Abstract
Multiple sclerosis (MS) involves dysregulation of innate immune cells including monocytes, especially in progressive MS. Fatty acid binding proteins (FABP) are essential for fatty acid transport and metabolism in multiple cell types. FABP7, a brain-FABP, maintains metabolic function in astrocytes and neural stem cells, but the effect of FABP7 on monocytes is unknown. Here we find elevated levels of FABP7 in the serum and cerebrospinal fluid of patients with secondary progressive MS. Elevated serum FABP7 levels positively correlate with higher disability scores, brain lesion volumes, and lower brain volumes. FABP7 levels are increased in astrocytes from MS postmortem brain lesion. Mechanistically, in vitro treatment of FABP7 induces CD16, CD80 and IL-1β expression in monocytes via increased glycolysis. FABP7-induced gene expression reflects enhanced inflammation, chemotaxis and glucose metabolism in monocytes. In conclusion, we find that FABP7 induces pro-inflammatory profiles in monocytes, correlates with disability and represents a potential biomarker and therapeutic target for progressive MS.
Similar content being viewed by others
Introduction
Multiple sclerosis (MS) is an immune-mediated demyelinating and neurodegenerative disorder of the central nervous system (CNS), causing both episodic relapses as well as progressive disability worsening1,2. The primary trigger for this disease has not yet been identified. However, genetic, lifestyle, and environmental factors are recognized risk factors for the disease3,4,5,6,7. In the current model of disease pathogenesis, peripheral immune cells, including T cells, B cells, and circulating monocytes, play a significant role in inflammatory-mediated damage to the CNS8,9,10,11. MS lesions are associated with monocyte infiltration, proliferation, and differentiation into macrophages12,13,14,15 which induce demyelination and neuronal damage16. Moreover, infiltrating monocytes, macrophages, and resident microglia play a critical role in the pathogenesis of progressive MS and predominate in cortical lesions, which are associated with disability10.
The chemotactic and patrolling nature of circulating monocytes on the vascular endothelium facilitates their role as immediate responders to the site of inflammation, including CNS17. Once these monocytes reach an inflammatory site, they differentiate into pro-inflammatory (M1) or homeostatic (M2) macrophages depending on the local stimuli. An increased M1 to M2 ratio is associated with pro-inflammatory microenvironments18. Inhibition of monocyte recruitment can block experimental autoimmune encephalitis (EAE) progression19. Brain lesions from MS patients show inflamed and degenerating brain resident cells, including neurons and astrocytes20,21,22. These cells release inflammatory cytokines, apoptotic debris, and cytosolic contents which promote an inflammatory microenvironment in part by promoting a pro-inflammatory phenotype in monocytes, macrophages, and microglia15,23,24,25,26. Inflammatory monocytes and macrophages have differential metabolic preferences compared to their non-inflammatory counterparts27. A switch from oxidative phosphorylation to the glycolytic pathway induces a switch to an M1, pro-inflammatory, phenotype in monocytes and macrophages28,29. M1 macrophages also demonstrate altered lipid synthesis, evidenced by increased transcription factors such as sterol regulatory element binding protein (SREBP), which induce the production of inflammatory molecules such as interleukin-1beta (IL-1β)30. Differentiated M1-type macrophages release inflammatory cytokines, including IL-1β, resulting in damage to astrocytes, neurons, and microglia31, thereby releasing mediators that further increase this destructive cascade32. However, the specific mediators of this destructive cascade in progressive forms of MS need to be further elucidated.
Fatty acid binding proteins (FABPs), also known as intracellular lipid chaperones, play an essential role in fatty acid transportation and lipid regulation. Several types of FABPs have been associated with MS, including FABP4, FABP5, and FABP733,34. FABP4 and FABP5 are secreted by adipocytes and macrophages and are related to chronic disease progression in diabetes mellitus, obesity, insulin resistance, and other cardiovascular complications35. A recent study shows that adipocyte-produced FABP4 elicits a T cell inflammatory response36. We have previously found serum levels of FABP4 to be associated with increased disability scores in patients with MS34. In addition, FABP4 is increased in secondary progressive MS37 and pediatric MS patients compared to controls38.
FABP7, also known as brain type FABP or brain lipid binding protein, is expressed predominantly in astrocytes of the adult brain39. FABP7 is known to maintain neurogenesis and regulate the function of neural stem cells and astrocytes40. However, increased expression of FABP7 has been observed in several CNS-associated diseases, including EAE, a mouse model of MS39,41,42.
FABP7 serum levels are elevated in Alzheimer’s Disease, Parkinson’s Disease, and dementia patients43. Increased mRNA levels of FABP7 were observed in postmortem brain tissue of autism spectrum disorder and schizophrenia patients44,45. Another study shows an abundance of FABP7 and several other proteins in cerebrospinal fluid (CSF) of patients with Amyotrophic Lateral Sclerosis (ALS), a progressive neurodegenerative disease.46. Unfiltered CSF from these patients induces gross motor deficit in the mouse model of ALS. In the SOD1 knockout (KO) ALS animal model, FABP7 enhances astrocytic inflammation via the NF-κB pathway, leading to neurotoxicity39. Further, FABP7 levels are also increased in the CSF of traumatic brain injury patients and correlate with astroglial injury47.
Astrocyte inflammation and destruction are prominent in active MS lesions22,24, which are also populated by infiltrated monocytes and macrophages, and resident microglia48. A study in the mouse model of EAE has shown that FABP7 is increased in astrocytes at the initiation and peak of disease42. Deficiency of FABP7 significantly reduces the clinical score in the EAE mice42. A recent study demonstrates that a synthetic inhibitor of FABP5 and FABP7 reduced EAE-related symptoms in mice33. Moreover, the presence of FABP7 is increased in the CNS of MS patients and can function as an autoantigen for T cells49.
Thus, there is strong evidence that FABP7 is released by dysfunctional or dying astrocytes in neurologic diseases; however, there is limited knowledge on the effects of FABP7 on MS immunopathogenic mechanisms and, in particular, its effects on monocytes and macrophages, which play a key role in progressive MS. In the present study, we examine the role of FABP4 and FABP7 as biomarkers of progressive MS and evaluate the mechanisms by which they affect monocyte function.
Results
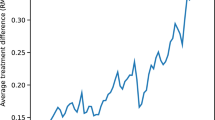
Serum levels of FABP7 were higher in SPMS and showed correlation with EDSS score and brain volumetrics in MS patients
We first analyzed the serum levels of FABP7 and FABP4 in a clinical cohort of relapsing remitting MS (RRMS) and secondary progressive MS (SPMS) patients (demography in Table 1) and compared them with the age-matched healthy controls (HC). We found that the serum level of FABP7 was significantly increased in SPMS patients compared to the age-matched HC (p = 1.42e-05) and RRMS (p = 6.46e-08) patients (Fig. 1a). FABP4 was insignificantly (p = 0.0925) increased in SPMS patients compared to the age-matched HC (Fig. 1b), however, significantly to RRMS (p = 0.0007).
a, b Bars (mean ± SD) show serum levels of (a) FABP7 and (b) FABP4 in RRMS, and age-matched healthy controls (HC1) and in SPMS and age-matched healthy controls (HC2). One-way ANOVA was performed following correction with Šidák multiple comparison test, and P-values are shown above the comparisons (n = 32 of each group). c A negative correlation of FABP7 with BPF (n = 34). d, e A positive correlation of FABP7 with T1LV and T2LV (n = 34). f A positive correlation of serum FABP7 with EDSS score (n = 98). Two-sided simple linear regression analysis was performed for correlation and shown as a regression line (shaded area represents error of the regression line), data is adjusted for age and sex, and p-values were adjusted using Holm’s correction for (c–f). RRMS Relapsing remitting MS, SPMS Secondary progressive MS, BPF Brain Parenchymal Fraction, T1LV whole brain T1 hypointense white matter lesion volume, T2LV whole brain T2 hyperintense white matter lesion volume.
Next, we asked whether FABP7 and FABP4 are associated with MRI measures of disease severity, including reduction in various brain regions such as the global pallidus and deep gray matter (DGM), changes in brain parenchymal fraction (BPF), and increases in lesion volume within our MRI cohort (demography in Table 2). We evaluated the correlation of serum levels of FABPs with the brain MRI volumetrics (Table 3). These results showed a significant correlation of the serum levels of FABP4 and FABP7 with the volume of different regions of the brain. FABP4 levels were significantly negatively correlated with BPF, globus pallidus, thalamus and total DGM volumes and positively correlated with T1 hypointense (T1LV) and T2 hyperintense (T2LV) lesion volumes. This is in line with our prior findings showing that FABP4 is associated with disability in MS patients34. FABP7 had a strong negative correlation with BPF and a positive correlation with T1LV and T2LV (Table 3 and Fig. 1c–e). Next, we assessed the correlation of serum FABP7 levels with expanded disability status scale (EDSS) in our combined clinical and MRI cohorts. We found that serum FABP7 levels positively correlated with the EDSS score (β = 0.14, p = 0.000005) in MS patients (Fig. 1f).
FABP7 increased CD16, CD80 and IL-1β in monocytes
The negative correlation of FABP4 and FABP7 with brain volume and the positive correlation with disability scores and lesion volumes suggests that they may have an important role as serum biomarkers in MS patients. Moreover, to explore whether these proteins play a role in modulating cellular physiology and inducing inflammation in immune cells, we studied monocytes isolated from frozen PBMCs of HC and SPMS (male and female) patients. First, in order to validate the use of monocytes from frozen PBMCs as an experimental in vitro model, we isolated monocytes from freshly drawn blood (termed as fresh monocytes) and from frozen PBMCs (termed as thawed monocytes) from same HCs (age 32.5 ± 3.2 years, 50% male and 50% female) using pan monocyte isolation kit (Supplementary Table 1). We analyzed monocyte subpopulations and expression of HLA-DR as a functional marker for monocytes (Supplementary Fig 1a–d). We did not find a significant difference in classical (CD14+ CD16-), intermediate (CD14+ CD16+), and non-classical (CD14- CD16+) monocyte populations in fresh vs thawed monocytes. Further, the HLA-DR expression was also similar in fresh and thawed monocytes, and there was no difference in viability of these cells.
