Abstract
The mitochondrial glutamine transporter SLC1A5_var plays a central role in the metabolic reprogramming of cancer cells by facilitating glutamine import into mitochondria for energy production and redox homeostasis. Despite its critical function, the development of effective and selective inhibitors targeting SLC1A5_var has remained a significant challenge. Here, we introduce iMQT_020, a selective allosteric inhibitor identified through structure-based screening. iMQT_020 disrupts the trimeric assembly of SLC1A5_var, causing metabolic crisis in cancer cells and selectively suppressing their growth. Mechanistically, iMQT_020 reduces glutamine anaplerosis and oxidative phosphorylation, resulting in a broad disruption of cancer metabolism. Additionally, iMQT_020 treatment epigenetically upregulates PD-L1 expression, enhancing the efficacy of combination therapies with anti-PD-L1 immune checkpoint inhibitors. These findings highlight the therapeutic potential of targeting SLC1A5_var as a critical metabolic vulnerability in cancer and demonstrate that targeting allosteric interprotomer interactions is a novel and promising therapeutic strategy for cancer treatment.
Similar content being viewed by others
Introduction
Mitochondrial glutaminolysis is frequently elevated in many cancer cells1,2,3 and is driven by SLC1A5_var, which transports glutamine into the mitochondria4. Glutaminolysis fuels cancer metabolism by supplying TCA cycle metabolites along with glutamine-derived metabolites4. The level of SLC1A5_var is notably increased in various cancers and is correlated with decreased overall survival rates and poor prognosis4. SLC1A5_var inhibition has potential clinical relevance for addressing glutamine dependency in tumor cells5, necessitating further research into selective mitochondrial glutamine transporter inhibitors.
Although targeting mitochondrial glutaminolysis in cancer cells has been a key anticancer strategy6,7,8,9, developing effective and safe inhibitors of mitochondrial glutaminolysis has been challenging. Notably, CB-839, a glutaminase inhibitor, has failed in clinical trials10, highlighting the need for better options. Recently, V-9302 and JHU-083, which block SLC1A5 and glutamine metabolism, were proven to be effective in treating cancer, albeit to a limited degree, as their effects were compensated with alternative metabolites11,12. Our discovery of SLC1A5_var as a promising target offers superior effectiveness and selectivity. Genetic inhibition of this transporter reduces mitochondrial glutamine uptake and inhibits tumor growth4. Although it was anticipated that existing glutamine analogs could be utilized to inhibit SLC1A5_var5, the mechanistic nature of these analogs raises multiple complications. However, allosteric inhibitors present promising avenues for developing potent SLC1A5_var inhibitors with increased effectiveness and selectivity.
Here, we present iMQT_020 as the first reported allosteric SLC1A5_var inhibitor. Biochemical and structural data revealed that iMQT_020 binds to the trimer interaction site of SLC1A5_var, preventing nonselective inhibition of different plasma membrane glutamine transporters. As a result, iMQT_020 potently inhibits the growth of various types of cancer cells without affecting normal epithelial cells. Moreover, iMQT_020 not only has global antitumor effects by inducing metabolic crisis through blocking the essential mitochondrial glutamine transport function of SLC1A5_var for cancer metabolism, but also increases the antitumor immune function of T cells in combination with anti-PD-L1 via epigenetic modulation of PD-L1 expression in pancreatic cancer cells. These results suggest that allosteric inhibition of SLC1A5_var is among the most promising therapeutic strategies.
Results
iMQT_020 is an allosteric inhibitor of mitochondrial SLC1A5_var
SLC1A5_var, a mitochondrial glutamine transporter, drives the metabolic reprogramming of cancer cells, and its genetic inhibition suppresses cancer growth4. Therefore, pharmacological targeting of SLC1A5_var could have potent anticancer effects. First, we aimed to determine the structure of SLC1A5_var through biochemical studies. Crosslinking experiments with mitochondrial extracts containing endogenous or exogenous SLC1A5_var or purified eGFP-tagged SLC1A5_var protein revealed the homotrimeric conformation of SLC1A5_var (Supplementary Fig. 1A–C). To further validate this structure, we conducted two-step immunoprecipitation. Interactions among Myc-tagged, His-tagged, and Flag-tagged SLC1A5_var were consistently observed, as evidenced by the presence of SLC1A5_var monomers in the second immunoprecipitation eluate (Supplementary Fig. 1D). Notably, progressive deletions from the C-terminus of SLC1A5_var, especially deletion of the TM3 domain, led to a significant decrease in interprotomer interactions (Supplementary Fig. 1E), suggesting the crucial role of the TM3 domain in maintaining the homotrimeric structure of SLC1A5_var.
We subsequently attempted to determine the homotrimeric structure of SLC1A5_var. Using AlphaFold and RoseTTAFold13,14, we input the amino acid sequence of SLC1A5_var, considering its homotrimeric nature (Fig. 1A). Using MMseqs2, which identifies homologous sequences, we examined homotrimeric models of SLC1A5_var predicted by AlphaFold2 and RoseTTAFold2. These models closely resemble the structure of SLC1A5. However, they revealed interactions between monomers in the TM1, TM2, and TM4 regions, contradicting our biochemical findings (Supplementary Fig. 1E). Therefore, we utilized AlphaFold and RoseTTAFold to predict a reliable monomeric structure and then employed the MZ-Dock server, a program for estimating protein‒protein interactions, to predict the homotrimeric structure15. We assessed the top 20 models and chose a trimeric structure derived from the structure of the monomeric model predicted by RoseTTAFold, which revealed interactions between the HP1 and TM3 domains (Fig. 1A). Initial models with bond clashes were refined with the Desmond molecular dynamics function16. We subsequently investigated the TM3 domain to identify the residues responsible for interprotomer interactions. Visual inspection revealed strong interactions involving F97, I104, and L105 (FIL) (Fig. 1B). Maintaining the trimeric interaction requires one monomer to engage with the two other monomers. This interaction pattern, in which F97 of one monomer interacts with A98, G101, and K102 of another, and vice versa, was consistently observed within the SLC1A5_var trimer (Fig. 1B). Similar interactions occur between I104 of one monomer and L105 of another (Fig. 1B). Cavities accommodating these residues facilitate connections among the monomers. F97, A98, G101, K102, I104, and L105 were identified as conserved residues across species (Supplementary Fig. 1F) that play crucial roles in homotrimeric interactions.
A Discovery scheme for a selective allosteric inhibitor of SLC1A5_var, iMQT_020. B Structural model of the SLC1A5_var homotrimer. C Mitochondrial glutamine uptake measured by tritium-labeled glutamine in isolated mitochondria with 10 μM of 103 compounds. D Chemical structure of iMQT_020. E SLC1A5_var residues critical for monomer-monomer interaction upon iMQT_020 binding; electrical surface gradient shown. F Virtual docking of iMQT_020 to the allosteric site. G, H (G) Microscale thermophoresis analysis and (H) Normalized fluorescence of SLC1A5_var WT or SLC1A5_var FIL/AAA at different concentrations of iMQT_020. I, J Circular dichroism spectra of (I) SLC1A5_var WT and FIL/AAA at different molar ratios and (J) SLC1A5_var WT and FIL/AAA incubated with vehicle or 4 μM iMQT_020 were recorded at 20 °C. All spectra are buffer corrected. K Size exclusion chromatography of eGFP-tagged SLC1A5_var WT and FIL/AAA incubated with vehicle or 10 μM iMQT_020. The fractions from 8 to 13 mL were blotted. L Concentration-dependent mitochondrial glutamine transport measured with tritium-labeled glutamine after treatment with V-9302 (green) and iMQT_020 (red) (n = 3, biological replicates). M Lineweaver-Burk plot for V-9302, GPNA, and iMQT_020 (n = 3, biological replicates). 3H-glutamine was utilized as a substrate for competitive analysis. N, O Inhibition of (N) mitochondrial and (O) cellular amino acid uptake via V-9302 (green) and iMQT_020 (red). 3H-labeled glutamine, asparagine, serine, leucine, proline, and glutamate were incubated with isolated mitochondria or the medium of MIA PaCa-2 cells treated with vehicle, 10 μM V-9302 (Mitochondrial Uptake; p = 0.012; p = 0.015; p = 0.007; p = 0.021; p = 0.499; p = 0.888), (Cellular Uptake; p = 0.003; p = 0.024; p = 0.031; p = 0.020; p = 0.293; p = 0.051) or 10 μM iMQT_020 (Mitochondrial Uptake; p = 0.004; p = 0.041; p = 0.019; p = 0.854; p = 0.029; p = 0.134) (Cellular Uptake; p = 0.247; p = 0.795; p = 0.401; p = 0.503; p = 0.158; p = 0.264) (n = 3, biological replicates). Amino acid uptake was calculated relative to the vehicle. In K–O experiments were repeated independently three times. In K, representative images/data from one experiment among similar three results are shown. In L–O, all data are presented as the means ± SDs from three biological replicates per group. All p values were determined by Student’s unpaired two-tailed t test. Source data are provided as a Source Data file.
We then assessed the biochemical characteristics to validate the accuracy of the MZ-dock homotrimeric model generated by RoseTTAFold. While the protein‒protein interaction between SLC1A5_var WT and the F97A/I104A/L105A mutant (FIL/AAA) was moderately preserved, the interaction between two of the FIL/AAA mutant proteins was notably reduced (Supplementary Fig. 1G). Furthermore, we mutated G101A and K102A (GK/AA) to analyze nearby conserved residues. These interactions were compared with those of the D186A mutant (D/A), a functionally inert SLC1A5_var mutant4, and another control protein (LRKY/AAAA) that contained the L116A, R128A, K129A, and Y132A mutations. The LRKY/AAAA mutants were predicted to interact with other monomers on the basis of the model but were located in the HP1 and TM4 regions. Compared with the D/A and LRKY/AAAA mutants, the FIL/AAA and GK/AA mutants presented significantly weaker protein‒protein interactions, further confirming the importance of FIL residues but not residues in the TM4 region in homotrimer formation (Supplementary Fig. 1H).
Next, we evaluated the importance of trimeric interactions for mitochondrial glutamine uptake. Upon the downregulation of endogenous SLC1A5_var expression via SLC1A5_var specific siRNAs, the overexpression of SLC1A5_var WT, but not FIL/AAA, restored glutamine transport levels (Supplementary Fig. 1I). Moreover, in SLC1A5_var-downregulated MIA PaCa-2 cells, SLC1A5_var WT overexpression increased cell viability, whereas FIL/AAA overexpression did not (Supplementary Fig. 1J). To further confirm the necessity of the trimeric structure for the metabolic function of SLC1A5_var, we examined alterations in oxygen consumption levels. Although SLC1A5_var FIL/AAA overexpression did not affect the basal oxygen consumption rate (OCR) compared with that of the control (Con), the basal OCR was restored by SLC1A5_var WT overexpression (Supplementary Fig. 1K). Since SLC1A5_var mediates glutamine-derived αKG production4, we hypothesized that the disruption of SLC1A5_var function by the FIL/AAA mutation would reduce αKG production. Overexpressing SLC1A5_var WT increased αKG levels compared with those in the Con group, whereas overexpressing FIL/AAA had no significant effect on αKG levels (Supplementary Fig. 1L). Therefore, we concluded that the interprotomer interaction site is crucial for maintaining the homotrimeric structure and essential for mitochondrial glutamine transport activity.
Our next goal was to identify an allosteric inhibitor capable of selectively binding SLC1A5_var. To achieve this goal, we aimed to develop a compound that inhibits glutaminolysis without affecting the substrate-binding site. Our model of SLC1A5_var revealed the presence of an inhibitor-binding cavity at the trimer interaction site (Fig. 1A). Through initial in silico virtual screening, 1.2 million compounds from the ChemBridge database were subjected to grid-based ligand docking. Our focus was on the trimer interaction site near residues F97, I104, and L105, where we identified a binding pocket. Using Maestro (Schrodinger, USA), we prepared the protein structure, and the LigPrep module of Schrodinger software was utilized for chemical compound docking with the application of the OPLS3 force field. Consequently, the top 103 compounds were selected after visual inspection for druggability and binding conformation states. These compounds were subjected to secondary screening via a mitochondrial 3H-labeled glutamine uptake assay in isolated mitochondria to measure SLC1A5_var functional inhibition. Among the compounds, iMQT_020 had the greatest inhibitory effect at a concentration of 10 μM (Fig. 1C). iMQT_020 consists of a benzyl-quinazoline-dione structure with diverse nucleophiles and electrophiles in its side chain (Fig. 1D). In silico binding of iMQT_020 to the SLC1A5_var model yielded an inhibition constant of 19.77 μM and a Gibbs free energy of -6.42 kJ/mol (Fig. 1E and Supplementary Table 1), indicating a stable energy state upon the binding of iMQT_020 to the protein. The quinazoline-dione structure of iMQT_020 engaged in a hydrophobic interaction with F97, one of the three residues crucial for trimer interactions. It also maintained hydrogen bonding with R263 and I87, which formed bonds with the hydroxyl group and the chloride group of the chloro-fluorophenol side chain of iMQT_020, respectively (Fig. 1F). Additionally, other residues, such as V83, M90, V93, and L260, formed hydrophobic bonds with the carbon molecules of the chloro-fluorophenol side chain (Fig. 1F).
Next, we investigated whether iMQT_020 directly binds to the SLC1A5_var protein and disrupts its homotrimer formation. To confirm the direct binding of iMQT_020 to SLC1A5_var, microscale thermophoresis (MST) was performed using purified eGFP-tagged SLC1A5_var WT and FIL/AAA proteins. The fluorescence intensity changes with varying concentrations of iMQT_020 indicated direct binding to SLC1A5_var WT (Fig. 1G, left), whereas no binding was observed with the FIL/AAA mutant (Fig. 1G, right). MST data further revealed a dissociation constant (Kd) of 4.473 μM for iMQT_020 binding to SLC1A5_var WT (Fig. 1H). To investigate the structural changes induced by iMQT_020, circular dichroism (CD) spectroscopy was conducted. CD spectra of SLC1A5_var WT and FIL/AAA proteins at different molar ratios revealed distinct patterns of reduced ellipticity between 190 nm and 270 nm as monomeric FIL/AAA became dominant (Fig. 1I). Treatment with 4 μM iMQT_020 caused a substantial decrease in the CD signal within this wavelength range for SLC1A5_var WT, whereas no spectral changes were observed for FIL/AAA (Fig. 1J). These results confirm that iMQT_020 directly binds to SLC1A5_var WT and induces structural alterations. To validate the on-target engagement of iMQT_020, we used SLC1A5_var-knockdown cell lines overexpressing one of four constructs: control, wild-type SLC1A5_var (WT), a sodium-binding site mutant (SLC1A5_var D/A), or a trimer interaction mutant (SLC1A5_var FIL/AAA). To eliminate the contribution of endogenous SLC1A5_var, cells were first transfected with SLC1A5_var-specific siRNAs, followed by overexpression of the exogenous WT or mutant constructs (Supplementary Fig. 2A). Mitochondria isolated from these cells were analyzed for mitochondrial 3H-glutamine uptake after 24 h of treatment with either vehicle or 10 μM iMQT_020 (Supplementary Fig. 2A). In control siRNA-transfected cells, overexpression of SLC1A5_var WT significantly increased mitochondrial glutamine uptake, whereas D/A and FIL/AAA mutants failed to do so. This WT-specific increase was completely abolished by iMQT_020 treatment. These results demonstrate that the inhibitory effect of iMQT_020 is specifically mediated through SLC1A5_var activity.
To confirm the inhibition of SLC1A5_var trimer formation by iMQT_020, size exclusion chromatography-based FPLC was conducted. eGFP-tagged SLC1A5_var WT and FIL/AAA were used to assess whether iMQT_020 dissociates from the SLC1A5_var trimer (Fig. 1K). eGFP-tagged SLC1A5_var_WT or FIL/AAA protein lysates were passed through a Superdex 200 Increase 10/300 column in the presence or absence of iMQT_020. Without iMQT_020, eGFP-tagged SLC1A5_var_WT exhibited a peak at 11.2 mL (gray), whereas eGFP-tagged SLC1A5_var_FIL/AAA exhibited a peak at 13.4 mL (green). Upon the introduction of iMQT_020, eGFP-tagged SLC1A5_var_WT exhibited two peaks (red), one at 11.4 mL and one at 13.3 mL. In contrast, iMQT_020 induced no change in the fluorescence peaks exhibited by eGFP-tagged SLC1A5_var_FIL/AAA (purple). Fractions collected from 8 mL to 14 mL were analyzed to detect SLC1A5_var. Interestingly, the SLC1A5_var band was detected in early fractions with eGFP-tagged SLC1A5_var_WT, whereas others showed the SLC1A5_var band in the eleventh and twelfth fractions (Fig. 1K). Building on these observations, we next assessed the impact of iMQT_020 on SLC1A5_var interprotomer interactions and its direct binding affinity. The interaction between Myc-tagged and His-tagged SLC1A5_var_WT proteins was progressively weakened by iMQT_020 in a concentration-dependent manner, with an IC50 of 1.594 μM (Supplementary Fig. 2B). In contrast, iMQT_020 did not inhibit interprotomer interactions in SLC1A5 (Supplementary Fig. 2B). To confirm the trimer-disrupting effect and direct target engagement of iMQT_020 in a cellular context, we established a BRET-BiFC system17 that enables real-time monitoring of SLC1A5_var within the mitochondrial membrane. We first transfected cells with constructs encoding N-terminal or C-terminal halves of YFP-tagged SLC1A5_var for split YFP complementation. Co-localization with the mitochondrial marker, Mitotracker, confirmed mitochondrial localization of the tagged proteins (Supplementary Fig. 2C, D). Trimer formation was subsequently evaluated using a Renilla luciferase (RLuc)-tagged SLC1A5_var, which facilitates bioluminescence resonance energy transfer (BRET) to the reconstituted YFP fluorescence. Notably, treatment with iMQT_020 led to a consistent and pronounced reduction in BRET-BiFC ratio, indicating the disruption of SLC1A5_var trimerization within the mitochondrial compartment (Supplementary Fig. 2E). These findings suggest that iMQT_020 acts as a selective allosteric inhibitor of SLC1A5_var, effectively impeding the formation of its trimeric structure.
