORF72exp~800 G4C2, SOD1+/L144F, TDP43G298S) iPSC~MN lines and one sALS iPSC~MN line to reflect MN degeneration (Fig. 8a–d). Then, we treated these lines with FB23-2, an inhibitor of FTO (an m6A eraser)50, to see if this approach could be applied to rescue MN degeneration (Fig. 8a). First, we differentiated the ALS iPSCs under defined conditions to cause MN degeneration through a selective ER stressor, CPA (Fig. 8a, b). Subsequently, we applied FB23-2 to determine if doing so could elevate m6A levels and thereby restore the m6A repertoire to rescue MN degeneration in different contexts of ALS. As expected, we observed a consistently significant increase in m6A levels upon applying FB23-2, albeit to varying degrees (Fig. 8c). By using SMI32 to assess MN degeneration and neurite complexity, we observed that FB23-2 promotes MN survival upon CPA stressor treatment for both familial and sporadic ALS MNs (Fig. 8d). Thus, our results indicate that fortifying basal m6A levels by adding a m6A eraser inhibitor can rescue human ALS iPSC~MNs from degeneration.
a Schematic illustration of the m6A biogenesis pathway and the applied FTO inhibitor (FB23-2) with their corresponding targeting pathways. Created in BioRender. Chen, J. (2025) https://BioRender.com/3nizfu7. b Representative images of FB23-2 rescuing the MN degeneration associated with ALS. Scale bar, 200 µm. Quantifications of m6A mRNA methylation levels (c) and degeneration index values (d) at an indicated time point and compared to the CPA treatment. Note the significant rescue of the degeneration index upon applying FB23-2 to CPA-stressed C9ORF72exp, SOD1+/L144F, TDP43G298S, and sALS iPSC~MNs. Data are presented as mean ± SD, n = 3, significant P values from two-way ANOVA. N.S. non-significant. e Heatmaps of normalized expression level between stress-treated (CPA) ALS-relevant lines with or without subsequent FB23-2 treatment, revealing restorations of many ALS risk genes with m6A modifications (highlighted with rectangles) to control (vehicle) levels for the FB23-2-treated groups. A z-score normalization was performed on the normalized read counts across samples for each gene after stress treatment (CPA) with or without subsequent FB23-2 treatment. Samples were normalized to the vehicle control to reveal the normalized expression level. Notably, following FTO inhibitor treatment, these genes were restored to levels comparable to controls among those ALS iPSC~MNs are highlighted in bold purple. Source data are provided as a Source data file.
To determine if the neuroprotective effects of the FTO inhibitor on MN degeneration in ALS are mediated through its regulation of m6A-modified ALS risk genes (Fig. 6g, h), we performed RNA-seq analysis on ALS iPSC~MNs treated with FB23-2. Differential expression analysis revealed that following FB23-2 treatment, several m6A-modified genes involved in synaptic function, RNA metabolism, and chromatin and histone modifications were restored to levels similar to controls (Supplementary Fig. 11). Notably, the expression of multiple m6A-modified ALS risk genes was returned to control-like levels (vehicle-treated), suggesting that FTO inhibition may mitigate MN degeneration in ALS by modulating m6A-modified gene expression (Fig. 8e). These findings indicate that enhancing m6A levels in ALS iPSC~MNs using small molecules could help restore the balance of the m6A epitranscriptome and preserve MN integrity.
Fto gene therapy extends the lifespan of SOD1 G93A mice and ameliorates their MN degeneration
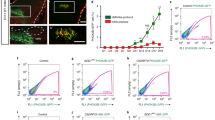
Although human iPSCs may be used as an effective platform for drug screening, they preclude further investigation of treatment efficacy at the tissue/organ/behavioral levels. Moreover, the penetrance of FTO inhibitors through the blood-brain barrier is poor and so systematic appliance of such inhibitors might cause adverse effects51. To circumvent this issue, we adopted gene therapy as an alternative approach and delivered self-complementary adeno-associated vector serotype 9 (scAAV9) of Fto-shRNA to knock down Fto expression in the spinal cord (Fig. 9a, b and Supplementary Fig. 12). We first verified the knockdown efficiency of three Fto-shRNAs and their restoration of m6A level in C2C12 cells (Supplementary Fig. 12a–c). Then we subcloned the Fto-shRNAs into scAAV9 driven by the H1 promoter (scAAV9-shFto)38. To test the efficiency of viral infection, we injected scAAV9-EGFP into control mice and observed sustained GFP expression in their spinal MNs and in some dorsal cells at 40 days post-injection (Supplementary Fig. 12d). Subsequently, scAAV9-shFto was injected into SOD1G93A mice and their wild-type littermates. Then we verified a dramatic reduction in Fto level in the mouse spinal cords (Fig. 9c). Prominently, scAAV9-shFto gene therapy delayed the disease onset (Fig. 9d) and robustly prolonged the median survival of SOD1G93A mice to ~14 days (Fig. 9e). In the SOD1G93A ALS mouse model, we observed a reduction in m6A methylation by m6A immunostaining (Fig. 9f). Moreover, we also found that m6A levels can be upregulated via intrathecal delivery of scAAV9-shFto in the SOD1G93A mouse model (Fig. 9f). The enhanced MN survival (Fig. 9g, h) with a significant reduction of gliosis (Fig. 9g, i) is also shown in SOD1G93A; scAAV9-shFto mouse model.
a, b Overview of the experimental strategy. Created in BioRender. Chen, J. (2025) https://BioRender.com/3m2vxum. c Western blot reveals that Fto protein level is reduced in mouse lumbar spinal cords after scAAV9-shFto injection (n = 3 mice). d and e Kaplan-Meier survival curves with log-rank test revealing prolongation of the onset of weight decline in SOD1G93A mice (from ∼140 to ∼155 days), with lifespans extended by ∼10% (from ∼170 days to ∼187 days), following scAAV9-shFto injection. m6A levels is upregulated (f), MN number is rescued (f, with quantification in h), and gliosis is reduced (g, with quantification in i) after scAAV9-shFto injection of SOD1G93A mice at P140. Scale bars, 100 µm. (mean ± SD, n = 4 mice; two-tailed t-tests). j The CMAP amplitude is reduced in SOD1G93A mice at P60 and gradually declines further over time, whereas scAAV9-shFto treatment significantly ameliorates neuromuscular function at P160 (mean ± SD, Ctrl: n = 5/1, 4/3, 3/3, and 4/3 mice, Ctrl; scAAV9-shFto: n = 5/1, 5/2, 5/2, and 5/2 mice; SOD1G93A: n = 2/3, 5/6, 4/5, and 4/3 mice; SOD1G93A; scAAV9-shFto: n = 3/2, 5/3, 4/3, and 4/3 mice at P60, 120, 140, and 160 from males/females respectively; two-tailed t-tests) (right). k, l Motor coordination and muscle strength are enhanced by scAAV9-shFto injection, as assayed by rotarod test at P60~P160 (k) (mean ± SD, Ctrl: n = 6/2, 4/3, 4/3, and 4/3 mice, Ctrl; scAAV9-shFto: n = 5/1, 5/2, 5/2, and 5/2 mice; SOD1G93A: n = 3/4, 5/6, 5/6, and 5/5 mice; SOD1G93A; scAAV9-shFto: n = 5/2, 5/3, 5/3, and 5/3 mice at P60, 120, 140, and 160 from males/females respectively; two-tailed t-tests), and by grip strength test (l) (mean ± SD, Ctrl: n = 6/2, 4/3, 4/3, and 4/1 mice, Ctrl; scAAV9-shFto: n = 5/1, 5/2, 5/2, and 5/2 mice; SOD1G93A: n = 4/4, 5/6, 5/6, and 4/3 mice; SOD1G93A; scAAV9-shFto: n = 5/2, 5/3, 5/3, and 5/3 mice at P60, 120, 140, and 160 from males/females respectively; two-tailed t-tests). N.S. non-significant. Illustrations in j–l were created in BioRender. Chen, J. (2025) https://BioRender.com/mrtnwlk. Source data are provided as a Source data file.