Expression of IL-1β is a well-known marker of inflammation in monocytes and macrophages50. To investigate the effect of these FABPs in inflammation in monocytes, we isolated monocytes from SPMS patients (age 62.9 ± 2.7 years, 55% female and 45% male) and treated them with various doses of FABP7 (20, 50, 100, and 300 ng/ml) for 24 h and 48 h and performed flow cytometry using various antibodies (Supplementary Table 2). The subsets of monocytes were evaluated using the gating strategy shown in supplementary Fig 2. We found that compared to the control group, FABP7 decreased the percentage of CD14+ CD16- monocytes and increased CD14+ CD16+ monocytes at both 24 h and 48 h (Supplementary Fig 3a). FABP7 at all doses increased CD80 expression at 24 h and in doses ≥ 50 ng/ml at 48 h compared to the respective controls. CD86 was increased at 24 h, however, no significant difference was observed at 48 h. Further, CD163 was decreased at higher doses of FABP7 at 48 h. HLA-DR was also found to be decreased at 48 h at all the doses of FABP7, but an opposite effect was observed at 24 h. We did not find changes in IL-10 expression, however, TNFα was increased at 24 h and no difference was observed at 48 h. Importantly, the IL-1β expression was significantly upregulated at 50 ng/ml and above doses of FABP7 at both time points (Supplementary Fig 3a,b). Overall, the effect of FABP7 at higher doses and 48 h time point were found suitable and consistent across the experiments.
To further investigate the effect of FABP7 and FABP4, we stimulated monocytes from HCs (age 30 ± 4.5 years, 50% male and 50% female) with 100 or 300 ng/ml doses of FABP7 and 25 or 100 ng/ml doses of FABP4 for 48 h. We chose these in vitro doses of FABP7 based on our serum studies demonstrating levels in the range of 20 ng/ml or higher in SPMS patients, and with the assumption that local CNS levels of FABP7 would be much higher, due to the gradient from the CNS to the serum. The relative levels of glial fibrillary acidic protein (GFAP) and neurofilament light chain in CSF and serum of MS patients is an example where CSF level of these proteins are many times elevated compared to their serum levels51. Further, FABP4 is mostly expressed by adipocytes and is not specific to the CNS therefore, we used in vitro doses of FABP4 comparable to their serum levels. Figure 2a and supplementary Fig 4a showed that FABP7 at the doses of 100 and 300 ng/ml significantly decreased the percentage of the CD14+ CD16- population (p = 0.0007, and p = 0.0044, respectively) while increased the CD14+ CD16+ population (p = 0.0007, and p = 0.0045, respectively), compared to the control group. FABP7 at both doses also significantly increased the expression of CD80 (p = 0.0001, and p = 0.0009, respectively), and IL-1β (Fig. 2a, b) (p = 0.0004, and p = 0.0001, respectively), compared to the control group. Further, we observed that FABP7 at 100 ng/ml decreased IL-10 expression (p = 0.01) however, we did not observe significant changes in other markers associated with inflammatory monocytes, such as CD86 and TNFα and anti-inflammatory monocytes, such as CD163 and CD206, at the protein level. We also did not observe significant effects of FABP4 on monocyte subset population or cytokine production (Fig. 2a, b).
Monocytes were treated in vitro with FABP7 or FABP4 for 48 h. Flow cytometry was performed, and data is represented as (a) bars (mean ± SD, n = 12) indicating percentage of cell population gated from CD14+ cells from HCs and (b) zebra plot showing expression of IL-1β based on FMO. One-way ANOVA was performed following correction with Dunnett’s multiple comparison test for (a). c Bars (mean ± SD, n = 9) showing cell populations gated from CD14+ cells in HCs and SPMS. Two-way ANOVA was performed following correction with Šidák multiple comparison test. d zebra plot showing expression of IL-1β. P-values are shown above the comparisons. FMO Fluorescent minus one.
Next, we compared the effect of FABP7 on monocytes of SPMS patients (age 62.9 ± 2.7 years, 55% female and 45% male) and age-matched HCs (age 56.7 ± 1.5 years, 55% female and 45% male) and analyzed whether FABP7 has a differential effect on SPMS monocytes (Fig. 2c, d and supplementary Fig 4b). Since SPMS patients are older, therefore, we used an age-matched cohort of HC, which are different from the cohort of HC used in Fig. 2a. FABP7 at both doses significantly upregulated CD16, CD80, CD86, and IL-1β expression while downregulated HLA-DR expression in monocytes of SPMS and age-matched HCs. However, we did not find differences when comparing SPMS to their respective groups of HC monocytes. We also did not find a significant difference in CD14+ CD16-, CD14+ CD16+ or CD14- CD16+ population and HLA-DR expression in monocytes immediately isolated from PBMCs of these SPMS and HC (Supplementary Fig 5a–e).
FABP7 increased basal ECAR and inhibition of glycolysis by 2DG reduced the effect of FABP7 in monocytes
Increased glycolysis is one of the most prominent features of inflammatory monocytes and macrophages27. For example, LPS has been shown to increase glycolysis in monocytes as well as macrophages, thereby promoting strong IL-1β production and other inflammatory responses27. To investigate if FABP4 and FABP7 have any effect on glycolysis, we performed a mitochondrial stress test in monocytes of HC (age 29 ± 3.5 years, 50% female and 50% male) using the standard Seahorse XF Cell Mito Stress kit. Figures 3b, e indicated that FABP7 at the dose of 100 or 300 ng/ml strongly elevated the basal extracellular acidification rate (ECAR) (p = 0.0048, and p = 9.42e-07, respectively) in monocytes after 48 h of incubation, whereas FABP4 at 100 ng/ml marginally increased basal ECAR (Fig. 3g). Further, there was a non-significant increase in basal respiration (Fig. 3a, c), and a significant increase in ATP-linked respiration (Fig. 3d) (p = 0.0313) at 300 ng/ml of FABP7 was observed. Furthermore, FABP4 treatment did not show changes in the oxygen consumption rate (OCR) (Fig. 3f). Moreover, we did not observe any significant effect of FABP4 on monocyte metabolism or inflammatory phenotype as previously mentioned. Consequently, we decided not to include FABP4 in our subsequent experiments.
Monocytes were treated with FABP7 and FABP4 for 48 h, and a seahorse assay was performed. a–e Graphs (mean ± SD) indicate (a) OCR and (b) ECAR in monocytes treated with FABP7, bar diagram represents (c) basal respiration, (d) ATP-linked respiration/ATP production and (e) basal ECAR (n = 8 for control and FABP7 and n = 5 for LPS, biological replicates). f, g Graphs (mean ± SD) indicate OCR and ECAR in monocytes treated with FABP4 (n = 8 for control and FABP4 and n = 5 for LPS, biological replicates). h Monocytes were treated with FABP7 in the presence or absence of 2DG for 48 h, and flow cytometry was performed, data is represented as bars (mean ± SD, n = 24) indicating the percentage of cell population gated from CD14+ cells, and (i) zebra plot showing expression of IL-1β. j Effect of FABP7 on ROS production in monocytes of HC. ROS assay was performed in Monocytes treated with FABP7 for 6 h using flow cytometry, and bars represent MFI of ROS positive cells (mean ± SD, n = 5), a representative histogram is also shown for ROS. k Mitochondrial superoxide was determined using flow cytometry after treating the HC monocytes with FABP7 for 48 h, the bar (mean ± SD) represents MFI of MitoSox positive cells (n = 6), and a representative histogram is also shown for MitoSox. One-way ANOVA was performed following correction with Dunnett’s multiple comparison test for (c–e, j, and k), and correction with Šidák multiple comparison test for (h). P-values are shown above the comparisons. OCR Oxygen consumption rate, ECAR Extracellular acidification rate, ROS reactive oxygen species.
Following our observation that FABP7 increased basal glycolysis in monocytes, we next asked whether inhibition of glycolysis can affect FABP7-induced inflammation and the induction of an M1 phenotype (CD16, CD80, CD86, IL-1β, TNFα) or M2 phenotype (CD206, CD163, IL-10) in monocytes. Therefore, we used 2-deoxy-D-glucose (2DG), a glucose analog that inhibits glycolysis (by inhibiting hexokinase), in the presence or absence of FABP7 (300 ng/ml) using monocytes of HC (age 43.1 ± 14.14 years, 50% female and 50% male). As shown in Fig. 3h, treatment with 2DG in the presence of FABP7 increased the CD14+ CD16- population (p = 0.0111) compared to the FABP7 alone group. Further, 2DG treatment in combination with FABP7 decreased the CD14+ CD16+ population (p = 0.0111), i.e., to near control value when compared to FABP7 alone. Furthermore, the expression of CD80 and CD86 were reduced (p = 0.0136, and p = 1.29e-05, respectively), in the FABP7 group treated with 2DG compared to the FABP7 alone group. Interestingly, the expression of IL-1β (Fig. 3h, i) was decreased (p = 0.0089) in the FABP7 group treated with 2DG compared to the FABP7 group alone. However, we did not find any effect of 2DG in the presence of FABP7 on the expression of CD163, IL-10, or TNFα when compared to FABP7 alone (Supplementary Fig 6). However, CD206 was found to be significantly decreased (p = 3.17e-09) in the group treated with 2DG in the presence of FABP7 when compared to FABP7 alone.
FABP7 has no effect on ROS production in monocytes
Reactive oxygen species (ROS) generation is a well-known phenomenon during inflammation in monocytes and is known to induce IL-1β expression30. Studies have indicated that increased glycolysis in monocytes and macrophages induces ROS52. Therefore, we also analyzed if FABP7 can induce ROS production in monocytes. Contrary to our hypothesis, we did not observe increased ROS production after treating the cells with FABP7 (Fig. 3j) in monocytes of HC (age 32.1 ± 3.5 years, 60% female and 40% male). We further investigated whether FABP7 had any effect on mitochondrial superoxide production using monocytes of HC (age 32.6 ± 3.1 years, 50% female and 50% male). However, we did not find any change in mitochondrial superoxide production after treating the monocytes with FABP7 (Fig. 3k).
