Abstract
In brain development, neural stem cells (NSCs) undergo asymmetric cell divisions to replicate themselves and meanwhile produce differentiating siblings. It remains obscure how NSCs preserve their self-renewing fate across mitosis. Even less is known how cell fate memory is differentially propagated to sibling daughter cells adopting distinct cell fates. Here we found that key differentiation genes are dually bookmarked by pioneer factor GAF (GAGA factor) and H3K27ac in asymmetrically-dividing Drosophila central brain NSCs. In daughter cells adopting NSC fate, GAF promotes self-renewal through timely inhibiting differentiation genes via HDAC1-mediated H3K27 deacetylation, whereas in sibling daughter cells adopting neural progenitor fate, GAF occupancy is replaced by competitor SWI/SNF complex, allowing retention of H3K27ac mark and fast activation of differentiation genes. Thus, our study unveils a paradigm by which cell fate memory can be differentially transmitted to sibling daughter cells via dual antagonistic mitotic bookmarking and selective molecular competition mechanism.
Similar content being viewed by others
Introduction
Brain is the most complex organ in our body. A typical human brain processes 86 billion neurons of nearly 3000 types, all derived from a limited number of neural stem cells (NSCs) in early fetal development1,2. To generate all these neurons of enormous number and great diversity, NSCs need to continuously undergo asymmetric cell divisions to replicate themselves and meanwhile produce differentiating siblings such as neural progenitors, neural precursors or post-mitotic neurons3.
However, each round of mitotic division poses a huge challenge for preserving cell fate in a precise and timely manner. Once a cell enters mitosis, chromatin is highly condensed to form compact chromosomes4. As a consequence, the vast majority of chromatin-associating proteins, including transcription factors (TFs), RNA polymerase II, chromatin remodelers and histone modifying enzymes, are expelled or degraded5,6. Cell fate-specific transcriptional programs are hence almost completely halted and three-dimensional (3D) chromatin architecture fundamentally altered7,8, presenting a daunting challenge for a cell to reestablish its characteristic transcriptional programs and 3D chromatin architecture with high fidelity and speed upon mitotic exit.
Mitotic bookmarking has been posited as an important strategy for solving this conundrum9,10,11,12. That is, specific transcriptional factors or chromatin remodelers retained on highly condensed chromosomes can “bookmark” key target genes and foster their fast transcriptional reactivation, thereby ensuring faithful propagation of cell fate memory13,14,15,16,17,18,19. However, whether mitotic bookmarking plays a significant role in neural development remains enigmatic.
More importantly, if mitotic bookmarking indeed plays a crucial role in preserving NSC fate memory, a new puzzle emerges. That is, after asymmetric cell divisions, how NSC-specific self-renewal fate memory is preserved in one daughter cell but selectively erased in its sibling cell adopting a differentiating fate? Since previous studies on mitotic bookmarking are mainly conducted in symmetrically dividing cells, how differential mitotic bookmarking is achieved remains unexplored.
GAF (GAGA factor) is a pioneer factor that plays an essential role in Drosophila early embryogenesis20,21,22,23. As a BTB-ZF (broad-complex, tramtrack and bric-à-brac-zinc finger) transcription factor, GAF has been found to perform vital functions in a range of biological processes including heat shock response, silencing of satellite repeats, position-effect variegation and maternal-to-zygotic transition22,24,25,26,27,28,29. Significantly, previous studies revealed that GAF is mitotically retained at specific target genes in dividing cells of fly early embryos and is essential for fast transcriptional reactivation upon mitotic exit of embryonic cells22,28,30.
The SWI/SNF complex is an evolutionarily conserved ATP-dependent chromatin remodeler that controls transcription through modulating chromatin accessibility at enhancers or promoters31,32,33. Mutations in SWI/SNF subunits have been found in more than 20% of human cancers and several types of neurodevelopmental disorders34,35. Intriguingly, earlier studies indicated that inactivation of SWI/SNF subunits leads to dedifferentiation of fly neural progenitors back to NSC-like cells and brain tumor formation36.
The formation of membraneless biomolecular condensates allows cells to concentrate specific proteins and increase local reaction rate37,38. For example, the formation of TF condensates has been linked to increased rate of transcription activation39,40. Biomolecular condensates have been found to play crucial roles in a wide range of biological processes41,42,43,44. However, it remains unclear whether condensate formation ability is important for mitotic bookmarking and cell fate memory propagation.
The type II NSCs (so called neuroblasts) in the central brain area of Drosophila larvae provide an ideal model system for understanding the molecular mechanisms and functions of mitotic bookmarking in neural development. Firstly, type II NSC lineages, controlled by highly conserved regulatory molecules and signaling pathways, are anatomically and functionally similar to mammalian NSC lineages yet with much simpler lineage composition (Fig. 1a)3,45,46,47,48. Secondly, type II NSCs divide very fast, with each mitotic cell division spanning only about two hours49,50, making it relatively easier to observe the distribution of mitotically-retained proteins on dividing chromosomes; Thirdly, type II NSCs, a well-characterized model for studying asymmetric stem cell division51,52,53,54, each harboring only four pairs of chromosomes, allow an easy assessment whether the segregation of a mitotic bookmarker into sibling daughter cells are symmetric. Finally, powerful fly genetics combined with fast-developing multi-omics tools greatly facilitate a mechanistic understanding of mitotic bookmarking in neural development55.
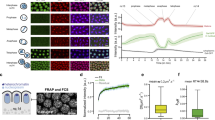
a A schematic drawing of a fly type II neural stem cell (NSC) lineage. Created in BioRender. Li, Y. (2025) https://BioRender.com/pcljp15b GFP-GAF-KI (knock-in) retains at the chromosomes of dividing NSCs throughout cell cycle. Chromosome regions decorated by big-sized GAF foci and uniform distribution of GAF are indicated by hollow and solid arrowheads respectively. Scale bar, 5 μm. c, d Quantification of mitotic retention index of GFP-GAF-KI at H3K9me3- euchromatin regions (c) and H3K9me3+ heterochromatin regions (d) of NSCs. n = 10 (c), n = 11 (d). Data are presents as mean values ± SD (standard deviation). e–g In vivo FRAP analysis of GAF-GFP droplets highlighted by the white box in interphase (top) or metaphase (bottom) NSCs of ex vivo larval brain (e). Quantification of FRAP of GFP-GAF condensates over indicated time course is shown in (f, g). n = 5 (f), n = 9 (g). Time 0 indicates the start of recovery after photobleaching. Scale bar, 5 μm. h OptoDroplet assay in HEK293T cells. Cells expressing the optoIDR constructs were subjected to blue light activation. Scale bar, 5 μm. i Larval brain lobes of indicated genotypes were stained for Dpn (Deadpan, red) and Ase (Asense, blue) to identify type II NSCs (Dpn+ Ase-), neural progenitors (Dpn+ Ase+) and neural precursors (Dpn- Ase+). In close-up images (bottom panel), type II NSC lineages are outlined by white dashed lines. Scale bar, 50 μm (top), 10 μm (bottom). j, k Quantification of total type II NSC number per brain lobe (j) and total progenitor or precursor number per type II NSC lineage (k) of indicated genotypes. **p = 0.0028 (j), ****p < 0.0001 (k). n = 5, 10 (j), n = 15, 13 (k). l, m MARCM clonal analysis of WT (wild type), GAFKO (GAF knockout), and GAF.FL (GAF full-length) expression in GAFKO background (l). MARCM clones are marked by CD8-GFP (gray). Quantification of total cell number in each clone is shown in (m). ****p < 0.0001, ns, not significant. n = 6, 5, 10. Scale bar, 10 μm. Source data are provided as a Source Data file.
Through an in vivo unbiased RNAi-based genetic screen for regulators promoting fly NSC fate decision49, we identified the pioneer factor GAF as a strong hit. Considering that GAF has been found to be associated with mitotic chromosomes in fly embryos21,28, we sought to investigate whether GAF acts as a mitotic bookmarking factor in preserving transcriptional memory of NSCs.
Here, we found that GAF, via condensate formation, acts as a mitotic bookmarker to propagate NSC fate memory and promote NSC self-renewal. More importantly, we revealed that, after NSC asymmetric cell division, cell fate memory can be differentially preserved or acquired in sibling daughter cells. Such differential prorogation of cell fate memory is achieved through dual antagonistic mitotic bookmarking by GAF and H3K27ac coupled with selective molecular competition by the SWI/SNF complex.
Results
Mitotic retention of GAF in dividing neural stem cells
To examine whether GAF protein exhibits mitotic retention in neural stem cells, we first transiently expressed FLAG-tagged GAF transgenes in NSC lineages at the central brain area of larval brains. There exist two major GAF splicing isoforms, RA and RB56,57. Throughout cell cycle, FLAG-GAF.RA faintly decorates the whole dividing chromosomes of NSCs and meanwhile accumulates as discrete foci at pericentromeric regions (Supplementary Fig. 1a).
To assess the expression and dynamics of endogenous GAF protein in NSCs, we generated GFP-tagged GAF.RA knock-in (KI) lines via the CRISPR-Cas9 system (Supplementary Fig. 1b–d). Consistent with its transgenic distribution, GFP-GAF.RA-KI forms foci and is weakly retained at the whole mitotic chromosomes of NSCs throughout the cell cycle (Fig. 1b–d). We hereafter focused on GAF.RA to investigate the significance of mitotic bookmarking in preserving NSC fate memory.
GAF forms phase separation-based condensates in NSCs
We next investigated whether GAF foci exhibit liquid demixing properties. Fluorescence recovery after photobleaching (FRAP) experiments were first performed to probe the dynamics of GAF foci in vivo. In interphase NSCs transiently expressing GAF-GFP, GAF-GFP signal rapidly recovers after photobleaching (Fig. 1e, f, Supplementary Fig. 1e), indicating that GAF protein is highly dynamic and freely exchanges with its cytoplasmic pool. Time-lapse live imaging further revealed that the GAF-GFP major foci are highly mobile and occasionally fuse with each other and round up (arrowheads in Supplementary Fig. 1f). In dividing NSCs transiently overexpressing GAF-GFP, however, GAF-GFP signal cannot fully recover after photobleaching (Fig. 1e, g, Supplementary Fig. 1e, g), indicating that GAF protein decorating mitotic chromosomes is less dynamic and contains immobile fraction. Importantly, these results indicate that GAF-GFP exhibits distinct biophysical properties in NSCs at different cell cycle stages (Supplementary Fig. 1e). Furthermore, the big-sized GFP-GAF.RA-KI foci largely overlap with the heterochromatin mark H3K9me3 (Supplementary Fig. 2a).
Next, optoDroplet assay58 was carried out to directly assess the liquid signatures of GAF in HEK293T cells. Upon blue light illumination, while Cry2 protein itself does not show any clustering, fusing the DNA-binding domain-deleted form of GAF (GAF.ΔZF) to Cry2 (GAF.ΔZF-mCherry-Cry2) leads to rapid assembly of spherical, droplet-like clusters (Fig. 1h). Taken together, GAF forms phase separation-based condensates in NSCs, with big-sized condensates overlapping with pericentromeric heterochromatin.
