Abstract
Genomics can inform both tissue-of-origin (TOO) and precision treatments for patients with cancer of unknown primary (CUP). Here, we use whole genome and transcriptome sequencing (WGTS) for 72 patients and show diagnostic superiority of WGTS over panel testing (386–523 genes) in 71 paired cases. WGTS detects all reportable DNA features found by panel as well as additional mutations of diagnostic or therapeutic relevance in 76% of cases. Curated WGTS features and a CUP prediction algorithm (CUPPA) trained on WGTS data of known cancer types informs TOO in 71% of cases otherwise undiagnosed by clinicopathology review. WGTS informs treatments for 79% of patients, compared to 59% by panel testing. Finally, WGS of cell-free DNA (cfDNA) from patients with a high cfDNA tumour fraction (>7%), enables high-likelihood CUPPA predictions in 41% of cases. WGTS is therefore superior to panel testing, broadens treatment options, and is feasible using routine pathology samples and cfDNA.
Similar content being viewed by others
Introduction
Cancers of unknown primary (CUP) are metastatic tumours for which a tissue-of-origin (TOO) cannot be identified after standardised diagnostic investigations1. Although improvements in cancer diagnostics have reduced the incidence of CUP in recent years, they still account for 1–3% of all new cancer diagnoses2. While a minority (~20%) of CUP patients have a favourable outcome corresponding to treatment-responsive cancer types, most CUP patients have an unfavourable prognosis and empirical chemotherapy is generally ineffective1.
DNA sequencing can identify therapeutically actionable mutations in a third or more of CUP tumours3. The CUPISCO clinical trial showed that patients with unfavourable CUP who received molecular or immune-checkpoint targeting therapies based on tumour profiling data had improved progression-free survival following three cycles of induction chemotherapy when compared to platinum-chemotherapy treated controls4. Therefore, applying molecularly targeted treatments in CUP patients irrespective of the cancer’s origin is plausible, but despite this, targeted treatments can still show variable efficacy between cancer types5, while drug access is also often restricted to specific cancer types. Genomic tests that can resolve the primary cancer diagnosis and direct targeted treatments are therefore needed.
Genomic profiling has been used to predict a primary cancer TOO in CUP, with a number of studies utilising RNA expression6,7,8, DNA methylation9 or DNA mutations10,11,12,13. The application of DNA mutation data for TOO diagnosis is attractive as the same data also informs precision treatments. Some gene-specific mutations are enriched among certain cancer types, while the detection of mutational signatures, associated with ultraviolet light and tobacco smoking, can also provide additional diagnostic evidence14,15,16. Indeed, the detection of diagnostic features using panel sequencing was shown to assist in resolving a likely TOO in a third of CUP patients17. Applying whole-genome sequencing (WGS) is expected to have superior diagnostic yield as the number of genome-wide features captured is much greater. One particularly promising application of WGS is the use of machine learning to predict TOO by using a combination of genome-wide driver and passenger mutation features, many of which cannot be reliably detected using panel sequencing10,11,18. Despite this, no systematic comparison of WGS to panel testing has been described in CUP patients to date, and when using formalin-fixed paraffin-embedded (FFPE) samples11,19.
Here, we show the feasibility of combining whole-genome and transcriptome sequencing (WGTS) in a retrospective series of 73 CUP tumours from 72 patients recruited to an Australian CUP clinical genomics study. For 59/73 (81%) tumours, we utilise FFPE samples for WGTS. In 71/73 (97%) cases, we also perform comprehensive cancer panel testing, enabling direct comparison of mutation features between platforms. Furthermore, a CUP prediction algorithm (CUPPA)11 is applied to the WGTS data to compare algorithmic TOO prediction against a pathologist’s favoured diagnosis informed by individual genome features curated from the data. Finally, given tissue availability for molecular profiling is an issue for many CUP patients, we evaluate WGS of blood plasma cell-free DNA (cfDNA) from CUP patients where patients have high cfDNA tumour fraction.
Results
Feasibility of clinical WGTS applied to a national CUP programme
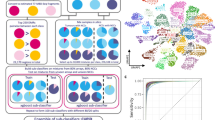
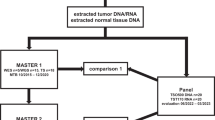
To assess the feasibility and diagnostic yield of WGTS we initially selected 75 tumours from 74 CUP patients who were recruited to a national genomics study. (Fig. 1a). One case (CUP 1209) had two metastases sequenced, which were later identified as independent tumours based on their discordant molecular profiles. (Supplementary Fig. 1, Supplementary Data 1). WGS data was very poor quality for two tumours and these were excluded from further analysis, leaving WGTS data for a total of 73 tumours from 72 patients. (Fig. 1a). DNA panel testing was done prior to attempting WGTS for 71/73 (97%) tumours. A custom 386-gene cancer panel (CCP17) was used in 24/71 (34%) cases where both tumour and matched germline DNA were sequenced. A commercial 523-gene tumour-only DNA panel with RNA-seq capture for fusion detection (Illumina TSO500) was used for the remaining 47/71 (66%) cases (Fig. 1a).
a Flow chart of CUP cases profiled in this study. Fresh or FFPE tumour samples were either diagnostic core needle biopsies or surgical resections accessed from pathology archives. Seventy-five samples from 74 CUP patients had WGTS performed, where 73 WGS and 69 WTS were successful. Seventy-one of these had matched panel sequencing: 24 using a custom cancer panel and 47 a commercial cancer panel. Created in BioRender. (License: Tothill, R. (2025) https://BioRender.com/h69j379). b Seventy-five tumours were sampled from a range of anatomical sites for molecular profiling in 74 CUP patients. Non-recurrent sites were combined as ‘other less common sites’. Created in BioRender. (License: Tothill, R. (2025) https://BioRender.com/r31l964). c Fraction of CUP tumours clinicopathology-resolved or clinicopathology-unresolved pre-genomic testing, after a centralised pathology review. Clinicopathology-unresolved CUPs were categorised based on a modified MSKCC OncoTree classification. Source data are provided as a Source Data file.
Most tumour samples (61/75, 81%) were FFPE tissues (Fig. 1a and Supplementary Data 2) obtained by core needle biopsy (71/75, 95%), with the most common biopsy sites being lymph node or liver representing 22/75 (29%) and 11/75 (15%) of cases, respectively (Fig. 1b and Supplementary Data 1). Fresh tissues were sequenced in 14/75 (19%) cases, and paired fresh and FFPE was available in three cases. Whole transcriptome data could not be generated in four cases due to poor RNA quality or yield.
FFPE WGS libraries had shorter fragment lengths (FFPE: median 437 base pairs (bp), vs. Fresh: median 618 bp) and higher sequence duplication rates (FFPE: median 25%, vs Fresh: median 7%), necessitating additional sequencing to achieve sufficient sequence coverage (Supplementary Fig. 2a–c and Supplementary Data 2). Consistent with a prior study20, copy-number variants (CNV) were noisier for FFPE samples, as indicated by a higher number of samples with a copy-number noise warning (PURPLE tool, see ‘Methods’) (Pearson’s chi-square test, *P < 0.0001, Supplementary Fig. 2d and Supplementary Data 2). However, the ability to histologically review and then dissect tumour regions from FFPE samples was advantagous and may have accounted for the higher average tumour fraction estimated by WGS among FFPE cases compared to fresh tissues (Supplementary Fig. 2e, f and Supplementary Data 2).
According to ESMO criteria, 61/74 (82%) CUP patients had an unfavourable outcome profile (Supplementary data 1). A summary of clinical characteristics for the patient cohort is shown in Table 1. Centralised clinical and pathology review in the setting of a specialised cancer hospital with a dedicated CUP clinic21 enabled a single TOO diagnosis in 16/75 (21%) cases before any genomics data was considered, and these cases were termed clinicopathology-resolved (Fig. 1c). The remainder of cases were otherwise clinicopathology-unresolved and were assigned a modified Memorial Sloan Kettering Cancer Center (MSKCC) OncoTree classification using CK7, CK20 and CDX2 immunohistochemistry to help define additional CUP subsets, as previously described17. The most common histological subtypes were adenocarcinoma (Ad) and poorly-differentiated carcinoma (PDC), representing 26/75 (35%) and 24/75 (32%) of tumours, respectively (Fig. 1c). Neuroendocrine carcinoma (NEC, 3/75, 4%) and squamous cell carcinoma of unknown origin (SCC, 3/75, 4%) were minor subsets. (Fig. 1c, Supplementary data 1).