FABP7 increased expression of inflammatory genes and gene enrichment associated with inflammatory pathways and glucose metabolism
We have found that FABP7 increases inflammatory phenotypes in monocytes along with an increase in the ECAR, and inhibition of glycolysis by 2DG decreased this effect. This observation indicates that glycolysis is an essential pathway for FABP7-induced inflammation in monocytes. To further investigate this observation and considering that the CNS lesion microenvironment may have higher concentration of FABP7, we treated monocytes of HC (age 30.6 ± 4 years, 50% female and 50% male) with FABP7 (300 ng/ml) for 24 h and performed bulk RNA sequencing. However, lower doses such as 100 ng/ml may also have a similar effect on the expression of inflammatory markers in monocytes. Figure 4a showed transcriptomic changes in control vs FABP7 treated monocytes. The heatmap (Fig. 4b) showed the top 50 differentially expressed genes, where upregulated genes are found to be associated with inflammation and chemotaxis, including IL1A, IL6, CXCL8, CCL3L1, CCL7, CCL4L1, IL1B, IL2RA, CCL4, CCL3, CXCL1, CCL19 etc. Further, Gene Ontology pathway analysis (ClusterProfiler) for the biological process of all DEGs indicated that genes were significantly enriched for chemotaxis, cell migration, and inflammatory pathways, including IL1 (Fig. 4c). These results show that FABP7 skews overall gene expression in monocytes towards an inflammatory phenotype. Furthermore, to analyze gene enrichment in cellular metabolic pathways, we selected all significant DEGs and analyzed them using Enrichr, an online tool for gene list enrichment analysis53,54,55. We selected the Human Cyc 2016, which is known for metabolic pathway analysis. As shown in Fig. 4d (and Supplementary Table 3), these significant DEGs were enriched for glucose metabolism, represented as a superpathway of conversion of glucose to acetyl CoA and entry into the TCA cycle. This observation aligns with our findings, where we have shown that FABP7 increased metabolic preference towards glycolysis, indicated as increased ECAR. However, the role of highly DEGs needs to be further investigated.
a Volcano plot showing the distribution of overall transcriptomics in monocytes after FABP7 treatment. DESeq2 with a two-sided Wald test was performed for comparison. Data was adjusted for sex. b Heat map showing top 50 DEGs. c Dot plot of gene set enrichment analysis of GO biological pathways using ClusterProfiler. The pathways related to inflammation and chemotaxis were highlighted in the red box. d Pathway analysis of all significant DEGs using Human Cyc 2016 in Enrichr indicates gene enrichment for glucose metabolism. The tests conducted were two-sided, and p-values were adjusted using Benjamini-Hochberg to correct for multiple hypothesis testing for (c, d). DEGs Differentially expressed genes.
FABP7 induced inflammatory profiles in monocytes of secondary progressive multiple sclerosis patients
Based on the available literature and our current observations where we found increased levels of FABP7 in SPMS patients, we hypothesized that the increased presence of FABP7 in the CNS milieu of SPMS patients may induce inflammation in infiltrating monocytes. To investigate this, we isolated peripheral monocytes from untreated SPMS patients (60.9 ± 5.7-year-old, 55% female and 45% male), stimulated with FABP7 (300 ng/ml) for 48 h, and performed Seahorse assay and flow cytometry. Figure 5a, c indicated that FABP7 marginally and insignificantly increased basal respiration or ATP-linked respiration (Fig. 5d). Interestingly, FABP7 significantly increased basal ECAR (Fig. 5b, e) (p = 0.0041) in monocytes of SPMS patients. Further, Fig. 5f showed that FABP7 induced changes in the CD14+ CD16-, CD14+ CD16+, CD80, CD86, and IL-1β were similar as observed previously (Fig. 2c, d). Furthermore, treatment with 2DG along with FABP7 increased the CD14+ CD16- monocyte population (p = 0.0327) while it decreased the CD14+ CD16+ monocyte population (p = 0.0328) along with CD80 (p = 0.0325), and CD86 (p = 0.0312) expression, compared to the FABP7 alone group. However, we did not observe a significant difference in IL-1β expression in the 2DG group treated with FABP7 compared to FABP7 alone in the monocytes of SPMS patients.
a–e SPMS monocytes were treated with FABP7 for 48 h, and a seahorse assay was performed. Graphs (mean ± SD) indicate (a) OCR and (b) ECAR, bar diagram represents (c) basal respiration, (d) ATP-linked respiration and (e) basal ECAR (mean ± SD, n = 11, biological replicates). f SPMS monocytes were treated with FABP7 in the presence or absence of 2DG for 48 h and flow cytometry was performed, data is represented as bars (mean ± SD, n = 11) indicating the percentage of cell population gated from CD14+ cells. (g) Structure and predicted interaction of FABP7 with TLR4-MD2 complex. h, i Isolated monocytes from HC (n = 12) were incubated with anit-TLR4 antibody following FABP7 treatment for 48 h, and IL-1β was analyzed by flow cytometry and represented as bars (mean ± SD) and zebra plot. A paired two-sided t test was performed for (c–e), and one-way ANOVA was performed following correction with Šidák multiple comparison test for (f and h). P-values are shown above the comparisons. OCR Oxygen consumption rate, ECAR extracellular acidification rate.
In addition to glycolysis, we investigated whether FABP7 has any effect on fatty acid oxidation in monocytes of SPMS patients (age 59 ± 1.7 years, 50% female and 50% male). As shown in Supplementary Fig 7a,b, FABP7 at 300 ng/ml marginally increased fatty acid oxidation in monocytes (p = 0.0546).
FABP7-induced IL-1β is partially regulated by glucose transporter and TLR4
We have observed that FABP7 increased glycolysis and induced a pro-inflammatory phenotype in monocytes. From bulk RNA sequencing (Fig. 4a), we found that FABP7 increased mRNA expression of glycolytic enzymes, including HK2, PKM, and PFKFB3 along with glucose transporters such as SLC2A3 and SLC2A6 which regulate glycolysis. To confirm if FABP7 regulates inflammation in monocytes through modulating the expression of these proteins, we silenced these genes using siRNA in monocytes from HCs. We found that silencing SLC2A6 and HK2 near significantly decreased FABP7-induced IL-1β (p = 0.0519 and p = 0.0720, respectively) expression in monocytes of HC (Supplementary Fig 7c,d).
Inflammasome activation is also known to induce IL-1β production. Therefore, to investigate if FABP7 also acted through this pathway, we performed inflammasome activation assay in monocytes of SMPS in the presence of FABP7. First, we validated the assay in our experimental system using LPS stimulation for 4 h in the presence or absence of inflammasome inhibitor (YVAD-CHO) (Supplementary Fig 7e). Next, we stimulated SPMS monocytes with FABP7 for 4 h and observed no changes in inflammasome activation (Supplementary Fig 7f). We also did not observe any difference in inflammasome activation between age-matched HC and SPMS monocytes after 4 h of culturing without any stimulation (Supplementary Fig 7g).
Further, to understand how FABP7 could interact with monocytes and mediate its effect, we utilized a protein receptor interaction approach. Toll-like receptors (TLRs) mediated signaling is a well-known mechanism that induces transcriptional changes associated with inflammation. TLR4 is a prominent receptor that binds a wide complement of ligands, including pathogen associated molecules, non-LPS molecules, and cellular proteins56. TLR4 pathway can induce the release of IL-1β independent of NLRP3/Casp-1 activation57. To understand if FABP7 interacts with TLR4 and induces inflammation in monocytes, we performed a protein interaction study between TLR4 and FABP7 (Fig. 5g). All of the top 10 best scoring models generated in HDOCK had docking (< 200) and confidence scores (> 0.7) suggesting a high likelihood of binding between FABP7 and the TLR4-MD2 complex. The predicted interface residues, those within a 5 Å distance between the two proteins of the highest scoring model, are summarized in Supplementary Table 4. To further confirm this, we blocked TLR4 using an anti-TLR4 antibody and stimulated the monocytes of HC (age 33.8 ± 2.9 years, 50% female and 50% male) with FABP7 in vitro. Interestingly, blocking TLR4 following FABP7 stimulation (100 ng/ml and 300 ng/ml) significantly decreased IL-1β expression (p = 0.0278 and p= 0.0002, respectively) in monocytes compared to the FABP7 alone groups (Fig. 5h, i).
FABP7 is increased in the CSF of MS patients, highly expressed by astrocytes, and associated with inflammatory immune cell phenotypes in the MS brain lesion
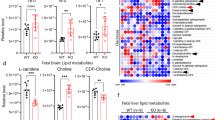
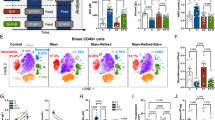
To better understand how FABP7 might contribute to brain volume loss in MS patients, we first investigated the FABP7 level in CSF of SPMS patients (n = 14; age = 58.3 ± 7.6 years, 64% female and 36% male) on Foralumab treatment, and compared it with non-MS controls (n = 10; age 29.5 ± 7.5 years, 100% female and 0% male), using ELISA. We found an increased level (p = 0.0005) of FABP7 in CSF of MS patients (Fig. 6a). We next used postmortem brain tissue of an MS patient (50-year-old female) and healthy control (85-year-old female) and analyzed the FABP7 expression. Interestingly, we observed that FABP7 expression was increased (p = 0.0002) in the brain lesion compared to the healthy control (Fig. 6b, c and Supplementary Fig 8). The FABP7 was mostly found localized in the astrocytes (Fig. 6b). Next, we examined FABP7 expression in postmortem brain tissue using single-nucleus RNA sequencing. Datasets from 20 white matter tissue compartments derived from 5 patients with progressive MS and 3 patients without neurologic disease58 were uniformly processed into a harmonized atlas of MS brain cell types. The tissue compartments include MS lesions (subdivided further into lesion core, lesion rim, and periplaque) and control white matter. Integrated analysis revealed six broad classifications of cell types in these tissues, consistent with previously published results: oligodendrocytes, oligodendrocyte precursor cells (OPCs), neurons, astrocytes, endothelial/vascular cells, and immune cells (Supplementary Fig 9). FABP7 expression was most reliably detected in astrocytes and OPCs (Supplementary Fig 10). However, we found that FABP7 was increased and mostly localized in astrocytes of MS lesions as shown in Fig. 6b. When plotting astrocyte FABP7 expression by tissue compartment, a striking pattern emerged. FABP7 expression was significantly elevated in MS lesion-containing tissue, with significant enrichment at the rims of chronic lesions compared to periplaque and healthy control tissue (Fig. 6d, e). These rims, identifiable in vivo with iron-sensitive MR imaging, have previously been characterized as hypercellular/microglia-rich regions of smoldering inflammation and axonal degeneration; they are suspected to perpetuate disease activity and/or neurodegenerative processes in progressive MS59,60.
a FABP7 levels were analyzed in CSF of SPMS and non-MS controls using ELISA (n = 10 for non-MS control, and n = 14 for SPMS). b Brain tissue showing lesion in MS and immunofluorescence images captured at 200X magnification (a small area was further enlarged) showing FABP7 expression (green), GFAP positive astrocytes (bright purple) and nucleus (DAPI: blue) in HC and MS participants. c FABP7 mean fluorescence intensities from five different random areas were quantified and represented as bar (mean ± SD, n = 5), unpaired t-test was performed for (a and c). d–j Brain tissue section showing Single cell transcriptome datasets of 20 postmortem brain tissue compartments from 5 MS patients and 3 healthy controls were examined for FABP7 expression and corresponding immune cell gene expression signatures. d UMAP dimensionality reduction plot of 10,767 astrocytes. e FABP7 gene expression in astrocytes from healthy and disease white matter tissue compartments. Indicated p-values of differential gene expression are from the two-sided Wilcoxon Rank Sum test, and p-values were adjusted for multiple testing using Bonferroni correction. f UMAP plot of 5148 immune cells from the same tissue compartments as (d and e). g Immune cell marker analysis and cell subset classifications. h Percent change in the relative proportion of immune cell subsets in MS tissue compartments versus healthy control tissue. The percentage of CD14/CD16 double-positive cells is indicated by the color strip on the left. i Statistical associations between astrocyte FABP7 expression and proportions of immune cell subsets from 18 brain tissue microenvironments. Depicted values are Spearman’s rho statistic. j Scatter plots and statistical associations (mean ± SEM) of homeostatic (left) and inflamed (right) microglia with astrocyte FABP7 expression by tissue microenvironments in paired samples was performed using two-sided Spearman’s rho test with no adjustment for multiple correction.
