Abstract
Eosinophils are major effector cells in type 2 immune responses, contributing to host defense and allergic diseases. They also contribute to maintaining tissue homeostasis by regulating various immune cell types, including neutrophils. Here we show that eosinophils directly associate with neutrophils in the lungs of asthma-induced mice. Eosinophil-specific deficiency of the short-chain fatty acid receptor, GPR43, results in hyperactivation of eosinophils and increases the expression of neutrophil chemoattractants and PECAM-1, thereby enhancing the interaction between eosinophils and neutrophils. This interaction exposes neutrophils to eosinophil-derived IL-4 and GM-CSF, which induce the conversion of conventional neutrophils into more pathogenic, Siglec-Fhi neutrophils capable of enhancing Th17 cell differentiation and aggravating asthma symptoms in mouse models. Our results thus implicate GPR43 as a critical regulator of eosinophils, and describe eosinophil-mediated modulation of neutrophil differentiation and function.
Similar content being viewed by others
Introduction
Eosinophils are multifunctional granulocytes that are primarily recruited during type 2 immune responses. Once activated, they degranulate and release pre-synthesized cationic proteins, cytokines, chemokines, and lipid mediators to protect the host from infectious agents. However, this potent response can also cause tissue damage in allergic diseases1,2,3. Moreover, eosinophils are constitutively present in several peripheral organs, where they maintain tissue homeostasis4,5. For example, eosinophils in adipose tissue are the major source of IL-4, which sustains the alternatively activated macrophages that control glucose metabolism6. Intestinal eosinophils not only dominate the myeloid cells in the gastrointestinal tract but also prevent inflammation by suppressing Th1 and Th17 cells7,8. In addition, eosinophils in the lung protect it from allergen-induced airway inflammation by suppressing dendritic cells (DCs)9. Notably, these local eosinophils markedly differ from newly infiltrating inflammatory eosinophils in terms of morphology and functions9. Thus, eosinophils have diverse functions and make complex interactions with other immune cell types.
The development, migration, and activation of eosinophils are regulated by common β chain cytokines, such as IL-3, IL-5, and GM-CSF1,10. In addition, recent studies suggest that tissue environmental factors and nutrient-derived signals can control eosinophil functions. For example, aryl hydrocarbon receptor signaling in intestinal eosinophils induces transcriptional programming that permits their adaptation to the local environment, extends their life span, and confers their unique characteristics11,12. Similarly, the programming of intestinal eosinophils is also driven directly by retinoic acid: this signaling is specifically required for maintaining a villus-resident subpopulation of intestinal eosinophils13. Moreover, the binding of neuromedin-U, a neuropeptide produced in the gut, to its receptor on eosinophils promotes their intestinal accumulation under normal conditions and induces goblet cell differentiation during helminth infection14.
An interesting nutrient-derived factor is short-chain fatty acids (SCFAs), which are mainly produced by the gut microbiota via dietary fiber fermentation and circulated throughout the body15,16,17. SCFAs are recognized by GPR43, which is expressed by many non-immune and immune cells, including eosinophils18,19. Significantly, numerous studies show that SCFAs exert anti-inflammatory effects, and the lack of GPR43 signaling aggravates inflammatory diseases such as food allergy, colitis, rheumatoid arthritis, and asthma18,20,21. Various GPR43-bearing cells have been found to mediate this anti-inflammatory activity, including DCs, neutrophils, type 3 innate lymphoid cells, and intestinal epithelial cells18,20,21,22,23. However, whether eosinophils also have GPR43-mediated anti-inflammatory activities is unknown.
Here, we show with mice lacking GPR43 in eosinophils that GPR43 prevents eosinophils from becoming hyperactivated in allergic airway inflammation. Interestingly, this reduces direct eosinophil interactions with neutrophils in the lung, thereby suppressing the emergence of a pathogenic, Th17-promoting neutrophil subset. Thus, this study shows that GPR43 regulates eosinophil functions, and that eosinophils have a previously little-known ability to directly modulate neutrophil differentiation and activities in lung inflammation.
Results
GPR43 deficiency in eosinophils exacerbates asthma in association with increased lung neutrophils and Th17 cells
GPR43 is encoded by free fatty acid receptor 2 (FFAR2). As previously observed14,24,25,26,27,28, FFAR2 is constitutively expressed at high levels in eosinophils from various organs in humans and mice. By contrast, other FFAR genes (FFAR1 and FFAR3) are poorly expressed (Supplementary Fig. 1a, b). Ffar2 is also expressed in eosinophils in the lungs of mice in which asthma was induced with house dust mite extracts (HDM) (Supplementary Fig. 1c). Moreover, publicly available single-cell RNA-sequencing (scRNA-seq) data from patients with eosinophilic esophagitis show that eosinophils in the inflamed esophagus and duodenum highly express FFAR229(Supplementary Fig. 1d, e). Given that GPR43 has well-known anti-inflammatory functions and eosinophils drive airway inflammation in asthma, we asked whether GPR43 regulates eosinophils in asthma.
To address this, GPR43 was specifically deleted in eosinophils by crossing Gpr43fl/fl mice with Epx-Cre mice, thus generating EpxCre/+Gpr43fl/fl mice. Eosinophil-specific Cre-mediated recombination was confirmed with EpxCre/+Rosa26-eYFPfl/+ reporter mice. As reported previously7, YFP was exclusively expressed in eosinophils, of which 60-70% expressed YFP (Supplementary Fig. 1f–h, 15). Notably, despite the apparently incomplete recombination induced by Epx-Cre in the reporter mice, GPR43 mRNA was not detected in eosinophils from EpxCre/+Gpr43fl/fl mice (Supplementary Fig. 1i). Moreover, acetate, an endogenous GPR43 ligand, did not induce Ca2+ influx in eosinophils from EpxCre/+Gpr43fl/fl mice (Supplementary Fig. 1j). Thus, GPR43 signaling is efficiently abrogated in GPR43-deficient eosinophils.
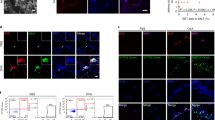
EpxCre/+Gpr43fl/fl and control (EpxCre/+) mice were then subjected to allergic airway inflammation induced by house dust mite extracts (Fig. 1a). To minimize variability arising from hormonal fluctuations, all experiments were initially conducted using male mice. Compared to EpxCre/+ mice, EpxCre/+Gpr43fl/fl mice had significantly more immune cells, including eosinophils and neutrophils, in the bronchoalveolar space (Fig. 1b and Supplementary Fig. 16a). Histological analysis revealed more inflammation, mucus accumulation, and fibrosis in the lungs of EpxCre/+Gpr43fl/fl mice (Fig. 1c). Accordingly, EpxCre/+Gpr43fl/fl mice showed worse airway hyperresponsiveness compared to control mice (Fig. 1d). Thus, GPR43 deletion in eosinophils aggravated airway inflammation and asthma symptoms. The bronchoalveolar lavage fluid (BALF) from asthmatic EpxCre/+Gpr43fl/fl mice also contained higher type 2 cytokine levels (IL-4, IL-5, and IL-13) (Fig. 2a) and their lungs bore more GATA3- and type 2 cytokine-expressing CD4 T cells (Fig. 2b–d and Supplementary Fig. 16b–d). Notably, the BALF from EpxCre/+Gpr43fl/fl mice also contained significantly more IL-17A, and the lungs bore higher frequencies of RORγt- and IL-17A-expressing CD4 T cells (Fig. 2a–d). These results show that GPR43 deficiency in eosinophils enhances both Th2 and Th17 immune responses in asthma.
a Experimental scheme of HDM-induced asthma. b Total immune cell, eosinophil, and neutrophil counts in BALF. EpxCre/+ (n = 5), EpxCre/+Gpr43fl/fl (n = 6). Data are representative of more than three independent experiments. c Histological analysis of immune cell infiltration (H&E), mucus production (PAS), and fibrosis (Sirius red) in asthmatic lungs. Scale bar, 100 µm. Data are representative of two independent experiments. d Airway resistance. EpxCre/+ (n = 9), EpxCre/+Gpr43fl/fl (n = 9). Data pooled from two independent experiments. Data are presented as mean ± s.e.m. Statistical analysis was performed using an unpaired two-tailed Student’s t-test (b) or two-way ANOVA (d).
a Cytokine levels in BALF. b Percentages of CD4 T cells expressing each transcription factor in asthmatic lungs. c Percentages of CD4 T cells producing each cytokine in asthmatic lungs. EpxCre/+ (n = 11), EpxCre/+Gpr43fl/fl (n = 12). Data in (a–c) were pooled from two independent experiments. d Cytokine levels in culture supernatants of asthmatic lung cells stimulated with anti-CD3 and anti-CD28. EpxCre/+ (n = 5), EpxCre/+Gpr43fl/fl (n = 6). Data in (d) are representative of three independent experiments. Data are presented as mean ± s.e.m. Statistical analysis was performed using an unpaired two-tailed Student’s t-test.
Pathogenic Siglec-Fhi neutrophils accumulate in the asthmatic lungs of Epx Cre/+ Gpr43 fl/fl mice
To comprehensively examine the effect of eosinophil-specific GPR43 deletion on the lung immune cell landscape during airway inflammation, CD45+ cells from HDM-induced asthmatic lungs of EpxCre/+ and EpxCre/+Gpr43fl/fl mice were subjected to scRNA-seq. Unsupervised cell clustering revealed nine groups bearing conventional immune cell type markers (Fig. 3a and Supplementary Fig. 2a). Notably, eosinophils were not detected. This was also observed in publicly available scRNA-seq datasets of healthy and asthmatic lung30,31 and probably reflects high RNase levels in eosinophils30,31,32. The most notable difference between EpxCre/+ and EpxCre/+Gpr43fl/fl mice was the neutrophils: they were highly enriched in EpxCre/+Gpr43fl/fl mice (Fig. 3b). Fine clustering of T cells showed that GPR43 deletion in eosinophils increased Gata3-, Il4-, Il5-, and Il13-expressing Th2 cells and Rorc- and Il17-expressing Th17 cells (Supplementary Fig. 2b–d). These findings are consistent with our flow cytometry results (Fig. 2b, c), despite the lack of statistical analysis in this transcriptomic data.
a–e CD45+ cells from three mice were pooled per group and subjected to single-cell RNA sequencing. UMAP plots of immune cells in asthmatic lungs of EpxCre/+ and EpxCre/+Gpr43fl/fl mice (a). Ratio of the immune cell types in each group (b). UMAP plots of neutrophils showing two distinct clusters (c). Dot plot of marker genes for each neutrophil cluster (d). Ratios of each group in the individual neutrophil clusters (e). f Percentages of Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi cells among neutrophils in asthmatic lungs. EpxCre/+ (n = 6), EpxCre/+Gpr43fl/fl (n = 8). g Percentages of Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi cells among neutrophils in blood, lung, and BALF. Steady-state EpxCre/+ (n = 5), asthmatic EpxCre/+ (n = 5), asthmatic EpxCre/+Gpr43fl/fl (n = 4). h Flow cytometric analysis of neutrophils in steady-state lung, asthmatic lung, and asthmatic BALF after intravenous administration of anti-CD45 antibodies. i, Representative histograms showing Siglec-F and PECAM-1 expression in CD45 i.v.- and CD45 i.v.+ neutrophils from HDM-induced asthmatic lungs of EpxCre/+ and EpxCre/+Gpr43fl/fl mice. Data in (f–i) are representative of more than three independent experiments. Data are presented as mean ± s.e.m. Statistical analysis was performed using an unpaired two-tailed Student’s t-test (f, g). ns, not significant.