GAF is critical for preserving neural stem cell self-renewal and proliferation capacity
To investigate the functional significance of GAF in NSCs, we specifically knocked down GAF in type II NSC lineages. Knockdown of GAF through RNAi (with GAF-IR-1) leads to reduced number of type II NSC lineages in each brain lobe from 8 in wild type to on average 5.6 (Fig. 1i, j), indicating that GAF is essential for preserving type II NSC self-renewal capacity. In accordance, all the remaining type II NSC lineages in GAF-IR-1 brains drastically shrink, with the number of neural progenitors and neural precursors in each lineage dramatically reduced (Fig. 1i, k). Knockdown of GAF with another RNAi line GAF-IR-2 results in strong lineage shrinkage of all 8 type II NSC lineages and drastic reduction in the number of neural progenitors and precursors (Supplementary Fig. 2b–d), demonstrating that GAF is also critical for NSC proliferation. Fly FUCCI (fluorescent ubiquitination-based cell cycle indicator) analysis59, cell polarity assay and P35 rescue experiments further indicate that the NSC and progenitor loss phenotypes induced by GAF knockdown are not due to impaired cell cycle progression, defective asymmetric cell division or apoptosis (Supplementary Fig. 2e–k). Taken together, GAF is essential for both NSC self-renewal and proliferation.
To further confirm the GAF loss-of-function phenotypes in type II NSC lineages, we generated GAF knockout (GAFKO) fly line using the CRISPR-Cas9 system (Supplementary Fig. 1b). Since GAFKO line is homozygous lethal at embryonic stages, we assessed the GAFKO phenotypes by carrying out MARCM (Mosaic analysis with a repressible cell marker) analysis55. Consistent with GAF knockdown phenotypes, GAFKO MARCM clones derived from individual NSCs in type II DM5 or DM6 lineages exhibited strong NSC proliferation defects (Fig. 1l, m, Supplementary Fig. 2b, l, n). The expression of FLAG-GAF transgene in GAFKO MARCM clones completely rescue the NSC proliferation defects (Fig. 1l, m), confirming that GAF is critical for preserving NSC self-renewal and proliferation capacity.
Mitotic retention ability is crucial for GAF to promote NSC self-renewal and proliferation
We next sought to investigate whether the mitotic retention ability of GAF underlies its function in promoting NSC self-renewal and proliferation. We first assessed the distribution pattern of various truncated versions of GAF in NSCs. Despite the fact that the polyQ domains have been found to be important for protein oligomerization, including GAF60, GAF.ΔpolyQ forms discrete foci and decorates the mitotic chromosomes of NSCs as full-length GAF (GAF.FL) (Fig. 2a–d, Supplementary Fig. 3a, b). In sharp contrast, GAF.ΔBTB and GAF.ΔZF exhibit weak or no mitotic retention in NSCs (Fig. 2a–d, Supplementary Fig. 3a, b). Importantly, the expression levels of variant GAF transgenes in NSC lineages are comparable (Supplementary Fig. 3c). Therefore, both the BTB domain and the DNA-binding ability of GAF are important for its mitotic retention in vivo.
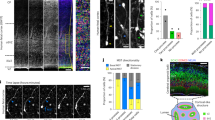
a A schematic summary of the mitotic retention and MARCM rescue ability of GAF variants. b–d FLAG-GAF variants were transiently expressed in NSCs and stained for FLAG and DAPI. Chromosome regions decorated with GAF foci and uniform distribution of GAF are indicated by hollow and solid arrowheads, respectively (b). Quantification of mitotic retention index of GAF-GFP derivatives at H3K9me3- (c) and H3K9me3+ (d) regions of NSCs. Scale bar, 5 μm. e, f OptoDroplet assay of indicated protein fragments in HEK293T cells. A schematic summary of the optoDroplet formation ability of GAF variants is shown in (e). Representative images are shown in (f). Scale bar, 5 μm. g–i Expression pattern of GAF-GFP variants in interphase NSCs (g). GAF condensates are indicated by white arrowheads. Quantification of the percentage of NSCs with condensates is shown in (h). Quantification of the relative fluorescent intensity (F.I.) of GAF-GFP variants is shown in (i). Scale bar, 5 μm. n = 13, 10, 8. (j) A schematic drawing depicting the binding of GAF condensates at target genome loci. Scale bar, 5 μm. k Quantification of the number of neural progenitors (left) or precursors (right) per MARCM clone of the indicated genotypes. Related images are in Supplementary Fig. 3g. l Quantification of the percentage of cells with condensates of indicated genotypes at different time points in the optoDroplet assay. Nuclear foci with a diameter greater than 0.5 μm are counted. Related images are in Supplementary Fig. 4a. n = 20, 50. m, n N7-GAF.ΔBTB forms larger droplets (arrowheads) than GAF.FL in progenitors and precursors (m). Quantification of the diameter of the largest droplet of indicated genotypes in NSCs (left) and progenitors (right) is shown in (n). n = 5, 19. Scale bar, 5 μm. o, p Quantification of type II NSC number per brain lobe of indicated genotypes. N7-GAF.ΔBTB overexpression is induced in NSC lineages (o; pntP1-GAL4) or neural progenitors (p; 12E09-GAL4). *p = 0.0177, n = 5, 13 (o). *p = 0.0396, n = 5, 9 (p). Related images are in Supplementary Fig. 4h, i. Source data are provided as a Source Data file.
To test whether truncation of the BTB domain attenuates the transcription activity of GAF per se, we performed a luciferase reporter assay in HEK293T cells utilizing the GAL4-UAS system (Supplementary Fig. 3d, e). Full-length or truncated versions of GAF were fused with GAL4, which binds to the promoter carrying GAL4-binding site UAS (upstream activation sequence) and activates luciferase expression (Supplementary Fig. 3d). GAF.ΔBTB exhibited transcription repression activity as potent as GAF.FL or EnRD, a repressor domain from the Engrailed protein serving here as a positive control (Supplementary Fig. 3e). Therefore, bypassing DNA-binding ability, the BTB domain is dispensable for the transcription repression activity of GAF.
To test the idea that the condensate formation ability underlies GAF mitotic retention, we next investigated whether the BTB domain is critical for GAF condensate formation using optoDroplet assay. GAF.ΔZF exhibits a potent phase separation ability, comparable to DDX4N (Fig. 2e, f, a well-characterized domain driving phase separation58. In sharp contrast, upon deletion of the BTB domain, GAF.ΔBTB.ΔZF loses its ability to form liquid droplets upon blue light illumination (Fig. 2e, f), strongly suggesting that the condensate formation ability of GAF is driven by BTB domain. We then asked whether restoring phase separation ability per se can rescue the mitotic retention defects of GAF.ΔBTB. We fused GAF.ΔBTB with 1) the BTB domain of the insulator protein Mod(mgd4) (M.BTB), which drives protein oligomerization but shares limited amino acid sequence similarity with GAF.BTB (Supplementary Fig. 3f), 2) the N7 motif of an evolutionarily conserved homeodomain transcription factor Prospero (Pros.N7) that has been known to drive LLPS, or 3) Pros.N7.5 m, a phase separation-defective version of Pros.N7 due to five amino acid mutagenesis (Fig. 2e, f)61. M.BTB and Pros.N7 but not Pros.N7.5 m fusions are indeed effective in restoring the phase separation ability of GAF (Fig. 2e, f). To further validate that the BTB domain drives GAF condensate formation in NSCs, GFP-tagged GAF.FL, GAF.ΔBTB, and Pros.N7-GAF.ΔBTB with comparable expression levels are expressed in NSCs. While GAF.FL and Pros.N7-GAF.ΔBTB form discrete condensates, the condensate formation ability of GAF.ΔBTB is drastically weaker (Fig. 2g–i). Thus, the BTB domain underlies the condensate formation of GAF in NSCs. Remarkably, both M.BTB-GAF.ΔBTB and Pros.N7-GAF.ΔBTB but not Pros.N7.5m-GAF.ΔBTB display mitotic retention upon their transient expression in NSCs (Fig. 2a–d, Supplementary Fig. 3a, b). Therefore, phase separation drives mitotic retention of GAF in NSCs (Fig. 2h).
To directly assess the functional significance of GAF mitotic retention in NSCs, we carried out MARCM clonal analysis. Upon transient expression within GAFKO MARCM clones, mitotic retention-competent GAF.FL or GAF.ΔpolyQ fully rescues the GAFKO NSC proliferation defects (Fig. 2k, Supplementary Fig. 3g). In comparison, mitotic retention-defective GAF.ΔBTB or GAF.ΔZF fails to restore NSC proliferation-promoting ability (Fig. 2k, Supplementary Fig. 3g). Significantly, fusing M.BTB domain with GAF.ΔBTB completely restored the NSC proliferation-promoting ability of GAF (Fig. 2k, Supplementary Fig. 3g). Given that M.BTB mediates distinct protein-protein interaction in vivo from GAF.BTB and that GAF.BTB failed to functionally substitute M.BTB for the insulator function of Mod(mgd4)62, which is the ability of the BTB domain in driving phase separation, but not in modulating cofactor-binding, that is critical for GAF to promote NSC proliferation. Taken together, condensate formation driven by the BTB domain underlies mitotic retention of GAF, which in turn ensures NSC self-renewal and proliferation.
GAF with excessively potent condensate formation ability induces neural progenitor fate conversion and tumorigenesis
If condensate formation ability is indeed a prerequisite for GAF to remain on mitotic chromosomes and execute its self-renewal and proliferation-promoting function, we reasoned that upregulation of GAF phase separation ability might lead to further enhanced self-renewal ability of neural progenitors than normal and hence progenitor dedifferentiation. Intriguingly, while testing whether fusing Pros.N7 with GAF.ΔBTB (Pros.N7-GAF.ΔBTB) can restore the condensate formation and mitotic retention ability of GAF, we noted that Pros.N7-GAF.ΔBTB.ΔZF exhibits a more potent phase separation ability than GAF.ΔZF in optoDroplet assay (Fig. 2l, Supplementary Fig. 4a). In accordance, Pros.N7 displays a stronger condensate formation ability than GAF.BTB (Supplementary Fig. 4b, c). More importantly, Pros.N7-GAF.ΔBTB condensates are prominently larger than GAF.FL condensates in interphase neural progenitors (Fig. 2m, n), strongly suggesting that replacing the BTB domain of GAF with Pros.N7 significantly enhanced the condensate formation ability of GAF.
We next attempted to examine whether Pros.N7-GAF.ΔBTB has a further enhanced self-renewal-promoting ability than wild-type GAF. Since we noted that overexpression of GAF transgenes above certain threshold results in cell toxicity and NSC apoptosis (Supplementary Fig. 4d–g), GAF transgenes are expressed at relatively low levels in NSC lineages for their functional analysis. Interestingly, while weak expression of GAF.FL in type II NSC lineages does not cause any noticeable NSC phenotypes, expression of Pros.N7-GAF.ΔBTB in NSC lineages at similar levels leads to neural progenitor dedifferentiation and tumorigenesis (Fig. 2o, p, Supplementary Fig. 4h–m). Comparing with the control brain lobe that harbors 8.0 type II NSCs, Pros.N7-GAF.ΔBTB overexpressing brain lobe contains on average 32.1 type II NSCs (Fig. 2o). Therefore, under physiological conditions, the condensate formation ability of GAF needs to be tightly controlled to prevent neural progenitor fate reversion back to NSC-like cells and hence a supernumerary NSC phenotype.