We hypothesized that immune cells within lesion rims might exhibit phenotypic similarities with in vitro FABP7-treated monocytes, given the potentially elevated local FABP7 concentration in these regions. To investigate further, we used optimal-resolution clustering to define immune cell subsets from the broader immune cell cluster (Fig. 6f and Supplementary Fig. 11). Several categories of homeostatic and activated microglia, as well as infiltrating peripheral immune cells were detected in the MS tissue (Fig. 6g). MS lesion rims exhibited a marked increase in the relative proportions of T cells, B cells, infiltrating macrophages, and activated microglial subtypes compared to control tissues (Fig. 6h). A relative depletion of homeostatic microglia was also observed. Furthermore, cells co-expressing CD14 and FCGR3A, the genes encoding surface markers CD14 and CD16, respectively, were enriched in the rims of MS lesions (Supplementary Fig 12). Interestingly, the immune cell subsets most strongly correlated with our in vitro bulk RNA-seq of FABP7-treated monocytes were clusters 2 and 10, which have previously been defined as “microglia inflamed in MS” (MIMS) and MIMS-iron (Supplementary Fig 13). This observation suggests that immune cell populations specific to chronic MS lesions, consisting of CD14-expressing microglia and/or infiltrated monocyte populations, may express CD16 in a high FABP7 environment, which warrants further investigation. This transcriptional phenotype is consistent with our in vitro results of FABP7-treated peripheral monocytes, however, the precise origin of these brain immune cells, whether peripheral or central, remains unknown.
For a more granular view of the association between astrocyte FABP7 expression and immune cell phenotypes in the brain, we correlated the pseudobulk expression of astrocyte FABP7 with the proportion of immune cell subsets for each individual library (Fig. 6i). We reasoned that each individual library represents a tissue “microenvironment” – the cells within the library were in close physical proximity at the time of autopsy – and thus spatial relationships can be inferred. This analysis revealed a strong association between astrocyte FABP7 expression and inflammatory immune cell phenotypes. Specifically, Spearman’s rank-based correlation test yielded a rho statistic of 0.67 and p-value of 0.0021 for the association between astrocyte FABP7 and the proportion of inflammatory microglia in the microenvironment (Fig. 6j). There was a modest negative correlation between FABP7 and homeostatic microglial populations. When considered alongside our in vitro results, the correlative postmortem transcriptomics data support a model that astrocyte FABP7 expression perpetuates inflammatory immune activity in chronic MS lesions, potentially contributing to brain atrophy over time in progressive MS patients.
Discussion
The present study, for the first time, highlighted the role of FABP7 in promoting inflammation in monocytes in MS. First, we have shown that increased levels of serum FABP7 serve as a biomarker for SPMS, increasing disability, and worsening of brain volumetrics. Second, we found that FABP7 induces a pro-inflammatory phenotype in monocytes in healthy as well as SPMS patients. Importantly, FABP7 increased IL-1β and other M1 macrophage-associated markers CD16 and CD80. We also observed that FABP7 increased glycolysis, which induced a pro-inflammatory phenotype in monocytes associated with increased IL-1β. RNA sequencing analysis of FABP7-treated monocytes showed that enriched gene expression is associated with inflammation, chemotaxis, and cell migration and metabolic pathways associated with glucose metabolism. Further, we found that FABP7-induced IL-1β is partially mediated by the TLR4 pathway. In addition, FABP7 was increased at the protein level in CSF and astrocytes in the brain lesion of MS patients. Single nuclei sequencing analysis of postmortem brain tissue of MS patients showed that FABP7 expressing microenvironment have elevated CD14+ CD16+ immune cells in chronic active lesions, suggesting FABP7 may promote an inflammatory phenotype in infiltrated monocytes contributing to inflammation in the brain of MS patients.
There has been much recent work focused on identifying potential biomarkers for MS61. Depending on their source and ability to cross the BBB, these biomarkers may be present in blood or CSF. The presence of Ig oligoclonal bands in CSF, Ig index in CSF/serum, GFAP, chitinase-3-like-1, and neurofilaments in CSF and blood are some well-known indicators of inflammation and neurodegeneration62,63. In this study, we explored the presence of FABP4 and FABP7 in the serum of MS patients where FABP4 and FABP7 were significantly correlated with progressive MS, increased disability and brain atrophy in MS patients and thus FABPs may be incorporated into biomarker panels and algorithms correlating with the clinical stage of MS and potentially subsequent disease course. Interestingly, the in vitro addition of Docosahexaenoic acid (DHA) stimulates FABP7-mediated induction of GFAP, which is a commonly used astrocytic marker of disease progression in MS64.
FABP7 levels are increased in several neurodegenerative diseases, which may reflect shared pathobiology15. FABP7 is mainly expressed in astrocytes in the brain65, and its increased production induces an astrocytic inflammatory response which causes toxicity in the neurons39. A recent study has shown the presence of FABP7 in the CNS of MS patients, functions as an autoantigen for T cells49. In the present study, our focus was on how the released FABP7 in the CNS milieu affects the infiltrated immune cells. Although studies have shown that reactive astrocytes increase the presence of inflammatory macrophages66,67, it was unclear how FABP7 affected the macrophages and monocytes. It is possible that FABP7 is secreted in greater amounts by reactive or dying astrocytes, which could cause inflammation in infiltrating monocytes. Our studies add to the emerging roles of FABP7 by showing that FABP7 in vitro increased the M1 inflammatory markers, IL-1β, CD16, CD80, and CD86 in monocytes from controls and SPMS patients and induces a glycolytic switch in these cells, relevant to the mechanisms of progressive MS. Here, we did not observe a prominent dose dependent effect of FABP7 in monocytes on the expression of CD16, CD80 or CD86, however, IL-1β was increased in a dose dependent manner. Age of the patients could be an important factor in FABP7-induced inflammation in monocytes68. We also observed a slightly different percentage of monocyte markers in different cohorts of HCs as shown in Fig. 2a,c. Further, we aimed to analyze the sole effect of FABPs in in vitro culture of monocytes without stimulation of GM-CSF or any interleukins which results in decreased viability after 24 or 48 h and is considered a limitation in this study and can be studied further.
In this study, we used monocytes isolated from thawed PBMCs and did not find a change in the phenotypes of thawed and fresh monocytes immediately post-monocyte isolation. However, they may acquire different phenotypes when stimulated with FABP7 or other stimuli, including LPS and can be studied further. In addition, we did not observe the difference in CD16 or HLA-DR expression in HC and SPMS monocytes, we believe that there would be a difference when analyzing various other markers using transcriptomic studies as shown in other studies69,70. Therefore, the differences in FABP7 serum levels between HC and SPMS may not accurately represent the phenotypes in their respective monocytes, as our study only analyzed a limited number of phenotypic markers. A detailed study may address this, where serum-matched monocytes can be used in HC and SPMS participants.
Differentiation of monocytes to M1 macrophages preferentially upregulates glycolytic pathways to compensate for the high energy demand and requirements of various intermediate metabolites, known as the Warburg effect71. Also, increased glycolysis in macrophages has been shown to promote inflammation52,72. We found that FABP7 affected monocyte metabolic pathways by increasing glycolysis dependency (Figs. 3a–i and 5a–f). Moreover, analysis from bulk RNA sequencing demonstrated that FABP7 increased gene enrichment for pathways involving glucose metabolism in monocytes (Fig. 4d). Importantly, we found that inhibition of glycolysis using 2DG significantly reversed the inflammatory effect of FABP7 in monocytes, suggesting a potential mechanism of M1 phenotype conversion, mediated by FABP7 induction of a glycolytic switch. However, inhibiting glycolysis did not affect the IL-1β expression in monocytes of SPMS patients. This suggests that there are other mechanisms involved, such as fatty acid induced IL-1β expression73 in these monocytes, which should be further explored. We also observed increased fatty acid oxidation in SPMS monocytes in the presence of FABP7, however, further experiments are needed to confirm these effects.
FABP7 also interacts with proteins and regulates cellular function. For example, FABP7, as an intracellular fatty acid chaperone, actively interacts with ATP citrate lyase, triggering an acetyl-CoA metabolism in astrocytes and glioma74. However, in monocytes, FABP7-mediated alterations need further investigation. Our in vitro transcriptomic data (Supplementary Table 5) does indicate that FABP7 significantly increases glucose transporter genes such as SLC2A3, SLC2A6, and SLC2A14, key glycolytic enzyme genes such as ADPGK, PFKFB3, HK2 and PKM and lipid synthesis genes such as SREBF2, INSIG1, and INSIG2. Although, we found that FABP7-mediated IL-1β is partially regulated by increased expression of SLC2A6 and HK2. However, their role in the regulation of glycolysis and other mechanisms for IL-1β production in the presence of FABP7 further needs to be validated. Interestingly, inflammatory macrophages in the brain fail to clear debris due to loss of the CD36 molecule, a marker of macrophages associated with the clearance of cellular debris75. We also observed that FABP7 causes downregulation of CD36, which further supports our hypothesis that FABP7-induced changes in monocytes play an important role in the pathobiology of neurodegenerative diseases.