WGTS is superior to panel sequencing for the detection of reportable features
The availability of paired panel and WGTS data enabled a direct comparison of clinically reportable features. Small somatic variants (single-nucleotide variants, SNVs; Insertions/deletions, Indels), CNVs and structural variants (SVs) were curated independently for WGTS and panel sequencing platforms. As curation protocols varied between platforms and also over time, variant annotation was retrospectively and centrally reviewed to enable direct comparison of therapeutic and diagnostic features according to standardised AMP/ASCO/CAP consensus guidelines (see ‘Methods’) (Fig. 2a).
a Total number of therapeutic and diagnostic variants in CUP cases detected in both matched panel and WGTS or by WGTS only (n = 73 tumours). Variants are also categorised by type (SNV/Indel, SV and CNV). b Schematic showing comparison of SNV-96 SBS mutational signatures where a dominant SBS signature was found by WGS and there was matched panel data (COSMICv2, n = 29/73). Number of cases with dominant SBS signatures(SBS4, SBS7, SBS3 and SBS6) that are true positive, false positive or false negative as determined by panel using matched WGS data as a known truthare shown in the histogram. Asterisks (*) represent cases with greater than 50 somatic SNVs that were detected by panel. TMB (muts/Mb), HRD (HRDetect and CHORD tools) and MSI (Indels/Mb) scores from WGS data are presented alongside for each case. Source data are provided as a Source Data file.
With respect to curated SNVs and small Indels, there was 89% (181/203) concordance between paired panel and WGTS data, but all small somatic variants detected by panel sequencing were also found by WGS (Supplementary Data 3). SNV/Indel somatic alterations missed by panel testing were most likely explained by the absence of DNA capture probes (using the CCP panel) for these regions of interest, such as the TERT promoter. Despite differences in the design of the two panels, only six variants detected by WGTS were missed by the smaller CCP panel and these could have theoretically been captured using the TSO500 panel. Panel design was therefore not deemed a major factor in considering the overall differences in diagnostic yield between panel and WGTS data. Detection of SVs and CNVs was a major advantage of WGTS over panel, with most CNVs (37/60, 62%) and nearly all SVs (53/54, 98%) detected only by WGTS (Fig. 2a, Supplementary Data 3).
Although discordancy between WGS and panel can be due to sampling and tumour heterogeneity, the same tumour nucleic acid extracts were used for panel and WGTS in 66/71 (93%) cases. Analysis of WGTS data generated from independent fresh and FFPE tissues in three cases showed differences in the molecular profile between samples where tumour heterogeneity was likely a factor, but the reportable therapeutic and diagnostic features were largely concordant, with only one additional SV of clinical interest found in one tumour of a paired case (Supplementary Fig. 2g–i).
Considering all available sequence data, WGTS identified 90 therapeutic and 105 diagnostic variants in 66/73 (90%) tumours, which was superior to panel testing, detecting 59 therapeutic and 79 diagnostic variants in 56/71 (79%) of tumours. Considering only patients with matched data, WGTS detected additional reportable variants in 54/71 (76%), where 25/71 (35%) were of known therapeutic/diagnostic relevance to a potential cancer type (Fig. 2a).
WGS improves detection of diagnostic and therapeutic mutation signatures
Somatic single-base substitution (SBS) signatures can be of high diagnostic and therapeutic value22. With respect to diagnostic signatures, WGS detected a dominant SBS4 (tobacco smoking) in 11/73 (15%) and SBS7 (ultraviolet light) in 2/73 (3%) cases, consistent with lung and skin cancer aetiology, respectively. (Fig. 2b). Comparatively, while panel sequencing could detect SBS7 in the two presumed skin-cancer CUPs, it could only detect SBS4 in 8/11 (73%) presumed lung-CUPs. (Fig. 2b). Furthermore, as there was propensity for false-positive detection of dominant mutational signatures using a panel in tumours with a low mutation burden, we set a minimum threshold of 50 mutations, which further reduced the sensitivity for SBS4 detection by panel to only 7/11 (64%) cases.
Homologous recombination deficiency (HRD) can predict response to platinum chemotherapy or Poly-ADP-ribose polymerase (PARP) inhibition14,23,24. Two algorithmic HRD-prediction tools (CHORD25 and HRDetect26) were applied to WGS data. CHORD/HRDetect-positivity was detected in six cases (Fig. 2b). Importantly, CHORD/HRDetect-positivity corresponded with somatic or germline mutations in HRD-related genes in four cases, and two cases (CUP 1092 and CUP 1208) were also deemed HR-deficient in the absence of HR-gene mutations. Detection of HRD would have made these patients eligible for a clinical trial open at the time of retrospective analysis (such as IMPARP-HRD: NCT04985721) (Fig. 2b). HR-Detect and CHORD have not been validated using cancer panel data, therefore, this was not attempted. Dominant SBS3 is also associated with HR-deficient cancers14,16 and was detected in 8/73 (11%) CUPs by WGS and three cases by panel, although this was reduced to only one case by panel when applying a minimum threshold for the number of mutations detected (≥50 mutations). Furthermore, it is known that SBS3 lacks specificity for predicting HRD when used in isolation27, and this was clearly demonstrated for two CUP cases (CUP 1227 and CUP 2016) that had dominant SBS3 but were otherwise deemed HR-proficient by CHORD/HRDetect.
Microsatellite instability (MSI) and high tumour mutational burden (TMB) can predict clinical response to immune checkpoint blocking antibodies28,29,30. An MSI caller was applied to WGS and TSO500 panel data (see ‘Methods’), but not CCP panel given it was not designed for MSI detection (Fig. 2b). Three of 73 CUP cases (4%) were MSI-High by WGS (CUPs 1085, 1137, W021) and harboured additional supporting MSI features: e.g. MLH1 protein loss of expression in tumour cells by immunohistochemistry and high TMB (>10 mut/Mb)14 (Fig. 2b). Interestingly, in these three MSI-High CUPs, the TSO500 MSI score was below the threshold for calling MSI-High. However, a dominant SBS6 signature, which is a known feature of MSI, was detected by both panel and WGS, but is not typically used for reporting MSI, and again a dominant SBS6 signature also had a propensity to be false by panel testing (e.g. CUP D007 and CUP 1226) when less than 50 somatic mutations were detected (Fig. 2b).
Overall, TMB was highly correlated between WGS and with panel sequencing (CCP vs WGS, Spearman’s r = 0.82; TSO500 vs WGS, r = 0.65) (Supplementary Fig. 3a, Supplementary Data 4). Panel sequencing generated a higher TMB estimate in 54/71 (76%) of cases (Supplementary Fig. 3b, Supplementary Data 4). Considering the TMB threshold for FDA-approved immune checkpoint treatments (≥10 muts/Mb), 17/71 (24%) of tumours had high TMB by panel, although only 12 of these were deemed TMB-high by WGS.
As expected, WGS was therefore the more sensitive test for the detection of genome-wide mutational signatures of diagnostic and therapeutic importance, which included tobacco smoking, HRD and MSI signatures, whereas a high TMB was more often reported by panel sequencing.
