Abstract
Asian people are under-represented in population-based, clinical, and genomic research. To address this gap, we have initiated the Health for Life in Singapore (HELIOS) longitudinal cohort study, comprising comprehensive behavioural, phenotypic, and genomic measurements from 10,004 Asian men and women of Chinese, Indian or Malay background. Phenotyping has been carried out using validated approaches, that are internationally interoperable. Health record linkage enriches both baseline phenotyping and evaluation of prospective outcomes. The integrated multi-omics data include whole-genome and RNA sequencing, quantification of DNA methylation, and metabolomic profiling. Our data reveal extensive lifestyle, physiological, genomic, and molecular diversity between the distinct Asian ethnic groups, and the biological interconnectivity between functional layers. This includes characterisation of divergent patterns of genome regulation between Asian individuals, that correlate with differences in educational attainment, dietary quality, and adiposity, and which overlap transcription factors and DNA methylation sites linked to the development of diabetes and other chronic diseases. Our unique HELIOS Asian Precision Medicine cohort study represents a state-of-the-art platform to enable biomedical researchers to understand the aetiology and pathogenesis of diverse disease outcomes in Asia, and to generate insights that have the potential to improve health outcomes for Asian populations globally.
Similar content being viewed by others
Introduction
Age-related chronic diseases such as diabetes, cardiovascular and respiratory diseases, cancer, and cognitive decline are leading causes of morbidity and mortality across all regions of the world1. Driven by demographic transitions, increasing urbanisation, and adoption of unfavourable lifestyle choices, the Asia Pacific region in particular is facing a rapid increase in chronic disease burden that contrasts stable or falling disease rates in Europe and North America2. There are already 296 million Asian people living with diabetes, and this is expected to rise to 412 million by 20453. Cardiovascular disease (CVD) deaths in Asia have nearly doubled from 5.6 million in 1990 to 10.8 million in 20194. Addressing the rising burden of these major chronic disorders in Asian populations is a high priority for national and international stakeholders, including policymakers and healthcare providers.
Chronic diseases are complex and multi-factorial, and arise through the interaction of lifestyle, environment, and genomic factors5. Longitudinal population studies, in which people are characterised at baseline and followed up over time for health outcomes, play a unique role in identification of the proximal and upstream processes and aetiological mechanisms underlying chronic disease. However, existing prospective cohort studies with comprehensive phenotypic and particularly genotypic measurements are predominantly based on populations of European ancestry6,7,8. This not only represents an important global health inequity, but also a major opportunity for discoveries relevant to the health of the ~4.8 billion Asian people living worldwide9,10.
Singapore, a city-state in Southeast Asia, is home to 5.6 million people, most of whom are of Chinese, Malay, or Indian ancestries. The presence of these three population groups living side-by-side, provides a unique opportunity to explore the diverse lifestyle and genetic profiles of people from East Asia, Southeast Asia, and South Asia, and to relate these to health trajectories. Endowed with highly advanced healthcare and research infrastructure, Singapore is ideally positioned to advance precision medicine and population health research, relevant to global Asian communities.
Here we describe the motivation, design, and early results of the Health for Life in Singapore (HELIOS) Study, a longitudinal population resource focussed on understanding the diseases and health states that are important to Asian populations. We show how the HELIOS Study combines state-of-the-art clinical, molecular, and genetic epidemiological approaches, enriched with information derived from national health data, and highlight the extensive opportunities for transformative research. Our companion papers describe specific discoveries and innovations achieved using the study data, including findings directly relevant to health outcomes of people living in Asia11,12,13.
Results
We recruited 10,004 Asian men and women aged 30 to 84 years to the HELIOS Study between 2018 and 2022 (www.healthforlife.sg). Participants were recruited from the Singapore general population11,13,14. The cohort includes 6784 people who identified as Chinese or other East Asian background, 1807 people who are of Indian or other South Asian background, and 1354 people of Malay or other South-East Asian heritage. There were 59 participants from other ethnicities (Supplementary Data 1). Compared to national averages in Singapore, our cohort has a higher proportion of women, lower cigarette smoking rates, and greater total years in education amongst study participants. However, the prevalence rates for diabetes, hypertension, hypercholesterolemia and obesity among HELIOS participants approximate those reported in national statistics, indicating a minor healthy cohort effect overall (Supplementary Data 2). Figure 1 briefly summarised our study design.
DEXA dual energy X-Ray absorptiometry, ECG electrocardiogram, OCT optical coherence tomography, OCTA optical coherence tomography angiography. This figure was created in BioRender https://BioRender.com/gtyffu3.
Disparities in health outcomes amongst Asian populations in Singapore
Despite similar age and sex distributions, our three Asian ethnic groups exhibit distinct profiles for health, including disease burden and distributions for clinically relevant exposures and endophenotypes. We highlight that Indian and Malay participants have a higher prevalence for hypertension, obesity, and type 2 diabetes (T2D), compared to Chinese participants, and a higher frequency of symptoms for depression and anxiety. Waist circumference, waist-to-hip ratio, and visceral fat mass are also highest in Indian and Malay people (Supplementary Data 1). This is accompanied by increased levels of triglycerides, haemoglobin A1c (HbA1c), fasting plasma glucose, insulin, C-reactive protein (CRP) and other traits related to adiposity and insulin resistance (Supplementary Data 1). The health disparities in major chronic conditions observed between the ethnic groups in our study mirror the health differences seen between the populations of China, Malaysia and India (Supplementary Data 3). We also find evidence for differences in healthcare reach, across a wide range of actionable disease diagnoses. For example, compared to Chinese, Indians and Malays were more likely to have undiagnosed diabetes, while undiagnosed osteoporosis was common in all ethnic groups (Supplementary Fig. 1). These illustrations provide insights into the potential health gains that might be achieved through improved uptake and reach of healthcare interventions in our multiethnic Asian population.
Behavioural and upstream exposures relevant to chronic disease in Asian populations
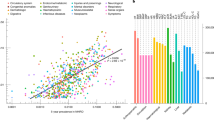
Our study design enables exploration of ‘upstream’ behavioural, environmental, and social factors relevant to health in Asian communities. As an initial illustration, we show that density of food related amenities and the ratio of public to private housing correlate closely with the prevalence of diabetes in our cohort, a key exemplar of major chronic disease risk in the population (r = 0.5 and 0.5, respectively; P < 0.05, Fig. 2). Self-reported food intake of study participants shows divergent consumption of food items, nutrient composition, and differences in diet quality indices between the ethnic groups, that align closely with traditional Asian dietary habits (Fig. 3a–d). While diet quality scores are associated with multiple cardiovascular and metabolic phenotypes within our Asian population groups, we highlight that dietary habit does not fully explain the differences between the ethnic groups. For example, while ‘favourable’ DASH (dietary approaches to stop hypertension) dietary quality score is highest amongst Indians, this directly contrasts with their high rates of obesity and adverse metabolic profiles compared to Chinese participants (P < 2 × 10-6). Similarly, both self-reported and accelerometer-based objective measurements identify that total physical activity is higher, amongst Indians and Malays, despite their unfavourable patterns of adiposity and metabolic performance (Fig. 3e–h and Supplementary Data 1, P < 2.2 × 10-16). Our data thus reveal striking variation between communities in behavioral, environmental, and social factors important for health. Interestingly, while our results confirm expected relationships with key clinical traits within ethnic groups, they do not fully explain health differences between populations. Our observations provide a strong motivation for deeper clinical and molecular epidemiological research focused on Asian populations.
a The Singapore population density across planning area overlayed with HELIOS study participant postcode; b The distribution of diabetes cases; c The distribution of food amenities, including hawker centres, shopping malls, restaurants, and convenience stores; and d the ratio of public/ private residence, overlayed with public parks, connectors, and nature reserves. The built environment has impact health outcomes, as illustrated by the correlations of diabetes case proportions and e the density of food amenities/ area, and f the public/private residence ratio, weighted by population numbers.
a Longitudinal changes in macronutrient trends nationally from 1998 to inception of HELIOS study in 2022 (n = 10,004). Ethnic variations in b macronutrients (proportion to total energy intake, n = 8119), c diet quality represented by a modified DASH score (range from 0 for low quality to 35 for high quality, data points represent mean DASH score; error bars represent standard errors, n = 8119) and d top 20 FFQ foods (servings/day) significantly different across ethnicities within HELIOS study. e Physical activity (data points represent mean total MET-minutes per week; error bars represent standard errors, n = 8119) and f Accelerometer-based physical activity according to the levels of self-reported physical activity (R = 0.23, two-sided p = 1.7 × 10-11 from Kruskal-Wallis rank sum test). g The proportion of people who meet the WHO guideline of physical activity by ethnicity and the proportion of physical activity across different domains (n = 7853). h The relationships between lifestyle factors and cardiometabolic phenotypes are heterogeneous across ethnic groups. Data points represent the coefficients derived from linear regression model; error bars represent 95% confidence intervals, n = 7307. The associations of DASH with DBP were similar to those with SBP, the effect size were: β(SE)DASH->SBP = −0.04 (0.03) in Indian group, compared to −0.02 (0.01) and −0.01 (0.03) in Chinese and Malay groups). DASH dietary approaches to stop hypertension, IPAQ International Physical Activity Questionnaire, MET metabolic equivalent of task, MUFA monounsaturated fat, PUFA polyunsaturated fat, SE standard error, SFA saturated fat, WHO World Health Organization.