We then perform a proof-of-concept experiment to validate iMQT_020 using V-9302 as a comparative control. The IC50 for mitochondrial glutamine transport by iMQT_020 was determined to be 6.156 μM (Fig. 1L). In contrast, the IC50 of V-9302 was 16.406 μM, indicating that V-9302 also suppresses SLC1A5_var. To confirm that iMQT_020 acts as a noncompetitive allosteric inhibitor of SLC1A5_var, we generated Lineweaver–Burk plots of iMQT_020, V-9302, and L-γ-glutamyl-p-nitroanilide (GPNA) at 10 μM (Fig. 1M). The results indicated that iMQT_020 exerts its inhibitory effect in a noncompetitive manner, where Vmax decreased but Km was unchanged, suggesting that the compound does not bind the substrate-binding site but rather binds the allosteric site of SLC1A5_var. On the other hand, V-9302 and GPNA were shown to be competitive inhibitors with the substrate glutamine, as indicated by increased Km values and unchanged Vmax values (Fig. 1M). Further investigation was conducted to assess whether iMQT_020 could inhibit other substrates of SLC1A5_var. The rates of inhibition of mitochondrial uptake of asparagine and serine, known substrates of SLC1A5_var4, were 84.01% and 62.14%, respectively. However, the uptake of leucine, proline, and glutamate, which are not recognized substrates of SLC1A5_var, was not significantly inhibited or even increased (Fig. 1N, red). As glutamate transporters such as SLC25A22 support cancer cell survival18,19, the inhibition of the mitochondrial glutamine transporter was hypothesized to be compensated for by the increased uptake of glutamate into the mitochondria. However, iMQT_020 did not affect the level of glutamate transport into mitochondria, indicating that glutamate does not compensate for the inhibition of the mitochondrial glutamine transporter. Unlike iMQT_020, V-9302 inhibited the mitochondrial uptake of leucine, asparagine, and serine (Fig. 1N, green). Furthermore, the cellular uptake of glutamine, asparagine, serine, leucine, proline, and glutamate did not change upon treatment with 10 μM iMQT_020, whereas V-9302 affected the cellular uptake of glutamine, asparagine, serine, and leucine (Fig. 1O). To evaluate the off-target effects of iMQT_020, a transporter activity assay employing a halide-sensitive YFP reporter, which is commonly used for constructing sodium-dependent amino acid transporter assays20, was conducted. For comparison, the activity of V-9302 was also assessed alongside iMQT_020 across various concentrations against a panel of transporters and channels, including SLC1A5, SLC38A1, SLC38A2, SLC6A14, SLC6A19, SLC26A3, SLC26A4, SLC26A6, SLC26A7, SLC26A9, and ANO1 (Supplementary Fig. 2F). While V-9302 significantly inhibited YFP quenching mediated by SLC1A5, SLC38A2, and SLC6A19 (Supplementary Fig. 2F, bottom), iMQT_020 did not affect YFP quenching in any of these transporters or channels (Supplementary Fig. 2F, top), indicating its selectivity for SLC1A5_var. Interestingly, V-9302 was found to increase the activity of the calcium-activated chloride channel ANO1 (Supplementary Fig. 2F, bottom). Further validation of the selectivity of iMQT_020 was performed via a thermal shift assay of 13 transporters and channels (Supplementary Fig. 2G, H). Building upon previous research21, this assay measures changes in the stability of membrane proteins upon ligand binding. iMQT_020 selectively destabilized SLC1A5_var, reducing its melting temperature (Tm) from 26.1 °C to 21.0 °C (Supplementary Fig. 2H). In contrast, V-9302 significantly altered the Tm values of SLC1A5, SLC38A2, SLC6A19, and ANO1. These findings collectively confirm that iMQT_020 functions as a selective allosteric inhibitor of mitochondrial SLC1A5_var.
Mitochondrial SLC1A5_var inhibition impairs glutamine anaplerosis and redox balance in pancreatic cancer cells
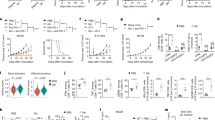
To investigate the role of mitochondrial glutamine transport in cancer metabolic reprogramming, we treated cancer cells with iMQT_020 and assessed its effects on glutamine metabolism via 13C5-labeled glutamine (Fig. 2A). MIA PaCa-2 cells were treated with 10 μM iMQT_020 for 0, 12, or 24 h, followed by analysis of the 13C5-labeled glutamine metabolites via LC‒MS/MS. Treatment with iMQT_020 reduced the levels of glutamine-derived TCA cycle metabolites and glutaminolysis downstream products (Fig. 2B–K). Specifically, the levels of glutamine-derived glutamate and α-ketoglutarate (αKG), key mitochondrial glutaminolysis products, were significantly reduced (Fig. 2B, C). We further investigated the effects of iMQT_020 on glutamine-derived TCA cycle metabolites. Notably, the levels of glutamine-derived succinate (Fig. 2D), fumarate (Fig. 2E), malate (Fig. 2F), oxaloacetate (Fig. 2G), and citrate (Fig. 2H) significantly decreased with iMQT_020 treatment. Additionally, we evaluated downstream metabolites, including those from reductive carboxylation22, such as reduced glutathione (GSH), proline, and palmitate (Fig. 2A). Glutamine-derived glutathione levels (Fig. 2I) and proline levels (Fig. 2J) were significantly reduced in iMQT_020-treated cells. Although most palmitic acid is not derived from glutamine, the levels of glutamine-derived palmitic acid significantly decreased with iMQT_020 treatment (Fig. 2K). These results indicate that inhibiting mitochondrial glutamine transport with iMQT_020 markedly suppresses glutamine anaplerosis, a crucial pathway for cancer metabolism. To further elucidate the effects of iMQT_020, we explored the downstream consequences of inhibiting mitochondrial glutamine transport. Initially, we observed GSH levels (Fig. 2L) after iMQT_020 treatment via a GSH/GSSG assay. The addition of dimethyl-alpha-ketoglutarate (DM-αKG), a cell permeable form of metabolite downstream of glutaminolysis, restored glutathione levels, even in iMQT_020-treated cells. Additionally, in MIA PaCa-2 cells, iMQT_020 treatment increased the levels of cellular reactive oxygen species (ROS), which decreased with DM-αKG supplementation (Fig. 2M). iMQT_020 hindered mitochondrial metabolism, increasing mitochondrial ROS (MitoROS) levels, but mitochondrial metabolism was restored with DM-αKG supplementation (Fig. 2N). Since glutaminolysis fuels oxidative phosphorylation for ATP production, we expected iMQT_020 to lower ATP levels. Indeed, iMQT_020 suppressed ATP production, which was restored by DM-αKG supplementation (Fig. 2O). These results demonstrate how inhibiting mitochondrial glutamine transport compromises energy production and redox balance in cancer cells.
A Diagram of 13C isotopomer patterns with [U-13C] glutamine as a tracer. The incorporation of 13C atoms is denoted as m + n, where n is the number of 13C atoms. B, C Abundance of direct [U-13C] glutamine-derived metabolites in MIA PaCa-2 cells treated with 10μM iMQT_020 for 0, 12, or 24 h, (B) glutamate (p = 0.0077) or (C) αKG (p = 0.0005) total ion counts (n = 3, biological replicates). D–H Abundance of [U-13C] glutamine-derived TCA metabolites in MIA PaCa-2 cells treated with 10 μM iMQT_020 for 0, 12, or 24 h, (D) succinate (p = 0.0454), (E) fumarate (p = 0.0129), (F) malate (p = 0.0018), (G) oxaloacetate (p = 0.0236), or (H) citrate (p = 0.0060) total ion counts (n = 3, biological replicates). I–K Abundance of [U-13C] glutamine-derived metabolites downstream of glutaminolysis in MIA PaCa-2 cells treated with 10 μM iMQT_020 for 0, 12, or 24 h, (I) glutathione (p = 0.0025), (J) proline (p = 0.00007), or (K) palmitate (p = 0.0039) total ion counts (n = 3, biological replicates). L–O Effects of 24 h of 10 μM iMQT_020 and the restorative effects of 2 mM DM-αKG on cellular (L) GSH levels (p = 0.0004, p = 0.0057), (M) cellular ROS levels (p = 0.0008, p = 0.0009), (N) mitochondrial ROS levels (p = 0.0059, p = 0.0032), and (O) ATP levels (p = 0.0299, p = 0.0601) (n = 3, biological replicates). In B–O experiments were repeated independently three times. In B–O all data are presented as the means ± SDs from three biological replicates per group. All p values were determined by Student’s unpaired two-tailed t test; **** p < 0.0001, *** p < 0.001, ** p < 0.01, * p < 0.05 (B-O). Source data are provided as a Source Data file.
We next examined the broader metabolic effects of inhibiting mitochondrial glutamine uptake through iMQT_020 treatment. MIA PaCa-2 cells were treated with 10 μM iMQT_020 for 0, 12, or 24 h, and glucose metabolic flux was analyzed via 13C6-labeled glucose (Supplementary Fig. 3A). Notably, iMQT_020 treatment significantly inhibited glycolysis, as evidenced by reduced levels of 13C6-labeled glucose metabolites, including fructose-1,6-bisphosphate (Supplementary Fig. 3B) and lactate (Supplementary Fig. 3C). Additionally, the production of glucose-derived TCA cycle metabolites, such as citrate, αKG, and succinate, was decreased (Supplementary Fig. 3D–F). Furthermore, iMQT_020 treatment inhibited the pentose phosphate pathway, as shown by a reduction in the 6-phosphogluconate level (Supplementary Fig. 3G). Interestingly, the hexosamine biosynthesis pathway was slightly upregulated. While the level of 13C6-labeled GlcNAc-1-P slightly increased (Supplementary Fig. 3H), the level of 13C6-labeled UDP-GlcNAc significantly increased (Supplementary Fig. 3I), indicating a shift in the metabolic fate of cytosolic glucose. These findings suggest that the inhibition of SLC1A5_var not only disrupts the critical glutaminolysis pathway but also impairs glucose metabolism, highlighting the interconnectedness of these metabolic pathways in cancer cells.
We then explored the metabolic fate of cytosolic glutamine following SLC1A5_var inhibition (Supplementary Fig. 3J). Given the crucial role of glutamine as a nitrogen donor in the cytoplasm9, we performed LC‒MS/MS analysis using 15N2-labeled glutamine to comprehensively elucidate the metabolic pathways by which accumulated cytosolic glutamine is utilized. The nitrogen moiety of glutamine can be incorporated into the purine or pyrimidine pool during de novo nucleotide synthesis. Upon iMQT_020-mediated inhibition of SLC1A5_var, we observed a reduction in the levels of AMP and UMP derived from 15N2-labeled glutamine (Supplementary Fig. 3K, L). In addition, the nitrogen moiety of glutamine can be catabolized to produce asparagine and UDP-GlcNAc. Notably, the relative abundances of 15N-labeled asparagine and 15N-labeled UDP-GlcNAc derived from 15N2-labeled glutamine increased (Supplementary Fig. 3M, N). Interestingly, even the levels of nonglutamine-derived asparagine and UDP-GlcNAc slightly increased compared with those in the 0-h treatment group (Supplementary Fig. 3O, P). These results suggest that under metabolic stress, excess cytosolic glutamine is redirected toward asparagine and hexosamine biosynthesis.
Mitochondrial SLC1A5_var inhibition rewires metabolic reprogramming in pancreatic cancer cells
We assessed the impact of iMQT_020 on cellular metabolic flux by overexpressing SLC1A5_var WT and the FIL/AAA mutant in cells to evaluate the specific metabolic effects of iMQT_020 on its target. After 24 h of treatment with 10 μM iMQT_020, Seahorse analysis was performed. Cells overexpressing SLC1A5_var WT presented a significant increase in the oxygen consumption rate (OCR) (Fig. 3A, B). In contrast, the OCRs of cells overexpressing the FIL/AAA mutant were similar to those of control (Con)-expressing cells. Treatment with iMQT_020 reduced the OCR in SLC1A5_var WT-overexpressing cells to match those of Con cells. These findings indicate that iMQT_020 specifically targets SLC1A5_var and that its metabolic effects are not due to off-target effects. To further confirm this, we investigated the effect of glutamine on the OCR. Glutamine-induced increases in the OCR were inhibited by iMQT_020 in SLC1A5_var WT-overexpressing cells, but cells overexpressing the FIL/AAA mutant maintained low stable OCRs regardless of glutamine supplementation (Fig. 3C, D). In the presence of 2 mM DM-αKG, mitochondrial respiration was maintained irrespective of vehicle or iMQT_020 treatment (Fig. 3E, F). However, iMQT_020 treatment alone induced mitochondrial stress, significantly reducing the OCR in SLC1A5_var WT-overexpressing cells (Fig. 3E, F). These results demonstrate that iMQT_020 directly targets SLC1A5_var, impairing mitochondrial respiration and highlighting the specific action of the compound on SLC1A5_var.
A–P Effects of vehicle or 10 μM iMQT_020 for 24 h on the metabolic phenotype of MIA PaCa-2 cells and human cancer cell lines. A, B Mitochondrial stress OCR in MIA PaCa-2 overexpressing Con, SLC1A5_var WT, or FIL/AAA plotted (A) over time or (B) in a bar graph (p = 0.00005, p = 0.29994, p = 0.00107, p = 0.18525). C, D 4 mM glutamine-induced OCR in MIA PaCa-2 overexpressing SLC1A5_var WT or FIL/AAA plotted (C) over time or (D) in a bar graph (p = 0.00695, p = 0.67442, p = 0.00009, p = 0.62542). E, F With or without 2 mM DM-αKG on OCR in MIA PaCa-2 overexpressing Con, SLC1A5_var WT, and FIL/AAA plotted (E) over time or (F) in a bar graph (p = 0.549, p = 0.009, p = 0.150, p = 0.056, p = 0.283, p = 0.064). G, H 4 mM glutamine-induced ECAR in MIA PaCa-2 overexpressing SLC1A5_var WT or FIL/AAA plotted (G) over time or (H) in a bar graph (p = 0.003, p = 0.941, p = 0.007, p = 0.826). I, J 4.5 g/L glucose-induced OCR in MIA PaCa-2 overexpressing SLC1A5_var WT or FIL/AAA plotted (I) over time or (J) in a bar graph (p = 0.042, p = 0.980, p = 0.004, p = 0.374). K, L 4.5 g/L glucose-induced ECAR in MIA PaCa-2 plotted (K) over time or (L) in a bar graph (p = 0.034, p = 0.179, p = 0.005, p = 0.391). M, N 4 mM glutamine-induced OCR in isolated mitochondria of MIA PaCa-2 transfected with siCon, siSLC1A5_var and Con, siSLC1A5_var and SLC1A5_var WT, and siSLC1A5_var and SLC1A5_var FIL/AAA plotted (M) over time or (N) in a bar graph (p = 0.0004, p = 0.0831, p = 0.0001, p = 0.4132, p = 0.0003, p = 0.0166, p = 0.0001, p = 0.015. O, P Effects of vehicle, 10 μM iMQT_020, or 10 μM iMQT_020 and 2 mM DM-αKG for 24 h on (O) OCRs (p = 0.00005, p = 0.00035) and (P) the metabolic phenotype of 25 cancer cell lines. In A–P experiments were repeated independently three times. In A–P all data are presented as the means ± SDs from three biological replicates per group. All p values were determined by Student’s unpaired two-tailed t test; **** p < 0.0001, *** p < 0.001, ** p < 0.01, * p < 0.05, ns nonsignificant (B, D, F, H, J, L, N, O). Source data are provided as a Source Data file.