To evaluate the mice in a more clinically relevant setting, we assayed MN and gastrocnemius (GA) muscle connectivity by measuring the compound motor action potential (CMAP). We performed the CMAP assay from P60 (directly before AAV treatment) to P160 (Fig. 9j). Consistent with a previous study, CMAP amplitude is already reduced in SOD1G93A mice at P60 and gradually declines further over time, whereas scAAV9-shFto treatment mildly ameliorates neuromuscular function at P160. (Fig. 9j). This moderate improvement was also reflected by the enhanced behavioral performance of SOD1G93A mice from P120 following scAAV9-shFto treatment (Fig. 9k, Supplementary Movie 4), with a substantial increase of muscle strength (Fig. 9l), a major clinical measurement for ALS motor score.
To determine if scAAV9-shFto treatment rescues SOD1G93A mice through m6A-mediated molecular changes, such as histone modifications in H3K9me3 and γH2AX, we assessed these markers in four groups—wild-type control (Ctrl), Ctrl treated with scAAV9-shFto (Ctrl; scAAV9-shFto), SOD1G93A, and SOD1G93A treated with scAAV9-shFto (SOD1G93A; scAAV9-shFto)—at postnatal day 140 (P140), i.e., when SOD1G93A mice are at an early symptomatic stage (Fig. 10). Remarkably, H3K9me3 levels were significantly elevated in the ventral horn MNs of the SOD1G93A mice and were restored to control levels following scAAV9-shFto treatment (Fig. 10), suggesting that epigenetic dysregulation may be a novel hallmark of ALS pathogenesis mediated by hypo-m6A. In contrast, no significant changes in γH2AX were observed in the spinal cords of the SOD1G93A mice (Supplementary Fig. 13).
a Representative image illustrating H3K9me3 marks (yellow arrowheads) in the ventral horn of different sets of Ctrl and SOD1G93A mice, with scAAV9-shFto intrathecal injections. b Quantifications of lumbar H3K9me3on MNs (Ctrl: n = 7 mice, Ctrl; scAAV9-shFto: n = 3 mice, SOD1G93A: n = 4 mice, SOD1G93A; scAAV9-shFto: n = 4 mice, quantification for all MN nuclei from all views of captured images; Scale bars, 20 µm). Data are presented as mean ± SD with significant P values from two-way ANOVA. N.S. non-significant. Source data are provided as a Source data file.
Overall, our results support that m6A hypomethylation promotes ALS and, significantly, augmenting the m6A reservoir can mitigate the disease phenotypes of sporadic and familial human ALS iPSC~MNs. In a SOD1-linked ALS context, enhancing m6A levels in adult MNs either via small molecule treatment or scAAV9 delivery delays the onset of MN degeneration and enhances motor function. The discoveries we have presented herein reveal m6A as a potential prognostic indicator for MN degeneration and a promising therapeutic candidate in individuals afflicted with ALS. We discuss the significance and implications of our study in detail below.
Discussion
In this study, we generated two conditional spinal MN-Mettl14 knockout mouse lines, using the Olig2-Cre; Mettl14floxed mice to impair m6A homeostasis from the embryonic stage and the ChAT-Cre; Mettl14floxed mice to disrupt m6A levels from the late embryonic/early postnatal stage. Although Olig2-Cre; Mettl14floxed mice exhibit early postnatal lethality (~P24 to P28), their MNs are relatively normal. The ChAT-Cre; Mettl14floxed mice also display normal MN development and ordinary gross appearance at the postnatal and juvenile stages. These results indicate that m6A homeostasis appears to be less critical for embryonic spinal MN development. This scenario is different to conditional Mettl14 knockout in mouse neural progenitor cells (NPCs) using Nestin-Cre that impairs NPC differentiation, prolongs cell cycle progression of radial glia, and extends cortical neurogenesis into postnatal stages18,19. Thus, our results, together with these previously published results, emphasize the neuron-type context-dependent role of m6A homeostasis, prompting the notion that the m6A epitranscriptome is both dynamic and diverse, and it operates in a cell and developmental context-dependent manner. Our Olig2-Cre; Mettl14floxed mice display a shivering phenotype, likely a reflection of compromised oligodendrocytes, consistent with a previous study using a different Olig2-Cre line40. We have shown previously that the Olig2-Cre line we deployed in this study displays more efficient floxed allele removal activity than the other line52 used in that previous study, so it is understandable that we observed a more lethal shivering postnatal phenotype than the other study that only showed oligodendrocyte defects at the adult stage40.
Despite sporadic ALS cases predominating in the real world, current animal models often rely on familial genetic mutations. The expanding genetic spectrum of ALS has prompted continuous efforts to establish rodent models that emulate human physiological deficits and pathological manifestations30,48,53. Nevertheless, translation of effective treatment outcomes in rodent models to clinical success in humans remains limited, emphasizing the crucial need for an authentic rodent ALS model that mirrors human symptoms. Thus, even though rodent models of familial ALS are prevalent, creating sporadic ALS animal models is pivotal for pathomechanistic insights and establishing novel therapeutics. Various sporadic ALS pathologies in MNs have been simulated in mice, such as reduced ADAR2 protein levels or filament alterations54,55. Environmental toxin exposure, including metals and β-N-methylamino-L-alanine, has also been explored alongside patient-derived fluid injections56. These models partially replicate ALS features, but their status as true sporadic ALS models is arguable. Here, our ChAT-Cre; Mettl14floxed mice exhibit several salient features of human ALS pathologies and symptoms (Supplementary Fig. 14a). At the phenotypic level, our ChAT-Cre; Mettl14floxed mice exhibit late-stage paralysis with early adult lethality, diminished body weight, kyphosis with muscle atrophy, as well as coordination and motor deficits. At the pathological level, our ChAT-Cre; Mettl14floxed mice display gradual MN loss, neuroinflammation, axonopathy, neuromuscular junction abnormality, and muscle denervation. Of paramount significance, our novel mouse model distinctly manifests early-onset cytoplasmic aggregations of Tardbp (Tdp43) and Fus, representing a principal hallmark observed in the post-mortem tissues of both familial and sporadic ALS cases, yet this phenotype has been conspicuously rare in existing rodent models. Moreover, we observed consistent m6A hypomethylation in several familial and sporadic ALS lines we tested, congruent with a recent study indicating that the expression of m6A methyltransferases and levels of m6A RNA modification are down-regulated in C9ORF72-ALS/FTD patients36. Thus, taken together, we assert that m6A hypomethylation leads to ALS and that impairing m6A modification in adult MNs has provided us with one of the most reliable rodent models of ALS. Although ChAT-Cre; Mettl14floxed mice exhibited several ALS-like phenotypes, we were unable to determine whether these effects were specifically attributable to C-boutons synapsing on the soma and proximal dendrites of MNs, the MNs themselves, or both. This is an intriguing question, as recent reports have suggested that spinal inhibitory neurons, including C-boutons, might degenerate before MNs in a mouse model of ALS57,58. Currently, the lack of a Cre driver that specifically targets adult MNs limits examining this question. However, our single-nuclei ATAC-seq data offer a promising avenue for identifying enhancers specific to adult cholinergic INs and MNs. Generating Cre lines driven by these enhancers could allow for a more precise dissection of the respective contributions of INs and MNs to the observed phenotypes.
As a hallmark of ALS, why do Tardbp (Tdp43) and Fus move out of the nucleus to form cytoplasmic aggregations in the ChAT-Cre; Mettl14floxed mice? A previous study has revealed that a lack of specific RNA modifications may affect global and/or local translation rates, consequently increasing protein aggregation59. Thus, it has been proposed that RNA modifications serve as conduits of information linking a cell’s metabolic condition with its translational productivity60. Consequently, any disruption in regulating RNA modifications could potentially perturb the equilibrium between metabolic processes and protein synthesis. Further work is needed to disentangle the causal relationship between dysregulation of RNA modifications and Tdp43 translocation. Additionally, the nuclear m6A reader YTHDC1 has been shown to exert an important role in modulating many biological processes and contributing to disease, especially cancers. YTHDC1 might be the main mediator of a series of m6A readers, thereby controlling their activity in neuronal functions61. Moreover, YTHDC1 RNAs have been discovered as binding to TDP43 protein in human SH-SY5Y neuroblastoma cells62, and peripheral blood sample transcriptional profiling of a huge heterogeneous ALS cohort (not only sporadic cases) revealed YTHDC1 to be differentially expressed63. These results point to YTHDC1-mediated m6A nuclear events and nucleocytoplasmic trafficking as contributing to ALS. Further experiments are warranted to dissect this potential disease mechanism in detail.