DNA sequence variation and functional genomic diversity
The disparate clinical profiles across the three ethnic groups are mirrored by extensive and structured covariation in molecular genotypes. Whole-genome sequencing (30x depth) reveals 252 million variants, including 239.7 million autosomal variants with 206.1 million Single Nucleotide Polymorphisms (SNPs) and 33.6 million short indels (Fig. 4a). Principal Component Analysis (PCA) and admixture analysis helped identify and cluster the dataset into three distinct population clusters corresponding to people of Chinese, Indian or Malay ancestry, as well as individuals who were admixed (Fig. 4b and Supplementary Fig. 2). The majority of variants identified are rare (minor allele frequency, MAF < 1 %; 95% and 90% of autosomal SNPs and short indels respectively), while greater than 50% of variants were observed only in one of the populations. Functional annotation using Variant Effect Predictor (VEP)15 identifies 1,530,701 coding variants anticipated to impact protein structure (Supplementary Data 4). Among the coding variants, 1,373,981 are non-synonymous SNPs, 47,976 are stop codon mutations (stop-gain and stop loss), and 108,744 are indels (frameshift and non-frameshift mutations, Fig. 4a).
a Number of variants annotated in each functional coding mutation category, by ancestry. The darker shades indicate unique variants observed only in the specific ancestry. b 2-dimensional PCA genomic variants by ancestry group. c Distribution of PRS scores for six complex traits in the three major ancestry groups. Comparison across the three ancestries was done using two-sided ANOVA test. d Forest plot displaying association PRS scores with hypertension (n = 8237), obesity (n = 8269), type 2 diabetes (n = 8244), coronary artery disease (n = 8261), depression (n = 8361) and osteoporosis (n = 8261) by ancestry group, and overall [C: Chinese, M: Malay, I: Indian]. Data points represent odds ratios; error bars represent 95% confidence intervals. Two-sided P-value was calculated using logistic regression. ANOVA Analysis of Variance, PCA Principal Component Analysis, PRS Polygenic Risk Scores.
The genetic separation between the ancestral groups provides the basis for population specific sequence variation and allelic heterogeneity with potential clinical relevance, as well as opportunities to reveal new biological relationships12. As an exemplar, we show that carrier frequencies for actionable pathogenic variants in the three primary Familial Hypercholesterolaemia genes (APOB, PCSK9 and LDLR) differ between the three ethnic groups, and also show allelic heterogeneity (Supplementary Fig. 3 and Supplementary Data 5). The frequency of ClinVar supported pathogenic variants is highest amongst Chinese individuals (0.82%) compared to Malay (0.31%) and Indians (0.13%). This difference is reduced by inclusion of predicted pathogenic variants, suggesting under-representation of Malay-specific variants in curated databases, and consistent with the limited inclusion of South East Asian populations in global genomic datasets.
Polygenic risk scores (PRS) confirm a strong relationship of genetic variation with quantitative traits and complex diseases, overall and in each of the three Asian ethnic groups, with little evidence for heterogeneity of effect (Phet > 0.05, Fig. 4). In contrast to their three-fold difference in risk of T2D, mean values for trans-ethnic T2D PRS are comparable between the Asian subgroups (Mean [SD]: Chinese, −0.03 [0.98]; Malay, 0.03 [0.99]; Indian, 0.09 [1.07]; ANOVA P = 0.0001; Fig. 4 and Supplementary Data 6). Similarly, there is no systematic difference in effect allele frequency, or weighted effect allele frequency between the populations (Supplementary Fig. 4). Based on the difference in T2D PRS between the populations, and the observed risk ratios for T2D, we estimate that the small increase in PRS observed in Indian and Malay populations accounts for only ~5% of their three-fold excess risk of T2D compared to Chinese participants (Indians: 5.7 [95%CI: −1.3 to 12.6]%; Malay: 3.1 [95%CI: −5.4 to 11.6]%, Supplementary Methods). Although we cannot exclude residual confounding based on differences in LD structure, our results suggest that the genetic factors identified by current trans-ethnic genome-wide association studies do not make a material contribution to the higher risk of T2D observed amongst Indian and Malay individuals, compared to people of Chinese ancestry.
Quantification of DNA methylation in genomic DNA from whole blood (N = 837,722 CpG sites), as a marker of genomic regulation, reveals 16,444 unique CpG sites that are highly differentiated between the three Asian ethnic groups (P < 2.9 × 10-8). These population specific methylation disturbances are enriched for location in DNase hypersensitivity sites (DHS), histone marks, enhancer and promoter regions, indicating ethnic specific patterns of genome regulation (Supplementary Fig. 5a and Supplementary Data 7). The population-stratified methylation markers are enriched for location in the binding sites for specific, documented transcription factors across multiple cell lines (Supplementary Fig. 5b and Supplementary Data 8). These include Pleiomorphic Adenoma Gene 1 (PLAG1) and Eleven-Nineteen Lysine-Rich Leukaemia Protein (ELL) (both P < 10-4, Supplementary Data 9). PLAG1 is a nuclear transcription factor subject to maternal imprinting16, and which is implicated in pancreatic genesis, insulin secretion, and diabetes in neonates and adult organisms. We also note that the ethnically divergent methylation patterns strongly overlap CpG sites that predict future diabetes, providing evidence for nuclear regulatory disturbances that may contribute to the divergent metabolic outcomes observed between ethnic groups (Supplementary Fig. 5c).
We used PCA to explore potential processes driving genome regulation in the population. We show that the perturbations in DNA methylation are enriched for association with multiple aspects of dietary intake, including the Alternate Mediterranean Diet (aMED), DASH and the Alternative Healthy Eating Index (aHEI) diet quality scores, adiposity and cardiometabolic health, based on directly measured exposures (P < 0.0015, based on Bonferroni correction for an estimated 32 independent phenotypes, Fig. 5, Supplementary Fig. 6, and Supplementary Data 10), supporting an important effect of environmental exposures on genomic regulation, independent of genetic effects. The perturbations in DNA methylation were also associated with educational attainment and adiposity based on genetically inferred exposures. Directly measured and genetically inferred relationships of adiposity with methylation were highly concordant. Smoking and alcohol intake were not associated with methylation PCs in our samples, but we note that these exposures are of low prevalence in our Asian population (Supplementary Data 11). Our results thus shine new light on the fundamental roles that these key modifiable social, behavioural, and physiological factors play, as primary, interlinked drivers of genomic regulation and health outcomes in diverse human populations.
a–c Display the methylation diversity for individuals from the three ethnic groups (PC1 on x axis, and PC2 on y axis). Overlaid are the effect sizes of the measured and genetically inferred exposures that are associated with PC1 and PC2 of the DNA methylation (two-sided P < 0.0015 multiple testing correction for 32 independent tests; all exposures are Z-standardised to enable comparison of effect sizes); a clinical traits; b dietary exposures assessed objectively by circulating metabolites. Specifically, fermented rice/lentil items refer to idli and thosai. c Polygenic Risk Scores (PRS). The beta weights for PC1 and PC2 are scaled along the top x-axis and the right y-axis respectively. The results identify the directly measured and genetically inferred exposures that may relate to population level epigenetic variation between Asian ethnic groups. d The effect sizes and the directions for the beta coefficients derived from regression analysis of measured and genetically inferred exposures on PC1 to PC5 (Variance explained as percentage of top 100 variance components; PC1 – 17.6%, PC2 – 9.9%, PC3 – 5.1%, PC4 – 2.2%, PC5 – 2%, two-sided P < 0.0015 multiple testing correction for 32 independent tests). *P < 0.05, **P < 0.0015. CIMT carotid intimamedia thickness, PCA Principal Component Analysis, WHR waist hip ratio.
Metabolic variation in Asian populations
Metabolomic profiling of plasma by high-throughput semi-quantitative mass spectrometry enabled us to quantify plasma concentrations of 1073 discrete metabolites. We show that dietary patterns of our Asian participants intersect closely with their metabolic variation, enabling identification of metabolite sets that are representative of Asian dietary patterns; these associate closely with perturbations in regulatory pathways, and predict multiple chronic diseases13. We also find that 153 of the 1073 plasma metabolites characterized show marked divergence between all three Asian ethnic groups (P < 1 × 10-5, Fig. 6a and Supplementary Data 12); of these, 128 metabolites are of known identity. In general, amongst the 153 highly differentiated metabolites, Indians and Malays had lower levels of lipid metabolites, independent of cholesterol lowering treatment, and higher levels of amino acids and nucleotides compared to Chinese (Fig. 6b, Supplementary Data 12, and Supplementary Data 13). 63% of lipid metabolites were inversely associated with the presence of hypertension, obesity, T2D, or CVD (P < 4.7 × 10-5), and 16% were inversely associated with all four phenotypes. Age, sex, genetic ancestry, diet, and body mass index (BMI) were each determinant of plasma concentrations for the highly differentiated metabolites (Fig. 6c), but with substantial differences in their contribution on a metabolite specific basis. For example, BMI accounted for 18% of the variation in glutamate, while age accounted for 20% of the variation in the androgenic steroid dehydroepiandrosterone sulphate. Strong effects for genetic ancestry on metabolic variation were seen for 1-margaroyl-2-arachidonoyl-GPC (17:0/20:4), a phosphatidylcholine derived from eggs, fish, and meat17,18,19. We show that concentrations of this metabolite are positively associated with self-reported intakes of red meat (P = 5.3 × 10-57), fish (P = 6.5 × 10-38), dairy (P = 1.4 × 10-20), and poultry (P = 1.1 × 10-17). Levels are also associated with chapati consumption, which is common in Indian communities (P = 1.6 × 10-3). Circulating 1-margaroyl-2-arachidonoyl-GPC levels are strongly influenced by genetic variants in the FADS1/FADS2 gene cluster, a highly pleiotropic region that is linked to multiple lipids, cardiometabolic, inflammatory traits, skin diseases and pregnancy outcomes20. FADS1/FADS2 variants are also known to be stratified between Asian populations, and recognised to influence metabolic responses to dietary intake, and may provide the basis for genomically determined ‘Precision Nutrition’20. Our observations further highlight the important roles for both genetic and lifestyle factors in driving divergent metabolite profiles and health outcomes amongst Asian people.