GAF preserves NSC fate through transcriptional inhibition of key differentiation genes
To understand the molecular mechanisms by which GAF promotes NSC self-renewal, we performed transcriptome analysis comparing gene expression profiles in control versus GAF-knockdown NSCs. Interestingly, few self-renewal genes are downregulated in NSCs upon GAF knockdown (Fig. 3a, b, Supplementary Fig. 5a). Instead, a subset of differentiation genes specifically expressed in neural progenitors to ensure their timely differentiation, including ham (hamlet), erm (earmuff), opa (odd-paired) and ey (eyeless), are enriched in GAF-knockdown NSCs (Fig. 3a, b, Supplementary Fig. 5b).
a volcano plot of differentially regulated gene expression from RNA-seq analysis comparing GAF-IR with the control. IR: RNAi. (Wald test, p.adj <0.05, log2[fold-change] > 0.5). Differentially expressed genes related to cell proliferation (left) and differentiation (right) are labeled. b Heat maps showing upregulated and downregulated genes upon GAF knockdown by RNA-seq. Selective genes related to differentiation (upper) or proliferation (bottom) are labeled. c Heat maps showing reduced GAF binding to target genomic loci upon GAF knockdown in NSCs by CUT&Tag. d (GA)n motif enrichment within GAF-specific binding peaks, as reported by MEME. One-sided Fisher’s exact test. e Venn diagram showing the overlap of GAF-specific binding genes and upregulated genes upon GAF knockdown. f GO analysis of overlapping genes identified in (e). g Quantification of type II NSC number per brain lobe of indicated genotypes. Related images are in Supplementary Fig. 5g. h, i Larval brain lobes of indicated genotypes were stained for Dpn (red) and Ase (blue) (h). NSCs are marked by white circles. Some NSCs out of focal plane are not labeled in the image. Quantification of type II NSC number of indicated genotypes is shown in (i).***p = 0.0004, ****p < 0.0001. Scale bar, 50 μm. j Genome browser image of CUT&Tag tracks for GAF at key differentiation gene loci in control (dark green) and GAF-IR (light green) NSCs, embryo (purple30) and wing disc (yellow80). The NSC-specific binding peaks of GAF are marked by gray boxes. Illustrations are created in BioRender. Li, Y. (2025) https://BioRender.com/pcljp15. k, l Quantification of the percentage of type II NSCs with erm.R9D10-GFP reporter expression (k). Expression pattern of erm.R9D10 enhancer-GFP reporter in type II NSC lineages of indicated genotypes (l). n = 11, 19. Scale bar, 10 μm. m, n Expression pattern of control or GACA.m form of erm.R9D10-GFP reporter in type II NSC lineages (m). Quantification of the percentage of NSCs with ectopic erm reporter expression is shown in (n). n = 23, 18. Scale bar, 10 μm. Source data are provided as a Source Data file.
To identify GAF target genes in NSCs, we performed an in vivo CUT&Tag assay, utilizing a GAF antibody generated in this study. To ensure the specificity of the GAF antibody and its valid application in CUT&Tag assay, we carried out a series of control experiments. Firstly, western blotting using brain extracts revealed that our GAF antibody detects a specific band with correct molecular weight in the brain extracts (Supplementary Fig. 5c). Importantly, this band is diminished upon GAF knockdown, strongly demonstrating the specificity of the GAF antibody (Supplementary Fig. 5c). Secondly, the immunostaining signal detected by this GAF antibody was substantially reduced in GAF-knockdown NSCs (Supplementary Fig. 5d). Finally, we carried out anti-GAF CUT&Tag analysis with both control and GAF-IR type II NSCs. The majority of the GAF binding peaks are remarkably reduced in GAF-knockdown NSCs (Fig. 3c). Together, this GAF antibody exhibits great specificity and is applicable for in vivo CUT&Tag assay.
Almost all identified GAF-bound sites from our CUT&Tag assay are located in promoter and enhancer regions and are enriched for the canonical GAF-binding motif GAGAG (Fig. 3d, Supplementary Fig. 5e, f). 231 gene loci, including differentiation genes erm, ham and opa, are found to be specifically bound by GAF in NSCs and whose transcription is upregulated upon GAF knockdown in NSCs (Fig. 3e). Consistently, our Gene Ontology (GO) analysis showed that these genes are remarkably enriched for gene sets associated with cell fate commitment, especially neuronal differentiation (Fig. 3f). Consistent with previous results, ectopic overexpression of Ham, Erm or Opa in type II NSC lineages leads to NSC precocious differentiation and drastic reduction in NSC number (Fig. 3g, Supplementary Fig. 5g). More importantly, erm and ham knockdown effectively mitigates NSC loss or underproliferation phenotypes induced by GAF knockdown (Fig. 3h, i, Supplementary Fig. 5h, i), indicating that these differentiation genes are functionally critical downstream target genes of GAF. In accordance, GAF binds to the promoters and enhancers of these key differentiation genes, and the binding peaks are remarkably reduced upon GAF knockdown (Fig. 3j). Intriguingly, via comparing the binding of GAF in type II NSCs to that of early embryos and wing discs, we identified enhancer regions that are specifically bound by GAF in NSCs, such as the ones at erm, ham and opa gene loci (Fig. 3j). These enhancers likely represent NSC-specific gene regulation mediated by GAF.
To examine the functional significance of these NSC-specific enhancers, we focused on the GAF-bound enhancer in the erm locus. R9D10 is a previously characterized erm enhancer in type II NSC lineages63. Indeed, the expression of an erm.R9D10-GFP reporter generated in this study is inhibited in the type II NSCs but immediately activated in neural progenitors (Fig. 3k), largely resembling the expression pattern of erm64. Interestingly, erm.R9D10-GFP is ectopically expressed in type II NSCs upon GAF knockdown (Fig. 3k, l), supporting the notion that GAF normally binds to this enhancer to suppress erm transcription in type II NSCs. To further validate the functional significance of direct binding of GAF to erm.R9D10 enhancer, we mutagenized the canonical GAF binding motif GAGA in this enhancer to GACA (Fig. 3m). As expected, the resulting mutant erm enhancer GACA.m indeed exhibited ectopic expression in type II NSCs (Fig. 3m, n). Taken together, our results indicate that GAF binds to NSC-specific enhancers of key differentiation genes to inhibit their transcription and hence promote NSC self-renewal and proliferation.
GAF bookmarks key differentiation gene loci in mitotic NSCs
As GAF exhibits mitotic retention ability in NSCs, according to previous studies about mitotic bookmarker, we raised the tantalizing possibility that GAF acts as a mitotic bookmarking factor to label these key differentiation gene loci in dividing NSCs and ensure their timely transcriptional reinhibition. To explore this possibility, we attempted to perform in vivo mitotic CUT&Tag analysis. In order to purify mitotic type II NSCs via FACS (Fluorescence Activated Cell Sorting), type II NSCs were dissociated from brain lobes, mildly fixed, and stained with pH3 (phospho-histone H3) (Fig. 4a). pH3-positive (mitotic) and pH3-negative (interphase) type II NSCs were FACS purified (with 98%-100% purity) before anti-GAF CUT&Tag analysis being carried out (Fig. 4a, Supplementary Fig. 6a, b). Importantly, the key differentiation gene loci such as erm and ham are indeed bound by GAF in mitotic NSCs (Fig. 4b). GAF binding sites were then classified into three clusters: interphase only, mitosis only, and mitotically retained (Fig. 4b, c). We found that 448 gene loci are bound by GAF in both interphase and mitosis, accounting for around 18.7% of the total GAF interphase binding sites (Fig. 4b, c). The remaining 81.3% sites are interphase-only sites, bound by GAF in interphase but not mitosis (Fig. 4b, c). Therefore, GAF indeed bookmarks the key differentiation gene loci in mitotic NSCs to ensure their rapid transcriptional reinhibition upon NSC mitotic exit.
a Experimental workflow of mitotic and interphase NSC sorting in a drug synchronization-free manner followed by CUT&Tag analysis. Created in BioRender. Li, Y. (2025) https://BioRender.com/pcljp15. b Genome browser examples of CUT&Tag for GAF tracks at key differentiation gene loci in interphase and mitotic NSCs. Representative examples of GAF binding peaks (mitotically retained, interphase only, and mitosis only) are marked by gray boxes. c Venn diagram showing GAF interphase only, mitotically retained, and mitosis only peaks. d A schematic drawing depicting the asymmetric division of NSC produces sibling daughter cells with distinct cell fates. e Genome browser images of CUT&Tag for GAF tracks at key differentiation gene loci in NSCs versus progenitors. Decreased NSC-specific binding peaks of GAF in progenitors are marked by gray boxes. f Genome browser images of CUT&Tag tracks for H3K27ac, H3K14ac, and H3K4me at key differentiation gene loci in NSCs versus progenitors. Increased H3K27ac binding peaks at key differentiation gene loci in progenitors in comparison to NSCs are marked by gray boxes. Here pnt and tub gene loci serve as negative controls.
To further confirm the functional significance of GAF mitotic retention per se, we attempted to isolate a GAF variant with defective mitotic retention in mitosis but normal transcriptional regulatory ability in interphase. By generating a series of GAF variants that weaken the phase separation ability of the BTB domain to different degrees, we obtained GAF.L112Q, which mutagenizes leucine at position 112 to glutamine (Supplementary Fig. 6c). Importantly, while GAF.L112Q fails to retain on mitotic chromosomes in mitosis, it exhibits relatively normal condensate formation ability in interphase (Supplementary Fig. 6d–h). Compared with GAF.FL, overexpression of GAF.L112Q exhibits a significantly reduced transcriptional inhibition activity in the in vivo erm.9D10-lacZ reporter assay (Supplementary Fig. 6i, j). Therefore, mitotic retention is crucial for GAF to exert its full transcriptional inhibition of key differentiation genes in NSCs.
Key differentiation gene loci exhibit increased H3K27ac deposition in neural progenitors
We next attempted to understand how GAF inhibits transcription of key differentiation genes in NSCs but allows their transcriptional activation in neural progenitors (Fig. 4d). One possibility is that GAF binding at the NSC-specific enhancers of key differentiation genes is significantly reduced in neural progenitors than it is in NSCs. We therefore performed anti-GAF CUT&Tag analysis with type II NSCs versus neural progenitors. Indeed, GAF binding at the enhancers of erm, ham, and opa loci is substantially reduced upon differentiation of type II NSCs into neural progenitors (Fig. 4e).
If GAF inhibits target gene transcription via modulating histone modification(s) of the NSC-specific enhancers, we reasoned that such histone modification(s) would also exhibit differential deposition at the enhancers in NSCs versus neural progenitors. Therefore, we carried out CUT&Tag analysis with type II NSCs versus neural progenitors to evaluate the levels of histone marks, including H3K27ac, H3K14ac, and H3K4me, on key differentiation genes. Although the deposition of H3K14ac and H3K4me at erm, ham and opa loci is essentially the same in NSCs versus neural progenitors, the levels of H3K27ac at these key differentiation gene loci are substantially higher in neural progenitors than they are in NSCs (Fig. 4f, Supplementary Fig. 7a–d). Furthermore, ATAC-seq analysis with NSCs versus neural progenitors confirmed that the higher levels of H3K27ac deposition at differentiation gene loci in neural progenitors correspond to higher levels of chromatin accessibility (Supplementary Fig. 7e). These results strongly suggested that GAF regulates transcriptional inhibition of key differentiation genes in NSCs by decreasing H3K27ac deposition at these gene loci, likely through recruiting HDAC(s) (histone deacetylases), the eraser(s) of histone acetylation.