Mutation features can inform a clinicopathological diagnosis
We previously showed that genomic features enriched in specific cancer types, including somatic driver mutations, mutational signatures and oncoviral DNA can be used to support a TOO diagnosis when used in conjunction with a clinicopathology review17. Using the same approach in our current CUP series, panel testing enabled 19/56 (34%) cases initially clinicopathology-unresolved to subsequently be assigned a single TOO (Fig. 3a, b). Comparatively, WGTS detected additional features to inform TOO in 31/58 (53.4%) cases. The two most recurrent diagnoses were non-small cell lung cancer (NSCLC) (n = 11) and cholangiocarcinoma (n = 6) with smaller numbers of breast (n = 2) and kidney cancers (n = 2) as well as single examples across a range of other cancer types (Fig. 3b, c). Furthermore, a narrowed diagnostic differential was possible for 7/58 (12%) CUP cases. (Fig 3bii). For example, detection of HPV16/18 (HPV) indicated a likely primary site of either anogenital or head and neck regions, while driver mutations in genes enriched in only some cancers such as KRAS may have also informed a reduced differential (Supplementary Fig. 4). However, there was insufficient molecular evidence in such cases to resolve to a single TOO diagnosis.
a Number of clinicopathology-unresolved CUP tumours that had a genomics-informed change in diagnosis due to panel or additional WGTS features. (n = 58). b Bubble chart showing clinicopathology-unresolved CUPs categorised by MSKCC OncoTree cancer classification before and after panel and WGTS. (n = 58). Post-sequencing, tumours are grouped as: (i) putative diagnosis (n = 31), (ii) narrowed anatomical differential (n = 7), or (iii) remains unresolved (n = 20) and are colour coded by the assay that changed their diagnosis. c Schematic of all putatively diagnosed clinicopathology-unresolved CUPs grouped by favoured cancer type showing useful diagnostic molecular features, including mutational signatures and viral DNA, in addition to immunohistochemical features. (n = 31). Source data are provided as a Source Data file.
Algorithmic TOO prediction complements data curation and clinicopathology
CUPPA uses orthogonal DNA features from WGS to classify unknown tumours into 36 defined cancer classes11. For the current study, we extended CUPPA to include WTS data, enabling independent RNA classification as well as combined ‘DNA + RNA’ predictions (Fig. 4a and see ‘Methods’). We tested the accuracy of the CUPPA DNA + RNA by leave-one-out cross-validation (LOOCV) using a training dataset of 6106 primary and metastatic tumours of known origin (PCAWG and HMF datasets) (Supplementary Fig. 5a–c). The CUPPA DNA + RNA classifier had a LOOCV-accuracy of 91.3%, which was superior to using DNA-only or RNA-only classifiers that had LOOCV-accuracies of 86.6% and 82.7%, respectively. The accuracy of the CUPPA DNA + RNA classifier improved to 98.2% by setting a high-likelihood threshold (≥0.8), with high-likelihood predictions representing 83.5% of all training set samples in LOOCV (Supplementary Fig. 5b, c).
a Schematic representation of CUPPA that classifies CUP samples toward one of 36 defined cancer types by using a large reference dataset of known cancers. CUPPA (version 1.4) calculates five orthogonal variant feature scores for each class: three that are combined into an overall DNA score and two that combine into an overall RNA score. A combined DNA + RNA score is used for tissue samples when tumour fraction determined by WGS is ≥30%, otherwise, DNA-only classification is used. Created in BioRender. (License: Tothill, R. (2025) https://BioRender.com/w69c793). b Confusion matrix of CUPPA prediction scores against a pathologist’s favoured diagnosis, limited to CUPPA classes, colour coded by high (≥0.8) or low (<0.8) likelihood, where out of scope and yet unresolved tumours are separate categories. (n = 73). c Box plot showing CUPPA prediction scores using classification toward a single site TOO prediction, categorised by concordance with a genomics-informed, but CUPPA-blinded, pathology review (concordant n = 35, discordant n = 5). Cases with a favoured origin not represented in CUPPA training data were separated into a third group: out of scope cancer (n = 6). Whiskers indicate the minima and maxima, the centre lines represent the median, the box upper and lower bounds represent the 75th and 25th percentile, respectively. d Box plot of unresolved CUPs (n = 27) and schematic of ten of these CUP samples that were resolved with CUPPA and their high-likelihood (DNA + RNA or DNA only) predicted CUPPA classes. Whiskers indicate the minima and maxima, the centre lines represent the median, the box upper and lower bounds represent the 75th and 25th percentile, respectively. Bold tumour types in the pathologist differential are concordant with the CUPPA prediction. e Sankey plot of all CUP tumours (n = 56/73) with a resolved or putative diagnosis flowing toward the method by which they were resolved and cancer type. Source data are provided as a Source Data file.
We next applied CUPPA to all WGTS data from our CUP patient series. WTS data was unavailable in four cases, therefore, the CUPPA DNA-only classifier was used instead. Despite tumour fraction not greatly affecting the accuracy of CUPPA DNA or DNA + RNA classifiers during LOOCV (Supplementary Fig. 5d, e), we excluded the use of RNA data in classifying CUPs that had <30% tumour fraction, as including RNA data for these cases reduced the number of high-likelihood classifications (supplementary Fig. 5f, g). Importantly, we observed DNA-only classification was superior to RNA-only classification among lung-CUPs with respect to the proportion of high-confidence predictions made (Supplementary Fig. 6). These observations are similar to those made in our previous study using a Nanostring gene-expression classifier, where poor RNA classication for many lung-CUPs suggested these tumours have an atypical transcriptional profile17. High likelihood CUPPA predictions were made for 10/15 (67%) clinicopathology-resolved CUP tumours and 29/58 (50%) clinicopathology-unresolved CUPs, or 39/73 (53%) overall (Fig. 4b, c and Supplementary Fig. 6a, c). Again, the two most recurrent high-likelihood cancer type predictions were NSCLC (7) and bile duct/gall bladder, which includes cholangiocarcinoma (6). CUPPA predictions were mostly consistent with the pathologist’s favoured TOO based on genomics-informed pathology review, which was done blinded to CUPPA results (Fig. 4b, c). Importantly, all cases with high-likelihood CUPPA predictions were concordant with genomics-informed pathology review, indicating the CUPPA algorithm had high specificity (Fig. 4c and Supplementary Fig. 6c, d). Conversely, among 17 low-likelihood CUPPA predictions (<0.8), six were still concordant with pathology review, while 11 were discordant. Six of the 11 discordant CUPPA predictions were thought to be rare cancers not represented in the CUPPA training data set (e.g. CUP 1174, a malignant melanotic nerve sheath tumour) (Fig. 4c and Supplementary Data 1). Ten CUP tumours that had high-likelihood CUPPA predictions could not be conclusively resolved by genomics-informed pathology review. These included a spectrum of cancer type classifications to bile duct/gall bladder (2), breast (2), pancreas (2) and liver cancer (2) and these CUPPA predictions were all within the pathologist’s favoured diagnostic differential and therefore plausibly correct (Fig. 4b, d).
Considering combined results from all high-likelihood CUPPA predictions as well as independent genomics-informed pathology review, a total of 41/58 (71%) clinicopathology-unresolved cases could be assigned a likely TOO, compared to only 19/56 (34%) by panel testing and genomics-informed review (Fig. 4e).
Potential to improve treatment options using WGTS
We next retrospectively assessed potential treatment options for all patients considering therapeutically actionable mutations detected and predicted TOO (Fig. 5a). Treatment options were assessed for the entire cohort at single time point based on open phase I–II clinical trials in Australia and approved standard-of-care (SOC) treatments (Fig. 5b). Importantly, at the time of our analysis a TOO diagnosis would have been a requisite for drug access in many cases. As CUP 1209 had two synchronous primary tumours (Mesothelioma and Ad CK7 + CK20−), this patient would not have been considered for a targeted therapy, and was therefore excluded from analysis.
a Sankey plot showing all therapeutic features captured in matched panel and WGTS or WGTS only assays (n = 73 CUP tumours from 72 patients), separated by variant type and gene or genome wide feature, flowing to eligible inhibitor (drug class and name). b Sankey plot of 72 CUP patients showing the number and proportion (% of total) that had one or more therapeutic features detected, whether a putative TOO was assigned and whether a SOC treatment and/or clinical trial could be considered to gain access to treatment. Source data are provided as a Source Data file.
WGTS identified one or more therapeutic features in 53/72 (74%) patients, while a putative diagnosis was found in a partially overlapping group. (Fig. 5b). The combined diagnostic and treatment-related information would have directed SOC treatment for 51/72 (71%) patients and/or consideration for clinical trial in 39/72 (54%). (Fig. 5b and Supplementary Data 5). In contrast, panel sequencing detected therapeutic features in 49/71 (69%) but fewer patients would have had access to a SOC therapy (30/71, 42%) or clinical trials (29/71, 41%) (Supplementary Fig. 7 and Supplementary Data 4).