Subclustering of all neutrophils in our scRNA-seq datasets resulted in two distinct clusters (Fig. 3a, c). Both expressed Cxcr2 and S100a8 but other genes typically associated with neutrophils (e.g. Mmp8 and Retnlg) demonstrated dramatic downregulation in Cluster 1. Moreover, several other genes were almost exclusively expressed in Cluster 1, namely, Siglecf, Pecam1, Havcr2, Scimp, Gpr84, and Tmem273 (Fig. 3d and Supplementary Fig. 3a). Siglec-F was particularly interesting because, while it often serves as an eosinophil marker, recent studies have identified Siglec-F-expressing neutrophil subsets in diverse diseases, including lung adenocarcinoma, myocardial infarction, and diesel exhaust particle (DEP)-induced lung inflammation33,34,35. Transcriptomic analyses revealed that many genes upregulated in Siglec-Fhi neutrophils (relative to Siglec-Flo neutrophils) in these conditions were also highly enriched in our Cluster 1 neutrophils, suggesting potential functional similarities (Supplementary Fig. 3b). In DEP-induced lung inflammation, Siglec-Fhi neutrophils actively produced neutrophil extracellular traps (NETs) and promoted the accumulation of type 2 cytokine-producing cells in the lung35. Moreover, neutrophils expressing Siglec-8 (the human paralog of murine Siglec-F) are present in the sputum of asthma patients, and their frequency in peripheral blood positively correlates with asthma severity35. These observations suggest that the Siglec-F-expressing neutrophils enriched in the asthmatic lungs of EpxCre/+Gpr43fl/fl mice may also be pathogenic.
Neutrophils in Cluster 0 (Siglec-Flo) were derived from both EpxCre/+ and EpxCre/+Gpr43fl/fl mice at approximately a 1:1 ratio, whereas the majority of Cluster 1 (Siglec-Fhi) neutrophils originated from EpxCre/+Gpr43fl/fl mice (Fig. 3e). This enrichment of Siglec-F-expressing neutrophils in asthmatic lungs of EpxCre/+Gpr43fl/fl mice was confirmed by flow cytometry (Fig. 3f and Supplementary Fig. 16e). Because Pecam1 was among the most highly upregulated genes in Cluster 1 neutrophils compared to Cluster 0 neutrophils (Fig. 3d and Supplementary Fig. 3a), we also examined PECAM-1 protein expression. PECAM-1-expressing neutrophils were increased in the asthmatic lungs of EpxCre/+Gpr43fl/fl mice relative to EpxCre/+ mice, and a large proportion of Siglec-F-expressing neutrophils co-expressed PECAM-1 (Fig. 3f). These neutrophils expressing either Siglec-F or PECAM-1 (hereafter referred to simply as Siglec-Fhi neutrophils) were almost exclusively found in asthmatic lungs and BALF, whereas few were present in healthy lungs or in the peripheral blood of either healthy or asthmatic mice (Fig. 3g). Additionally, the frequencies of Siglec-Fhi neutrophils were significantly higher in the BALF than in the lungs (Fig. 3g). Thus, Siglec-Fhi neutrophils likely differentiate from Siglec-Flo neutrophils in the lung during airway inflammation and infiltrate the airway more efficiently. This was supported by in vivo labeling of circulating neutrophils with intravenously injected CD45 antibodies just before sacrifice: none of Siglec-Fhi neutrophils in the asthmatic lungs and BALF were labeled with anti-CD45 antibodies (Fig. 3h, i). Thus, during airway inflammation, Siglec-Fhi neutrophils are exclusively localized in the lung tissue and exudate, but not in its microvasculature36.
We also examined the role of GPR43 in eosinophils using an OVA-induced asthma model. Similar to the HDM-induced asthma model, eosinophil-specific deletion of GPR43 exacerbated OVA-induced airway inflammation, which was accompanied by enhanced Th17 responses and increased frequencies of Siglec-Fhi neutrophils in the lungs (Supplementary Fig. 4), indicating that this phenomenon is not restricted to a specific model of airway inflammation. Furthermore, global GPR43 knockout mice also exhibited more severe HDM-induced airway inflammation, characterized by simultaneous increases in Th2 and Th17 responses as well as elevated proportion of Siglec-Fhi neutrophils in the lungs (Supplementary Fig. 5). Under steady-state conditions, the abundance of Siglec-Fhi neutrophils, as well as Th2 and Th17 cells in the lung, was minimal and comparable between EpxCre/+ and EpxCre/+Gpr43fl/fl mice (Supplementary Fig. 6).
GPR43-deficient eosinophils directly induce the differentiation of Siglec-Fhi neutrophils
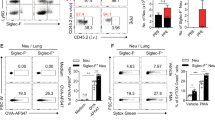
We next asked how the lack of GPR43 in eosinophils caused the emergence of Siglec-Fhi neutrophils during airway inflammation. First, we compared control (EOSCtrl) and GPR43-deficient eosinophils (EOS△GPR43) from the asthmatic lungs of EpxCre/+ and EpxCre/+Gpr43fl/fl mice, respectively. EOS△GPR43 were highly vacuolized and exhibited disorganized nuclear shapes (Fig. 4a–c). Eosinophils from patients with severe asthma also have these features37, which suggests that EOS△GPR43 were hyperactivated. The transcriptomic analysis by bulk-RNA sequencing revealed that activated eosinophils expressed multiple chemokines (Supplementary Fig. 7a and Supplementary Data 1). Among them, Ccl6 and Cxcl2, both encoding potent neutrophil chemoattractants38,39, were the two most highly expressed chemokine genes, suggesting that eosinophils may directly recruit neutrophils in asthmatic lungs. ELISA of culture supernatants from asthmatic lung eosinophils showed that EOS△GPR43 produced significantly higher levels of CCL6 and CXCL2 than EOSCtrl, indicating a greater capacity to attract neutrophils (Fig. 4d, e and Supplementary Fig. 17a). In addition, production of CCL24, a representative eotaxin and another highly expressed chemokine in asthmatic lung eosinophils, was also elevated in EOS△GPR43 cells compared with EOSCtrl, suggesting that the increased eosinophil numbers observed in EpxCre/+Gpr43fl/fl mice may be, at least in part, mediated by eosinophil-derived signals (Fig. 4f).
a Representative images of H&E-stained asthmatic lung eosinophils from EpxCre/+ and EpxCre/+Gpr43fl/fl mice. Scale bar, 10 µm. b Representative images of propidium iodide-stained asthmatic lung eosinophils. Scale bar, 10 µm. c Percentages of asthmatic lung eosinophils showing vacuolation (left) and disorganized nuclei (right). Percentages were calculated from three independent H&E images for each group. Total eosinophil numbers are indicated in the centers of the donut plots. d CCL6 levels in culture supernatants of asthmatic lung eosinophils. EOSCtrl (n = 5), EOS△GPR43 (n = 4). e CXCL2 levels in culture supernatants of asthmatic lung eosinophils after PMA/Ionomycin stimulation. EOSCtrl (n = 4), EOS△GPR43 (n = 5). f CCL24 levels in culture supernatants of asthmatic lung eosinophils. EOSCtrl (n = 6), EOS△GPR43 (n = 6). g Neutrophil migration assay scheme. Created in BioRender. Yu, J. (2025) https://BioRender.com/lohw7oc. h Neutrophil counts within 100x HPF images of the bottom plates. Asthmatic lung eosinophils were pooled from three mice per group for sorting, and assays were performed in technical replicates. NEU (n = 3), NEU + EOSCtrl (n = 3), NEU + EOS△GPR43 (n = 4). i Representative images showing CTV-labeled neutrophils migrated toward eosinophils in the bottom plate. Scale bar, 200 µm. j Images of eosinophil-neutrophil co-culture. Scale bar, 100 µm. k Immunofluorescence images of eosinophil-neutrophil co-culture. Scale bar: 100 µm. l Percentages of eosinophils associated with neutrophils (top) and neutrophils associated with eosinophils (bottom). Percentages were calculated from four and three independent fluorescence images for the EOSCtrl and EOS△GPR43 groups, respectively. m–o Percentages of Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi cells among neutrophils. Bone marrow neutrophils were co-cultured with either steady-state eosinophils (n = 5) or asthmatic lung eosinophils (n = 9) (m); with EOSCtrl (n = 6) or EOS△GPR43 (n = 5) (n); or with EOSCtrl in the presence or absence of 0.4 μm transwell (n = 6) (o). p gMFI of PECAM-1 in eosinophils from steady-state (n = 5) and asthmatic lungs (n = 5). q gMFI of PECAM-1 in asthmatic lung eosinophils. EpxCre/+ (n = 10), EpxCre/+Gpr43fl/fl (n = 10). r Percentages of Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi cells among neutrophils after co-culture with eosinophils in the presence or absence of anti-PECAM-1. EOSCtrl (n = 10), EOS△GPR43 (n = 9). Data in (m, q, r) were pooled from two independent experiments. Data are representative of more than two independent experiments. Data are presented as mean ± s.e.m. Statistical analysis was performed using one-way ANOVA (h, m–o), two-way ANOVA with Tukey’s multiple comparison test (r), or unpaired two-tailed Student’s t-test (d–f, p, q). NEU: neutrophils.
Next, we directly tested whether asthmatic lung eosinophils could attract neutrophils. When EOSCtrl or EOS△GPR43 were cultured with bone marrow-derived neutrophils in a transwell system (Fig. 4g and Supplementary Fig. 17b), neutrophil migration was enhanced by both eosinophils, but especially by EOS△GPR43 (Fig. 4h, i). Moreover, unlike neutrophils cultured alone, neutrophils co-cultured with EOSCtrl or EOS△GPR43 formed aggregates, especially with EOS△GPR43 (Fig. 4j). Immunofluorescence imaging of the aggregates revealed direct contact between neutrophils and eosinophils, especially with EOS△GPR43 (Fig. 4k, l). Immunofluorescence imaging of the asthmatic lung of EpxCre/+ mice also showed physical interactions between the neutrophils and eosinophils, especially around the airways (Supplementary Fig. 7b). Thus, eosinophils in the asthmatic lung actively attract and make contact with neutrophils, and this interaction is augmented by GPR43 deficiency in eosinophils. Notably, reanalysis of publicly available transcriptomic data showed that when peripheral blood eosinophils from asthma patients were activated in vitro with IL-33, many neutrophil chemotaxis-related genes were upregulated, implying that human eosinophils can also recruit neutrophils (Supplementary Fig. 7c).
The physical association between eosinophils and neutrophils in asthmatic lungs led us to ask whether eosinophils directly caused Siglec-Flo neutrophils to differentiate into Siglec-Fhi neutrophils. Thus, bone marrow neutrophils, which are Siglec-Flo, were co-cultured with eosinophils from asthmatic or healthy lungs of control mice for 2 days. In the absence of eosinophils or in the presence of steady-state lung eosinophils, few Siglec-Fhi neutrophils arose (Fig. 4m and Supplementary Fig. 18). In contrast, eosinophils from asthmatic lungs markedly promoted Siglec-Fhi neutrophil differentiation, and this effect was further augmented by GPR43 deficiency in eosinophils (Fig. 4m, n). These findings suggest that eosinophils in asthmatic lungs directly drive the emergence of Siglec-Fhi neutrophils and that GPR43 deletion in eosinophils exacerbates this process.