a PCA plot of 153 significantly differentiated metabolites across three age-sex matched ethnic cohorts of 1146 individuals each (The first two PCs explain 23% of variation). Selection criteria for 153 metabolites include: 1) significantly associated with ethnicity in the discovery cohort (70% participants, two-sided linear regression analysis P < 1 × 10-5 after multiple-test correction), and 2) significantly associated with ethnicity in the test cohort (30% participants, two-sided test P < 0.05 and same direction of estimates as in the discovery cohort). b Circos plot of 128 well-characterized and known metabolites (in sequence from outermost to innermost layer): 1) metabolite super-pathways, 2) significant associations with HT, Obs, T2D, and CVD, denoted by a black dot, 3) estimates of regression coefficient for association with ethnicities (CI: Indian compared to Chinese, CM: Malay compared to Chinese, IM: Malay compared to Indian). Curved lines at the centre highlight significant pairwise correlation between metabolites. Grey lines represent pairwise correlations within the same super-pathway; blue lines represent pairwise correlations across sub-pathways but within the same super-pathway; green lines represent pairwise correlations across super-pathways. c Violin plot showing contribution (as partial r-squared values) of age, dietary PCs, BMI, sex, and genetic PCs on variation of plasma abundance of 153 metabolites in 7546 individuals, with the outline indicating kernel probability density, the width of the shaded area represents the proportion of data located there. The embedded boxplot shows the median and interquartile range (box), with whiskers extending to 1.5 times the interquartile range. The inset plot zooms in on the partial r-squared distribution between 0.0–0.1. BMI body mass index, CVD cardiovascular disease, HT hypertension, Obs Obesity, PCA Principal Component Analysis, T2D type 2 diabetes.
Diversity of gene expression in Asian populations
Quantification of gene expression from whole blood using RNA Sequencing (N = 12,434 genes) identified 2972 genes that are differentially expressed across the three ethnic groups (P < 1.34 × 10-6, Fig. 7a–c, Supplementary Data 14). 13 genes showed differential expression across all three pairwise comparisons. These include ASNS, NSF and SCRN1 (Effect size (average difference in gene expression per group measured in standardized transcripts per million [TPM]): 0.22 to 0.36, P < 1.34 × 10-6). Gene set enrichment analysis identified that these 2972 stratified genes are enriched in inflammatory processes, as well as thermogenesis and COVID-19 susceptibility (FE: 1.9 – 3.3, P < 1.5 × 10-4, N = 34–82 genes, Fig. 7d and Supplementary Data 15). We also observed enrichment for neurological disease associated gene sets such as Amyotrophic lateral sclerosis, Alzheimer’s disease, Huntington disease and Parkinson’s disease (FE: 1.67–1.75, P < 8 × 10-6, N = 66–94 genes, Supplementary Data 15). Genomic association analysis of the 2972 genes differentially expressed between populations identified that 1177 are influenced by cis-acting genetic variants significantly (P < 3.1 x 10-9). Amongst these, we identify cis-eQTLs for 76 genes (44 protein coding) that are not reported in the largest publicly available blood based eQTL database, eQTLgen21. The strongest identified eQTLs were for NBPF8 (P = 2.57 × 10-131) and TMEM126A (P = 9.74 × 10-63, Fig. 7e). We also identify eQTLs for TXNIP (rs7212, P = 1.97 × 10−15, Fig. 7e), encoding a major regulator of cellular glucose uptake. Epigenomic regulation of TXNIP is closely associated with future T2D in Asian populations22,23. Our observations further highlight the opportunities to reveal new biological insights through our multi-molecular Asian Precision Medicine datasets.
Volcano plot of differential gene expression across the three Asian populations in a pairwise comparison between Chinese and Indians (a), Chinese and Malays (b), and Indians and Malays (c) after adjusting for Age, Sex, RIN and top six PEER factors. The x-axis is the effect size estimate of association per standard deviation change in gene expression and the y-axis is the two-sided log10P value of the association from the regression model. d Gene set enrichment analysis of the 2972 unique differentially expressed genes in at least one pairwise comparison across populations. Enrichment was performed using gprofiler2 tool in three pathway databases (Kyoto Encyclopedia of Genes and Genomes (KEGG), Reactome (REAC) and Wiki Pathways (WP). Each dot is an independent gene set and the y-axis is the one-sided hypergeometric P-value of enrichment per gene set. e Regional plots of three cis-eQTLs identified in the HELIOS cis-eQTL analyses in 1228 individuals. The first two plots are the top two strongest associations (NBPF8 and TMEM126A) and the last one is for the TXNIP gene identified as an important gene linked to diabetes via DNA methylation in the Asian population. The y-axis in the regional plots represents the –log₁₀ of the two-sided P-value from the cis-eQTL regression analysis.
Potential for discovery through molecular epidemiological studies of Asian populations
The clinical, molecular, behavioural, and environmental diversity between the Asian ethnic groups provides robust new opportunities for discovery relevant to human biology and health outcomes. To illustrate this, we carried out genome-wide association of the 153 ethnically diverse plasma metabolites. We identify 365 independent genetic variants in 140 genomic loci, that are significantly associated with 113 metabolites at a genome wide significance threshold (P < 5 × 10-8) (Fig. 8a). We observe a strong degree of genetic pleiotropy at multiple loci, in particular the FADS1/FAD2 gene locus which was associated with 39 metabolites (Fig. 8b). Summary-data-based Mendelian Randomisation (SMR) analysis of metabolites with cis-eQTLs identified 1176 significant gene-metabolite pairs after multiple testing correction (P < 4 × 10-5), comprising 585 genes and 104 metabolites (Supplementary Data 16). We were able to replicate 166 gene-metabolite associations and identify 51 additional associations using cis-eQTL information obtained using the HELIOS transcriptomics data (Supplementary Data 16 and Supplementary Data 17). Colocalization analysis reveals shows that 79 of these gene-metabolite pairs are likely to share a common causal variant (coloc-H4 P > 0.7). This includes the finding that plasma concentrations of dopamine 3-O-sulfate are influenced by genetic variants at the cis-eQTL locus for SMAD5, a transcriptional regulator protein involved in the TGF-Beta pathway (Fig. 8c) and implicated in the development of dopaminergic neurones24. Similarly, variation in plasma levels of metabolite X-11381 is determined by genetic variation found at the cis-eQTL locus for Nephrocystin 4 (NPHP4, Fig. 8d), which plays an important role in renal tubular development and function. X-11381 is also associated with raised blood pressure and cardiovascular disease in our cohort (false discovery rate, FDR– P < 0.05; Supplementary Data 12). Our rich multi-omics data thus provide multiple opportunities to improve understanding of the molecular pathways influencing metabolic performance and other pathways leading to chronic disease in Asian populations.
a Manhattan plot for summary of all associations between plasma metabolite levels and genetic loci (N = 8234). Only genetic variants with P < 5 × 10-8 (two-sided test in linear mixed regression model) are coloured based on the strongest associated metabolite group for the specific variant. Genes for top 50 loci identified through SMR are annotated. b Distribution of number of associated metabolites per locus, demonstrating the pleiotropy of genetic effects on metabolites. The loci with at least 5 associated metabolites are annotated with the SMR associated gene. c Regional plot highlighting the shared causal variant between Dopamine 3-O-sulfate and SMAD5. d Regional plot highlighting the shared causal variant between X-11381 and NPHP4. The y-axis in the regional plots represents the –log₁₀ of the two-sided P-value from the GWAS analysis. GWAS Genome-Wide Association Study, SMR Summary-data-based Mendelian Randomisation.
Linkage to national health and administrative records
With participant consent, we link HELIOS research phenotypic data securely to their national health data, using the NRIC, a unique national identifier that is held by Singaporean citizens and Permanent Residents. De-identified linked research, health and administrative data were made available through the Trusted Research and Real-World Data Utilisation and Sharing Tech (TRUST) platform (https://trustplatform.sg). National Health and Administrative Records were identified for 95% of study participants, and include national disease registry records, disease diagnoses, national insurance claims, medications, laboratory tests, radiology, surgical procedures, and death registry records from 1998 to 2020. The linked national health and administrative records for our 10,004 participants include 1.6 million laboratory test results, 776,505 prescriptions and 131,211 diagnostic episode codes. Using diabetes as a case study, we show that the national health data recapitulate age stratified, ethnic disparity disease risk, and enable identification of incident diabetes cases, with greatest risk amongst participants who are older, obese and impaired fasting glucose (Fig. 9). These linked national data thus provide deep opportunities to extend baseline health assessment of participants, and to identify future health trajectories, including incident disease.
CI confidence interval, HDL High-density lipoproteins, NHD national health data, OR odds ratio, WHR Waist hip ratio.
Reproducibility of measurements
We demonstrate the reproducibility of our research phenotypic characterisation, by carrying out repeat assessment of 398 participants, one year after enrolment (range 58 to 1073 days). We show moderate to strong intra-class correlations for measures made, in all domains of assessment, a performance that is similar to or better than those reported by UK Biobank25 and other major population studies26. In general, objective physiological measurements were more reproducible than self-reported lifestyle and cognitive measurements (Fig. 10 and Supplementary Data 18). High data completeness and reproducibility further support the validity of our unique multiethnic Asian dataset.
