Abstract
Comprehensive in situ structures of macromolecules can transform our understanding of biology and advance human health. Here, we map protein synthesis inside human cells in detail by combining automated cryo-focused ion beam (FIB) milling and in situ single-particle cryo electron microscopy (cryo-EM). With this in situ cryo-EM approach, we resolved a 2.2 Å consensus structure of the human 80S ribosome and unveiled 23 functional states, nearly all better than 3 Å resolution. Compared to in vitro studies, we observed variations in ribosome structures, distinct environments of ion and polyamine binding, and associated proteins such as EDF1 and NACβ that are typically not enriched with purified ribosomes. We also detected additional peptide-related density features on the ribosome and visualized ribosome–ribosome interactions in helical polysomes. Finally, high-resolution structures from cells treated with homoharringtonine and cycloheximide revealed a distinct translational landscape and a spermidine that interacts with cycloheximide at the E site, one of the numerous polyamines that also bind native ribosomes. These results underscore the value of high-resolution in situ studies in the native environment.
Similar content being viewed by others
Introduction
Protein synthesis is a universally conserved and critical process in which ribosomes translate genetic information into proteins. The eukaryotic 80S ribosome is comprised of the small and large ribosomal subunits (40S and 60S subunits, respectively), and translates mRNA into protein through a sequence of initiation, elongation, termination, and recycling steps1,2,3,4. During elongation, synthesis of the nascent peptide requires intricate concerted processes, including small subunit rotation, cofactor coordination, tRNA translocation, and movement of mobile structural elements2,4. Extensive in vitro studies of purified ribosomes using X-ray crystallography and cryo-EM have provided much of our current understanding of translation1,4. Affinity purification and in vitro assembly have been used to study specific ribosomal complexes5,6,7,8,9,10,11. Many dynamic states have been captured in vitro using ribosomal inhibitors, providing a detailed view of the translation elongation cycle11,12,13,14,15. Despite extensive studies, it remains unclear to what extent the purified ribosomes differ from those in native cell environments.
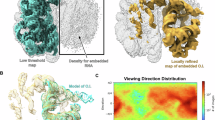
Structure determination of biological macromolecules at high resolution within their native cellular environment is an enticing pursuit. Advances in cryogenic electron tomography (cryo-ET) have enabled steady progress toward this goal16,17,18. Recent applications of cryo-ET to thin cellular samples, such as small bacteria or cellular lamellae milled by a focused ion beam (FIB)19,20,21,22, have revealed remarkable aspects of ribosome dynamics inside cells, including drug interactions and their impact on translation12,23,24,25. To complement the time-intensive cryo-ET method, an in situ single-particle cryo-EM approach (Fig. 1a) has emerged as a promising technique to interrogate large cellular complexes inside the cell with higher throughput and potentially greater resolution26,27,28. However, the full potential of in situ single-particle cryo-EM for biological discovery—such as identifying transient or weakly bound factors, low-abundance conformational states, or native distributions of ions and post-translational modifications within cells—has yet to be fully demonstrated.
a Schematic of the in situ single-particle cryo-EM pipeline. b Consensus map of human 80S ribosome at 2.2 Å resolution (left) and the local resolution map (right). c Selected features of the 80S ribosome structure. The cryo-EM map is shown as a semitransparent surface with the fitted model in stick representation. rRNA modifications are marked with arrows. d Well-resolved cryo-EM map of the P site tRNA (P-tRNA). e Cryo-EM density of mRNA (a polyU model, blue sticks) in A, P, and E sites of the 40S subunit (bottom) and its interaction with the P-tRNA (right inset). The well-resolved density for mRNA is an average of all translating mRNAs. A schematic of the interactions is shown at the top. f Density for the nascent peptide (red) in the 60S subunit. The ribosome is shown as a gray surface clipped at the nascent peptide exit tunnel. g Cryo-EM densities of homoharringtonine (HHT) and cycloheximide (CHX) (shown as sticks) resolved in cells treated with both drugs.
In this study, using in situ single-particle cryo-EM, we resolve the consensus structure of human 80S ribosomes inside HEK293A cells at 2.2 Å resolution. We uncover 23 distinct 80S ribosome conformations at high resolutions, revealing previously unobserved structural features of ribosomal components, protein factors, and small-molecule ligands. Our results provide a detailed view of the translation cycle and the interfaces between ribosomes in polysomes inside cells. Using this pipeline, we also resolve translational dynamics in HEK293A cells treated with homoharringtonine and cycloheximide, revealing differences in the translation landscape due to drug presence. Our results underscore the potential of the in situ single-particle cryo-EM approach for high-resolution structural studies of biological complexes in their native environment.
