Abstract
Mitochondrial diseases (MtD) represent a significant public health challenge due to their heterogenous clinical presentation, often severe and progressive symptoms, and lack of effective therapies. Environmental exposures, such bacterial and viral infection, can further compromise mitochondrial function and exacerbate the progression of MtD. However, the underlying immune alterations that enhance immunopathology in MtD remain unclear. Here we employ in vitro and in vivo approaches to clarify the molecular and cellular basis for innate immune hyperactivity in models of polymerase gamma (Polg)-related MtD. We reveal that type I interferon (IFN-I)-mediated upregulation of caspase-11 and guanylate-binding proteins (GBP) increase macrophage sensing of the opportunistic microbe Pseudomonas aeruginosa (PA) in Polg mutant mice. Furthermore, we show that excessive cytokine secretion and activation of pyroptotic cell death pathways contribute to lung inflammation and morbidity after infection with PA. Our work provides a mechanistic framework for understanding innate immune dysregulation in MtD and reveals potential targets for limiting infection- and inflammation-related complications in Polg-related MtD.
Similar content being viewed by others
Introduction
Mitochondrial diseases (MtD) are the most common inborn error of metabolism and are caused by deleterious variants in genes that function in oxidative phosphorylation (OXPHOS) and mitochondrial DNA (mtDNA) maintenance1. MtD can present at any age, vary in severity, and display clinical heterogeneity with differing degrees of organ system involvement2,3. Although the genetic basis of MtD is well understood, the role of environmental factors in disease progression remains unclear. Notably, viral and bacterial infections may unmask MtD or hasten the stepwise progression of disease by triggering metabolic decompensation, a dangerous deterioration in mitochondrial function4. Clinical evidence suggests that patients with MtD are highly susceptible to infection by opportunistic respiratory pathogens. Moreover, pneumonia and microbial sepsis are the primary causes of early death in pediatric patients, and some adults, with MtD5,6. Thus, there is ample evidence highlighting the damaging toll of infection in MtD7. However, the underlying immune alterations that drive immunopathology and prevent proper control of infection represent a significant knowledge gap that complicates treatment and enhances morbidity and mortality in this vulnerable patient population.
In addition to their bioenergetic functions, mitochondria are key metabolic and signaling hubs in the mammalian immune system8. Mitochondria are also important sources of damage-associated molecular patterns (DAMP), such as mtDNA, which engage innate immune signaling when released from the organelle and are linked to inflammation in an increasing number of human diseases9,10. There is growing appreciation for a bidirectional relationship between MtD and inflammation, where mitochondrial dysfunction in MtD activates the innate immune system leading to inflammatory responses that further compromise mitochondrial activity11. In agreement with this hypothesis, elevated inflammatory cytokines have been observed in patients with Polymerase gamma (Polg)-related Alpers-Huttenlocher syndrome12, Barth syndrome13, and Friedreich ataxia14,15. Animal models have also revealed innate immune system dysregulation as a frequent feature of MtD. For example, the Ndufs4−/− mouse model of neurodegeneration and Leigh syndrome exhibits increased circulating pro-inflammatory cytokines along with macrophage and microglial activation16,17,18,19.
We recently uncovered that the PolgD257A mutator model of mtDNA instability exhibits a hyperinflammatory innate immune status due to chronic engagement of the cyclic GMP-AMP synthase (cGAS)—Stimulator of Interferon Genes (STING)—type one interferon (IFN-I) axis20. Notably, we reported that persistent IFN-I signaling in these mice causes widespread repression of the antioxidant and anti-inflammatory transcription factor nuclear factor erythroid 2-related factor 2 (Nrf2)21,22 in myeloid immune cells and organs of the PolgD257A mouse. Consequently, PolgD257A mice are more sensitive to septic shock caused by intraperitoneal injection of a high dose of lipopolysaccharide (LPS)20. Although PolgD257A mutator mice do not phenocopy any particular human MtD, these animals present pathology overlapping the MtD spectrum, including hearing loss, cardiomyopathy, progressive anemia, and lymphopenia23,24,25. Moreover, increased susceptibility to septic shock and elevated inflammatory and IFN-I signatures are common features of both Polg mutator mice and patients with diverse MtD11.
Overall, our prior work provided foundational evidence linking mitochondrial dysfunction to hyperinflammatory innate immune signaling in the PolgD257A model of MtD. However, there remains a critical need to expand these findings into relevant infection models, given the high rate of infections that frequently lead to innate immune hyperactivity and sepsis in MtD. Pseudomonas aeruginosa (PA) is a common opportunistic pathogen of concern in MtD, as infection can lead to sepsis and pneumonia6. PA is a robust activator of innate immunity in the respiratory tract, and lung colonization by PA drives persistent recruitment of monocytes and neutrophils leading to pneumonia, lung injury, and/or systemic bacterial spread26,27. PA is especially problematic in MtD due to the prevalence of antibiotic resistant strains, abundance in hospitals, and high rate of recurrence26,28,29. Thus, it is important to define how PA and other opportunistic bacteria engage the innate immune system in MtD to define immunomodulatory strategies that boost anti-bacterial immunity, while limiting hyperinflammation that may hasten metabolic decompensation and disease progression.
In this study, we utilize in vitro and in vivo PA infection models to characterize the underlying innate immune mechanisms governing hyperinflammation in PolgD257A mice. We also generate and characterize a mouse model that incorporates a pathogenic Polg variant observed in humans. We reveal that both Polg mutant models exhibit excessive macrophage inflammasome activation, cytokine secretion, and pyroptotic cell death, which contribute to inflammation, acute lung injury, and morbidity after infection with PA. These findings further advance knowledge of how mitochondrial dysfunction impacts innate immune responses during bacterial infection and Polg-related diseases and reveals innate immune targets for limiting infection- and inflammation-related complications in MtD.
Results
Polg D257A mutant macrophages exhibit increased IFN-I and pro-inflammatory responses after bacterial challenge
Our recent work uncovered that elevated IFN-I signaling augments innate immune activity and inflammation in PolgD257A mice challenged with the Toll-like receptor (TLR) agonist, LPS20. We hypothesized that the hyperinflammatory phenotype of PolgD257A mice might be further exacerbated during bacterial infection. Therefore, we challenged bone marrow-derived macrophages (BMDM) with Pseudomonas aeruginosa strain O1 (PAO1) (Fig. 1a) and performed RNA sequencing (RNAseq) and pathway analysis (Fig. 1b). Similar to BMDMs treated with LPS20, PAO1-challenged PolgD257A BMDMs displayed heightened expression of pro-inflammatory cytokines, IFN-I, interferon-stimulated genes (ISG), and innate immune signaling factors (Fig. 1c, d; Supplementary Fig. 1a). Gene Set Enrichment Analysis confirmed PAO1-infected PolgD257A BMDMs exhibit more robust pro-inflammatory cytokine, chemokine, and IFN-I gene expression relative to wild-type (WT) BMDMs (Supplementary Fig. 1b). Additional experiments substantiated RNAseq datasets, revealing heightened expression of ISGs after challenge of PolgD257A BMDMs with PAO1 between six to twenty-four hours (Fig. 1e, f). Proteomic analysis of PAO1-challenged BMDMs revealed strong concurrence with RNAseq data, showing elevated IFN-I and inflammatory responses in PolgD257A BMDMs (Fig. 1g, h, Supplementary Fig. 1c). Moreover, multi-analyte cytokine analysis by LegendPlex showed a global elevation of proinflammatory cytokines produced by PolgD257A peritoneal macrophages (PerMac) six hours after PAO1 challenge (Fig. 1i) and ELISA showed a more rapid and sustained release of these cytokines over a one-day time course (Fig. 1j). In sum, our data highlight that PolgD257A macrophages are hyperactivated by both LPS and PAO1 challenge.
a Workflow for preparing RNA from Polg mutant BMDMs after Pseudomonas aeruginosa (PA) strain O1 (PAO1) challenge. Created in BioRender. Vanportfliet, J. (2025) https://BioRender.com/g7t8646. b Ingenuity Pathway Analysis (IPA) of RNA-seq from bone marrow-derived macrophages (BMDMs) showing Z-scores of pathways in PolgD257A homozygous mutant BMDMs compared to wild-type (WT) after LPS or PAO1 challenge. hpc = hours post challenge. N = 3 biological replicates/genotype/condition. c Log2 fold changes of representative genes in IPA pathways from (b) in PolgD257A BMDMs compared to WT after LPS or PAO1. d Protein abundance of interferon-stimulated genes (ISGs) in BMDMs exposed to PAO1 at a multiplicity of infection (MOI) of 5 at the indicated hours post challenge (hpc). e qRT-PCR analysis of ISGs in WT and PolgD257A BMDMs after PAO1. N = 3 biological replicates, 2 technical replicates/genotype/condition. f Western blot analysis of interferon stimulated genes (ISGs) over time course infection. Representative blots from N = 3 experiments. g Ingenuity pathway analysis showing z-scores of upregulated pathways by proteomics from PolgD257A (PAO1/Un)/WT (PAO1/Un) BMDM. N = 4 biological replicates. h IPA Upstream Regulator analysis showing Z-scores in PolgD257A (PAO1/Un)/WT (PAO1/Un). i Legendplex pro-inflammatory cytokines analysis of PerMac supernatants 24 h post challenge (hpc). N = 3 biological replicates. j Proinflammatory cytokines in supernatant of peritoneal macrophages (PerMacs) measured by ELISA. MOI = multiplicity of infection, N = 8 biological replicates over 4 time points. k IL-1β in supernatant of peritoneal macrophages (PerMacs) measured by ELISA. MOI = multiplicity of infection, N = 8 biological replicates over 4 time points. l Ingenuity pathway analysis of proteomics from PolgD257A (PAO1, 8hpc/Un)/WT (PAO1, 8 hpc/Un) BMDM proteins in the Pyroptotic Signaling Pathway. N = 4 biological replicates. Statistics: (e, i, j) Two-Way ANOVA. Error bars represent SEM. Source data are provided as a Source Data file.
Polg D257A mutant macrophages exhibit potentiated pyroptotic cell death upon bacterial challenge
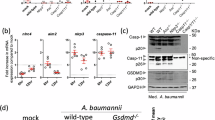
As we observed increased IL-1β secretion from PolgD257A PerMacs (Fig. 1i, k), we next investigated inflammatory cell death pathways during PAO1 infection. RNAseq and pathway analysis revealed significant enrichment of genes involved in inflammasome signaling, immunogenic cell death, and death receptor signaling in PolgD257A BMDMs after PAO1 challenge (Supplementary Fig. 1d, e). Pyroptotic cell death proteins were also highly upregulated in proteomic analysis (Fig. 1l). To further confirm a role for pyroptosis in hyperinflammation and increased cell death in PolgD257A macrophages, we incubated BMDMs with Ac-YVAD-cmk (AcYVAD), a selective caspase-1 inhibitor. AcYVAD nearly abrogated IL-1β secretion from PolgD257A macrophages, while also reducing the levels of inflammasome-independent inflammatory cytokines TNFα and IL-6 upon PAO1 challenge (Fig. 2a). In contrast, QVD-OPH, a more potent inhibitor of apoptotic caspases, had minor effects on PAO1-induced cytokine secretion in PolgD257A BMDMs (Fig. 2a). Moreover, we did not observe striking differences in necroptotic signaling proteins during PAO1 challenge, and necrostatin-1 failed to blunt elevated cytokine secretion from PolgD257A BMDMs (Supplementary Fig. 2a, b).
a ELISA analysis of inflammatory cytokines in BMDM supernatant after PAO1 or PAO1 plus caspase inhibitors. N = 4 biological replicates over 2 timepoints. b Cytation5 image analysis of PI positive BMDMs after PAO1 or PAO1 plus caspase-1 inhibitor. N = 6 biological replicates/genotype/condition. c Percent PerMacs dying after PAO1 and PAO1 plus caspase-1 inhibitor from IF (d). N = 4 IF images/genotype/condition. d Representative IF of PerMacs after PAO1 or PAO1 plus caspase inhibitor. Cells were stained with DAPI to mark the nucleus, CellMask to mark the total cell volume, and anti-GFP to mark GFP tagged PAO1. White arrows highlight dying cells characterized by condensed nuclei and loss of CellMask staining. Representative images from N = 3 experiments. e Western blot analysis of lysates from BMDMs challenged with PAO1. Representative blots from N = 3 experiments. f LegendPlex IL-1 cytokine analysis of BMDM supernatant 24 hpc. All undefined comparisons between WT and WT+AcYVAD are not significant. N = 3 biological replicates/genotype/condition. g LegendPlex pro-inflammatory cytokine analysis of BMDM supernatant 24 hpc. All undefined comparisons between WT and WT+AcYVAD are not significant. N = 3 biological replicates/genotype/condition. h Cytation5 image quantification of PI positive BMDMs after LPS priming plus LPS transfection, and LPS priming plus LPS transfection plus caspase-1 inhibitor. N = 3–5 biological replicates/genotype/condition. i ELISA analysis of IL-1β in BMDM supernatant after LPS priming plus LPS transfection in the presence or absence of caspase-1 inhibitor. N = 4 biological replicates. j qRT-PCR analysis of Caspase-11 RNA expression in BMDMs after PAO1. N = 3 biological replicates, 3 technical replicates/genotype/condition. k Cytation5 image quantification of PI positive cells in BMDMs after LPS priming plus LPS transfection, and incubated with αIFNAR blocking antibody or αIgG control. N = 4–6 biological replicates/genotype/condition. Statistics: (c, f, g, i) One-Way ANOVA (a, b, h, j, k) Two-Way ANOVA. Error bars represent SEM. Source data are provided as a Source Data file.