Increased cellular ROS-mediated induction of IL-1β and inflammation in monocytes and macrophages has been demonstrated in various disease models30,76. However, in our study, we did not find a correlation between the increased presence of FABP7 and intracellular ROS or mitochondrial superoxide production in monocytes (Fig. 3j, k). A study of patients with chronic granulomatous diseases showed that IL-1β expression in monocytes could be independent of ROS77, which supports our observation that FABP7 induces IL-1β without generating ROS. Further, we observed that FABP7 partially induced IL-1β via TLR4 interaction, however, without inflammasome activation. It is possible that FABP7 acts via TLR4-mediated signaling and activates NF-κB following increased gene expression associated with inflammation in monocytes78. Being a fatty acid binding protein, FABP7 could also transport arachidonic acid for its metabolism by COX2, which produces prostaglandin that increases NF-κB79. Our transcriptomic analysis (Supplementary Table 5) further supports this hypothesis as we observed an increased expression of inflammation associated transcription factors including NF-κB (NFKB1) and HIF-1α (HIF1A) which could be involved in increased IL-1β expression as shown at the transcriptomics and protein level. Moreover, HIF-1α is highly associated with increased glycolysis and IL-1β production in monocytes and macrophages80,81. Several other mechanisms, such as calcium influx and calmodulin interaction have been identified through which IL-1β is released in primary monocytes82. These observations indicated that FABP7 changes gene expression patterns in monocytes and skews them towards an inflammatory M1 phenotype. However, further validation is required to confirm an association between increased glycolysis and the above-mentioned molecules in FABP7-treated monocytes.
Once we observed the effect of FABP7 in monocytes, we wanted to explore whether monocytes in the brains of MS patients have similar phenotypes. Firstly, there was an increased level of FABP7 in the CSF of SPMS patients and increased expression of FABP7 in astrocytes of MS brain lesion. We expected that CSF FABP7 would be much higher compared to what we observed, however, the treatment status in these patients could affect the level of FABP7 in CSF. Secondly, in our analysis of postmortem MS brain tissue, we found the presence of inflamed microglia (CD14+ CD16+) in the microenvironment of FABP7-expressing astrocytes. A robust statistical association of our in vitro transcriptomics of FABP7-treated monocytes with the inflamed microglia signature further supports our observation that the presence of FABP7 in CSF could modulate the monocyte function. However, this observation does not confirm the FABP7 association with inflamed microglia in the brain and needs further investigation. It remains unclear how astrocyte-expressed FABP7 may encounter brain-resident and invading peripheral immune cells, it is possible that damaged astrocytes release their contents into the extracellular milieu. The reliability of the astrocyte protein GFAP as a serum biomarker for progressive MS is evidence of this phenomenon63. However, the effect of FABP7 on infiltrated immune cells within the CNS microenvironment needs to be explored. In the present study, we have used 100 or 300 ng/ml doses of FABP7 for in vitro study. However, tissue concentrations of FABP7 in the lesion microenvironment may be higher than the CSF or serum levels and different from our in vitro doses and can be considered as a limitation of this study, as there was no previous finding suggesting the in vitro doses of FABP7.
In conclusion (Fig. 7), we have shown for the first time increased serum levels of FABP7 as a biomarker for progressive MS, disability, and brain atrophy. We found that FABP7 increases inflammation in monocytes via promoting glycolysis. Moreover, we show that FABP7 presence in CNS lesions is associated with pro-inflammatory infiltrated monocytes. This adds to previously published mechanisms of FABP7 in promoting CNS inflammation, including the induction of neurotoxic astrocytes and acting as a T cell antigen. Therefore, FABP7 serves as a potential therapeutic target for the prevention of MS progression and other autoimmune diseases that involve CNS inflammation. Further studies are required to validate the role of FABP7 in the CNS using in vivo model system.
(1) Inflammation and BBB damage cause infiltration of monocytes and other immune in the brain during MS. (2) Activated or dying astrocytes release FABP7 which creates an inflammatory environment for the infiltrating monocytes. (3) The release of FABP7 increases glycolysis in monocytes and triggers TLR4-signaling, leading to increased expression of M1 macrophage associated markers. (4) These M1 macrophages increase the inflammation burden by producing interleukins such as IL-1β and promote demyelination in neurons leading to loss of brain volume. (5) The released FABP7 may cross the disrupted BBB and serve as a biomarker for brain atrophy in MS patients. (Created in BioRender. Patel, R. (2025) https://BioRender.com/i41h044).
Methods
Chemicals and reagents
Chemicals and reagents are mentioned in Supplementary Table 1, and various conjugated antibodies are listed in Supplementary Table 2.
Human participants
PBMC samples from HC and patients with a diagnosis of SPMS meeting the McDonald 2017 criteria and enrolled in the ‘The Biorepository for Comprehensive Longitudinal Investigations of Multiple Sclerosis at the Brigham and Women’s Hospital’ (CLIMB bio), IRB: 2017P001169, were utilized in this study. All the participants signed an informed consent form. The sex of the participant was considered in the experiment and determined by a self-reported method. The HC participants were provided with $50 for donating each blood sample and the parking cost was covered for their visits. The parking cost of extra visits for SPMS participants for sample collection was covered, however, these participants were not provided with any cost for sample collection. The CLIMB bio study is approved by the Mass General Brigham Human Research Committee. We used different cohorts that included MS patients and HC. Our clinical cohort consists of RRMS (n = 32, age 37.9 ± 7.2 years, 65.6% female and 34.4% male), RRMS age-matched HC (n = 32, age 33.9 ± 7.8 years, 68.7% female and 31.3% male), SPMS (n = 32, age 58 ± 8 years, 65.6% female and 34.4% male), and SPMS age-matched HC (n = 32, age 55.3 ± 9 years, 65.6% female and 34.4% male). Most of the participants were untreated (Table 1). Our MRI cohort consists of RRMS (n = 30, age 43.5 ± 9.3, 63.3% female and 36.7% male) and SPMS (n = 4, age 51.4 ± 11.6 years, 75% female and 25% male) patients. In this cohort, most of the participants were on treatment (Table 2). All the PBMC samples used were from HC and untreated SPMS patients. All the experimental n are distinct and mentioned if otherwise.
PBMCs isolation and monocyte culture
PBMCs were isolated from the HC or untreated SPMS patients’ blood by the ficoll gradient method and cryopreserved until utilization for the experiments83. To isolate PBMCs, the blood was first layered on top of ficoll (2:1 blood to ficoll ratio) and centrifuged for 20 min at RT with 7 acceleration and 1 deceleration using an Eppendorf centrifuge. Plasma was removed, and PBMCs layer was collected and transferred to another tube containing 30 ml of HBSS buffer. The cell suspension was centrifuged, followed by a rewash with HBSS buffer. Finally, the cell pellet was resuspended in freezing medium (90% Heat inactivated FBS and 10% DMSO) and cryopreserved in − 80 °C for 24 h followed by storing in vapor phase of liquid nitrogen. Further, to isolate monocytes, PBMCs were thawed at 37 °C and proceeded to negative pan monocyte isolation using the manufacturer’s protocol. To isolate monocytes from freshly isolated PBMCs, the PBMCs were immediately proceeded for monocyte isolation using the same kit. We used this negative pan monocyte isolation for all our in vitro experiments. Isolated monocytes were counted using the Cellometer Auto 2000 Cell Counter, and transferred to fresh RPMI medium containing 10% heat inactivated FBS and grown in CO2 incubator at 37 °C with 5% CO2 and 90% humidity. Monocytes were then stimulated with various treatment conditions as per the requirements of the experiment.
Brain MRI
MRI acquisition
Brain MRI acquisition protocol was performed on a 3 T unit (Siemens Skyra)84, using a 20-channel head coil, comprised of 3 sagittal high-resolution sequences, covering the whole head with 1 mm3 isotropic voxel sizes. This included a 3D T1-weighted gradient echo, 3D T2 spin echo, and 3D T2-fluid-attenuated inversion-recovery (FLAIR). The sequences were optimized in contrast for depicting brain-CSF interfaces and white matter lesions.
Whole brain volume
We developed a fully automated quantitative analysis pipeline to measure whole brain volumes84. This involved the co-registration of three MR sequences, anatomical parcellation with heuristic misclassification correction, and an expectation-maximization algorithm. The output provided brain parenchymal fraction (BPF), a normalized measure of whole brain volume, and a surrogate of whole brain atrophy85. This pipeline showed high accuracy and precision84.
Deep gray matter volumes
We used two automated methods for determining normalized deep gray nuclei volumes86. The 3D T1 series was applied to two pipelines: FSL-FIRST (v.5.0, https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/) and SIENAX (v. 5.0), The Analysis Group, Oxford, UK, http://fsl.fmrib.ox.ac.uk/fsl/fslwiki/SIENA) to derive raw volumes of the caudate, putamen, globus pallidus, and thalamus, also summed as total deep gray matter (FSL-FIRST), and a normalization factor used to adjust for intrasubject head size (SIENAX). Gray matter volumes were thus normalized within each participant by multiplying the raw volume by the whole brain SIENAX normalization factor and were provided in ml. We have shown high scan-rescan reliability using this method86.
White matter lesion volumes
Whole brain white matter T1LV and T2LV were quantified by an expert using manual identification on the sagittal T1 and FLAIR images, respectively. The expert employed a semi-automated edge-finding tool in Jim 7.0 software (Xinapse Systems Ltd., West Bergholt, UK; http://www.xinapse.com) to determine contours/volumes of lesions, with manual adjustment as needed. The Jim software then calculated the total lesion volume on each sequence.
ELISA
Serum from MRI matched patients’, HC, RRMS, and SPMS were obtained from CLIMB bio. CSF from SPMS were obtained from the biorepository under Intermediate-Size Patient Population Expanded Access Treatment Protocol at Brigham and Women’s Hospital and non-MS control (Idiopathic intracranial hypertension (IIH) patients) CSF samples were obtained from the Clinical Research Center, Beth Israel Deaconess Medical Center, Boston. All the participants signed an informed consent form. ELISA was performed to detect the level of FABPs by following the manufacturer’s protocol. Briefly, for FABP4, 100 µl of diluted serum (1:10 dilution) per well was incubated for 60 min at room temperature (RT). After washing, the wells were incubated with Biotin-labeled antibody solution for 60 min at RT. This was followed by washing and incubation with Streptavidin-HRP conjugate for 30 min at RT, followed by substrate solution incubation for 10 min and stop solution was added in the final step. For FABP7, 100 µl of serum or diluted CSF (1:2.5 dilution) was used and incubated for 90 min at 37 °C followed by Biotin labeled antibody for 60 min at 37 °C. After washing, HRP-conjugated solution was used for 30 min at 37 °C followed by substrate reagent for 15 min at 37 °C and stop solution was added in the final step. Optical density was observed at 450 nm using a GloMax Explorer microplate reader and concentration of each protein was calculated based on standard curves.