Interestingly, eosinophil-mediated differentiation of Siglec-Fhi neutrophils was completely abolished when the two cell types were physically separated using a 0.4 μm pore transwell, indicating that direct cell–cell contact is essential (Fig. 4o). Based on this observation, we hypothesized that cell surface adhesion molecules mediating eosinophil–neutrophil interactions may contribute to Siglec-Fhi neutrophil differentiation. Notably, we found that asthma induction upregulated PECAM-1 expression in eosinophils, mirroring the upregulation observed in neutrophils (Fig. 4p). Furthermore, eosinophil PECAM-1 expression was higher in EOS△GPR43 than in EOSCtrl (Fig. 4q). PECAM-1 is an adhesion molecule that can mediate the binding of neutrophils to other cells in a homophilic manner40,41. To determine whether PECAM-1 participates in eosinophil-induced Siglec-Fhi neutrophil differentiation, EOS△GPR43 or EOSCtrl were co-cultured with bone marrow-derived neutrophils in the presence of PECAM-1-blocking antibody. The antibody reduced the ability of EOS△GPR43 to induce Siglec-Fhi neutrophil differentiation but did not inhibit EOSCtrl-induced Siglec-Fhi neutrophil differentiation (Fig. 4r). This suggests that (i) PECAM-1 proteins overexpressed on EOS△GPR43 contribute to the ability of these GPR43-deficient eosinophils to enhance Siglec-Fhi neutrophil differentiation, likely by strengthening eosinophil-neutrophil adhesion, but (ii) other mechanisms also mediate these interactions, making PECAM-1 dispensable for Siglec-Fhi neutrophil differentiation by EOSCtrl.
Next, we sought to identify the eosinophil-derived mediators that promote Siglec-Fhi neutrophil differentiation. Bone marrow neutrophils were stimulated with various cytokines or LPS. GM-CSF and, to a lesser extent, IL-33, TNF, and LPS, induced Siglec-F expression. By contrast, IL-4 and LPS, and to a lesser extent, IL-33, GM-CSF, and TGFβ, promoted PECAM-1 expression in neutrophils (Fig. 5a). We then combined the cytokines that most strongly stimulated Siglec-F (GM-CSF) and PECAM-1 (IL-4) expression: this combination increased the frequencies of Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi neutrophils in a highly synergistic manner (Fig. 5b). This efficient GM-CSF + IL-4-mediated induction of Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi neutrophils was also observed when neutrophils from healthy lungs were treated with these cytokines (Fig. 5c). Consistent with these findings, differentiation of neutrophils into Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi neutrophils was significantly inhibited by neutralizing antibodies against both IL-4 and GM-CSF, indicating that eosinophil-derived IL-4 and GM-CSF drive Siglec-Fhi-neutrophil differentiation (Fig. 5d). IL-4 secretion by asthmatic lung eosinophils was confirmed by ELISA (Supplementary Fig. 7d), whereas GM-CSF was detectable by intracellular cytokine staining but not by ELISA (Supplementary Fig. 7e,19). Unlike neutrophil chemokines, IL-4 and GM-CSF production was comparable between EOSCtrl and EOS△GPR43 (Supplementary Fig. 7d, e), suggesting that the enhanced capacity of GPR43-deficient asthmatic lung eosinophils to promote Siglec-Fhi-neutrophil differentiation is mainly attributable to their increased neutrophil recruitment and stronger physical interactions.
Shown are the percentages of Siglec-Fhi, PECAM-1hi, and Siglec-FhiPECAM-1hi neutrophils. a Bone marrow neutrophils stimulated with various cytokines or LPS (n = 5 for untreated and GM-CSF; n = 3 for IL-2, IL-12, and IL-33; n = 4 for all other groups). Data were pooled from five independent experiments. b Bone marrow neutrophils stimulated with IL-4, GM-CSF, or both cytokines (n = 3). Data were pooled from three independent experiments. c Steady-state lung neutrophils were stimulated with the combination of IL-4 and GM-CSF (n = 4). Data are representative of two independent experiments. d Bone marrow neutrophils were co-cultured with asthmatic lung eosinophils in the presence of anti-IL-4 plus anti-GM-CSF antibodies or isotype controls (n = 5). Data are representative of three independent experiments. Data are presented as mean ± s.e.m. Statistical analysis was performed using one-way ANOVA with Tukey’s multiple comparison test (a, b, d) and unpaired two-tailed Student’s t-test (c).
Siglec-Fhi neutrophils enhance Th17 differentiation
To better characterize Siglec-Fhi neutrophils in the HDM-induced asthmatic lung, Cluster 0 (Siglec-Flo) and Cluster 1 (Siglec-Fhi) neutrophil transcriptomes were further compared. Relative to Siglec-Flo neutrophils, Siglec-Fhi neutrophils demonstrated (i) strong upregulation of cytokine (e.g. Csf1, Tnf, Il1a), chemokine (Ccl3, Ccl4), degranulation (Cd63), and lipid-mediator synthesis (Ptgs1, Ltc4s) genes, and (ii) downregulation of classical neutrophil function genes (Lyz2, Mmp8, S100a8, S100a9, Ly6g, Retnlg) and a gene related to homing to secondary-lymphoid organs (Sell) (Fig. 6a). Gene set enrichment analysis also showed that ‘cytokine activity’, ‘TNF signaling pathway’, and ‘NF-kappa B signaling pathway’ genes were highly enriched in Siglec-Fhi neutrophils, as were genes that relate to rheumatoid arthritis and Coronavirus disease-COVID-19 (Fig. 6b and Supplementary Fig. 8). Moreover, Siglec-Fhi neutrophils demonstrated enrichment in pattern recognition receptor pathways such as ‘NOD-like receptor signaling pathway’ and ‘C-type lectin receptor signaling pathway’ along with’Phagosome’, ‘Lysosome’, ‘Ribosome’, ‘Oxidative phosphorylation’, and ‘Apoptosis’ genes (Supplementary Fig. 8). Thus, compared to conventional Siglec-Flo neutrophils, Siglec-Fhi neutrophils were more activated and likely performed more pro-inflammatory functions.
a Volcano plot displaying differential gene expression between Siglec-Flo and Siglec-Fhi neutrophils (FDR < 0.05, absolute logFC > 2 highlighted). b Gene set enrichment analysis of differentially expressed genes. c, d UMAP plots (c) and violin plots (d) for cytokine genes related to Th17 differentiation. e Correlation between the proportion of Siglec-Fhi neutrophils and RORγt+ or IL-17A+ CD4 T cells in asthmatic lungs. EpxCre/+ (n = 14), EpxCre/+Gpr43fl/fl (n = 15). Data were pooled from three independent experiments. f Experimental scheme for co-culture of neutrophils and naïve CD4 T cells under Th17 differentiation conditions. Created in BioRender. Yu, J. (2025) https://BioRender.com/j8a0ji3. Siglec-Flo and Siglec-Fhi neutrophils were prepared by culturing bone marrow neutrophils in the absence or presence of IL-4 and GM-CSF, respectively. g Percentages of RORγt+ and IL-17A+ CD4 T cells after the co-culture. Siglec-Flo neutrophils (n = 4), Siglec-Fhi neutrophils (n = 4). Data are representative of three independent experiments. Data are presented as mean ± s.e.m. Statistical analysis was performed using a two-sided permutation test with false discovery rate (FDR) correction for multiple comparison (b), the Wilcoxon rank-sum test with Benjamini–Hochberg correction (d), simple linear regression (e), and one-way ANOVA with Tukey’s multiple comparison test (g). ns, not significant.
Closer examination of cytokine expression in Siglec-Fhi neutrophils revealed notable increases in Il1a, Il23a, Tgfb1, and Tnf, whereas two other Th17-inducing cytokines were either expressed at similar levels in Siglec-Fhi and Siglec-Flo neutrophils (Il1b) or were poorly expressed in both (Il6) (Fig. 6c, d and Supplementary Fig. 9a). Greater expression of TNF and IL-1α in Siglec-Fhi neutrophils, along with high but comparable levels of IL-1β in Siglec-Fhi and Siglec-Flo neutrophils, was confirmed by ELISA (Supplementary Fig. 9b). Interestingly, of all immune cells in asthmatic lungs, neutrophils were largely the only producers of Il1a, Il23a, and Tnf (Supplementary Fig. 9a). Thus, Siglec-Fhi neutrophils may drive Th17 responses. Indeed, Siglec-Fhi-neutrophil frequencies in asthmatic lungs correlated positively with RORγt+ and IL-17A+ CD4 T cell frequencies (Fig. 6e). In contrast, the frequencies of Siglec-Flo neutrophils and dendritic cells showed no correlation with RORγt+ and IL-17A+ CD4 T cell frequencies (Supplementary Fig. 9c, d).
To determine whether Siglec-Fhi neutrophils can directly drive Th17 differentiation, we generated Siglec-Fhi neutrophils from Siglec-Flo neutrophils with IL-4 + GM-CSF (Fig. 6f). Like Siglec-Fhi neutrophils in asthmatic lungs, these in vitro-generated Siglec-Fhi neutrophils expressed high levels of Il1a, Il23a, and Tnf compared to Siglec-Flo neutrophils (Supplementary Fig. 9e). They also expressed more IL-1α and TNF, as determined by ELISA (Supplementary Fig. 9f). Thus, Siglec-Fhi neutrophils generated in vitro with cytokines closely resembled Siglec-Fhi neutrophils in asthmatic lungs. Importantly, co-culture with naïve CD4 T cells showed that Siglec-Fhi neutrophils significantly enhanced Th17 cell differentiation compared to Siglec-Flo neutrophils, as indicated by higher RORγt+ and IL-17A+ cell frequencies (Fig. 6f, g).
Eosinophils have been shown to modulate CD4 T cell differentiation directly or indirectly by regulating DC functions7,8,42,43. Because the frequencies of RORγt+ and IL-17A+ CD4 T cells in asthmatic lungs also positively correlated with eosinophil frequencies (Supplementary Fig. 9c, d), we next asked whether eosinophils, like Siglec-Fhi neutrophils, can directly augment Th17 differentiation. However, co-culturing naïve CD4 T cells with EOSCtrl or EOS△GPR43, with or without DCs, showed that while DCs increased Th17 differentiation, eosinophils could neither directly nor indirectly enhance it (Supplementary Fig. 10a, b). This is consistent with the comparable expression in EOSCtrl and EOS△GPR43 of antigen presentation- and Th17 differentiation-related genes (Supplementary Fig. 10c, d), as well as their similar levels of IL-1-receptor antagonist (IL-1Ra), which can suppress Th17 cell differentiation8 (Supplementary Fig. 10e). Moreover, DCs in asthmatic lungs from EpxCre/+ and EpxCre/+Gpr43fl/fl mice exhibited similar expression of antigen presentation-, Th17 activation-, and Th17 differentiation-related genes (Supplementary Fig. 10f). Collectively, these results suggest that lack of GPR43 in eosinophils induces their hyperreactivity in the asthmatic lung, leading to the generation of Siglec-Fhi neutrophils, which then can promote Th17 cell responses.
GPR43 in eosinophils is required for acetate-mediated suppression of Siglec-Fhi neutrophils in female mice
Given the well-documented influence of sex on asthma pathogenesis and severity44,45,46, we investigated whether female EpxCre/+Gpr43fl/fl mice also display exacerbated airway inflammation and increased frequencies of Siglec-Fhi neutrophils compared with female EpxCre/+ controls. When asthma was induced by HDM in male and female mice in parallel, females exhibited greater immune cell infiltration in the BALF and higher lung Siglec-Fhi neutrophil frequencies, indicating stronger overall immune responses than males (Supplementary Fig. 11a, b). Moreover, unlike males, female EpxCre/+ and EpxCre/+Gpr43fl/fl mice showed no differences in BALF immune cell counts or lung Siglec-Fhi neutrophil frequencies. Th2 and Th17 responses were also comparable between the two female genotypes (Supplementary Fig. 11c, d). Similarly, systemic GPR43 deficiency did not alter BALF immune cell counts, lung Siglec-Fhi neutrophil frequencies, and Th2 and Th17 responses in female mice (Supplementary Fig. 12).