Consistent with cell death gene expression patterns, PolgD257A BMDMs more rapidly became propidium iodide (PI) positive during PAO1 challenge (Fig. 2b). PolgD257A PerMacs stained weakly for the nuclear and plasma membrane dye CellMask HCS Deep Red and exhibited rapid condensation of nuclei after PAO1 challenge (Fig. 2c, d). Reduced CellMask staining was also observed in ASC positive WT BMDMs after LPS priming and nigericin treatment (Supplementary Fig. 2c, d), indicating that low CellMask staining marks cells undergoing pyroptosis. PolgD257A BMDMs also released more lactate dehydrogenase (LDH) after PAO1 challenge (Supplementary Fig. 2e). We also noted elevated release of PAO1 into the culture supernatant from PolgD257A BMDMs, concomitant with lower numbers of intracellular bacteria (Supplementary Fig. 3a). Reduced intracellular bacteria was not the result of impaired phagocytosis, as PolgD257A BMDMs displayed no impairment in uptake of LPS coated fluorescent microspheres (Supplementary Fig. 3b, c). AcYVAD treatment was sufficient to blunt pyroptotic phenotypes of PolgD257A macrophages (Fig. 2b–d) after incubation with PAO1, further supporting a caspase-1-dependent cell death pathway.
In addition to activating NLRP3, PA can also activate the NLRC4 canonical inflammasome and the non-canonical caspase-11 inflammasome27,30,31,32. Culture of PolgD257A BMDMs with non-pathogenic E. coli (DH5α), which lack flagellin and therefore can only activate NLRP3 or caspase-11, also promoted pyroptotic cell death (Supplementary Fig. 3d, e) and increased IL-1β secretion (Supplementary Fig. 3f), while boosting TNFα and IL-6 secretion (Supplementary Fig. 3g). Collectively, these data indicate that elevated cell death and cytokine secretion in PolgD257A macrophages during bacterial challenge are not due to PAO1 virulence factors but are likely caused by potentiated activation of NLRP3 and/or caspase-11 inflammasomes.
Elevated IFN-I enhances expression and activation of caspase-11 and caspase-1 in Polg D257A macrophages
In WT macrophages, PAO1 mainly engages the NLRC4 inflammasome, while activation of NLRP3 and caspase-11 inflammasomes occur at later stages of infection30,33,34,35. Given elevated pyroptosis in PAO1-infected PolgD257A BMDMs, we next performed isolated challenge experiments to evaluate hyperactivation of particular inflammasome pathways in PolgD257A macrophages. PolgD257A BMDMs treated with LPS alone did not exhibit increased PI uptake, suggesting that priming is not sufficient to potentiate cell death (Supplementary Fig. 4a). Surprisingly, LPS priming followed by flagellin transfection to activate the NLRC4 inflammasome revealed that PolgD257A macrophages exhibit reduced PI uptake and IL-1β secretion (Supplementary Fig. 4b, c). LPS priming followed by the addition of ATP also revealed modestly impaired NLRP3 inflammasome activation in PolgD257A BMDMs, as measured by PI uptake, IL-1β secretion, and caspase-1 cleavage (Supplementary Fig. 4d–f). LPS priming followed by nigericin treatment revealed similar impairments in NLRP3 inflammasome activity (Supplementary Fig. 3g–i). OXPHOS has recently been shown to be vital for NLRP3 activation in response to nigericin and other stimuli36. PolgD257A BMDMs displayed dysfunctional mitochondrial morphology by TEM as well as reduced basal and maximal respiration at rest and after LPS challenge (Supplementary Fig. 4j–m), suggesting that reduced canonical NLRP3 activation is likely due to OXPHOS impairment in PolgD257A BMDMs.
Although PolgD257A BMDMs exhibited reduced NLRP3 and NLRC4 responses when triggered with pure inflammasome activators, mutant BMDMs displayed elevated caspase-1 and Gasdermin D (GSDMD) cleavage, as well as increased caspase-11 and IL-1β expression, after incubation with PAO1 as compared with WT controls (Fig. 2e). Moreover, AcYVAD treatment inhibited the hypersecretion of IL-1β and other pro-inflammatory cytokines in PolgD257A BMDMs challenged with PAO1 for 24 h (Fig. 2f, g). Notably, IL-1α secretion was markedly higher in PolgD257A macrophages after PAO1 challenge (Figs. 1i and 2f). IL-1α is an alarmin that feeds forward to augment production of other cytokines, and the robust secretion of IL-1α from myeloid cells involves caspase-1137,38,39. Thus, given elevated caspase-11 expression in PolgD257A BMDMs, we next explored its involvement in cell death and cytokine secretion. After LPS priming followed by LPS transfection, PolgD257A BMDMs rapidly became PI positive and secreted increased levels of IL-1β (Fig. 2h, i). Both phenotypes could be markedly inhibited by treatment with AcYVAD, indicating that heightened activation of caspase-11 feeds forward to activate caspase-1 more robustly in PolgD257A BMDMs (Fig. 2h, i). Inhibition of NLRP3 with MCC950 reduced cell death and hyperinflammation in PolgD257A BMDMs after PAO1 challenge (Supplementary Fig. 5a–d). NLRP3-dependent caspase-1 activation can drive GSDMD to permeabilize mitochondrial membranes40. Therefore, we isolated mitochondria from unchallenged and PAO1-challenged BMDMs and examined GSDMD accumulation in mitochondria-enriched fractions. Although there was no difference in the mitochondrial accumulation of GSDMD between genotypes, we observed markedly increased GSDMD cleavage in whole cell lysates of PolgD257A BMDMs rapidly upon PAO1 challenge (Supplementary Fig. 5e).
As caspase-11 activation requires IFN-I priming41 and PolgD257A macrophages and mice exhibit elevated IFN-I signaling20, we reasoned that IFN-I is responsible for caspase-11 expression and activation in PolgD257A BMDMs. In agreement, we observed potentiated caspase-11 mRNA and protein after PAO1 infection (Figs. 1l, 2e, j). Notably, IFN alpha receptor (IFNAR) blockade using the MAR1-5A3 monoclonal antibody markedly reduced PI uptake after caspase-11 activation by transfected LPS (Fig. 2k). Collectively, these data indicate that the IFN-I-dependent potentiation of caspase-11 drives non-canonical caspase-1 activation and hyperinflammation in PolgD257A macrophages exposed to PAO1.
Elevated IFN-I signaling increases GBP expression and inflammation in Polg D257A macrophages
We next hypothesized that elevated IFN-I may contribute to the hyperinflammatory response of PolgD257A macrophages during PAO1 infection. Priming macrophages with an anti-IFNAR antibody prior to PAO1 challenge reduced cell death and IL-1β release in PolgD257A BMDMs (Fig. 3a–c). IFNAR blockade also reduced secretion of IL-6 and ISG expression (Fig. 3d, e). Prior work has shown that IFN-I signaling during bacterial infection drives the expression of guanylate binding proteins (GBP)42, which facilitate innate immune responses and non-canonical inflammasome activation by disrupting membranes of pathogen-containing vacuoles, or pathogens themselves, to liberate LPS or other PAMPs31,43,44,45. Consistent with elevated IFN-I signaling, RNAseq analysis revealed more robust GBP expression in PolgD257A BMDMs after challenge with PAO1 (Supplementary Fig. 5f). Proteomic analysis also showed elevated GBPs, with marked increases in GBP2 and GBP5 in PolgD257A BMDMs six hours post challenge with PAO1 (Supplementary Fig. 5g). Immunofluorescence microscopy revealed increased GBP5 expression in resting PolgD257A BMDMs (Supplementary Fig. 5h–i), as well as during PAO1 challenge (Fig. 3f–g). Notably, we observed a more robust colocalization of GBP5 with PAO1 in PolgD257A macrophages after staining with an antibody specific for PA LPS. Moreover, GBP5 staining was more aggregated on PAO1-LPS positive bacteria than in WT BMDMs (Fig. 3h). These findings suggest that elevated IFN-I in PolgD257A macrophages augments GBP expression, likely enhancing caspase-11-dependent sensing of LPS that contributes to potentiated cell death and pro-inflammatory cytokine release.
a Percent BMDMs dying after PAO1 and PAO1 plus anti-IFNAR blockade from (b) representative IF. N = 5–6 IF images/genotype/condition. c ELISA analysis of pyroptotic cytokine in BMDM supernatant after PAO1 or PAO1 plus anti-IFNAR blockade. N = 4 biological replicates over 2 timepoints. d ELISA analysis of inflammatory cytokine, IL-6, in BMDM supernatant after PAO1 or PAO1 plus anti-IFNAR blockade. N = 4 biological replicates over 2 timepoints. e qRT-PCR analysis of ISGs in WT and PolgD257A BMDMs after PAO1 and PAO1 plus anti-IFNAR blockade. N = 3 technical replicates/genotype/condition. f Representative IF of BMDMs 2 hpc with PAO1 at a multiplicity of infection (MOI) of 5. Cells were fixed, permeabilized, and stained with DAPI to mark the nucleus, anti-LPS antibody to stain PAO1 LPS, and anti-GBP5 antibody. Representative images from N = 3 experiments. g Sum intensity of GBP5 per cell area and (h) PAO1-LPS localized with GBP5 in BMDMs after PAO1 in (f) IF. N = 7–8 images/genotype. Statistics: (g, h) Two-Sided Student’s t-test (a, e) One-Way ANOVA (c, d) Two-Way ANOVA. Error bars represent SEM. Source data are provided as a Source Data file.
Caspase-11 activation, accentuated by increased GBP expression, controls pyroptosis and subsequent inflammation in Polg D257A macrophages
Our data suggest that potentiation of caspase-11-mediated pyroptosis drives excessive cell death and inflammation in PolgD257A macrophages challenged with PAO1. Therefore, we hypothesized that elevated levels GBPs may contribute to the enhanced release of phagolysosomal PAO1 LPS for sensing by caspase-11 in the cytoplasm. We next used siRNA to silence caspase-11 or GBP2 and GBP5 in WT and PolgD257A BMDMs to repress this pathway. Knockdown efficiency of siRNAs was confirmed by qPCR of RNA at the time of infection (Supplementary Fig. 6a). Notably, caspase-11 silencing reduced elevated cell death, heightened cleavage of both GSDMD caspase-1, and increased secretion of IL-1 cytokines from PolgD257A BMDMs 2 h post challenge with PAO1, while having little effect on WT BMDMs (Fig. 4a–d). Similar results were obtained by silencing GBP2 and GBP5 (Fig. 4a–d). Furthermore, heightened caspase-11 association with LPS in PolgD257A BMDMs was reduced upon either caspase-11 or GBP silencing, although to a lesser extent after GBP knockdown (Fig. 4e). Caspase-11 or GBP knockdown reduced secretion of proinflammatory cytokines in PolgD257A BMDMs at the 24 h timepoint (Fig. 4f). Finally, we utilized an LPS-deficient PAO1 mutant to further investigate caspase-11 activation. The PA algC gene product is important for synthesis of the LPS core, and therefore this mutant displays altered lipid A levels and lower LPS overall46, which we confirmed by LPS staining intensity as measured with an anti-PAO1 LPS antibody (Supplementary Fig. 6b). Indeed, PolgD257A BMDMs challenged with PAO1-algC displayed reduced cell death, inflammasome activation, IL-1β release, IFN-I secretion, and pro-inflammatory cytokine production (Supplementary Fig. 6c–g), further supporting that increased detection of PAO1 LPS by caspase-11 is a predominant driver of hyperinflammatory phenotypes in PolgD257A macrophages.
a Percent BMDMs dying after being given an siRNA control or siRNA to knock down caspase-11 or GBPs and then challenged with PAO1 from (b) representative IF. N = 8–9 IF images/genotype/condition. c Western blot analysis of lysates from BMDMs challenged with PAO1 after siRNA knock down of caspase-11. d ELISA analysis of IL-1 cytokines in BMDM supernatant after being given an siRNA control or siRNA to knock down caspase-11 or GBPs and then challenged with PAO1. N = 3–4 biological replicates/genotype/condition. e Quantification of caspase-11 associated with LPS in BMDMs after knock down of caspase-11 or GBPs by siRNA and then challenge with PAO1. N = 5 IF images/genotype/condition. f ELISA analysis of inflammatory cytokines in BMDM supernatant after being given an siRNA control or siRNA to knock down caspase-11or GBPs and then challenged with PAO1. N = 4 biological replicates/genotype/condition. Statistics: (a, d, e, f) One-Way ANOVA. Error bars represent SEM. Source data are provided as a Source Data file.