Flow cytometry
Approximately, 75 × 103 monocytes/well were seeded in a U-bottom 96-well cell culture plate and incubated with various treatment conditions for 24 or 48 h in 200 µl of complete medium. In the last 2 h of incubation, cells were stimulated with PMA, Ionomycin, and Golgi stop (PIG) in a complete medium to detect the expression level of cytokines. This was followed by washing twice with 1X PBS and incubation with the live/dead assay dye in 1X PBS for 15 min at 4 °C. After washing twice with 1X PBS, cells were stained against various surface receptor markers (Supplementary Table 2) in FACS buffer (1X PBS containing 0.5% FBS and 2 mM EDTA) for 45 min at 4 °C followed by cell fixation and permeabilization using Fix/perm buffer for 30 min at 4 °C. Cells were then washed twice with permeabilization buffer and incubated with antibodies against cytokines in permeabilization buffer for 45 min at 4 °C. Finally, cells were washed in permeabilization buffer, resuspended in FACS buffer and run in a flow cytometry analyzer (BD LSRFortessa). The FCS files were analyzed using FlowJo software V10, and gating was done based on fluorescence minus one (FMO).
Seahorse assay
To test oxygen consumption rate (OCR) and extracellular acidification rate (ECAR) in monocytes, we performed mitochondrial stress tests using a Seahorse XFe 96 analyzer (Agilent). Approximately, 200,000 monocytes/well were seeded in a U-bottom 96-well cell culture plate and incubated with the treatment conditions for 48 h in 300 µl of complete medium. Cells were washed with XFe RPMI medium and resuspended in 50 µl/well of XFe RPMI medium supplemented with 1 mM pyruvate, 2 mM glutamine, and 10 mM glucose. Cells were then transferred to poly-D-lysine (PDL)-coated (50 µg/ml) XFe 96-well plate in a 50 µl medium/well. Plate was then centrifuged at 200 × g for 5 min at 2 acceleration and 0 deceleration and incubated at 37 °C for 15 min followed by the addition of 130 µl of XFe RPMI medium in each well. The plate was then incubated in a non-CO2 incubator at 37 °C for around 45 min before running the assay. 2 µM Oligomycin, 1.5 µM FCCP, and 0.5 µM Rot/AA were used for the mitochondrial stress test. After running the assay, protein concentration was determined by the BCA assay and used for data normalization.
ROS detection and mitochondrial superoxide detection assay
To analyze ROS level, we performed the ROS detection assay following manufacturer’s protocol. First, we isolated the monocytes and incubated them with ROS detection reagent for 60 min (for internalization of reagent into the cells) in complete medium. Cells were then centrifuged, washed once with 1X PBS, and resuspended in fresh medium. Following this, cells were incubated with FABP7, LPS (100 ng/ml), or H2O2 (1 mM) for 6 h. This was followed by washing and staining the cells with Live/dead assay dye and fixing them in 4% paraformaldehyde for 20 min at 4 °C. Cells were resuspended in FACS buffer and run in a flow cytometry analyzer. The FITC channel was used to detect ROS intensity.
To determine mitochondrial superoxide in monocytes, we first treated the cells with FABP7 or LPS, or H2O2 (200 µM) for 48 h (1 mM H2O2 was found to be extremely toxic by the end of 48 h). This was followed by incubating the cells with Mitosox dye (5 µM) for 30 min in CO2 incubator. After washing with 1X PBS, cells were stained with Live/Dead dye as mentioned in methods for flow cytometry. Cells were fixed in 4% paraformaldehyde for 20 min at 4 °C. Cells were then washed and resuspended in FACS buffer and run in a flow cytometry analyzer87. The PE channel was used to detect mitosox dye intensity.
TLR4 antibody blocking
Approximately, 75 × 103 monocytes/well were seeded in a U-bottom 96 well cell culture plate. The cells were first incubated with 10 µg/ml of TLR4 blocking antibody or IgG isotype control for 1 h in 100 µl/well in CO2 incubator at 37 °C. This was followed by addition of FABP7 and making up the final volume to 200 µl/well. The cells were then incubated for 48 h and flow cytometry was performed as mentioned in methods for flow cytometry.
Fatty acid oxidation assay
To detect fatty acid oxidation in monocytes, we used FAO blue (fatty acid oxidation detection reagent). First, approximately 75 × 103 monocytes/well were incubated with FABP7 or LPS in a total of 200 µl complete medium for 48 h in a U-bottom 96-well cell culture plate. In the last 4 h of incubation, the cells with media were transferred to a PDL-coated 18-well glass bottom µ-Slide (ibidi). The slides were centrifuged at 200 × g with 2 acceleration and 0 deceleration for 5 min to assist cell adherence to the slide surface. Post incubation, media was removed, and cells were washed twice with 100 µl of HEPES Buffered saline solution (HBSS) followed by incubation with 10 µM of FAO blue in HBSS for 1 h in CO2 incubator at 37 °C. After the incubation was completed, cells were washed thrice with HBSS and 100 µl of HBSS was added in each well and images at the DAPI channel were captured immediately. The mean fluorescence intensity was quantified using ImageJ (Fiji) and normalized with the cell numbers per microscopic image.
siRNA transfection
Approximately 75 × 103 monocytes/well were seeded in a U-bottom 96 well cell culture plate in 30 µl of serum free media without antibiotic. The siRNA transfection complex was also prepared in serum-free and antibiotic free media. To prepare the transfection complex, 40 nM siRNA (for final volume in the well) was mixed in media and 6 µl of HiPerfect/ 100 µl complex was added and vortexed for 5 s. This was kept at RT for 8–10 min to form transfection complex and 30 µl of complex was added dropwise on top of the cells. The cells were then incubated in CO2 incubator at 37 °C. After 4 h of incubation, 120 µl media containing FBS (final 10%) was added and incubated for 24 h. This was followed by replacing the media with fresh complete media and incubated for an additional 12 h. Cells were then treated with FABP7 for 48 h, and flow cytometry was performed as mentioned in methods for flowcytometry.
Total RNA isolation and RT-qPCR
Cells were washed twice with 1X PBS and lysed in RL buffer. Total RNA was isolated, and cDNA was synthesized following the manufacturer’s protocol. Briefly, 150 ng of total RNA was used to prepare 20 µl of cDNA. One µl of cDNA was used in a 10 µl of reaction mixture containing TaqMAN master mix and primer (TaqMAM assay) for RT-qPCR. The reaction was heated at 95 °C for 20 s following 40 cycles of 2-step application, 95 °C for 1 s, and 60 °C for 20 s in QuantStudio 7 Flex (Applied Biosystem). Ct values were utilized for calculating differential gene expression. Beta-2-microglobulin (B2M) gene expression was used as a loading control. Standard formula 2- ΔΔCT was used to calculate relative gene expression.
Bulk RNA sequencing
Approximately, 75 × 103 monocytes/well were seeded in a U-bottom 96 well cell culture plate and treated with FABP7 in 200 µl of complete medium and incubated for 24 h. After completion of the incubation, cells were washed twice with 1X PBS. Cells were then lysed in TCL buffer containing 1% BME at a density of 1000 cells/ 5 µl of TCL buffer and stored at − 80 °C until processed for sequencing. The SmatSeq2 population sequencing was performed at the Genomics facility at Broad Institute, Cambridge, MA, USA. Post-sequencing, the raw sequencing reads were processed through FASTQC for quality control assessment. The reads were further aligned to the reference genome GRCh38 using the hisat2 aligner. The transcripts were quantified using the featureCounts function of the subread Bioconductor library. Post sequencing, the analysis was performed using R packages V4.3.
HDOCK in silico protein-protein interaction analysis
To investigate the likelihood of binding between FABP7 and TLR4, models of putative docking interactions were generated in HDOCK (PMID: 32269383) using the hybrid template based and template free algorithm. Crystal structures for modeling were sourced from the PDB. Structure 7E25 (PMID: 34196621) was selected for FABP7, and structure 3FXI (PMID: 19252480) was selected for the TLR4-MD2 complex. Full structures were uploaded to HDOCK for the FABP7 and the TLR4-MD2 complex as the ligand and receptor, respectively. All parameters were set to default values. Visualizations of resulting models were generated in PyMOL.
Immunofluorescence
Fresh-frozen brain tissue samples from MS patient and HC were used for cryosectioning. The brain samples were obtained from The Netherlands Brain Bank (NBB), Netherlands Institute for Neuroscience, Amsterdam (https://www.brainbank.nl/). All Material has been collected from donors for or from whom a written informed consent for a brain autopsy and the use of the material and clinical information for research purposes had been obtained by the NBB.
The sections with 10-micron thickness were transferred to Superfrost Plus Microscope Slides and processed for staining. Sections were fixed with 4% formaldehyde at RT for 15 min and washed thrice for 5 min each with 1X PBS. Tissue sections were permeabilized with 0.4% Triton X-100 for 15 min at RT and washed thrice with 1X PBS. This was followed by blocking the sections with blocking buffer (Immunofluorescence Application Solutions Kit) for 60 min at RT. Sections were then incubated overnight at 4 °C with antibodies against FABP7 and GFAP in antibody dilution buffer. The sections were washed thrice with 1X PBS and completely dried. The sections were mounted with Prolong Gold Antifade Reagent with DAPI and kept overnight at RT before taking the images. A confocal microscope (Zeiss LSM710) was used to capture images at 200X magnification. Images at 5 random locations within the lesion or normal appearing white matter (NAWM) in HC were captured. Images were analyzed using ImageJ (Fiji) software, and mean fluorescence intensity was quantified.