Nonetheless, administration of acetate, an endogenous GPR43 agonist, during asthma induction in female mice reduced lung Siglec-Fhi neutrophil proportions in a GPR43-dependent manner, indicating that GPR43 in eosinophils can suppress Siglec-Fhi neutrophil differentiation in females under specific conditions such as elevated SCFA levels (Supplementary Fig. 13). To determine more directly whether GPR43 intrinsically regulates female eosinophil function without potential confounding effects of the in vivo environment, we analyzed chemokine production by bone marrow–derived eosinophils (BMDEs) from EpxCre/+ and EpxCre/+Gpr43fl/fl mice. Consistent with findings in male asthmatic lung eosinophils, GPR43 deficiency increased chemokine production in male BMDEs stimulated with IL-33 (Supplementary Fig. 14). Likewise, GPR43-deficient female BMDEs secreted higher levels of CCL6 and CCL24, although CXCL2 production was unaffected (Supplementary Fig. 14). Collectively, these results demonstrate that GPR43 intrinsically modulates eosinophil function in both sexes, but functional differences between control and GPR43-deficient female eosinophils can be masked in vivo by cell-extrinsic factors, especially under highly inflammatory conditions.
Discussion
GPR43 plays anti-inflammatory roles in various inflammatory diseases, and multiple cell types can mediate this activity. Here, we demonstrate that GPR43 also plays anti-inflammatory roles in eosinophils. Upon induction of allergic airway inflammation, eosinophils lacking GPR43 became hyperactive, leading to enhanced type 2 immune responses. This was reflected by increased frequencies of both eosinophils and Th2 cells, which together contributed to elevated type 2 cytokine levels. Our in vitro and in vivo observations further suggest that eosinophil hyperactivity also drove excessive differentiation of Siglec-Fhi neutrophils, which in turn increased Th17 cells in the lung and exacerbated airway inflammation.
Eosinophils interact with and control diverse immune cell types. However, whether eosinophils can regulate neutrophils, particularly directly, has been poorly studied47,48, likely because eosinophils and neutrophils play canonical roles in type 2 and type 3 immunity, respectively49. Our study showed that eosinophils can, in fact, directly regulate neutrophils in several ways. First, even normal lung eosinophils (with GPR43) from asthmatic mice could recruit neutrophils by producing neutrophil chemoattractants. Moreover, our re-analysis of publicly available bulk RNA-seq data showed that when human eosinophils are treated with the asthma-promoting cytokine IL-33, they express multiple neutrophil chemoattractants. This potential for direct neutrophil attraction by eosinophils has also been observed in other inflammatory settings50. Second, we observed that eosinophils interacted physically with neutrophils, both in vitro and in vivo, and that this led to their differentiation into Siglec-Fhi neutrophils. Both mechanisms were elevated when eosinophils lacked GPR43, which suggests that GPR43 constitutively prevents eosinophils from recruiting neutrophils and converting them into more pathogenic Siglec-Fhi neutrophils.
The eosinophil-induced differentiation of Siglec-Fhi neutrophils was likely mediated by eosinophil-derived IL-4 and GM-CSF, since differentiation was largely blocked by antibodies against these cytokines. Surprisingly, despite the involvement of these soluble factors, direct physical interaction between the two cell types was indispensable for Siglec-Fhi neutrophil differentiation. Given that asthmatic lung eosinophils produced relatively low levels of IL-4 and GM-CSF, direct contact may represent the most efficient means of delivering these limited cytokines. Moreover, the comparable IL-4 and GM-CSF levels in control and GPR43-deficient eosinophils suggest that stronger chemoattraction and an increased likelihood of physical contact are the key factors driving the enhanced differentiation observed in the absence of GPR43 in eosinophils.
In vivo, additional mechanisms are likely to contribute. For example, the increased eosinophil numbers in EpxCre/+Gpr43fl/fl mice could elevate total IL-4 and GM-CSF levels within the asthmatic lung environment. Other cellular sources, including Th2 cells and ILC2s, may also supply these cytokines; notably, IL-4–producing CD4 T cells were increased in EpxCre/+Gpr43fl/fl mice. Furthermore, other soluble factors may also contribute. Previous studies have shown that ATP and TGFβ can also promote Siglec-Fhi neutrophil differentiation35,51. We also found that IL-33 and TNF can generate Siglec-Fhi neutrophils, albeit less well than GM-CSF. Further identification of the signals required for in vivo generation of Siglec-Fhi neutrophils in allergic airway inflammation will be important.
We found that many Siglec-Fhi neutrophils in the asthmatic lungs and BALF co-expressed PECAM-1. Siglec-FhiPECAM-1hi neutrophils were also generated in vitro from Siglec-FloPECAM-1lo neutrophils by asthmatic lung eosinophils, especially GPR43-deficient eosinophils. Notably, PECAM-1 co-expression by Siglec-Fhi neutrophils has not been observed elsewhere, including in lung adenocarcinoma33, myocardial infarction34, and DEP-induced lung inflammation35. Since IL-4 was by far the most potent PECAM-1-upregulating cytokine among the 16 cytokines tested, PECAM-1 co-expression by Siglec-Fhi neutrophils may be unique to type 2 cytokine-rich inflammatory conditions. However, we found that LPS, GM-CSF, and IL-33 also induced PECAM-1 expression. Thus, PECAM-1 could potentially be upregulated in Siglec-Fhi neutrophils in other inflammatory conditions.
Since asthmatic lung eosinophils also express PECAM-1, and this is upregulated by GPR43 deletion, we speculated that homophilic PECAM-1 interactions induce binding of neutrophils to eosinophils and contribute to Siglec-Fhi neutrophil induction. However, PECAM-1-blocking antibody only suppressed the enhanced ability of EOS△GPR43 to induce Siglec-Fhi neutrophil differentiation: it did not suppress baseline EOSCtrl-induced Siglec-Fhi neutrophil differentiation. Therefore, other adhesion molecules appear to play a greater role in the physical interaction between eosinophils and neutrophils, especially in the early phase of their interactions, while overexpressed PECAM-1 in EOS△GPR43 further enhances the interactions. Another possibility is that PECAM-1-mediated signaling events participate in the eosinophil-induced neutrophil differentiation: the cytoplasmic tail of PECAM-1 bears immunoreceptor tyrosine-inhibitory motifs, and PECAM-1 can modulate immune cell intracellular signaling41,52. Further studies are needed to identify additional molecules mediating the direct eosinophil-neutrophil interaction and to determine the exact mechanism(s) by which PECAM-1 regulates eosinophil-induced Siglec-Fhi-neutrophil differentiation.
Notably, asthmatic lungs also bore neutrophils that expressed PECAM-1 but not Siglec-F (Fig. 3f). Moreover, none of the PECAM-1hi or Siglec-Fhi neutrophils were found in the circulation (Fig. 3g–i). In addition, genes related to Th17 differentiation53,54 were upregulated in Siglec-Fhi neutrophils, and they could enhance Th17 cell differentiation. These findings, together with the production of neutrophil chemoattractants by eosinophils in asthmatic lungs, suggest the following sequence of immunological events in asthmatic lungs: (1) PECAM-1lo neutrophils enter the lung from blood, partly due to eosinophil-derived chemokines such as CCL6 and CXCL2, (2) the IL-4-rich local environment induces neutrophil expression of PECAM-1, (3) the eosinophil-derived chemokines further encourage the neutrophils to approach and physically connect with the eosinophils and the local eosinophil-derived IL-4 further increases PECAM-1 expression on the neutrophils, (4) eosinophil-derived IL-4 + GM-CSF induces Siglec-F expression (i.e. differentiation) of the neutrophils, and (5) Siglec-Fhi neutrophils promote Th17 differentiation (Fig. 7). This whole process can potentially occur in the asthmatic EpxCre/+ lung but significantly worsens when GPR43 is deleted in eosinophils because the deletion increases eosinophil-mediated neutrophil chemoattraction and eosinophil PECAM-1 expression. Consequently, EOS△GPR43 attracts more neutrophils and converts them to Siglec-Fhi neutrophils. Therefore, eosinophil-expressed GPR43 downregulates crosstalk between eosinophils and neutrophils in asthma-induced lung inflammation, thus minimizing eosinophil-mediated induction of Siglec-Fhi neutrophils.
(Top) In wild-type mice, GPR43 restrains eosinophil activation and the production of neutrophil chemoattractants, resulting in limited neutrophil recruitment and minimizing the generation of Siglec-Fhi neutrophils. (Bottom) In eosinophil-specific GPR43-deficient mice, infiltrating neutrophils are exposed to elevated levels of IL-4 and upregulate PECAM-1 expression. GPR43-deficient eosinophils are hyperactivated, produce more neutrophil chemoattractants, and express higher levels of PECAM-1 compared with wild-type eosinophils. These changes lead to enhanced neutrophil recruitment and stronger eosinophil-neutrophil interactions, promoting the induction of Siglec-Fhi neutrophils via eosinophil-derived IL-4 and GM-CSF. Siglec-Fhi neutrophils, in turn, promote Th17 cell differentiation. Created in BioRender. Yu, J. (2025) https://BioRender.com/d0uv25k.
Siglec-Fhi neutrophils have been found to play pathogenic roles in many diseases33,34,35,36. Similarly, Siglec-Fhi neutrophils in our HDM-induced asthma model highly expressed proinflammatory genes, especially Il1a and Il23a, promoting Th17 cells53,54. Since Th17 cells and IL-17A can, in turn, promote neutrophil-mediated immune responses55,56, it is possible that there is a positive feedback loop between Siglec-Fhi neutrophils and Th17 cells that results in exacerbated asthma symptoms. Moreover, this feedback loop can be strengthened when GPR43 is deleted in eosinophils.
While most of these mechanisms were studied in male mice, enhanced production of chemokines, particularly CCL6 and CCL24, was also observed in GPR43-deficient female eosinophils. Moreover, under acetate-treated asthma conditions, female EpxCre/+Gpr43fl/fl mice exhibited higher frequencies of Siglec-Fhi neutrophils than control females, indicating that the role of eosinophil GPR43 is conserved across sexes. Nonetheless, several other phenotypic differences observed between male EpxCre/+ and EpxCre/+Gpr43fl/fl mice were not evident in females, revealing notable sex-specific variations. Sex-based differences in asthma pathogenesis, including more severe inflammation, have been widely reported44,45,46. Proposed contributing factors include hormonal influences57,58,59, structural differences in lung anatomy60, and gut microbiota composition61,62,63. In addition, systemic SCFA levels differ between males and females under both steady-state and disease conditions64,65,66, although whether such differences occur during asthma remains unknown. These variables may influence the magnitude or context-dependence of GPR43 signaling. Further investigation is needed to elucidate how sex-specific physiological factors intersect with GPR43-mediated regulation of eosinophil–neutrophil interactions in allergic airway inflammation.