Increased intracellular availability of LPS enhances caspase-11 activation and initiation of IL-1 signaling in Polg D257A macrophages
Transmission electron microscopy (TEM) revealed that while PolgD257A BMDMs have similar nuclear and cellular area at baseline (Supplementary Fig. 7a, b), PolgD257A BMDMs present condensed nuclei and enlarged phagolysosomes 2 h post challenge with PAO1 (Fig. 5a, b and Supplementary Fig. 7c). Mitochondrial dysfunction can drive lysosomal defects and leakiness47,48,49, and we observed abnormal phagolysosomes in PolgD257A BMDMs by TEM two hours after PAO1 (Fig. 5a, b). We also observed increased presence of vesicles adjacent to PAO1 in PolgD257A phagolysosomes, which resembled bacterial outer membrane vesicles (OMV)50,51. At eight hours post PAO1 challenge, PolgD257A BMDMs contained more intact bacteria with OMV-like structures that were contained in phagolysosomes with poorly defined membranes (Supplementary Fig. 7e). This was accompanied by increased PAO1 protein expression as assessed by proteomics analysis (Supplementary Fig. 7f). Oxidative stress is a critical driver of PA OMV generation, and we noted increased reactive oxygen species20 and nitrite production from PolgD257A BMDMs after PAO1 challenge (Supplementary Fig. 7g). Therefore, we hypothesized that caspase-11 hyperactivity in PolgD257A macrophages may be due to increased sensing of LPS on OMVs or lingering bacteria, leading to more robust recruitment and activation of caspase-11. Quantitation of IF imaging showed a greater intensity of caspase-11 and GBP5 colocalizing with PAO1 in PolgD257A BMDMs (Fig. 5c, d), which was not observed after challenge with LPS-deficient PAO1-algC (Supplementary Fig. 7h, i). To determine if these phenotypes were in part due to LPS uptake and/or degradation kinetics, we utilized a fluorescently labeled LPS. Notably, WT and PolgD257A BMDMs took up comparable amounts of LPS after LPS priming followed by LPS transfection at acute timepoints (Supplementary Fig. 8a, b). However, we observed more fluorescent LPS in the cytosol seven hours after LPS transfection of LPS primed BMDMs (Supplementary Fig. 8c, d), as well as significantly more caspase-11 associated with LPS by IF analysis (Supplementary Fig. 8e, f). WT and PolgD257A BMDMs contained similar levels of fluorescent LPS signal immediately after passive uptake from the culture media (Supplementary Fig. 8g–h), and Polg mutant BMDMs did not release IL-1β at early timepoints (Supplementary Fig. 8k). However, 48 h after the addition of LPS to the cell culture media, PolgD257A BMDMs had a marked increase of cell-associated LPS (Supplementary Fig. 8i, j) and released small amounts of IL-1β (Supplementary Fig. 8k). Together, these data suggest that elevated levels of GBPs and caspase-11 in PolgD257A macrophages, coupled with increased bacterial OMVs and LPS in the cytosol, promote enhanced caspase-11 activation, leading pyroptosis and hyperinflammation.
a Quantitation of PAO1 and lysosomes in BMDMs from (b) TEM images. Black arrows highlight outer membrane derived vesicles. N = 9 images/condition. c Quantification of capase-11 and GBP5 intensity in the area containing caspase-11, GBP5, and PAO1 from representative IF images (d). N = 10–12 IF images/genotype/condition. e ELISA analysis of IL-1 cytokines in BMDM supernatant after PAO1 or PAO1 plus anti-IL-1R blockade 2 hpi. N = 4 biological replicates/genotype/condition. f Percent BMDMs dying after PAO1 and PAO1 plus anti-IL-1R blockade from (Supplementary Fig. 9b) representative IF. N = 11–13 IF images/genotype/condition. g ELISA analysis of IL-1 cytokines in BMDM supernatant after PAO1 or PAO1 plus anti-IL-1R blockade 24 hpi. N = 4 biological replicates/genotype/condition. h ELISA analysis of inflammatory cytokines in BMDM supernatant after PAO1 or PAO1 plus anti-IL-1R blockade 24 hpi. N = 4 biological replicates/genotype/condition. Statistics: (a, c) Two-Sided Student’s t-test (e–h) Two-Way ANOVA. Error bars represent SEM. Source data are provided as a Source Data file.
In WT macrophages, caspase-11 and caspase-1 are not directly involved in the generation of nuclear-factor kappa B (NF-κB)-dependent cytokines such as TNFα and IL-6. Therefore, we hypothesized that elevated caspase-11 activity may potentiate autocrine or paracrine signaling via the early release of alarmins that hyperactive PA-induced IFN-I or NF-κB. A kinetic stimulation study revealed increased phosphorylation of factors involved in IFN-I signaling (i.e., TBK1, STAT1) in PolgD257A BMDMs at early timepoints after PAO1 challenge, as well as elevated phospho-IκBα levels at early timepoints (Supplementary Fig. 9a). As mentioned above, IL-1α release is associated with caspase-11 activation and GSDMD cleavage52. IL-1α signaling through the IL-1 receptor (IL-1R) activates NF-κB and induces transcription of pro-IL-1β, inflammasome components including NLRP3, and other proinflammatory cytokines including TNFα and IL-653,54,55. To determine if early caspase-11-dependent IL-1α signaling feeds forward to potentiate cytokine secretion in PolgD257A BMDMs at later timepoints, we used a neutralizing antibody to block IL-1R signaling. Notably, the IL-1R antibody did not reduce elevated IL-1 cytokine release, cell death, or GSDMD and caspase-11 cleavage in PolgD257A BMDMs two hours post challenge (Fig. 5e, f, Supplementary Fig. 9b, c). However, IL-1R blockade was sufficient to markedly decrease IL-1 cytokines, IL-6, and TNFα from PolgD257A BMDMs twenty-four hours post PAO1 challenge (Fig. 5g, h). Overall, these data indicate that the rapid, caspase-11-dependent release of IL-1α after PAO1 challenge drives feed forward NF-κB and IFN signaling to potentiate inflammatory cytokine production in PolgD257A macrophages.
A Polg R292C mouse model incorporating a deleterious patient variant exhibits macrophage mtDNA depletion, elevated IFN-I responses, increased pyroptosis, and hyper-inflammation after challenge with PAO1
To determine if our findings in PolgD257A macrophages are more broadly applicable to other models of OXPHOS deficiency due to mtDNA perturbations, we leveraged a model of Polg-related mitochondrial disease harboring a single amino acid missense mutation (Arg292Cys) in exon 4 of the exonuclease domain of Polg (Supplementary Fig. 10a), which is homologous to the human Arg309Cys (c.925 C > T) pathogenic variant56,57. We generated these mice using CRISPR/Cas9 editing of embryonic stem (ES) cells and verified germline transmission of the mutation by sequencing (Supplementary Fig. 10b). The human Arg309Cys variant alone or in combination with other Polg variants causes reduced mtDNA copy number, mtDNA deletions, and severe phenotypes including mitochondrial myopathy, encephalopathy, lactic acidosis and stroke-like episodes (MELAS), peripheral neuropathy, ataxia, and epilepsy in patients. Although being born below expected Mendelian ratios, homozygous PolgR292C mice are viable and display reduced mitochondrial DNA copy number and elevated ISGs in multiple tissues (Supplementary Fig. 10c–e).
BMDMs derived from PolgR292C mice exhibited mtDNA depletion at rest (Supplementary Fig. 11a), as well as lower numbers of nucleoids and fragmented mitochondrial networks after stimulation with LPS (Supplementary Fig. 11b, c). In line with our data from the PolgD257A mutator mice, PolgR292C BMDMs displayed a constitutive elevation of ISGs at rest and potentiated ISG expression after LPS stimulation (Supplementary Fig. 10d, e). We also noted markedly reduced basal and maximal respiration after LPS stimulation (Supplementary Fig. 11f–h), indicating that PolgR292C macrophages exhibit OXPHOS deficiencies resembling those of PolgD257A BMDMs (Supplementary Fig. 4l, m).
After challenge with PAO1, PolgR292C BMDMs exhibited increased ISG and GBP expression (Fig. 6a), while also secreting more cytokines over a twenty-four-hour time course (Fig. 6b). Similar to PolgD257A BMDMs, PolgR292C exhibited elevated rates of PAO1-induced cell death, IL-1 cytokine secretion, and proinflammatory cytokine production, which were blunted by treatment with caspase-1 inhibitor AcYVAD (Fig. 6c–e, Supplementary Fig. 12) or IFNAR blockade (Fig. 6f–h). Finally, silencing of caspase-11 in PolgR292C BMDMs (Fig. 6i) alleviated acute cell death (Fig. 6j–k), IL-1 release (Fig. 6l), and secretion of IL-6 and TNFα at late timepoints (Fig. 6m). These data suggest that elevated IFN-I, hyperinflammation, and pyroptosis are common features of macrophages with deficient Polg exonuclease activity, including the MtD-relevant Arg292Cys variant.
a qRT-PCR analysis of ISGs in BMDMs after PAO1, N = 3 technical replicates/genotype. b ELISA analysis of inflammatory cytokines in BMDM supernatant after PAO1. N = 12 biological replicates over 3 time points. c Percent BMDMs dying after PAO1 and PAO1 plus caspase-1/11 inhibitor. N = 7 IF images/genotype/condition. d ELISA analysis of IL-1 cytokines in BMDM supernatant after PAO1 or PAO1 plus caspase-1/11 inhibitor. N = 3 biological replicates/genotype/condition. e ELISA analysis of proinflammatory cytokines in BMDM supernatant after PAO1 or PAO1 plus caspase-1/11 inhibitor. N = 3 biological replicates/genotype/condition. f Percent BMDMs dying after PAO1 and PAO1 plus anti-IFNAR blockade from. N = 7 IF images/genotype/condition. g ELISA analysis of IL-1 cytokines in BMDM supernatant after PAO1 or PAO1 plus anti-IFNAR blockade. N = 4 biological replicates/genotype/condition. h ELISA analysis of pro-inflammatory cytokines in BMDM supernatant after PAO1 or PAO1 plus anti-IFNAR blockade. N = 4 biological replicates/genotype/condition. i Quantification of caspase-11 in BMDMs after knock down by siRNA and PAO1 challenge. N = 5 IF images/genotype/condition. k Percent BMDMs dying after PAO1 and PAO1 plus caspase-11 knock down by siRNA from (j) representative IF. N = 8 IF images/genotype/condition. l ELISA analysis of IL-1 cytokines in BMDM supernatant after PAO1 or PAO1 caspase-11 knock down by siRNA. N = 3 biological replicates/genotype/condition. m ELISA analysis of pro-inflammatory cytokines in BMDM supernatant after PAO1 or PAO1 caspase-11 knock down by siRNA. N = 3 biological replicates/genotype/condition. Statistics: (a) Two-Sided Students t-test. (b) Two-Way ANOVA. (c–i, k–m) One-Way ANOVA. Error bars represent SEM. Source data are provided as a Source Data file.
Intratracheal instillation of a non-lethal dose of PAO1 elicits greater lung cytokine production and alterations in lung immune cell populations in Polg D257A mice
Patients with primary MtD often experience recurrent viral and bacterial infections of the respiratory tract, including infections by PA5. These events can trigger metabolic decompensation and can progress to systemic inflammatory endpoints and/or sepsis. However, few studies have examined infection kinetics and lung immune responses in Polg-related mitochondrial disorders. We therefore employed intratracheal instillation protocols to administer LPS or PAO1 into the lungs of PolgD257A mutator mice. Histological analysis of resting PolgD257A lungs did not reveal obvious differences in morphology or cellular composition (Supplementary Fig. 13a–c), although consistent with our earlier findings20, basal expression of ISGs were elevated compared to WT littermates (Supplementary Fig. 13d). After LPS was instilled into lungs through cannulation of the trachea, PolgD257A mice secreted more TNFα and IL-6 at all timepoints and increased IL-1β at the 12-h timepoint in the bronchial alveolar lavage fluid (BAL) compared to WT mice (Supplementary Fig. 13e).
To next characterize lung innate immune responses to PAO1 infection in the PolgD257A mice, we challenged mice with a non-lethal dose of PAO158. Both WT and PolgD257A mice infected with 1 × 104 colony forming units (CFU) of PAO1 via intratracheal instillation lost weight initially but returned to normal weight three days post infection (Supplementary Fig. 14a). Flow cytometric analysis of lung cells in BAL fluid revealed an overall decrease in the percentage of CD45+ immune cells at rest and after infection, consistent with general leukopenia noted in PolgD257A mice20,25 (Supplementary Fig. 14b, c). Analysis of specific immune cell markers on lung CD45+ cells revealed few changes in BAL populations (Supplementary Fig. 14d–l) between uninfected WT and PolgD257A mice, although there were more SiglecF+ and F4/80+ cells at rest perhaps indicating elevated numbers of lung macrophages (Supplementary Fig. 14e, g). Six hours post infection, there was a marked reduction in SiglecF+ cells in PolgD257A mice relative to WTs and an increase in CD3+ T cells (Supplementary Fig. 14e, i). Cells positive for the neutrophil marker Ly6G were dramatically increased at 6 h, although there were no differences between WT and PolgD257A mice (Supplementary Fig. 14k). At 48 h post infection, neutrophils remained the most predominant CD45+ population in both genotypes, but we observed a reversion toward increased numbers of cells staining positive for macrophage markers SiglecF+ and F4/80+ in PolgD257A mice (Supplementary Fig. 14e, g, k). Despite modest changes in cellular composition of the BAL fluid, PolgD257A mice had elevated numbers of CD45+ cells expressing TNFα both at rest and six hours post infection (Supplementary Fig. 14m), consistent with increased TNFα production after LPS challenge.