Based on four lines of evidence, we believe m6A hypomethylation is one of the key factors leading to ALS. First, the reduced presence of the m6A writer complex (METTL3/METTL14) is manifested in a random yet comprehensive (n > 800) selection of familial and sporadic ALS iPSC~MNs and postmortem tissues, consistent with a recently published study36. Second, a global reduction in the m6A epitranscriptome of several familial ALS iPSC~MNs precedes MN degeneration. Third, ChAT-Cre; Mettl14floxed mice phenocopy ALS symptoms at molecular, cellular, and phenotypic levels. Fourth, restoring the m6A repertoire significantly mitigates ALS pathology in human iPSC and mouse models. However, the discrepancy between the current study’s findings (i.e., the impact of hypo-m6A) and another recent study reporting m6A hypermethylation in the spinal cord of sporadic ALS patients warrants critical examination35. Several plausible explanations may account for this disparity. For instance, experimental model limitations, with previous studies predominantly utilizing cellular models and postmortem tissues, thereby potentially limiting direct correlations to in vivo pathology. The timing of sample collection, particularly at the end stage of the disease, might not adequately capture the initial contributors to ALS pathology. We suggest that studies involving animal models are necessary to establish a more direct link between manipulating m6A levels and ALS progression. Alternatively, the discrepancy could be attributable to the multifaceted and complex nature of sALS, encompassing diverse pathological mechanisms. To detect m6A levels, different assays have been used by our study and those of others, including antibody-based pull down, dot blot, ELISA, and microarray method51. We have further applied Oxford Nanopore direct RNA-seq to uncover m6A stoichiometry, which appears to provide better sensitivity. Each of these methods has advantages and disadvantages in terms of the sensitivity and feasibility of large-scale studies51. We advocate screening m6A levels in a larger cohort of sALS cases using advanced techniques such as direct Mass Spectrometry in future. Our study acknowledges the pivotal role of an optimal m6A reservoir in maintaining adult MN function, highlighting that both m6A hypomethylation and hypermethylation could be detrimental3. This nuanced perspective underscores the need for a balanced understanding of m6A modifications in ALS pathophysiology and it calls for further investigation using diverse methodologies and models to comprehensively understand the role of m6A dysregulation in ALS progression.
In our proposed model (depicted in Supplementary Fig. 14b), we postulate that the aberrant down-regulation of the two m6A methyltransferases, namely METTL3 and METTL14, in ALS reflects an anomaly likely associated with the natural aging process compounded by unidentified exacerbating factors29. This reduced expression of m6A methyltransferases has multifaceted implications. Firstly, it instigates widespread dysregulation across the mRNA transcriptome, notably enriching pathways associated with ALS risk genes, synaptic activity, and neuronal functional pathways. Simultaneously, m6A modification of chromatin regulators, crucial for modulating heterochromatin repression, is diminished. This global dysregulation of m6A-mediated epitranscriptomic processes disrupts RNA stability and perturbs the repressive chromatin landscape, potentially exacerbating the degeneration of MNs in the context of ALS. Strikingly, we observed a significant increase in closed chromatin regions in the ChAT-Cre; Mettl14floxed mice, with only a small fraction of these chromatin changes correlating with dysregulated gene expression. Given that previous studies have shown FTO-mediated LINE1 RNA m6A demethylation plays a role in regulating chromatin state and gene expression during mouse oocyte and embryonic development64, and that retrotransposon reactivation has been observed in some ALS postmortem tissues62, it would be intriguing to explore in future studies whether Mettl14-mediated m6A impairment leads to retrotransposon reactivation through its interaction with chromatin state.
In this study, we adopted two approaches to test possible treatments of ALS by bolstering the m6A reservoir. First, by applying a small molecule FTO inhibitor (FB23-2) in our familial and sporadic ALS iPSC~MNs, we prevented the MNs from degenerating. FTO inhibition notably restored to control levels the expression patterns of several genes associated with ALS risk and chromatin-regulated pathways. This outcome strongly suggests that the FTO inhibitor’s efficacy in preserving MNs from degeneration primarily stems from its ability to restore m6A-modified genes relevant to ALS pathology. Second, we observed a significant enhancement of motor and neuromuscular functions, together with a significant delay in disease onset following scAAV9-shFto intrathecal injection in our SOD1G93A mice. Building on this exciting outcome, we are currently testing some new synthesized FTO inhibitors with stronger activity and better blood-brain barrier penetrance. We knocked down FTO using a ubiquitous promoter as an initial attempt. Future optimization to knock down FTO in a cell type-specific manner might minimize possible adverse effects, although targeting FTO might also represent a relatively safe treatment strategy as Fto knockout mice are morphologically normal and only display mild learning defects65,66. Future experiments to develop a new drug cocktail by combining small molecules together with gene therapy to fortify the m6A reservoir of both familial and sporadic ALS patients offer tantalizing treatment prospects. While we have validated several ALS risk genes with m6A-modified sites that are dysregulated in the ChAT-Cre; Mettl14floxed mice, the direct contribution of these genes as a cohort to MN degeneration in ALS remains to be systematically tested. Furthermore, FTO is not only an m6A demethylase, but also functions as a demethylase for the m2-isoform (N6,2′-O-dimethyladenosine, m6Am) in small nuclear RNAs (snRNAs)14 and mRNAs. Although we have demonstrated that FTO inhibition rescues the expression of many ALS pathway- and chromatin regulator-related genes, it remains to be investigated if this therapeutic effect involves m6Am-modified snRNAs.
Additionally, though we identified an increase in H3K9me3 and closed chromatin regions as potential new features of ALS, the precise mechanism of how hypo-m6A induces these chromatin changes remains unclear. It is also uncertain if the increase in H3K9me3 directly correlates with closed chromatin regions. Future large-scale investigations using human ALS iPSC~MNs will be crucial to further explore these potential pathological features. Moreover, it would be valuable to assess if combinatorial treatments targeting both H3K9me3 and m6A modifications could offer enhanced benefits in preventing MN degeneration. Extending beyond the challenge of ALS, perturbations of the m6A reservoir is a recurring theme of other neurodegenerative conditions, including Alzheimer’s disease, Parkinson’s disease, and multiple sclerosis3. As these pioneering paradigms relating to m6A modification ripple through the scientific landscape, we envisage their transformative potential will extend to diverse ailments in the future.
Methods
Ethical compliance
All mice experimental procedures were performed in accordance with guidelines approved by the Institutional Animal Care and Use Committee (IACUC) at Academia Sinica (protocol number 13-06-559 and 23-07-2022).
Human iPSC culture
The SOD1+/L144F ALS iPSC mutant line (female) and a healthy control line were acquired from the Harvard Stem Cell Institute iPSC Core Facility. TDP43G298S (male), C9ORF72exp (male), an isogenic control line of C9ORF72exp, and sporadic ALS iPSC lines (male) were acquired from the Answer ALS project of Cedars-Sinai. Cells were maintained in feeder-free Essential 8 (Life Science) conditions and subcultured by 0.5 mM EDTA treatment. Cells were cultured in a 5% CO2 humidified atmosphere at 37 °C. The influence of sex was not assessed in this study. Rather the effect of CPA treatment was compared between isogenic lines; the FB23-2 treatment to rescue ALS iPSC lines was compared with the vehicle control.
Differentiation and survival assay of human iPSC~MNs
Human iPSCs were differentiated into MNs using an improved protocol67. Specifically, iPSCs were dissociated into single cells using accutase (Gibco) at day 0. Next, 1.5~2 × 105 cells were resuspended in 10 mL N2/B27 medium [1:1 of DMEM-F12 and Neurobasal medium containing N2 (Life Technologies), B27 (Life Technologies), 1% penicillin-streptomycin, 200 mM Glutamax, 0.2 mM 2-mercaptoethanol, and 0.5 mM ascorbic acid (Sigma-Aldrich)], supplemented with 10 mM Y-27632 (STemGent), 20 mM SB431542 (Merck), 0.1 mM LDN 193189 (Sigma-Aldrich), 3 mM CHIR-99021 (Merck), and 10 ng/mL bFGF (Peprotech). Small embryoid bodies (EBs) should become visible after two days of differentiation. SB431542 and LDN 193189 were kept for four days and supplemented with 100 nM retinoic acid and 0.5 mM smoothened agonist from day 2 to day 16. Then, 10 ng/mL BDNF (Peprotech) was added from day 7, and 10 mM DAPT (Calbiochem) was included from day 9 to day 16. After HB9on nascent MNs had been generated (day 11), EBs were dissociated utilizing accutase, and the dissociated MNs were plated onto poly-L-Ornithine/laminin-coated four-well plates at a density of 5 × 104 cells per well and 7.5 × 105 cells per 6 cm plate. Dissociated MNs were maintained in MN culture medium [CultureOne Supplement medium (Thermo Fisher Scientific Inc.) containing 10 mM Y-27632, 10 ng/mL BDNF, 10 ng/mL GDNF (Peprotech) and 10 mM 5-fluoro-20-deoxyuridine/Uridine (to inhibit proliferating cells) (Merck)]. The medium was replenished every three to five days.