Spearman correlation analyses were used to calculate the correlation coefficients between z-scored baseline and retest measurements (N = 398). Correlations are presented as box plots, where the upper bound marks the 75th percentile, and the lower bound marks the 25th percentile. BP blood pressure, DEXA dual-energy X-ray absorptiometry, ECG electrocardiogram, FFQ food frequency questionnaire, HR heart rate, IPAQ International Physical Activity Questionnaire.
Discussion
Asian populations are widely recognised to be under-represented in global genomic and health-related research cohorts, compared to their European counterparts. This represents an important impediment to identification of the population specific behavioural, environmental, genomic, and molecular exposures and processes that impact Asian health. The limited ethnic diversity of existing population studies also represents a major obstacle to the development of effective and evidence-based approaches for accurate diagnosis and therapeutic intervention, that address the health needs of Asians.
To advance beyond current state-of-the-art, we have established the HELIOS Study, a deeply phenotyped, longitudinal population cohort comprising 10,004 men and women from the multi-ethnic Asian population of Singapore. Our participants underwent extensive clinical, behavioural, environmental, and molecular characterisation, adopting techniques that are validated, aligned to best practices, and directly interoperable with international precision medicine cohort studies. The HELIOS Study includes people of Chinese (East Asian), Malay (Southeast Asian) and Indian (South Asian) background with heterogeneous lifestyle behaviours, genetic background and health outcomes. The inclusion of these three major Asian ethnic groups provides an opportunity for precision medicine research, that has the potential for relevance beyond Singapore, and across the wider Asia-Pacific region. The clinical characteristics of the cohort are broadly representative of the population from which they were recruited and are notable for the high rates of diabetes and related metabolic disturbances, that are recognised to be highly prevalent amongst Asian people.
We observed patterns of dietary quality in Indian Asians, compared to Chinese and Malay participants that are anticipated to be favourable for health outcome. This contrasts with the adverse cardiometabolic health outcomes, including diabetes and coronary artery disease that are well documented amongst people from Indian ethnic groups. This implies that dietary scores such as aHEI, DASH and aMED, that have been developed in North American and European settings may not accurately identify disease outcomes amongst Asians populations. Our observation illustrates the need for Precision Nutrition studies that are focussed on Asian communities, and the potential opportunities for transformational insights from the HELIOS programme of research. Our multi-ethnic, longitudinal population study will enable us to bring better understanding of behavioural choices in Asia, and their links to health outcomes. As an example, our complementary studies using the HELIOS data, show how metabolite variation provides a quantitative measure of dietary habit, and the potential for an objective framework that can support Precision Nutrition in Asian populations13.
Whole genome sequencing demonstrates the genetic diversity of the population. While the majority of individuals cluster in one of the three main ancestral groups, there is also evidence for recent population admixture between each of these three groups. We show that known genetic variation identified by large trans-ethnic genome-wide association studies, does not explain the three-fold excess risk of T2D in Indian and Malay individuals. Nevertheless, the unique multi-ethnic population genetic architecture provides the basis for the presence of functionally and clinically relevant DNA sequence variation, that is specific to Asian subgroups. We observed a larger number of unique SNPs and short INDELs in the Chinese population, compared to the other two groups, which is likely attributed to the larger sample size for people of Chinese ancestry in the cohort (~68%). Characterisation and interpretation of this population genetic variation are anticipated to provide opportunities for new discoveries relevant to disease aetiology and are also essential prerequisites for the application of genomic medicine in Asian populations.
Our comprehensive characterisation of study participants is specifically designed to capture a wide spectrum of exposures relevant to health, as well as to reveal the systems biology that links these exposures to phenotypic variation and health outcomes in Asia. In keeping with this approach, we demonstrate the presence of variation in genome regulation and metabolic performance between the three Asian ethnic groups. For example, we identify extensive, ethnic-specific perturbations in DNA methylation that intersect with PLAG1, a nuclear transcription factor linked to pancreatic biology and diabetes27, which overlay CpG sites that are linked with obesity and diabetes28, mirroring the divergent metabolic outcomes between ethnic groups. Stratified genomic regulation correlates with socio-economic factors, population specific dietary habits, physical activity, adiposity, and genetic diversity; these observations shine new light on the fundamental roles that social, behavioural, and inherited factors play as interlinked drivers of health outcomes in diverse human populations. We further use metabolomic profiling to demonstrate extensive variation in metabolite concentrations between our Asian groups, that reflects their specific patterns of diet, the differing levels of adiposity, the presence of genetic variation and transcriptional control. In parallel, metabolite profiling of our unique Asian population cohort has enabled identification of previously unrecognised pathways underlying cholesterol transport and cardiovascular risk, and potential opportunities for novel therapeutic approaches to cardiovascular disease prevention12. We note that for downstream analysis such as SMR and Colocalization with gene expression data, the publicly available large-scale datasets such as eQTLGen are dominated by samples from individuals from European ancestry, which could lead to biased results. To address this potential concern, we show replication of the majority of the metabolite-gene associations using eQTL data generated from our Asian participants of the HELIOS Study.
Linkage to national health data relevant to health provides powerful opportunities to enrich baseline phenotyping of participants in population-based cohorts, as well as to identify longitudinal health outcomes efficiently and accurately. Longitudinal population cohorts in Europe and North America have a long tradition of successful record linkage that has accelerated health-related research in these settings. In contrast, record linkage has been uncommon amongst the available Asian population cohorts, reflecting both limited implementation of national health data, as well as the nascent state of regulatory frameworks to enable safe data integration. Here we demonstrate the ability to achieve linkage to health and administrative records amongst the multi-ethnic Asian populations of Singapore, using a secure platform for linkage, deidentification and analysis, hosted by the Singapore Ministry of Health (the TRUST platform). We use this framework to retrieve extensive medication, laboratory, and diagnostic data for our study participants. With diabetes as a use case, we demonstrated the ability to identify accurately people with diabetes both cross-sectionally and prospectively, and to show expected longitudinal risk relationships. This successful approach to secure record linkage is unrivalled in the Asia Pacific region and will be instrumental in advancing the research goals of the study, for the benefit of Asian people living in Singapore and other global settings.
With rich, multi-layered baseline data and long-term follow-up through linkage, the HELIOS Study provides a world class resource for biomedical researchers from a wide range of disciplines, to investigate the behavioural, environmental, genomic, and molecular factors impacting health in Asian populations, with a level of detail that has not been previously possible. The successful approaches to population-based research established in the HELIOS study also provide the blueprint for ongoing efforts to create a precision medicine cohort comprising 100,000 people, the SG100K population study, to enable national efforts to advance precision medicine for Asian populations.
Methods
The HELIOS Study is a prospective population-based cohort, comprising men and women aged 30 to 84 years, living in Singapore (www.healthforlife.sg, https://www.instagram.com/heliossg100k/, https://www.facebook.com/HELIOSSG100K/, IRB approval by Nanyang Technological University: IRB-2016-11-030). Study design was informed by initial pilot studies (N = 184, recruited between January 3, 2018 and March 21, 2018, Supplementary Data 19), which enabled development of community engagement and involvement activities, study protocols and training programs. Participants were recruited between April 2, 2018 and January 7, 2022. Assessment of reproducibility was carried out in 398 participants (recalled between September 3, 2019 and January 28, 2022, Supplementary Data 20 and Fig. 10). The study is a template for future efforts with increased sample size (SG100K, target 100,000 participants).
Recruitment. Study participants were recruited from the Singapore general population through a range of community outreach programs to ensure participation from ethnic minority groups, as well as people across socio-economic groups. Community engagement included language-specific recruitment drives conducted in the worship places, religious associations, and community associations across Singapore; multilingual study advertisement and documents (English, Chinese, Malay, and Tamil); and collaboration with a range of employers and occupational groups. Individuals were excluded if they were pregnant or breastfeeding, or had acute illness, major surgery within the previous 3 months, current participation in a drug trial, or cancer treatment in the past year.
Consent. HELIOS asks permission from participants to use the data and samples that they contribute, for clinical and molecular epidemiological research focussed on improving human health. This includes the application of ‘untargeted’ molecular profiling techniques that assess genomic, proteomic, transcriptional, metabolomic and other ‘omic’ variation in the biological samples collected. Participant consent also includes permission for linkage to disease registers, medical records, social care datasets and other health-related datasets held by Singapore’s public bodies. Linkage is enabled by the Singapore NRIC, a unique national identifier allocated at birth, and with universal coverage. Consent provides permission for use of the data and samples from participants, by both academic and industry researchers, and for recontact of participants, including recontact based on phenotypic or genotypic characteristics. The HELIOS Study operates under the governance framework of the Nanyang Technological University, and with Institutional Review Board approval (Ref: 2016-11-030).
Baseline examination. At enrolment, participants complete a comprehensive physiological, clinical, and behavioural assessment, carried out in a single visit (Supplementary Data 21). The electronic health and lifestyle questionnaires collect demographic, lifestyle, reproductive history, and other potentially health-related information. In addition, a broad range of physiological measurements, including a state-of-the-art imaging module comprised of a 3-D carotid ultrasound, dual energy X-ray absorptiometry (DEXA) scans for bone density and body composition, and comprehensive optical imaging, are performed. These imaging technologies will enable the identification of pre-clinical disease phenotypes that will aid prognostic and preventative research. Participants also complete a physical fitness test and have physical activity monitored using accelerometer devices over a 7-day period. Biological samples (blood, urine, saliva, stool, and skin tapes) are also collected. The assessment process, biological samples collection and storage, quality management, return of assessment findings, ethics and data security are described in detail in Supplementary Methods.