To explore the effects of iMQT_020 on the glutamine-dependent glycolytic rate, we assessed its impact on metabolic pathways. Previous studies have demonstrated a strong interconnection between glutamine and glucose metabolism23,24,25. Our earlier research highlighted the critical role of SLC1A5_var in oxidative phosphorylation and glucose metabolism through the regulation of glutamine flux4. The modulation of glycolysis by glutamine flux has been attributed to the malate-aspartate shuttle26. Consistent with this mechanism, iMQT_020 treatment was found to reduce glutamine-derived malate production (Fig. 2F) and glucose-derived lactate production (Supplementary Fig. 3C). These findings were corroborated by measuring the extracellular acidification rate (ECAR) in response to glutamine and iMQT_020. While glutamine supplementation slightly increased the ECAR, iMQT_020 treatment did not increase the ECAR (Fig. 3G, H), suggesting that the inhibition of mitochondrial glutaminolysis by iMQT_020 disrupts glycolytic activity. iMQT_020 also decreased the basal ECAR of SLC1A5_var WT-expressing cells. The effects of iMQT_020 on glucose-dependent metabolic phenotypes were further validated through comprehensive metabolic analysis. Treatment with iMQT_020 for 24 h suppressed glucose-induced increases in both the OCR (Fig. 3I, J) and the ECAR (Fig. 3K, L). iMQT_020 significantly reduced the basal and maximal OCRs in the presence of glucose in SLC1A5_var WT-overexpressing cells (Fig. 3I). However, the OCR of SLC1A5_var FIL/AAA-overexpressing cells was not different from that of vehicle- and iMQT_020 treated cells. Additionally, iMQT_020 partially inhibited the glucose-stimulated increase in the ECAR in SLC1A5_var WT-overexpressing cells (Fig. 3K). In SLC1A5_var FIL/AAA-overexpressing cells, the glucose-stimulated increase in the ECAR was unresponsive via iMQT_020. To confirm the mitochondria-specific effects of iMQT_020, we assayed mitochondria isolated from MIA PaCa-2 cells. Knockdown of SLC1A5_var in mitochondria resulted in a reduction in the glutamine-induced OCR (Fig. 3M, N). The expression of Con and SLC1A5_var FIL/AAA was not able to restore the OCR, and iMQT_020 was ineffective, resulting in a nonsignificant difference in the OCR. However, the restoration of SLC1A5_var WT expression reversed the OCR suppression caused by iMQT_020, confirming that its effects are mediated through the inhibition of SLC1A5_var function in mitochondria. Further analysis of the OCR in cancer cell lines revealed that treatment with iMQT_020 reduced the OCR, a phenotype that was rescued by supplementation with DM-αKG (Fig. 3O). Metabolic profiling of cancer cells under iMQT_020 treatment demonstrated a shift from an energetic to a quiescent metabolic state due to mitochondrial SLC1A5_var inhibition. This metabolic transition was reversible upon DM-αKG supplementation (Fig. 3P). These findings demonstrate that allosteric inhibition of mitochondrial SLC1A5_var by iMQT_020 reprograms cancer cell metabolism, impairing both glycolysis and oxidative phosphorylation.
Given that iMQT_020 disrupts oxidative phosphorylation in cancer cells (Fig. 3A, B), we evaluated its effects on mitochondrial health. Initially, we investigated whether iMQT_020 influences the localization of its target, SLC1A5_var. Immunofluorescence analysis revealed that the colocalization of SLC1A5_var with MitoTracker remained unaffected after treatment with iMQT_020 (Supplementary Fig. 4A, B). Similarly, mitochondrial fractionation analysis revealed that the cofractionation of SLC1A5_var with Tim23, an inner mitochondrial membrane marker, was not affected by iMQT_020 (Supplementary Fig. 4C), indicating that the mitochondrial localization of SLC1A5_var was not disrupted by the treatment. Next, we examined the impact of iMQT_020 on mitochondrial biogenesis and dynamics. MitoTracker staining revealed altered mitochondrial morphology, with fewer fragmented mitochondria observed in iMQT_020-treated cells (Supplementary Fig. 4A, D). This morphological change suggests the induction of mitochondrial oxidative stress and fragmentation. In support of this finding, TMRE staining revealed a significant decrease in the mitochondrial membrane potential in iMQT_020-treated cells (Supplementary Fig. 4E). These findings collectively indicate that iMQT_020 significantly alters mitochondrial dynamics and functionality in cancer cells through the inhibition of mitochondrial glutamine transport activity.
Mitochondrial SLC1A5_var inhibition causes metabolic crisis in pancreatic cancer cells
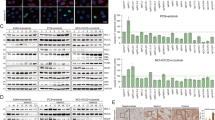
To gain insight into the global impact of iMQT_020 on cellular metabolism in cancer cells, we investigated changes in mRNA expression after 24 h in 10 μM iMQT_020-treated MIA PaCa-2 cells. mRNA-seq analysis revealed significant changes in more than 279 genes. Subsequent KEGG pathway analysis revealed significant downregulation of genes related to metabolism in iMQT_020-treated cells. Given that iMQT_020 profoundly alters cancer cell metabolism, we conducted a detailed examination of metabolism-related genes. Most notably, genes that govern amino acids, carbohydrates, energy metabolism, lipids, and nucleotides were markedly downregulated (Supplementary Fig. 5A). Glutathione metabolism, amino sugar and nucleotide sugar metabolism, fatty acid metabolism, and purine metabolism exhibited notably reduced enrichment scores for each metabolic pathway, with significant p values.
To validate the transcriptomic findings, we compared them with protein expression data obtained from Western blot analysis. Following iMQT_020 treatment, no significant change in SLC1A5_var protein expression was observed (Supplementary Fig. 5B). Consistent with a previous report that the inhibition of SLC1A5_var suppressed mTORC14, we detected a reduction in phospho-S6K levels. Moreover, we observed decreased expression of enzymes involved in amino acid, carbohydrate, lipid, and nucleotide metabolism, including GPX4, IDH2, HK1, PKM1, ACADVL, ACADM, APRT, NME2, PPAT, and PFAS (Supplementary Fig. 5B). In contrast, the levels of UDP-GlcNAc and ASNS were elevated, corroborating both the transcriptomic and metabolic flux data. The increased ASNS expression was hypothesized to result from the integrated stress response (ISR). To investigate this, we analyzed the expression of ATF4 and its downstream targets. The expression levels of ASNS, ATF4, CHOP, SHMT2, and CTH were increased (Supplementary Fig. 5B). Additionally, the KRAS- and PAR4-associated proteins ERK and RhoA were examined to confirm that the effects of iMQT_020 were specifically mediated through the inhibition of SLC1A5_var but not off-target interactions (Supplementary Fig. 5B).
To investigate the systemic impact of iMQT_020 on cancer metabolism, we conducted an unbiased metabolomic analysis comparing MIA PaCa-2 cells treated with iMQT_020 to untreated controls. Using Q Exactive Orbitrap LC‒MS/MS, we identified 615 metabolites, of which 188 exhibited significant changes across 6 Metabolika Superpathways (p < 0.05, Welch’s two-sample t test) (Supplementary Fig. 5C, D). Notable alterations were observed in purine metabolism, glutathione metabolism, and ammonia recycling pathways, which are key processes associated with glutamine metabolism (Supplementary Fig. 5C). Metabolites derived from glutamine, including glutathione, glutamate, αKG, and N-acetyl-glutamate, were significantly reduced, as were ATP- and NAD-related metabolites, upon iMQT_020 treatment (Supplementary Fig. 5C). Given the critical role of glutamine in the nucleotide salvage pathway, we observed a marked decrease in the levels of purine- and pyrimidine-associated metabolites27,28,29 (Supplementary Fig. 5C). Among the lipid metabolites, fatty acid-related metabolites, such as stearic acid, lauric acid, and oleic acid, were significantly reduced (Supplementary Fig. 5C). Interestingly, asparagine and UDP-GlcNAc levels were elevated, suggesting a diversion of cytosolic glutamine toward asparagine or hexosamine biosynthesis. Increased levels of DL-histidine30, hydroxyethyl-methacrylate (HEMA)31, and vitamin C32 were observed, indicating compensatory upregulation of alternative antioxidants in the absence of glutathione. Additionally, elevated D-pantothenic acid levels (Supplementary Fig. 5C) point to an adaptive response involving increased coenzyme A (CoA) production via the Krebs cycle33,34 and increased antioxidant synthesis35. This integrative analysis aligns with biochemical findings, providing deeper insights into the molecular mechanisms underlying the metabolic effects of iMQT_020.
Inhibition of mitochondrial SLC1A5_var leads to cancer cell death in vitro
To investigate the role of SLC1A5_var in cancer cell survival, we performed genetic validation of this target. Analysis of SLC1A5_var mRNA expression revealed low levels in normal epithelial cells of the pancreas, lung, breast, and ovary, as well as in normal astrocyte cells. In contrast, its expression was significantly elevated in most cancer cells (Fig. 4A). Notably, other plasma membrane glutamine transporters, including SLC1A5, SLC38A1, and SLC38A2, did not exhibit consistent differences in expression between normal and cancer cells (Fig. 4A). As glutamine fuels mitochondrial activity in cancer cells, particularly under the metabolic constraints of the Warburg effect, SLC1A5_var plays an essential role in supporting their survival. To test this hypothesis, we downregulated SLC1A5_var via SLC1A5_var-specific siRNAs in both normal and cancer cells derived from the pancreas, colon, and lung. While the viability of normal cells remained unaffected, cancer cell viability was significantly reduced (Supplementary Fig. 6A, B). This result underscores the critical role of SLC1A5_var in maintaining cancer cell survival by supplying TCA cycle metabolites and supporting ATP production.
A Expression of SLC1A5_var, SLC1A5, SLC38A1, SLC38A2, and GAPDH in pancreatic, lung, colon, brain, breast, and ovarian cells measured by RT‒PCR. B A panel of 43 cancer and 7 normal epithelial cell lines were exposed to a single concentration of iMQT_020 or V-9302 (10 μM, 48 h), and the results of WST-1-based viability assay are shown (n = 3, biological replicates). Cells with p < 0.05 according to two-sided Student’s t test compared to the vehicle are shown below the dotted line. C, D SLC1A5_var expression in 50 cell lines were evaluated and correlated with either (C) iMQT_020 sensitivity or (D) V-9302 sensitivity. E Expression of SLC1A5_var, SLC1A5, SLC38A1, SLC38A2, and GAPDH in PDAC patient-derived organoids measured by RT‒PCR. F Effects of exposure to 10 μM V-9302, 10 μM iMQT_020, or 10 μM iMQT_020 with 2 mM DM-αKG for 48 h on the viability of PDAC patient-derived organoids (p = 0.003, p = 0.011, p = 0.0009, p = 0.044) (n = 3, biological replicates). Representative brightfield photomicrographs at 10x (top) and cell viability (bottom) are presented. G SLC1A5_var expression in 10 organoids were evaluated for their correlation with iMQT_020 or V-9302 sensitivity. H Relative viability of MIA PaCa-2 cells treated with vehicle (p = 0.0004) or 10 μM iMQT_020 in the presence of 2 mM DM-αKG (p = 0.0076), 2 mM glutamate (p = 0.0567), 2 mM fructose-1,6-bisphosphate (p = 0.9086), 2 mM 6-phosphogluconate (p = 0.2246), 1 μM disulfiram (p = 0.2688), 10 μM ferrostatin-1 (p = 0.0206), 10 μM necrostatin-1 (p = 0.4754), 10 μM Z-VAD-FMK (p = 0.0045), or a combination of ferrostatin-1 and Z-VAD-FMK (p = 0.0004) for 48 h (n = 3, biological replicates). Significance test was conducted compared with iMQT_020 treated group. In A, B, E, F, H, experiments were repeated independently three times. In A, E, representative images from one experiment among similar three results are shown. In B, F, H, all data are presented as the means ± SDs from three biological replicates per group. All p values were determined by Student’s unpaired two-tailed t test; **** p < 0.0001, *** p < 0.001, ** p < 0.01, * p < 0.05, ns nonsignificant (C, D, and F–H). Source data are provided as a Source Data file.
On the basis of these findings, we hypothesized that iMQT_020 selectively hinders SLC1A5_var, thereby reducing cancer cell viability while sparing normal cells and triggering a metabolic crisis in cancer cells only. To test this hypothesis, we performed an in vitro efficacy assessment by treating 11 pancreatic cell lines and 39 additional different cell lines (6 normal cell lines and 33 cancer cell lines derived from 5 different tissues) with iMQT_020 for 48 h (Fig. 4B). The mutation status of the cancer cells did not significantly influence the efficacy of iMQT_020 (Fig. 4B). Normal cells, including HPDE, BJ, 16HBE, FHC, MCF10A, CHO-K1, and astrocytes, exhibited no significant changes in viability, whereas most cancer cell lines presented a substantial reduction in viability. Notably, V-9302 significantly reduced the viability of normal cells. The IC50 values for iMQT_020 could not be determined in normal cell lines (Supplementary Fig. 6C–H), suggesting minimal toxicity. In contrast, cancer cell lines presented IC50 values ranging from 5 to 40 μM. Specific IC50 values were determined for the pancreatic cancer cell line MIA PaCa-2 (13.15 μM; Supplementary Fig. 6C), the lung cancer cell line NCI-H460 (7.612 μM; Supplementary Fig. 6D), the colon cancer cell line COLO205 (19.18 μM; Supplementary Fig. 6E), the glioblastoma cell line SNB-19 (28.21 μM; Supplementary Fig. 6F), the breast cancer cell line BT-20 (9.993 μM; Supplementary Fig. 6G), and the ovarian cancer cell line OVCAR3 (10.61 μM; Supplementary Fig. 6H). To investigate the variability in sensitivity to iMQT_020, we examined the correlation between its effectiveness and SLC1A5_var expression levels (Fig. 4C). Analysis of 50 cell lines across six cancer types revealed a strong negative correlation between iMQT_020 efficacy and SLC1A5_var expression, with a Pearson correlation coefficient of -0.6881 (Fig. 4C). In contrast, the correlation between V-9302 efficacy and SLC1A5_var expression was much weaker, with a Pearson correlation coefficient of -0.1964 (Fig. 4D). These findings suggest that the expression level of SLC1A5_var is a key determinant of iMQT_020 sensitivity. We next evaluated the effect of iMQT_020 on human pancreatic ductal adenocarcinoma (PDAC) patient-derived organoids with varying levels of SLC1A5_var expression (Fig. 4E). Treatment with iMQT_020 significantly inhibited organoid growth, and DM-αKG supplementation rescued viability, consistent with a glutamine metabolism-dependent mechanism (Fig. 4F). Correlation analysis revealed a strong inverse relationship between iMQT_020 sensitivity and SLC1A5_var expression in PDAC organoids, with a Pearson correlation coefficient of -0.6740 (Fig. 4G).
Next, we investigated the mechanism underlying the anti-proliferative effect of iMQT_020. The observed reduction in GPX4 levels (Supplementary Fig. 5B) and increased in ROS production (Fig. 2M) led us to hypothesize that ferroptosis may be a key contributor to the decreased cancer cell viability. To test this, we silenced SLC1A5_var in MIA PaCa-2 cells and assessed cell viability following treatment with RSL3 (ferroptosis inducer) or ferrostatin-1 (Fer-1, ferroptosis inhibitor) (Supplementary Fig. 6I). Co-treatment with RSL-3 and SLC1A5_var knockdown led to synergistic reductions in viability, while Fer-1 partially rescued it. To directly evaluate lipid peroxidation, a hallmark of ferroptosis, we conducted BODIPY-based lipid oxidation assays. Treatment with iMQT_020 significantly increased oxidized BODIPY levels, indicating enhanced lipid peroxidation (Supplementary Fig. 6J). These findings suggest that iMQT_020 promotes ferroptosis by elevating lipid oxidative stress. Considering the possibility of additional cell death mechanisms, we performed cell cycle analysis and observed significant G1 phase arrest in response to iMQT_020 in both a time- and dose-dependent manner (Supplementary Fig. 6K). Notably, short-term and low-dose exposure primarily induced G1 arrest, whereas prolonged and high-dose treatment led to a marked increase in the Sub-G1 population, indicative of apoptotic cell death. RNA-seq analysis further revealed strong upregulation of pro-apoptotic gene signatures, and downregulation of anti-apoptotic, cell cycle-related, and ferroptosis-regulating gene sets (Supplementary Fig. 6L). To directly assess the involvement of distinct cell death pathways, we measured cell viability following 48-h co-treatment with iMQT_020 and specific inhibitors of ferroptosis (Fer-1), apoptosis (Z-VAD-FMK), pyroptosis (disulfiram), and necroptosis (necrostatin-1), as well as a combination of Fer-1 and Z-VAD (Fig. 4H). Neither Fer-1 nor Z-VAD alone significantly rescued cell viability, but combined treatment markedly restored it (Fig. 4H). Additionally, DM-αKG supplementation restored cell viability, further supporting the mechanism of action of iMQT_020, whereas other control metabolites (glutamate, fructose-1,6-bisphosphate, and 6-phosphogluconate) failed to rescue viability (Fig. 4H). These results suggest that both ferroptosis and apoptosis contribute to iMQT_020-induced cytotoxicity, with apoptosis likely playing a dominant role, as Z-VAD alone elicited a greater rescue effect than Fer-1.
To validate these findings at the protein level, we performed immunoblotting for key regulators of the cell cycle (Cyclin D1, CDK4), anti-apoptotic proteins (Bcl-2, Bcl-xL, Mcl-1), ferroptosis markers (GPX4, FTH, Nrf2), and apoptosis markers (cleaved PARP, cleaved Caspase-3) (Supplementary Fig. 6M). Fer-1 and Z-VAD selectively reversed protein changes associated with ferroptosis and apoptosis, respectively; however, neither treatment fully restored the affected pathways (Supplementary Fig. 6M). Notably, Cyclin D1 expression was rescued only by combined treatment, while CDK4 levels were partially restored by either treatment alone (Supplementary Fig. 6M). Collectively, these results demonstrate that iMQT_020 exerts its cytotoxic effects through coordinated induction of apoptosis and ferroptosis, accompanied by cell cycle arrest.