FTO inhibitor treatments
To accelerate MN degeneration, the CultureOne Supplement medium of day 31 MN culture was replaced with N2/B27 medium only (no BDNF and GDNF), for which CPA was supplemented to accelerate degeneration for another seven days. FB23-2 is a small molecule that inhibits the demethylase activity to elevate m6A levels. FB23-2 (1 nM) was treated together with the CPA (50 µM) for 15~30 days (change the small molecules and medium every two days) in ALS iPSC~MNs with the increased m6A mRNA methylation level and extended disease onset or rescue of the degenerative process. Then MNs were revealed by SMI32 immunostaining and captured by an ImageXpress® Micro XLS High-Content Imaging System (Molecular Devices). The degeneration index was calculated at the indicated time point after CPA treatment.
Quantification of neurite degeneration
To ensure accurate measurement of neurites, images were captured from blindly selected regions with well-separated axon tracts. Neurite fragmentation was then quantified using an automated image analysis method. The extent of neurite degeneration was expressed as a degeneration index (DI), defined as the ratio of the fragmented neurite area to the total neurite area. To process images for DI calculation, gray intensity in images was first normalized using the auto-level function of GNU Image Manipulation Program (GIMP) software, ensuring consistent background intensity across all images. Subsequently, ImageJ and Ilastik software were employed to binarize the images and remove cell bodies, resulting in a black-and-white rendering of neurites. While intact neurites exhibit continuity, degenerating neurites display disrupted, particulate structures due to blebbing and fragmentation. To quantify the fragmented areas of these degenerating neurites, the Particle Analyzer algorithm in ImageJ was used, with detection parameters set for size (20–10,000 pixels) and circularity (0.2–1.0). The total area of identified neurite fragments was then divided by the overall black neurite area to calculate the DI. Consistent with previous studies, the DI ranged from 0, representing completely intact neurites, to 1.0, indicating complete degeneration into fragmented particles68,69.
m6A methylation and quantification
The total RNA or mRNA was extracted from samples using Trizol (Life Technologies) and m6A methylation was quantified using the m6A RNA Methylation Assay Kit (ab185912, Abcam), with absorbance read at 450 nm in biological triplicates using an EnSpire Multimode reader and EnSpire software. To quantify relative m6A RNA methylation, the percentage m6A content in total RNA was calculated as:
m6A dot blot assay
Total RNA was extracted with Trizol (Life Technologies) and purified into mRNA using Invitrogen PolyA+ RNA selection (Dynabeads mRNA Purification Kit). After mRNA purification, 100~200 ng mRNA was dropped onto the Hybond-N+ membrane (Amersham Hybond™ -N+ Membranes) for UV crosslinking (1200 μJ X100; UV Stratalinker 2400). Hybridized mRNA was blocked in 5% milk in 0.5% Triton X-100/PBS for one hour and then incubated with anti-m6A antibody (1:1,000, Synaptic) in a blocking solution at 4 °C overnight. After three TBST washes, the mRNA was incubated with an anti-HRP secondary antibody (1:10,000; Santa Cruz) in a blocking solution for 30 minutes at room temperature. The signal was developed with enhanced chemiluminescence (ImageQuant LAS 4000). The hybridized mRNA was stained with 0.2% methylene blue in 0.3 M sodium acetate (pH 5.2) as the loading control of mRNA.
Mouse crosses
Mice carrying the mutant human SOD1G93A transgene (B6SJL-Tg(SOD1*G93A)1Gur/J) were purchased from the Jackson Laboratory (JAX:002726). Mettl14floxed mice were obtained from Chuan He in the University of Chicago18, and Olig2-Cre was a gift from Tom Jessell in Columbia University. ChAT-IRES-Cre (Δneo) (Jackson Laboratory stock: 031661), in which the neomycin cassette was removed to avoid the ectopic expression sometimes observed in the ChAT-IRES-Cre line, were bred with the Mettl14floxed allele to generate ChAT-Cre; Mettl14f/+ and ChAT-Cre; Mettl14floxed mice. WT and conditional knockout mice were generated by crossing ChAT-Cre; Mettl14f/+ males and Mettl14floxed females. CAG-Sun1/sfGFP mice were purchased from JAX (stock no. 021039; B6;129-Gt(ROSA)26Sortm5(CAG-Sun1/sfGFP)Nat/J) and crossed with ChAT-Cre; Mettl14floxed mice. F1 heterozygous reporter mice were maintained to P100~P120 and then sacrificed for subsequent single-nucleus multiome experiments. All live animals were kept in an SPF animal facility, housed at ~ 55% humidity, 25 °C, on a 12:12-hour light/dark cycle, and approved and overseen by IACUC, Academia Sinica.
Immunostaining
All adult Spinal cord sections were permeabilized in 0.5% Triton X-100/PBS for one hour and then blocked in 3% bovine serum albumin (BSA) in 0.5% Triton X-100/PBS for one hour. The sections were incubated with indicated primary antibodies in blocking solutions at 4 °C for two days. After five PBS washes, the sections were incubated with secondary antibodies and DAPI in a blocking solution for 1.5~2 h at room temperature. After five PBS washes, the sections were mounted with Aqua-Poly/Mount (18606-5; Polysciences Inc.). Dissociated MNs from iPSC differentiation were fixed and permeabilized in 0.1% Triton X-100/PBS for 5~10 min and then blocked in 10% FBS in 0.1% Triton X-100/PBS for 30 min. The sections were incubated with indicated primary antibodies in a blocking solution at 4 °C overnight. After three PBS washes, the sections were incubated with secondary antibodies and DAPI in a blocking solution for 45 min at room temperature.
Spinal motor neuron quantification
At each indicated stage, ChATon MNs in the lumbar ventral horns with DAPI nuclear signal were counted on one side of a 20-μm-thick sectioned spinal cord. MNs that did not show regular nuclear shapes were excluded. The quantification bar charts represent average MN counts from at least three sections per mouse (n ≥ 3 mice) of the same age and genotype.
Survival analyses
Mice were assessed weekly for baseline weight from P60. The onset of weight decline is defined as the age at which the animal lost 5% of peak body weight. We used the Kaplan–Meier method with a log-rank test to compare onset-free survival for each group. The disease end-point was defined as the day when mice could not right themselves within 15 s70, when they were sacrificed. Survival analysis was performed using Kaplan–Meier analysis.
NMJ analysis
To reveal neuromuscular junctions (NMJs) in whole-mount muscles, tissues were permeabilized and blocked in 3% bovine serum albumin (BSA) in 2% Triton X-100/PBS at room temperature for two hours, followed by incubation with anti-neurofilament antibody (DSHB, 1: 250) together with anti-SV2 (DSHB, 1: 500) to label axonal endfeet in blocking buffer at 4 °C for three days. After five 2% Triton X-100/PBS washes, the muscle tissues were incubated overnight at 4 °C with secondary antibodies, together with Alexa Fluor 555 labeled a-bungarotoxin [α-BTX, Thermo Fisher Scientific, B13422] to detect nicotinic acetylcholine receptors (AChR). Muscles were teased apart and flattened before mounting on slides. Z-stack images were acquired using a Zeiss LSM780 confocal microscope. All figures containing confocal images are projections of Z stacks. Innervated NMJs were counted when the AChRon endplates overlapped with axon terminals (as revealed by neurofilament staining), whereas the denervation ratio was calculated according to the colocalization between neurofilament, SV2 and the α-BTX signals of each picture, with values normalized against total endplate area. All NMJ areas were analyzed in ImageJ using threshold adjustment. Herein, we only examined the total denervation ratio, as represented by marker non-colocalization in NMJs.
Behavioral assays
Locomotor activity was measured according to open field, rotarod, and treadmill tests. Age-matched wild-type (WT) mice from the littermate control line were used for experimental comparisons. Both sexes of indicated ages (P40, P70, P100, P130, and P160 for ChAT-Cre; Mettl14floxed mice) were used in this study. The rotarod, grip strength, and CMAP tests were conducted in SOD1G93A mice one week prior to intrathecal injection (P60) and subsequently assessed at the specified time points (P120, P140, and P160). The experimenters conducting all behavioral assays were blind to mouse genotypes.