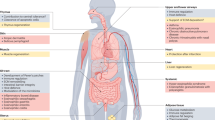
Beyond these mechanistic and sex-specific insights, our findings are of clinical interest because asthma is broadly classified on the basis of immune cell profiles as eosinophilic and non-eosinophilic asthma67,68. Eosinophilic asthma is driven by Th2 responses and is generally readily managed with corticosteroids. Non-eosinophilic asthma includes the neutrophilic and paucigranulocytic endotypes and associates with either Th1 or Th17 responses. It is often resistant to corticosteroid treatment. Non-eosinophilic asthma also includes the mixed-granulocytic endotype, which involves both eosinophils and neutrophils and is driven by Th2 and Th17 response: this endotype is associated with severe symptoms69,70,71. Since we found that eosinophils directly regulate neutrophils and GPR43 is a critical suppressor of eosinophil hyperactivation, our results offer valuable insights into mechanisms that may underlie the development of mixed-granulocytic asthma. For example, our findings suggest that insufficient GPR43 signaling due to antibiotic overuse or inadequate dietary fiber intake could induce dysfunctional eosinophils that aberrantly recruit and activate neutrophils in response to asthmogenic signals, which in turn trigger powerful Th17 responses. Further studies on the GPR43-eosinophil-Siglec-Fhi neutrophil-Th17 axis in not only asthma but also other diseases that involve both eosinophils and neutrophils (e.g., chronic obstructive pulmonary disease, infections, and tumors) are warranted.
Methods
Experimental animals
Gpr43fl/fl mice were generated using ES cells (Ffar2tm1a(EUCOMM)Hmgu, Clone # HEPD0778_6_G06) from EUCOMM. Epx-Cre mice72 and Rosa26-eYFPfl/fl mice (Jackson laboratory, 006148)73 were obtained from the late James J. Lee (Mayo Clinic, Rochester, Minnesota, USA) and Suk-jo Kang (Korea Advanced Institute of Science and Technology (KAIST), Daejeon, Republic of Korea), respectively. GPR43-/- mice were previously described74. All mice were maintained on a C57BL/6 background. Control and experimental groups were co-housed from three weeks of age until the end of the experiments. Unless otherwise stated, all experiments were performed using eight to thirteen-week-old male mice. Mice were provided with standard chow ad libitum and maintained under specific pathogen-free conditions at 22 ± 2 °C, 50 ± 10% humidity, with a 12-h light-dark cycle in the KAIST Laboratory Animal Resource Center. All mouse experiments were approved by the KAIST Institutional Animal Care and Use Committee (IACUC).
Allergic airway inflammation models
For the HDM-induced asthma model, mice received intranasal administration of 10 µl of 1 mg/ml house dust mite (HDM) extract (Greer, USA, Catalogue#XPB82D3A2.5) dissolved in PBS on days 0, 7, 8, 9, 10, and 11. For acetate treatment, sodium acetate (Sigma-Aldrich, USA, Catalogue#S5636) was dissolved in water, adjusted to pH 7.4, and administered intraperitoneally at 0.5 g/kg body weight 30 minutes before HDM administration on days 7-11. For the OVA-induced asthma model, mice were sensitized intraperitoneally on day 0 with 200 µl of 1 mg/ml OVA (Sigma-Aldrich, USA, Catalogue#A5503) and Alum (Imject, USA, Catalogue#77161) mixed in a 1:1 ratio. Mice were then challenged intranasally with 20 µl of 2.5 mg/ml OVA in PBS on days 7, 8, and 9. For all intranasal administration, mice were anesthetized by intraperitoneal injection of a mixture containing ketamine (80 mg/kg) and rompun (4 mg/kg) in PBS. In both models, mice were euthanized using CO2 one day after the last challenge.
Measurement of airway resistance
Airway resistance was measured using FlexiVent (SCIREQ, Canada). Mice were anesthetized by intraperitoneal injection of pentobarbital sodium solution in PBS (30 mg/kg) and intubated with 20-gauge catheters by tracheostomy. To block spontaneous breathing, 100 µl of 1 µg/ml pancuronium bromide (Sigma-Aldrich, Catalogue#P1918) was injected intraperitoneally. Mice were mechanically ventilated at a tidal volume of 10 ml/kg and a respiratory rate of 150/min. Respiratory resistance (Rrs) values were acquired after administering increasing doses (0, 5, 10, 20 mg/ml) of aerosolized methacholine (Sigma-Aldrich, Catalogue# A2251).
Histological analysis of asthmatic lung
Half of the left lung tissues were excised, fixed with 3.7% formaldehyde (Sigma-Aldrich, Catalogue#F1635) at 4 °C overnight, and embedded in paraffin using a tissue processor (TP1020 Automatic Benchtop Tissue Processor / HistoCore Arcadia, Leica). The paraffin block was cut into 10 µm-thick sections with the microtome (RM2235, Leica) and stained with hematoxylin (Merck, Germany, Catalogue#1.09253.1000) and eosin (Merck, Catalogue#1.09844.1000) using a standardized protocol. Periodic Acid Schiff (PAS) and Picro Sirius Red staining were performed using kits (Abcam, UK, Catalogue#ABC-AB150680, ABC-AB150681) according to the manufacturer’s protocols. Images were obtained using a slide scanner (Pannoramic SCAN II, 3DHISTECH, Hungary) in the EM & histology core facility at the KAIST biomedical research center.
Immunofluorescence imaging of asthmatic lung
After euthanizing the mice, 1 ml OCT solution (Sakura Finetek, USA, Catalogue#4583) was injected into the lungs through the trachea, and the lungs were extracted. The extracted lungs were immersed in OCT solution and preserved at -20 °C. OCT-embedded lungs were sectioned into 14 µm-thick slices and attached to the slide glasses. After fixation with 4% paraformaldehyde (Sigma-Aldrich, Catalogue#P6148) and permeabilization with 0.3% Triton X-100 in PBS, anti-Siglec-F (E50-2440, BD Biosciences, Catalogue#55125) and anti-myeloperoxidase (P11247, R&D Systems, Catalogue#AF3667) antibodies were used to stain eosinophils and neutrophils, respectively. Anti-rat IgG-AF594 (Thermo Fisher Scientific, Catalogue#A-21209) and anti-goat IgG-AF488 (Thermo Fisher Scientific, Catalogue#A-11055) antibodies were used as the secondary antibodies. After nuclear staining with DAPI (Thermo Fisher Scientific, Catalogue#D1306) and adding a fluorescence mounting medium (Dako, Denmark, Catalogue#S3020), a cover glass was mounted. Confocal imaging was performed using Nikon A1 HD25 microscope (Japan), and images were processed with NIS-Elements BR 4.60. software.
Lung and BALF cell preparation
Lungs were chopped into 1 mm3 fragments and digested with 400 MandlU/ml collagenase D (Roche, Switzerland, Catalogue#11088882001) and 10 µg/ml DNase I (Roche, Catalogue#10104159001) in RPMI1640 media (Welgene, South Korea, Catalogue#LM011-01) containing 3% FBS (Atlas biologicals, USA, Catalogue#EF-0500-A), 20 mM HEPES (Welgene, Catalogue#BB001-01), 1 mM non-essential amino acids (Welgene, Catalogue #LS005-01), 100 U/ml penicillin, 100 µg/ml streptomycin (Welgene, Catalogue#LS 202-02), and 1 mM sodium pyruvate (Welgene, Catalogue#LS013-01) at 37 °C for 30 min with stirring. The enzyme reaction was stopped by adding 10 µl of 0.5 M EDTA (Sigma-Aldrich, Catalogue#E6758) and incubating the cells for an additional 5 min. After vigorous resuspension, the cells were filtered through 100 µm strainers (SPL, South Korea, Catalogue#93100). Red blood cells were lysed with ACK lysis buffer containing 0.15 M NH4Cl (Daejung, South Korea, Catalogue#1060-4405), 10 mM KHCO3 (Daejung, Catalogue#6580-4405), and 0.1 mM EDTA (Sigma-Aldrich, Catalogue#E6758) for 3 min at room temperature. BALF was collected by flushing the airways with 1 ml PBS. To evaluate the tissue localization of immune cells by flow cytometry, mice were injected intravenously with 1 µg of CD45-PE antibody (30-F11, Biolegend, Catalogue#103106) 5 min before sacrifice.
Flow cytometry
Cells were first stained with Ghost dye violet 510 (Tonbo, USA, Catalogue#13-0870-T500) for live/dead staining and then with fluorescent dye-labeled antibodies in PBS-based buffer supplemented with 3 % FBS, 1 mM EDTA, and 20 mM HEPES on ice for 20 min. For intracellular cytokine staining, the cells were stimulated with a cocktail containing 81 nM PMA (Sigma-Aldrich, Catalogue#79346), 1.34 µM ionomycin (Sigma-Aldrich, Catalogue#I9657), 2 µM monensin (Sigma-Aldrich, Catalogue#M5273), and 10.6 µM brefeldin A (Sigma-Aldrich, Catalogue#B7651) at 37 °C for 4 h. The cells were then fixed and permeabilized with 3.7% formaldehyde (Sigma-Aldrich, Catalogue#F1635) and 0.5% saponin (Sigma-Aldrich, Catalogue#47036), respectively. To stain the transcription factors in CD4 T cells, a FoxP3 staining kit (eBioscience, Catalogue#00-5521-00) was used according to the manufacturer’s protocol. All antibodies used for flow cytometry analysis are detailed in Supplementary Table 1. Flow cytometric analysis was performed using BD Biosciences LSR Fortessa X-20 and data were analyzed with BD Biosciences Flowjo v.10.10.0. Gating strategies can be found in Supplementary Figs. 15–19.
Purification of lung eosinophils and morphological analysis
After staining lung cells with fluorescent dye-labeled antibodies, eosinophils were purified by fluorescence-activated cell sorting using BD Biosciences Aria Fusion. Dead cells were excluded using propidium iodide staining. The purity of the sorted eosinophils was over 95% (Supplementary Fig. 17a). For morphological analysis, 50,000 cells were attached to a slide glass using Cytospin 4 (Thermo Fisher Scientific). After overnight drying, cells were fixed with 3.7% formaldehyde, stained with hematoxylin and eosin using a standardized protocol, and mounted with xylene mounting solution (BBC Biochemical, USA). To analyze their nuclear shape, cells were stained with propidium iodide and mounted with a fluorescence mounting medium (Dako). The images were taken with Pannoramic SCAN II.
Purification of bone marrow and lung neutrophils
To purify bone marrow neutrophils, bone marrow cells were collected by flushing the tibia and femur with RPMI1640-based media containing 2% FBS (Atlas Biologicals), 10 mM HEPES, 100 U/ml penicillin, and 100 µg/ml streptomycin using 26 G syringes. The bone marrow and lung cell preparations were enriched for neutrophils by negative selection with biotin-conjugated antibodies listed in Supplementary Table 2 by using EasySep Mouse Streptavidin RapidSpheres isolation kit (STEMCELL Technologies, Catalogue#ST19860). The neutrophils were further stained with fluorescent dye-labeled antibodies and purified using an Aria Fusion cell sorter (BD Biosciences). Their purity was over 95% (Supplementary Fig. 17b).