Multianalyte analysis of the BAL fluid using Legendplex assays revealed two to four fold increases in lung pro-inflammatory cytokines in PolgD257A mice 6 h post PAO1 infection (Fig. 7a, Supplementary Fig. 15a). Consistent with resolution of infection in both genotypes, cytokine differences in the BAL of WT and PolgD257A mice were less pronounced at 48 h post infection (Supplementary Fig. 15b). However, IFNβ and inflammatory cytokines associated with the skewing and differentiation of T cell Th1 (IL-27) and Th17 subsets (IL-23) remained elevated in PolgD257A mice in an IFNAR-dependent fashion (Supplementary Fig. 15b). Analysis of lung protein extracts revealed elevated caspase-1 cleavage, as well as increased caspase-11, NLRC4, and NLRP3 expression in PolgD257A mice 48 h post intratracheal instillation of 1 × 104 CFU of PAO1. GSDMD, GBP5, and other ISGs were also elevated in PolgD257A lungs, consistent with sustained IFNβ expression observed by Legendplex (Supplementary Fig. 15c).
a LegendPlex pro-inflammatory cytokine analysis of bronchial alveolar lavage (BAL) 6 h post infection (hpi) with 1 × 104 colony forming units (CFU) of PAO1. N = 5 mice/genotype. b Ingenuity Pathway Analysis (IPA) of RNA-seq from lung showing Z-scores of pathways in PolgD257A homozygous mutant BMDMs compared to wild-type (WT) after PAO challenge. N = 3 biological replicates/genotype/condition. c IPA Upstream Regulator analysis showing Z-scores in PolgD257A homozygous mutant lungs compared to WT after 6 h PAO1 challenge. d Log2 fold changes of representative genes in IPA pathways from (a) in PolgD257A lungs compared to WT after PAO1. e Protein expression or WT and PolgD257A lung homogenates 6 hpi with 1 × 105 CFU of PAO1 and densitometry. N = 4 mice/genotype. f Colony counts from homogenized tissue infected with 1 × 105 PAO1 for 6 h. N = 4 mice/genotype. g Colony counts from homogenized tissue infected with 1 × 105 PAO1-algC for 6 h. N = 4 mice/genotype. h ELISA analysis of pro-inflammatory cytokines in BAL of mice infected with 1 × 105 PAO1- algC for 6 h. N = 4 mice/genotype. i Representative hematoxylin and eosin (H&E) staining of lung sections 20 hpi with 1 × 105 CFU of PAO1. j Graphs of pathology scoring from H&E stained lungs in (i) histology. N = 5 mice/genotype. k Protein expression or WT and PolgD257A lung homogenates 20 hpi with 1 × 105 CFU of PAO1. l Illness scoring of WT and PolgD257A mice over PAO1 infection time course. N = 4–5 mice/genotype. (m) Kaplan-Meier survival curve after infection with 1 × 105 CFU of PAO1. N = 4–5 mice/genotype. Statistics: (a, e–h, j) Two-Sided Students t-test (l) Two-Way ANOVA (m) Kaplan-Meijer Survival Curve. Error bars represent SEM. Source data are provided as a Source Data file.
Given that recurrent respiratory tract infections are often observed in patients with MtD, we next infected mice with 1 × 104 CFU PAO1, followed by a 7-day rest period, then re-infected with 1 × 104 CFU PAO1. Histological analysis of lungs 24 h following re-infection revealed monocyte and granulocyte infiltration into lungs, areas of vascular hemorrhage, and more eosin positive cells in PolgD257A lungs relative to WT (Supplementary Fig. 16a, b). Lung IL-12 family cytokines and IFNβ were significantly higher in PolgD257A mice at the 24-h harvest point following re-infection, while other cytokines were trending higher in PolgD257A mice, although they did not reach statistical significance (Supplementary Fig. 16c). We also observed higher bacterial burdens in the lungs and spleens of PolgD257A mice compared to WT controls (Supplementary Fig. 16d). Together, these data indicate that PolgD257A mice exhibit hyperinflammation and impaired antibacterial immunity upon re-infection with PAO1.
PAO1 elicits greater caspase-11 expression, increases inflammation, and reduces survival of Polg mutant mice
We next utilized a higher dose of PAO1 to further assess innate immune responses Polg mutant mice. RNAseq of lung tissue collected six hours after intratracheal instillation of 1 × 105 CFU PAO1 revealed a striking upregulation of many inflammatory-, IFN-, and programmed cell death-related pathways in PolgD257A mice relative to WT littermates (Fig. 7b, c, Supplementary Fig. 17a). In agreement with macrophage data, transcriptional signatures of pyroptosis were apparent, and Casp11, Nlrp3, Il1a, and Il1b were some of the most abundantly enriched transcripts in PolgD257A versus WT lung lysates (Fig. 7d). PolgR292C lung homogenates also displayed upregulation in transcriptional pathways indicative of hyperinflammation, elevated IFN signaling, and potentiated pyroptotic cell death 6 h post PAO1 infection (Supplementary Fig. 17b–d). Western blotting of PolgD257A lung homogenates reinforced RNAseq data, showing increased protein expression of pyroptotic factors including caspase-11 and cleaved GSDMD (Fig. 7e), as well as a depletion in OXPHOS Complex I and IV subunits 6 h post infection (Supplementary Fig. 17e). Moreover, PolgD257A mice exhibited elevated bacterial burdens in the lung, liver, and spleen (Fig. 7f). Increased bacterial dissemination was accompanied by elevated plasma proinflammatory cytokines, as well as elevated ISG expression with reduced OXPHOS in PolgD257A liver homogenates (Supplementary Fig. 17f–g). We next utilized the PAO1-algC mutant to further define whether reduced LPS sensing via caspase-11 would alter response to infection in PolgD257A mice. Compared to WT PAO1, PAO1-algC did not disseminate (Fig. 7g), or trigger increased expression of pyroptotic proteins and cytokines in PolgD257A lungs relative to WTs (Fig. 7h, Supplementary Fig. 17h). Overall, these data suggest Polg mutant mice display increased caspase-11-medated sensing of PAO1 LPS, elevated bacterial burdens, and hyperinflammatory responses during acute PAO1 infection
At later timepoints, PolgD257A lungs exhibited elevated neutrophil influx and inflammation via histological analysis (Fig. 7i, j, Supplementary Fig. 17i), despite harboring similar bacterial burdens to WT mice (Supplementary Fig. 17j). However, caspase-11 and IL-1β remained elevated in lung lysates of PolgD257A mice compared to WT littermates twenty hours post infection (Fig. 7k). Using a comprehensive health scoring system to track breathing, hydration, body appearance, and activity, we noted that PolgD257A mice appeared sicker and continued to decline in health, whereas WT animals recovered from infection (Fig. 7l). Even though we observed a steady decline in the health of PolgD257A mice at 48 h after infection, PAO1 were largely cleared in both PolgD257A and WT animals (Supplementary Fig. 17k). However, PolgD257A mice sustained elevated ISG expression in peripheral tissues, suggesting sustained IFN-I signaling (Supplementary Fig. 17l, m). Finally, we observed that a greater number of PolgD257A mice reached a moribund state after infection with 1 × 105 CFU of PAO1 relative to WT littermates (Fig. 7m). Overall, these in vivo data suggest that PolgD257A mice are more prone to excessive inflammation and acute lung injury following infection with PAO1.
Discussion
Patients with MtD experience a greater frequency of bacterial and viral infections that can rapidly progress to sepsis or systemic inflammatory response syndrome (SIRS), leading to premature death5. Recurrent infections can also accelerate MtD progression by promoting metabolic decompensation, a life-threatening condition that hastens deterioration of mitochondrial function59,60,61. Polg is the sole mitochondrial DNA polymerase, and variants in the POLG gene are a prevalent cause of MtD62. There are several clinical reports of bacterial or viral infections unmasking Polg-related MtD symptoms and hastening disease progression63,64. Moreover, one study revealed the presence of inflammatory cytokines in the cerebrospinal fluid of a patient with Polg-related MtD12,19. As cytokine-mediated toxicity is an important driver of metabolic decompensation7, there is a critical need to understand immune responses in patients with Polg-related disorders and other MtD to manage infections and inflammation-related complications. This study builds upon our prior work to characterize innate immune responses in mouse models of MtD20,65 and leverages bacterial infection, cytokine profiling, and in vitro and ex vivo cellular assays to define molecular and cellular nodes governing hyperinflammatory immune phenotypes in two models of Polg-related MtD (Supplementary Fig. 18).
Our prior work revealed that elevated IFN-I responses in PolgD257A mutator mice are mediated by chronic mtDNA release and cGAS-STING signaling20. Using our disease-relevant PolgR292C mouse model, we observed increased IFN-I signatures basally and after challenge with PAO1, similar to PolgD257A. Analogous variants in human POLG at R309 affect proofreading activity66, and therefore, we hypothesize that the PolgR292C exonuclease mutation also impairs replication fidelity, leading to mtDNA release and chronic IFN-I signaling. A recent study reported that fibroblasts from patients carrying the POLG1 c.2243G>C, p.W748S mitochondrial recessive ataxia syndrome (MIRAS) allele display reduced IFN-I signaling in response to viral infection and nucleic acid stimuli67. The MIRAS variant affects POLG holoenzyme processivity but not the exonuclease domain, and thus impaired exonuclease activity leading to mtDNA mutations and deletions may selectively trigger IFN-I in the PolgD257A and PolgR292C backgrounds. Despite reduced IFN-I, mice homozygous for the MIRAS variant do exhibit increased necroptosis, elevated liver immune cell infiltration, and potentiated inflammatory cytokine secretion during viral infection67. Thus, inflammatory cell death and hyperinflammation are common features in both patient cells and mouse models of Polg-related MtD.
Our data reveal that IFN-I signaling is a key driver of caspase-11-dependent pyroptosis and hyperinflammation in Polg mutant macrophages. Pre-treatment with an anti-IFNAR neutralizing antibody was sufficient to reduce both augmented PI uptake after LPS transfection and cytokine hypersecretion after challenge with PAO1. IFN-I is critical to license caspase-11 expression41, but also enhances expression of interferon-inducible GTPases like GBPs, which help to expose cytosolic LPS for detection31,68,69,70,71. We have found that macrophages from both the PolgD257A and our MtD-relevant PolgR292C model exhibit enhanced expression of multiple GBPs at rest and after challenge with PAO1. Moreover, we observed that GBP5 strongly co-localizes with intracellular PAO1 and caspase-11 in PolgD257A BMDMs. GBPs are important in orchestrating caspase-11 activation to bacterial outer membrane vesicles (OMV)50,51 and can localize to damaged cell membranes45. Although the mechanisms by which GBPs sense intracellular pathogens and facilitate caspase-11 activation are not fully understood, silencing of caspase-11 or GBP2 and GBP5 markedly reduced pyroptosis and the secretion of IL-1 and other cytokines from both Polg mutant BMDMs. TEM imaging also revealed altered PAO1-containing phagolysosomal membranes and increased bacteria-associated vesicles in PolgD257A BMDMs. Thus, it is likely that GBPs and caspase-11 more readily sense PAO1 LPS due to phagolysosomal leakiness or elevated OMV generation, yet more work is required to fully define the underlying mechanisms.
Our data reveal that Polg mutant macrophages undergo rapid pyroptosis upon PAO1 challenge in a NLRP3- and caspase-1-dependent fashion. Although caspase-1 cleavage is elevated in PolgD257A compared to WT macrophages during PAO1 challenge, and its inhibition is sufficient to block augmented PA-induced cell death and IL-1β secretion, we found that PolgD257A BMDMs are less responsive to direct activation of the NLRP3 and NLRC4 inflammasomes by their respective ligands. In contrast, activation of the caspase-11 inflammasome via LPS priming followed by transfection of LPS into the cytosol drove more PI uptake and IL-1β expression in PolgD257A BMDMs. Therefore, sensing of cytosolic PA LPS by the non-canonical inflammasome leads to more robust activation of the NLRP3 for caspase-1 dependent cell death and cytokine secretion. Notably, a PAO1-algc mutant lacking robust LPS synthesis did not hyper-stimulate pyroptosis and cytokine secretion, providing further support for caspase-11 as the predominant driver of inflammatory cell death in Polg mutant mice.