Rotarod
A commercially available rotarod apparatus (47600 Rota-Rod, Ugo Basile, Italy) with a rotating rod of 5 cm diameter was used. Mice were transferred to the testing room and habituated in the home cage at least 15 minutes before testing. In the training phase, three trials with a constant speed of 4 rpm and a 60-s cut-off time were used to ensure that all test mice could stay on the rod for a training trial before moving to the test phase. After a 30-minute rest interval, the mice were evaluated during the test phase, with the rod accelerating from 4 to 40 rpm with a 300-second cut-off time in a series of three trials. The longest falling latency, as well as the rotating speed when the mouse fell off the apparatus, were used to represent the motor coordination of each mouse71.
Grip strength
A grip strength meter (MK-380CM/R; Muromachi) was used to measure forelimb grip strength. As a mouse grasped the bar, the peak pull force in grams was recorded on a digital force transducer. During the test, a mouse was allowed to grasp the bar mounted on the force gauge. We performed four consecutive measurements per test at one-minute intervals.
Open-field test
A square arena with opaque walls (area 48 × 48 cm and height 35 cm) was used. Mice were transferred to the testing room and habituated in the home cage one hour before testing and then allowed to explore the test chamber for another hour during which all behaviors were videotaped and tracked using a video-tracking system mounted on top of the arena (Clever System, Reston, VA). Total distance and average velocity were analyzed for each mouse71.
Treadmill locomotion analysis
A TreadScan apparatus (CleverSys, Reston, VA) was used to analyze gait. Mice were placed on a stationary treadmill for acclimation and trained at a speed of 10 cm/s for 5 minutes before testing. Four test speeds were analyzed (10, 15, 20, and 25 cm/s) for each trial, which were recorded at 79 frames/s for 10 s using TreadScan software. For data analyses, the successful trials in which a mouse was able to maintain treadmill speed with continuous locomotion for each 10-s recording was selected and further analyzed using TreadScan software. Only gait analyses from the trials conducted at 15 cm/s are shown in the present study. The gait parameters of stride for each limb were automatically and unbiasedly calculated, and average values were used for statistical analysis.
Mouse motor neuron differentiation
Mouse embryonic stem cells were differentiated into MNs following published protocols38,41. Specifically, in this condition, we cultured the 3D embryoid bodies for five days, then attached the culture of the embryoid bodies to poly-ornithine and laminin-coated plates with an enrichment of neurotrophic factors (30 ng/mL GDNF, Peprotech) and cultured them for an extra 5 days (Day12). Change the medium every two days. Mouse ESC~MNs were used for imaging-based experiments, qPCR, and Nanopore direct RNA-seq.
Quantification of neurite thickness, neurite complexity, and synaptic puncta numbers
Using an automated image analysis method, the synaptic puncta labeled with SynI and the neurite labeled with Smi32 were measured. For synapse quantification, images were processed with ImageJ and ilastik software to binarize the images and eliminate the background and embryonic body, resulting in an image featuring black synapses and neurites on a white background. The Particle Analyzer algorithm in ImageJ was utilized to quantify the number of synaptic puncta within the neurites, setting parameters for size (0–100 pixels) and circularity (0.2–1.0). The neurite length was measured using the Analyze Skeleton algorithm in ImageJ. The total number of the identified synaptic puncta was then divided by the overall neurite length to calculate the average number of synapses. For the neurite complexity and thickness measurement, the neurite outside the embryonic body was imaged. Neurite diameter was measured manually at the segment of each neurite within the fields with ImageJ’s line tool. The image was processed with ImageJ and ilastik software to quantify the branch point and analyzed using the Analyze Skeleton algorithm in ImageJ. The total number of branch points was then divided by the overall neurite length to calculate the complexity.
Nanopore direct RNA-seq and data analysis
Nanopore direct RNA-seq was conducted following instructions provided by Oxford Nanopore Technologies (Oxford, UK) using a Direct RNA Sequencing Kit (SQK-RNA002) and MinION flowcells (FLO-MIN106 R9 version). After live base-calling using Guppy (22.10.7) in MinKNOW, reads that passed the quality threshold were subjected to post-run base-calling with the latest version of Guppy (6.3.9) under default parameters. A reference transcriptome file was generated from the mouse genome reference and corresponding annotation. The Nanopore data provided in FASTQ format were initially aligned to the mouse genome reference from GenCode M20 using the minimap2 aligner. The resulting SAM format output was then converted to a sorted BAM format using samtools. To predict m6A modification sites and stoichiometry, we employed both EpiNano42 and m6Anet43. For EpiNano prediction, the output was first filtered with the DRACH motif. Then, the probability of modification was taken from the ‘ProbM’ column and the results from the ‘prediction’ column, and then values > 0.5 were filtered for the results. For m6Anet prediction, the probability was yielded from the ‘probability_modified’ column and then filtered for values > 0.9 for the results. Each tool predicted m6A sites independently, which were subsequently converted into BED format for further analysis. To refine our predictions, we cross-referenced predicted sites from EpiNano and m6Anet for each sample. We then identified the shared sites across EpiNano and m6Anet, finalizing our list of predicted m6A sites. Each m6A modified site was annotated using Homer v4.11 to determine its genomic location, categorizing them as Transcription Start Site (TSS), Transcription Termination Site (TTS), Exon (Coding), 5′ UTR Exon, 3′ UTR Exon, Intronic, or Intergenic. To estimate the predicted site with stoichiometry, the modification results from the “mod_ratio” column to quantify the percentage of reads in a given site.
m6A immunoprecipitation and quantitative real-time PCR
To verify m6A sites, total RNA was isolated from mESC~MNs using Trizol (Life Technologies). Total RNA samples were pooled to a total RNA amount of ~75 µg/sample and processed using Invitrogen PolyA+ RNA selection (Dynabeads mRNA Purification Kit, Cat. #61006) according to the manufacturer’s protocol. Input mRNA (1%) was reserved for reverse transcription. Magnetic A beads (#88802, Thermofisher Scientific) were prepared and washed, adding 2 µl of N6-Methyladenosine Antibody/per sample. m6A-modified transcripts of interest were pulled down using a rabbit polyclonal anti-m6A antibody (Synaptic Systems). The m6A-modified mRNAs were competitively eluted from beads using N6-Methyladenosine 5′-monophosphate sodium salt. Input m6A pull-down mRNA was reverse transcribed by using the SuperScript III First-Strand Synthesis System for RT-PCR (Thermo Fisher). cDNA was then used for SYBR-green-based quantitative real-time PCR. Enrichment of m6A-modified genes in m6A pull-down over input was calculated by comparing relative concentrations using Ct values (2-Ct) and dividing each concentration by the relative concentration of the input. The concentrations of the immunoprecipitated RNA were then divided by the concentration in the input RNA and multiplied by 100, to obtain the percentage of transcripts in the m6A immunoprecipitate relative to the input. This value was then normalized to low enrichment in m6A-modified genes, which was also calculated using relative concentrations to determine a percentage of the input. Primers used are listed in Supplementary Data 6.
Nuclei collection for single-nucleus multiome
Mice were euthanized with isoflurane according to IACUC Academica Sinica guidelines. Mice were decapitated and spinal cords were extracted using a hydraulic extrusion approach. To do so, a blunt 25G ¼ inch needle filled with ice-cold 1 × PBS was placed into the caudal end of the vertebral column and a rapid hydraulic pressure was introduced to extract the entire spinal cord. Lumbar segments were dissected based on morphology and dissociated two at a time in 2 mL pre-chilled lysis buffer with a 7 mL Dounce Homogenizer (pestle A: eight strokes; pestle B: 5 strokes). An additional 3 mL lysis buffer was added, and the entire homogenate was filtered through a 100 µm strainer. Then, 5 mL of 50% iodixanol was added to create a 10 mL suspension of 25% iodixanol. The suspension was inverted to mix well before equally splitting into two 15 mL tubes. Iodixanol (40%, 2 mL) was layered gently at the bottom of the homogenate and centrifuged at 1000 × g for 12 minutes at 4 °C, with a swing bucket centrifuge (brand). The interphase nuclei were transferred to a new 15 mL tube and resuspended with 1:1 volume of 10 × wash buffer (10 mM Tris-HCl pH7.4, 10 mM NaCl, 3 mM MgCl2, 1% BSA, 0.1% Tween 20, 1 mM DTT, and 1 U/µL RNase inhibitor), followed by centrifuging at 500 g for 10 minutes at 4 oC. The supernatant was discarded and the pellet was resuspended in 800 µL diluted nuclei buffer (20× Nuclei Buffer from Chromium Next GEM Single Cell Multiome ATAC + Gene Expression Kit supplemented with 1 mM DTT, and 1 U/µL RNase inhibitor), before conducting FANS (fluorescence-activated nuclei sorting) using a BD FACS Aria III cell sorter (BD BioSciences, USA). GFPon nuclei were collected in diluted nuclei buffer and centrifuged at 500 × g for 7 minutes at 4 °C. Nuclei were adjusted to an appropriate concentration following the manufacturer’s protocol (10x Genomics, 1000285) with diluted nuclei buffer. The quality of nuclei was examined using an AxioImager Z1 upright microscope (Zeiss, Germany) at 40-fold magnification.