Importantly, resolving TOO diagnosis alone would have provided access to SOC immunotherapy in 31/72 (43%) patients, according to Australian treatment guidelines for metastatic NSCLC (11), cholangiocarcinoma (9), cutaneous SCC (1), Merkel cell carcinoma (1), colorectal carcinoma (1), clear-cell renal cell carcinoma (2), unresectable mesothelioma (1), hepatocellular carcinoma (2), triple negative breast cancer (2) and cervical cancer (1). (Fig. 5b and Supplementary Data 5). These results highlight the growing use of immunotherapy for several cancer types but where a TOO diagnosis is still necessary for drug access.
In summary, WGTS would have directed a potential treatment option for 79% of patients considering either SOC or clinical trial, compared to 59% of patients if considering only the panel data.
Assessing the utility of cfDNA-WGS for TOO prediction
To explore the application of blood plasma cfDNA as a potential tissue substitute for WGS testing, we collected blood from 144 CUP patients (Fig. 6a). Of these cases, 76/144 (52.8%) yielded minimum amount of cfDNA (20 ng) to generate libraries suitable for WGS. (Fig. 6a).
a Flow chart diagram showing the workflow for WGS of cfDNA from CUP patients. Created in BioRender. (Licence: Tothill, R. (2025) https://BioRender.com/v92f950.) b CfDNA yield per mL of blood plasma and ctDNA tumour fraction estimates calculated by the ichorCNA method across 76 patients. Dotted lines demarcate median tumour fraction at 7% and cfDNA content of 18.3 ng, respectively. c Bar graph showing distribution of unique or common SNVs in eight cases with matched ctDNA and tissue WGS. d SNV-96 mutational signatures (COSMICv2) across 8 cases with matched ctDNA and tissue WGS. Tumour fraction (ichorCNA estimated), TMB, and the presence or absence of a dominant signature (greater than 20% abundance) or diagnostic/therapeutically relevant signature are shown alongside for each sample. e Schematic of the 22 CUP-cfDNA cases showing immunohistochemistry profiles of matched biopsied tissues alongside known genomic features from the tumour that were detected in cfDNA-WGS data. Percentage known driver variant allele frequencies (VAF) and ichorCNA estimated ctDNA fraction for each sample are shown alongside. CUPPA predictions from cfDNA-WGS data are shown alongside for each case, as well as whether tissue and cfDNA CUPPA DNA-only predictions were concordant. Source data are provided as a Source Data file.
We first applied low-coverage whole genome sequencing (lcWGS) to determine the relative fraction of circulating tumour DNA (ctDNA) in total blood plasma cfDNA in individual samples (0.6x coverage, ichorCNA method31). The estimated ctDNA fraction across the cohort ranged from 0-47%, and a median of 3.75% across the 76 samples (Fig. 6b). The median yield of cfDNA from these CUP cases was 17.3 ng/mL (range 3.6–834 ng/mL) (Fig. 6b).
Twenty-two CUP cfDNA cases with ≥7% ctDNA were subjected to deeper sequencing (median 118x coverage). (Fig. 6a). Of these cases, 8/22 (36.3%) had matched tissue-based WGS data available, while 14/22 (63.6%) had cancer panel data available. We first processed the cfDNA-WGS data through our standard clinical reporting pipeline to compare SNV detection and the similarity of SBS mutational signatures (Fig. 6c, d). The average percentage of SNVs commonly detected in paired tissue and cfDNA WGS data was 39.3% (range 4.9-68.4%). Importantly, for the cfDNA-WGS analysed samples with matched CCP or TSO500 panel data (14/22), we identified the same tissue-reported driver mutations, in all of these cases, confirming the ctDNA had originated from same tumour and that variant detection was sensitive using the cfDNA-WGS data (Supplementary Data 6). The same dominant SBS signatures were also found in matched cfDNA and tissue WGS data for 6/8 (75%) cases, including some signatures of diagnostic utility. (Fig. 6d). For example, dominant SBS4 (tobacco smoking) was identified in a suspected lung-CUP case (CUP 1198). We also detected SBS17 in CUP 1201, a resolved oesophageal cancer, and this is a signature with unknown aetiology that has been reported at high frequency in oesophageal tumours16,32. (Fig. 6d).
Finally, we applied CUPPA to the 22 cfDNA-WGS cases to compare predictions to CUPPA on tissue WGS data as well as TOO based on clinicopathology review (Fig. 6e). CUPPA classified 9/22 (41%) cfDNA cases with high-likelihood prediction, and these were all concordant with the favoured TOO or were within a likely diagnostic differential. Of the 13 cases with low-likelihood CUPPA predictions, only two cases were discordant with the pathologist favoured diagnosis. Importantly, CUPPA using cfDNA-WGS made a high-likelihood prediction for four cases that were previously unresolved using panel data and where WGS was not possible due to limited tissue availability (CUP 1127 & CUP 1112: NSCLC, CUP 1190: Colorectum/Appendix/SI and CUP 1132: Pancreas).
Altogether, this demonstrates there is sufficient cfDNA with high enough tumour fraction to perform WGS in ~25% of all CUP patients enabling high-likelihood CUPPA predictions in 41% of patients tested.
Discussion
Genomic profiling is increasingly being used as part of clinical testing in CUP patients, supported by recent recommendations for tumour mutation profiling in CUP guidelines1,33. Although comprehensive panel sequencing can detect many approved therapeutic targets, WGS increases the diagnostic yield for treatment-decision making34. In the current study, we have also shown that WGTS is feasible using real-world FFPE clinical samples and can be as sensitive as panel testing. Our systematic comparison of panel and WGTS platforms confirmed that WGTS significantly increased the number of therapeutic and diagnostic features identified. Furthermore, algorithmic TOO prediction using WGTS data could assist resolving the TOO of many CUP tumours. Importantly, we show this method may also be extended to use on high-tumour-fraction cfDNA samples, thereby increasing accessibility of testing for CUP patients.
Several studies have described the feasibility of WGTS for paediatric and adult solid cancers, but few have supported the use of FFPE tissue specimens35,36,37,38,39,40,41,42. A pilot study of the 100,000 Genomes Project found WGS of FFPE nucleic acids was technically feasible in 52 cases20. Data quality was negatively impacted by formalin fixation, leading to increased copy-number noise, but this was improved by modifying the DNA reverse cross-linking protocol required for DNA extraction. The use of FFPE samples for clinical WGS was also shown in another larger pan-cancer study42. Although FFPE data was found to be noisier with identifiable artefacts detected, employing a quality control method (FFPEImpact) facilitated the interpretation of sequence data and a high concordance in actionable variants was shown (r = 0.86) in 51 matched fresh-FFPE pairs. In the current study, we achieved a high WGTS success rate (97%) using FFPE samples when we applied DNA quality control prior to library preparation and good concordance of reportable features was also observed between three paired fresh-FFPE WGS cases. High concordance was also seen in matched cancer panel sequencing for 71 cases, where no loss of sensitivity was observed in panel-captured regions. Although analysis of fresh tissue is still considered the gold standard for WGS, the collection and processing of fresh tissue samples remains logistically challenging in many centres and utilising FFPE samples would aid more equitable testing for patients while eliminating the need for additional invasive tissue biopsy procedures.
We have demonstrated that the yield of reportable treatment-related and diagnostic findings was higher for WGTS compared to panel testing. A previous study used in-silico comparison of WGS, whole-exome and multi-gene testing, showing panel sequencing was sufficient for detecting most treatment-related features, but WGS detected more features required for clinical trials43. Similarly, we found an increase in clinical trial eligibility using WGS data, with 20% more patients eligible for access to SOC therapy and/or phase I-II clinical trials. Importantly, we found that large structural events (CNV, SV) constituted most of the additional diagnostic information found by WGTS. SVs can occasionally involve pathognomonic drivers, such as TMPRSS2-ERG in prostate cancer and SS18-SSX in synovial sarcoma, as highlighted in recent ESMO guidelines1. These events can also be detected by some commercial panels or histological methods like fluorescence in situ hybridisation; however, non-recurrent SV events involving tumour suppressor genes are often missed by panel testing. For example, among our CUP tumours profiled, PBRM1 mutations and SVs were identified in four of seven CUPs thought to be intrahepatic cholangiocarcinomas by pathology review and confirmed by CUPPA prediction. PBRM1 events can be found in ~8%44,45 of cholangiocarcinomas and ~39% renal cell carcinomas46, therefore both cancers may be considered; however, these tumour types have distinct histological appearances, therefore, highlighting the need to interpret mutation data together with available clinical and pathological information.