In contrast to iMQT_020, the in vitro efficacy of V-9302 was limited (Fig. 4B). This discrepancy may stem from the mechanism of V-9302, which nonspecifically targets multiple glutamine transporters and acts as a competitive inhibitor, rendering its efficacy highly dependent on glutamine levels in the culture medium (Supplementary Fig. 7A). Indeed, V-9302 exhibited significantly greater potency under low-glutamine conditions (Cohen’s d = 1.60) compared to high-glutamine conditions. Whereas, the efficacy of iMQT_020 was minimally affected by glutamine concentration (Cohen’s d = 0.48) in pancreatic cancer (Supplementary Fig. 7B). A similar trend was observed in other cancer types, with V-9302 showing enhanced activity in low-glutamine media (Cohen’s d = 1.15), while iMQT_020 remained largely insensitive to glutamine availability (Cohen’s d = 0.68) (Supplementary Fig. 7C). The IC50 values further highlighted the superior selectivity of iMQT_020. In normal HPDE cells, viability remained above 50%, even at 300 μM iMQT_020, whereas the IC50 value for Panc-1 cells was 13.81 μM (Supplementary Fig. 7D). By contrast, V-9302 showed IC50 values of 63.1 μM in HPDE and 29.79 μM Panc-1 cells, underscoring its lower selectivity (Supplementary Fig. 7D), Similarly, in patient-derived pancreatic cancer organoids, iMQT_020 demonstrated greater potency than V-9302, with efficacy showing a weaker and distinct correlation with SLC1A5_var expression (Fig. 4G, Supplementary Fig. 7E, F). Together, these results highlight the superior efficacy and selectivity of iMQT_020, which induces a metabolic crisis in glutamine-dependent cancer cells while sparing normal cells.
Inhibition of mitochondrial SLC1A5_var inhibits tumor growth in vivo
We next evaluated the in vivo efficacy and safety profile of iMQT_020. Pharmacokinetic analysis revealed a mean residence time of 6.9 h (extrapolated to infinity), with the maximum steady-state concentration achieved 0.6 h after intraperitoneal injection and a half-life of 3.8 h (Supplementary Fig. 8A). To assess safety, hERG toxicity and Ames tests were performed. iMQT_020 did not inhibit hERG channels, and Ames tests revealed no mutagenic effects (Supplementary Fig. 8B-D). Chronic exposure to iMQT_020 was assessed by monitoring in vivo toxicity and mouse weight following treatment with vehicle, 75 mg/kg V-9302, or 75 mg/kg iMQT_020 (Supplementary Fig. 8E). No significant weight changes were observed in any treatment group (Supplementary Fig. 8E). To further evaluate systemic toxicity, plasma glucose and glutamine levels were measured in healthy mice treated with vehicle, V-9302, or iMQT_020. Compared with the vehicle, iMQT_020 did not alter plasma glucose or glutamine levels, whereas V-9302 reduced glutamine levels, which is consistent with previous findings12 (Supplementary Fig. 8F, G). H&E staining of lung, liver, and kidney tissues revealed no significant tissue damage from iMQT_020 or V-9302 treatment, which was further supported by nonsignificant changes in plasma AST, ALT, and BUN levels (Supplementary Fig. 8H, I). To investigate potential effects on normal immune cells, the viability and activation of mouse splenocytes and thymocytes were analyzed following treatment with iMQT_020 or V-9302. Neither compound affected cell viability, but both activated immune cells, as confirmed by increased CD44 expression (Supplementary Fig. 8J, K). This finding is consistent with reports suggesting that inhibition of glutamine metabolism enhances T-cell antitumor activity36,37.
Following confirmation of the safety profile of iMQT_020, we next evaluated its in vivo antitumor efficacy. Athymic nude mice bearing MIA PaCa-2 xenografts were treated with 75 mg/kg of either iMQT_020 or V-9302 for 35 days, resulting in significant reductions in tumor size, volume, and weight (Fig. 5A–D). In orthotopic PDAC xenograft models, iMQT_020 treatment also significantly suppressed tumor growth, as evidenced by decreased radiant efficiency (Fig. 5E–G). The antitumor activity of iMQT_020 was further validated in xenograft models of colon cancer (COLO 205) and lung cancer (NCI-H1299), where treatment led to substantial reductions in tumor volume and weight (Fig. 5H–O). Immunohistochemical (IHC) analysis of tumor tissues revealed a marked increase in cleaved caspase-3 and 4-HNE-positive, along with a pronounced decrease in Cyclin D1 and Ki-67-positive cells in the iMQT_020-treated group. These findings indicate enhanced apoptosis, ferroptosis, and cell cycle arrest in MIA PaCa-2, NCI-H1299, and COLO 205 tumors (Fig. 5P–R), consistent with our in vitro observations. These results establish the robust in vivo efficacy of iMQT_020 in multiple cancer models, highlighting its potential as a versatile therapeutic agent.
A–O, in-vivo efficacy of V-9302 and iMQT_020 in human cancer xenograft models. A–D, Athymic nude mice bearing MIA PaCa-2 subcutaneous xenografts (n = 5) were treated daily with vehicle, V-9302, or iMQT_020 (75 mg/kg) for 35 days, starting 7 days after tumor cell injection. Shown are the (A) treatment regimen, (B) representative tumor images, (C) tumor growth curves, and (D) final tumor weights (V-9302, p = 0.00027; iMQT_020, p = 0.00029). E–G Athymic nude mice bearing orthotopic luciferase-expressing MIA PaCa-2 xenografts (n = 5) received the same regimens. Shown are the (E) treatment design, (F) longitudinal bioluminescence quantification, and (G) representative luminescence images. H–K, Athymic nude mice bearing NCI-H1299 lung cancer xenografts (n = 5) were treated daily for 35 days. Shown are the (H) treatment regimen, (I) representative tumor images, (J) tumor growth curves, and (K) final tumor weights (V-9302, p = 0.000017; iMQT_020, p = 0.00018). L–O, Athymic nude mice bearing COLO 205 colon cancer xenografts (n = 5) were treated as above. Shown are the (L) treatment regimen, (M) representative tumor images, (N) tumor growth curves, and (O) final tumor weights (V-9032, p = 0.00067; iMQT_020, p = 0.00019). P–R Immunohistochemical staining for cCas3, 4-HNE, Cyclin D1, and Ki-67 in vehicle-, V-9302-, and iMQT_020-treated xenografts of MIA PaCa-2 (p), NCI-H1299 (q), COLO 205 (r). Representative photomicrographs (top) and quantitative results of cCas3-positive (MIA PaCa-2, p = 0.00077; NCI-H1299, p = 0.00052; COLO 205, p = 0.000068 4-HNE-positive (MIA PaCa-2, p = 0.0099; NCI-H1299, p = 0.0111; COLO 205, p = 0.00829), Cyclin D1-positive (MIA PaCa-2, p = 0.00056; NCI-H1299, p = 0.00006; COLO 205, p = 0.00018), and Ki-67-positive (MIA PaCa-2, p = 0.00002; NCI-H1299, p = 0.00032; COLO 205, p = 0.00005) cells (bottom) are shown (n = 6). For, P–R Box-and-whisker plots show mean (centre), 25th and 75th percentiles (bounds of box), and whiskers extending to the minima and maxima values. All data are presented as the means ± SDs. All p values were determined by Student’s unpaired two-tailed t test; **** p < 0.0001, *** p < 0.001, * p < 0.05 (D, K, O–R). For D, K, and O, p values were calculated compared with the vehicle-treated tumor weight. Source data are provided as a Source Data file.
Inhibition of mitochondrial SLC1A5_var upregulates PD-L1
Pancreatic cancer patients exhibit limited responsiveness to immune checkpoint inhibitors due to the presence of immunosuppressive cells and a low tumor mutational burden; thus, combinatorial therapies, that could enhance the efficacy of immune checkpoint inhibitors, have been developed38,39. Notably, the inhibition of glutaminolysis has been shown to increase PD-L1 expression in cancer cells and enhance the expression of immune checkpoint receptors40,41. Since glutaminolysis produces a critical substrate for histone-modifying enzymes42,43, we hypothesize that SLC1A5_var is involved in glutamine-mediated epigenetic regulation of immune checkpoint receptors. To investigate this hypothesis, we assessed the impact of SLC1A5_var knockdown on PD-L1 expression. Knockdown of SLC1A5_var significantly elevated PD-L1 mRNA levels, an effect that was reversed upon supplementation with DM-αKG or inhibition of KDM5 with CPI-455 (Fig. 6A). Similarly, treatment with iMQT_020, a selective SLC1A5_var inhibitor, increased PD-L1 protein levels in human PDAC cell lines expressing high levels of SLC1A5_var (Fig. 6B). Furthermore, in mouse cancer cell lines (KPC, LLC, and MC-38), treatment with iMQT_020 for 24 h elevated PD-L1 at both the mRNA and protein levels (Fig. 6C). This effect was reversed by the addition of DM-αKG or CPI-455, indicating that the observed PD-L1 induction is linked to glutamine metabolism-dependent epigenetic regulation via H3K4me3 modification. Flow cytometry analysis of KPC cells confirmed increased membrane expression of PD-L1 upon iMQT_020 treatment, which was mitigated by αKG-mediated activation of KDM5 (Fig. 6D). To further elucidate the mechanism underlying PD-L1 induction, chromatin immunoprecipitation (ChIP) assays were performed, which revealed that iMQT_020 treatment significantly increased H3K4me3 levels in the PD-L1 promoter region in KPC cells (Fig. 6E). Concurrently, among the downregulated genes, the oncogene MYC and its target genes exhibited a marked decrease in expression. This reduction was reversed by cotreatment with GSK126, an EZH2 inhibitor (Supplementary Fig. 9A). Additional ChIP assays demonstrated increased H3K27me3 enrichment in the MYC promoter region upon iMQT_020 treatment, which was correlated with suppressed MYC expression. Supplementation with DM-αKG reversed this effect, reducing H3K27me3 enrichment and restoring MYC expression (Supplementary Fig. 9B). These findings illustrate that iMQT_020 modulates glutamine-dependent epigenetic modifications in cancer cells. This modulation leads to increased PD-L1 expression and suppressed MYC expression, highlighting the potential of iMQT_020 to increase the efficacy of immunotherapy by altering the tumor immune and metabolic landscape.
A mRNA expression levels normalized to beta-actin levels in KPC (p = 0.005, p = 0.888, p = 0.013), LLC (p = 0.0009, p = 0.0536, p = 0.012), and MC-38 (p = 0.046, p = 0.872, p = 0.041) mouse cancer cell lines transfected with siCon or siSLC1A5_var supplemented with 2 mM DM-αKG or 10 μM CPI-455 (n = 3, biological replicates). B Relative level of PD-L1 mRNA expression normalized to that of beta-actin and Western blotting of PD-L1 and beta-actin after treatment with vehicle or 10 μM iMQT_020 in SU 86.86. (p = 0.051), SW1990 (p = 0.444), BxPC-3 (p = 0.008), HPAF-II (p = 0.006), Panc-1 (p = 0.009), and MIA PaCa-2 (p = 0.047) pancreatic cancer cells (n = 3, biological replicates). C mRNA expression levels normalized to beta-actin and protein levels of PD-L1 and beta-actin in KPC (p = 0.004, p = 0.042, p = 0.043), LLC (p = 0.011, p = 0.042, p = 0.044), and MC-38 (p = 0.009, p = 0.002, p = 0.004) mouse cancer cell lines treated with vehicle or 10 μM iMQT_020 supplemented with 2 mM DM-αKG or 10 μM CPI-455 for 24 h (n = 3, biological replicates). D (top) Histogram of PD-L1-PE-labeled fluorescence levels of vehicle, 10 μM iMQT_020, or 10 μM iMQT_020 supplemented with 2 mM DM-αKG or 10 μM CPI-455 KPC cells. (bottom) PD-L1-positive KPC cell percentage after treatment with vehicle or 10 μM iMQT_020 supplemented with 2 mM DM-αKG or 10 μM CPI-455 for 24 h. E Chromatin immunoprecipitation assay of the promoter region of PD-L1 after treatment with vehicle, 10 μM iMQT_020, or 10 μM iMQT_020 and 2 mM DM-αKG in the KPC cell line (p = 0.0008, p = 0.0171, p = 0.018, p = 0.033, p = 0.011, p = 0.006) (n = 3, biological replicates). H3K4me3 enrichment was found to be increased upon iMQT_020, which led to increased expression of PD-L1 compared with the input level. In A–C, E, experiments were repeated independently three times. In B, C, representative images from one experiment among similar three results are shown. In A–C, E, all data are presented as the means ± SDs from three biological replicates per group. All p values were determined by Student’s unpaired two-tailed t test; **** p < 0.0001, *** p < 0.001, ** p < 0.01, * p < 0.05, ns nonsignificant (A–C and E). Source data are provided as a Source Data file.
Inhibition of mitochondrial SLC1A5_var synergizes with immune checkpoint inhibitor in vivo
To evaluate the in vivo synergistic effects of combining iMQT_020 with immune checkpoint inhibition, we conducted experiments using subcutaneous xenograft models established with KPC, LLC, and MC-38 mouse cancer cells. The mice were treated with 200 μg of an αPD-L1 antibody every three days, 25 mg/kg iMQT_020 daily, 25 mg/kg V-9302 daily, a combination of αPD-L1 and V-9302, or a combination of αPD-L1 and iMQT_020 (Fig. 7A, D, G). Compared with either monotherapy, combination therapy significantly reduced tumor volume and weight, with superior effects observed in KPC and LLC tumor cells compared with MC-38 tumors (Fig. 7A–I). These results underscore the enhanced therapeutic potential of pairing iMQT_020 with αPD-L1 treatment. IHC staining revealed an increase in cleaved caspase-3-positive cells and a decrease in Ki-67-positive cells within tumor tissues, further confirming the efficacy of the combination therapy in promoting apoptosis and reducing proliferation (Fig. 7J–L). Flow cytometry and IHC analyses revealed increased PD-L1 expression, decreased PD-1+ T cells, and increased infiltration of cytotoxic CD8+ T cells in tumor tissues treated with the combination therapy (Fig. 7M–P, Supplementary Fig. 10A–E). Additionally, tumor digestion analysis revealed elevated populations of cytotoxic CD8+ T cells expressing Granzyme B and IFNγ following iMQT_020 treatment (Fig. 7Q, R). Further evaluation of the tumor microenvironment revealed an increase in CD4+ T cells, a decrease in regulatory T (Treg) cells, and a significant decrease in Ly6G+ and Ly6C+ myeloid-derived suppressor cells (MDSCs) (Fig. 7S–W). These results suggest that iMQT_020 modulates the tumor microenvironment to increase susceptibility to immune-mediated destruction. This pattern of enhanced cytotoxic T cell infiltration and reduced immunosuppressive cells was consistently observed across the LLC and MC-38 xenograft models (Supplementary Fig. 10F–AA). Ki-67 and cleaved caspase-3 staining further highlighting decreased tumor cell proliferation and increased apoptosis following combination therapy (Supplementary Fig. 10F, G). The treatment also resulted in an increase of PD-L1 expression, reduced immunosuppressive cell populations near the tumor tissues, and an increase in cytotoxic T cell infiltration (Supplementary Fig. 10H–AA). These findings demonstrate that targeting SLC1A5_var with iMQT_020 in combination with αPD-L1 therapy effectively reduces tumor growth and promotes cancer cell death. This dual therapeutic approach highlights the potential of targeting glutaminolysis to enhance immunotherapy outcomes, offering a promising metabolic strategy for cancer treatment.
A–I 21-day treatment regimen (with vehicle, αPD-L1 200 μg every 3 days, V-9302 25 mpk daily, or iMQT_020 25 mpk daily) in C57BL/6 N mice bearing (A) KPC, (D) LLC, and (G) MC-38 mouse xenografts (n = 5). (B, D, G) Analysis of tumor volume over the course of all treatments in KPC, LCC, and MC-38 mouse cell xenograft experiments. (C) Final weights of the pancreatic (p = 0.304, p = 0.309, p = 0.426, p = 0.0004, p = 0.0003) (n = 5), lung (p = 0.437, p = 0.967, p = 0.383, p = 0.0001, p = 0.00007) (n = 5), and colon(p = 0.287, p = 0.737, p = 0.496, p = 0.0010, p = 0.0004) (n = 5) xenograft tumors on day 21 after all the treatments. J–L, (J) Immunohistochemistry showing Ki-67 and cCas3 in KPC pancreatic xenografts. The quantitative values of (K) Ki-67- (p = 0.0011) and (L) cCas-3-positive (p = 0.0017) cells per field are shown (n = 6, biological replicates, six independent experiments). M, Percentage analysis of PD-L1-positive cells in KPC cancer tissue. N–W Flow cytometry analysis of the immune population of tumor tissues from all treatment regimens (n = 4). (N) PD-L1-positive-CD45-negative tumor cells were measured (p = 0.012, p = 0.179, p = 0.0001, p = 0.0225, p = 0.0003). T-cell population analysis of (O) PD-1-positive-CD45-positive-CD3-positive (p = 0.0006, p = 0.157, p = 0.0076, p = 0.0001, p = 0.0003) (P) -CD8-positive (p = 0.0001, p = 0.209, p = 0.0024, p = 0.0003, p = 0.0001), and (S) -CD4-positive (p = 0.085, p = 0.915, p = 0.688, p = 0.0076, p = 0.0004) T cells in digested tumor tissues. Quantification of (Q) IFNγ-positive (p = 0.024, p = 0.505, p = 0.035, p = 0.0005, p = 0.0002) and (R) Granzyme B-positive (p = 0.010, p = 0.099. p = 0.267, p = 0.002, p = 0.006) CD8-positive cytotoxic-T-cells. (T) FoxP3-positive-CD25-positive-TCR-β-positive (p = 0.006, p = 0.607, p = 0.927, p = 0.0037, p = 0.0038) regulatory T cells in CD4-positive T cells, (U) -CD11b-positive-Ly6G-positive (p = 0.0001, p = 0.023, p = 0.0003, p = 0.00002, p = 0.000002) and (V) CD11b-positive-Ly6C-positive positive (p = 0.000001, p = 0.624, p = 0.0001, p = 0.00007, p = 0.00004) MDSCs, and (W) CD206-positive (p = 0.0033, p = 0.1301, p = 0.0581, p = 0.0014, p = 0.0015) TAMs are shown. For K, L, N–W, Box-and-whisker plots show mean (centre), 25th and 75th percentiles (bounds of box), and whiskers extending to the minima and maxima values. All data are presented as the means ± SDs. All p values were determined by Student’s unpaired two-tailed t test; **** p < 0.0001, *** p < 0.001, ** p < 0.01, * p < 0.05, ns nonsignificant (C, F, and I–L). For N–W, p values were calculated compared with vehicle-treated cell population. Source data are provided as a Source Data file.