Follow-up. The HELIOS Study will follow up participants long-term to identify any event of interest. This design allows the investigation of the causes and nature history of a broad range of diseases and health conditions. Participants enrolled in the HELIOS Study will be followed up through routine health record linkage, re-contact with participants, Singapore Cancer Registry, Ministry of Health, and Health Promotion Board records, where available, for medical records, ongoing behaviours and built environment exposure.
Analysis of biological samples
Clinical chemistry
This includes assessment of fasting glucose, insulin, and lipid profile, as well as HbA1c and CRP. Fasting glucose, HbA1c and lipid profile were measured from fasting blood samples by the accredited laboratory (QuestLab, Singapore, SAC–SINGLAS ISO 15189:2012). Fasting insulin and CRP were measured with immunoassays using the ADVIA Centaur XPT Immunoassay System and ADVIA 1800 Chemistry System, respectively (Siemens Healthcare, Erlangen, Germany).
Whole Genome sequencing (WGS)
Whole genome sequencing was carried using the Novaseq platform, with data processing using DRAGEN v3.7.8. Individual sample Variant Call Format (VCF) files were transformed into HAIL matrix tables29. Multi-allelic sites were efficiently split into multiple rows of bi-allelic sites, ensuring a comprehensive representation of the genetic variation. Samples were merged in batches of 1000, to create a unified HAIL matrix table representing the sample cohort, with 258,062,302 genetic variants. Stringent variant and sample quality control (QC) parameters were employed to ensure the accuracy and reliability of the genomic data. These included a number of q30_bases (threshold 77.5GB high quality bases), as well as ratios for transition /transversion, heterozygous/homozygous variation, and insertion/deletion, applying a threshold of 6x Median Absolute Deviation (MAD) for each. Samples exhibiting more than 1% cross-contamination, call rate <95%, autosomal coverage <95% at 15X, or discordant sex information (reported vs genetically determined) were also flagged. The QC metrics were added as annotations to the HAIL Matrix table, which was then converted to a merged VCF file of 10,000 samples. The VCF file was also converted and stored as PLINK230 binary files to perform downstream analysis.
Methylation profiling
Bisulfite conversion of genomic DNA was performed using the EZ DNA methylation kit (Zymo Research, Orange, CA), with DNA methylation quantified using the Illumina Infinium MethylationEPIC BeadChip® array (EPIC) (Illumina, Inc, CA, USA) according to manufacturer protocols. Bead intensity was retrieved using the minfi package in R, with a detection P < 0.01 used for marker calling. Of the 846,604 positions assayed by the array, we excluded markers with call rates <95% (N = 8882). In total 58 samples were excluded, 2 for array scanning failure, 39 for sex inconsistency and 17 duplicates. None of the samples failed sample call rate (<95%). This left us with 837,722 CpG sites and 2342 samples for analysis. We analysed epigenome-wide data in R using minfi and other R scripts, in accordance with the CPACOR pipeline31. In brief, marker intensities were normalised by quantile normalisation, with white blood cell subsets imputed32.
RNAseq
RNAseq libraries were prepared using samples of whole blood (n = 1234) collected in PaxGene RNA tubes at enrolment. RNAseq libraries were prepared from at least 1 mg of total RNA using NEBNext® Ultra™ II Directional RNA Library Prep (New England Biolabs, Inc.), with GLOBINClear (Thermo Fisher Scientific) for depletion of globin gene RNA and Ribosomal RNA (rRNA). The libraries were sequenced on a NovaSeq6000, using a paired-end run of 2 x 150 bp. We aimed for at least 30 M aligned reads per library (~9 Gb of data). Adapter and quality trimming were performed in TrimGalore33 whereas SortMeRNA was used for the removal of rRNA34. Alignment to the reference genome (GRCh38) was done using STAR version 2.7.9a35, followed by quantification of reads with RSEM version 1.3.336, which identified a total of 60,708 genes. Gender mismatch check was performed by interrogating for anomaly across 5 genes – namely XIST, RPS4Y1, EIF1AY, DDX3Y, and KDM5D. A total of 6 samples had failed this check, resulting in a total of 1228 samples for downstream analysis. Genes with TPM ≥ 1 and read count ≥6 in at least 20% of the samples were retained; resulting in a remaining total of 12,434 genes. Finally, the genes were normalized using the Trimmed Mean of the M-values (TMM) approach37.
Metabolite profiling
The Metabolon Global Discovery Panel was used for untargeted mass-spectrometry-based metabolic profiling of 10,000 fasting EDTA plasma samples. Samples were initially stored at −80 °C, then thawed, aliquoted, and shipped on dry ice to Metabolon. Samples were prepared and extracted for assay using four methods: two separate reverse-phase (RP)/UPLC-MS/MS methods with positive ion mode electrospray ionization (ESI), RP/UPLC-MS/MS with negative ion mode ESI, HILIC/UPLC-MS/MS with negative ion mode ESI. All methods utilized a Waters ACQUITY ultra-performance liquid chromatography (UPLC) and a Thermo Scientific Q-Exactive high resolution/accurate mass spectrometer interfaced with a heated electrospray ionization (HESI-II) source and Orbitrap mass analyzer operated at 35,000 mass resolution. Several recovery and internal standards, and controls (blanks and pooled matrices) were added for quality control (QC) purposes. Experimental samples were randomized across the platform run with QC samples spaced evenly among the injections. Five samples failed Metabolon QC standards and were removed from analysis. Peak area-under-curve was used for metabolite quantification, and data across inter-day batches were normalized by median scaling. Data corresponding to (i) 235 samples from a second visit of the same participant, and (ii) 9 outlier samples (greater than 6 standard deviations from the mean of first and second principal components) were excluded. Metabolites missing in more than 25% of the data were removed, and the remaining were imputed with minimum value, then log-transformed and standardized before further data analysis.
Statistical analyses
Clinical definitions
The clinical conditions were defined as those who self-reported the conditions or met the following criteria: Hypertension: i. the International Society of Hypertension (ISH) threshold: blood pressure ≥140/90 mmHg38; ii. ethnicity-specific threshold: blood pressure ≥123/90 mmHg in Indian39; blood pressure ≥140/90 mmHg in all other groups. Obesity: i. WHO Asian threshold: BMI ≥ 27.5 kg/m40; ii. ethnicity-specific threshold: BMI ≥ 26.9 kg/m2 in Chinese; BMI ≥ 23.9 kg/m2 in Indian41; BMI ≥ 27.5 kg/m2 in Malay; BMI ≥ 30.0 kg/m2 in others. Type 2 diabetes: fasting glucose ≥7.0 mmol/L. Cardiovascular disease: subclinical atherosclerosis defined as presence of atherosclerotic plaque or mean carotid intima-media thickness (cIMT) ≥ 0.8. Depressive symptoms: Patient Health Questionnaire-9 (PHQ-9) score ≥10. Anxiety symptoms: General Anxiety Disorder-7 (GAD-7) score ≥10. Osteoporosis: lumbar spine bone mineral density T-score of −2.5 or below.
Comparison between ethnicities
Comparisons between the ethnic groups were performed using generalized linear models for categorical measurements and linear regression for continuous measurements, adjusting for age and sex. A p-value threshold of P < 0.0011 (0.05/44 comparisons) was applied.
Dietary habit and nutrition
Ethnic variations in dietary intakes (foods and macronutrients) and diet quality (DASH score)42 were assessed using the validated Food Frequency Questionnaire (FFQ)43. Food items were recorded as servings/day, and macronutrients were expressed in kcal/day after accounting for type of macronutrients/serving and weight/portion of each food, and subsequently aggregated to derive % contribution to total daily energy intake. For macronutrients, % macronutrient was scaled and visualised as radar plot (R package fmsb) across ethnicity. For food items, daily servings were log-transformed and analysed using linear regression, adjusted for age, sex and ethnicity, with Chinese being the reference. We applied Bonferroni-Hochberg corrected p-value threshold of P < 1 × 10-100 and selected the top 20 foods significantly higher in Malay and in Indian subgroups. Foods were grouped into 4 categories based on animal source or relevance to ethnicity. For DASH, a modified score was derived from 7 components (fruits, vegetables, wholegrains, nuts and legumes, low-fat dairy, red and processed meats, and sweetened beverages), ranging from 0 (low quality) to 35 (high quality). Difference across ethnicity was analysed using linear regressions.
Physical activity
Physical activity in various domains and intensity levels and sedentary behaviour were derived based on the validated long International Physical Activity Questionnaire (IPAQ)44. Participants with at least 150–300 min of moderate-intensity or at least 75–150 min of vigorous-intensity physical activity or an equivalent combination of moderate- and vigorous-intensity activity throughout the week and with at least 2 or more days on moderate- and vigorous-intensity activities a week were deemed as meeting the WHO guidelines for recommended physical activity45. Accelerometry data were collected as an optional assessment amongst the first 1000 participants in the next phase of the HELIOS Study. Participants wore Axivity AX3 wrist-worn triaxial accelerometer on their non-dominant wrist continuously for 7 consecutive days, including during sleep. Raw accelerometry data were calibrated to local gravitational acceleration46,47 following which movement-related acceleration was expressed using the Euclidean Norm Minus One (ENMO) metric (https://github.com/MRC-Epid). This method has been validated against energy expenditure in free-living conditions46,47 to generate mean Euclidean Norm Minus One (ENMO). The data of 867 out of 1000 participants were suitable for analysis. Comparison across ethnicities were performed using Kruskal-Wallis rank sum and Chi-square tests.