GAF inhibits transcription of differentiation genes in NSCs through HDAC1 recruitment
We therefore carried out an in vivo RNAi-based HDAC screen to investigate whether knockdown of any individual HDAC leads to type II NSC underproliferation phenotypes. Interestingly, NSC lineage-specific downregulation of HDAC1 but no other HDACs resulted in drastic shrinkage of type II NSC lineages (Fig. 5a, b, Supplementary Fig. 8a–c), demonstrating that HDAC1 plays an important role in promoting NSC proliferation, in line with previous observations65.
a, b Larval brain lobes of indicated genotypes were stained for Dpn and Ase (a). Quantification of progenitor number per type II NSC lineage is shown in (b). ****p < 0.0001. c, d Volcano plot of differentially expressed genes from RNA-seq analysis comparing HDAC1 knockdown with control (Two-sided Wald test, |log2 fold change | > 0.5, p-value < 0.05) (c). Heat maps displaying upregulated and downregulated genes upon HDAC1 knockdown by RNA-seq are shown in (d). Selective genes related to differentiation (top) or self-renewal (bottom) are labeled. e Genome browser images of CUT&Tag tracks for H3K27ac at key differentiation gene loci in control versus HDAC1 knockdown NSCs. f, g Larval brain lobes of indicated genotypes were stained for Dpn and Ase (f). Quantification of type II NSC number per brain lobe is shown in (g). ****p < 0.0001. Scale bar, 50 μm. h, i Quantification of the percentage of NSCs with ectopic Opa expression (h) or ectopic R9D10-11 erm-GFP reporter expression (i) of indicated genotypes. j Coimmunoprecipitation (CoIP) of HA-GAF and FLAG-HDAC1 in HEK293T cells. Schematic drawings of full-length and truncated versions of HDAC1 are shown in the upper panel. HDAC1.N and HDAC1.C are fused with Maltose-Binding Protein (MBP) to increase the protein stability. k, l Genome browser images of CUT&Tag tracks for HDAC1 and GAF in NSCs (k) or for HDAC1 in control versus GAF-IR NSCs (l) at key differentiation genes loci. Co-occupancy of HDAC1 and GAF (k) or decreased HDAC1 binding peaks upon GAF knockdown (l) at NSC-specific enhancer regions are marked by gray boxes. m Distribution of H3K27ac (upper) and HDAC1 (lower) in dividing NSCs. Scale bar, 5 μm. n Genome browser images of CUT&Tag tracks for H3K27ac and HDAC1 at key differentiation gene loci in interphase versus mitotic NSCs. NSC-specific binding peaks of GAF are marked by gray boxes. Related images are shown in Supplementary Fig. 8. Source data are provided as a Source Data file.
We next attempted to determine whether HDAC1 is recruited by GAF to key differentiation gene loci to erase histone acetylation marks. Firstly, we performed RNA-seq analysis with control versus HDAC1-IR type II NSCs and found that erm, ham and opa are ectopically transcribed in NSCs upon HDAC1 knockdown (Fig. 5c, d). Secondly, we carried out H3K27ac CUT&Tag analysis with control versus HDAC1-knockdown NSCs and noted that H3K27ac deposition at erm, ham and opa loci is indeed substantially increased upon downregulation of HDAC1 (Fig. 5e). Thirdly, we carried out genetic experiments to test whether GAF and HDAC1 functionally synergize with each other in inhibiting transcription of key differentiation genes. As expected, although single knockdown of GAF (with GAF-IR−2) or HDAC1 barely affects type II NSC self-renewal, double knockdown of GAF and HDAC1 leads to drastic reduction in type II NSC number (from 8 in wild type to on average 3.36 in GAF, HDAC1 double knockdown) (Fig. 5f, g). Concordantly, ectopic expression of opa (as indicated by FLAG-Opa-KI) or erm (as indicated by erm.R9D10-11 reporter) in type II NSCs is only detectable upon simultaneous downregulation of GAF and HDAC1 (Fig. 5h, i, Supplementary Fig. 8d–f). Fourthly, we performed coimmunoprecipitation (coIP) analysis in HEK293T cells to test whether GAF and HDAC1 physically interact with each other. HA-tagged GAF is specifically and effectively coimmunoprecipitated by FLAG-tagged full-length or the C-terminus of HDAC1 but only marginally coimmunoprecipitated by the N-terminus of HDAC1 (Fig. 5j, Supplementary Fig. 8g), strongly demonstrating that HDAC1 specifically interacts with GAF through its C-terminal domain. Importantly, our coIP experiments further demonstrated that GAF.ΔBTB interacts with HDAC1 with similar affinity as full-length GAF (Supplementary Fig. 8h), confirming that it is the reduced ability of mitotic retention but not the weakened ability in HDAC1 recruitment that accounts for the compromised capacity of GAF.ΔBTB in promoting NSC self-renewal. Lastly, we carried out CUT&Tag analysis with NSCs transiently expressing HDAC1-V5 to analyze the binding peaks of HDAC1 in NSCs. We indeed noted substantial overlap between GAF and HDAC1 binding peaks at erm and opa loci (Fig. 5k). More importantly, the enrichment of HDAC1 at specific enhancers is remarkably reduced upon GAF knockdown (Fig. 5l), demonstrating that HDAC1 is recruited to the specific enhancers by GAF.
Taken together, GAF inhibits transcription of key differentiation genes in NSCs by specifically recruiting HDAC1 and hence erasing H3K27ac from these gene loci.
Mitotic bookmarking of H3K27ac but not HDAC1 in dividing NSCs
We next attempted to understand why GAF needs to remain on NSC mitotic chromosomes to preserve NSC self-renewal fate memory. Our staining results showed that H3K27ac decorates chromosomes in dividing NSCs (Fig. 5m, Supplementary Fig. 9a), By contrast, immunostaining with a specific HDAC1 antibody revealed that HDAC1 is not mitotically retained at NSC chromosomes (Fig. 5m). In accordance, HDAC1-V5 transgene exhibits uniform distribution in dividing NSCs excluding mitotic chromosomes (Supplementary Fig. 9b). These results strongly suggested the tantalizing possibility that it is the high enrichment of H3K27ac at key differentiation genes in dividing NSCs that necessitates timely GAF-HDAC1-mediated deacetylation of H3K27ac in early G1 to avoid ectopic activation of these differentiation genes in NSCs.
To vigorously test this idea, we next performed anti-H3K27ac CUT&Tag with FACS-purified mitotic versus interphase NSCs. Strikingly, H3K27ac is indeed strongly and specifically enriched at key differentiation gene loci in dividing NSCs (Fig. 5n). Such H3K27ac mitotic bookmarking can foster fast activation of these key differentiation genes in future neural progenitors and ensure their differentiating fate lock-in. Importantly, H3K27ac peaks at these key differentiation gene loci are markedly reduced in interphase NSCs (Fig. 5n), explaining why these key differentiation genes are completely shut down in NSCs. This also explains why mitotic retention of GAF is crucial for preserving NSC self-renewal fate memory. That is, mitotic bookmarking by GAF at key differentiation gene loci can ensure timely HDAC1 recruitment and hence rapid erasure of H3K27ac mark in early G1, avoiding ectopic transcription of these differentiation genes in NSCs. Strongly supporting this notion, anti-HDAC1 CUT&Tag with mitotic versus interphase NSCs confirmed that HDAC1 does not bind key differentiation genes in dividing NSCs but is specifically recruited to these gene loci to erase H3K27ac in interphase (Fig. 5n, Supplementary Fig. 9c).
Interestingly, other than erm and ham, 36 more genes involved in neuronal differentiation or cell fate commitment are also dually bookmarked by GAF and H3K27ac in dividing NSCs, transcriptionally inhibited by GAF in interphase NSCs and transcriptionally activated in neural progenitors (Supplementary Fig. 9d, e), indicating that such dual mitotic bookmarking mechanism provides an effective and powerful tactic to accomplish differential propagation of cell fate memory.
Differential GAF bookmarking is achieved by selective competition of the SWI/SNF complex
We next sought to understand how differential GAF bookmarking in neural stem cells versus progenitors is achieved. One likely possibility is that, after NSC asymmetric cell divisions, the mitotic bookmarker GAF is asymmetrically segregated into two daughter cells, with future NSCs inheriting more GAF than future neural progenitors. However, no noticeable difference in terms of mitotic retention of GFP-GAF-KI or FLAG-GAF transgene on sister chromatids can be observed in dividing type II NSCs in anaphase or telophase (Fig. 6a, Supplementary Fig. 1a). Given that the enrichment of GAF at key differentiation gene loci is significantly higher in NSCs than it is in neural progenitors (Fig. 4e), we then considered the possibility that the lower bookmarking ability of GAF in neural progenitors than it is in NSCs might be due to the presence of a transcription activator in neural progenitors, which displaces GAF from target gene loci and hence derepresses differentiation genes. Strongly supporting this notion, specific knockout of GAF-bound enhancer at erm locus led to neural progenitor dedifferentiation and tumorigenesis66, indicating that this enhancer is bound by a critical transcription activator in neural progenitors (Fig. 6b–d).
a Distribution of GFP-GAF-KI in anaphase (left) and telophase (right) NSCs. Scale bar, 5 μm. b–d Deletion of NSC-specific binding enhancers of GAF in the key differentiation gene, erm, leads to tumorigenesis. The locations of NSC-specific enhancers R9D11 and R9D11S are indicated in (b). Larval brain lobes of indicated genotypes stained for Dpn and Ase is shown in (c). Quantification of type II NSC number per brain lobe of the indicated genotypes is shown in (d). **p = 0.0054, ***p = 0.0002, n = 6, 4, 5. Scale bar, 50 μm. e, f Larval brain lobes of indicated genotypes were stained for Dpn and Ase (e). Overexpression of Osa leads to type II NSC loss. Type II NSCs are indicated by white circles. Quantification of type II NSC number per brain lobe is shown in (f). ****p < 0.0001, n = 4, 5. Scale bar, 50 μm. g, h Volcano plot of differentially expressed genes from RNA-seq analysis comparing Osa-overexpression with control NSCs (two-sided Wald test, | log2 fold change|> 0.8, p-value<0.05) (g). Heat maps displaying upregulated and downregulated genes upon Osa overexpression by RNA-seq are shown in (h). Selective genes related to differentiation (top) or self-renewal (bottom) are labeled. i Genome browser images of CUT&Tag tracks for Osa and GAF in NSCs at key differentiation gene loci upon Osa overexpression. j, k Venn diagram (j) and GO analysis (k) of the overlap of GAF-specific binding and Osa binding peaks from CUT&Tag analysis. l, m Larval brain lobes of indicated genotypes were stained for Dpn and Ase (l). Quantification of type II NSC number per brain lobe of the indicated genotypes is shown in (m). ****p < 0.0001. Scale bar, 50 μm. Source data are provided as a Source Data file.
Intriguingly, previous studies found that ARID1B/Osa, a critical subunit of the SWI/SNF chromatin remodeler complex, binds and transcriptionally activates key differentiation genes including erm, ham and opa in neural progenitors but not NSCs36. Furthermore, ChIP-qPCR experiments indicated that Osa binding sites at these differentiation genes are enriched for GAGA motif36, rendering Osa an ideal candidate for a GAF competitor in neural progenitors. Strongly supporting the idea that Osa acts as a transcriptional activator for key differentiation genes, overexpression of Osa in type II NSC lineages phenocopies GAF-knockdown phenotypes, leading to drastic NSC loss (Fig. 6e, f). Consistently, our RNA-seq analysis revealed marked upregulation of differentiation genes including erm, ham and opa upon Osa overexpression (Fig. 6g, h). These results indicate that ARID1B/Osa is a pivotal subunit of the SWI/SNF complex for ensuring functional complex assembly and determining target gene specificity. Our results further revealed that although Osa does not bind to key differentiation genes in NSCs under physiological conditions, it does so upon overexpression.
Significantly, our CUT&Tag assay further showed that overexpression of Osa in NSCs markedly reduces the binding of GAF at specific enhancers of key differentiation genes (Fig. 6i–k), providing direct evidence supporting the competition between Osa and GAF for target gene binding. Consistently, inactivation of the SWI/SNF complex, as a result of osa knockdown, leads to neural progenitor dedifferentiation and brain tumor formation (Fig. 6l, m). We therefore reasoned that the osa knockdown phenotype is due to the fact that the removal of Osa allows GAF to bind to the enhancers of key differentiation genes in neural progenitors and hence transcriptional inhibition of these genes. If this hypothesis is correct, we would expect that double knockdown of GAF and osa would de-repress the key differentiation genes and thereby rescue osa-knockdown-induced brain tumor phenotypes. Indeed, downregulation of GAF strongly rescues the neural progenitor dedifferentiation phenotypes of osa knockdown to almost wild type (Fig. 6l, m).