Postmortem brain tissue and Single Nuclei RNA sequencing
Single-nucleus RNA-seq datasets from postmortem MS brain tissue58 – categorized as normal appearing periplaque, lesion core, lesion rim, and white matter from non-neurologically-diseased healthy controls – were downloaded from GEO under accession number GSE180759. Raw data was processed using Cell Ranger v7.1.0 and Seurat 4.4. Our analysis pipeline deviated slightly from that of the original authors in accordance with recently developed computational techniques. These improve upon data quality over former snRNA sequencing analysis best practices, which often employ arbitrary and uniform thresholds88 during quality control (QC). We opted for data-driven (QC): specifically, an initial low-quality droplet filtering step was applied based on mitochondrial gene content < 3 median absolute deviations (MADs) above the mean. Minimum and maximum UMI thresholds were determined for each library individually according to the observed distribution for that library. Suspected doublets/multiplets were identified and labeled for each library using DoubletFinder89. Seurat v4 was then used to integrate all datasets90. After integration, QC metrics were assessed by library, patient, and cell cluster. In order to estimate the optimal number of clusters in the snRNA sequencing data for each broad category of cell type, we used MultiK91. Briefly, MultiK assess candidate numbers of clusters after subsampling and performing Louvain clustering92 with a range of resolution parameters. Features are re-selected during each run. Next, MultiK generates a consensus matrix with the same K groups regardless of the resolution parameter (Supplementary Fig 14). The relative proportion of ambiguous clustering PAC (rPAC) score is used to identify an optimal K. Next, we used an iterative clustering-based approach to identify and remove doublets. The percentage of doublets (from DoubletFinder) was calculated for each cluster. Clusters with excessive doublets were removed, and data were re-clustered. This process was repeated until all clearly doublet clusters were removed. Cell type identification was then performed manually, with details for individual cell types documented in the supplement. Code for processing raw data, running all analyses, and generating figures is available on GitHub. All these analyses were performed using R packages V4.3.
Statistical analysis
For FABPs serum levels and brain atrophy in MS patients, we fit a simple linear regression for each FABP level as an outcome and individual brain volumes as the predictor to assess the relationship between the FABPs and brain volumes. For differential gene expression (DGE) analysis between groups was performed using the Bioconductor DESeq2 library. The analysis was performed for all the samples by adjusting for sex and then stratified by sex.
In in vitro experiments, we used GraphPad Prism V10 to perform statistical comparisons. A paired or unpaired parametric t test was performed to compare between two groups, as mentioned for each result. For multiple comparisons, where comparing all the groups to control, one-way ANOVA was performed, followed by correction with Dunnett’s multiple comparisons test. To compare two groups in multiple comparisons, one-way ANOVA or two-way ANOVA was performed, followed by correction with Šidák multiple comparisons test and mentioned for each result. Further, p-value ≤ 0.05 was statistically considered significant. Whenever possible, for the p-values less than 0.0001, we used R V4.3 and conducted the same analysis to obtain the exact p-value.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the Supplementary Information or available from the corresponding author, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. The bulk RNA sequencing data have been deposited in the ‘Sequence Read Archive’ under the accession number PRJNA1259057. The snRNA sequencing data for postmortem brain tissue were obtained from the ‘Gene Expression Omnibus’ under the accession number GSE180759 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA749443). Source data are provided in this paper.
Code availability
The code generated for analyzing bulk RNA sequencing and published snRNA sequencing analysis are available at GitHub (https://github.com/TNRC-code/FABP7/).
References
Baecher-Allan, C., Kaskow, B. J. & Weiner, H. L. Multiple sclerosis: mechanisms and immunotherapy. Neuron 97, 742–768 (2018).
Reich, D. S., Lucchinetti, C. F. & Calabresi, P. A. Multiple sclerosis. N. Engl. J. Med. 378, 169–180 (2018).
Olsson, T., Barcellos, L. F. & Alfredsson, L. Interactions between genetic, lifestyle and environmental risk factors for multiple sclerosis. Nat. Rev. Neurol. 13, 25–36 (2017).
International Multiple Sclerosis Genetics, C. Multiple sclerosis genomic map implicates peripheral immune cells and microglia in susceptibility. Science 365, https://doi.org/10.1126/science.aav7188 (2019).
Taan, M., Al Ahmad, F., Ercksousi, M. K. & Hamza, G. Risk factors associated with multiple sclerosis: A case-control study in Damascus, Syria. Mult. Scler. Int. 2021, 8147451 (2021).
Bhagavati, S. Autoimmune disorders of the nervous system: Pathophysiology, clinical features, and therapy. Front. Neurol. 12, 664664 (2021).
Bjornevik, K. et al. Longitudinal analysis reveals high prevalence of Epstein-Barr virus associated with multiple sclerosis. Science 375, 296–301 (2022).
Jäckle, K. et al. Molecular signature of slowly expanding lesions in progressive multiple sclerosis. Brain 143, 2073–2088 (2020).
Lassmann, H. Pathogenic mechanisms associated with different clinical courses of multiple sclerosis. Front. Immunol. 9, 3116 (2018).
Kuhlmann, T. et al. Multiple sclerosis progression: time for a new mechanism-driven framework. Lancet Neurol. 22, 78–88 (2023).
Huang, X., Hussain, B. & Chang, J. Peripheral inflammation and blood-brain barrier disruption: effects and mechanisms. CNS Neurosci. Ther. 27, 36–47 (2021).
Biber, K., Möller, T., Boddeke, E. & Prinz, M. Central nervous system myeloid cells as drug targets: current status and translational challenges. Nat. Rev. Drug Discov. 15, 110–124 (2016).
Prineas, J. W. & Parratt, J. D. E. Multiple sclerosis: Microglia, monocytes, and macrophage-mediated demyelination. J. Neuropathol. Exp. Neurol. 80, 975–996 (2021).
Häusler, D. et al. CNS inflammation after natalizumab therapy for multiple sclerosis: A retrospective histopathological and CSF cohort study. Brain Pathol. 31, e12969 (2021).
Chitnis, T. & Weiner, H. L. CNS inflammation and neurodegeneration. J. Clin. Invest. 127, 3577–3587 (2017).
Yamasaki, R. et al. Differential roles of microglia and monocytes in the inflamed central nervous system. J. Exp. Med. 211, 1533–1549 (2014).
Varvel, N. H. et al. Infiltrating monocytes promote brain inflammation and exacerbate neuronal damage after status epilepticus. Proc. Natl. Acad. Sci. USA 113, E5665–E5674 (2016).
Parisi, L. et al. Macrophage polarization in chronic inflammatory diseases: Killers or builders?. J. Immunol. Res. 2018, 8917804 (2018).
Ajami, B., Bennett, J. L., Krieger, C., McNagny, K. M. & Rossi, F. M. V. Infiltrating monocytes trigger EAE progression, but do not contribute to the resident microglia pool. Nat. Neurosci. 14, 1142–1149 (2011).
Dowling, P. et al. Cell death and birth in multiple sclerosis brain. J. Neurol. Sci. 149, 1–11 (1997).
Trapp, B. D. et al. Axonal transection in the lesions of multiple sclerosis. N. Engl. J. Med. 338, 278–285 (1998).
Prineas, J. W. & Lee, S. Multiple sclerosis: Destruction and regeneration of astrocytes in acute lesions. J. Neuropathol. Exp. Neurol. 78, 140–156 (2019).
Kennedy, P. G. E., George, W. & Yu, X. The possible role of neural cell apoptosis in multiple sclerosis. Int. J. Mol. Sci. 23, https://doi.org/10.3390/ijms23147584 (2022).
Linnerbauer, M., Wheeler, M. A. & Quintana, F. J. Astrocyte crosstalk in CNS inflammation. Neuron 108, 608–622 (2020).
Tang, D. et al. Extracellular vesicle/macrophage axis: Potential targets for inflammatory disease intervention. Front. Immunol. 13, 705472 (2022).
Zhu, X. et al. Frontline Science: Monocytes sequentially rewire metabolism and bioenergetics during an acute inflammatory response. J. Leukoc. Biol. 105, 215–228 (2019).
Viola, A., Munari, F., Sánchez-Rodríguez, R., Scolaro, T. & Castegna, A. The metabolic signature of macrophage responses. Front. Immunol. 10, 1462 (2019).
Lee, M. K. S. et al. Glycolysis is required for LPS-induced activation and adhesion of human CD14(+)CD16(-) monocytes. Front. Immunol. 10, 2054 (2019).
Kaushik, D. K. et al. Enhanced glycolytic metabolism supports transmigration of brain-infiltrating macrophages in multiple sclerosis. J. Clin. Invest. 129, 3277–3292 (2019).
Varghese, J. F., Patel, R. & Yadav, U. C. S. Sterol regulatory element binding protein (SREBP) -1 mediates oxidized low-density lipoprotein (oxLDL) induced macrophage foam cell formation through NLRP3 inflammasome activation. Cell Signal 53, 316–326 (2019).
Wang, W. Y., Tan, M. S., Yu, J. T. & Tan, L. Role of pro-inflammatory cytokines released from microglia in Alzheimer’s disease. Ann. Transl. Med. 3, 136 (2015).
Meyer-Arndt, L. et al. Inflammatory cytokines associated with multiple sclerosis directly induce alterations of neuronal cytoarchitecture in human neurons. J. Neuroimmune Pharmacol. 18, 145–159 (2023).
Cheng, A., Jia, W., Kawahata, I. & Fukunaga, K. A novel fatty acid-binding protein 5 and 7 inhibitor ameliorates oligodendrocyte injury in multiple sclerosis mouse models. EBioMedicine 72, 103582 (2021).
Bove, R. et al. Fatty acid binding protein-4 is associated with disability in multiple sclerosis patients. Mult. Scler. 25, 344–351 (2019).
Furuhashi, M. et al. Independent and distinct associations of FABP4 and FABP5 with metabolic parameters in type 2 diabetes mellitus. Front. Endocrinol.11, 575557 (2020).
Xiao, Y. et al. Fatty acid binding protein 4 promotes autoimmune diabetes by recruitment and activation of pancreatic islet macrophages. JCI Insight 6, https://doi.org/10.1172/jci.insight.141814 (2021).
Messina, S. et al. Increased leptin and A-FABP levels in relapsing and progressive forms of MS. BMC Neurol. 13, 172 (2013).
Keyhanian, K. et al. Adipokines are associated with pediatric multiple sclerosis risk and course. Mult. Scler. Relat. Disord. 36, 101384 (2019).
Killoy, K. M., Harlan, B. A., Pehar, M. & Vargas, M. R. FABP7 upregulation induces a neurotoxic phenotype in astrocytes. Glia 68, 2693–2704 (2020).
Sharifi, K. et al. FABP7 expression in normal and stab-injured brain cortex and its role in astrocyte proliferation. Histochem. Cell Biol. 136, 501–513 (2011).