Results
High-resolution in situ human 80S ribosome structures
To investigate the structure of the human ribosome inside cells, we used cryo-FIB milling to section 193 lamellae from human cells (HEK293A), from which we collected 16,636 micrographs using a single-particle cryo-EM data collection procedure (Supplementary Fig. 1a, b). Particle picking and initial orientation determination were first performed by template matching with a 60S ribosomal subunit reference map29, followed by further search with the refined 80S ribosome map, using the programs GisSPA23,30 and cisTEM31 (Supplementary Fig. 3). 3D classification and refinement using the single-particle pipeline in CryoSPARC32 yielded a consensus cryo-EM map of the 80S ribosome at 2.2 Å resolution, with the highest local resolution better than 2 Å (Fig. 1b, Supplementary Fig. 1c and Supplementary Table 1). The density for ribosomal proteins and ribosomal RNA (rRNA) was resolved at a level that has not been previously achieved in situ, including densities for numerous rRNA modifications consistent with those previously observed for purified ribosomes (Fig. 1c)29,33,34,35. Focused classification on the tRNA, mRNA, and nascent peptide allowed us to resolve densities for these features in 80S ribosomes (Fig. 1d–f). The path of the mRNA through the A (aminoacyl), P (peptidyl), and E (exit) tRNA-binding sites is clearly visible, including well-resolved densities for nucleotide bases that are averaged from all translating mRNAs in the cell (Fig. 1e). Certain mRNA regions, especially those not base-paired with tRNAs, exhibit reduced resolution due to increased conformational flexibility. We also applied this pipeline to cells treated with homoharringtonine (HHT) and cycloheximide (CHX) and obtained a consensus structure (2.4 Å resolution) of the 80S ribosome with the two drugs bound. The quality of the map allowed us to unambiguously model HHT and CHX (Fig. 1g).
Compared to in vitro studies, we observed conformational differences for regions of the ribosome and its bound ligands (Fig. 2 and Supplementary Figs. 1d, 2). Intriguingly, in all 23 states, the 5.8S rRNA nucleotides C109, U110, and U111 were observed with a loop conformation drastically different from that observed for purified ribosomes (Fig. 2a). This loop is buried inside the 60S subunit, and the different conformation may affect the stability of the ribosome. In another interesting case, nucleotide G3922 of the 28S rRNA located in the tRNA-binding groove in between the P and E sites was observed to adopt two conformations depending on the translation state (Figs. 2a, 3d, e, and Supplementary Fig. 6b), suggesting functional consequences of the conformations. By contrast, only conformation 1 has been reported in in vitro studies.
a Example of nucleotide conformation differences of 5.8S rRNA nucleotides C109U110U111 (top) and 28S rRNA nucleotides G3922 (bottom). A sliced view of the 80S ribosome showing the buried positions of these nucleotides is shown in the side panels. b Examples of polyamine densities. The locations of all polyamine molecules identified are shown in the ribosome as schematics. Selected polyamine densities are shown with stick models in the insets. Densities of all identified polyamines are shown in Supplementary Fig. 2a. c, d Examples of amino acid side chain differences caused by different ion or ligand occupations. e Examples of observed differences in ion occupation. Densities of all identified ion differences are shown in Supplementary Fig. 2b. For all figures, cryo-EM maps are shown in semitransparent surfaces, and atomic models are shown in sticks.
a 14 classes of human 80S ribosome in the translation elongation cycle, clipped to show tRNAs and factors bound. Solid arrows indicate pathways that have previously been reported. Dashed arrows are proposed pathways. The percentages indicate the proportion of particles in each class. The 60S and 40S subunits are shown in light and dark gray, respectively. The tRNAs are colored by position. b Comparison of the positions of tRNAs and translation elongation factors in each class, colored as in 2a. c 9 classes of hibernating states, with the E-tRNA bound rotated hibernating states (7a-11a) grouped together. d Translation state-dependent conformations of 28S rRNA G3922 (sticks). e Relative occupancy of the two conformations in different states.
The structure and function of the ribosome strongly depend on the proper concentration of Mg2+, K+, Zn2+, NH4+and polyamine molecules36,37,38,39. Ion concentrations either too low or too high inhibit translation40,41,42,43,44. The binding of cations and polyamines has been extensively studied using purified ribosomes at high resolution35, although it is not clear how these ions associate with ribosomes inside the cell. Through our in situ ribosome structure, we identified 21 polyamine molecules, including spermine, spermidine, and putrescine, bound to the ribosome, often coordinating with rRNA backbone phosphates (Fig. 2b, Supplementary Fig. 2a, and Supplementary Table 7). Some of these interactions have been observed previously in vitro structures of purified ribosomes, but the presence of these molecules is inconsistent across the literature, possibly because the ligands and ions observed depend on the buffer system being used29,45,46. We have compared all Mg2+, K+, and Zn2+ binding sites previously identified in vitro35, and found numerous differences (Fig. 2e and Supplementary Fig. 2b). Importantly, in several cases, the different polyamines and ion occupation result in changes to the conformation of ribosomal protein and rRNA (Fig. 2c, d). These small-molecule ligands may play important roles in fine-tuning translation elongation.