It is interesting to note that treatment of Polg mutant macrophages with caspase-1 inhibitor dramatically reduced nearly all cytokine levels in culture supernatants after challenge with PAO1. These include inflammasome-independent cytokines such as TNFα, IL-6, IFNγ, and IL-12 family members as well as the DAMP IL-1α, which does not require caspase-1 cleavage for secretion but is downstream of caspase-11 and GSDMD37,72. This is in contrast with cytokine secretion in WT BMDMs, where AcYVAD treatment only reduced IL-1β levels during challenge with PAO1. Similarly, anti-IFNAR blockade or caspase-11 and GBP knockdown reduced hyper-secretion of IL-1β levels and reduced IL-6 and TNFα in Polg macrophages, while having marginal effects on WT BMDM cytokine profiles. While blocking the IL-1R did not reduce early cell death in Polg mutant macrophages, it was sufficient to blunt elevated cytokines a day after PAO1 challenge. This suggests that early IL-1α release downstream of capsase-11 and GSDMD synergizes with IFN-I signaling to sustain hyperinflammation in PAO1-challenged Polg mutant BMDMs. Our data do not definitively rule out contributions for canonical inflammasome-mediated IL-1β in potentiating NF-κB-mediated cytokine release. However, the link between caspase-11 activation and IL-1α suggests that the early release of this alarmin may prime neighboring cells for activation of canonical inflammasomes, further promoting increased cell death and inflammation in Polg mutant macrophages.
Although it is appreciated that bacterial and viral infections are problematic in MtD, few studies have examined innate immune responses in relevant mouse models of disease. Using an intratracheal instillation model, we found that PolgD257A mice exhibit hyperactive innate immune responses resulting in elevated cytokine production, changes in lung immune cell dynamics recruitment, and increased morbidity following PAO1 infection. Notably, we observed elevated expression of inflammasome proteins including caspase-11, ISGs including GBPs, and increased caspase-1 cleavage in PolgD257A lung homogenates after infection. Remarkably, RNAseq analysis of PA infected lungs from 3-month-old PolgR292C mice also revealed increased pyroptosis, interferon, and hyperinflammatory signaling. These data agree with our in vitro findings implicating IFN-I as an overarching regulator of caspase-1/11 activation and hyperinflammation in our models. Additional studies are warranted to define the cellular populations that govern lung hyperinflammation and to determine the degree to which caspase-11 shapes inflammatory signaling and morbidity in Polg mutant mice in vivo.
While PolgD257A mice had higher bacterial burdens in the lung and peripheral tissues at acute timepoints, they ultimately managed to control the infection by 20 h post infection and cleared bacteria by 48 h. However, lungs from PolgD257A mice were more inflamed at 20 h post infection and the mice had more severe sickness scores, resulting in higher rates of morbidity. This suggests that the adverse outcomes observed in Polg mutant animals may be due to chronic and damaging hyperinflammation as opposed to impaired bacterial clearance. Therefore, therapeutic targeting of IFN-I and caspase-11 may represent promising avenues to restore proper immune balance patients with Polg-related MtD. Although our work does not include clinical data, two recent publications revealed elevated IFN-I signatures in the whole blood and peripheral blood mononuclear cells (PBMCs) from patients with diverse MtD disease variants11,73. Moreover, gene ontology analysis revealed heightened IL-1β signatures in MtD PBMCs11. Our study provides important insight into the innate immune mechanisms governing infection-related hyperinflammation in mouse models of MtD. The consistency of our discoveries across two different Polg mutant models strengthens the relevance of our findings to human disease and suggest that IFN-I and inflammasome hyperactivity may be critical determinants of increased rates of sepsis and SIRS in patients with MtD.
Methods
Mouse husbandry and strains
C57BL/6 J (JAX Stock # 000664), PolgD257A/D257A (JAX Stock # 017341), and Ifnar1−/− (JAX Stock # 028256) mice were obtained from The Jackson Laboratory and bred and maintained in multiple vivaria at Texas A&M University and The Jackson Laboratory. Experimental Polg+/+ (WT) and PolgD257A/D257A littermates were obtained from PolgD257A/+ maintenance colony crosses of male PolgD257A/+ to female Polg+/+. This breeding strategy was adopted to limit the accumulation and maternal transmission of mutant mtDNA molecules to heterozygous breeders. PolgD257A/D257A mice and littermate controls used for macrophage experiments were aged for 12–16 months. PolgD257A/D257A mice and littermate controls used for in vivo infections were aged from 7 to 12 months.
The PolgR292C/R292C mouse line (B6N(Cg)-Polgem5(F290F,R292C)Lutzy/Mmjax) was generated using CRISPR/Cas9 endonuclease-mediated genome editing in mouse embryonic stem cells (mESCs). A single guide RNA (CTCCCATCCACAGGCTGCGC) targeting exon 4 of Polg was cloned into pX459, which expresses Cas9 nuclease. This plasmid was co-delivered to JM8A3 ESCs via electroporation with a single-stranded oligonucleotide donor (ssODN) that included the missense R292C (CGC>TGC) and an additional silent mutation (F290F; TTC>TTT) to reduce recutting of the targeted allele74. Following puromycin selection, resistant clones were screened for proper targeting and to exclude integration of the pX459 plasmid. Multiple clones were selected for expansion and karyotyping, and euploid clones were injected into C57BL/6J-albino blastocysts (B6(Cg)-Tyrc-2J/J; JAX STOCK# 000058) to produce chimeras. Chimeras were bred to C57BL/6NJ (JAX Stock #005304) mice, and a single germline clone was selected for expansion and characterization. The line is available through the MMRRC repository (071656-JAX). All PolgR292C/R292C mice and littermate controls used for macrophage experiments were aged for 5–8 months. PolgR292C/R292C mice and littermate controls used for in vivo experiments were aged for 3–4 months. Breeding colonies of PolgR292C mice were maintained similarly to PolgD257A models.
Mice were housed in 12 h dark, 12 h light cycles, at a temperature of 70 °F and 60% humidity. Sex was not considered in mouse experiments due the lack of observed or reported sex differences in the Polg mutant mice or patients with Polg-related MtD. All animal experiments were approved by the Institutional Animal Care and Use Committees (IACUC) at Texas A&M University and The Jackson Laboratory.
Pseudomonas aeruginosa (PAO1) in vivo infection
7–12 month old mice were moved into BSL2 procedure space and allowed to acclimate for 1–2 days. Mice were anesthetized with an intraperitoneal injection of 100–10 mg/kg Ketamine-Xylazine cocktail. Mice were then intubated with a 24 G catheter. Sterile PBS (sham), 0.5 mg/kg LPS (E. Coli serotype 0111:B4; Invivogen), or the specified dose of PAO1-GFP was pipetted into the catheter in a 25–30 µL volume. A bolus of air was delivered using a syringe to ensure the dose evenly coated the lungs. Mice were kept warm until fully recovered from anesthesia and monitored daily until experimental timepoints were reached.
Bronchial alveolar lavage (BAL) fluid collection
Mice were euthanized using intraperitoneal injection of 200 mg/kg Pentobarbital Sodium to preserve the lung environment and structure of the tracheal. Upon confirmation of death the chest cavity and throat were opened. The trachea was canulated using a 23 G needle threaded with tubing and the lung was washed with 3 mL cold Hanks’ Balanced Salt Solution (HBSS, Thermo Fisher Scientific, 14175-129) containing 100 µM EDTA (VMR Life Science, E522) and used in subsequent flow cytometry and LegendPlex assays.
Flow cytometry
Bronchial alveolar lavage fluid (BAL) was spun at 350 g for 5 min to pellet the cells. Cells were transferred to a V bottom plate for staining. Cells were stained with viability dye then Fc receptors were blocked with anti-mouse CD16/CD32 (2.4G2) antibody. Surface proteins were stained with antibodies specified in Supplementary Table 6. Cells were permeabilized with Foxp3/Transcription Factor Staining Buffer Kit (TNB-0607-KIT, Tonbo) and stained with antibodies against intracellular proteins. Flow cytometry analysis was done using a 5 L Cytek Aurora and FlowJo v10.9.0. A schematic of the gating strategy that was used is included in Supplementary Fig. 19.
Colony forming unit counts from macrophages and tissues
BMDMs or tissues were homogenized in 0.1% NP40 in sterile PBS using an immersion homogenizer for 15 s per tissue. Serial dilutions were plated on soy agar containing 100 μg/mL carbenicillin. Plates were incubated at 37 °C overnight and colonies were counted.
Histology
Tissues were collected in cassettes, then were washed in PBS, incubated for 24 h in 10% formalin, and transferred to 70% ethanol. Tissue embedding, sectioning, and staining were performed at AML Laboratories in Jacksonville, FL, or at the histology core at Texas A&M University. Images of hematoxylin and eosin (H&E) staining were collected using an Olympus slide scanner with a 20X objective. Slides were analyzed and scored by a pathologist, Dr. Jessica Wong, at the Jackson Laboratory in Sacramento, CA. Briefly, semi-quantitative histologic scoring from 0 to 4 severity was performed on neutrophilic and mononuclear cell inflammation. 0 = no inflammation, 1 = minimal inflammation including individual cells or small clusters, 2 = mild inflammation including small clusters and increased perivascular and peribronchiolar cuffing, 3 = moderate inflammation including multifocal to coalescent areas and perivascular cuffing, and 4 = severe inflammation including coalescent to patchy-diffuse inflammation.
Body condition scoring
Mice were monitored three times daily and were assigned a body condition score based on an additive score of appearance, behavior, hydration, and respiration. A score of 0 corresponds to a healthy mouse, and a score of 10 corresponds to a moribund animal requiring euthanasia. Animals were given an appearance score of 0 if normal, 1 if they lacked grooming or ruffled fur, 2 if they displayed piloerection, and 3 if they displayed hunched posture. Animals were given a behavior score of 0 if they were socially active, 1 if they had decreased mobility, 2 if they showed increased isolation and only moved when prodded, and 10 if they did not ambulate with prodding. Animals were given a hydration score of 0 if the skin tent immediately returned to normal, 1 if the tent took 1–2 s to return to normal, 2 if the tent took greater than 2 s to return to normal, and 3 if this occurred accompanied by sunken and dry eyes. Animals were given a respiration score of 0 if they displayed a normal breathing pattern, 1 if they had an increased breathing rate with an abdominal component, 3 if their breathing appeared labored, and 10 if their breathing was agonal.
Cell culture
L929 cells were obtained from ATCC and maintained in DMEM (Sigma-Aldrich, D5796) and 10% fetal bovine serum (FBS; VWR, 97068-085). Primary bone marrow-derived macrophages (BMDMs) were isolated from femurs of 6–12-month-old WT, PolgD257A, and 2–6-month-old PolgR292C mice as described20. Briefly, whole femurs were crushed in media and passed through a 70 µm filter. Red blood cells were lysed through osmosis by resuspending in ACK lysis buffer (Quality Biological, 118-156-101) for 2 min. Remaining cells were passed through a 40 µm filter, counted, and plated on non-tissue culture treated dishes. Macrophages were differentiated over for 7 days using 30% L929 conditioned medium. Peritoneal macrophages (PerMacs) were collected from the peritoneal cavity via gavage 4 days after intraperitoneal injection of 3% brewer thioglycolate medium (Sigma-Aldrich, B2551) and plated in medium with 10% FBS. Unless stated, 6 × 105 BMDMs and 1.2 × 106 PerMacs per millimeter were used in in vitro experiments. Cells were plated and left to adhere overnight prior to challenge.
Pseudomonas aeruginosa (PAO1) in vitro infection
GFP-expressing PAO1 was provided by Dr. Carolyn Cannon and was grown on Soy agar plates (BD, 211043) with 100 μg/mL carbenicillin (Dot Scientific, 46000-5) overnight from a glycerol stock. A single colony was incubated in 5 mL Mueller Hinton broth (BD, 275730) with carbenicillin at 37 °C spinning overnight. Overnight culture was back diluted 1:5 and grown for 2 h until bacteria reached log-phase. Bacteria were washed with sterile phosphate-buffered saline (PBS) and opsonized for 20 min in mouse serum (Fisher, PI31880). After two more washes with PBS macrophages were infected at the corresponding MOI and spun at 250 g for 5 min to synchronize infection. For inhibitor experiments, cells were primed with 10 µM Ac-YVAD-cmk (Invivogen, inh-yvad), 10 µM Q-VD-OPh (Millipore Sigma, 551476), or 10 µM MCC950 (InvivoGen inh-mcc) for 3 h prior to challenge. For antibody blockade experiments, cells were primed with 20 µg/mL anti-IFNAR antibody (Bio X Cell, BE0241), anti-IL-1R (Bio X Cell, BE0256), or anti-IgG control (Bio X Cell, BE0083 or BE0091) antibodies for 4 h prior to challenge. One hour after infection, wells were washed and treated with 300 μg/mL gentamicin (VWR, 0304) for 30 min to inactivate extracellular bacteria. The remainder of the experiment was completed with media supplemented with 50 μg/mL gentamicin and inhibitor or blocking antibody when described.