Koga, M. et al. Plasma fatty acid-binding protein 7 concentration correlates with depression/anxiety, cognition, and positive symptom in patients with schizophrenia. J. Psychiatr. Res. 144, 304–311 (2021).
Kamizato, K. et al. The role of fatty acid binding protein 7 in spinal cord astrocytes in a mouse model of experimental autoimmune encephalomyelitis. Neuroscience 409, 120–129 (2019).
Teunissen, C. E. et al. Brain-specific fatty acid-binding protein is elevated in serum of patients with dementia-related diseases. Eur. J. Neurol. 18, 865–871 (2011).
Shimamoto, C. et al. Functional characterization of FABP3, 5 and 7 gene variants identified in schizophrenia and autism spectrum disorder and mouse behavioral studies. Hum. Mol. Genet. 24, 2409 (2015).
Watanabe, A. et al. Fabp7 maps to a quantitative trait locus for a schizophrenia endophenotype. PLoS Biol. 5, e297 (2007).
Venkatraman, V. et al. Filtered cerebrospinal fluid from patients with amyotrophic lateral sclerosis displays an altered proteome and affects motor phenotype in a mouse model. Cureus 14, e32980 (2022).
Halford, J. et al. New astroglial injury-defined biomarkers for neurotrauma assessment. J. Cereb. Blood Flow. Metab. 37, 3278–3299 (2017).
Waschbisch, A. et al. Pivotal role for CD16+ monocytes in immune surveillance of the central nervous system. J. Immunol. 196, 1558–1567 (2016).
Bronge, M. et al. Identification of four novel T cell autoantigens and personal autoreactive profiles in multiple sclerosis. Sci. Adv. 8, eabn1823 (2022).
Fiedler, S. E. et al. Analysis of IL-6, IL-1β and TNF-α production in monocytes isolated from multiple sclerosis patients treated with disease modifying drugs. J. Syst. Integr. Neurosci. 3, https://doi.org/10.15761/jsin.1000166 (2017).
Abdelhak, A., Huss, A., Kassubek, J., Tumani, H. & Otto, M. Serum GFAP as a biomarker for disease severity in multiple sclerosis. Sci. Rep. 8, 14798 (2018).
Lee, M. K. S. et al. Glycolysis is required for LPS-induced activation and adhesion of human CD14 + CD16 − monocytes. Front. Immunol. 10, https://doi.org/10.3389/fimmu.2019.02054 (2019).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with enrichr. Curr. Protoc. 1, e90 (2021).
Wang, Y. et al. TLR4/MD-2 activation by a synthetic agonist with no similarity to LPS. Proc. Natl. Acad. Sci. USA 113, E884–E893 (2016).
Eltom, S. et al. TLR4 activation induces IL-1β release via an IPAF dependent but caspase 1/11/8 independent pathway in the lung. Respir. Res. 15, 87 (2014).
Absinta, M. et al. A lymphocyte-microglia-astrocyte axis in chronic active multiple sclerosis. Nature 597, 709–714 (2021).
Gillen, K. M., Mubarak, M., Nguyen, T. D. & Pitt, D. Significance and in vivo detection of iron-laden microglia in white matter multiple sclerosis lesions. Front. Immunol. 9, 255 (2018).
Pukoli, D. & Vécsei, L. Smouldering lesion in MS: Microglia, lymphocytes and pathobiochemical mechanisms. Int. J. Mol. Sci. 24, https://doi.org/10.3390/ijms241612631 (2023).
Mathur, D. et al. Potential biomarkers associated with multiple sclerosis pathology. Int. J. Mol. Sci. 22, https://doi.org/10.3390/ijms221910323 (2021).
Ziemssen, T., Akgün, K. & Brück, W. Molecular biomarkers in multiple sclerosis. J. Neuroinflamm. 16, 272 (2019).
Barro, C. et al. Serum GFAP and NfL levels differentiate subsequent progression and disease activity in patients with progressive multiple sclerosis. Neurol. Neuroimmunol. Neuroinflamm. 10, https://doi.org/10.1212/nxi.0000000000200052 (2023).
Tripathi, S. et al. Docosahexaenoic acid up-regulates both PI3K/AKT-dependent FABP7-PPARgamma interaction and MKP3 that enhance GFAP in developing rat brain astrocytes. J. Neurochem. 140, 96–113 (2017).
Aharoni, R., Eilam, R. & Arnon, R. Astrocytes in multiple sclerosis-essential constituents with diverse multifaceted functions. Int. J. Mol. Sci. 22, https://doi.org/10.3390/ijms22115904 (2021).
Mayo, L. et al. Regulation of astrocyte activation by glycolipids drives chronic CNS inflammation. Nat. Med. 20, 1147–1156 (2014).
Giovannoni, F. & Quintana, F. J. The role of astrocytes in CNS inflammation. Trends Immunol. 41, 805–819 (2020).
Cao, Y. et al. Phenotypic and functional alterations of monocyte subsets with aging. Immun. Ageing 19, 63 (2022).
Almsned, F., Lipsky, R. H. & Jafri, M. S. Transcriptomic analysis of multiple sclerosis patient-derived monocytes by RNA-sequencing for candidate gene discovery. Inform. Med. Unlocked 23, 100563 (2021).
Chiricosta, L., Blando, S., D’Angiolini, S., Gugliandolo, A. & Mazzon, E. A comprehensive exploration of the transcriptomic landscape in multiple sclerosis: A systematic review. Int. J. Mol. Sci. 24, https://doi.org/10.3390/ijms24021448 (2023).
Kelly, B. & O’Neill, L. A. J. Metabolic reprogramming in macrophages and dendritic cells in innate immunity. Cell Res. 25, 771–784 (2015).
Palmer, C. S., Cherry, C. L., Sada-Ovalle, I., Singh, A. & Crowe, S. M. Glucose metabolism in T cells and monocytes: New perspectives in HIV pathogenesis. EBioMedicine 6, 31–41 (2016).
Pillon, N. J. et al. Saturated fatty acids activate caspase-4/5 in human monocytes, triggering IL-1β and IL-18 release. Am. J. Physiol. Endocrinol. Metab. 311, E825–e835 (2016).
Kagawa, Y. et al. FABP7 Regulates acetyl-CoA metabolism through the interaction with ACLY in the nucleus of astrocytes. Mol. Neurobiol. 57, 4891–4910 (2020).
Grajchen, E. et al. CD36-mediated uptake of myelin debris by macrophages and microglia reduces neuroinflammation. J. Neuroinflamm. 17, 224 (2020).
Santos, D. et al. IL-1β Production by intermediate monocytes is associated with immunopathology in Cutaneous Leishmaniasis. J. Invest. Dermatol. 138, 1107–1115 (2018).
van de Veerdonk, F. L. et al. Reactive oxygen species-independent activation of the IL-1beta inflammasome in cells from patients with chronic granulomatous disease. Proc. Natl. Acad. Sci. USA 107, 3030–3033 (2010).
Hamilton, H. L. et al. FABP7 drives an inflammatory response in human astrocytes and is upregulated in Alzheimer’s disease. GeroScience 46, 1607–1625 (2024).
Needham, H. et al. A dichotomous role for FABP7 in sleep and Alzheimer’s disease pathogenesis: A hypothesis. Front. Neurosci. 16, 798994 (2022).
Del Rey, M. J. et al. Hif-1α knockdown reduces glycolytic metabolism and induces cell death of human synovial fibroblasts under normoxic conditions. Sci. Rep. 7, 3644 (2017).
Tannahill, G. M. et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1α. Nature 496, 238–242 (2013).
Ainscough, J. S., Gerberick, G. F., Kimber, I. & Dearman, R. J. Interleukin-1β processing is dependent on a Calcium-mediated interaction with Calmodulin. J. Biol. Chem. 290, 31151–31161 (2015).
Saxena, S. et al. Identification of TNFAIP3 as relapse biomarker and potential therapeutic target for MOG antibody associated diseases. Sci. Rep. 10, 12405 (2020).
Meier, D. S. et al. Dual-sensitivity multiple sclerosis lesion and CSF segmentation for multichannel 3T brain MRI. J. Neuroimaging 28, 36–47 (2018).
Sharma, J. et al. Whole-brain atrophy in multiple sclerosis measured by automated versus semiautomated MR imaging segmentation. AJNR Am. J. Neuroradiol. 25, 985–996 (2004).
Chu, R., Hurwitz, S., Tauhid, S. & Bakshi, R. Automated segmentation of cerebral deep gray matter from MRI scans: effect of field strength on sensitivity and reliability. BMC Neurol. 17, 172 (2017).
Yang, Y. et al. A flow-cytometry-based protocol for detection of mitochondrial ROS production under hypoxia. STAR Protoc. 2, 100466 (2021).
Hippen, A. A. et al. miQC: An adaptive probabilistic framework for quality control of single-cell RNA-sequencing data. PLoS Comput. Biol. 17, e1009290 (2021).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: Doublet detection in single-Cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Liu, S., Thennavan, A., Garay, J. P., Marron, J. S. & Perou, C. M. MultiK: an automated tool to determine optimal cluster numbers in single-cell RNA sequencing data. Genome Biol. 22, 232 (2021).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory Exp. 2008, P10008 (2008).
Acknowledgements
This study was funded by the Department of Defense (W81XWH1810648 to T.C.) and the National Institutes of Health (R21NS123826 to T.C.). We thank the NeuroTechnology Studio at Brigham and Women’s Hospital for providing the ‘Agilent Seahorse XFe96 Analyzer’ instrument access and consultation on data acquisition and data analysis.
Author information
Authors and Affiliations
Contributions
R.P. and T.C. designed the study and experimental setup. R.P. performed experiments, analyzed the results, and wrote the manuscript. T.C. identified patients, analyzed the results, and added to the manuscript writing. D.K. performed analysis of single-nuclei sequencing and added in manuscript writing. B.L. helped analyzing single-nuclei sequencing. H.L. performed data analysis and added to the manuscript writing. D.C. and J.F.V. performed experiments and analyzed data, S.S. and A.Z. performed experiments. R.B. and K.W. analyzed the MRI data and added in manuscript writing. M.A.B. identified patients and arranged sample collections. All the authors contributed to reading, editing, and approving the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sara De Biasi, Heinz Wiendl and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Patel, R., King, D., LaBarre, B. et al. FABP7 is increased in progressive multiple sclerosis and induces a pro-inflammatory phenotype in monocytes through a glycolytic switch. Nat Commun 16, 6049 (2025). https://doi.org/10.1038/s41467-025-60747-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60747-9