If our competitor model is correct, we reasoned that when N7-GAF.ΔBTB, the GAF variant with excessive phase separation ability, is overexpressed in neural progenitors, it is likely to outcompete SWI/SNF for key differentiation gene binding, resulting in transcriptional inhibition of these genes and hence neural progenitor dedifferentiation (Supplementary Fig. 10a). Indeed, specific expression of erm reporter in neural progenitors is almost completely silenced upon N7-GAF.ΔBTB expression (Supplementary Fig. 10b).
In sum, our results unveiled a previously uncharacterized paradigm whereby asymmetrically-dividing NSCs differentially propagate cell fate memory to sibling daughter cells through a dual antagonistic mitotic bookmarking and selective molecular competition mechanism (Fig. 7).
In dividing NSCs, key differentiation genes are dually bookmarked by phase-separated GAF condensates (orange) and H3K27ac mark (lime green), keeping NSCs at a poised state ready for binary cell fates. Upon mitotic exit, GAF rapidly recruits the deacetylase HDAC1 (red) and removes H3K27ac, inhibiting transcription of key differentiation genes (red traffic light). As a result, NSC self-renewal fate memory is successfully propagated to the newly-born NSCs. By contrast, in the newly-born neural progenitors, due to the activation of the competitor SWI/SNF complex (green), GAF is displaced from its binding sites at key differentiation gene loci. As a consequence, high H3K27ac deposition retains key differentiation genes transcriptionally re-activated (green traffic light), propagating the differentiating fate memory to the newly-born progenitors. Created in BioRender. Li, Y. (2025) https://BioRender.com/pcljp15.
Discussion
Since the first discoveries revealing that mitotic chromosomes still harbor accessible regions, an ever-increasing number of TFs, chromatin remodelers, or histone marks have been found to bind to mitotic chromosomes of various cells. Drug-treatment induced cell cycle synchronization combined with mitosis-specific degron system provided compelling evidence that many of these mitotically-retained factors or histone marks act as mitotic bookmarkers to preserve cell fate memory across cellular generations by ensuring rapid transcriptional reactivation of key cell fate genes. However, whether mitotic bookmarking plays such a vital role in organ-specific developmental settings, especially the complex neurodevelopmental process, remains largely unexplored.
Here, we found that the pioneer factor GAF, through condensate formation, is retained on mitotic chromosomes of NSCs and specifically marks key differentiation genes. Importantly, GAF preserves NSC fate memory by promoting rapid transcriptional reinhibition instead of transcriptional reactivation of these key differentiation genes. Therefore, our studies unraveled that, for mitotic bookmarkers to facilitate the re-establishment of cell fate-specific gene expression profiles upon mitotic exit, transcriptional reinhibition is equally important as transcriptional reactivation.
Interestingly, upon being transiently expressed in NSCs, GAF.RB exhibits weaker mitotic retention than GAF.RA (Supplementary Fig. 11), suggesting that the mitotic retention ability of a bookmarking factor can be modulated by RNA splicing.
Immediately after NSC asymmetric cell division, key differentiation genes need to be quickly expressed in neural progenitors to ensure their progenitor fate commitment and avoid their cell fate reversion65. However, in the daughter cells that adopt the NSC fate, the transcription of key differentiation genes needs to be completely shut down. How can continuously dividing NSCs resolve such a dilemma?
Here we found that part of the answers to this conundrum lies in the dual mitotic bookmarking by transcription factor GAF and histone mark H3K27ac at these key differentiation gene loci in dividing NSCs. Utilizing our pipeline for enriching mitotic versus interphase NSCs and performing in vivo CUT&Tag67,68, we found that key differentiation genes in dividing NSCs are bookmarked by GAF, and meanwhile, enrich high H3K27ac peaks. Such dual bookmarking by GAF and H3K27ac keeps NSCs in a poised status, ready to produce two sister daughter cells with distinct cell fates. In newly-born neural progenitors, GAF condensates are displaced by newly-activated SWI/SNF competitor, leaving high H3K27ac peaks at key differentiation genes intact, driving timely neural progenitor fate lock-in to a differentiating status. By contrast, in newly-born NSCs, mitotically-retained GAF condensates quickly recruit HDAC1 to erase H3K27ac mark from key differentiation genes and epigenetically silent these genes, preserving self-renewal fate memory.
It remains unclear, when NSCs entering next round of mitosis, how the erased H3K27ac marks at key differentiation genes are specifically added back by writer HAT(s) (histone acetyltransferases).
Previous studies have revealed that mitotically-retained transcription factors help to preserve transcriptional memory of mammalian cultured cells, such as mouse ESCs and HeLa cells69,70, or Drosophila synchronized early embryonic cells30. However, since most of these cells do not undergo asymmetric cell divisions, it remains enigmatic whether mitotic bookmarking factors are asymmetrically segregated into sibling daughter cells, and if not, how symmetric inheritance of mitotic bookmarkers leads to asymmetric cell fate determination.
Our study unveiled a paradigm whereby cell fate memory can be differentially propagated and preserved in sibling daughter cells through differential bookmarking via a molecular competition mechanism. In this study, although the mitotic bookmarker GAF is symmetrically segregated into NSCs and their progenitor siblings, its occupancy at key differentiation genes is specifically reduced in neural progenitors by the competitor SWI/SNF complex. As a consequence, HDAC1 is not specifically recruited to the key differentiation genes, which hence retain their H3K27ac mark. On top of that, the chromatin remodeler SWI/SNF complex, once being recruited, increases local chromatin accessibility and hence stimulates the transcriptional activation of key differentiation genes, further driving neural progenitor differentiating fate commitment.
It remains obscure why ARID1B/Osa exhibits much stronger target binding affinity in neural progenitors than it is in NSCs, despite its comparable expression levels in two sibling cells. The lack of a suitable Osa antibody for performing CUT&Tag and mass spectrometry analysis limited our further investigation. The molecular mechanisms underlying specific activation of SWI/SNF in neural progenitors warrant future investigation. Since transcriptional activation of differentiation genes in neural progenitors occurs immediately after NSC asymmetric division, we speculated that post-translational modification(s) might play a role in the rapid activation of SWI/SNF in neural progenitors.
To our knowledge, our work provided one of the first examples demonstrating that phase separation ability of a mitotic bookmarker needs to be tightly controlled – excessive or defective condensation of a mitotic bookmarker both results in devastating consequences.
On one hand, GAF requires sufficient condensate formation ability to enable it to retain on mitotic chromosomes, serving as a mitotic bookmarker to label target genes and rapidly inhibit their transcription upon mitotic exit. If its condensate formation ability is defective, GAF will be largely expelled from the mitotic chromosomes, unable to prevent ectopic transcription of key differentiation genes. As a result, NSCs prematurely differentiate and partially lose self-renewal ability. On the other hand, if the condensate formation ability of GAF is excessive, the competitor SWI/SNF complex in neural progenitors presumably fails to outcompete this ultra-potent version of GAF. As a consequence, transcription of key differentiation genes in neural progenitors is inhibited by GAF, eventually causing cell fate revision of neural progenitors back to NSCs and tumorigenesis.
Intriguingly, we noted that GAF condensates exhibit distinct mobility in NSCs at different cell cycle stages (Fig. 1e–g). The liquid-like mobility of GAF condensates in interphase NSCs are likely to better support their function in transcriptional regulation, with cofactors such as HDAC1 being quickly recruited and released from the GAF condensates. By contrast, the increased immobility of GAF condensates on mitotic chromosomes of dividing NSCs allows GAF to remain on highly condensed chromosomes and bookmark key target genes. This observation is in line with previous studies, which revealed that GAF binds chromatin with long residence times in fly hemocytes and embryos29,30. How the biophysical properties of a mitotic bookmarker are delicately orchestrated to facilitate its distinct roles in interphase versus mitosis warrants further investigation.
In sum, our results unveiled that a mitotic bookmarking factor plays an important role in preserving NSC self-renewing fate memory across mitosis, and it does so through rapid transcriptional reinhibition of key differentiation genes, instead of fast transcriptional reactivation of self-renewal genes. More importantly, our results revealed a previously uncharacterized paradigm whereby asymmetrically-dividing NSCs differentially propagate cell fate memory to future NSCs versus progenitors through dual mitotic bookmarking by GAF condensates and H3K27ac as well as a selective molecular competition mechanism.
Given that both BTB and ZF domains are essential for GAF mitotic retention and that the BTB-ZF proteins belong to a large and evolutionarily conserved protein family71, it would be tempting to speculate that other BTB-ZF proteins may serve as mitotic bookmarkers in preserving cell fate memory at other brain regions in neural development, as well as in other tissues and organs. Indeed, BTB-ZF protein Th-POK (T-helper-inducing POZ/Krüppel-like factor), GAF homolog in mammals, has been found to 1) partner with HDAC and associate with mitotic chromosomes in cultured murine fibroblasts72; and 2) be both necessary and sufficient to drive the commitment of thymocytes to the CD8 killer instead of the CD4 helper T-cell lineages73. It is tempting to speculate that, in asymmetrically-dividing stem cells, other TFs, especially BTB-ZF proteins, utilize a similar “dual mitotic bookmarking and selective competition” tactic to achieve differential propagation of cell fate memory to sibling daughter cells.
Our studies revealed that differential bookmarking of GAF in NSCs versus neural progenitors can be achieved via a selective competition mechanism. However, it remains enigmatic why the competitor SWI/SNF complex exhibits much stronger target binding and, hence, GAF-displacement ability in neural progenitors than in NSCs. The molecular mechanisms underlying the selective activation of SWI/SNF in neural progenitors warrant future investigation.
Methods
Fly Genetics
Fly culture and crosses were performed according to standard procedures. Drosophila stocks used in this study include: insc-GAL4; pntP1-GAL4; erm.R9D10-lacZ49; UAS-GAF-IR−1 (VDRC #106433); UAS-GAF-IR−2 (BDSC #67265); UAS-HDAC1-IR (BDSC #34846); UAS-HDAC1-V5 (BDSC #32242); erm.9D11KO (BDSC #94625); erm.9D11SKO (BDSC #94279); UAS-Erm-HA (BDSC# 83308); UAS-ham-IR (BDSC# 32470); UAS-erm-IR (BDSC# 26778); UAS-Dicer2 (VDRC# 60008); UAS-His2A-RFP (BDSC# 56555); brat[k06028]74 and 12E09-GAL4 (BDSC #48510). hsFLP; tubulin-FRT-stop-FRT-GAL4; UAS-GFP (gift from Dr. Daniel St Johnston); Full-length or truncated versions of UAS-FLAG-GAF.PA or UAS-GAF-GFP, UAS-FLAG-Mod.BTB-GAF.ΔBTB, UAS-FLAG-Pros.N7-GAF.ΔBTB, GAFKO, GFP-GAF.PA-KI, FLAG-Opa-KI, erm.9D10-GFP, erm.9D10.GACA.m-GFP, erm.9D10-11-GFP, UAS-Osa-V5, UAS-Opa-Myc and UAS-FLAG-Ham were generated in this study. Flies were not sexed, and a mixed population of male and female larval brains was analyzed.