Discussion
Despite extensive research, no FDA-approved inhibitors specifically targeting glutaminolysis are currently available. Challenges such as off-target effects and isoform specificity, particularly for SLC1A5 and glutaminases, have hindered the clinical development of inhibitors targeting glutamine metabolism44,45. The ubiquitous expression of ASCT2/SLC1A5 in normal tissues and the pivotal role of glutaminases in immune function pose additional barriers. This study presents iMQT_020, a first-in-class selective allosteric inhibitor of the mitochondrial glutamine transporter that effectively inhibits glutaminolysis, which restricts glutamine dependency in cancer. Our results underscore the therapeutic potential of targeting SLC1A5_var to disrupt glutamine metabolism in cancer cells, thereby suppressing tumor growth and enhancing antitumor immune responses, particularly when combined with immune checkpoint inhibitors.
Targeting the mitochondrial glutamine transporter has emerged as a critical strategy in cancer therapy5. This is because there are several problems with the original targets and inhibitors. First, among potential targets, SLC1A5_var shows particular promise over plasma membrane glutamine transporters such as SLC1A5, SLC38A1, and SLC38A2 because of the functional redundancy of these plasma membrane transporters46. For example, depletion of SLC1A5 induces compensatory upregulation of SLC38A1 and SLC38A2, indicating that inhibition of a single transporter may be insufficient to fully block glutamine uptake into the cytoplasm47,48,49. While SLC1A5 is overexpressed in various tumors, its ablation suppresses tumor growth only under glutamine-limiting conditions50. Conversely, genetic inhibition of SLC1A5_var effectively impairs cell viability under both glutamine-limiting and standard culture conditions4. Knockdown of SLC1A5_var disrupts amino acid homeostasis by inhibiting glutaminolysis, an effect distinct from that of redundant plasma membrane transporters4. These findings highlight the superiority of SLC1A5_var as a therapeutic target for blocking mitochondrial glutamine metabolism. Second, given the critical role of glutamine in numerous cellular processes, direct inhibition of glutamine-dependent enzymes or pathways may lead to substantial off-target effects, indicating that allosteric inhibition of SLC1A5_var is an exclusive strategy. Targeting the glutamine-binding site poses risks of affecting multiple glutamine-utilizing enzymes, potentially causing toxicity. Moreover, the high abundance of glutamine in serum challenges the efficacy of competitive inhibitors (Supplementary Fig. 7A–C). iMQT_020 overcomes these limitations by selectively and allosterically inhibiting SLC1A5_var, suggesting a noncompetitive approach to targeting mitochondrial glutamine transport (Fig. 1M). Unlike earlier compounds such as GPNA or V-9302, which act on the glutamine-binding site, iMQT_020 disrupts the interprotomer interaction required for SLC1A5_var trimer formation, thereby inhibiting mitochondrial glutamine uptake (Fig. 1K, L). This unique mechanism enhances specificity and minimizes off-target effects (Supplementary Fig. 2D–F), as demonstrated by the selective inhibition of SLC1A5_var without affecting normal cell viability (Fig. 4B).
The loss of cancer cell viability upon iMQT_020 treatment is driven by two key mechanisms: increased reactive oxygen species (ROS) levels and reduced ATP production (Fig. 2M, O). ROS accumulation is linked to reduced glutathione (GSH) synthesis (Fig. 2L), a consequence of impaired glutaminolysis, and decreased NADPH production due to metabolic rerouting of glucose metabolism to the hexosamine biosynthesis pathway (Supplementary Fig. 3B–I). Concurrently, ATP production is impaired through two distinct pathways: depletion of TCA cycle intermediates resulting from glutaminolysis inhibition (Fig. 2B–K) and disruption of the malate–aspartate shuttle, which impedes mitochondrial NADH utilization, leading to a decrease in ATP production from glucose metabolism (Fig. 2O). Therefore, supplementation with glucose-derived metabolites failed to restore cell viability, indicating that the rewiring of glucose metabolism represents an adaptive response to iMQT_020 treatment (Fig. 4H). Collectively, these effects induce profound metabolic stress, leading to G1 phase arrest and a subsequent increase in the sub-G1 population, thereby reducing cancer cell viability primarily through apoptosis, with a contributory role from ferroptosis (Fig. 4H and Supplementary Fig 6I–K). Consistent with our findings, previous studies on glutaminolysis inhibition have reported the activation of multiple cell death pathways, inducing cell cycle arrest51,52, ferroptosis53,54, and apoptosis55,56. Further investigations into the combined use of SLC1A5_var inhibition and ferroptosis inducers may uncover synergistic vulnerabilities in cancer cells and further elucidate the therapeutic potential of targeting glutamine metabolism.
Beyond its direct impact on cancer cell metabolism, iMQT_020 influences the tumor microenvironment by modulating glutamine-dependent epigenetic regulation. Specifically, iMQT_020 treatment enhances PD-L1 expression through the enrichment of H3K4me3 at the PD-L1 promoter region (Fig. 6E). In combination with anti-PD-L1 immune checkpoint inhibitors, iMQT_020 promotes the infiltration of cytotoxic CD8+ T cells (Fig. 7P–R) and decreased the presence of immunosuppressive cell populations, such as tumor-associated macrophages and regulatory T cells (Fig. 7T–W). These effects align with recent studies in renal and lung cancers36,37, highlighting the dual role of iMQT_020 in selectively inducing cancer cell death while amplifying antitumor immune responses. The therapeutic potential of iMQT_020 in combination with immune checkpoint inhibitors is particularly relevant for pancreatic ductal adenocarcinoma (PDAC), a malignancy where no FDA-approved immune checkpoint inhibitors are currently available57. Although immune checkpoint inhibitors have demonstrated efficacy in certain solid tumors58, their success in PDAC remains limited due to low tumor immunogenicity–evidenced by the downregulation of PD-L1 in TCGA datasets59 and the immunosuppressive tumor microenvironment characterized by lactate accumulation and hypoxia60. Despite these challenges, emerging research and novel combination strategies are identifying approaches to overcome resistance to immune checkpoint inhibitors, providing optimism for their use in PDAC treatment61. Several ongoing clinical trials are evaluating immune checkpoint inhibitors in combination with anticancer agents as second-line or later therapies. In this context, the synergistic interaction between iMQT_020 and αPD-L1 in the KPC mouse model demonstrates significant therapeutic benefits, including reductions in tumor weight and size (Fig. 7A–C) and modulation of immune cell populations (Fig. 7O–W). These findings highlight the potential of this combination therapy to overcome resistance to immune checkpoint inhibitors in pancreatic cancer. Collectively, these results support the clinical application of iMQT_020 and αPD-L1 as a promising treatment strategy for aggressive and refractory solid tumors, particularly those characterized by low PD-L1 expression and poor response to conventional therapies.
Despite these promising results, several challenges remain. Cancer cells may develop adaptive resistance to SLC1A5_var inhibition through compensatory pathways, such as the upregulation of alternative amino acid transporters such as SLC25A22. Preliminary data suggesting increased levels of UDP-GlcNAc and asparagine after iMQT_020 treatment suggest potential metabolic adaptations (Supplementary Fig. 2H, I). Further studies are warranted to explore these mechanisms and evaluate the long-term effects of iMQT_020 in advanced in vivo models.
In conclusion, iMQT_020 represents a promising therapeutic agent with high specificity for SLC1A5_var, suggesting a novel approach for targeting glutamine metabolism in cancer. Its ability to synergize with immune checkpoint blockade highlights its potential as part of a combination therapy for resistant cancers. Future research should focus on understanding resistance mechanisms, optimizing combination strategies, and advancing clinical development to fully realize the potential of iMQT_020 as a precision medicine tool.
Methods
Ethics statement
All experiments were conducted in accordance with relevant ethical regulations and guidelines at the Korea National Institute of Health (BEC-CAN-2016-001 and 002), and all studies were approved by the Institutional Animal Care and Use Committee of Yonsei University (IACUC-A-202204-1455-02, IACUC-A-202405-1856-01).
Cell lines
Lung cancer cell lines (NCI-H358, NCI-H596, NCI-H226, NCI-H441, NCI-H1299, A549, and NCI-H460), colon cancer cell lines (HT-29, HCT 116, DLD-1, NCI-H508, SW620, LS 174 T, LS1034, COLO 205, SW480, and MC-38), pancreatic cancer cell lines (SU.86.86., SW1990, BxPC-3, CFPAC-1, Panc10.05, AsPC-1, HPAF-II, Capan02, Panc-1, and MIA PaCa-2), glioblastoma cell lines (U-251MG, A-172, D-54 MG, T98G, SNB-19, and U-343 MG), breast cancer cell lines (MDA-MB-231, HCC1395, HCC1937, and BT-20), ovarian cancer cell lines (TOV-112D, SK-OV-3, SNU-119, and OVCAR3), FHC cells, and BJ cells were obtained from ATCC. Astrocytes and MCF10A, HEK293T, HCC44, HCC15, and NCI-HCC2108 cells were obtained from the Korean Cell Line Bank. 16HBE cells were obtained from Merck. The human pancreatic duct epithelial cell line (HPDE) was provided by Dr. Ming-Sound Tsao. The KPC mouse cell line was provided by the National Cancer Center. The LLC mouse cell line was obtained from Ubigene. All cells were grown in medium at 37 °C with 5% CO2. MIA PaCa-2, Panc-1, HEK293T, astrocyte, LLC, and MC-38 cells were cultured in DMEM supplemented with 10% FBS. HPDE cells were grown in serum-free medium supplemented with EGF and bovine pituitary extract. CFPAC-1 cells were cultured in Iscove’s modified Dulbecco’s medium (IMDM). HPAF-II and BJ cells were cultured in minimum essential medium (MEM) supplemented with 10% FBS. Capan2 cells were cultured in McCoy’s 5 A medium supplemented with 10% FBS. SW1990 cells were cultured in Leibovitz’s L-15 medium supplemented with 10% FBS. FHC were cultured in complete growth medium (DMEM:F12 nutrient mix = 1:1; 10% FBS, extra HEPES at a final concentration of 25 mM; cholera toxin, 10 ng/mL; insulin, 5 μg/mL; transferrin, 5 μg/mL; hydrocortisone, 100 ng/mL; EGF, 20 ng/mL). MCF10A cells were cultured in DMEM supplemented with insulin (5 μg/mL), and 16HBE cells were cultured in DMEM/F12 supplemented with 10% FBS. NCI-H358, NCI-H596, NCI-H226, NCI-H441, NCI-H1299, A549, NCI-H460, HT-29, HCT 116, DLD-1, NCI-H508, SW620, LS 174 T, LS1034, COLO 205, SW480, SU.86.86., BxPC-3, Panc 10.05, AsPC-1, U-251MG, A-172, D-54 MG, T98G, SNB-19, U-343 MG, MDA-MB-231, HCC1395, HCC1937, BT-20, TOV-112D, SK-OV-3, SNU-119, OVCAR3, HCC44, HCC15, NCI-HCC2108, and KPC cells were cultured in RPMI medium supplemented with 10% FBS.
Mice
For in vivo subcutaneous and orthotopic tumor xenograft assays, five-week-old male athymic NCr-nu/nu mice (RRID:MGI:5652489, Koatech Co., Pyeongtaek, Republic of Korea) and five-week-old male C57BL/6 N mice (RRID:C57BL/6NCrlOri, Orient Co., Seongnam, Republic of Korea) were used. The mice were housed at 4 per cage at 24 °C under a 12-h light/dark cycle. The mice had access to chow diet (38057, Purina) and water ad libitum in a pathogen-free facility, and their health status was monitored via culture, serum analysis, and microscopic examination. The maximal tumor size permitted by the Institutional Animal Care and Use Committee of Yonsei University was 3000 mm3 which was not exceeded during this study.
Cell lysis and immunoblotting
The cells were washed once with DPBS and first lysed with lysis buffer (40 mM HEPES (pH 7.4), 0.5% Triton X-100, 10 mM β-glycerol phosphate, 10 mM pyrophosphate, and 2.5 mM MgCl2) supplemented with a protease inhibitor (5 μg/mL aprotinin, 10 μg/mL leupeptin, and 250 μM PMSF). The lysate was subsequently sonicated on ice. Unless mentioned otherwise, samples containing SLC1A5_var were never heated to 40 °C or frozen whenever they were used for immunoblotting. The total protein concentration was measured with a BCA assay (Intron), and the proteins were denatured in Laemmli sample buffer immediately before SDS‒PAGE. A total of 30 μg of protein was subjected to SDS‒PAGE and transferred to a PVDF membrane (Merck) via the Trans-BlotR Turbo Blotting System (Bio-Rad). The membranes were subsequently blocked in TBST buffer containing 2% skim milk and incubated with the appropriate primary antibodies for 18 h. The membranes were incubated with the appropriate secondary antibodies (anti-mouse or anti-rabbit IgG conjugated with horseradish peroxidase, Invitrogen) for 1 h. The immunoblot signals were detected with a MicroChemi instrument (DNR Bioimaging Systems) with enhanced chemiluminescence (EzWestLumi, ATTO) and quantified via densitometric analysis of the protein bands with the DNR Bioimaging Tool. The immunoblot images are representative of three independent experiments.
Transfection with cDNA constructs and siRNAs
All vectors utilized in this study were produced from SLC1A5, SLC38A1, and SLC38A2 cDNA provided by the Korean Human Gene Bank (Medical Genomics Research Center, KRIBB). For expression studies, the coding sequences of human SLC1A5_var were cloned and inserted into the pIRESpuro3 vector with Myc, His, and 3XFlag tags. The SLC1A5_var serial deletion sequence was also cloned and inserted into pIRESpuro3. The TM5 Del, HP2 Del, TM4 Del, HP1 Del, and TM3 Del constructs consisted of aa 1-252, aa 1-208, aa 1-169, aa 1-130, and aa 1-90, respectively. The EGFP-fused SLC1A5_var protein was cloned as previously described4. FIL/AAA, GK/AA, D/A, and LRKY/AAAA were generated via PCR via primers containing point mutations. For the transfection of these cDNAs, cells were grown at 1.0 × 106 cells/well in 6-well plates, and after overnight incubation, the cells were transfected with the respective cDNA cloning vectors utilizing TurboFect transfection reagent (Thermo Fisher Scientific, #R0531). The cells were used for the experiments twenty-four hours prior to transfection. For the siRNA experiments, the siRNAs were transfected via Lipofectamine RNAiMAX reagent (Invitrogen) according to the manufacturer’s instructions.
Submitochondrial fractionation
Mitochondria were isolated as previously described62. All the experiments were performed on ice with prechilled buffers and equipment. The cells cultured in 15-cm dishes ( > 90% confluence) were washed twice with KPBS (100 mM KCl, 36 mM NaCl, 10 mM KH2PO4, pH 7.2) and then gently scraped into 1 mL of KPBS. The cells were centrifuged at 900 × g for 3 min, the supernatant was discarded, and the cell pellets were resuspended in 1 mL of KPBS supplemented with protease inhibitors (5 mg/mL aprotinin, 10 mg/mL leupeptin, and 250 mM PMSF). The cells were then lysed by homogenization with 50 strokes inside a 2-mL Dounce homogenizer to avoid introducing air bubbles into the samples (cell integrity was carefully examined under a microscope after 50 strokes to confirm that at least 80% of the total cells were damaged). The homogenate was centrifuged at 600 × g for 5 min, and the pellet was retained for further experiments. The supernatant was centrifuged at 7000 × g for 10 min, after which the supernatant was discarded. Next, the pellet was resuspended with a Dounce homogenizer (rather than pipetting). Finally, the mitochondrial suspension was centrifuged at 10,000 × g for 10 min. The total mitochondrial protein content was determined with a BCA assay kit (Intron).