Mutational signatures are another DNA feature that can contribute to diagnosis, where WGS was also superior. While SBS4 (tobacco smoking) and SBS7 (UV light damage) can both be detected by panel, these signatures can be missed due to a lower number of variants detected by panel sequencing. Aside from being of diagnostic value, mutational signatures are of high therapeutic importance, and we found WGS was superior not only for detecting HRD but also, surprisingly, MSI, although the patient numbers were small. Although targeted capture sequencing methods have been recently developed for detection of HRD and are used for cancer types such as ovarian (HRDscar47, Myriad Genetics, USA), breast and pancreatic cancer (HRDsig48, Foundation Medicine, USA), these tests would not typically be applied to CUP patients. For instance, we found HRD and a BRCA2 mutation in a CUP tumour that was ultimately resolved as a cholangiocarcinoma. Although it is plausible to detect HRD in a sequential manner to DNA panel testing, the competitive affordability of using targeted sequencing is eroded when there is a need to apply multiple panel tests.
WGTS was able to resolve a likely TOO in most CUP patients using a combination of curated DNA features to inform clinicopathology review and CUPPA. We previously reported that panel testing can assist TOO diagnosis in 31% of cases17. In an independent cohort of 58 CUPs, we now show that WGTS can aid in resolving up to 71% of cases. CUPPA alone made high-confidence predictions in 51% of cases, and all predictions were either concordant with the pathologist’s opinion or within a favoured diagnostic differential. These results agree with Schipper et al., who used CUPPA (DNA-only) for 72 CUP tumours, assigning a putative diagnosis in 64% of cases11. Although a limitation of our study was that a single pathologist performed centralised review of all cases, the review was done by an experienced pathologist in a hospital specialised in reviewing and treating CUP patients21. A good concordance was observed between CUPPA predictions and the pathologist’s independent opinion, enabling reciprocal validation of each approach. Importantly, we found that using genomic features to inform pathology review was important for rare cancers that are out of scope of the current CUPPA method, and equally, CUPPA did not make any high-likelihood misclassifications of rare cancers, showing its high specificity. Deep-learning classification has also been recently applied to panel data12, but high-confidence classifications were made in only 41% of CUP cases, again highlighting the increased diagnostic benefit of WGS over panel testing. Notably, we found that CUPPA classifier accuracy was improved by combining DNA and RNA data. However, gene expression alone can be unreliable for classification of some CUP cases, including lung-CUPs that have atypical transcriptional patterns, reflected by the absence of the diagnostic lung marker TTF-117. These tumours frequently harbour disruptive mutations in the SWI-SNF chromatin modifier SMARCA4 (BRG1), and we found SMARCA4 mutations in four of 11 lung-CUPs in the current series.
Despite the potential to use FFPE samples for WGTS, accessing tissue from pathology archives can still be slow and lead to extensive delays in testing49. In many cases, tissue samples are completely exhausted after immunohistochemistry testing. CUP patients are known to have high amounts of ctDNA in their blood, with panel testing detecting reportable findings in up to 80% of patients50. We found approximately a quarter of our CUP patients had enough cfDNA from a 20 mL blood draw for library preparation and a ctDNA fraction high enough to be compatible with current WGS pipelines. Further improvements are needed to increase the sensitivity of the approach, which may be achieved by simply increasing sequence coverage or modifying analytical pipelines. As others have shown, TOO classification is also possible using DNA methylation profiling9; therefore, incorporating DNA methylation into cfDNA classification may improve CUPPA classifier performance. Simultaneous mutation and DNA methylation detection is also now possible with some platforms51, which may be advantageous when limited tissue DNA or cfDNA prevents running two independent tests.
The adoption of WGTS into routine diagnostic workup is likely to increase with reduced cost of sequencing. We have shown the clinical validity of this technology in the work-up of CUP patients using real-world clinical samples, increasing the sensitivity for detecting therapeutic targets and identifying diagnostic features that can aid a TOO diagnosis. Further improvements will increase the sensitivity of the CUPPA method to improve classifier performance, especially among rare cancers, as well as when using cfDNA samples. Although the diagnostic and therapeutic applications of WGTS for CUP are apparent in our retrospective analysis, wider acceptance of such genomic tests for cancer diagnosis and treatment decision-making will require prospective validation to determine clinical impact.
Methods
CUP clinicopathological review
All CUP patients were recruited to the SUPER study from 11 Australian sites between 02/10/2017 and 20/10/2021 with informed consent under an approved protocol at Peter MacCallum Cancer Centre (PMCC) human research ethics committee (HREC protocol: 13/62). No direct patient identifying information has been included in this study and all patients are referenced with anonymous identifiers, where identities are only known to the study team. Patient consent was obtained in written and signed form to report indirect identifiers such as age and sex. No sex or gender specific analysis was carried out as this was not relevant to the study. Sex was self-reported but consistent with genomic data. Blinded histopathology review for the attempted tissue WGTS set (n = 74 patients and n = 75 distinct tumours) was performed by a single pathologist (O.W.J.P) (supplementary Data 1). Cases were assigned a favoured TOO upon registration to the study, also considering clinical presentation and cancer imaging and then again assigned a revised favoured TOO based on a curated panel and WGTS reports (Supplementary Data 1). The diagnosis was reassessed retrospectively, alongside as clinicopathology-unresolved or putatively diagnosed for all cases. This was done unblinded to genomics curated reports but blinded to the CUPPA result in all cases (n = 87, tissue and ctDNA predictions). When classification could not be reached, a modified version of the Memorial Sloan Kettering Cancer Centre (MSKCC) Oncotree classification criteria52 for CUPs was used to subclassify malignancies, and these were: undifferentiated malignant neoplasms (UDMN); poorly differentiated carcinoma (PDC); adenocarcinoma, not otherwise specified (ADNOS); neuroendocrine tumours, not otherwise specified; neuroendocrine carcinomas, not otherwise specified; and squamous cell carcinomas, not otherwise specified (SCCNOS). ADNOS were further subdivided based on cytokeratin 7 (CK7) and cytokeratin 20 (CK20) IHC staining, where CK7 was negative and CK20 had positive staining, caudal type homeobox 2 (CDX2) positivity was annotated. SCCNOS were subclassified based on p16INK4A (p16) IHC staining positivity.
Tissue and blood collection and nucleic acid extraction
Where possible, fresh tissue biopsy specimens were collected and stored in ‘RNAlater’ (Thermo Fisher, USA, cat. #AM7020) for 24 h before DNA and RNA extraction. Otherwise, an archived (FFPE) tissue specimen was used. Representative sections were reviewed by a pathologist and tumour regions macro-dissected before nucleic acid extraction. Regions > 30% tumour content were thought to be most suitable for WGTS, but there was no strict threshold applied. DNA and RNA extraction were done using the AllPrep DNA/RNA FFPE kit (QIAgen, USA, #80234). Alongside, for each recruited patient, a 25 mL whole blood sample was collected in EDTA tubes (5 mL) or Streck DNA Blood Collection Tubes (Streck, USA). Germline DNA was exclusively derived from whole blood in EDTA tubes. Blood plasma was derived only from Streck tubes and stored at −80 °C as 1 mL aliquots before cfDNA extraction with QIAamp circulating nucleic acid kit (QIAgen, USA, cat. #55114).
Cancer gene panel DNA sequencing
All cancer panel sequencing was performed by the medical laboratory Nexomics at PMCC. Thirty-four samples (24 with matched tissue WGTS) were sequenced with a custom cancer panel (CCP) targeting 386 genes (Agilent SureSelect), involving sequencing of both tumour and matched germline DNA, as previously described53. The CCP gene list is provided in supplementary information (Supplementary Table 2). Briefly, libraries were prepared and enriched using SureSelect XT enrichment (Agilent) and indexed libraries were pooled and sequenced to a median on-target tumour DNA coverage of 676x (range: 422–1086x) using 2 × 75 bp reads on an Illumina NextSeq500 instrument. Seqliner v0.7 (http://bioinformatics.petermac.org/seqliner/) was used to generate aligned reads against the hg38 human reference genome. Somatic variants were detected with Mutect v2.2 (https://github.com/broadinstitute/mutect) and filtered with GATK FilterMutectCalls v4.1.8.1 (https://github.com/broadinstitute/gatk). TMB estimation was described previously17 using the ensemble variant caller bcbio-nextgen (BCBio) cancer somatic variant calling pipeline (version 1.1.3a) (https://github.com/bcbio/bcbio-nextgen). Fifty tumour samples (47 with matched tissue WGS) were sequenced with the TruSight Oncology 500 panel (TSO500, Illumina), targeting 523 genes for DNA mutation detection and 55 cancer genes for RNA fusion and splice variant detection. TSO500 Libraries were prepared as per the manufacturer’s instructions and sequenced to median on-target coverage of 883x (range of 309–1473x) on an Illumina NextSeq500 instrument. Illumina Software TSO500 v2.0 Local App performed read alignment and variant calling against the hg37 human reference genome.