For in vivo FRAP experiments, as shown in Fig. 1e, Supplementary Fig. 1e and 1f, eggs were collected and kept at 22 °C until being shifted to 29 °C for 12-14 h before larval brain dissection. For in vivo droplet assays, as shown in Fig. 2g, eggs were collected and kept at 18 °C until brain dissection at late 3rd instar. For experiments shown in Figs. 1b, 5m, 6a and Supplementary Fig. 2a, eggs were collected and kept at 25 °C until brain dissection at late 3rd instar. For transient expression of GAF as shown in Figs. 2b, m, 6c, Supplementary Figs. 1a, 3b, 4h, 4i, 4k, 10b, eggs were collected and kept at 22 °C until brain dissection at late 3rd instar. For GAF excessive overexpression experiments, eggs were collected and kept at 22 °C until being shifted to 29 °C for 24-50 h before larval brain dissection. For functional experiments as shown in Figs. 1i, 3h, l, m, 5a, f, 6e, l, Supplementary Figs. 2c, 2e, 2h, 5d, 5g, 5h, 8a, 8c, 8d, 8f, eggs were collected for 4-6 hr at 25 °C, shifted to 29 °C after larval hatching and dissected at late 3rd instar. For MARCM clonal analysis as shown in Fig. 1j, Supplementary Fig. 1l, 3g, eggs were collected for 4-6 h at 25 °C, heat-shocked at 37 °C for 2 h after larval hatching, and kept at 22 °C until dissection at late 3rd instar. For experiments shown in Supplementary Fig. 6d, 6g, 6h, newly-hatched larva were shifted to different temperatures to achieve comparable expression levels.
Cell lines
HEK293T cells were grown in DMEM containing 10% (v/v) fetal bovine serum, 1% non-essential amino acids at 37 °C with 5% CO2.
Molecular biology
CRISPR-Cas9-based GAFKO line was generated as previously described75. A 376-bp GAF genomic DNA sequence including partial 5’UTR, ATG, first exon and a small portion of intron was replaced with attP-loxP-3xP3-RFP-loxP to generate GAFKO line. attP-loxP-3xP3-RFP-loxP DNA fragment was amplified from the pEASY-attP-loxp-3xP3-RFP-loxP plasmid, assembled with GAF 5’ arm and 3’ arm via Gibson Assembly and fully sequenced. Synthesized gRNAs (ATACGCTAGCAACGCTGGGGAGG, AGCGAATACGCTAGCAACGTGG) and (CCACAAGATAGTCCTGTGCGCCG, CTAAAGGTAAGTGCTCGGTTCGG) and pBSK(GAF 5’arm-attP-loxP-3xP3-RFP-loxP-GAF 3’arm) plasmid were injected into Cas9 fly embryos. The 3xP3-RFP+ candidates were further confirmed by genomic DNA PCR.
GAFKO line was used as a host for generating the site-directed GFP-GAF.RA-KI line. To generate RIV(EGFP-GAF.RA, mini-white) reintegration plasmid, 97 bp 5’UTR sequence deleted in GAFKO lines, EGFP cDNA, full-length of GAF-RA cDNA, along with GAF.PA.3’UTR and downstream 506 bp sequence as terminator were assembled together into RIV(MCS; mini-white). The reintegration plasmid was verified by DNA sequencing before germline transformation.
FLAG-Opa-KI line was generated using CRISPR-Cas9-based genome engineering, as previously described64. Briefly, dsDNA (double-stranded DNA) donor was created using Gibson Assembly with 1.5 kb homology arms flanking 3×FLAG coding sequence immediately downstream of the start codon of opa. The gRNA sequences (ATCAAGGTAATTAGCATGGAATC) and (GGATTTTGACTGGCATGTATTT) were cloned into pBSK vector under the U63 promoter. The dsDNA donor plasmid and gRNA plasmid were injected into Cas9 fly embryos. The 3xFLAG+ candidates were further confirmed by genomic DNA PCR.
For transgenic fly lines, full-length, truncated or mutated versions of FLAG-GAF or GAF-GFP were cloned into the pUAST vector for transformation. For erm.R9D10-GFP reporter lines, WT or mutated erm R9D10 enhancer fragments were cloned into the pH-Stinger vector. R9D10 fragment and R9D10.GACA.m fragment with 14 GAGA mutagenized to GACA were synthesized. GAGA motifs predicted to be PntP1 binding sites were not mutated to avoid interfering with putative PntP1 regulation.
For BTB domain swapping assay, DNA fragments encoding the BTB domain of Mod(mdg4) was synthesized. Pros.N7 motif (aa 812-870) was amplified from pcDNA3.1-GFP-Pros-FL61. The replacement DNA fragments were added to the N terminus of GAF.ΔBTB via Gibson assembly.
For luciferase assay, full-length GAL4 was added to the N-terminus of EnRD (Engrailed repression domain), VP16 or variant versions of GAF, before being cloned into the pcDNA3.1 vector.
For coimmunoprecipitation experiments, full-length or truncated versions of GAF or HDAC1 were cloned into the pcDNA3.1 vector respectively and fully sequenced.
Immunohistochemistry
For larval brain immunostaining, larvae were dissected in Schneider’s Insect Medium (Sigma-Aldrich) and proceeded as previously described49,76. Briefly, larval brains were fixed with 4% paraformaldehyde in PEM buffer (100 mM PIPES at pH 6.9, 1 mM EGTA, 1 mM MgCl2) for 21 min at room temperature. Brains were washed several times with PBST buffer (1x PBS plus 0.1% Triton X-100) and were incubated with appropriate primary antibody overnight at 4 °C, labeled with secondary antibodies according to standard procedures, and mounted in Vectashield (Vector Laboratories). Images were obtained on a Leica TCS SP8 AOBS confocal microscope and were processed with LAS AF (Leica) and Adobe Photoshop CS5.
Primary antibodies used for immunohistochemistry were rabbit anti-GAF (1:400), rabbit anti-Ase (1:400, gift from Dr. Y.N. Jan), rat anti-Dpn (1:100, Abcam, ab195173), rabbit anti-phospho-Histone H3 [1:200, CST (Cell Signaling Technology), 9701], mouse anti-phospho-Histone H3 (1:200, CST, 9706 s), rabbit anti-FLAG (1:200, Abcam, ab205606), mouse anti-FLAG (1:200, Sigma-Aldrich, F1804), rabbit anti-H3K9me3 (1:100, Abcam, ab8898), rabbit anti-H3K27ac (1:100, Abcam, ab4729), rabbit anti-HDAC1 (1:100, Abcam, ab1767), rabbit anti-V5 (1:100, Sigma-Aldrich, V8137), rabbit anti-cleaved Caspase-3 (1:100, CST, 9661) and mouse anti-β-galactosidase (1:100, DSHB, 40-1a). Rabbit anti-GAF antibody [His-tagged fusion of aa 1-312, affinity purified (ABclonal Biotech.), used at 1:400] was generated in this study.
Larval brain lobes were stained for NSC and neural progenitor marker Dpn (Deadpan) and neural progenitor and precursor marker Ase (Asense) to identify type II NSCs (Dpn+ Ase−), neural progenitors (Dpn+ Ase+) and neural precursors (Dpn− Ase+). In images showing larval brain lobes, the white dashed line marks the boundary between the optic lobe and the central brain area, and the white dotted lines encircle each type II NSC lineage. In close-up images, type II NSC lineages are outlined by white dashed lines, whereas NSCs, progenitors, and precursors are marked with brackets, yellow arrowheads, and white arrowheads, respectively. The cell cortex is indicated by membrane-bound CD8-GFP. In a type II NSC lineage, NSC can be identified as Dpn+ Ase- big-sized cell, whereas in a type I NSC lineage, NSC can be identified as Dpn+ Ase+ big-sized cell.
Fluorescence recovery after photobleaching (FRAP) analysis
Third instar larval brains were dissected in explant solution and mounted on a gas-permeable culture dish. FRAP was performed using the Leica SP8 system, with a laser at 488 nm. Laser power for bleaching was attenuated to 40%. For each experiment, 4-6 images were acquired before an area of 3.84 × 3.84 µm was bleached. Images were collected every 10 s for 40 s after photobleaching. Note that some data that were obviously deviated due to changes of focal plane were not included in the statistics.
OptoDroplet assay
The optoDroplet assay was adapted from a previous study58. Plasmids were co-transfected with polyethylenimine (PEI) transfection reagent in 293 T cells. For transfection, 293T cells were plated 24h prior to transduction in a 4-chamber glass-bottom dish (Cellvis). After transfection, the dishes were cultured in 37 °C incubator for 20-24 h. Imaging was performed on Leica SP8 confocal. For global activation and imaging, cells were imaged typically by use of two laser wavelengths. Droplet formation was induced by 488 nm light for Cry2 activation with the varying activation intervals about 10 s, 20 s, 30 s…up to 60 s. mCherry imaging was stimulated with 568 nm light. Cry2 (mCherry-Cry2 alone) and DDX4N (DDX4N-mCherry-Cry2) served as negative and positive controls respectively.
MARCM clonal analysis
Type II NSC MARCM clones were generated as previously described55. Briefly, newly-hatched larvae were heat-shocked at 37 °C for 2 h and further aged at 22 °C for indicated time before dissection.
GAFKO MARCM clones induced at type II DM1-DM4 lineages barely exhibited NSC lineage shrinkage phenotypes, presumably due to heavier maternal loading of GAF mRNA and protein in these NSCs.
Flip-out clonal analysis
Type II NSC lineage flip-out clones were generated as previously described76. Briefly, newly-hatched larvae were heat-shocked at 37 °C for 2 h and further aged at 22 °C for indicated time before larval brain dissection.
Dual-luciferase reporter assay
Luciferase assay was performed as previously described61. Briefly, HEK293T cells were transfected with 0.5 µg experimental plasmids, luciferase reporter construct pGL4.35 (Promega) together with 50 ng Renilla luciferase reporter plasmid (as the internal control) using standard polyethylenimine (PEI) protocol. Luciferase assay was performed at 24 h post-transfection using the Dual Luciferase Assay system (Promega) following the manufacturer’s instructions.
Coimmunoprecipitation
48 h after transfection, HEK293T cells were harvested, washed and resuspended in lysis buffer [50 mM Tris-HCl (pH 8.0); 120 mM NaCl; 5 mM EDTA; 1% NP-40; 10% glycerol; protease inhibitor cocktail (Sigma-Aldrich); 2 mM Na3VO4] and kept on ice for 20 min. Cell extracts were sonicated with Bioruptor Plus (Biosense) at 4 °C, clarified by centrifugation, and proteins immobilized by binding to anti-FLAG M2 (Sigma-Aldrich) affinity gel for 4 h at 4 °C. Beads were washed and proteins recovered directly in SDS-PAGE sample buffer. Rabbit anti-FLAG (1:1000, CST), mouse anti-FLAG (1:1000, CST) and rabbit anti-HA (1:1000, CST) were used for western blot analysis. Unprocessed blots are provided in the Source Data.
In vivo CUT&Tag
Since wild type larval brains contain only 8 type II NSCs per brain lobe, we performed in vivo CUT&Tag and RNA-seq using type II NSCs derived from brat (brain tumor) mutant larvae. brat mutant larval brains contain up to 1000 type II NSCs per brain lobe and hence provides a large and pure pool of NSCs, ideal for omics analysis36,67,68,77.
Cell preparation for CUT&Tag was performed as previously described78,79. Briefly, larval brains were dissected and digested for 1 h at 30 °C with Dissociation solution (450 μL of complete Schneider’s culture medium with 25 μL of collagenase I and 25 μL of papain). After washing with Rinaldini’s solution and complete Schneider’s culture medium, the samples were dissociated into single cells. The CUT&Tag assays were carried out using Hyperactive Universal CUT&Tag Assay Kit for Illumina (TD903, Vazyme). The libraries were sequenced on an Illumina NovaSeq PE150.