Escherichia coli (E. coli) in vitro infection
E. Coli containing empty pcDNA3.1 vector for antibiotic resistance was grown on LB agar plates (Fisher, DF0401-17) with 100 μg/mL carbenicillin (Dot Scientific, 46000-5) overnight from a glycerol stock. A single colony was incubated in 5 mL LB broth (Fisher, BP1427-500) with carbenicillin at 37 °C spinning overnight. Overnight culture was back diluted 1:5 and grown for 2 h until bacteria reached log-phase. Bacteria were washed with sterile phosphate-buffered saline (PBS) and opsonized for 20 min in mouse serum. After two more washes with PBS macrophages were infected at the corresponding MOI and spun at 250 g for 5 min to synchronize infection. One hour after infection, wells were washed and treated with 300 μg/mL gentamicin (VWR, 0304) for 30 min to kill extracellular bacteria. The rest of the experiment was completed in DMEM + 10% FBS supplemented with 50 μg/mL gentamicin.
Inflammasome challenge
BMDMs were primed for 4 h with 20 ng/mL LPS-B5 Ultrapure (InvivoGen, tlrl-pb5lps). When used, caspase inhibitor Ac-YVAD-cmk (InvivoGen, inh-yvad) was added an hour after LPS, to allow for 3 h priming, at 10 μM concentration. To trigger the non-canonical and NLRC4 inflammasomes, respectively, polyethyleneimine (Alfa Aesar, 43896) was used to transfect 2 μg/mL LPS75 and 10 ng/mL PA flagella76 into the cytosol. To trigger the NLRP3 inflammasome, 5 μM ATP (Sigma, A6419) or 40 μM Nigericin (Cayman Chemical Company, 11437) was added to the media. When used, 20 µg/mL anti-IFNAR antibody (Bio X Cell, BE0241) or anti-IgG control (Bio X Cell, BE0083) were added to cells 5 h prior to LPS priming and added again upon LPS transfection.
Immunoblotting
Cells and tissues were lysed in 1% NP-40 lysis buffer supplemented with protease inhibitor and then centrifuged at 15,000 g for 10 min at 4 °C, and the supernatant was collected as the cellular lysate. A bicinchoninic acid assay (BCA) protein assay (Thermo Fisher Scientific, PI23235), was used to quantify protein, and 10−40 μg total protein was loaded into 10% SDS-polyacrylamide gels and transferred onto 0.22 μM polyvinylidene difluoride membranes. Membranes were dried to return them to a hydrophobic state and then incubated in primary antibodies diluted in 1x PBS containing 1% casein overnight at 4 °C rolling. Membranes were washed and incubated with horseradish peroxidase (HRP)- conjugated secondary antibodies for 1 h at room temperature, shaking. Membranes were developed with Luminata Crescendo Western HRP Substrate (Millipore, WBWR0500). Primary antibody information can be found in Supplementary Table 1.
LegendPlex multianalyte cytokine analysis
Supernatants from macrophages were subjected to centrifugation prior to the assay to remove detached cells and debris from the sample. The assay was completed using the LegendPlex Mouse Inflammation Panel (BioLegend, 740446) according to the included protocol. Samples were analyzed by flow cytometry using a 5 L Cytek Aurora and the BioLegend analysis software. Samples were normalized to total protein content of each sample determined by a bicinchoninic acid assay (BCA) protein assay, and MFI from the blank control beads was subtracted to exclude background.
Enzyme-linked immunosorbent assay (ELISA)
Supernatants were collected from macrophages plated on glass coverslips. Macrophages were stained for IF analysis while supernatants were subjected to centrifugation to remove detached cells and debris from the sample. Samples were stored at −80 °C until use in this assay and were thawed on ice upon use. The assay was completed in 96-well half area clear, flat bottom plates (USA Scientific, 5667-5061), and wells were incubated overnight at 4 °C in capture antibodies. Blocking was completed for one hour with PBS containing 10% FBS at room temperature. Standards and samples were added to wells for two hours. All sample dilutions were made in blocking buffer. Next, wells were incubated in detection antibody for one hour. Avidin-HRP and trimethyl boron (TMB) were used for quick color development. Plates were washed with PBS containing 0.05% Tween 20 between each step. Capture and detection antibodies were used in the following pairs from BioLegend (Supplementary Table 2). Samples were normalized to corresponding nuclei counts from 20X IF images taken on an EDLIPSE Ti2 (Nikon). Nuclei counts were determined using the automated bright spot counting function in NIS-Elements on DAPI, and an average of 3 images was used to normalize ELISA. Normalization of BAL from PAO1-algC infected mice was normalized to protein concentration. ELISA concentrations were determined using an epoch plate reader (BioTek) and BioTek Gen5 software. Normalized results were graphed in GraphPad.
Quantitative polymerase chain reaction (qPCR)
Relative gene expression with measured by qRT-PCR in cells. Total cellular RNA was isolated using Quick-RNA microprep kit (Zymo Research, R1051). Approximately 0.5 µg of RNA was used to make cDNA with XLT qScript cDNA Synthesis Kit (Quantabio, 95047). cDNA was then used in a qPCR assay using PerfeCTa Green FastMix (Quantabio, 95071). Two to three biological replicates and two to three technical replicates were normalized to Rpl37 cDNA using the 2−∆∆CT method. All primer sequences can be found in Supplementary Table 3.
Immunofluorescence (IF) imaging
Cells were seeded at 2.5 × 105 cell/well in a 24 well plate and allowed to grow overnight on glass coverslips sterilized with ethanol. After challenge or infection, cells were washed in PBS and fixed with 4% paraformaldehyde for 20 min. Next cells were permeabilized with 0.1% Triton X-100 in PBS for 5 min, blocked with PBS containing 5% FBS for 30 min, stained with primary antibodies (Supplementary Table 4) for 1 h, stained with secondary antibodies for 1 h, and stained with 200 ng/mL 4′,6-diamidino-2-phenylindole (DAPI) (ThermoFisher Scientific, 62247) for 2 min. When described, coverslips were stained with 2 µg/mL CellMask (ThermoFisher Scientific, C10046) for 10 min after secondary antibody staining and washed multiple times in PBS before DAPI staining. Coverslips were mounted with ProLong Diamond Antifade Mountant (Invitrogen, MP33025) and sealed with nail polish. Cells were imaged with a 40x or 60x oil-immersed objective with an ECLIPSE Ti2 widefield microscope (Nikon) and analyzed with the NIS-Elements AR 5.21.02 software.
Quantification was done from multiple intensity projection (MIP) images generated from Z-stacks. MIPs were made, and analysis was done using NIS-Elements AR 5.21.02. Specifically, cell death was determined using the automated bright spot counting function in NIS-Elements on DAPI to count the number of nuclei. Dying cells were manually determined by identifying cells with condensed nuclei and low CellMask staining. Percentage of dying cells was determined and graphed in GraphPad.
Protein intensities were quantified by thresholding a mask over each channel and exporting sum intensities and area of masks. The sum intensities were normalized to area or cell numbers, where described. When normalized to cell number, cells were counted using the automated bright spot counting function in NIS-Elements on DAPI. To quantify intensities of colocalized proteins, a mask was generated over each protein. Next, a new mask was made for the overlap of these channels, and the intensity and area of each channel in this mask was exported and graphed.
To quantify nucleoids, whole cells were outlines and made into a region of interest (ROI), and these ROIs were numbered. A thresholding mask was made on the channels with TFAM and DNA staining, and channel intensities were determined for these masks and graphed. A new mask was made based on the overlap of TFAM and DNA. The area of this mask and number of spots per ROI were exported and graphed in GraphPad. Graphs and statistics were generated in GraphPad. Huygens Essential software (v21.4.0) was used to deconvolve representative images from z-stacks taken with a 60X objective in oil immersion.
Lactate dehydrogenase (LDH) release
Lactate dehydrogenase (LDH) was measured in the supernatant of PAO1 challenged macrophages using a LDH-Cytox Assay Kit (Biolegend, 426401) according to manufacturer’s recommendations. A set of cells was treated with 1% Triton X-100, and the supernatant was used to determine total cellular LDH.
Propidium Iodide (PI) uptake
BMDMs in 96-well half-area glass-bottom imaging plates were seeded at 6 × 104 cells/well in 50 µL of Phenol-Red free DMEM + 10% FBS. The next day, cells were subjected to inflammasome challenge or infection with PAO1. Propidium iodide (PI, Alfa Aesar, J66584) was added to media and plates were immediately imaged every ten minutes using a BioTek Cytation V imager which maintains 37 °C and 5% CO2. PI positive nuclei were identified and counted using BioTek Gen5 software. Control wells treated with PI and Triton X-100 (to permeabilize cells) to for normalization. The difference between PI positive cells compared to the initial reading were graphed in line graphs using GraphPad Prism.
Cell fractionation and mitochondrial enrichment
Macrophages were lifted from cell culture dishes in PBS using a cell scraper. Cells were washed with PBS and then resuspended in Resuspension buffer (10 mM NaCl, 1.5 mM CaCl2, 10 mM Tris-Hcl, pH 7.5) with phosphatase and protease inhibitors. A dounce homogenizer was used to break cell membranes and liberate organelles. Mitochondria were bound to magnetic beads coated with TOMM20 antibodies isolated using a mitochondria isolation kit (Miltenyi Biotec, 120-009-492). The remaining cell components and unbound mitochondria were pelleted and run as the cellular fraction. All samples were lysed in 1% SDS lysis buffer and run on a western blot for analysis.
Extracellular flux assay
Mitochondrial respiration and glycolysis in BMDMs were measured using an Agilent XFe96 analyzer. Macrophages were plates at a denisity of 5 × 105 cells/well in DMEM supplemented with 10% FBS and 5% L929 in an Agilent Seahorse XF96 Cell Culture Microplate. When described, 20 ng/mL LPS was added 16 h prior to analysis. Cells were placed in XF assay medium immediately before analysis. ECAR and OCR measurements were recorded through sequential addition of 10 mM glucose, 1.5 µM oligomycin, 1.5 µM carbonyl cyanide p-trifluoromethoxy phenylhydrazone with 1 mM sodium pyruvate, and finally 833 nM antimycin A plus 833 nM rotenone. Data was analyzed and calculations were made according to77.
siRNA knock down in bone marrow derived macrophages
Bone marrow was isolated from mice as described above. On day four, macrophages were moved to 10 cm dishes. On day five, macrophages were transfected with 25 nM of Negative Control (51-01-14-03), mm.Ri.Casp4.13.1, mm.Ri.Gbp2.13.1, or mm.Ri.Gbp5.13.1 DsiRNAs (IDT) using Lipofectamine RNAiMAX (ThermoFisher Scientific, 13778100) in OPTIMEM (Gibco, 51985-034) according to company recommendations. Six hours after transfection, media was removed and replaced with fresh DMEM containing 10% FBS, 5% L929, and 20 µg/mL anti-IFNAR antibody (Bio X Cell, BE0241) to reduce IFN-I-mediated toxicity due to siRNA transfection. A day after transfection, cells were replated in cell culture plates in macrophage media without anti-IFNAR antibody, then used in infection experiments the next day as described above.
mtDNA copy number analysis
DNA was isolated from cell pellets or crushed tissues by boiling for 30 min in 50 mM NaOH lysis buffer and then neutralizing using 1 M Tris, pH8 solution. DNA was diluted to 2 ng/uL and utilized in qRT-PCR reaction (described above) with mtDNA specific primers. Two to three biological replicates and two to three technical replicates were normalized to nuclear encoded Tert using the 2−∆∆CT method. All primer sequences can be found in Supplementary Table 5.
Transmission electron microscopy (TEM)
Macrophages were lifted from cell culture dishes and pellets were fixed for 1 h in 4% PFA. After fixation, cells were rinsed in 0.1 M cacodylate buffer and then post-fixed in 1% osmium tetroxide (OsO4) in 0.1 M cacodylate buffer for 1 h at room temperature. Cells were rinsed again in the same buffer then spun down into pellets with 2% liquid agarose in distilled water and placed in a 4 °C refrigerator for 30 min, allowing the agarose to harden. After trimming the agarose cell pellets, samples were transferred through graded ethanol (40% ETOH to 100% ETOH), propylene oxide, and then infiltrated with 1:1 propylene oxide:Embed 812 resin (Electron Microscopy Sciences, Hatfield, PA) overnight, 1:3 propylene oxide:Embed 812 resin for 4 h, and finally 100% Embed 812 resin overnight. Tissues were embedded in 100% resin in beam capsules and cured at 65 °C for 48 h prior to sectioning. Thin sections (90 nm) were cut on a Leica EM UC7 ultramicrotome (Leica Microsystems, Buffalo Grove, IL.) with a diamond knife, placed onto 300 mesh copper grids, and stained using 2% uranyl acetate and Reynolds lead citrate. Samples were evaluated at 80 kV using a JEOL JM-1230 transmission electron microscope (JEOL, Tokyo, Japan) and images collected with an AMT Nanosprint15 MK-11 sCMOS digital camera (Advanced Microscopy Techniques, Woburn, MA.). Image quantification of cellular and PAO1 areas was completed in using NIS-Elements AR 5.21.02.