Whole genome and transcriptome sequencing
FFPE-derived DNA were assessed for WGTS suitability based on a modified GAPDH multiplex PCR assay that qualitatively estimates proportion of DNA fragments amplifiable between 100 bp and 400 bp in a given sample54. We modified this to also include additional primer pairs to amplify 500, 600 and 800 bp fragments (Supplementary information, Table 1), where scores of 1–8 were assigned to a sample depending on the largest amplifiable fragment (minimum ‘% Integrated Area’ of 10% as visualised by TapeStation 4200 D1000 electropherogram). Samples were deemed suitable for library preparation when they scored at least 4/8, indicating minimum ‘% Integrated Area’ of 10% was achieved for 400 bp length fragments as the largest fragment size. DNA Libraries were prepared using the Illumina TruSeq Nano library (Illumina, USA) method using 200 ng of input DNA. All libraries were quality controlled using the TapeStation high-sensitivity D5000 or D1000 ScreenTape (Agilent). Indexed libraries were pooled and sequenced, aiming for a depth of 50x for normal and 100x for tumour using 150 bp paired reads on an Illumina Novaseq 6000 platform (Illumina, USA). Sequence reads were aligned to hg37 and processed using the Hartwig Medical Foundation (HMF) pipeline v5 (https://github.com/hartwigmedical/pipeline5). WGS data was re-processed for all cases using hg38 retrospectively by the Illumina DRAGEN pipeline (https://doi.org/10.1101/2023.03.23.534011) and umccrise workflow (https://github.com/umccr/umccrise). For WTS, RNA samples were subjected to ribosomal RNA depletion using the NEBNext rRNA Depletion kit (New England Biolabs) according to the manufacturer’s instructions. ~100 million reads were generated per RNA sequencing library. WTS data was aligned using the STAR aligner55 (https://github.com/alexdobin/STAR) and feature counts were obtained as library-composition adjusted Transcripts Per Kilobase Million (adjTPM) using the Isofox algorithm (https://github.com/hartwigmedical/hmftools/tree/master/isofox). Fusions were confirmed in RNA-seq data using arriba (https://github.com/oicr-gsi/arriba).
TMB was calculated as outlined previously56. A ‘high’ TMB was called if the TMB was over ≥10 mutations per Mb, and ‘low’ if <10 mutations per Mb. MSI was estimated using the Personal Cancer Genome Report tool57 which utilises a statistical MSI classifier from somatic mutation profiles that separate MSI-high from microsatellite stable tumours (https://rpubs.com/sigven/msi_classification_v3), a model based on MSIseq58. COSMIC V2 mutational signatures were assigned using ‘MutationalPatterns’ (v3.12.0)59. Dominant signatures were defined as those with >20% of total somatic mutations within a sample.
HRD was independently confirmed by HRDetect26 and CHORD25. HRDetect and CHORD consider mutational patterns (SNVs, InDels and SVs) characteristic of HR-deficient tumours. HRDetect grants a score from 0 to 1; tumour samples with a score > 0.7 are categorised as HR-deficient. CHORD classifies tumours into BRCA1-deficient and BRCA2-deficient categories; tumours with a combined probability of these HRD categories >0.5 are categorised as homologous recombination-deficient.
Clinical and diagnostic features curation
A curation team at the PMCC reviewed the findings from CCP and TSO500. For CCP, Pathos v.1360 (https://github.com/PapenfussLab/PathOS) was used to annotate variants and filter for non-synonymous variants. Curation of germline variants was limited to 76 genes with evidence for cancer predisposition61. For TSO500, Clinical Genomics Workspace from PierianDx was used to annotate, filter and report mutations of diagnostic or therapeutic relevance. The genomic events reported include single-nucleotide and multi-nucleotide variants, small Indels, CNVs, MSI and TMB. Dominant mutational signatures were calculated using MutationalPatterns v3.12.0 (https://github.com/AlexandrovLab/SigProfilerExtractor).
A curation team at the University of Melbourne Centre for Cancer Research reviewed all the findings for each case that were clinically reported. Curation of SVs and CNVs in WGS data was limited to a custom list of 1246 cancer-related genes (https://github.com/umccr/umccrise/blob/master/workflow.md#key-cancer-genes) that were assembled using various sources. Manual inspection of DNA and RNA sequence reads was done to validate somatic variant calls using Integrative Genomics Viewer62 to visualise sequence coverage. Structural variants and breakpoint ends proximal to called CNV segment ends provided further supporting evidence. Gene fusions were supported by both WGS and WTS data. WTS data also supported genome amplification or deletion of genes by comparing the normalised expression (z-score) of an altered gene in the CUP sample to a large reference of unrelated cancer samples—the TCGA pan-cancer data set (https://github.com/umccr/RNAsum).
Regardless of sequencing approach, SNVs and Indels found in the tumour sample were prioritised into a four-tiered structure for clinical reporting based on actionable findings. This tiered structure was adopted from the joint consensus by AMP/ASCO/CAP on evidence-based variant categorisation63. We retrospectively reviewed and harmonised tumour variants based on therapeutic potential according to clinical trials available in January of 2024 and diagnostic potential according to significantly enriched variants within AACR Project GENIE cohorts17 and/or WGS landscape papers of the indicated cancer type. Briefly, therapeutic variants belonged to AMP/ASCO/CAP consensus tier 2c, while diagnostic variants were AMP/ASCO/CAP consensus tier 2 d if function was known or tier 3 if function was unknown but not/unlikely benign, therefore a variant could have been both therapeutic and diagnostic for a cancer type. To assign a standard of care (SOC) treatment or clinical trial, we used the EviQ knowledgebase, which is a free resource of evidence-based, consensus-driven cancer treatment protocols and information for the use of medical oncologists at the point of care (https://www.eviq.org.au/). EviQ references the Australian Pharmaceutical Benefit Scheme guide (updated 1 June 2024) to match government-funded treatments based on cancer type and molecular features. The Australian clinical trials database (https://www.australianclinicaltrials.gov.au/) was used to match CUP patients based on molecular evidence required for trial eligibility, with cancer type eligibility based on the combined genomics and clinicopathology evidence of cancer type. Although panel testing impacted enrolment of two patients onto a clinical trial, the outcomes of these trials have not been reported and all the WGTS and panel analysis in this manuscript are retrospective only. No information or data related to unpublished trial outcomes is reported in this manuscript.
Circulating tumour DNA extraction and sequencing
cfDNA was extracted from plasma using the QIAamp circulating nucleic acid kit (QIAgen, cat. #55114), following the standard extraction protocol using a QIAvac vacuum manifold (QIAgen, cats. #19413, #19419 and #84020) with one modification. Our laboratory prepares a stock concentration of 1 µg/µL of carrier RNA for use in the plasma sample lysis step instead of 0.2 ng/µL. Up to 5 mL of plasma was used as input into a single extraction for a sample. cfDNA was quantitated on an Agilent TapeStation 4200, using cfDNA tapes (cat# 5067–5630). We determined sample concentration from a size-limited range of 50–700 bp length DNA and cases were selected for library preparation if they had a minimum of 20 ng DNA. Whole-genome sequencing libraries were prepared using the NEB NEXT Ultra II kit (New England Biolabs, #E7645). Indexed libraries were pooled and sequenced, aiming for 1–2x coverage using paired 150 bp reads on the Illumina Novaseq 6000 platform (Illumina, USA). Tumour fraction was estimated using ichorCNA31 on each sample to determine suitability of libraries for deeper sequencing.