Single-nucleus RNA and ATAC sequencing
Transposition, nuclei isolation, and single-nucleus multiome libraries were prepared following the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression protocol CG000338 Rev F (10x Genomics, Pleasanton, CA). The generated snRNA and ATAC libraries were sequenced to a minimum depth of 25,000 and 35,000 mean paired-end reads per nuclei, respectively, using a Nextseq 500 sequencer (RNA: 28 – 10 – 10 – 90 bps (base pairs), ATAC: 50 – 8 – 8 using an Illumina Nextseq 500 sequencer (Illumina, Inc., USA) and 16 – 49 bps). Raw sequencing data were processed using the standard Cell Ranger ARC pipeline (version 7.0.0, 10x Genomics) for demultiplexing, mapping to the mm10 reference genome, barcode, and UMI (unique molecular identifier) counting, and generating the gene count matrix.
Joint processing of snRNA and snATACseq data
Analysis was performed in RStudio and R (version 3.6.1) using multiple packages: Seurat, Signac, DoubletFinder, ClusterProfiler, ggplot2. Sample quality was examined to ensure the nuclei conformed to the following criteria: based on gene expression data - nFeature_RNA < 10,000, nFeature_RNA > 200, and nCount_RNA > 500; and based on chromatin accessibility data - pct_reads_in_peaks > 15, blacklist_fraction <5, nucleosome_signal <4, and TSS.enrichment > 1. Doublets were estimated using DoubletFinder and removed from the analysis. The ATAC counts were normalized with TFIDF (term frequency-inverse document frequency), followed by dimensionality reduction with SVD (singular value decomposition). Clustering was performed using the second to tenth LSI (Latent Semantic Indexing) components. The gene expression counts were normalized whereby the expression value of each gene was divided by the total expression in each cell, multiplied by 10,000, and log-transformed. Highly variable genes across nuclei were selected and scaled. PCA (principal component analysis) was performed, and the top 30 principal components were selected for downstream clustering, as determined using an elbow plot. ATAC and RNA libraries for each sample were integrated using reciprocal LSI and MNN-CCA (Mutual Nearest Neighbor - Canonical Correlation Analysis). A WNN (weighted nearest neighbor) graph represented a weighted combination of RNA and ATAC assays with the first 60 and the second to tenth dimensions, respectively. Unbiased clustering of the RNA-ATAC integrated data at a resolution of 0.3 was used for cluster annotation. Major clusters were annotated using Seurat label transfer prediction scores with published single-cell RNA-seq clusters as reference45,46.
Differential expression and chromatin accessibility
Based on gene expression data, differential expression analysis was performed using the Seurat function FindMarkers and applying a likelihood-ratio test. Genes with Bonferroni-adjusted p-values of less than 0.05 were considered to be differentially expressed between Sun1sfGFP; ChAT-Cre; Mettl14floxed and littermate control. Differential chromatin accessibility analysis, based on chromatin accessibility data, was conducted using the Presto function wilcoxauc. In Presto, the p-value of the Wilcoxon rank sum test is computed based on Gaussian approximation and further adjusted by the Benjamini-Hochberg method. A peak is considered to be open in one condition and closed in the other if the adjusted p-value is less than 0.01, the logarithmic (base 2) fold-change is greater than 0.1, and the percentage of cells with a non-zero value in the first condition is greater than 5. Otherwise, the peak was denoted as neutral.
Peak annotation
Peak annotation, i.e., to determine if a peak is in the transcription start site (TSS), transcription termination site (TTS), Exon, 5′ UTR Exon, 3′ UTR Exon, intronic, or intergenic region was performed by HOMER using the annotatePeaks.pl command on the mm10 genome reference.
Identification of gene-to-peak linkage
Identification of gene-to-peak linkage was performed for each cell type using ArchR’s framework. First, to reduce the sparsity of chromatin data, aggregation of similar cells was performed using the k-nearest neighbor graph based on LSI reduction, and the cell aggregate-by-peak and cell aggregate-by-gene matrices were created and log-normalized. Aggregates with greater than 80% overlap with any other aggregates were filtered out. Then, all possible peak-to-gene pairs were selected if the peak was located within ± 250 kilobase pairs flanking the gene start. Finally, the Pearson correlation was calculated across all cell aggregates for each possible peak-gene pair. Peak-to-gene linkages were then defined as those with Pearson correlation coefficients no less than 0.45 and adjusted p-values (by Benjamini–Hochberg method) no greater than 0.0001.
H3K9me3 and γH2AX quantification
At the P120 stage, the signal of H3K9me3 or γH2AX was quantified (ImageJ) after training by machine learning (ilastik), and divided into the area of DAPI in each ChATon MN. Both the strength of the laser and the exposure time for H3K9me3 or γH2AX were fixed to facilitate quantification. In Figs. 7c, e, 10b, and Supplementary 13b, each point represents the quantified result for each ChATon MN, and those quantifications were counted from at least three mice of the same age and genotype.
Bulk RNA-seq
For bulk RNA-seq, RNA was extracted from ALS iPSC~MNs, and its quality was evaluated using a Bioanalyzer 2100 RNA pico kit. The cDNA libraries were prepared from the human iPSC~MNs according to the manufacturer’s instructions for the TruSeq Stranded mRNA library prep kit (Illumina). The concentration and size distribution of the completed libraries were determined using an Agilent Bioanalyzer 2100 DNA high-sensitivity kit and Qubit fluorometry (Invitrogen). Libraries were sequenced by following Illumina’s standard protocol using the Illumina NextSeq500 HighOutput kit V2.5. For the gene expression z score analysis (including published data revealed in Supplementary Fig. 1) by aligning raw RNA-seq data to the GRCh38 genome using the STAR aligner. Subsequently, gene expression was estimated using the GenCode v41 gene annotation file (gtf), focusing on gene types categorized as protein_coding and lncRNA. Genes originating from the mitochondrial genome (chrM) were also excluded from further analysis. The expression levels of each gene were then quantified using the salmon tool. Next, differential expression analysis was conducted using the edgeR package. Raw read count information was processed to identify genes showing significant changes in expression across different conditions. The filterByExpr function was applied to remove genes with low expression levels to ensure robust results. The remaining genes were subjected to TMM normalization and dispersion estimation methods to identify differentially expressed genes accurately. Subsequently, genes were categorized as upregulated or downregulated based on fold change values. The expression trends of the same genes across different samples were analyzed by calculating z-scores, allowing for the observation of consistent or divergent expression patterns among compared samples. This analysis pipeline enabled the identification of differentially expressed genes and provided insights into their expression trends across various conditions.
scAAV9 plasmid construction
To construct scAAV9-shFto plasmid, a 65 basepair short hairpin sequence targeting the fat mass and obesity-associated (Fto) gene (RNAi # TRCN0000183897) was subcloned into scAAV9-H1-CB-EGFP plasmid. A scAAV9-EGFP plasmid with the same backbone was provided by the AAV Core Facility in Academia Sinica. To verify the expression of the scAAV9-shFto construct, we used pLKO.1-shFto and pLKO.1-sh-Scramble (as a control group) and packaged them into lentivirus from the RNAi Core Facility of Academia Sinica. We transduced lentivirus (at a multiplicity of infection of 100) with 8 mg/mL protamine sulfate (Sigma-Aldrich) in a growth medium (DMEM) containing 20% fetal bovine serum. After 48 h of transduction, the medium was changed with the selection drug (8 µg/mL puromycin) for one week. We collected RNA/protein of puromycin-selected C2C12 cells, and levels of Fto were determined by qPCR/Western blotting, with m6A mRNA methylation levels being determined by m6A ELISA.
scAAV9 virus preparation and intrathecal injection
scAAV9-shFto and scAAV9-EGFP were packaged by the AAV Core Facility in Academia Sinica. For intrathecal injection, mice at P60 were anesthetized by isoflurane. The lumbar spine was exposed through a 1.5 cm window surgically cut on the back of the mice. We injected 20 µL of either scAAV9-EGFP or scAAV9-shFto (5 × 109 vg/µL) at a rate of 4 µL/minute into the groove between the L6 and S1 segments using a 27 G needle. A flick of the mouse’s tail indicated a successful injection.