In vitro differentiation of Siglec-Fhi neutrophils
For cytokine-induced differentiation of Siglec-Fhi neutrophils, 0.2 million neutrophils purified from the bone marrow were cultured in RPMI-based media supplemented with 10% FBS (Hyclone, Catalogue#SH30919.03), 1 mM non-essential amino acids, 5 mM HEPES, 100 U/ml Penicillin, 100 µg/ml Streptomycin, 1 mM sodium pyruvate, 50 µg/ml gentamicin (Gibco, USA, Catalogue#15750-060), 55 µM β-mercaptoethanol (Sigma-Aldrich, Catalogue#M6250), and 2 mM L-glutamine (Gibco, Catalogue#25030-081) for 2 days in the presence of 10 ng/ml of each cytokine, including IL-1α (Catalogue#400-ML), IL-1β (Catalogue#401-ML), IL-1Ra (Catalogue#480-RM), IL-2 (Catalogue#402-ML), IL-4 (Catalogue#404-ML), IL-5 (Catalogue#405-ML), IL-8 (Catalogue#208-IL-010), IL-10 (Catalogue#417-ML), IL-12 (Catalogue#419-ML), IL-17A (Catalogue#421-ML), IL-33 (Catalogue#3626-ML), IFNγ (Catalogue#485-ML) (all from R&D Systems), IL-6 (BD Biosciences, Catalogue#554582), TNF (BD Biosciences, Catalogue#554589), GM-CSF (Peprotech, Catalogue#315-03), M-CSF (Sigma-Aldrich, Catalogue#SRP3110), and TGFβ (Miltenyi Biotec, Catalogue#130-095-066), or 10 µg/ml LPS (Sigma-Aldrich, Catalogue#L4391).
For eosinophil-induced differentiation of Siglec-Fhi neutrophils, 0.2 million bone marrow neutrophils were co-cultured with 0.1 million lung eosinophils in the culture media described above in 96-well round-bottom plates for 2 days. For each steady-state eosinophil sample, lung eosinophils from four mice were pooled, whereas each asthmatic lung eosinophil sample was isolated from an individual mouse.
For antibody-mediated blocking of Siglec-Fhi neutrophil differentiation, asthmatic lung eosinophils were pretreated for 5 min with either a combination of 5 µg/ml anti-IL-4 (BVD4-1D1, BD Biosciences, Catalogue#554387) and 5 µg/ml anti-GM-CSF (MP122E9, R&D Systems, Catalogue#MAB415) or 1 µg/ml anti-PECAM-1 (MEC13.3, Biolegend, Catalogue#102501) and then co-cultured with bone marrow neutrophils. The following isotype control antibodies were used as controls: purified Rat IgG2a, κ (e-bioscience, Catalogue#16-4321-85), and purified Rat IgG2b, κ (BD Biosciences, Catalogue#553985). To achieve physical separation between eosinophils and neutrophils, 0.2 million bone marrow neutrophils were cultured in 24-well flat-bottom plates with 0.4 million asthmatic lung eosinophils placed in 0.4 μm transwell inserts (Corning, Catalogue#CLS3470) for 2 days.
Analysis of eosinophil-neutrophil interaction
Bone marrow neutrophils (0.2 million) were cultured in the presence or absence of asthmatic lung eosinophils (0.1 million) in a 96-well flat bottom plate for 2 days, and cells were imaged using a Nikon ECLIPSE Ts2R microscope (Japan). For fluorescence imaging, bone marrow neutrophils labeled with carboxyfluorescein succinimidyl ester (Thermo Fisher Scientific, Catalogue#C34570) and asthmatic lung eosinophils (0.1 million each) were co-cultured in a 96-well round bottom plate for 4 h. Without any resuspension, cells were carefully transferred into the cytofunnel and attached to a slide glass using Cytospin 4 (Thermo Fisher Scientific). After overnight drying, the cells were fixed with 4% paraformaldehyde, permeabilized with 0.3% Triton X-100 in PBS, and stained with an anti-CD63-APC antibody (NVG-2, Biolegend, Catalogue#143905). Fluorescence images were taken using a Nikon A1 HD25 confocal microscope.
Neutrophil transwell migration assay
Bone marrow neutrophils were labeled with 5 µM CellTrace Violet (CTV, Invitrogen, Catalogue#C34557) at 37 °C for 20 min. CTV-labeled neutrophils (0.1 or 0.2 million) and asthmatic lung eosinophils (0.7 million) were placed in a 3 µm transwell insert (Corning, Catalogue#3385) and the bottom 96-well plate, respectively, and cultured for 24 h. The cells in the bottom plate were resuspended to dissociate the aggregated cells, and CTV-labeled neutrophils were imaged using a Nikon ECLIPSE Ts2R fluorescence microscope.
In vitro Th17 differentiation
To determine the ability of neutrophils to induce Th17 differentiation, Siglec-Fhi and Siglec-Flo neutrophils were prepared by culturing bone marrow neutrophils for 2 days in the presence or absence of 10 ng/ml GM-CSF (Peprotech, Catalogue#315-03) and 10 ng/ml IL-4 (R&D Systems, Catalogue#404-ML), respectively. Following differentiation, neutrophils were washed to remove residual IL-4 and GM-CSF, and live cell numbers were recounted before co-culture with naïve CD4 T cells. To prepare naïve CD4 T cells, spleens and peripheral lymph nodes were ground through 100 µm strainers using the piston of 1 ml syringes. After lysing red blood cells with ACK lysis buffer, naïve CD4 T cells were isolated by negative selection with biotin-conjugated antibodies listed in Supplementary Table 2 by using EasySep Mouse Streptavidin RapidSpheres isolation kit (STEMCELL Technologies). Naïve CD4 T cells (0.1 million) were co-cultured with Siglec-Fhi or Siglec-Flo neutrophils (0.05 million) in the presence of 5 ng/ml TGFβ (Miltenyi Biotec, Catalogue#130-095-066), 2 ng/ml IL-6 (BD Biosciences, Catalogue#554582), 5 µg/ml anti-IL-4 (BVD4-1D1, BD Biosciences, Catalogue#554387), 2.5 µg/ml anti-IL-2 (S4B6, BD Biosciences, Catalogue#554375), 5 µg/ml anti-IFNγ (R4-6A2, Invitrogen, Catalogue#14-7312-85) in a 96-well flat bottom plate pre-coated with 2.5 µg/ml anti-CD3 (145-2C11, BD Biosciences, Catalogue#553057) and 1 µg/ml anti-CD28 (37.51, BD Biosciences, Catalogue#553294). After 3 days, the frequencies of RORγt or IL-17A-expressing CD4 T cells were analyzed by flow cytometry.
To assess the ability of eosinophils to induce Th17 differentiation, naïve CD4 T cells (0.1 million) were cultured as described above, except that neutrophils were replaced with asthmatic lung eosinophils (0.1 million), and IL-6 was supplied at 20 ng/ml under Th17 differentiation conditions. To assess whether eosinophils indirectly influence Th17 differentiation via dendritic cells, splenic dendritic cells were added to the naïve CD4 T cell-eosinophil co-culture. To prepare the splenic dendritic cells, spleens were chopped into 1 mm3 fragments and digested with 400 MandlU/ml Collagenase D and 10 µg/ml DNase I in RPMI1640 media containing 3% FBS (Atlas biologicals), 20 mM HEPES, 1 mM non-essential amino acids, 100 U/ml penicillin, 100 µg/ml streptomycin, and 1 mM sodium pyruvate at 37 °C for 30 min with stirring. The cells were centrifuged over 17.5 % Accudenz solution at 805 × g for 20 min at room temperature, and the cells at the interface were collected. The dendritic cells were further stained with fluorescent dye-labeled antibodies and purified using BD Biosciences Aria Fusion cell sorter to a purity of over 95%.
Cytokine and chemokine ELISA
To measure cytokine production from T cells in the lung, total asthmatic lung cells (0.5 million) were cultured in flat-bottom 96-well plates coated with 1 µg/ml of anti-CD3 (145-2C11, BD Biosciences, Catalogue#553057) and 1 µg/ml anti-CD28 (37.51, BD Biosciences, Catalogue#553294) for 3 days, and the culture supernatants were collected. To measure neutrophil cytokine production, bone marrow neutrophils (0.3 million) were cultured in the presence of 10 ng/ml IL-4 (R&D Systems, Catalogue#404-ML) or 10 ng/ml GM-CSF (Peprotech, Catalogue#315-03) in 96-well round-bottom plates for 2 days, and the culture supernatants were collected. On the second day of the culture, neutrophils were either stimulated or not with 100 nM PMA (Sigma-Aldrich, Catalogue#79346) and 1 µg/ml ionomycin (Sigma-Aldrich, Catalogue#I9657). To assess cytokine production by asthmatic lung neutrophils, Siglec-Flo and Siglec-Fhi neutrophils were sorted from asthmatic lungs based on Siglec-F expression, and eighty thousand cells were cultured in 96-well round-bottom plates for 24 hours without stimulation. To measure cytokine production by asthmatic lung eosinophils, sorted cells (0.5 million) were cultured in 96-well round-bottom plates for 24 hours either without stimulation (for CCL6 and CCL24) or in the presence of 100 nM PMA and 1 µg/ml ionomycin (for CXCL2). To measure cytokine production by bone marrow-derived eosinophils (BMDEs), 0.3 million cells were cultured in 96-well round-bottom plates for 24 hours with stimulation by 10 ng/ml IL-33 (R&D Systems, Catalogue#3626-ML). Antibodies and kits for cytokine ELISA were listed in Supplementary Table 3. Half-Area 96-well microplates (Corning, Catalogue#3690) were coated with 2 µg/ml capture antibodies in PBS at 4 °C overnight and blocked with blocking buffer containing 2% BSA fraction V (Roche, Catalogue#10735108001) in PBS-T (0.05% Tween-20 (Daejung, Catalogue#8571-1405) in PBS) for 1 h at room temperature. Subsequently, samples were added and incubated for 1 h. Thereafter, 1 µg/ml biotin-conjugated antibodies were added for 45 min, followed by treatment with 50 ng/ml streptavidin-horseradish peroxidase (R&D Systems, Catalogue#DY998) for 20 min. Between each step, plates were washed at least 3 times with PBS-T. To develop the reaction, 50 µl of tetramethylbenzidine solution (Surmodics, USA, Catalogue#TMBW) was added, and the reaction was stopped by adding 50 µl of 0.5 M H2SO4 (Daejung, Catalogue#7683-4100). Absorbance at O.D.450 nm was read using SpectraMax M2 Microplate Reader (Molecular Devices, USA).
Preparation of bone marrow-derived eosinophils
Bone marrow-derived eosinophils (BMDEs) were differentiated by following the previously described protocol75. In brief, bone marrow cells (3 ×106/ml) were differentiated with 100 ng/ml SCF (Peprotech, Catalogue#250-03), and 100 ng/ml FLT3 ligand (R&D Systems, Catalogue#427-FL-025/CF) in RPMI1640 media containing 20 % FBS (Hyclone, Catalogue#SH30919.03), 1 mM non-essential amino acids, 25 mM HEPES, 100 U/ml Penicillin, 100 µg/ml streptomycin, 1 mM sodium pyruvate, 50 µM β-mercaptoethanol, and 2 mM L-glutamine in 6-well plates for 4 days. The medium was then changed to new culture medium containing 10 ng/ml IL-5 (R&D Systems, Catalogue#405-ML). On days 8 and 11, the medium was changed again with new IL-5-containing medium, and the cell number was adjusted to 1 × 106/ml. On day 14, ELISA, Ca2+ flux assay, and RNA extraction were performed.
Ca2+ flux assay
Cells were loaded with 2 µg/ml Fluo-3 AM (Thermo Fisher Scientific, Catalogue#F23915) and 5 µg/ml Fura Red AM (Thermo Fisher Scientific, Catalogue#F3021) in RPMI1640 medium containing 2% FBS, 25 mM HEPES, 2.5 mM probenecid (Sigma-Aldrich, Catalogue#P8761), and 0.02% pluronic F-127 (Sigma-Aldrich, Catalogue#P2443) at 37 °C for 20 min. After washing, the cells were analyzed using a flow cytometer (LSR Fortessa X-20, BD Biosciences). Basal fluorescence was recorded for 30 seconds, and changes in the fluorescence were recorded for an additional 3 min after stimulating the cells with 10 mM acetate.