Environment
OneMap APIs (https://onemap.gov.sg) were called within the R environment to generate the latitude and longitude of each participant’s postal code and planning area. All geospatial Singapore data with relevant attribute tables were extracted from the national open data collection (https://beta.data.gov.sg). The extracted tables include planning area, population census by subzones; subzones by type of dwelling; and parks and nature reserves. Open-sourced QGIS v3.32.1 software was used to project geospatial data and population density. Geospatial tags for shops selling food and beverages and shopping centres, as well as amenities for sustenance, were extracted using QuickOpenStreetMap plug-ins. OpenStreetMap IDs representing food amenities (n = 9901) were tagged to the respective planning area in Singapore using OneMap API. To generate bubble plots linking environmental factors with disease outcome, the area (m2) per planning area polygons was calculated to derive population density using sf and lwgeom package.
Annotation of genetic variants
Variants were annotated using the VEP tool15, extracting the most severe consequence for each variant. Potential novel variants present in our dataset were identified after comparing the dbSNP v156 database48 of all reported variants.
Population structure analysis and clustering
To understand the genetic structure and stratify our population, we applied strict filters to the data excluding variants with MAF < 5% (i.e. present in less than ~500 out of the 10,000 genotyped individuals), Hardy-Weinberg equilibrium (HWE) P < 0.1%, sample and variant missingness <2% and removed variants in the MHC region as well as the Chr8 Inversion region. Duplicated samples (n = 3) and samples with reported ancestry labelled as “Others” (non-Southeast Asian [n = 60]) were removed for the current analysis. Linkage Disequilibrium (LD) based pruning was performed for the final filtered data with an LD-r2 of 0.1 within a 200KB window. Genetic relatedness was estimated using the genome function to determine the pi_hat estimate for all pairs of individuals in our dataset. We performed PCA to extract the top 50 genetic PCs from our data. Given the complex structure of our data, we use a data-driven approach to determine and cluster the individuals belonging to specific ethnic groups. The results of the PCA analysis were used to perform K-means clustering (K = 3) to group the individuals into three super populations (Chinese, Indian and Malay). These ancestry labels were used to estimate the ancestry-stratified allele frequency file, which is used as input to run supervised admixture analysis using SCOPE49. The results from the admixture analysis were used to determine the 6 final ancestry clusters using a semi-supervised K-means clustering approach. Additionally, to understand the genetic structure of our data with reference to 1000 genomes50, we merge the LD independent SNPs with 1KG data from four super populations and perform PCA again with the merged set of samples and variants after applying the same filters as above. All the filtering, PCA, LD and relatedness analysis were performed using the PLINK2 tool30 and the k-means clustering was done using R.
Rare variant analysis
We analysed rare variants in three genes strongly linked Hypercholesterolemia namely PCSK9, APOB, and LDLR—using data from the HELIOS cohort. Variants in these genes with a minor allele frequency (MAF) below 1% in at least one of the three ethnic groups were selected and annotated using Ensembl’s VEP15. The most severe transcript consequence was retained per variant. We classified variants into five pathogenicity-based sets: (i) strict reported pathogenic (ClinVar: pathogenic or likely pathogenic, excluding benign/uncertain labels), (ii) permissive reported pathogenic (including uncertain significance), (iii) predicted pathogenic (based on concordance across four in silico scores: SIFT < 0.05, PolyPhen > 0.85, ClinPred > 0.5, and REVEL > 0.5), and two combined sets including both reported and predicted variants (Supplementary Data 5). Carrier burden per gene was defined as the sum of individual variant dosages, and individuals were labelled as carriers if burden > 0. Carrier frequencies were calculated within each group (Chinese, Indian, Malay) as the proportion of individuals carrying at least one qualifying variant divided by the sample size of the ethnic group. To assess phenotypic impact, we compared LDL cholesterol levels (mmol/L) between carriers and non-carriers within each variant set and gene, using two-sided t-tests stratified by statin use.
PRS
Summary statistics for estimating PRS for the genomic and the epigenomic variation analysis were obtained from the PGS Catalog (Supplementary Data 6 and 22 respectively)51, selecting the study with best possible trans-ancestry base data and validation. PRS was estimated using the score function in PLINK230, in each cohort participant simultaneously, using the same set of PRS-specific genetic variants which were available in our QC-ed genomic dataset for all ancestries. We then applied Z-score standardization, to enable relationships to disease risk, and differences between ancestral groups to presented on a per standard deviation (SD) scale. This approach facilitates comparisons of effect size and ancestry-based risk scores between the PRS evaluated. The performance of PRS was tested separately for each ethnic group while adjusting for age and sex, and meta-analysed to determine trans-ancestry performance. The metrics for performance include Odd’s ratio, Observed R2 and R2 on the liability scale. For the PRS used in the methylation analysis, scores were estimated together for all the individuals with methylation data available.
DNA methylation
We first identified CpG sites that were considered significantly differentially methylated between any pair of Asian ethnic subgroups at a p-value threshold of 2.9 × 10-8. This cutoff was obtained via a two-step process. Firstly, we defined epigenome-wide significance as P < 8.62 × 10-8, which was obtained via permutation testing and is also close to what would have been obtained via Bonferroni correction. We then performed a second Bonferroni adjustment for the multiple testing between the three pairs of ethnic subgroups (Chinese versus Malay, Chinese versus Indian, and Malay versus Indian), which brings us to P < 2.9 × 10-8. To further assess the relationship between DNA methylation and metabolic outcomes, we focused on 315 sentinel CpG sites that are significantly associated with incident T2D based on our epigenome-wide association testing performed in age-, sex- and ethnicity-matched controls in the Translating Omics into A Stratified approach for prevention of T2D (TOAST) study. As one of the CpG sites was not found in HELIOS, this left us with 314 CpG sites for the analyses. DNA methylation was measured using baseline samples collected before onset of T2D, with primary analysis of epigenome-wide data performed as described previously in ref. 31. In brief, the association of each autosomal CpG site with incident T2D was tested using logistic regression, adjusted for confounders such as age, sex and further adjusted for imputed white blood cells (WBC) proportion and PC1-30 of control probe intensities. To assess the association of these CpG loci with BMI in the HELIOS participants, we then performed linear regression with the same covariate adjustments. Correlation between CpG sites were assessed using Pearson correlation analyses, with the circos plot generated by the circlize package.
Functional Annotation of Sentinel CpG
We perform functional overlap analysis and annotation of the sentinel CpGs using eFORGEv2.052 analyzing the 16,444 CpG sites for enrichment across DNase I hotspots, 5 histone marks and 15 chromatin states across 39 cell types from the Roadmap Epigenomic Consortium53. We determine the number of Sentinel CpGs overlapping with the annotated regulatory and chromatin regions in the different cell types. The enrichment of our sentinel CpG set was evaluated by comparing it to 1000 background sets that contain an equal number of sites as the input. The background sets were matched using gene annotation and CpG island annotation and the mean overlap for the background sets was calculated. We used the background sets to calculate the fold enrichment as observed count /mean (expected counts) and obtained an empirical P value from the distribution of the background sets.
Transcription Factor (TF) Enrichment
The binding site information for the 1210 human TFs tested was obtained from the Remap database, 2022 release (https://remap.univ-amu.fr/)54. We used the homo sapiens Cis Regulatory Modules (CRM) peaks for this analysis. We first determine how many of our sentinel CpGs overlap with the binding sites of the different TFs, and then estimated the fold enrichment; p-value for enrichment was calculated by comparing the overlap of our sentinels to the overlap of CpG probes from the background set of all CpGs. The p-value for enrichment was obtained using hypergeometric test and corrected for multiple testing at a False Discovery Rate threshold of 0.05 (Supplementary Data 9).
Enrichment across behavioural, lifestyle and genetically inferred traits
We tested the associations between the 16,444 ethnically differentiated CpGs, and 191 trait-exposures, including directly measured phenotypes as well as PRS to derive genetically inferred exposures (Supplementary Data 9). Linear or logistic regression was used, with adjustment for age, sex, ethnicity, methylation array control probe PCs, and white cell subset composition estimated by the Houseman method32. All traits were standardized using Z-score transformation before the analysis. We then performed the same analysis for all CpGs on the MethylEPIC array (837,722) to estimate background expectations. We then calculated enrichment (observed vs. background), using the hypergeometric test. We inferred statistical significance at P < 0.05/32, based on an estimate of 32 independent phenotypes derived from PCA of phenotypic covariation.
Epigenetic PCA analysis
To understand the genetic and environmental factors influencing genome regulation in our population, we also examined the relationship of 191 traits (Supplementary Data 11) with the principal components of variation in methylation at the 16,444 CpG sites that are differentiated between our Asian ethnic groups. We used PCA as a data reduction strategy to identify the primary axis of variation in the methylation at these CpG sites. We then tested the associations with the 5 PCs with potential exposure, adjusted for age, sex, ethnicity, methylation array control probe PCs, and white cell subset composition estimated by the Houseman method32. We again inferred statistical significance at P < 0.05/32, based on an estimate of 32 independent phenotypes derived from PCA of phenotypic covariation.