RNA sequencing and bioinformatic analysis
Total cellular RNA was prepared from uninfected BMDMs and infected BMDMs using Quick-RNA microprep kit (Zymo Research, R1051) and used for the next-generation RNA-sequencing procedure at Texas A&M University Bioinformatics Core. BaseSpace Sequence Hub (Illumina) was used for RNA seq analysis. Briefly, Spliced Transcripts Alignment to a Reference (STAR) algorithm of RNA-seq Alignment V2.0.0 software was used to align the results to reference genome Mus musculus/mm10 (RefSeq), and then RNA-seq Differential Expression V1.0.0 software was used to obtain raw gene expression files and determine statistically significant changes in gene expression in Polg mutant macrophages relative to WT. Ingenuity Pathway Analysis software (QIAGEN) was used to identify gene families and putative upstream regulators in the datasets. Gene Set Enrichment Analysis (GSEA) Software (Broad Institute) was used to generate enrichment plots and examine pathway enrichment. Heatmaps were generated using GraphPad Prism. Datasets of LPS-treated PolgD257A BMDMs used in Figs. 1–2 and Supplementary Fig. 1 were previously reported20 and are available under Gene Expression Omnibus (GEO) accession number GSE171960. Normalized gene counts from PAO1-infected PolgD257A BMDMs are supplied under GEO accession number GSE294537.
RNA sequencing of lungs was performed by isolating total cellular RNA from infected lungs using a Direct-zol RNA miniprep Plus kit (Zymo Research, R2072). Next-generation RNA-sequencing was performed by The Jackson Laboratory Genomic Technologies Core. Partek Flow (v12.3.0) was used for RNA seq analysis. Briefly, Spliced Transcripts Alignment to a Reference (STAR) algorithm of RNA-seq Alignment was used to align the results to reference genome Mus musculus/mm10 (RefSeq), and then RNA-seq Differential Expression software analysis was used to obtain raw gene expression files and determine statistically significant changes in gene expression in Polg mutant macrophages relative to WT. Ingenuity Pathway Analysis software (QIAGEN) was used to identify gene families and putative upstream regulators in the datasets. Heatmaps were generated using GraphPad Prism. Raw and normalized data are available under GEO accession numbers GSE294539 and GSE294540.
Macrophage proteomic and bioinformatic analysis
Cell sample protein extraction and quantification
The adherent cells were washed on the six-well culture plate with PBS (+calcium and magnesium) three times. After washing the cells, 250 µL of 6 M Urea in 50 mM HEPES, pH 8.0 was added to each well. Cells were scraped from the plate, and the solution was transferred to a 2 mL microcentrifuge tube. Samples were then snap frozen in liquid nitrogen and transferred to the Mass Spectrometry and Protein Chemistry Service at The Jackson Laboratory for further processing. Each tube had a pre-chilled 5 mm stainless steel bead (QIAGEN) added and was lysed with the Tissue Lyser II (QIAGEN) for 2 min at 30 1/s. A magnet was used to remove the stainless steel beads, and each sample was further lysed using a 1 mL syringe with a 23-gauge needle for 10 passes. Samples were then ice-waterbath sonicated at 37 Hz (100% power; Firsherbrand FB11207) for 5 min (30 s on, 30 s off for five cycles). The cell lysates were then centrifuged at 21,000 × g at 4 °C for 15 min, and the supernatant was transferred to a new 1.5 mL microcentrifuge tube. A small fraction of the sample was then used to create a 1:50 dilution (sample:50 mM HEPES) and a microBCA assay (Thermo) was run for protein quantification according to the manufacturer protocol.
For each of the quantified cellular protein lysates, 20 µg of protein was diluted in 50 mM HEPES, pH 8.0 to bring the urea concentration less than 2 M for the trypsin digest. Each sample was then reduced with 10 mM dithiothreitol (final concentration) for 30 min at 37 °C for 30 min in a ThermoMixer (500 rpm agitation). Samples were then cooled to room temperature and alkylated with 15 mM iodoacetamide for 20 min at 21 °C in a ThermoMixer (500 rpm agitation). Sequence grade modified trypsin (Promega) was then added at a 1:40 ratio (trypsin:protein) to each sample for digestion at 37 °C for 20 h. Millipore C18 zip-tips (Millipore, ZTC18S096) were then used to desalt and purify the peptide digests according to the manufacturer protocol78,79. All purified peptide samples were then dried using a vacuum centrifuge. Dried peptide digests were reconstituted in 20 µL of 98% H2O/2% ACN with 0.1% formic acid via pipetting up and down 10 times, followed by vortexing at max speed for 30 seconds. Reconstituted samples were centrifuged at low speed for 15 s on a tabletop minicentrifuge to collect the liquid from the walls in the bottom of the microcentrifuge tube. All liquid was then transferred to a mass spec vial (Thermo) and placed in the Vanquish Neo (Thermo) liquid chromatography autosampler set to 4 °C.
Liquid chromatography mass spectrometry analysis
Data independent acquisition mass spectrometry analysis (DIA-MS) was performed on a Thermo Orbitrap Astral mass spectrometer coupled to the Vanquish Neo liquid chromatography system with an EASY-Spray column (PepMap Neo C18, 75 µm x 150 mm, #ES75150PN) in the Mass Spectrometry and Protein Chemistry Service at The Jackson Laboratory. Each sample had two technical replicate runs consisting of an 8 µL injection of reconstituted sample. Before each sample injection the separation column underwent fast equilibration in the method using pressure control at 1500 bar and an equilibration factor of three column volumes. The trapping column underwent fast wash and equilibration using a was factor of 100, an automatic equilibration factor, and flow control at 120 uL/min. The method gradient was 25 min after the column equilibration using Buffer A (100% H2O with 0.1% formic acid) and Buffer B (100% acetonitrile with 0.1% formic acid) with a flow rate of 600 nL/min. The full gradient consisted of 2% B to 4% B from 0–2.5 min, 4% B to 6% B from 2.5–4.5 min, 6% B to 30% B from 4.5–22.5 min, and 30% B to 90% B from 22.5–24 min while ramping to 1 µL/min flow rate in the last step. The column was then washed with 90% B at 1 µL/min from 24–25 min. Orbitrap Astral settings included a static spray voltage using the NSI source at 2000 V in positive mode, an ion transfer tube temperature of 325 °C, an expected peak width of 10 s, advanced peak determination, a default charge state of 2, EASY-IC lock mass correction, and RunStart mode. Full scan settings in positive mode included an Orbitrap resolution of 240,000, scan range of 380–980 m/z, RF lens at 60%, normalized AGC target of 500% with an absolute AGC value of 5e6, and a maximum injection time of 5 ms. DIA method settings included a precursor mass range of 380–980 m/z, an isolation window of 2 m/z with no overlap, normalized HCD collision energy of 25%, detector type set to Astral with a scan range of 150–2000 m/z, RF lens at 60%, normalized AGC target of 500% with an absolute AGC value of 5e4, maximum injection time of 5 ms, and loop control set at 0.6 seconds.
DIA-MS data analysis
All Orbitrap Astral RAW data files underwent automatic processing through the Thermo Ardia Server and were directly sent to the Proteome Discoverer 3.1 workstation connected to the system. The RAW data files for each sample were searched against the UniProtKB Mus musculus (sp_tr_incl_isoforms TaxID=10090; v2024-05-01) protein database in Proteome Discoverer (version 3.1) using the CHIMERYS Ardia Server searching algorithm with the inderys 3.0 fragmentation prediction model. Search settings included trypsin digestion, a maximum missed cleavage of one, peptide length of 7–30 amino acids, peptide charge from 1–4, a fragment mass tolerance of 10 ppm, a dynamic oxidation of methionine modification (+15.995 Da), a static carbamidomethlyl modification of cysteine (+57.021 Da), and a false discovery rate of less than 0.05 for all matches; all other settings were set to manufacturer default recommendations. Total peptide amount was then utilized to normalize the ion signal in the samples. Quantification analyses settings included the use of unique peptides for quantification, and a protein abundance-based ratio calculation was used for the summed abundances, followed by a background based t-test. Strict parsimony principle was then applied for protein grouping, and master proteins were reported. The data was also run with no imputation and low abundance resampling imputation for comparison during the analysis process. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the Proteomics Identifications Database (PRIDE) partner repository with the dataset identifier PXD062877.
Statistical analysis
To ensure rigor and reproducibility, experiments were repeated at least three times with multiple biological replicates. Error bars displayed throughout the manuscript represent the mean +/− s.e.m. Statistical significance between 2 groups was determined via unpaired, two-tailed Student’s t-test or Mann-Whitney test, and differences between 3 or more groups was determined by two-way ANOVA with appropriate post-test. Survival plots were compared by the Log-rank test. Sample sizes were chosen by stated methods to ensure adequate power, and no randomization or blinding was used for animal studies. Significance was reported as a specific p-value or not significant (n.s.).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data needed to evaluate the conclusions drawn herein are present in the paper and Supplementary Materials. The RNA sequencing data generated from PAO1 challenged PolgD257A BMDMs in this study have been deposited in the GEO database under accession code GSE294537. The RNA sequencing data generated from PAO1 challenged PolgD257A lungs in this study have been deposited in the GEO database under accession code GSE294539. The RNA sequencing data generated from PAO1 challenged PolgR292C lungs in this study have been deposited in the GEO database under accession code GSE294540. The RNA-sequencing data deposited to the GEO database includes raw and normalized counts. The proteomic data used in this study are available in the PRIDE database under the dataset identifier PXD062877. The raw data and uncropped western blots generated in this study are provided in the Supplementary Information/Source Data file. Source data are provided with this paper.
References
Gorman, G. S. et al. Mitochondrial diseases. Nat. Rev. Dis. Prim. 2, 16080 (2016).
Suomalainen, A. & Battersby, B. J. Mitochondrial diseases: the contribution of organelle stress responses to pathology. Nat. Rev. Mol. Cell Biol. 19, 77–92 (2018).
Russell, O. M., Gorman, G. S., Lightowlers, R. N. & Turnbull, D. M. Mitochondrial diseases: hope for the future. Cell 181, 168–188 (2020).
Kruk, S. K. et al. Vulnerability of pediatric patients with mitochondrial disease to vaccine-preventable diseases. J. Allergy Clin. Immunol. Pr. 7, 2415–2418.e3 (2019).
Walker, M. A. et al. Predisposition to infection and SIRS in mitochondrial disorders: 8 years’ experience in an academic center. J. Allergy Clin. Immunol. Pr. 2, 465–468, 468.e1 (2014).
Eom, S. et al. Cause of death in children with mitochondrial diseases. Pediatr. Neurol. 66, 82–88 (2017).
Jestin, M. et al. Mitochondrial disease disrupts hepatic allostasis and lowers the threshold for immune-mediated liver toxicity. Mol. Metab. 37, 100981 (2020).
West, A. P., Shadel, G. S. & Ghosh, S. Mitochondria in innate immune responses. Nat. Rev. Immunol. 11, 389–402 (2011).
West, A. P. Mitochondrial dysfunction as a trigger of innate immune responses and inflammation. Toxicology 391, 54–63 (2017).
West, A. P. & Shadel, G. S. Mitochondrial DNA in innate immune responses and inflammatory pathology. Nat. Rev. Immunol. 17, 363–375 (2017).
Warren, E. B. et al. Inflammatory and interferon gene expression signatures in patients with mitochondrial disease. J. Transl. Med. 21, 331 (2023).
Hasselmann, O. et al. Cerebral folate deficiency and CNS inflammatory markers in Alpers disease. Mol. Genet. Metab. 99, 58–61 (2010).
Wilson, L. D., Al-Majid, S., Rakovski, C. S. & Md, C. D. Higher IL-6 and IL6:IGF ratio in patients with Barth syndrome. J. Inflamm. 9, 25 (2012).
Khan, W. et al. Neuroinflammation in the cerebellum and brainstem in Friedreich Ataxia: An [18 F]‐ FEMPA PET study. Mov. Disord. 37, 218–224 (2022).
Nachun, D. et al. Peripheral blood gene expression reveals an inflammatory transcriptomic signature in Friedreich’s ataxia patients. Hum. Mol. Genet. 27, 2965–2977 (2018).
Jin, Z., Wei, W., Yang, M., Du, Y. & Wan, Y. Mitochondrial complex I activity suppresses inflammation and enhances bone resorption by shifting macrophage-osteoclast polarization. Cell Metab. 20, 483–498 (2014).
Yu, A. K. et al. Mitochondrial complex I deficiency leads to inflammation and retinal ganglion cell death in the Ndufs4 mouse. Hum. Mol. Genet. 24, 2848–2860 (2015).
Stokes, J. C. et al. Leukocytes mediate disease pathogenesis in the Ndufs4(KO) mouse model of Leigh syndrome. JCI Insight 7, e156522 (2022).
Hanaford, A. & Johnson, S. C. The immune system as a driver of mitochondrial disease pathogenesis: a review of evidence. Orphanet J. Rare Dis. 17, 335 (2022).
Lei, Y. et al. Elevated type I interferon responses potentiate metabolic dysfunction, inflammation, and accelerated aging in mtDNA mutator mice. Sci. Adv. 7, eabe7548 (2021).
Saha, S., Buttari, B., Panieri, E., Profumo, E. & Saso, L. An overview of Nrf2 signaling pathway and its role in inflammation. Molecules 25, 5474 (2020).
Nezu, M., Suzuki, N. & Yamamoto, M. Targeting the KEAP1-NRF2 system to prevent kidney disease progression. Am. J. Nephrol. 45, 473–483 (2017).