RNA extraction
Asthmatic lung eosinophils, BMDEs, and bone marrow neutrophils were lysed in Qiazol (QIAzen, Germany, Catalogue#79306), and chloroform (Daejung, Catalogue#2548-4400) was added in a 5:1 ratio. After centrifugation, the clear upper phase was collected, and RNA was precipitated by adding the same volume of 100% isopropanol (Merck, Catalogue#1.09634.2511) and 20 µg glycogen (Sigma-Aldrich, Catalogue#G1767). After centrifugation, the pellet was washed with 75% ethyl alcohol (Merck, Catalogue#1.00983.2511) diluted in DEPC water (Biosesang, South Korea, Catalogue#WR2004-050-00) and air-dried. RNA was solubilized in nuclease-free water (Invitrogen, Catalogue#AM9930).
Quantitative PCR
RNAs were reverse-transcribed to cDNA using GoScript 5X reaction buffer (Catalogue#A500C), reverse transcriptase (Catalogue#A501C), ribonuclease inhibitor (Catalogue#N251B), dNTP (Catalogue#U151B), MgCl2 (Catalogue#A351H), and oligo(dT)15 primer (Catalogue#C110A) (all from Promega). Using cDNAs as templates, qPCR analysis of genes of interest was performed with SYBR Green Realtime PCR Master Mix (TOYOBO, Japan, Catalogue#QPK-201) and CFX Connect Real-Time System (Bio-Rad). Gene expression levels were calculated based on the comparative Cq method and normalized to Hprt gene expression. Primers are described in Supplementary Table 4.
Bulk-RNA sequencing
The integrity of asthmatic lung eosinophil RNA was measured using a Bioanalyzer 2100 (Agilent Technologies, USA). Ribosomal RNAs were removed using NEBNext rRNA Depletion Kit v2 (USA, Catalogue#E7405L), and the cDNA library was synthesized using NEBNext Ultra II Directional RNA Library Prep kit (Catalogue#E7765S) according to the manufacturer’s protocols. Sequencing was conducted on an Illumina HiSeq2500 with paired-ended reads of 100 base pairs by Macrogen (South Korea). Fastq files were trimmed with the package Fastp and aligned to the mm10 mouse reference genome sequences with STAR. Transcript per million (TPM) was calculated from the counts obtained with FeatureCounts, and differentially expressed genes (DEGs) were acquired from the raw counts with DESeq2 by using a p-value below 0.05. All processes were conducted through Galaxy (https://usegalaxy.org/). Additional pathway analysis was performed using pathfindR and Clusterprofiler in R v.4.3.1. Publicly available bulk-RNA sequencing data for eosinophils from mouse spleen26, peritoneal cavity26, lung24, colon25, skin14, blood14, bone marrow14, BMDE27, and human peripheral blood28 were analyzed with the same processes (Supplementary Table 5).
Single-cell RNA sequencing
Asthmatic lung cells from EpxCre/+ and EpxCre/+Gpr43fl/fl mice were isolated from 3 mice per genotype, pooled, and stained with Ghost dye Red780 (TONBO, Catalogue#13-0865-T100). Live cells, excluding FSChi SSChi cells, were sorted and suspended in PBS containing 0.04% BSA. The total of 28,320 and 25,620 cells from EpxCre/+ and EpxCre/+Gpr43fl/fl mice, respectively, were loaded onto Chromium-controller (10X Genomics), and scRNA-seq libraries were generated by using the Chromium Next GEM Single Cell 3ʹ Reagent Kits v 3.1 (10X Genomics, Catalogue#PN-1000268) according to the manufacturer’s protocol at the KAIST NGS core facility. Sequencing was performed on a HiSeqX platform by Macrogen (South Korea).
Raw sequencing files were initially processed with the Cell Ranger (v.4.0.0) pipeline, using the mm10 reference, and downstream analysis was conducted based on the Scanpy (v.1.8.2) pipeline. Low-quality cells meeting any one of the following criteria were excluded in the downstream analysis: (1) UMI gene count <1000, (2) number of genes detected <500, (3) number of genes detected > 7000, (4) fraction of mitochondrial genes > 10%, and (5) predicted doublets identified with the Scrublet (v.0.2.2) package. Raw counts were normalized in each cell (100,000 counts per cell) and log-transformed (log1p). Highly variable genes (n = 2,418) were selected by using the highly_variable_genes function in the Scanpy package. Principal component analysis (PCA) was conducted (n_PCs = 50) on the scaled expression values of the highly variable genes. Neighborhood graphs of the cells were constructed (n_neighbors = 15) based on PC coordinates, and the Uniform Manifold Approximation and Projection (UMAP) embedding of the cells was calculated. After the preprocessing steps, 3,989 cells from the EpxCre/+ sample and 4,871 cells from the EpxCre/+Gpr43fl/fl sample were retrieved and analyzed. Wilcoxon rank-sum test was used for the DEG analysis, and p-values were adjusted with the Benjamini-Hochberg method.
For gene set enrichment analysis (GSEA) of DEGs between Siglec-Flo and Siglec-Fhi neutrophils, a pre-ranked gene set was created by listing 3515 DEGs with an adjusted p-value of less than 0.05, ranked by test statistics score using GSEApy v.1.1.3. Normalized enrichment scores (NES) and false discovery rates (FDR) were obtained by using following parameters: (1) number of permutations = 1000, (2) permutation type: gene set, and (3) weighted enrichment statistics. Significant gene sets were those with an FDR < 0.25 and p-value < 0.05. Gene sets for the analysis were obtained from the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) databases. To compare the Siglec-Fhi neutrophils identified in our HDM-induced asthma model to those in other disease models using publicly available transcriptomic data33,34,35, another pre-ranked gene set was generated by ranking the same 3515 DEGs according to LogFC values. Gene lists for the Siglec-Fhi neutrophil signature genes in the other disease models were generated, using the top 350 DEGs between Siglec-Flo and Siglec-Fhi neutrophils in each disease model, and ranked based on LogFC values.
Analysis of publicly available scRNA-seq data was performed using Seurat v4.1.3. Matrix (mtx) files for esophagus and duodenum samples from 10 eosinophilic esophagitis patients were downloaded (GSE175930)29 together with feature and cell information files (Supplementary Table 5), and Seuratobject was created. Low-quality cells meeting any one of the following criteria were excluded in the downstream analysis: (1) min.cells <3, (2) min.features <200, (3) number of genes detected <200, (4) number of genes detected > 6000, and (5) fraction of mitochondrial genes > 20%. Seuratobjects were normalized, and highly variable genes were found by using the vst selection method. Each esophagus and duodenum sample was integrated separately (n_dimension = 50), and PCA was conducted (n_PC = 30). Neighborhood graphs of the cells were constructed (n_dimension = 30), UMAP was created, and the cells were clustered (esophagus: resolution = 0.3; duodenum: resolution = 0.1). Each cluster was manually annotated using known marker genes.
Statistical analysis
All data points represent biological replicates unless otherwise noted. Specifically, Fig. 4a–c show experimental replicates, and Fig. 6g and Supplementary Fig. 9e show technical replicates. For all technical replicate experiments, results were confirmed in at least three independent experiments. For comparisons between two groups, either a paired or unpaired two-tailed Student’s t-test was performed, depending on sample matching. One-way ANOVA with Tukey’s multiple comparisons test was used to compare the means of three or more groups with a single independent variable. To evaluate the effects of two independent variables and their interaction, two-way ANOVA with Tukey’s multiple comparisons test was conducted. Statistical analyses were performed with GraphPad Prism v8.4.2.
Reporting summary
Further information on the research design is available in the Nature Research Reporting Summary linked to this article.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw and processed bulk RNA-seq data (expression matrix) from asthmatic lung eosinophils have been deposited in the Gene Expression Omnibus under accession code GSE271832. In addition, a filtered expression matrix (genes with an average TPM ≥ 5 across samples) is also provided in Supplementary Data 1. The raw and processed single-cell RNA-seq data (expression matrix and associated metadata) from whole asthmatic lung cells are available under accession code GSE271835. Additional information on previously published datasets used in this study is provided in Supplementary Table 5. All data are included in this article, the Supplementary Information, and the Source Data file. Raw data files for flow cytometry analysis are available from the authors upon reasonable requests. Source data are provided with this paper.
References
Rothenberg, M. E. & Hogan, S. P. The eosinophil. Annu Rev. Immunol. 24, 147–174 (2006).
Weller, P. F. & Spencer, L. A. Functions of tissue-resident eosinophils. Nat. Rev. Immunol. 17, 746–760 (2017).
Arnold, I. C. & Munitz, A. Spatial adaptation of eosinophils and their emerging roles in homeostasis, infection and disease. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-024-01048-y (2024).
Lee, J. J., Jacobsen, E. A., McGarry, M. P., Schleimer, R. P. & Lee, N. A. Eosinophils in health and disease: the LIAR hypothesis. Clin. Exp. Allergy 40, 563–575 (2010).
Kanda, A. et al. The multiple functions and subpopulations of eosinophils in tissues under steady-state and pathological conditions. Allergol. Int 70, 9–18 (2021).
Wu, D. et al. Eosinophils sustain adipose alternatively activated macrophages associated with glucose homeostasis. Science 332, 243–247 (2011).
Arnold, I. C. et al. Eosinophils suppress Th1 responses and restrict bacterially induced gastrointestinal inflammation. J. Exp. Med 215, 2055–2072 (2018).
Sugawara, R. et al. Small intestinal eosinophils regulate Th17 cells by producing IL-1 receptor antagonist. J. Exp. Med 213, 555–567 (2016).
Mesnil, C. et al. Lung-resident eosinophils represent a distinct regulatory eosinophil subset. J. Clin. Invest 126, 3279–3295 (2016).
Dougan, M., Dranoff, G. & Dougan, S. K. GM-CSF, IL-3, and IL-5 Family of cytokines: Regulators of inflammation. Immunity 50, 796–811 (2019).
Diny, N. L. et al. The aryl hydrocarbon receptor contributes to tissue adaptation of intestinal eosinophils in mice. J. Exp. Med. 219, https://doi.org/10.1084/jem.20210970 (2022).
Wang, W. L. et al. The aryl hydrocarbon receptor instructs the immunomodulatory profile of a subset of Clec4a4(+) eosinophils unique to the small intestine. Proc. Natl Acad. Sci. USA 119, e2204557119 (2022).
Vassily, I. K., Korn, L. L. & Medzhitov, R. Nutrient-derived signals regulate eosinophil adaptation to the small intestine. Proc. Natl Acad. Sci. USA 121, e2316446121 (2024).
Li, Y. et al. Neuromedin U programs eosinophils to promote mucosal immunity of the small intestine. Science 381, 1189–1196 (2023).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Backhed, F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 165, 1332–1345 (2016).
Tan, J. K., Macia, L. & Mackay, C. R. Dietary fiber and SCFAs in the regulation of mucosal immunity. J. Allergy Clin. Immunol. 151, 361–370 (2023).
Yoon, J. H., Do, J. S., Velankanni, P., Lee, C. G. & Kwon, H. K. Gut microbial metabolites on host immune responses in health and disease. Immune Netw. 23, e6 (2023).