For mitotic CUT&Tag, the procedure is modified from our pipeline67,68. Briefly, larval brains were dissected and dissociated into single cells. The cell suspension was mildly fixed with 0.5% fresh paraformaldehyde at RT for 8 min, and quenched with 0.125 M glycine. Cells were permeabilized using 0.025% Tween-20 at RT for 8 min before being incubated with primary antibody buffer [mouse anti-pH3-647 antibody (Abcam, ab196698) 1:400 and DAPI 1:1000 in PBS buffer] for 2 h at 4 °C. After washing with cold PBS once, the cell suspension was filtered with a 30 μm mesh before being analyzed by the Aria SORP based on pH3, DAPI, CD8-RFP immunofluorescence, and cellular sizes. Primary antibodies used for CUT&Tag were rabbit anti-GAF, rabbit anti-H3K27ac (Abcam), rabbit anti-H3K14ac (Abcam), rabbit anti-H3K4me (Abcam), rabbit anti-V5 (Abcam), and rabbit anti-IgG (Abcam) as a negative control. 1 µg antibody was used for every reaction.
RNA-seq
RNA-seq analysis was performed using Discover-sc WTA Kit V2 (Vazyme #N711) according to the manufacturer’s protocol. Briefly, after larval brains being digested and dissociated into single cells, 1000 NSCs were collected by flow cytometry sorting according to cellular sizes. cDNA library with a peak size of 300 bp was constructed using TruePrep DNA Library Prep Kit V2 for Illumina (Vazyme #TD502) according to the manufacturer’s protocol. The libraries were sequenced on an Illumina NovaSeq PE150.
ATAC-seq
ATAC-seq was performed using Hyperactive ATAC-Seq Library Prep Kit for Illumina (Vazyme #TD711) according to the manufacturer’s protocol. Briefly, larval brains are digested and dissociated into single cells that were collected by flow cytometry sorting. After DNA extraction and amplification, DNA length between 200-2000 bp was selected to construct DNA libraries, which were sequenced on an Illumina NovaSeq PE150.
CUT&Tag analysis
Raw FASTQ reads were trimmed using Trim Galore (v0.6.10, with parameters --paired --quality 20 --stringency 3 -j 4) to eliminate sequencing adaptors and low-quality bases. Quality of reads before and after trimming were monitored using FastQC (v0.11.9) and MultiQC (v1.14). Trimmed reads were aligned to Drosophila genome (version: dm6) using Bowtie2 (v2.5.1) --very-sensitive-local mode. Picard tools (SortSam and MarkDuplicates) were used to mark and remove duplicate reads. Reads unmapped or with a mapping quality score <10 were further filtered using Samtools (v1.6). Filtered bam files were transformed into BigWig files for IGV visualization using deeptools bamCoverage (v3.5.1, with parameters --normalizeUsing RPKM --extendReads). Genome browser images shown in Figs. 4b, e, 5k, 6i subtracted IgG or V5 control using deeptools bigwigCompare (v3.5.1, with parameters --operation subtract). GAF peaks shown in Fig. 5k subtracted GAF-IR control using same parameters. Heatmaps were generated with deepTools (v3.5.1).
To identify GAF-specific binding loci (significantly altered upon GAF knockdown), for IgG, GAF, and GAF-IR, each replicate was down-sampled to the same read depth and then merged into a pseudo sample. This process was repeated twice with different random seeds and generated two pseudo samples for IgG, GAF and GAF-IR. GAF peaks were then called based on pseudo samples using MACS2 (v2.2.7.1, narrow-peak mode, q = 0.05), with IgG as controls. The overlap of these two peak sets (using bedtools intersect) was considered as GAF binding peaks. MACS2 bdgdiff (with parameters --cutoff 3.0) was then performed on every pair of pseudo samples to identify GAF binding regions significantly affected by GAF knockdown. The overlap of these two region sets (using bedtools intersect) was considered as loci with decreased GAF binding density. GAF binding peaks that decrease upon GAF knockdown were kept and defined as GAF specific binding peaks (with ENCODE blacklist regions removed).
To identify GAF binding peaks in mitotic versus interphase NSCs, peak calling was performed on each pair of replicate and control with MACS2 (v2.2.7.1, narrow-peak mode, q = 0.05). The overlap of replicate peak sets was kept, with ENCODE blacklist regions removed.
To identify genomic regions with higher H3K27ac levels in mitotic NSCs than in interphase NSCs, H3K27ac peak calling and differential analysis were performed. H3K27ac peaks were called with MACS2 (v2.2.7.1, narrow-peak mode, q = 0.001, no IgG control). The differential analysis was performed on peak sets of the two-stage using the R package MEDIPS (v1.46.0, pValue<0.05, logFC > 0.5, logCPM > 0). Mitotic H3K27ac peaks overlapped with differential regions were kept, with ENCODE blacklist regions removed.
To identify Osa binding loci, peak calling was performed on each pair of replicate and control with MACS2 (v2.2.7.1, narrow-peak mode, q = 0.05). The consensus peak sets were generated with Irreproducibility Discovery Rate (IDR), with ENCODE blacklist regions removed.
Differentially enriched regions of H3K27ac/H3K4me1/H3K14ac in neural stem cells versus progenitors were identified with DiffBind (v3.4.11). Regions with |logFoldChange | > 0.5 and FDR < 0.05 were considered as differentially enriched regions.
All the peak sets were loaded in R (v4.2.1) for further analysis. ChIPseeker (v1.34.1) and clusterProfiler (v4.8.2) were used for peak annotation and GO analysis. To identify enriched motifs, regions of ±250 bp around peak summits were extracted based on MACS2 peak files and used as input for MEME Suite (v5.5.4) STREME and SEA. Algorithm parameters were set by default.
RNA-seq analysis
Raw FASTQ reads were trimmed using Trim Galore (v0.6.10, with parameters --paired --quality 20 --stringency 3 -j 4) to eliminate sequencing adaptors and low-quality bases. Quality of reads before and after trimming were monitored using FastQC (v0.11.9) and MultiQC (v1.14). Trimmed reads were then aligned to Drosophila genome (Version: dm6) using HISAT2 (v2.2.1) and processed with Samtools (v1.6) to generate bam files. FeatureCounts (v2.0.3) were used to generate raw count matrix for ncbiRefSeq genes.
Differentially expressed genes (DEGs) were called using DESeq2 (v1.38.3). Variance stabilizing transformation (VST) of raw count matrix was used for heatmap visualization. GO analysis were performed using clusterProfiler (v4.8.2), org.Dm.eg.db (v3.16.0) and ggplot2 (v3.4.2) in R (v4.2.1).
ATAC-seq analysis
Raw FASTQ reads were trimmed using Trim Galore (v0.6.10, with parameters --paired --quality 20 --stringency 3 -j 4) to eliminate sequencing adaptors and low-quality bases. Quality of reads before and after trimming were monitored using FastQC (v0.11.9) and MultiQC (v1.14). Trimmed reads were aligned to the Drosophila reference genome (version: dm6) using Bowtie2 (v2.5.1)-very-sensitive-local mode. Duplicate reads were marked and removed using Picard tools. Mitochondrial and low-quality reads were filterer out using Samtools (v1.6). Filtered bam files were transformed into BigWig files for IGV visualization using deeptools bamCoverage (v3.5.1, with option --normalizeUsing RPKM --extendReads).
Quantification and statistical analysis
For NSC quantification, embryos of various genotypes were collected for 4-6 h and allowed to develop to the 3rd instar larval stage (96 hours or 120 hours after larval hatching). 10-20 larvae of each genotype were dissected and stained with anti-Dpn and anti-Ase antibodies to distinguish various cell types within NSC lineages. For quantification of the intensity of antibody staining, images were taken with the same confocal settings and the mean fluorescence intensity was measured with NIH Image J or Laica.
Quantification of the mitotic retention index of GAF is modified from a previous method calculating the chromosome enrichment levels16. Briefly, GFP-GAF-KI brains are stained for H3K9me3 and DAPI to mark the constitutive heterochromatin regions and the whole chromosomes respectively. Retention index at euchromatin and heterochromatin is calculated as log2 ratio of the mean fluorescence intensity of GAF in the H3K9me3- region and the H3K9me3+ region of the chromosomes over the mean fluorescence intensity of GAF outside of chromosomes, respectively.
The retention index of positive values indicates enrichment on the chromosomes, whereas the retention index of negative values indicates exclusion from the chromosomes. The mitotic retention index of GAF transgenes is quantified using a similar strategy.
For Go analysis, p-values are derived from a one-sided hypergeometric test with adjustment using the Benjamini-Hochberg (BH) method. The color indicates the ratio of the number of genes enriched in a specific pathway versus the number of all genes as input. Size of circles indicates the number of genes enriched in this pathway. Repetitive terms were deleted, and some terms related to cell fate decision with highly significant p values were selectively shown in all GO analyses.
Unpaired Student’s t-tests and one-way ANOVA were used for statistical analysis. Data are presented as mean values ± SD (standard deviation).
For each non-omics experiment, the exact n number, p value and the statistical test used for data analysis are provided in the Source Data file.
Statistics and reproducibility
Experiments in this study were independently repeated two to four times. Most of the experiments were performed three times as independent biological repeats. For those repeated twice, the results were highly reproducible. The number of biological replicates is indicated in the Source Data file. For statistical analyses of histograms, P-values greater than 0.05 are considered non-significant, P-values less than 0.05 are denoted with *, while those less than 0.01 are denoted with **, those less than 0.001 with ***, and those less than 0.0001 with ****.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data are accessible at the Gene Expression Omnibus (GEO) with accession codes GSE270071, GSE270072 and GSE292192. ChIP-seq data about GAF binding in embryo and wing disc are from reference under GEO accession number GSE180812 and GSE38594 respectively30,80. Source data are provided with this paper.
References
Siletti, K. et al. Transcriptomic diversity of cell types across the adult human brain. Science 382, eadd7046 (2023).
Braun, E. et al. Comprehensive cell atlas of the first-trimester developing human brain. Science 382, eadf1226 (2023).
Holguera, I. & Desplan, C. Neuronal specification in space and time. Science 362, 176–180 (2018).
Koshland, D. & Strunnikov, A. Mitotic chromosome condensation. Annu Rev. Cell Dev. Biol. 12, 305–333 (1996).
Martinez-Balbas, M. A., Dey, A., Rabindran, S. K., Ozato, K. & Wu, C. Displacement of sequence-specific transcription factors from mitotic chromatin. Cell 83, 29–38 (1995).
Palozola, K. C. et al. Mitotic transcription and waves of gene reactivation during mitotic exit. Science 358, 119–122 (2017).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359, eaao6135 (2018).
Michelotti, E. F., Sanford, S. & Levens, D. Marking of active genes on mitotic chromosomes. Nature 388, 895–899 (1997).
Raccaud, M. et al. Mitotic chromosome binding predicts transcription factor properties in interphase. Nat. Commun. 10, 487 (2019).
Palozola, K. C., Lerner, J. & Zaret, K. S. A changing paradigm of transcriptional memory propagation through mitosis. Nat. Rev. Mol. Cell Biol. 20, 55–64 (2019).
Zaidi, S. K. et al. Mitotic bookmarking of genes: a novel dimension to epigenetic control. Nat. Rev. Genet 11, 583–589 (2010).
Kadauke, S. et al. Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell 150, 725–737 (2012).
Caravaca, J. M. et al. Bookmarking by specific and nonspecific binding of FoxA1 pioneer factor to mitotic chromosomes. Genes Dev. 27, 251–260 (2013).
Liu, Y. et al. Widespread mitotic bookmarking by histone marks and transcription factors in pluripotent stem cells. Cell Rep. 19, 1283–1293 (2017).