Trifunovic, A. et al. Premature ageing in mice expressing defective mitochondrial DNA polymerase. Nature 429, 417–423 (2004).
Kujoth, G. C. et al. Mitochondrial DNA mutations, oxidative stress, and apoptosis in mammalian aging. Science 309, 481–484 (2005).
Chen, M. L. et al. Erythroid dysplasia, megaloblastic anemia, and impaired lymphopoiesis arising from mitochondrial dysfunction. Blood 114, 4045–4053 (2009).
Thomsen, K. et al. Animal models of chronic and recurrent Pseudomonas aeruginosa lung infection: significance of macrolide treatment. APMIS 130, 458–476 (2022).
Lavoie, E. G., Wangdi, T. & Kazmierczak, B. I. Innate immune responses to Pseudomonas aeruginosa infection. Microbes Infect. 13, 1133–1145 (2011).
Lister, P. D., Wolter, D. J. & Hanson, N. D. Antibacterial-resistant Pseudomonas aeruginosa: clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clin. Microbiol. Rev. 22, 582–610 (2009).
Lambert, M.-L. et al. Clinical outcomes of health-care-associated infections and antimicrobial resistance in patients admitted to European intensive-care units: a cohort study. Lancet Infect. Dis. 11, 30–38 (2011).
Sutterwala, F. S. et al. Immune recognition of Pseudomonas aeruginosa mediated by the IPAF/NLRC4 inflammasome. J. Exp. Med 204, 3235–3245 (2007).
Balakrishnan, A., Karki, R., Berwin, B., Yamamoto, M. & Kanneganti, T.-D. Guanylate binding proteins facilitate caspase-11-dependent pyroptosis in response to type 3 secretion system-negative Pseudomonas aeruginosa. Cell Death Discov. 4, 3 (2018).
Arlehamn, C. S. L. & Evans, T. J. Pseudomonas aeruginosa pilin activates the inflammasome. Cell Microbiol. 13, 388–401 (2011).
Minns, M. S. et al. NLRP3 selectively drives IL-1β secretion by Pseudomonas aeruginosa infected neutrophils and regulates corneal disease severity. Nat. Commun. 14, 5832 (2023).
Kroken, A. R. et al. Intracellular replication of Pseudomonas aeruginosa in epithelial cells requires suppression of the caspase-4 inflammasome. mSphere 8, e0035123 (2023).
Ramadan, A. et al. The NLRP3 inflammasome is required for protection against Pseudomonas keratitis. Investig. Ophthalmol. Vis. Sci. 64, 11 (2023).
Billingham, L. K. et al. Mitochondrial electron transport chain is necessary for NLRP3 inflammasome activation. Nat. Immunol. 23, 692–704 (2022).
Kumari, P., Russo, A. J., Wright, S. S., Muthupalani, S. & Rathinam, V. A. Hierarchical cell-type-specific functions of caspase-11 in LPS shock and antibacterial host defense. Cell Rep. 35, 109012 (2021).
Benjamin, J. T. et al. IL-1α and Not IL-1β drives IL-1R1–dependent neonatal murine sepsis lethality. J. Immunol. 201, 2873–2878 (2018).
Pyrillou, K., Burzynski, L. C. & Clarke, M. C. H. Alternative pathways of IL-1 activation, and its role in health and disease. Front. Immunol. 11, 613170 (2020).
Miao, R. et al. Gasdermin D permeabilization of mitochondrial inner and outer membranes accelerates and enhances pyroptosis. Immunity 56, 2523–2541.e8 (2023).
Rathinam, V. A. K. et al. TRIF licenses caspase-11-dependent NLRP3 inflammasome activation by Gram-negative bacteria. Cell 150, 606–619 (2012).
Kim, B.-H. et al. A family of IFN-γ-inducible 65-kD GTPases protects against bacterial infection. Science 332, 717–721 (2011).
Kim, B.-H. et al. Interferon-induced guanylate-binding proteins in inflammasome activation and host defense. Nat. Immunol. 17, 481–489 (2016).
Shenoy, A. R. et al. GBP5 promotes NLRP3 inflammasome assembly and immunity in mammals. Science 336, 481–485 (2012).
Coers, J. Sweet host revenge: galectins and GBPs join forces at broken membranes. Cell. Microbiol. 19, e12793 (2017).
Coyne, M. J., Russell, K. S., Coyle, C. L. & Goldberg, J. B. The Pseudomonas aeruginosa algC gene encodes phosphoglucomutase, required for the synthesis of a complete lipopolysaccharide core. J. Bacteriol. 176, 3500–3507 (1994).
Deus, C. M., Yambire, K. F., Oliveira, P. J. & Raimundo, N. Mitochondria-lysosome crosstalk: from physiology to neurodegeneration. Trends Mol. Med. 26, 71–88 (2020).
Newman, L. E. et al. Mitochondrial DNA replication stress triggers a pro-inflammatory endosomal pathway of nucleoid disposal. Nat. Cell Biol. 26, 194–206 (2024).
Nguyen, M. et al. Parkinson’s genes orchestrate pyroptosis through selective trafficking of mtDNA to leaky lysosomes. Preprint at https://doi.org/10.1101/2023.09.11.557213 (2023).
Santos, J. C. et al. LPS targets host guanylate‐binding proteins to the bacterial outer membrane for non‐canonical inflammasome activation. EMBO J. 37, e98089 (2018).
Finethy, R. et al. Inflammasome activation by bacterial outer membrane vesicles requires guanylate binding proteins. mBio 8, e01188-17 (2017).
Tsuchiya, K. et al. Gasdermin D mediates the maturation and release of IL-1α downstream of inflammasomes. Cell Rep. 34, 108887 (2021).
Weber, A., Wasiliew, P. & Kracht, M. Interleukin-1 (IL-1) pathway. Sci. Signal 3, cm1 (2010).
Malik, A. & Kanneganti, T.-D. Function and regulation of IL-1α in inflammatory diseases and cancer. Immunol. Rev. 281, 124–137 (2018).
Dinarello, C. A., Simon, A. & van der Meer, J. W. M. Treating inflammation by blocking interleukin-1 in a broad spectrum of diseases. Nat. Rev. Drug Discov. 11, 633–652 (2012).
Lam, C.-W. et al. Novel POLG mutation in a patient with sensory ataxia, neuropathy, ophthalmoparesis and stroke. Clin. Chim. Acta 448, 211–214 (2015).
Amiot, A. et al. Frequency of mitochondrial defects in patients with chronic intestinal pseudo-obstruction. Gastroenterology 137, 101–109 (2009).
Kinoshita, M. & Sawa, T. Serological immune response against major secretory proteins expressed by Pseudomonas aeruginosa upon intratracheal infection in mice. Respir. Investig. 60, 318–321 (2022).
Edmonds, J. L. et al. The otolaryngological manifestations of mitochondrial disease and the risk of neurodegeneration with infection. Arch. Otolaryngol. Head. Neck Surg. 128, 355 (2002).
Lee, W. S. & Sokol, R. J. Mitochondrial hepatopathies: advances in genetics, therapeutic approaches, and outcomes. J. Pediatrics 163, 942–948 (2013).
Mitochondrial, The et al. A modern approach to the treatment of mitochondrial disease. Curr. Treat. Options Neurol. 11, 414–430 (2009).
Rahman, S. & Copeland, W. C. POLG-related disorders and their neurological manifestations. Nat. Rev. Neurol. 15, 40–52 (2019).
Gaudó, P. et al. Infectious stress triggers a POLG-related mitochondrial disease. Neurogenetics 21, 19–27 (2020).
Al-Zubeidi, D. et al. Fatal human herpesvirus 6–associated encephalitis in two boys with underlying POLG mitochondrial disorders. Pediatr. Neurol. 51, 448–452 (2014).
Torres-Odio, S. et al. Loss of mitochondrial protease CLPP activates type I IFN responses through the mitochondrial DNA-cGAS-STING signaling axis. J. Immunol. 206, 1890–1900 (2021).
Buchel, G. et al. Structural basis for DNA proofreading. Nat. Commun. 14, 8501 (2023).
Kang, Y. et al. Ancestral allele of DNA polymerase gamma modifies antiviral tolerance. Nature 628, 844–853 (2024).
Liu, B. C. et al. Constitutive interferon maintains gbp expression required for release of bacterial components upstream of pyroptosis and anti-DNA responses. Cell Rep. 24, 155–168.e5 (2018).
Dickinson, M. S. et al. LPS-aggregating proteins GBP1 and GBP2 are each sufficient to enhance caspase-4 activation both in cellulo and in vitro. Proc. Natl Acad. Sci. USA 120, e2216028120 (2023).
Pilla, D. M. et al. Guanylate binding proteins promote caspase-11–dependent pyroptosis in response to cytoplasmic LPS. Proc. Natl Acad. Sci. USA 111, 6046–6051 (2014).
Meunier, E. et al. Caspase-11 activation requires lysis of pathogen-containing vacuoles by IFN-induced GTPases. Nature 509, 366–370 (2014).
Yazdi, A. S. & Drexler, S. K. Regulation of interleukin 1α secretion by inflammasomes: table 1. Ann. Rheum. Dis. 72, ii96–ii99 (2013).
Keshavan, N., Mhaldien, L., Gilmour, K. & Rahman, S. Interferon stimulated gene expression is a biomarker for primary mitochondrial disease. Ann. Neurol. 96, 1185–1200 (2024).
Pettitt, S. J. et al. Agouti C57BL/6N embryonic stem cells for mouse genetic resources. Nat. Methods 6, 493–495 (2009).
Bezbradica, J. S., Coll, R. C. & Boucher, D. Activation of the non-canonical inflammasome in mouse and human cells. in The Inflammasome (ed. Abdul-Sater, A. A.) vol. 2459 51–63 (Springer US, 2022).
Kamajaya, L. J. & Boucher, D. Gasdermin D cleavage assay following inflammasome activation. in The Inflammasome (ed. Abdul-Sater, A. A.) vol. 2459 39–49 (Springer US, 2022).
Van den Bossche, J., Baardman, J. & de Winther, M. P. J. Metabolic characterization of polarized M1 and M2 bone marrow-derived macrophages using real-time extracellular flux analysis. J. Vis. Exp. 53424 (2015) https://doi.org/10.3791/53424
Zhang, B. et al. Metabolic fitness of IgA+ plasma cells in the gut requires DOCK8. Mucosal Immunol. 17, 431–449 (2024).
Thatcher, K. et al. Temporal progression of aortic valve pathogenesis in a mouse model of osteogenesis imperfecta. J. Cardiovasc Dev. Dis. 10, 355 (2023).
Acknowledgements
We thank Robbie Moore at the Texas A&M College of Medicine Analytical Cytometry Core and Will Schott and Danielle Littlefield in The Jackson Laboratory Flow Cytometry Service for technical assistance. We thank Pete Finger in The Jackson Laboratory Histopathology and Microscopy Sciences Service for assistance preparing macrophage samples for TEM imaging. We thank Philipp Heinrich in The Jackson Laboratory Microscopy Service for assistance with confocal imaging. We would like to acknowledge the Mass Spectrometry and Protein Chemistry Service within Protein Sciences at The Jackson Laboratory for their guidance and support with the mass spectrometry analyses performed in this study. We acknowledge The Jackson Laboratory Center for Precision Genetics (JCPG) for generating the PolgR292C/R292C mouse line. Finally, we thank Vijay Rathinam for protocols and helpful discussions on non-canonical inflammasome biology. This research was supported by awards W81XWH-17-1-0052 to A.P.W. and W81XWH-20-1-0150 to C.C. and A.P.W. from the Office of the Assistant Secretary of Defense and for Health Affairs through the Peer Reviewed Medical Research Programs. Additional support was provided by NIH grants R01HL148153 to A.P.W., U54OD030187 to S.A.M., and a gift from the POLG Foundation. J.J.V. was supported by NIH NRSA F31AI179168, and Y. Lei was supported by American Heart Association Predoctoral Fellowship grant 825908.
Author information
Authors and Affiliations
Contributions
A.P.W. and J.J.V. designed experiments, analyzed data, and wrote the manuscript. Y.L. assisted with animal experiments and sample processing. M.R. assisted with sample processing. C.G.M. performed inflammasome challenge experiments. J.W. assisted with pathology scoring of histology samples. T.J.S. and B.R.H. performed proteomics and analysis. K.P. and R.S. maintained mouse lines and generated BMDMs. S.C.K. and S.A.M. generated the PolgR292C mouse model and provided samples. P.J.M. and C.L.C. assisted with data analysis and interpretation. A.P.W. conceived the project and provided overall direction.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jay H Chung and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
VanPortfliet, J.J., Lei, Y., Ramanathan, M. et al. Caspase-11 drives macrophage hyperinflammation in models of Polg-related mitochondrial disease. Nat Commun 16, 4640 (2025). https://doi.org/10.1038/s41467-025-59907-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59907-8
This article is cited by
-
Model organisms in POLG-related disorders: insights from yeast to multicellular systems
Cell Death & Disease (2025)
-
An inherited mitochondrial DNA mutation remodels inflammatory cytokine responses in macrophages and in vivo in mice
Nature Communications (2025)