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Theiler, A. et al. Butyrate ameliorates allergic airway inflammation by limiting eosinophil trafficking and survival. J. Allergy Clin. Immunol. 144, 764–776 (2019).
Tan, J. et al. Dietary fiber and bacterial scfa enhance oral tolerance and protect against food allergy through diverse cellular pathways. Cell Rep. 15, 2809–2824 (2016).
Mann, E. R., Lam, Y. K. & Uhlig, H. H. Short-chain fatty acids: linking diet, the microbiome and immunity. Nat. Rev. Immunol. 10.1038/s41577-024-01014-8 (2024).
Wu, W. et al. Microbiota metabolite short-chain fatty acid acetate promotes intestinal IgA response to microbiota which is mediated by GPR43. Mucosal Immunol. 10, 946–956 (2017).
Chun, E. et al. Metabolite-sensing receptor ffar2 regulates colonic group 3 innate lymphoid cells and gut immunity. Immunity 51, 871–884.e876 (2019).
Motomura, Y. et al. Basophil-derived interleukin-4 controls the function of natural helper cells, a member of ILC2s, in lung inflammation. Immunity 40, 758–771 (2014).
Grace, J. O. et al. Reuse of public, genome-wide, murine eosinophil expression data for hypotheses development. J. Leukoc. Biol. 104, 185–193 (2018).
Yoshida, H. et al. The cis-regulatory atlas of the mouse immune system. Cell 176, 897–912.e820 (2019).
Arnold, I. C. et al. The GM-CSF-IRF5 signaling axis in eosinophils promotes antitumor immunity through activation of type 1 T cell responses. J. Exp. Med. 217, https://doi.org/10.1084/jem.20190706 (2020).
Van Hulst, G. et al. Anti-IL5 mepolizumab minimally influences residual blood eosinophils in severe asthma. Eur. Respi.r J. 59, https://doi.org/10.1183/13993003.00935-2021 (2022).
Morgan, D. M. et al. Clonally expanded, GPR15-expressing pathogenic effector T(H)2 cells are associated with eosinophilic esophagitis. Sci. Immunol. 6, https://doi.org/10.1126/sciimmunol.abi5586 (2021).
Wang, L. et al. Single-cell transcriptomic analysis reveals the immune landscape of lung in steroid-resistant asthma exacerbation. Proc. Natl Acad. Sci. USA 118, https://doi.org/10.1073/pnas.2005590118 (2021).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Borrelli, C., Gurtner, A., Arnold, I. C. & Moor, A. E. Stress-free single-cell transcriptomic profiling and functional genomics of murine eosinophils. Nat. Protoc. 19, 1679–1709 (2024).
Engblom, C. et al. Osteoblasts remotely supply lung tumors with cancer-promoting SiglecF(high) neutrophils. Science 358, https://doi.org/10.1126/science.aal5081 (2017).
Calcagno, D. M. et al. SiglecF(HI) Marks late-stage neutrophils of the infarcted heart: A single-cell transcriptomic analysis of neutrophil diversification. J. Am. Heart Assoc. 10, e019019 (2021).
Shin, J. W. et al. A unique population of neutrophils generated by air pollutant-induced lung damage exacerbates airway inflammation. J. Allergy Clin. Immunol. 149, 1253–1269.e1258 (2022).
Borkner, L., Curham, L. M., Wilk, M. M., Moran, B. & Mills, K. H. G. IL-17 mediates protective immunity against nasal infection with Bordetella pertussis by mobilizing neutrophils, especially Siglec-F(+) neutrophils. Mucosal Immunol. 14, 1183–1202 (2021).
Muniz-Junqueira, M. I., Barbosa-Marques, S. M. & Junqueira, L. F. Jr. Morphological changes in eosinophils are reliable markers of the severity of an acute asthma exacerbation in children. Allergy 68, 911–920 (2013).
Lionakis, M. S. et al. Chemokine receptor Ccr1 drives neutrophil-mediated kidney immunopathology and mortality in invasive candidiasis. PLoS Pathog. 8, e1002865 (2012).
Grigolato, F., Egholm, C., Impellizzieri, D., Arosio, P. & Boyman, O. Establishment of a scalable microfluidic assay for characterization of population-based neutrophil chemotaxis. Allergy 75, 1382–1393 (2020).
Muller, W. A., Weigl, S. A., Deng, X. & Phillips, D. M. PECAM-1 is required for transendothelial migration of leukocytes. J. Exp. Med 178, 449–460 (1993).
Privratsky, J. R., Newman, D. K. & Newman, P. J. PECAM-1: conflicts of interest in inflammation. Life Sci. 87, 69–82 (2010).
Spencer, L. A. & Weller, P. F. Eosinophils and Th2 immunity: contemporary insights. Immunol. Cell Biol. 88, 250–256 (2010).
Akuthota, P., Wang, H. & Weller, P. F. Eosinophils as antigen-presenting cells in allergic upper airway disease. Curr. Opin. Allergy Clin. Immunol. 10, 14–19 (2010).
Ambhore, N. S., Kalidhindi, R. S. R., Loganathan, J. & Sathish, V. Role of differential estrogen receptor activation in airway hyperreactivity and remodeling in a murine model of asthma. Am. J. Respir. Cell Mol. Biol. 61, 469–480 (2019).
Shah, R. & Newcomb, D. C. Sex bias in asthma prevalence and pathogenesis. Front Immunol. 9, 2997 (2018).
Chiarella, S. E. et al. Sex differences in a murine model of asthma are time and tissue-compartment-dependent. PLoS One 18, e0271281 (2023).
Zhu, C. et al. Homeostatic and early-recruited CD101(-) eosinophils suppress endotoxin-induced acute lung injury. Eur. Respir. J. 56, https://doi.org/10.1183/13993003.02354-2019 (2020).
Jacobsen, E. A. et al. Eosinophil activities modulate the immune/inflammatory character of allergic respiratory responses in mice. Allergy 69, 315–327 (2014).
Hammad, H. & Lambrecht, B. N. The basic immunology of asthma. Cell 184, 1469–1485 (2021).
Davoine, F. & Lacy, P. Eosinophil cytokines, chemokines, and growth factors: emerging roles in immunity. Front Immunol. 5, 570 (2014).
Ryu, S. et al. Siglec-F-expressing neutrophils are essential for creating a profibrotic microenvironment in renal fibrosis. J. Clin. Invest. 132, https://doi.org/10.1172/JCI156876 (2022).
Marelli-Berg, F. M., Clement, M., Mauro, C. & Caligiuri, G. An immunologist’s guide to CD31 function in T-cells. J. Cell Sci. 126, 2343–2352 (2013).
Lee, Y. et al. Induction and molecular signature of pathogenic TH17 cells. Nat. Immunol. 13, 991–999 (2012).
Chung, Y. et al. Critical regulation of early Th17 cell differentiation by interleukin-1 signaling. Immunity 30, 576–587 (2009).
Fan, X., Shu, P., Wang, Y., Ji, N. & Zhang, D. Interactions between neutrophils and T-helper 17 cells. Front Immunol. 14, 1279837 (2023).
Shrestha, S. & Hong, C. W. Extracellular mechanisms of neutrophils in immune cell crosstalk. Immune Netw. 23, e38 (2023).
Radzikowska, U. & Golebski, K. Sex hormones and asthma: The role of estrogen in asthma development and severity. Allergy 78, 620–622 (2023).
Trivedi, S. et al. Progesterone amplifies allergic inflammation and airway pathology in association with higher lung ILC2 responses. Am. J. Physiol. Lung Cell Mol. Physiol. 327, L65–L78 (2024).
Laffont, S., Blanquart, E. & Guery, J. C. Sex differences in asthma: A key role of androgen-signaling in group 2 innate lymphoid cells. Front Immunol. 8, 1069 (2017).
Massaro, D. & Massaro, G. D. Estrogen receptor regulation of pulmonary alveolar dimensions: alveolar sexual dimorphism in mice. Am. J. Physiol. Lung Cell Mol. Physiol. 290, L866–L870 (2006).
Ekpruke, C. D., Alford, R., Parker, E. & Silveyra, P. Gonadal sex and chromosome complement influence the gut microbiome in a mouse model of allergic airway inflammation. Physiol. Genomics 56, 417–425 (2024).
Gao, A. et al. Sexual dimorphism in glucose metabolism is shaped by androgen-driven gut microbiome. Nat. Commun. 12, 7080 (2021).
Markle, J. G. et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339, 1084–1088 (2013).
LaBouyer, M. et al. Higher total faecal short-chain fatty acid concentrations correlate with increasing proportions of butyrate and decreasing proportions of branched-chain fatty acids across multiple human studies. Gut Microbiome (Camb.) 3, e2 (2022).
Sun, Q. H. et al. Sex-based differences in fecal short-chain fatty acid and gut microbiota in irritable bowel syndrome patients. J. Dig. Dis. 22, 246–255 (2021).
Stapleton, S., Welch, G., DiBerardo, L. & Freeman, L. R. Sex differences in a mouse model of diet-induced obesity: the role of the gut microbiome. Biol. Sex. Differ. 15, 5 (2024).
Lambrecht, B. N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Carr, T. F., Zeki, A. A. & Kraft, M. Eosinophilic and noneosinophilic asthma. Am. J. Respir. Crit. Care Med 197, 22–37 (2018).
Irvin, C. et al. Increased frequency of dual-positive TH2/TH17 cells in bronchoalveolar lavage fluid characterizes a population of patients with severe asthma. J. Allergy Clin. Immunol. 134, 1175–1186.e1177 (2014).
Liu, W. et al. Mechanism of T(H)2/T(H)17-predominant and neutrophilic T(H)2/T(H)17-low subtypes of asthma. J. Allergy Clin. Immunol. 139, 1548–1558.e1544 (2017).
Choy, D. F. et al. TH2 and TH17 inflammatory pathways are reciprocally regulated in asthma. Sci. Transl. Med 7, 301ra129 (2015).
Doyle, A. D. et al. Homologous recombination into the eosinophil peroxidase locus generates a strain of mice expressing Cre recombinase exclusively in eosinophils. J. Leukoc. Biol. 94, 17–24 (2013).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Kimura, I. et al. The gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 4, 1829 (2013).
Dyer, K. D. et al. Functionally competent eosinophils differentiated ex vivo in high purity from normal mouse bone marrow. J. Immunol. 181, 4004–4009 (2008).
Acknowledgements
The authors thank the scientific contributions and technical assistance by Dr. Yongsuk Hur at the EM & Histology Core Facility and Ms. Jiye Kim at the NGS Core Facility in the BioMedical Research Center, KAIST. This study was supported by grants from Korea Advanced Institute of Science and Technology and the National Research Foundation of Korea (2022R1A6A3A1306363412 to JY; RS-2024-00439160 and RS-2025-02232977 to YMK).
Author information
Authors and Affiliations
Contributions
Y.M.K. and J.Y. conceived the project and wrote the manuscript with comments from all authors. J.Y. performed the overall experiments and sequencing data analysis. S.K. and J.E.P. conducted single-cell RNA sequencing data analysis. H.S.S. performed confocal microscopy. J.K. and W.S. provided experimental support. I.K. provided critical resources. H.Y.K. offered technical advices on in vivo experiments and critical input on data interpretation. Y.M.K. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Francis Luscinskas, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yu, J., Kim, S., Song, HS. et al. GPR43 in eosinophils suppresses the emergence of pathogenic Siglec-Fhi neutrophils in allergic airway inflammation in mice. Nat Commun 16, 10548 (2025). https://doi.org/10.1038/s41467-025-65573-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65573-7