Teves, S. S. et al. A dynamic mode of mitotic bookmarking by transcription factors. Elife 5, e22280 (2016).
Deluz, C. et al. A role for mitotic bookmarking of SOX2 in pluripotency and differentiation. Genes Dev. 30, 2538–2550 (2016).
Pelham-Webb, B. et al. H3K27ac bookmarking promotes rapid post-mitotic activation of the pluripotent stem cell program without impacting 3D chromatin reorganization. Mol. Cell 81, 1732–1748 e1738 (2021).
Zhu, Z. et al. Mitotic bookmarking by SWI/SNF subunits. Nature 618, 180–187 (2023).
Biggin, M. D. & Tjian, R. Transcription factors that activate the Ultrabithorax promoter in developmentally staged extracts. Cell 53, 699–711 (1988).
Bhat, K. M. et al. The GAGA factor is required in the early Drosophila embryo not only for transcriptional regulation but also for nuclear division. Development 122, 1113–1124 (1996).
Gaskill, M. M., Gibson, T. J., Larson, E. D. & Harrison, M. M. GAF is essential for zygotic genome activation and chromatin accessibility in the early drosophila embryo. Elife 10, e66668 (2021).
Moshe, A. & Kaplan, T. Genome-wide search for zelda-like chromatin signatures identifies GAF as a pioneer factor in early fly development. Epigenetics Chromatin 10, 33 (2017).
Tsukiyama, T., Becker, P. B. & Wu, C. ATP-dependent nucleosome disruption at a heat-shock promoter mediated by binding of GAGA transcription factor. Nature 367, 525–532 (1994).
Farkas, G. et al. The Trithorax-like gene encodes the drosophila GAGA factor. Nature 371, 806–808 (1994).
Fuda, N. J. et al. GAGA factor maintains nucleosome-free regions and has a role in RNA polymerase II recruitment to promoters. PLoS Genet 11, e1005108 (2015).
Gaskill, M. M. et al. Localization of the Drosophila pioneer factor GAF to subnuclear foci is driven by DNA binding and required to silence satellite repeat expression. Dev Cell 58, 1610–1624 (2023).
Raff, J. W., Kellum, R. & Alberts, B. The drosophila GAGA transcription factor is associated with specific regions of heterochromatin throughout the cell cycle. EMBO J. 13, 5977–5983 (1994).
Tang, X. et al. Kinetic principles underlying pioneer function of GAGA transcription factor in live cells. Nat. Struct. Mol. Biol. 29, 665–676 (2022).
Bellec, M. et al. The control of transcriptional memory by stable mitotic bookmarking. Nat. Commun. 13, 1176 (2022).
Ahmad, K., Brahma, S. & Henikoff, S. Epigenetic pioneering by SWI/SNF family remodelers. Mol. Cell 84, 194–201 (2024).
Hargreaves, D. C. Chromatin openness requires continuous SWI/SNF activity. Nat. Genet 53, 263–264 (2021).
Iurlaro, M. et al. Mammalian SWI/SNF continuously restores local accessibility to chromatin. Nat. Genet 53, 279–287 (2021).
Valencia, A. M. et al. Landscape of mSWI/SNF chromatin remodeling complex perturbations in neurodevelopmental disorders. Nat. Genet 55, 1400–1412 (2023).
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet 45, 592–601 (2013).
Eroglu, E. et al. SWI/SNF complex prevents lineage reversion and induces temporal patterning in neural stem cells. Cell 156, 1259–1273 (2014).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855.e1816 (2018).
Yang, P. G. et al. G3BP1 is a tunable switch that triggers phase separation to assemble stress granules. Cell 181, 325 (2020).
Alberti, S. & Hyman, A. A. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat. Rev. Mol. Cell Biol. 22, 196–213 (2021).
Alghoul, E. et al. Compartmentalization of the SUMO/RNF4 pathway by SLX4 drives DNA repair. Mol. Cell 83, 1640–1658 e1649 (2023).
Zenk, F. et al. HP1 drives de novo 3D genome reorganization in early Drosophila embryos. Nature 593, 289–293 (2021).
Homem, C. C., Repic, M. & Knoblich, J. A. Proliferation control in neural stem and progenitor cells. Nat. Rev. Neurosci. 16, 647–659 (2015).
Brand, A. H. & Livesey, F. J. Neural stem cell biology in vertebrates and invertebrates: More alike than different? Neuron 70, 719–729 (2011).
Gaiano, N. & Fishell, G. The role of notch in promoting glial and neural stem cell fates. Annu Rev. Neurosci. 25, 471–490 (2002).
Hansen, D. V., Lui, J. H., Parker, P. R. & Kriegstein, A. R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561 (2010).
Liu, K. et al. The super elongation complex drives neural stem cell fate commitment. Dev. Cell 40, 537–551 e536 (2017).
Cabernard, C. & Doe, C. Q. Live imaging of neuroblast lineages within intact larval brains in drosophila. Cold Spring Harb. Protoc. 2013, 970–977 (2013).
Knoblich, J. A. Asymmetric cell division: recent developments and their implications for tumour biology. Nat. Rev. Mol. Cell Biol. 11, 849–860 (2010).
Yu, F., Kuo, C. T. & Jan, Y. N. Drosophila neuroblast asymmetric cell division: recent advances and implications for stem cell biology. Neuron 51, 13–20 (2006).
Siller, K. H. & Doe, C. Q. Spindle orientation during asymmetric cell division. Nat. Cell Biol. 11, 365–374 (2009).
Chia, W., Somers, W. G. & Wang, H. Drosophila neuroblast asymmetric divisions: cell cycle regulators, asymmetric protein localization, and tumorigenesis. J. Cell Biol. 180, 267–272 (2008).
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999).
Benyajati, C. et al. Multiple isoforms of GAGA factor, a critical component of chromatin structure. Nucleic Acids Res 25, 3345–3353 (1997).
Greenberg, A. J. & Schedl, P. GAGA factor isoforms have distinct but overlapping functions in vivo. Mol. Cell Biol. 21, 8565–8574 (2001).
Shin, Y. et al. spatiotemporal control of intracellular phase transitions using light-activated optodroplets. Cell 168, 159–171 e114 (2017).
Zielke, N. et al. Fly-FUCCI: A versatile tool for studying cell proliferation in complex tissues. Cell Rep. 7, 588–598 (2014).
Wilkins, R. C. & Lis, J. T. DNA distortion and multimerization: novel functions of the glutamine-rich domain of GAGA factor. J. Mol. Biol. 285, 515–525 (1999).
Liu, X. et al. Mitotic implantation of the transcription factor prospero via phase separation drives terminal neuronal differentiation. Dev. Cell 52, 277–293.e278 (2020).
Read, D., Butte, M. J., Dernburg, A. F., Frasch, M. & Kornberg, T. B. Functional studies of the BTB domain in the Drosophila GAGA and Mod(mdg4) proteins. Nucleic Acids Res 28, 3864–3870 (2000).
Pfeiffer, B. D. et al. Tools for neuroanatomy and neurogenetics in drosophila. Proc. Natl Acad. Sci. USA 105, 9715–9720 (2008).
Hamm, D. C. et al. A conserved maternal-specific repressive domain in Zelda revealed by Cas9-mediated mutagenesis in drosophila melanogaster. PLoS Genet 13, e1007120 (2017).
Janssens, D. H. et al. An Hdac1/Rpd3-poised circuit balances continual self-renewal and rapid restriction of developmental potential during asymmetric stem cell division. Dev. Cell 40, 367–380.e367 (2017).
Hildebrandt, K. et al. Enhancer analysis of the drosophila zinc finger transcription factor Earmuff by gene targeting. Hereditas 158, 41 (2021).
Shen, Y. et al. TBP bookmarks and preserves neural stem cell fate memory by orchestrating local chromatin architecture. Mol. Cell 85, 413–429.e410 (2025).
Shen, Y., Liu, J. & Song, Y. Protocol for identifying genomic binding sites of mitotic bookmarkers in Drosophila neural stem cells and cultured mammalian cells. STAR Protoc. 6, 103858 (2025).
Teves, S. S. et al. A stable mode of bookmarking by TBP recruits RNA polymerase II to mitotic chromosomes. Elife 7, e35621 (2018).
Blobel, G. A. et al. A reconfigured pattern of MLL occupancy within mitotic chromatin promotes rapid transcriptional reactivation following mitotic exit. Mol. Cell 36, 970–983 (2009).
Kelly, K. F. & Daniel, J. M. POZ for effect-POZ-ZF transcription factors in cancer and development. Trends Cell Biol. 16, 578–587 (2006).
Omori, M. et al. CD8 T cell-specific downregulation of histone hyperacetylation and gene activation of the IL-4 gene locus by ROG, repressor of GATA. Immunity 19, 281–294 (2003).
He, X. et al. The zinc finger transcription factor Th-POK regulates CD4 versus CD8 T-cell lineage commitment. Nature 433, 826–833 (2005).
Arama, E., Dickman, D., Kimchie, Z., Shearn, A. & Lev, Z. Mutations in the beta-propeller domain of the drosophila brain tumor (brat) protein induce neoplasm in the larval brain. Oncogene 19, 3706–3716 (2000).
Baena-Lopez, L. A., Alexandre, C., Mitchell, A., Pasakarnis, L. & Vincent, J. P. Accelerated homologous recombination and subsequent genome modification in Drosophila. Development 140, 4818–4825 (2013).
Song, Y. & Lu, B. Regulation of cell growth by notch signaling and its differential requirement in normal vs. tumor-forming stem cells in drosophila. Genes Dev. 25, 2644–2658 (2011).
Larson, E. D. et al. Cell-type-specific chromatin occupancy by the pioneer factor Zelda drives key developmental transitions in Drosophila. Nat. Commun. 12, 7153 (2021).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Kaya-Okur, H. S., Janssens, D. H., Henikoff, J. G., Ahmad, K. & Henikoff, S. Efficient low-cost chromatin profiling with CUT&Tag. Nat. Protoc. 15, 3264–3283 (2020).
Oh, H. et al. Genome-wide association of yorkie with chromatin and chromatin-remodeling complexes. Cell Rep. 3, 309–318 (2013).
Acknowledgements
We are grateful to Drs. Chris Q. Doe, Daniel St. Johnston, Yuh-Nung Jan, Zhengfan Jiang, Yulong Li, Liqun Luo, Yi Rao, Gerald M. Rubin, University of Iowa DSHB, VDRC, Bloomington Drosophila Stock Center, and the TRiP at Harvard Medical School and Tsinghua University for fly stocks and reagents. We thank the Flow Cytometry Core at the National Center for Protein Sciences at Peking University, particularly Ms. Huan Yang and Yinghua Guo, for technical help. We thank Ms. Yonghong Song and Qiuju Zhu for technical assistance, and members of the Song lab for discussions and help. This work was supported by the Ministry of Science and Technology of China (2022YFA1303100 to Y.S.), the National Natural Science Foundation of China (31771629 and 32371024 to Y.S.), and the Peking-Tsinghua Center for Life Sciences (to Y.S.). Part of the illustrations were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Y.S. and Z.R. conceived experiments. Z.R., Z.Z., C.Z., W.T., S.Y., L.Z., C.J., T.H. and Z.X. performed fly genetics, imaging, molecular biology, biochemical, and multi-omics experiments. J.L. performed bioinformatics analysis. Y.S. and Z.R. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hao Jiang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, R., Liu, J., Zhang, Z. et al. Dual mitotic bookmarking by GAF and H3K27ac orchestrates differential propagation of cell fate memory in neural development. Nat Commun 16, 7930 (2025). https://doi.org/10.1038/s41467-025-62974-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62974-6