Compound muscle action potential (CMAP)
Evoked CMAP of gastrocnemius muscle was measured on isoflurane-anesthetized mice at the indicated age (P60, P120, P140, P160). Before recording, hairs on the right hind limb and the lower back were shaved and completely removed using a depilatory cream. A reference recording electrode was then placed on the ankle of the right hind limb, and an active recording electrode was placed on the belly of the right gastrocnemius. To stimulate the sciatic nerve, a pair of stimulating anode and cathode needles were inserted near the sciatic nerve at the ipsilateral paraspinal site and the region of the proximal hind limb, respectively. A ground electrode was placed on the tail to minimize artifacts. To obtain maximal CMAP responses, we gradually increased the stimulus intensity from 3 mA and determined the supramaximal stimulation as ~120% of the stimulus intensity that no longer increased the response amplitude. The baseline-to-peak and peak-to-peak CMAP amplitudes evoked by supramaximal stimulation (mostly ~7 to 8 mA) were summed. We recorded at least 30 maximal CMAP responses for each mouse at each stage. The top five successful and maximal CMAP responses in each mouse were averaged and used for comparison between groups.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Requests for further information or resources should be directed to and will be filled by the lead authors, Y.-P.Y. and J.-A.C. All the sequencing data generated in this study have been deposited in the Gene Expression Omnibus repository under the GSE accession codes GSE290242, GSE290244, and GSE290245. Previously published human postmortem cortex RNA-seq and human iPSC~MNs are available under the accession codes GSE122649, GSE122650, GSE132972, and GSE173115. Transcriptome data from Answer ALS can be requested via the website: https://www.answerals.org/. Source data are provided with this paper.
Code availability
The codes and processed data are available on GitHub at https://github.com/jaclab-multiomic/Yen-et-al.-2025-Nat-Comm and figshare at https://figshare.com/projects/Yen-et-al_-2025-Nat-Comm/238025.
References
He, C. Grand challenge commentary: RNA epigenetics? Nat. Chem. Biol. 6, 863 (2010).
Li, X., Xiong, X. & Yi, C. Epitranscriptome sequencing technologies: decoding RNA modifications. Nat. Methods 14, 23 https://www.nature.com/articles/nmeth.4110#supplementary-information (2016).
Zhang, F., Ignatova, V. V., Ming, G. L. & Song, H. Advances in brain epitranscriptomics research and translational opportunities. Mol. Psychiatry https://doi.org/10.1038/s41380-023-02339-x (2023).
He, P. C. & He, C. m(6) A RNA methylation: from mechanisms to therapeutic potential. EMBO J. 40, e105977 (2021).
Liu, J. et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 10, 93–95 (2014).
Ping, X. L. et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Wang, X. et al. Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature 534, 575–578 (2016).
Ruzicka, K. et al. Identification of factors required for m(6) A mRNA methylation in Arabidopsis reveals a role for the conserved E3 ubiquitin ligase HAKAI. N. Phytol. 215, 157–172 (2017).
Wen, J. et al. Zc3h13 regulates nuclear RNA m(6)A methylation and mouse embryonic stem cell self-renewal. Mol. Cell 69, 1028–1038 e1026 (2018).
Yue, Y. et al. VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discov. 4, 10 (2018).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Mauer, J. et al. FTO controls reversible m(6)Am RNA methylation during snRNA biogenesis. Nat. Chem. Biol. 15, 340–347 (2019).
Wei, J. et al. Differential m(6)A, m(6)A(m), and m(1)A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol. Cell 71, 973–985 e975 (2018).
Deng, X. et al. RNA N6-methyladenosine modification in cancers: current status and perspectives. Cell Res. 28, 507–517 (2018).
Yen, Y. P. & Chen, J. A. The m(6)A epitranscriptome on neural development and degeneration. J. Biomed. Sci. 28, 40 (2021).
Yoon, K.-J. et al. Temporal control of mammalian cortical neurogenesis by m(6)A methylation. Cell 171, 877–889.e817 (2017).
Weng, Y.-L. et al. Epitranscriptomic m6A regulation of axon regeneration in the adult mammalian nervous system. Neuron 97, 313–325.e316 (2018).
Meyer, KateD. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Wu, Z. et al. m(6)A epitranscriptomic regulation of tissue homeostasis during primate aging. Nat. Aging 3, 705–721 (2023).
Yankova, E. et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature 593, 597–601 (2021).
Barbieri, I. et al. Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature 552, 126–131 (2017).
Vu, L. P. et al. The N(6)-methyladenosine (m(6)A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat. Med. 23, 1369–1376 (2017).
Batista, P. J. et al. m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Hsu, P. J., Shi, H. & He, C. Epitranscriptomic influences on development and disease. Genome Biol. 18, 197 (2017).
Jonkhout, N. et al. The RNA modification landscape in human disease. RNA 23, 1754–1769 (2017).
Chen, Y. G. et al. N6-methyladenosine modification controls circular RNA immunity. Mol. Cell 76, 96–109 e109 (2019).
Castro-Hernandez, R. et al. Conserved reduction of m(6)A RNA modifications during aging and neurodegeneration is linked to changes in synaptic transcripts. Proc. Natl Acad. Sci. USA 120, e2204933120 (2023).
Todd, T. W. & Petrucelli, L. Modelling amyotrophic lateral sclerosis in rodents. Nat. Rev. Neurosci. 23, 231–251 (2022).
Kim, G., Gautier, O., Tassoni-Tsuchida, E., Ma, X. R. & Gitler, A. D. ALS genetics: gains, losses, and implications for future therapies. Neuron 108, 822–842 (2020).
Mandrioli, J., Mediani, L., Alberti, S. & Carra, S. ALS and FTD: where RNA metabolism meets protein quality control. Semin. Cell Dev. Biol. 99, 183–192 (2020).
Droppelmann, C. A., Campos-Melo, D., Ishtiaq, M., Volkening, K. & Strong, M. J. RNA metabolism in ALS: when normal processes become pathological. Amyotroph. Lateral Scler. Frontotemporal Degener. 15, 321–336 (2014).
Chen, T. H. & Chen, J. A. Multifaceted roles of microRNAs: from motor neuron generation in embryos to degeneration in spinal muscular atrophy. Elife 8, https://doi.org/10.7554/eLife.50848 (2019).
McMillan, M. et al. RNA methylation influences TDP43 binding and disease pathogenesis in models of amyotrophic lateral sclerosis and frontotemporal dementia. Mol. Cell 83, 219–236.e217 (2023).
Li, Y. et al. Globally reduced N(6)-methyladenosine (m(6)A) in C9ORF72-ALS/FTD dysregulates RNA metabolism and contributes to neurodegeneration. Nat. Neurosci. 26, 1328–1338 (2023).
Thams, S. et al. A stem cell-based screening platform identifies compounds that desensitize motor neurons to endoplasmic reticulum stress. Mol. Ther. 27, 87–101 (2019).
Tung, Y. T. et al. Mir-17 approximately 92 confers motor neuron subtype differential resistance to ALS-associated degeneration. Cell Stem Cell 25, 193–209.e197 (2019).
Tung, Y. T. et al. Mir-17 approximately 92 governs motor neuron subtype survival by mediating nuclear PTEN. Cell Rep. 11, 1305–1318 (2015).
Xu, H. et al. m(6)A mRNA methylation is essential for oligodendrocyte maturation and CNS myelination. Neuron 105, 293–309.e295 (2020).
Patel, T. et al. Transcriptional dynamics of murine motor neuron maturation in vivo and in vitro. Nat. Commun. 13, 5427 (2022).