A previously described method was used for submitochondrial fractionation63. In brief, mitochondria purified as described above were suspended in 500 μL of swelling buffer (10 μM KH2PO4, pH 7.4) supplemented with digitonin (2 mg/mL) and homogenized with a Dounce homogenizer (Sigma). The mixture was incubated on ice for 1 h and then tapped every 10 min for swelling. Then, 1 volume of an iso-osmotic solution (32% sucrose, 30% glycerol, and 10 mM MgCl2) was added. The mixture of the two solutions was centrifuged at 10,000 × g for 10 min. The supernatant of the S1 fraction contained the outer mitochondrial membrane. The pellet (P1 fraction) contained the matrix and intact inner membrane. P1 was resuspended in 500 μL of swelling buffer, homogenized with a Dounce homogenizer (Sigma) and kept on ice for 1 h for swelling. Then, 1 volume of an iso-osmotic solution was added. The S1 fraction and the resuspended P1 fraction were centrifuged at 17,000 ×g for 1 h. The supernatant from fraction S1 contained the outer membrane. The supernatant from fraction P1 contained the matrix, and the pellet contained the inner membrane. Each fraction was analyzed by immunoblotting with the indicated antibodies.
Crosslinking
Mitochondria isolated from 1 × 106 cells were incubated with a formaldehyde gradient from 0 to 1% for 30 minutes on ice, and the reaction was quenched by the addition of 50 mM Tris-Cl (pH 8.0). Then, the crosslinked lysate was mixed with Laemmli buffer, separated on 10% SDS‒PAGE gels and subjected to immunoblotting. Then, 50 μL of purified eGFP-tagged SLC1A5_var was incubated with a formaldehyde gradient from 0 to 1% for 30 minutes on ice, and the reaction was quenched by the addition of the same volume of 50 mM Tris-Cl (pH 8.0). Then, the crosslinked lysate was mixed with Laemmli buffer, separated on 10% SDS‒PAGE gels and subjected to immunoblotting.
Purification of SLC1A5_var
eGFP-tagged SLC1A5_var was expressed in SF9 insect and HEK293T mammalian cells, and the cells that were transfected with eGFP-tagged SLC1A5_var were collected via centrifugation. The membranes were disrupted in buffer containing 20 μM HEPES (pH 7.5) and 150 mM NaCl with a homogenizer. The purified membranes were then solubilized in buffer containing 20 mM HEPES (pH 7.5), 150 mM NaCl, 0.5% (w/v) n-dodecyl-beta-D-maltopyranoside (DDM, Anatrace), and 0.2% (w/v) cholesterol hemisuccinate (CHS, Sigma) at 4 °C for 2 h. The solubilized protein was isolated by ultracentrifugation at 60,000 × g at 4 °C for 40 min and then incubated at 4 °C overnight with TALON resin. The resin was then washed with wash buffer (20 mM HEPES (pH 7.5), 100 mM NaCl, 0.05% DDM, 20 mM imidazole), and the protein sample was eluted with elution buffer (20 mM HEPES (pH 7.5), 100 mM NaCl, 0.05% DDM, 250 mM imidazole). For analysis, the buffer of the protein sample was exchanged for size exclusion chromatography.
Bioluminescence Resonance Energy Transfer (BRET)/Bimolecular Fluorescence Complementation (BiFC)
HEK293T and MIA PaCa-2 cells were transiently co-transfected with N-YFP-SLC1A5_var, C-YFP-SLC1A5_var, and RLuc-SLC1A5_var constructs. After 24 h, cells were detached and washed with HBSS buffer containing 10 mM glucose. The samples were normalized to the total protein concentration level determined by the Bradford assay kit. 10 μg of cell suspension was distributed in triplicate into white plates with a white bottom for bioluminescence readings and black plates with a transparent bottom for fluorescence readings. YFP fluorescence was calculated by subtracting the fluorescence background signals of cells expressing only RLuc-SLC1A5_var cells. BRET was determined by adding 5 mM coelenterazine and reading the bioluminescence signal after 1 min and 10 min. The ratio of light emitted by YFP (550 nm) over light emitted by RLuc (450 nm) were determined as BRET-BiFC ratio.
Microscale thermophoresis
Microscale thermophoresis experiments were performed with Monolith NT.115 (NanoTemper Technologies). eGFP-tagged SLC1A5_var proteins were purified as described above. The protein samples were diluted with 20 mM HEPES (pH 7.5), 100 mM NaCl, and 0.05% DDM and assayed with Monolith to confirm the absence of aggregation, revealing the optimal level of initial fluorescence. A series of dilutions with the same buffer were performed as the instrument guided, and each concentration of iMQT_020 was mixed with the protein at a ratio of 1:1 and incubated inside the Premium Capillary (Nanotemper, MO-K025). Red fluorescence was selected as the LED source, and 50% infrared-laser power was used at 25 °C. Analysis of the data was performed with MO Affinity Analysis 2.3.
Circular dichroism analysis
Circular dichroism measurements were performed with a Jasco J-1500 spectropolarimeter (Japan Spectroscopic Company). SLC1A5_var proteins were purified as described above. The protein samples were diluted to 0.5 mg/mL and scanned at 25 °C between 190 and 340 nm via a 1 mm optical path. The spectral data were corrected for background scattering by subtracting buffer or vehicle-added spectrum measurements. CD and absorbance were measured in the same cuvette to minimize errors in the determination of molar ellipticity. When samples with different molar ratios of WT and FIL/AAA mutant proteins were measured, the total concentration of the proteins was maintained at 0.5 mg/mL. CD spectra were measured with buffer containing 20 mM HEPES (pH 7.5) and 100 mM NaCl.
Coimmunoprecipitation
Different cDNA coding sequences were co-transfected at a ratio of 1:1 or 1:1:1 according to the experimental conditions. After twenty-four hours, the cells were harvested via lysis as described in the Cell Lysis section. The cell lysates were incubated for 4 h with 1 μg of the appropriate antibodies in the presence of Protein A/G agarose beads (InvivoGen) at 4 °C. The resulting complexes were washed four times with washing buffer containing 40 mM HEPES (pH 7.4), 0.5% Triton X-100, 10 mM β-glycerol phosphate, 10 mM pyrophosphate, and 2.5 mM MgCl2 and eluted with 2× Laemmli buffer with vigorous vortexing. The eluates were isolated through centrifugation and subjected to SDS‒PAGE for immunoblotting. For two-step co-IP, the cell lysates were incubated for 4 h with 1 μg of anti-3XFlag (Bethyl) antibody in the presence of Protein A/G agarose beads (InvivoGen) at 4 °C. The resulting complexes were washed four times with washing buffer. Then, 340 μM 3XFlag peptide (Sigma‒Aldrich, #F4799) was added to the tube containing the agarose-protein-3XFlag antibody complex. The mixture was incubated for 4 h at 4 °C. Agarose beads were separated through centrifugation, and the supernatant was added to a tube containing 1 μg of anti-His (Bethyl) antibody in the presence of Protein A/G agarose beads (InvivoGen) and incubated at 4 °C for 4 h. The complexes were subsequently washed four times with washing buffer and eluted with 2× Laemmli buffer with vigorous vortexing. The eluates were isolated through centrifugation and subjected to SDS‒PAGE for immunoblotting.
Size exclusion chromatography
The membrane fractions of lysates of HEK293T cells transfected with eGFP_SLC1A5_var_WT supplemented with vehicle and iMQT_020 or eGFP_SLC1A5_var_FIL/AAA supplemented with vehicle or iMQT_020 were applied to a Superdex 200 Increase 10/300 GL column supplemented with 20 mM Tris (pH 8.0), 300 mM NaCl, and 0.2% DDM. A Superdex 200 Increase 10/300 GL column was calibrated with apoferritin (440 kDa), alcohol dehydrogenase (150 kDa), conalbumin (75 kDa), ovalbumin (44 kDa), carbonic anhydrase (29 kDa), and ribonuclease (13.7 kDa), which were purchased from GE Healthcare. The eGFP-tagged proteins from the collected fractions were detected by a fluorescence detector with an excitation wavelength of 488 nm and an emission wavelength of 510 nm.
SLC1A5_var modeling
RoseTTAFold2.0 was used to construct a de novo-predicted model of the SLC1A5_var monomer. The input included the query sequence of SLC1A5_var, and no templates or MSAs from MMseqs2 were used. The RoseTTAFold2.0 prediction was run with the ColabFold code on a local GPU cluster. The output monomeric model was subsequently input to the MZ-Dock server to produce a homotrimeric structure. After coverage of the TM3 domain as the site of interprotomer interaction, homotrimeric structure modeling was performed. As the output homotrimer included bond clashes, additional refinement through molecular dynamics (MD) was performed. The MD simulation was conducted with Desmond with the default settings. The Desmond application was opened, and the predicted homotrimeric structure of SLC1A5_var was uploaded. Energy minimization was conducted with Desmond simulation. Structural relaxation was conducted with the relax panel of Desmond. Finally, NVIDIA GeForce GTX 1660 Super was selected for the final MD simulation.
In silico docking and virtual screening
Molecular docking-based virtual screening was used to screen the ChemBridge database (1.2 million compounds). The Glide software program (Schrodinger, LLC, USA) was used for virtual docking of the compounds, and grid-based ligand docking with an energetics algorithm was used. The homology model structure of the SLC1A5_var trimer was used as a starting model for virtual screening. A grid box was generated by selecting the center of the residues Phe97, Iso104, and Leu105 as the centroid of the grid box (20 Å × 20 Å × 20 Å) by utilizing the receptor grid generation panel implanted within the Schrodinger suite. The protein structure was corrected by adding hydrogen atoms, bond orders, and formal charges with the Protein Preparation Wizard tool of the Maestro software package (version 9.6, Schrodinger, LLC, USA). For ligand preparation, 1.2 million compounds were initially downloaded from the ChemBridge database in SDF format. The LigPrep module of the Schrodinger software suite was utilized for ligand preparation. The following criteria were used for ligand preparation: (i) an OPLS3 force field, (ii) the generation of all possible ionization states at pH 7.0 ± 2.0, (iii) the desalt option, (iv) the generation of a tautomer for all conformers, and (v) the generation of one low-energy conformer per ligand. The prepared ligands were subsequently subjected to a structure-based virtual screening process with the Glide module of the Schrodinger suite. HTVS mode was employed to filter the chemical entities suggested from the library of molecules, SP mode was applied for further screening, and XP mode was utilized to obtain more accurate docking calculations.
Cellular amino acid uptake assay
HEK293T and MIA PaCa-2 cells at 75% confluence were seeded in 12-well culture plates. Radiolabeled proline (L-[3H]Pro), serine (L-[3H]Ser), glutamine (L-[3H]Gln), asparagine (L-[3H]Asn), leucine (L-[3H]Leu), and glutamate (L-[3H]Glu) (Perkin Elmer, 3μCi/mL) were added to DMEM without the corresponding amino acids. Then, the cells were incubated at 37 °C for an h, and the reaction was terminated by removing the medium. The cells were washed with DPBS solution, extracted with lysis buffer (0.2% SDS), and mixed with AquaLight+ (HIDEX). All the measurements were analyzed with a liquid scintillation counter (HIDEX). The lysate was subsequently used to measure the total protein concentration of each sample with a BCA assay kit (Intron). The uptake of amino acids was calculated from the counts per minute per sample and the specific activity of each labeled amino acid, and the uptake of these amino acids was normalized to the total protein content.
Purification of mitochondria from cells and mitochondrial amino acid uptake assay
The mitochondria were prepared according to the methods of Wieckowski et al., with slight modifications62. For each experiment, HEK293T and MIA PaCa-2 cells that were stably transfected with 3XMyc-EGFP-OMP25 were immunoprecipitated with an anti-Myc antibody. Then, 100 μl of anti-Myc magnetic beads (Pierce) was prewashed three times with KPBS. All washes were performed by gentle pipetting via a wide-bore pipette tip, and the beads were collected with a magnetic stand (DynaMag Spin Magnet, Invitrogen). A total of 1 × 107 cells cultured in 15-cm dishes were washed twice with DPBS and then gently scraped into 1 mL of KPBS. The cells were subsequently centrifuged at 900 × g for 3 min, after which the pellet was resuspended in 1 mL of KPBS. The cell mixture was carefully homogenized with 50 strokes in a 2-mL Dounce homogenizer to avoid air bubbles. The homogenate was centrifuged at 1000 × g for 2 min, and the supernatant was immunoprecipitated with anti-c-Myc antibody-conjugated magnetic beads for 20 min. The c-Myc-tagged mitochondria were eluted with 250 mL of 0.25 mg/mL c-Myc peptide (Pierce). Mitochondrial amino acid uptake was measured by first diluting the purified mitochondria in 50 μL of KPBS. This solution was placed in a 1.5-mL tube with vehicle or chemicals for 24 h. Radiolabeled proline (L-[3H]Pro), serine (L-[3H]Ser), glutamine (L-[3H]Gln), asparagine (L-[3H]Asn), leucine (L-[3H]Leu), and glutamate (L-[3H]Glu) (Perkin Elmer) (3 μCi/mL) were used to assess mitochondrial amino acid uptake by the addition of labeled amino acids to the respective tubes, followed by incubation at 37 °C for one h. Uptake was stopped by incubating the tubes at 4 °C. Then, the tubes were centrifuged at 12,000 × g for 15 min, and the supernatant was removed and washed with KPBS twice. The cells were extracted with lysis buffer (0.2% SDS) and mixed with 3 mL of AquaLight+ (HIDEX). All measurements were analyzed by a liquid scintillation counter (HIDEX). The lysate was subsequently used to measure the mitochondrial protein concentration of each sample with a BCA assay kit (Intron). Amino acid uptake was calculated from the counts per minute per sample and the specific activity of each labeled amino acid, and the uptake of these amino acids was normalized to the mitochondrial protein content.
Transporter and channel protein activity test
SLC1A5-, SLC38A1-, SLC38A2-, SLC6A14-, SLC6A19-, SLC26A3-, SLC26A4-, SLC26A6-, SLC26A7-, SLC26A9-, or ANO1- and YFP-F46L/H148Q/I152L-expressing cells were plated in 96-well culture plates at a density of 2 × 104 cells/well and incubated for 24 h. Each well of the 96-well culture plates was washed 2 times with 100 μL of DPBS, and the wells were filled with 50 μL of HEPES-buffered solution. Each compound (vehicle, iMQT_020, or V-9302) was added at 1, 3, 10, 30, 100, or 300 μM before activity measurement. The 96-well culture plates were placed on a FLUOstar Omega Microplate Reader (BMG Labtech, Ortenberg, Germany) for fluorescence assays. Each well was assayed individually for iodide ion influx mediated by the control treatment or SLC1A5, SLC38A1, SLC38A2, SLC6A14, SLC6A19, SLC26A3, SLC26A4, SLC26A6, SLC26A7, SLC26A9, and ANO1 by continuously recording fluorescence (1 second per point) for 5 seconds (baseline). Then, 50 μL of NaI- or NaSCN-substituted HEPES-buffered solution (with NaI or NaSCN replacing NaCl from the original HEPES buffer) was added to the liquid injector at 5 seconds, and YFP fluorescence was recorded for 10 seconds. The activity of each transporter or channel was calculated by the gradient of YFP fluorescence quenching at the moment of halide salt injection.
Analysis of mitochondrial morphology
Mitochondrial morphology was analyzed as previously described64, and mitochondria were categorized into three groups: “elongated/tubular” mitochondria, with >90% of the mitochondria forming elongated interconnected networks; “intermediate” mitochondria, with mixed tubular and short mitochondria; and “fragmented” mitochondria, with >90% short punctiform mitochondria. The percentages of cells with elongated/tubular, intermediate, or fragmented mitochondria were measured, and at least 100 cells from each group of vehicle- and iMQT_020-treated cells were quantified in samples from three biologically independent experiments.
Measurement of the mitochondrial membrane potential
Changes in the mitochondrial membrane potential were analyzed by incubating cells treated with vehicle or iMQT_020 for 15 min with tetramethylrhodamine ethyl ester (TMRE 20 nM, Invitrogen), and the results were analyzed by flow cytometry (BD FACS ARIA III).
OCR/ECAR measurement
The OCR and ECAR were determined with an XFe24 extracellular flux analyzer (Agilent Technologies) as described in the manufacturer’s protocol. A total of 4.0 × 104 cells were seeded per well in 24-well micro cell culture plates for Seahorse (Agilent Technologies) in DMEM supplemented with 10% FBS and incubated at 37 °C in a 5% CO2 incubator overnight. Then, the DMEM was removed and replaced with bicarbonate-free DMEM containing phenol red (pH 7.4), and the cells were incubated at 37 °C in a non-CO2 incubator to equilibrate the CO2 concentration in the atmosphere for one h. Using the XFe24 analyzer, the OCR and ECAR were measured under baseline conditions and upon treatment with certain drugs/compounds, such as glucose (Glc, 4 mM), 2-deoxy-glucose (2-DG, 50 mM), glutamine (Gln, 2 mM), dimethyl-alpha-ketoglutarate (DM-αKG, 2 mM), oligomycin (2 mM), FCCP (carbonyl cyanide-p-trifluorome-thoxyphenylhydrazone, 0.5 mM), and rotenone/antimycin A (0.5 mM/0.5 mM) (these drugs were included in the Cell Mito Stress Test Kit, Agilent Technologies). For the metabolic phenotype assay, the basal OCR and basal ECAR of the cells were measured. To measure the maximum OCR and ECAR, the cells were exposed to oligomycin (2 mM) and FCCP (0.5 mM). Each measurement cycle consisted of 3 min of mixing, 3 min of waiting, and 4 min of measurement. The OCR and ECAR values were normalized to the number of cells and analyzed with WAVE software (Agilent Technologies).