CUP prediction algorithm (CUPPA) on tissue and cfDNA
The CUPPA tool, which is a TOO classifier that uses WGS and/or WTS data, was developed by Hartwig Medical Foundation and described previously11. We used version 1.4 of CUPPA on eligible WGS and WTS data (https://github.com/hartwigmedical/hmftools/tree/master/cuppa). We utilised CUPPA DNA and RNA combined classifier outputs for final classification in tissue when RNA was available (69/73 cases) or only considered the DNA combined classification when RNA was unavailable or tumour purity was estimated to be <30% by WGS (PURPLE tool) (Supplementary data 1).
We used CUPPA ‘DNA only’ classification for cfDNA samples. We observed a pattern of false-positive SNV calls in the eight tissue and cfDNA matched WGS cases. To reduce the false positive variants in cfDNA samples, we ran Strelka264 v2.9.2 (https://github.com/Illumina/strelka) and generated an ensemble of shared Strelka2 pass-filter calls and somatic SNVs from the HMF pipeline. In addition, we removed a customised panel of SNVs, which was constructed using somatic SNVs detected in two or more independent cfDNA samples but were absent in their matching tissues. HOTSPOT SNVs annotated by the HMF pipeline were not filtered.
Figure generation
Figure generation was performed using R (v4.2.0), using tidyverse (v2.0.0), ComplexHeatmap (v2.18.0), patchwork (v.1.2.0). Flow charts were created with BioRender (https://biorender.com). Bar charts, scatter plots were performed using GraphPad Prism version 10.0.0 for Windows, GraphPad Software, Boston, Massachusetts USA (www.graphpad.com). Sankey plots were made using SankeyMATIC (https://sankeymatic.com/). All other graphics were created with Affinity Designer 2 (https://affinity.serif.com/en-us/designer/).
Statistics and reproducibility
All statistical tests were performed using GraphPad Prism version 10.0.0, for Windows, GraphPad Software, Boston, Massachusetts, USA (www.graphpad.com). No statistical method was used to predetermine sample size. Samples and assays failing quality control measures for tumour content and nucleic acid quality were excluded from analysis and are specified in text. The experiments were not randomised. The investigators were not blinded to allocation during experiments, although pathologist O.W.J.P was blinded to CUPPA predictions when classifying CUP cases.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The genomics data generated in this study have been deposited in the European Genome-Phenome Archive (EGA) repository, EGAS50000000452. Raw data were deposited as GRCh38 aligned BAM files and includes whole genome sequencing (n = 73) and whole transcriptome sequencing (n = 69) of CUP tumours and matched panel (CCP or Illumina TSO500 panel, n = 71). As genomic data represent potentially identifying information, these data are under restricted access controlled by the Data Access Committee at the University of Melbourne Centre for Cancer Research. Access to sequence read-level data or germline variant data will require evidence of institutional human research ethics committee approval. Access can be obtained by application through the EGA portal and applications will be processed within four to six weeks. If approved, data access will be granted in perpetuity. The raw clinical data are protected and are not available due to data privacy laws. The HMF dataset that was used for CUPPA validation can be accessed via application to the Hartwig Medical Foundation. The curated variant data generated in this study are provided in both Supplementary Data and Source Data files. Source data are provided with this paper.
Code availability
Code to reproduce all relevant figures from supplementary and source data included in this article is available in FigShare. https://doi.org/10.6084/m9.figshare.2651866965.
References
Kramer, A. et al. Cancer of unknown primary: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann. Oncol. 34, 228–246 (2023).
Rassy, E. & Pavlidis, N. The currently declining incidence of cancer of unknown primary. Cancer Epidemiol. 61, 139–141 (2019).
Sivakumaran, T., Tothill, R. W. & Mileshkin, L. R. The evolution of molecular management of carcinoma of unknown primary. Curr. Opin. Oncol. 36, 456–464 (2024).
Kramer, A., et al. Molecularly guided therapy versus chemotherapy after disease control in unfavourable cancer of unknown primary (CUPISCO): an open-label, randomised, phase 2 study. Lancet 404, 527–539 (2024).
Hyman, D. M. et al. Vemurafenib in multiple nonmelanoma cancers with BRAF V600 mutations. N. Engl. J. Med. 373, 726–736 (2015).
Greco, F. A., Lennington, W. J., Spigel, D. R. & Hainsworth, J. D. Molecular profiling diagnosis in unknown primary cancer: accuracy and ability to complement standard pathology. J. Natl. Cancer Inst. 105, 782–790 (2013).
Tothill, R. W. et al. Development and validation of a gene expression tumour classifier for cancer of unknown primary. Pathology 47, 7–12 (2015).
Meiri, E. et al. A second-generation microRNA-based assay for diagnosing tumor tissue origin. Oncologist 17, 801–812 (2012).
Moran, S. et al. Epigenetic profiling to classify cancer of unknown primary: a multicentre, retrospective analysis. Lancet Oncol. 17, 1386–1395 (2016).
Nguyen, L., Van Hoeck, A. & Cuppen, E. Machine learning-based tissue of origin classification for cancer of unknown primary diagnostics using genome-wide mutation features. Nat. Commun. 13, 4013 (2022).
Schipper, L. J. et al. Complete genomic characterization in patients with cancer of unknown primary origin in routine diagnostics. ESMO Open 7, 100611 (2022).
Moon, I. et al. Machine learning for genetics-based classification and treatment response prediction in cancer of unknown primary. Nat. Med. 29, 2057–2067 (2023).
Darmofal, M. et al. Deep-learning model for tumor-type prediction using targeted clinical genomic sequencing data. Cancer Discov. 14, 1064–1081 (2024).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Alexandrov, L. B. et al. Mutational signatures associated with tobacco smoking in human cancer. Science 354, 618–622 (2016).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Posner, A. et al. A comparison of DNA sequencing and gene expression profiling to assist tissue of origin diagnosis in cancer of unknown primary. J. Pathol. 259, 81–92 (2023).
Jiao, W. et al. A deep learning system accurately classifies primary and metastatic cancers using passenger mutation patterns. Nat. Commun. 11, 728 (2020).
Mohrmann, L. et al. Comprehensive genomic and epigenomic analysis in cancer of unknown primary guides molecularly-informed therapies despite heterogeneity. Nat. Commun. 13, 4485 (2022).
Robbe, P. et al. Clinical whole-genome sequencing from routine formalin-fixed, paraffin-embedded specimens: pilot study for the 100,000 genomes project. Genet. Med. 20, 1196–1205 (2018).
van Mourik, A. et al. Six-year experience of Australia’s first dedicated cancer of unknown primary clinic. Br. J. Cancer 129, 301–308 (2023).
Van Hoeck, A., Tjoonk, N. H., van Boxtel, R. & Cuppen, E. Portrait of a cancer: mutational signature analyses for cancer diagnostics. BMC Cancer 19, 457 (2019).
Telli, M. L. et al. Homologous recombination deficiency (HRD) score predicts response to platinum-containing neoadjuvant chemotherapy in patients with triple-negative breast cancer. Clin. Cancer Res. 22, 3764–3773 (2016).
Tsang, E. S. et al. Homologous recombination deficiency signatures in gastrointestinal and thoracic cancers correlate with platinum therapy duration. NPJ Precis. Oncol. 7, 31 (2023).
Nguyen, L., W M Martens, J., Van Hoeck, A. & Cuppen, E. Pan-cancer landscape of homologous recombination deficiency. Nat. Commun. 11, 5584 (2020).
Davies, H. et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat. Med. 23, 517–525 (2017).
Schoenfeld, A. J. et al. The genomic landscape of SMARCA4 alterations and associations with outcomes in patients with lung cancer. Clin. Cancer Res. 26, 5701–5708 (2020).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Petrelli, F., Ghidini, M., Ghidini, A. & Tomasello, G. Outcomes following immune checkpoint inhibitor treatment of patients with microsatellite instability-high cancers: a systematic review and meta-analysis. JAMA Oncol. 6, 1068–1071 (2020).
Adalsteinsson, V. A. et al. Scalable whole-exome sequencing of cell-free DNA reveals high concordance with metastatic tumors. Nat. Commun. 8, 1324 (2017).