Liu, H., Begik, O. & Novoa, E. M. EpiNano: detection of m(6)A RNA modifications using oxford nanopore direct RNA sequencing. Methods Mol. Biol. 2298, 31–52 (2021).
Hendra, C. et al. Detection of m6A from direct RNA sequencing using a multiple instance learning framework. Nat. Methods 19, 1590–1598 (2022).
Spataro, R. et al. Mutations in ATP13A2 (PARK9) are associated with an amyotrophic lateral sclerosis-like phenotype, implicating this locus in further phenotypic expansion. Hum. Genomics 13, 19 (2019).
Blum, J. A. et al. Single-cell transcriptomic analysis of the adult mouse spinal cord reveals molecular diversity of autonomic and skeletal motor neurons. Nat. Neurosci. 24, 572–583 (2021).
Russ, D. E. et al. A harmonized atlas of mouse spinal cord cell types and their spatial organization. Nat Commun. 12, ARTN 5722 https://doi.org/10.1038/s41467-021-25125-1 (2021).
Bezprozvanny, I. Calcium signaling and neurodegenerative diseases. Trends Mol. Med. 15, 89–100 (2009).
Hung, S. T. et al. PIKFYVE inhibition mitigates disease in models of diverse forms of ALS. Cell 186, 786–802.e728 (2023).
Price, B. D. & D’Andrea, A. D. Chromatin remodeling at DNA double-strand breaks. Cell 152, 1344–1354 (2013).
Huang, Y. et al. Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell 35, 677–691 e610 (2019).
Cerneckis, J., Ming, G. L., Song, H., He, C. & Shi, Y. The rise of epitranscriptomics: recent developments and future directions. Trends Pharmacol. Sci. 45, 24–38 (2024).
Chen, J. A. & Wichterle, H. Apoptosis of limb innervating motor neurons and erosion of motor pool identity upon lineage specific dicer inactivation. Front. Neurosci. 6, 69 (2012).
Ma, X. R. et al. TDP-43 represses cryptic exon inclusion in the FTD-ALS gene UNC13A. Nature 603, 124–130 (2022).
Hideyama, T. et al. Induced loss of ADAR2 engenders slow death of motor neurons from Q/R site-unedited GluR2. J. Neurosci. 30, 11917–11925 (2010).
Beaulieu, J. M., Jacomy, H. & Julien, J. P. Formation of intermediate filament protein aggregates with disparate effects in two transgenic mouse models lacking the neurofilament light subunit. J. Neurosci. 20, 5321–5328 (2000).
Mishra, P. S. et al. Transmission of ALS pathogenesis by the cerebrospinal fluid. Acta Neuropathol. Commun. 8, 65 (2020).
Wells, T. L., Myles, J. R. & Akay, T. C-boutons and their influence on amyotrophic lateral sclerosis disease progression. J. Neurosci. 41, 8088–8101 (2021).
Montanana-Rosell, R. et al. Spinal inhibitory neurons degenerate before motor neurons and excitatory neurons in a mouse model of ALS. Sci. Adv. 10, eadk3229 (2024).
Nedialkova, D. D. & Leidel, S. A. Optimization of codon translation rates via tRNA modifications maintains proteome integrity. Cell 161, 1606–1618 (2015).
Helm, M. & Alfonzo, J. D. Posttranscriptional RNA modifications: playing metabolic games in a cell’s chemical Legoland. Chem. Biol. 21, 174–185 (2014).
Lence, T. et al. m(6)A modulates neuronal functions and sex determination in Drosophila. Nature 540, 242–247 (2016).
Tam, O. H. et al. Postmortem cortex samples identify distinct molecular subtypes of ALS: retrotransposon activation, oxidative stress, and activated glia. Cell Rep. 29, 1164–1177.e1165 (2019).
La Cognata, V., Gentile, G., Aronica, E. & Cavallaro, S. Splicing players are differently expressed in sporadic amyotrophic lateral sclerosis molecular clusters and brain regions. Cells 9 https://doi.org/10.3390/cells9010159 (2020).
Wei, J. et al. FTO mediates LINE1 m(6)A demethylation and chromatin regulation in mESCs and mouse development. Science 376, 968–973 (2022).
Ruud, J. et al. The fat mass and obesity-associated protein (FTO) regulates locomotor responses to novelty via D2R medium spiny neurons. Cell Rep. 27, 3182–3198.e3189 (2019).
Hess, M. E. et al. The fat mass and obesity associated gene (Fto) regulates activity of the dopaminergic midbrain circuitry. Nat. Neurosci. 16, 1042–1048 (2013).
Maury, Y. et al. Combinatorial analysis of developmental cues efficiently converts human pluripotent stem cells into multiple neuronal subtypes. Nat. Biotechnol. 33, 89–96 (2015).
Maor-Nof, M. et al. p53 is a central regulator driving neurodegeneration caused by C9orf72 poly(PR). Cell 184, 689–708 e620 (2021).
Kraemer, B. R. et al. A role for the p75 neurotrophin receptor in axonal degeneration and apoptosis induced by oxidative stress. J. Biol. Chem. 289, 21205–21216 (2014).
Scott, S. et al. Design, power, and interpretation of studies in the standard murine model of ALS. Amyotroph. Lateral Scler. 9, 4–15 (2008).
Chang, S. H., Su, Y. C., Chang, M. E. & Chen, J. A. MicroRNAs mediate precise control of spinal interneuron populations to exert delicate sensory-to-motor outputs. Elife 10, ARTN e63768 https://doi.org/10.7554/eLife.63768 (2021).
Acknowledgements
We appreciate Answer ALS for sharing the ALS patient’s omics data. We thank the genomics core facility at the Institute of Molecular Biology, Academia Sinica for the Illumina sequencing and nanopore direct RNA sequencing. The flow cytometry core facility and imaging core at the Institute of Molecular Biology, Academia Sinica for cell sorting service and confocal imaging. The Olig2-Cre line was a kind gift from Prof. Tom Jessell (University of Columbia, USA). We particularly thank Prof. Hongjun Song and Prof. Guo-Li Ming (University of Pennsylvania) for sharing details of the initial Mettl14floxed mouse strain. We thank the National RNAi Core Facility at Academia Sinica in Taiwan for providing Fto-shRNA-related plasmids and technical support for LV vector packaging. We also acknowledge the Taiwan Mouse Clinic and the AAV Core Facility of IBMS Academia Sinica for their technical support with mouse behavioral analyses (open field, rotarod, grip strength), X-ray imaging experiments, and scAAV9 vector packaging. We thank the Human Disease iPSC Service Consortium for iPSC generation and technical support. Y.-P.Y. is supported by an Academia Sinica Postdoctoral Fellowship (23-5g). The consortium is funded by the Ministry of Science and Technology (MOST) (MOST 110-2740-B-001-003). This work is funded by Academia Sinica (AS-GCP-113-L02 and AS-BRPT-113-01), the National Science & Technology Council (112-2326-B-001 -001 -), and the National Health Research Institutes (NHRI-EX113-11330NI).
Author information
Authors and Affiliations
Contributions
Conceptualization, J.-A.C. and Y.-P.Y.; Methodology, E.-S.L., T.-H.L.,C.-C.W., G.-L.H., F.-Y.H., M.C., and Y.-P.Y., Software and Formal Analysis, E.-S.L., T.-H.L.,C.-C.W., G.-L.H., F.-Y.H., Z.-D.Y., C.-Y.H., W.Z., J.-H.H., Q.N., and Y.-P.Y.; Investigation, T.-H.L.,C.-C.W., G.-L.H., F.-Y.H., M.C., and Y.-P.Y., and J.-A.C.; Resources, Z.Z. and C.H.; Writing – Original Draft, J.-A.C. and Y.-P.Y.; Writing – Review & Editing, J.-A.C., Y.-P.Y., and C.H.; Funding Acquisition, J.-A.C. and Y.-P.Y.; Supervision, J.-A.C. and Y.-P.Y.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests. J.-A.C. and Y.-P.Y. declare a PCT and a US patent application related to this work.
Peer review
Peer review information
Nature Communications thanks Ki-Jun Yoon, who co-reviewed with Sung-Min Lee, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yen, YP., Lung, TH., Liau, E.S. et al. The motor neuron m6A repertoire governs neuronal homeostasis and FTO inhibition mitigates ALS symptom manifestation. Nat Commun 16, 4063 (2025). https://doi.org/10.1038/s41467-025-59117-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59117-2