OCRs were measured in isolated mitochondria, MIA PaCa-2 cells transfected with siCon, siSLC1A5_var, siSLC1A5_var + SLC1A5_var WT, or siSLC1A5_var + SLC1A5_var FIL/AAA mutant were prepared, and their mitochondria were isolated with a Mitochondria Isolation Kit (Abcam). Isolated mitochondria were resuspended in mitochondrial assay solution (70 mM sucrose, 220 mM mannitol, 10 mM KH2PO4, 2.5 mM MgCl2, 2.5 mM NaCl, 2 mM HEPES, 1 mM EGTA, 4 mM ADP, and 0.2% (w/v) fatty acid-free BSA, pH 7). 2). A total of 6 μg of mitochondrial protein was seeded at 540 μL on a 24-well Seahorse culture plate, and the plates were spun down at 2000 × g for 20 minutes at 4 °C. 60 μL of 40 mM glutamine, 66 μL of 20 mM oligomycin, 72 μL of 5 mM FCCP, and 78 μL of 5 mM rotenone and 5 mM antimycin A were injected to analyze the mitochondrial metabolic state.
Measurement of GSH, ROS, and MitoROS
Cellular GSH contains a thiol group that can be derivatized with monobromobimane, a fluorescent tag (Ex/Em: 394/490 nm)65. The cells were incubated with monobromobimane (40 mM, 10 min), and the stained cells were analyzed via a flow cytometer (BD FACSAria III). The ratio of GSH to oxidized glutathione (GSSG) was measured with a GSH/GSSH assay kit (Abcam) according to the manufacturer’s instructions. The fluorescence at an excitation wavelength of 490 nm and an emission wavelength of 520 nm was recorded, and the GSH/GSSG ratio was calculated on the basis of the formula in the manufacturer’s instructions. Intracellular ROS levels were determined by staining cells with 1 mM 5-(and-6)-carboxy-2’,7’-dichlorodihydrofluorescein diacetate (H2DCFDA, Invitrogen) for 10 min according to the manufacturer’s protocol (Ex/Em: 492/517 nm). The stained cells were analyzed via flow cytometry (BD FACSAria III). MitoROS levels were determined by staining cells with the mitochondrial superoxide indicator MitoSOX Red at 5 mM (Ex/Em = 510/580 nm) for 10 min, and stained cells were analyzed via flow cytometry (BD FACSAria III).
Cell viability and growth measurements
Dialyzed FBS (Gibco) was used to exclude the effects of serum-derived nutrients. For the cell viability assays, cells were seeded into 96-well culture plates at a density of 10,000 cells per well in the appropriate medium supplemented with 10% dialyzed FBS. After successful cell attachment, the medium was replaced with conditioned medium. After 48 h, cell viability was measured via a WST-1 assay. The WST-1 assay uses an extremely sensitive, water-soluble tetrazolium salt to produce a water-soluble formazan dye upon reduction in the presence of mitochondrial dehydrogenases. The absorbance of each sample at 450 nm (reference wavelength 625 nm) was measured with a microplate reader (Infinite 200 PRO, Tecan) and normalized to that of the control. The cells were counted via an automated cell counter (ADAM-MC, NanoEntek).
Cell cycle analysis
Cells were seeded in a 60 mm plate and treated at the selected conditions for 24 h or 48 h. The cells were detached and washed with ice-cold PBS buffer three times and fixed with 70% ethanol for 30 min at ice. The cells were stained with 100 μg/mL of propidium iodide (Pi) and treated with 100 μg/mL RNase A for 30 min. The DNA content was measured through flow cytometry (BD FACSAria III)
Human patient-derived pancreatic tumor organoid culture and drug response
Human patient-derived organoids were prepared following a previous report66,67,68. Patient-derived sample collection and the study protocols were approved by the Institutional Review Board of the National Cancer Center of Korea (NCC2019-034, NCC2021-0232, NCC2021-0282). Written informed consent was obtained from all participants prior to enrollment. No monetary or other forms of compensation were provided. Pancreatic tumor tissues or ascites were obtained from 10 patients who (4 males and 6 females; age range: 49-81 years) visited the pancreatobiliary cancer clinic at the National Cancer Center, Korea (see Supplementary Table 2 for details). Samples were collected from surgical resections, endoscopic ultrasound-guided (EUS) biopsies, gun biopsies, and ascites drainage. Sex was determined from clinical records. However, sex and/or gender were not considered in the study design or analysis, as the primary aim was to establish patient-derived organoid models and evaluate drug responses. Organoids were cultured in growth media consisting of advanced DMEM/F12 medium (Invitrogen) containing B-27 (Invitrogen, USA), 1.25 mM N-acetylcysteine (Sigma), 10 mM nicotinamide (Sigma), 10 mM HEPES (Gibco), 1% GlutaMAX (Gibco), 50 ng/mL EGF (PeproTech, USA), 100 ng/mL FGF-10 (Peprotech), 50 ng/mL recombinant human R-spondin 1 (Qkine), 100 ng/mL WNT surrogate-Fc fusion protein (U-protein express BV), 1% noggin-Fc fusion protein-conditioned medium (U-protein express BV), 500 nM A83-01 (Tocris), and 1% primocin (InvivoGen) were added. For drug response, cells were seeded at 1 × 103 cells/well with 10% BME in 384-well plates. After 3 days of incubation for organoid formation, the organoids were treated with iMQT_020, V-9302 or iMQT_020 + DM-αKG. The medium used for drug analysis excluded N-acetylcysteine from the organoid growth medium. After 5 days, cell viability was measured via a Cell-Titer Glo® 3D viability assay kit (Promega Corporation). The luminescence intensity was determined via an Infinite 200 Pro spectrophotometer (Tecan).
RNA-seq analysis and GSEA
The quality of the raw RNA-seq data from each sample was controlled through FastQC (RRID:SCR_014583). TopHat (RRID:SCR_013035) was used to map reads from the RNA-seq experiment to a reference genome (Homo sapiens GRCh38.94). The mapped reads were subsequently counted via HTSeq (RRID:SCR_005514) with the following parameters: HTSeq-count -s no -m intersection-nonempty -f bam. Differential gene expression analysis was performed with the DESeq2 package (v. 1.24.0, RRID:SCR_000154), and the data were manipulated in R (v3.6.1). Genes with a low count ( < 10) were filtered out, and regularized log transformation was applied to minimize differences between samples with small counts. The fold change value was calculated based on the housekeeping gene GAPDH. GSEA (RRID:SCR_003199) was used to compare the data from vehicle-treated and iMQT_020-treated MIA PaCa-2 cells. We divided the subsets according to the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway gene sets and calculated the rlog FPKM FC values. For each gene set, the mean rlog FPKM FC value for each sample was subsequently calculated. This approach enabled an overall comparison of vehicle-treated and iMQT_020-treated MIA PaCa-2 cells. The normalized enrichment score was calculated for each KEGG pathway gene set and plotted.
LC/MS-based Metabolomics and Quantification of Metabolite Abundances
Whole-cell metabolite profiling was performed with 1 × 107 vehicle- or iMQT_020-treated MIA PaCa-2 cells for 0, 12, or 24 h treated with glutamine (L-[13C]Gln or L-[15N]Gln) for glutamine or with glucose (L-[13C]Glc) for glucose metabolomics profiling (n = 3). These cells were washed two times with DPBS, briefly washed with Optima LC/MS water and then scraped into 1 mL of 80% Optima LC/MS methanol in water. The samples were vortexed vigorously for 1 min and centrifuged at 17,500 × g for 15 min, after which the supernatants were analyzed via LC‒MS. For ultra-performance liquid chromatography‒triple quadrupole mass spectrometry (UPLC‒TQ‒MS), the metabolite extract was diluted with 20% (v/v) acetonitrile. An Ultimate 3000 RSLC instrument equipped with an Acquity UPLC BEH C18 (1.7 μm, 2.1 × 100 mm) and Q Exactive Orbitrap Plus MS (Thermo Fisher) was used for analysis. Compound Discoverer (ver 3.0, Thermo Fisher) software was used for data acquisition and analysis. MS was performed with the following settings: gas temperature, 37 °C; isolation window, 2.0 m/z; collision energy, 30; dynamic exclusion, 5 s; sheath gas flow rate, 50; aux gas flow rate, 13; and spray voltage, 2.5 kV. Metabolomic analysis was conducted with the Metabolika Pathway Analysis software Compound Discoverer (ver 3.0, Thermo Fisher). The detected metabolites were subjected to Metabolika pathway library comparison. All molecules present in the library were identified from the obtained data. Peaks were quantified via the area under the curve. The fold change was calculated, and statistical significance analyses were performed with GraphPad Prism 7 software.
Gene expression analysis
A MiniBEST Universal RNA Extraction Kit (TAKARA) was used to isolate RNA from cells according to the manufacturer’s instructions. Reverse transcription of 1,000 ng of RNA was conducted with the PrimeScript 1st Strand cDNA Synthesis Kit (TAKARA). The complete reaction mixture was incubated at 42 °C for 45 min, and the reverse transcriptase was inactivated by incubation at 90 °C for 5 min. For RT‒PCR, the resulting cDNA was diluted in nuclease-free water (1:1) and subjected to PCR using EmeraldAmp GT PCR Master Mix (TAKARA). The PCR products were subsequently analyzed via 1% agarose gel electrophoresis. Gene expression levels were determined with ImageJ and normalized to the expression level of the housekeeping gene GAPDH. For quantitative real-time PCR analysis, cDNA was diluted in nuclease-free water (1:4), and gene expression levels were analyzed with a Step One Plus system (Applied Biosystems). The expression levels were normalized to the expression level of the housekeeping gene ACTB. The primers used for the experiment are listed in Supplementary Table 8.
Glucose and glutamine measurements
The glucose concentration was determined with a Glucose Colorimetric Assay Kit II (BioVision), and the glutamine concentration was determined with a Glutamine Detection Assay Kit (BioVision) in accordance with the manufacturer’s instructions. The compounds of interest produced stable colorimetric signals that were observed by measuring the absorbance at a specific wavelength (Glucose Colorimetric Assay Kit II and Glutamine Detection Assay Kit: 450 nm) with a microplate reader (Infinite 200 PRO, Tecan). The absorbance of each control well was also measured for background subtraction.
Ames test and hERG toxicity test
Ames test strains (TA98 and TA100 cells) and media were purchased from Xenometrix (Basel, Switzerland). Test procedures were performed according to the manufacturer’s instructions. iMQT_020 and the positive control drugs were prepared at 10.2, 30.8, 92.5, 277.7, 833.3, and 2500 μg/mL, and 2 μg/mL 2-NF was prepared for TA98 cells without S9, 0.5 μg/mL 2-AA was prepared for TA98 cells with S9, 0.1 μg/mL 4-NQO was prepared for TA100 cells without S9, and 1.25 μg/mL 2-AA was prepared for TA100 cells with S9. Each chemical was dissolved in the exposure medium and incubated at 37 °C for 90 min. Indicator medium was added, and the samples were aliquoted into 284-well plates and cultured for 24 ~ 48 h. Mutagenicity was assessed according to the baseline indicated by the control group.
Cells expressing hERG were plated on 8-mm coverslips and incubated overnight. Each coverslip was placed on a microscope stage in a chamber and perfused at 1 mL/min with saline containing 160 mM NaCl, 4 mM KCl, 2 mM CaCl2, 1 mM MgCl2, 10 mM HEPES, and 10 mM glucose. All the recordings were conducted at room temperature. Currents were recorded in whole-cell recording mode. The cell membranes were held at -90 mV, and the currents were filtered at 67 Hz and sampled at 2.0 kHz with an EPC10/2 amplifier. Voltage-dependent activation curves were obtained by increasing the potential to -70 mV for 50 ms, increasing the command potential from -70 mV to -40 mV for 2 s and repolarizing the membrane by decreasing the voltage to -70 mV for 2 s, with the potential held at -90 mV every 5 s.
Serum profiles for biomarkers of liver and renal injury
Two months after chemical administration, blood samples from each animal were collected, and serum samples were acquired via centrifugation at 5000 × g for 20 min. Aspartate aminotransferase (AST), alanine aminotransferase (ALT), and blood urea nitrogen (BUN) levels were determined with AST (BioAssay System, EASTR-100), ALT (BioAssay System, EALT-100), and BUN (BioAssay System, DIUR-100) assay kits following the manufacturers’ protocols.
Test of iMQT_020 Plasma Stability
Whole-blood samples (n = 5 mice) were collected at 0, 0.08, 0.25, 0.5, 1, 2, 4, 6, 8, and 10 h after intravenous or intraperitoneal injection of iMQT_020 at 5 mg/kg or 75 mg/kg, respectively. LC‒MS/MS was performed via reverse-phase chromatography with a Q Exactive Orbitrap Plus MS (Thermo Fisher). Standard curves with quality control samples were generated by spiking iMQT_020 into fresh whole blood and then applying the spiked blood to 10 μL Mitra microsamplers. All the data were collected by Compound Discoverer and analyzed with its software.
In vivo subcutaneous xenograft and orthotopic xenograft models
All animal experiments were conducted in accordance with the Guide for the Care and Use of Laboratory Animals published by the Korea National Institute of Health (BEC-CAN-2016-001 and 002), and all studies were approved by the Institutional Animal Care and Use Committee of Yonsei University (IACUC-A-202204-1455-02, IACUC-A-202405-1856-01). Xenografts were generated with five-week-old male athymic NCr-nu/nu mice for MIA PaCa-2, NCI H1299, and COLO 205 and five-week-old male C57BL/6 N for KPC, LLC, and MC-38. After the subcutaneous injection of 5.0 × 105 MIA PaCa-2, NCI H1299, COLO 205, KPC, LLC, or MC-38 mouse cells in 200 μL of DPBS, when the tumor size reached approximately 30 mm3, the mice were treated with vehicle (10% DMSO, 10% Tween 20, 0.9% NaCl, 79.1% water, IP, every day), iMQT_020 (75 mg/kg, IP, every day), V-9302 (75 mg/kg, IP, every day) or anti-PD-L1 (200 μg, IP, every 3 days). Tumor volumes were measured with electronic calipers every day and calculated via the following formula: tumor volume = (length × width × height)/2.
MIA PaCa-2 cells stably expressing luciferase were orthotopically injected into five-week-old male athymic nu/nu mice. The mice were anesthetized with isoflurane, and a small incision was made in the left abdomen. A total of 5.0 × 105 cells were diluted with 100 μL of DMEM and injected into the tip of the mouse pancreas under isoflurane anesthesia (Pfizer). A swab was held at the injection site for 1 min to prevent leakage of the pancreatic tumor cells. The peritoneum and skin incisions were sequentially sutured with absorbable sutures. When the tumor volume reached approximately 30 mm3, the mice were treated with vehicle (10% DMSO, 10% Tween 20, 0.9% NaCl, 79.1% water, daily), iMQT_020 (75 mg/kg, IP, daily) or V-9302 (75 mg/kg, IP, every day). Every three days, the mice were injected with D-luciferin (3 mg/mouse, IP) for 10 min, and luminescence was measured with an IVIS instrument (Caliper Life Sciences). At the end of the study, the mice were sacrificed via CO2 asphyxiation, and the tumor volume (n = 5) was measured.
Chromatin immunoprecipitation assay
The cells were cultured in medium containing vehicle, iMQT_020, or DM-αKG for 24 h, and ChIP assays were performed with a ChIP kit (Abcam) according to the manufacturer’s protocols. In brief, 4 × 106 cells were fixed with 1.5% formaldehyde for 10 min. Nuclei were isolated, and DNA from 150-μg nuclear samples was sheared into fragments of 200 ~ 1000 bp by sonication. The resulting fragmented chromatin was subjected to ChIP with 5 μg of an anti-H3K4me3 antibody, anti-H3K27me3 antibody, or control IgG complexed with Protein A/G agarose beads. The formaldehyde crosslinked immunoprecipitated DNA was dissociated by heating (95 °C for 10 minutes). Proteins were removed via proteinase K treatment. As a control, non-enriched DNA samples were subjected to the same protocol. The resulting ChIP and input DNAs were subjected to qRT‒PCR using primers specific for the promoter regions of PD-L1 or MYC.
Immunohistochemistry staining
The tumor tissues were dissected after the mice were sacrificed. The tumor tissues were fixed with formalin and sectioned for IHC staining. IHC staining was carried out with a fully automated Ventana Discovery Ultra instrument (Roche Diagnostics International AG). The formalin-fixed tumor tissue sections were deparaffinized, and antigen retrieval was performed with CC1 buffer (Roche Diagnostics). The sections were subsequently incubated with cleaved caspase-3 (9661, Cell Signaling Technology), Ki-67 (ab16667, Abcam), PD-L1 (13684, Cell Signaling Technology), CD8 (98941 T, Cell Signaling Technology), IFNγ (MAB485-SP, R&D Systems), and granzyme B (46890S, Cell Signaling Technology) antibodies at 37 °C for 30 minutes. An Ultramap DAB Staining Kit (Roche Diagnostics) was used for tissue slide development according to the manufacturer’s instructions. The number of positive cells per field was determined in at least six areas at 10x magnification, and the values were averaged with ImageJ.