Metabolic variation
To explore the variation in metabolite levels across ethnicities, we randomly split the dataset into discovery (70%) and test (30%) cohorts. Using linear regression analysis, we estimated the association between variation in levels of 1073 metabolites and self-reported ethnicities (Malay compared to Chinese, Indian compared to Chinese, and Malay compared to Indian) in the discovery cohort, adjusted for age, sex, and shipment batch. We applied a Bonferroni-corrected p-value threshold of 1 × 10-5 to account for multiple testing (1073 metabolites x 3 pair-wise tests). We then repeated the same set of analyses for these 162 metabolites in the replication cohort, and a subset of 153 metabolites that met the following criteria: 1) significantly associated with ethnicity in the discovery cohort at P < 1 × 10-5, and 2) significantly associated with ethnicity in the test cohort at P < 0.05 and with the same direction of estimates. In an age and sex-matched cohort of 1,146 participants per ethnicity, we performed PCA of the 153 metabolites to assess the extent of clustering of individuals by ethnicity. Out of these 153 metabolites, 128 were well-characterized and known metabolites. We evaluated associations between these 128 metabolites and four common health outcomes: hypertension, obesity, T2D, and CVD, using logistic regressions adjusted for age, sex, and shipment batch. For each phenotype, we applied a Bonferroni-corrected p-value threshold of 4.7 × 10-5 to account for multiple testing (1073 metabolites). We also evaluated associations between a metabolite of interest (1-margaroyl-2-arachidonoyl-GPC) and FFQ foods, adjusted for age, sex, ethnicity and shipment batch, and reported Bonferroni-Hochberg corrected p-values for the top four foods. Furthermore, for each of these metabolites, we calculated partial R-squared values to estimate the contribution of genetic ancestry and various demographic and lifestyle factors on metabolic variation. Genetic ancestry was represented using the first 50 genetic PCs, and dietary habits using the first 10 PCs representing 169 food items and major macronutrients. Finally, pairwise correlation between metabolites was estimated using Pearson correlation and a significance p-value threshold of 1 × 10-6 was applied to account for multiple testing. Metabolites were grouped into 10 categories (super-pathways) and annotated to pathways or chemical classes based within each category (sub-pathways). The circos plot was generated using the circlize package.
Gene expression variation
We estimated the association between variation in levels of 12,434 genes (expression measured in standardized TPM) and self-reported ethnicities (Malay compared to Chinese, Indian compared to Chinese, and Malay compared to Indian) in the discovery cohort, adjusted for age, sex, RNA Integrity Number (RIN) and the top six Probabilistic Estimation of Expression Residuals (PEER) factors55. We applied a Bonferroni-corrected p-value threshold of P < 1.34 × 10-6 to account for multiple testing (12,434 transcripts x 3 pair-wise tests).
Gene set enrichment analysis
We performed GSEA on the 2972 genes that were differentially expressed in any one pairwise ancestry comparison. GSEA was performed using the gprofiler2 tool56 to determine enriched gene sets across 3 pathway databases. These include Kyoto Encyclopedia of Genes and Genomes (KEGG), Reactome (REAC) and Wiki Pathways (WP). Fold enrichment was determined using the ratio of sentinel gene overlap compared to total term size, and the p-value for enrichment was determined using hypergeometric test for over representation. Significantly enriched gene sets were determined after FDR testing for multiple testing correction (FDR P < 0.05).
Cis-eQTLs for gene expression
cis-Expression quantitative trait loci (eQTLs) for the 2972 differentially expressed genes were analysed using Matrix eQTL (R package MatrixEQTL56, with gene expression modelled as a regression model of genotypes and covariates, including age, sex, ethnicity, RIN (RNA integrity number) and the top 6 PEER (Probabilistic Estimation of Expression Residuals) factors57. For the identification of significant cis-eQTLs, a Bonferroni-corrected p-value threshold (P < 0.05 /16220560 tests) was applied.
Genome Wide Association Studies (GWAS)
To identify genetic variants associated with metabolite levels in the HELIOS dataset, we first divide the cohort to select only individuals having metabolite data and were clustered in our three main ancestry groups (Chinese, N = 5961; Indian: N = 1470; Malay: N = 838) that were determined by our data driven approach. The individuals in the three admixed group (n = 409) were not included in the analysis. We then perform GWAS QC and analysis for each group separately, followed by inverse variance meta-analysis to create summary statistics across the study population. GWAS variant QC filters were: MAF < 0.5%, HWE p-value < 1 × 10-6, Missingness <2%. Sample filters were pi_hat <0.75, IBC < | 0.2| and Sex-mismatch. We used PLINK230 to get the final set of samples and variants to be used for the analysis. Overall, 5940 Chinese, 1461 Indians and 833 Malays with 12.7, 16, and 14.7 million variants respectively were included in the analysis. For the GWAs of metabolites, we log transformed the metabolite data and removed individuals with the highest deviation (>5 SD from the mean). Age, sex, top 20 genetic PCs, and batch were used as covariates in the analysis. The individual GWAS for each ancestry was performed using REGENIE55. The subset of SNPs for REGENIE step 1 were chosen after filtering for MAF < 5%, HWE P < 1 × 10-6 in the ethnic group being analysed. We removed the MHC and chr8 inversion regions, followed by LD pruning at an r2 of 0.05 within a 200 kb window. Meta-analysis of the three summary statistics was performed using METAL with a fixed effect model controlling for genomic inflation across each dataset. Variants were filtered for being in at least two datasets, heterozygosity P > 0.05 and max difference between allele frequencies < 0.5.
SMR and colocalization
SMR21 was performed to identify pleiotropic association between gene expression (exposure) (from the eqtlgen dataset21) and metabolite levels (outcome) using GWAS summary statistics. To limit the number of tests, we include SNPs that pass genome-wide significance in our GWAS as well as in the cis-eQTL dataset. Analysis was performed using the SMR tool58. For the metabolite-gene pairs with significant SMR association, we performed colocalization analysis using the coloc package implemented in R59. The region of 1MB on each side of the SMR associated SNP was used for colocalization analysis under a single causal variant assumption and the default prior probabilities. Metabolite- Gene pairs with a coloc H4 posterior probability > 0.7 were considered to be colocalized and share a common causal variant.
Reproducibility assessment of measurements
The reproducibility of 107 measurements in 9 domains (Supplementary Data 18) between baseline test and the repeated study was assessed using correlation coefficients calculated from Spearman correlation analysis for z-scored transformed measurements.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The individual level data for study participants are available under restricted access due to data privacy and ethical considerations. Access request can be submitted to the HELIOS Data Access Committee (email: helios_science@ntu.edu.sg). The GWAS summary statistics for the metabolites generated in the present study have been deposited in the GWAS catalogue under accession codes: GCST90557814-GCST90557966 (https://www.ebi.ac.uk/gwas/). The cis-eQTL summary statistics for the significant genes generated in the HELIOS study have been deposited in our GitHub repository60 and are publicly available. Source data are provided with this paper.
Code availability
The analytic codes developed in this study have been deposited in our GitHub repository60 and are publicly available.
References
World Health Organization. Noncommunicable diseases fact sheets. https://www.who.int/news-room/fact-sheets/detail/noncommunicable-diseases (accessed 18 June 2023).
World Health Organization. Noncommunicable diseases in the South East Asia. World Health Organization. https://www.who.int/southeastasia/health-topics/noncommunicable-diseases (accessed 18 June 2023).
International Diabetes Federation. IDF Diabetes Atlas, 10th Edition. https://diabetesatlas.org/resources/previous-editions/ (accessed 22 June 2023).
Zhao, D. Epidemiological features of cardiovascular disease in Asia. JACC: Asia 1, 1–13 (2021).
Dans, A. et al. The rise of chronic non-communicable diseases in southeast Asia: time for action. Lancet 377, 680–689 (2011).
Popejoy, A. B. & Fullerton, S. M. Genomics is failing on diversity. Nature 538, 161–164 (2016).
Fatumo, S. et al. A roadmap to increase diversity in genomic studies. Nat. Med. 28, 243–250 (2022).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Wong, E. et al. The Singapore National Precision Medicine Strategy. Nat. Genet. 55, 178–186 (2023).
World Population Review. Asia population. World Population Review. https://worldpopulationreview.com/continents/asia-population (accessed 25 June 2023).
Mina, T. et al. Adiposity and metabolic health in Asian populations: an epidemiological study using dual-energy x-ray absorptiometry in Singapore. Lancet Diabetes Endocrinol. 12, 704–715 (2024).
Sadhu, N. et al. Metabolome-wide association identifies ferredoxin-1 (FDX1) as a determinant of cholesterol metabolism and cardiovascular risk in Asian populations. Nat. Cardiovasc. Res. 4, 567–583 (2025).
Low, D. Y. et al. Metabolic variation reflects dietary intake in a multi-ethnic Asian population. Nat. Metab. 7, 1939–1954 (2025).
Mina, T. et al. Adiposity impacts cognitive function in Asian populations: an epidemiological and Mendelian Randomization study. Lancet Reg. Health West. Pac. 33, 100710 (2023).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Stuppia, L. Chapter 18 - Transgenerational effects of imprinting. in Transgenerational Epigenetics (Second Edition) (ed. Tollefsbol, T. O.) vol. 13 389–400 (Academic Press, 2019).
Aldana-Hernández, P. et al. Dietary phosphatidylcholine supplementation reduces atherosclerosis in Ldlr − /− male mice2. J. Nutr. Biochem. 92, 108617 (2021).
Calder, P. C. Dietary arachidonic acid: harmful, harmless or helpful?. Br. J. Nutr. 98, 451–453 (2007).
Van Parys, A. et al. Food Sources Contributing to Intake of Choline and Individual Choline Forms in a Norwegian Cohort of Patients With Stable Angina Pectoris. Front. Nutr. 8, 676026 (2021).
Koletzko, B. et al. FADS1 and FADS2 Polymorphisms Modulate Fatty Acid Metabolism and Dietary Impact on Health. Annu. Rev. Nutr. 39, 21–44 (2019).
Võsa, U. et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
Jain, P. R. et al. Nuclear regulatory disturbances precede and predict the development of Type-2 diabetes in Asian populations. medRxiv 2025.02.14.25322264 https://doi.org/10.1101/2025.02.14.25322264 (2025).