Secrier, M. et al. Mutational signatures in esophageal adenocarcinoma define etiologically distinct subgroups with therapeutic relevance. Nat. Genet. 48, 1131–1141 (2016).
Referenced with permission from the NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) for Occult Primary V.1.2024. © National Comprehensive Cancer Network, Inc. 2024. All rights reserved. Accessed [02/03/2024]. To view the most recent and complete version of the guideline, go online to NCCN.org. (2024).
Samsom, K. G. et al. Feasibility of whole-genome sequencing-based tumor diagnostics in routine pathology practice. J. Pathol. 258, 179–188 (2022).
Wong, M. et al. Whole genome, transcriptome and methylome profiling enhances actionable target discovery in high-risk pediatric cancer. Nat. Med. 26, 1742–1753 (2020).
Byrjalsen, A. et al. Nationwide germline whole genome sequencing of 198 consecutive pediatric cancer patients reveals a high incidence of cancer prone syndromes. PLoS Genet. 16, e1009231 (2020).
Trotman, J. et al. The NHS England 100,000 genomes project: feasibility and utility of centralised genome sequencing for children with cancer. Br. J. Cancer 127, 137–144 (2022).
Tessier-Cloutier, B. et al. The impact of whole genome and transcriptome analysis (WGTA) on predictive biomarker discovery and diagnostic accuracy of advanced malignancies. J. Pathol. Clin. Res. 8, 395–407 (2022).
Samsom, K. G. et al. Optimized whole-genome sequencing workflow for tumor diagnostics in routine pathology practice. Nat. Protoc. 19, 700–726 (2023).
Pleasance, E. et al. Whole-genome and transcriptome analysis enhances precision cancer treatment options. Ann. Oncol. 33, 939–949 (2022).
Shukla, N. et al. Feasibility of whole genome and transcriptome profiling in pediatric and young adult cancers. Nat. Commun. 13, 2485 (2022).
Basyuni, S. et al. Large-scale analysis of whole genome sequencing data from formalin-fixed paraffin-embedded cancer specimens demonstrates preservation of clinical utility. Nat. Commun. 15, 7731 (2024).
Ramarao-Milne, P. et al. Comparison of actionable events detected in cancer genomes by whole-genome sequencing, in silico whole-exome and mutation panels. ESMO Open 7, 100540 (2022).
Zou, S. et al. Mutational landscape of intrahepatic cholangiocarcinoma. Nat. Commun. 5, 5696 (2014).
Jusakul, A. et al. Whole-genome and epigenomic landscapes of etiologically distinct subtypes of cholangiocarcinoma. Cancer Discov. 7, 1116–1135 (2017).
Scelo, G. et al. Variation in genomic landscape of clear cell renal cell carcinoma across Europe. Nat. Commun. 5, 5135 (2014).
Koskela, H. et al. HRD related signature 3 predicts clinical outcome in advanced tubo-ovarian high-grade serous carcinoma. Gynecol. Oncol. 180, 91–98 (2024).
Chen, K. T. et al. A Novel HRD signature is predictive of FOLFIRINOX benefit in metastatic pancreatic cancer. Oncologist 28, 691–698 (2023).
Huey, R. W. et al. Feasibility and value of genomic profiling in cancer of unknown primary: real-world evidence from prospective profiling study. J. Natl. Cancer Inst. 115, 994–997 (2023).
Kato, S. et al. Utility of genomic analysis in circulating tumor DNA from patients with carcinoma of unknown primary. Cancer Res. 77, 4238–4246 (2017).
Fullgrabe, J. et al. Simultaneous sequencing of genetic and epigenetic bases in DNA. Nat. Biotechnol. 41, 1457–1464 (2023).
Kundra, R. et al. OncoTree: a cancer classification system for precision oncology. JCO Clin. Cancer Inf. 5, 221–230 (2021).
McEvoy, C. R. et al. Improved next-generation sequencing pre-capture library yields and sequencing parameters using on-bead PCR. Biotechniques 68, 48–51 (2020).
van Beers, E. H. et al. A multiplex PCR predictor for aCGH success of FFPE samples. Br. J. Cancer 94, 333–337 (2006).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Chalmers, Z. R. et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 9, 34 (2017).
Nakken, S. et al. Personal cancer genome reporter: variant interpretation report for precision oncology. Bioinformatics 34, 1778–1780 (2018).
Huang, M. N. et al. MSIseq: software for assessing microsatellite instability from catalogs of somatic mutations. Sci. Rep. 5, 13321 (2015).
Blokzijl, F., Janssen, R., van Boxtel, R. & Cuppen, E. MutationalPatterns: comprehensive genome-wide analysis of mutational processes. Genome Med. 10, 33 (2018).
Doig, K. D. et al. PathOS: a decision support system for reporting high throughput sequencing of cancers in clinical diagnostic laboratories. Genome Med. 9, 38 (2017).
Rahman, N. Realizing the promise of cancer predisposition genes. Nature 505, 302–308 (2014).
Robinson, J. T., Thorvaldsdottir, H., Wenger, A. M., Zehir, A. & Mesirov, J. P. Variant review with the integrative genomics viewer. Cancer Res. 77, e31–e34 (2017).
Li, M. M. et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: a joint consensus recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J. Mol. Diagn. 19, 4–23 (2017).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Rebello, R. et al. Whole genome sequencing improves tissue-of-origin diagnosis and treatment options for cancer of unknown primary. FigShare https://doi.org/10.6084/m9.figshare.26518669 (2025).
Acknowledgements
We wish to thank SUPER study coordinators at participating hospitals for facilitating collection of biospecimens and data for the study. We wish to acknowledge the patients who have contributed to this study and the CUP consumer steering committee: Cindy Bryant (chair), Kym Sheehan, Christine Bradfield, Clare Brophy, Dale Witton, and Frank Stoss. The study was supported by funding from Australian Health Genome Alliance (AGHA) and National Health and Medical Research Council (NHMRC) (1113531) and the Medical Research Future Fund (MRFF, GHFMCDI000003). RWT was supported by funding from the Victorian Cancer Agency (TP828750). The Westmead, Blacktown, and Nepean study sites were supported by the Cancer Institute NSW 11/TRC/1-06, 15/TRC/1-01, and 15/RIG/1-16. We acknowledge the TAGC clinical genomics platform group for their valuable contributions.
Author information
Authors and Affiliations
Contributions
R.W.T. and L.M. conceived the study. R.J.R. and A.P. performed the analysis. R.J.R. and R.D. draughted the figures and tables. O.W.J.P. and C.M. undertook histopathology review of CUP. cases. T.S. and L.M. reviewed the clinical data, and R.J.R., C.B.M., W.Z. and T.S. retrospectively reviewed clinical trials eligibility based on biomarker data. C.F., K.F. and S.W. collected clinical research data for the study. L.M., T.S., H.W., A.D.F., N.W., B.G., C.St., M.S., I.M.C., C.S., M.W. and N.K. screened patients for eligibility and recruited them to the study. C.Sh. and P.P. developed the CUPPA classifier. V.D. provided critical experimental support. C.B.M., J.H.A.V., W.Z., S.N., A.Fe, H.X., S.B. and S.F. curated mutation profiling data. R.D., S.K., A.C. and A.Fl. performed the bioinformatic analysis. R.J.R. and R.T. co-wrote the manuscript and all authors critically reviewed the manuscript (including R.J.H., S.J.D. and S.B.F.). S.G., K.P.S. and O.H. oversaw the clinical genomic profiling and bioinformatic cohort profiling for this study. P.S., D.B., L.M. and R.T. were the principal investigators and obtained research funding to support the study.
Corresponding author
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Angela Rose Brannon, Rebecca Deyell and Lino Möhrmann for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rebello, R.J., Posner, A., Dong, R. et al. Whole genome sequencing improves tissue-of-origin diagnosis and treatment options for cancer of unknown primary. Nat Commun 16, 4422 (2025). https://doi.org/10.1038/s41467-025-59661-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59661-x
This article is cited by
-
Rethinking cancer of unknown primary: from diagnostic challenge to targeted treatment
Nature Reviews Clinical Oncology (2025)
-
Pathologist-initiated whole genome and transcriptome sequencing demonstrates diagnostic utility in resolving difficult-to-diagnose tumors
Genome Medicine (2025)