Flow cytometry analysis of PD-L1 expression and tumor-infiltrating immune cells
The tumor tissues were excised and washed with precooled PBS. Tumors were chopped and digested in RPMI medium containing 50 μg/mL DNase (10104159001, Roche) and 150 μg/mL Liberase (05401020001, Roche) at 37 °C for 15 min. Digestion was stopped by supplying 10% RPMI medium. The fragmented tissue was filtered through a 40 μm cell filter (431750, Corning). The pellet of the filtered cells was lysed with RBC lysis buffer (00-4333-57, Invitrogen). The reaction was stopped by adding 10 mL of complete RPMI media and washing. The tumor single-cell suspensions were blocked before being stained with an anti-mouse CD16/CD32 antibody (156604, Biolegend). The cells were then stained with anti-mouse FITC CD45 (S18009F, Biolegend), APC-Cy7 PD-1 (135223, Biolegend), PE PD-L1 (1242307, Biolegend), PE-Cy7 CD3 (100219, Biolegend), PE-Cy5 CD8 (100709, Biolegend), PerCP-Cy5 CD4 (100433, Biolegend), PE CD45 (147711, Biolegend), PerCP-Cy5 CD8 (140417, Biolegend), PE-Cy7 TCR-β (109221, Biolegend), FITC CD25 (102005, Biolegend), PerCP-Cy5 CD11β (101229, Biolegend), V450 Ly6G (127603, Biolegend), APC Ly6C (128005, Biolegend), PE CD206 (141705, Biologend), and PE-Cy7 F4/80 (157305, Biolegend) antibodies. After staining for cell surface antigens, the cells were fixed with fixation buffer (420801, Biolegend), permeabilized with permeabilization wash buffer (421002, Biolegend), and stained for intracellular antigens with anti-mouse APC FoxP-3 (320011, Biolegend), FITC Granzyme B (515403, Biolegend), and APC IFNγ (505809, Biolegend). Live and dead cells were excluded via PI (421301, Biolegend) or viability stain (AB_2869405, BD Biosciences). Finally, the samples were collected and analyzed via flow cytometry (BD FACSAria III) and Flow Jo (BD Biosciences). The gating strategy for the analysis of flow cytometry data was conducted as described in the Supplementary Information.
CD8+ T cell experiments
CD8+ T cells were isolated from the spleens and thymocytes of 8-week-old C57BL6/N mice in accordance with the Guide for the Care and Use of Laboratory Animals. The splenocytes and thymocytes were stimulated with anti-CD3 (5 μg/mL) and anti-CD28 (5 μg/mL) antibodies and activated recombinant IL-2 (10 ng/mL) for up to 7 days. After complete activation, the cells were stained with FITC-annexin V and incubated for 30 min at room temperature in the dark, followed by analysis via flow cytometry (BD FACS Aria III). To analyze the CD8+ T-cell activation state, CD44-PE/Cy5 was used to stain the cells for 30 min at room temperature in the dark, followed by analysis via flow cytometry (BD FACS Aria III).
Statistics & reproducibility
All the statistical analyses were performed with GraphPad Prism 7 software. The RT‒PCR data, immunofluorescence staining data, and OCR/ECAR data were statistically analyzed via Student’s t test, and the graphs show the means ± SDs. Metabolomic data were statistically analyzed via Welch’s two-sample t test. For the animal studies, the experiments were replicated across multiple batches. The mice were randomized into groups and treated according to the experimental design. The animal study data were statistically analyzed via Student’s t test, and the graphs show the means ± SDs. No statistical methods were used to predetermine sample sizes, but the group sizes used were similar to those commonly applied in mouse studies. No data were excluded from the analyses. The detailed methods and p values used to assess statistical significance are described in the Figure legends and detailed descriptions of the methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are provided in the article. Source data are provided with this paper. All RNA-seq data generated in this study have been deposited in the GSE database under accession code GSE248719. All LC-MS Metabolomics data used in this study are available in the MassIVE database under accession code MassIVE MSV000098820 [https://doi.org/10.25345/C5XP6VG8J]. Source data are provided with this paper.
References
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the warburg effect: The metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Xiao, Y. et al. Emerging therapies in cancer metabolism. Cell Metab. 35, 1283–1303 (2023).
Liberti, M. V. & Locasale, J. W. The warburg effect: How does it benefit cancer cells?. Trends Biochemical Sci. 41, 211–218 (2016).
Yoo, H. C. et al. A Variant of SLC1A5 is a mitochondrial glutamine transporter for metabolic reprogramming in cancer cells. Cell Metab. 31, 267–283.e212 (2020).
Stine, Z. E. & Dang, C. V. Glutamine skipping the Q into mitochondria. Trends Mol. Med. 26, 6–7 (2020).
Shen, Y.-A. et al. Inhibition of glutaminolysis in combination with other therapies to improve cancer treatment. Curr. Opin. Chem. Biol. 62, 64–81 (2021).
Wang, Z. et al. Targeting glutaminolysis: New perspectives to understand cancer development and novel strategies for potential target therapies. Front Oncol. 10, 589508 (2020).
Matés, J. M., Di Paola, F. J., Campos-Sandoval, J. A., Mazurek, S. & Márquez, J. Therapeutic targeting of glutaminolysis as an essential strategy to combat cancer. Semin. Cell Developmental Biol. 98, 34–43 (2020).
Jin, J., Byun, J.-K., Choi, Y.-K. & Park, K.-G. Targeting glutamine metabolism as a therapeutic strategy for cancer. Exp. Mol. Med. 55, 706–715 (2023).
Tannir, N. M. et al. CANTATA: Primary analysis of a global, randomized, placebo (Pbo)-controlled, double-blind trial of telaglenastat (CB-839) + cabozantinib versus Pbo + cabozantinib in advanced/metastatic renal cell carcinoma (mRCC) patients (pts) who progressed on immune checkpoint inhibitor (ICI) or anti-angiogenic therapies. J. Clin. Oncol. 39, 4501–4501 (2021).
Encarnación-Rosado, J. et al. Targeting pancreatic cancer metabolic dependencies through glutamine antagonism. Nat. Cancer 5, 85–99 (2023).
Schulte, M. L. et al. Pharmacological blockade of ASCT2-dependent glutamine transport leads to antitumor efficacy in preclinical models. Nat. Med. 24, 194–202 (2018).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pierce, B. G. et al. ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Bowers K. J., et al. Scalable algorithms for molecular dynamics simulations on commodity clusters. In: Proceedings of the 2006 ACM/IEEE conference on Supercomputing). Association for Computing Machinery (2006).
Gandia, J. et al. Detection of higher-order G protein-coupled receptor oligomers by a combined BRET-BiFC technique. FEBS Lett. 582, 2979–2984 (2008).
Shin, E. et al. Mitochondrial glutamate transporter SLC25A22 uni-directionally export glutamate for metabolic rewiring in radioresistant glioblastoma. Int J. Biol. Macromol. 253, 127511 (2023).
Zhou, Q. et al. Targeting of SLC25A22 boosts the immunotherapeutic response in KRAS-mutant colorectal cancer. Nat. Commun. 14, 4677 (2023).
Zielewicz, L. & Grewer, C. Genetically encoded halide sensor-based fluorescent assay for rapid screening of glutamate transport and inhibition. ACS Sens. 4, 2358–2366 (2019).
Ashok, Y., Nanekar, R. & Jaakola, V.-P. Defining thermostability of membrane proteins by western blotting. Protein Eng., Des. Selection 28, 539–542 (2015).
Mullen, A. R. et al. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature 481, 385–388 (2012).
McKeehan, W. L. Glycolysis, glutaminolysis and cell proliferation. Cell Biol. Int Rep. 6, 635–650 (1982).
Medina, M. A. & Núñez de Castro, I. Glutaminolysis and glycolysis interactions in proliferant cells. Int J. Biochem 22, 681–683 (1990).
Mirveis, Z., Patil, N. & Byrne, H. J. Experimental and computational investigation of the kinetic evolution of the glutaminolysis pathway and its interplay with the glycolysis pathway. FEBS Open Bio 14, 1247–1263 (2024).
Gaude, E. et al. NADH shuttling couples cytosolic reductive carboxylation of glutamine with glycolysis in cells with mitochondrial dysfunction. Mol. Cell 69, 581–593.e587 (2018).
McCauley, R., Kong, S.-E. & Hall, J. Review: Glutamine and nucleotide metabolism within enterocytes. J. Parenter. Enter. Nutr. 22, 105–111 (1998).
Boza, J. J. et al. Role of glutamine on the de novo purine nucleotide synthesis in Caco-2 cells. Eur. J. Nutr. 39, 38–46 (2000).
Wu, R. et al. De novo synthesis and salvage pathway coordinately regulate polyamine homeostasis and determine T cell proliferation and function. Sci. Adv. 6, eabc4275 (2020).
Wade, A. M. & Tucker, H. N. Antioxidant characteristics of L-histidine. J. Nutritional Biochem. 9, 308–315 (1998).
Krifka, S. et al. The influence of glutathione on redox regulation by antioxidant proteins and apoptosis in macrophages exposed to 2-hydroxyethyl methacrylate (HEMA). Biomaterials 33, 5177–5186 (2012).
Arrigoni, O. & De Tullio, M. C. Ascorbic acid: much more than just an antioxidant. Biochimica et. Biophysica Acta (BBA) - Gen. Subj. 1569, 1–9 (2002).
Leonardi R., Jackowski S. Biosynthesis of Pantothenic Acid and Coenzyme A. EcoSal Plus 2, https://doi.org/10.1128/ecosalplus.1123.1126.1123.1124 (2007).
Kreuzaler, P. et al. Vitamin B5 supports MYC oncogenic metabolism and tumor progression in breast cancer. Nat. Metab. 5, 1870–1886 (2023).
Fenech, M. et al. Low intake of calcium, folate, nicotinic acid, vitamin E, retinol, β-carotene and high intake of pantothenic acid, biotin and riboflavin are significantly associated with increased genome instability—results from a dietary intake and micronucleus index survey in South Australia. Carcinogenesis 26, 991–999 (2005).
Varghese, S. et al. The Glutaminase Inhibitor CB-839 (Telaglenastat) Enhances the Antimelanoma Activity of T-Cell–Mediated Immunotherapies. Mol. Cancer Therapeutics 20, 500–511 (2021).
Edwards D. N. et al. Selective glutamine metabolism inhibition in tumor cells improves antitumor T lymphocyte activity in triple-negative breast cancer. J. Clin. Invest. 131, (2021).
Sun, Q. et al. Immune checkpoint therapy for solid tumours: clinical dilemmas and future trends. Signal Transduct. Target. Ther. 8, 320 (2023).
Brahmer, J. R. et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N. Engl. J. Med. 366, 2455–2465 (2012).
Byun, J.-K. et al. Inhibition of glutamine utilization synergizes with immune checkpoint inhibitor to promote antitumor immunity. Mol. Cell 80, 592–606.e598 (2020).
Tang, Y. et al. Simultaneous glutamine metabolism and PD-L1 inhibition to enhance suppression of triple-negative breast cancer. J. Nanobiotechnology 20, 216 (2022).
Reid, M. A., Dai, Z. & Locasale, J. W. The impact of cellular metabolism on chromatin dynamics and epigenetics. Nat. Cell Biol. 19, 1298–1306 (2017).
Schvartzman, J. M., Thompson, C. B. & Finley, L. W. S. Metabolic regulation of chromatin modifications and gene expression. J. Cell Biol. 217, 2247–2259 (2018).
Katt, W. P., Lukey, M. J. & Cerione, R. A. A tale of two glutaminases: homologous enzymes with distinct roles in tumorigenesis. Future Med Chem. 9, 223–243 (2017).
Pochini, L., Scalise, M., Galluccio, M. & Indiveri, C. Membrane transporters for the special amino acid glutamine: structure/function relationships and relevance to human health. Front Chem. 2, 61 (2014).
Castro-Rodríguez, V. et al. Redundancy and metabolic function of the glutamine synthetase gene family in poplar. BMC Plant Biol. 15, 20 (2015).
Yoo, H. C., Sung, Y., Yu, Y. C. & Han, J. M. Glutamine reliance in cell metabolism. Exp. Mol. Med. 52, 1496–1516 (2020).
Hassanein, M. et al. Targeting SLC1a5-mediated glutamine dependence in non-small cell lung cancer. Int. J. Cancer 137, 1587–1597 (2015).
Choudhury, M. et al. Targeting Pulmonary Fibrosis by SLC1A5-Dependent Glutamine Transport Blockade. Am. J. Respiratory Cell Mol. Biol. 69, 441–455 (2023).
Bröer, A. et al. Ablation of the ASCT2 (SLC1A5) gene encoding a neutral amino acid transporter reveals transporter plasticity and redundancy in cancer cells. J. Biol. Chem. 294, 4012–4026 (2019).
Han, T. et al. A novel glutaminase inhibitor-968 inhibits the migration and proliferation of non-small cell lung cancer cells by targeting EGFR/ERK signaling pathway. Oncotarget 8, 28063–28073 (2017).
Hwang, Y. et al. Co-inhibition of glutaminolysis and one-carbon metabolism promotes ROS accumulation leading to enhancement of chemotherapeutic efficacy in anaplastic thyroid cancer. Cell Death Dis. 14, 515 (2023).
Gao, M., Monian, P., Quadri, N., Ramasamy, R. & Jiang, X. Glutaminolysis and Transferrin Regulate Ferroptosis. Mol. Cell 59, 298–308 (2015).
Kremer, D. M. et al. GOT1 inhibition promotes pancreatic cancer cell death by ferroptosis. Nat. Commun. 12, 4860 (2021).
Okabe, S., Tanaka, Y., Moriyama, M. & Gotoh, A. Inhibition of glutaminolysis alone and in combination with HDAC inhibitor has anti-myeloma therapeutic effects. Cancer Drug Resist 7, 25 (2024).
Villar, V. H. et al. mTORC1 inhibition in cancer cells protects from glutaminolysis-mediated apoptosis during nutrient limitation. Nat. Commun. 8, 14124 (2017).
Fitzsimmons, T. S. et al. Immune checkpoint inhibitors efficacy across solid cancers and the utility of PD-L1 as a biomarker of response: a systematic review and meta-analysis. Front. Med. 10, (2023).
Li, B., Chan, H. L. & Chen, P. Immune checkpoint inhibitors: Basics and challenges. Curr. Med Chem. 26, 3009–3025 (2019).
Birnbaum, D. J. et al. Prognostic value of PDL1 expression in pancreatic cancer. Oncotarget 7, 71198–71210 (2016).
Luo, Y. et al. Effects of lactate in immunosuppression and inflammation: Progress and prospects. Int. Rev. Immunol. 41, 19–29 (2022).
Tran L. C., Özdemir B. C. & Berger M. D. The role of immune checkpoint inhibitors in metastatic pancreatic cancer: current state and outlook. Pharmaceuticals (Basel) 16, (2023).
Wieckowski, M. R., Giorgi, C., Lebiedzinska, M., Duszynski, J. & Pinton, P. Isolation of mitochondria-associated membranes and mitochondria from animal tissues and cells. Nat. Protoc. 4, 1582–1590 (2009).
Hovius, R., Lambrechts, H., Nicolay, K. & de Kruijff, B. Improved methods to isolate and subfractionate rat liver mitochondria. Lipid composition of the inner and outer membrane. Biochimica et. Biophysica Acta (BBA) - Biomembranes 1021, 217–226 (1990).
Karbowski, M., Norris, K. L., Cleland, M. M., Jeong, S. Y. & Youle, R. J. Role of bax and bak in mitochondrial morphogenesis. Nature 443, 658–662 (2006).
Hedley, D. W. & Chow, S. Evaluation of methods for measuring cellular glutathione content using flow cytometry. Cytometry 15, 349–358 (1994).
Park, J. et al. YAP and AP-1 cooperate to initiate pancreatic cancer development from ductal cells in mice. Cancer Res. 80, 4768–4779 (2020).
Choi, W. et al. Establishment of patient-derived organoids using ascitic or pleural fluid from cancer patients. Cancer Res Treat. 55, 1077–1086 (2023).
Kang, S., Lee, M. R., Choi, W., Kong, S. Y. & Kim, Y. H. Protocol for generation and utilization of patient-derived organoids from multimodal specimen. STAR Protoc. 6, 104039 (2025).
Acknowledgements
This work was supported by Health Fellowship Foundation (Y.S.). This work was supported by grants from the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Korean government (MSIT) (2020M3A9A5036362, RS-2023-00219297, RS-2025-00573098, J.M.H.). This research was supported by the Korea Drug Development Fund, funded by the Ministry of Science and ICT, the Ministry of Trade, Industry, and Energy, and the Ministry of Health and Welfare (RS-2024-00334337, J.M.H.).
Author information
Authors and Affiliations
Contributions
Y.S., conception and experimental design, collection and analysis of the data, and manuscript writing; Y.C.Y., conception and experimental design, and analysis of the data; M.L., experimental design and collection and analysis of the data; S.L., Y.L., M.K., D.K., and A.P., collection and analysis of the data; J.C., experimental design, analysis of the data, and supervision; D.S.M., supervision; K.K., W.N., Y.H.K., experimental design, analysis of the data, and supervision; S.M.W., experimental design and supervision; N.D.K., collection and analysis of the data and supervision; H.C.Y., conception of the study, collection and analysis of the data and supervision; J.M.H., supervision and final approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sung, Y., Yu, Y.C., Lee, M. et al. Targeting cancer glutamine dependency with a first-in-class inhibitor of the mitochondrial glutamine transporter SLC1A5_var. Nat Commun 16, 9690 (2025). https://doi.org/10.1038/s41467-025-64730-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64730-2