Chambers, J. C. et al. Epigenome-wide association of DNA methylation markers in peripheral blood from Indian Asians and Europeans with incident type 2 diabetes: A nested case-control study. Lancet Diab. Endocrinol. 3, 526–534 (2015).
Meyers, E. A. & Kessler, J. A. TGF-β Family Signaling in Neural and Neuronal Differentiation, Development, and Function. Cold Spring Harb. Perspect. Biol. 9, a022244 (2017).
Rutter, C. E., Millard, L. A. C., Borges, M. C. & Lawlor, D. A. Exploring regression dilution bias using repeat measurements of 2858 variables in ≤49 000 UK Biobank participants. Int. J. Epidemiol. 52, 1545–1556 (2023).
Chen, Z. et al. China Kadoorie Biobank of 0.5 million people: Survey methods, baseline characteristics and long-term follow-up. Int. J. Epidemiol. 40, 1652–1666 (2011).
Declercq, J. et al. Increased β-Cell Mass by Islet Transplantation and PLAG1 Overexpression Causes Hyperinsulinemic Normoglycemia and Hepatic Insulin Resistance in Mice. Diabetes 59, 1957–1965 (2010).
Juma, A. R., Damdimopoulou, P. E., Grommen, S. V. H., Van de Ven, W. J. M. & De Groef, B. Emerging role of PLAG1 as a regulator of growth and reproduction. J. Endocrinol. 228, R45–R56 (2016).
Hail Team. Hail 0.2.13-81ab564db2b4. https://github.com/hail-is/hail/releases/tag/0.2.13 (accessed 25 June 2023).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, s13742-015-0047–8 (2015).
Lehne, B. et al. A coherent approach for analysis of the Illumina HumanMethylation450 BeadChip improves data quality and performance in epigenome-wide association studies. Genome Biol. 16, 1–12 (2015).
Houseman, E. A. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinforma. 13, 86 (2012).
Krueger, F. TrimGalore. https://github.com/FelixKrueger/TrimGalore (accessed 18 June 2024).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 323 (2011).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Unger, T. et al. 2020 International Society of Hypertension Global Hypertension Practice Guidelines. Hypertension 75, 1334–1357 (2020).
Su, D. et al. Ethnicity-specific blood pressure thresholds based on cardiovascular and renal complications: a prospective study in the UK Biobank. BMC Med. 22, 54 (2024).
WHO Expert Consultation. Appropriate body-mass index for Asian populations and its implications for policy and intervention strategies. The Lancet 363, 157–163 (2004).
Caleyachetty, R. et al. Ethnicity-specific BMI cutoffs for obesity based on type 2 diabetes risk in England: a population-based cohort study. Lancet Diab. Endocrinol. 9, 419–426 (2021).
Fung, T. T. et al. Adherence to a DASH-style diet and risk of coronary heart disease and stroke in women. Arch. Intern. Med. 168, 713–720 (2008).
Whitton, C. et al. Relative validity and reproducibility of a food frequency questionnaire for assessing dietary intakes in a multi-ethnic Asian population using 24-h dietary recalls and biomarkers. Nutrients 9, 1059 (2017).
Craig, C. L. et al. International physical activity questionnaire: 12-Country reliability and validity. Med. Sci. Sports Exerc. 35, 1381–1395 (2003).
World Health Organization. Physical activity fact sheets. https://www.who.int/news-room/fact-sheets/detail/physical-activity (accessed 12 July 2023).
White, T. et al. Estimating energy expenditure from wrist and thigh accelerometry in free-living adults: a doubly labelled water study. Int. J. Obes. 43, 2333–2342 (2019).
White, T., Westgate, K., Wareham, N. J. & Brage, S. Estimation of physical activity energy expenditure during free-living from wrist accelerometry in UK adults. PLoS One 11, e0167472 (2016).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Chiu, A. M., Molloy, E. K., Tan, Z., Talwalkar, A. & Sankararaman, S. Inferring population structure in biobank-scale genomic data. Am. J. Hum. Genet. 109, 727–737 (2022).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Lambert, S. A. et al. The Polygenic Score Catalog as an open database for reproducibility and systematic evaluation. Nat. Genet. 53, 420–425 (2021).
Breeze, C. E. et al. eFORGE v2.0: updated analysis of cell type-specific signal in epigenomic data. Bioinformatics 35, 4767–4769 (2019).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Hammal, F., de Langen, P., Bergon, A., Lopez, F. & Ballester, B. ReMap 2022: a database of Human, Mouse, Drosophila and Arabidopsis regulatory regions from an integrative analysis of DNA-binding sequencing experiments. Nucleic Acids Res. 50, D316–D325 (2022).
Mbatchou, J. et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat. Genet. 53, 1097–1103 (2021).
Kolberg, L., Raudvere, U., Kuzmin, I., Vilo, J. & Peterson, H. gprofiler2 - an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Res 9, 709 (2020).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Wallace, C. A more accurate method for colocalisation analysis allowing for multiple causal variants. PLoS Genet. 17, e1009440 (2021).
Mina, T. et al. HELIOS-SG100K-LKC/HELIOS-Study-design-and-baseline-characteristics: The Health for Life in Singapore (HELIOS) Study: delivering Precision Medicine research for Asian populations (v1.0.0). Zenodo (2025). https://doi.org/10.5281/zenodo.16875846.
Acknowledgements
This study is supported by Singapore Ministry of Health’s (MOH) National Medical Research Council (NMRC) under its OF-LCG funding scheme (MOH-000271-00), Singapore Translational Research (StaR) funding scheme (NMRC/StaR/0028/2017), the National Research Foundation, Singapore through the Singapore MOH NMRC and the Precision Health Research, Singapore (PRECISE) under the National Precision Medicine programme (NMRC/PRECISE/2020) and intramural funding from Nanyang Technological University, Lee Kong Chian School of Medicine and the National Healthcare Group. This research was also supported in part through the National Research Foundation, Prime Minister’s Office, Singapore under its Campus for Research Excellence and Technological Enterprise (CREATE) programme. RNA sequencing was partially funded by i) Ministry of Education Academic Research Fund Tier 1 Grant (RS09/20), ii) A*STAR-NHMRC Joint Grant Call (A20PRb0138), iii) Imperial - Nanyang Technological University Collaboration Fund (awarded to M.Loh [PI]). T.M. was funded by Dean’s Postdoctoral Fellowship from the Lee Kong Chian School of Medicine. This study made use of data generated by Ministry of Health (MOH) and Immigration and Checkpoints Authority (ICA). This study was supported by the Trusted Research and Real-World-Data Utilisation and Sharing Tech platform (“TRUST Platform”) developed by the Ministry of Health and Smart Nation and Digital Government Office, through the use of its research data analytics facilities. The views expressed are those of the author(s) are not necessarily those of the Government, MOH and ICA investigators or institutional partners. The computational work for this study was partially performed on resources of the National Supercomputing Centre, Singapore (https://www.nscc.sg). We thank all participants and research staff who made the study possible. We thank Dr. Tomas Gonzalez, Lewis Griffiths, Stefanie Hollidge, and Antony Siahaan (MRC Epidemiology Unit, University of Cambridge), Dr. Ian Goon and Samiul Hoque for assistance in accelerometry data extraction and analysis, Dr. Shaikh Fairul Edros Shaikh Ahmad (Earth Observatory Singapore, NTU) for guidance in geospatial data analysis, and Dr. Clare Whitton for the collaborative development of e-FFQ and the data collection platform.
Author information
Authors and Affiliations
Consortia
Contributions
J.C.C., P.E., E.R., J.N., L.E.S., J.L., T.M., K.T., L.Lee., L.T.H., and J.B., conceived and designed the HELIOS study. T.M., T.T.Y.Y., C.W.L., L.Lim., K.S.K., L.G.L., B.L.C.C., R.D., G.W., Y.Y.W., and R.M.D. implemented the study and collected data. T.M., N.S, D.L.Y.W., P.R.J, D.T., G.A.M, K.E.W, P.A.S, L.P.Y., Y.Z.X., N.B., C.B., M.H., P.G., E.J.L, S.B and H.S. curated epidemiological and molecular data. X.W., T.M., N.S., P.R.J., H.K.N., D.L.Y.W., D.T., R.D., M.Lam, and M.Loh performed the data analyses. J.C.C. supervised the study implementation, data curation and analyses. X.W., T.M., N.S., P.R.J., H.K.N., D.L.Y.W., M.Lam, M.Loh, P.T., G.X.L., and J.C.C. wrote the manuscript. The HELIOS study team was involved in sample collection and processing and data analysis. All authors reviewed and contributed to the revision of the submitted manuscript.
Corresponding author
Ethics declarations
Competing interests
B.L.C.C. receives honorarium for obesity-related presentations and/or participates in the advisory board of Novo Nordisk, Abbott Nutrition and DKSH, and all honorariums were paid to Khoo Teck Puat Hospital, Singapore. J.N. receives research funding from Astra Zeneca. J.L. participates in the advisory board of Boehringer Ingelheim and is a council member of National Council Against Drug Abuse, Singapore. G.A.M., K.E.W., and P.A.S. are employees of Metabolon. L.P.Y., and Y.Z.X. are employees of Ministry of Health, Singapore. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jeroen van Rooij and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, X., Mina, T., Sadhu, N. et al. The Health for Life in Singapore (HELIOS) Study: delivering precision medicine research for Asian populations. Nat Commun 17, 1 (2026). https://doi.org/10.1038/s41467-025-65774-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65774-0
This article is cited by
-
Adversity, adiposity, nutrition and metabolic well-being in multi-ethnic Asia
Nature Metabolism (2026)