Translation dynamics in human cells
In situ cryo-EM enabled us to capture a series of translationally active and inactive states of human ribosomes, from selection of the cognate aminoacyl-tRNAs to addition of the amino acids onto the growing nascent peptide chain. The distribution of states observed here differs from previous cryo-ET studies12,23,24, potentially due to variations in cell conditions. From nearly 2 million high-quality particles, we identified 23 distinct ribosome states, almost all resolved at better than 3 Å resolution (Fig. 3a, b, Supplementary Figs. 3 and 4a and Supplementary Tables 2–4). Of these, 14 states represent actively translating ribosomes, as indicated by clear density for the mRNA, tRNAs, and protein factors, and are consistent with those described in the literature (Fig. 3a and Supplementary Fig. 5a)12,22,23,24,47. The most prevalent states are the eEF1A-A/T-P-E, eEF1A-A/T-P-Z, and eEF1A-A/T-P states (states 2a, 2b, 2c, respectively; Fig. 2a), which together comprise 30.3% of all 80S ribosomes. The mRNA at the decoding site is disordered in these states, suggesting that these are codon sampling states where the mRNA has not yet formed stable hydrogen bonds with the cognate tRNA (Supplementary Fig. 5a). This might also reflect different mRNA conformations depending on the mRNA sequence. The higher overall occupancy of these eEF1A-A/T states is consistent with the notion that codon recognition is a rate-limiting step in translation elongation45. We also resolved the high-resolution structures of the ribosome with eEF1A in a post GTP hydrolysis state (states 2a’, 2b’, 2c’), where the mRNA at the decoding site is structured and interacts with the A/T-tRNA (Fig. 2a, Supplementary Fig. 5a). This is accompanied by a large rotation of the A/T-tRNA, which interacts with helix H43 and H44 before rotation and with helix H89 of the 28S rRNA after rotation (Supplementary Fig. 6a).
The detailed structures allowed us to observe a translation state-dependent difference in rRNA conformation (Fig. 3d, e). In states lacking both E- (or P/E-) and Z-tRNAs, 28S rRNA residue G3922 tends to adopt a conformation that stacks onto the G4541U4542G4543A4544 tetraloop close to the P site (From 1), consistent with previous in vitro studies (Fig. 3d, e and Supplementary Fig. 6b). By contrast, in states with either E-, P/E-,or Z-tRNA, G3922 tends to flip toward the E site to interact with the phosphate group of nucleotide G3705 (Form 2) (Fig. 2a, Fig. 3d, e, Supplementary Fig. 6b). Although G3922 does not directly contact any of the tRNAs, its location in the tRNA movement trajectory suggests possible roles in modulating or sensing the tRNA movement (Fig. 3d). The translation state-dependent conformations identified here have not been observed in previous in vitro studies or lower-resolution cryo-ET analyses.
Our data also provides insights into the functional roles of the recently discovered noncanonical Z site in the ribosome12,22,48. We resolved five distinct Z-tRNA-bound states, 1b (P-Z), 2b (eEF1A-A/T-P-Z), 2b’ (Post-eEF1A-A/T-P-Z), 3b (A-P-Z), and 7b (Hiber-rotate3-Z) (Fig. 3a, b), four of which contain mRNA density and thus might represent translating ribosomes. In agreement with a previous report48, we observed that the Z-tRNA interacts with both the 60S and 40S ribosomal subunits, and its position is incompatible with the presence of a tRNA in the E site (Supplementary Fig. 6c, d). The acceptor arm of the Z-tRNA interacts with eL42 and H68 of the 28S rRNA, while the anticodon loop is held by eS25 (Supplementary Fig. 6c). Three of the four Z-tRNA states (1b, 2b, and 2b’) have similar conformations, but the anticodon loop of the tRNA in the A-P-Z state (3b) and eS25 shift towards the solvent together by ~3.8 Å. The T-loop/elbow of the Z-tRNA is bound to the backbone of the L1 stalk, similar to how the E-tRNA contacts the L1 stalk, and the stalk moves with the tRNA toward the Z site (Supplementary Fig. 6d). Therefore, it is possible that the L1 stalk facilitates the displacement of tRNA from the E site through a rigid-body motion, guiding it toward the solvent via the Z site. Future studies will be needed to interrogate this possibility.
Additionally, we observed 9 non-translating ribosome states with bound eEF2 (Fig. 3c). Among these, two states have an unrotated 40S subunit (states 6a, 6b; Fig. 3c), with the translation reactivation factor CCDC124 bound in one of the states (6b). The remaining seven states are associated with SERPINE1 mRNA binding protein 1 (SERBP1) and exhibit varying degrees of 40S subunit body rotation and head swivel, shedding light on the action of SERBP1 (states 7-11; Fig. 3c and Fig. S7d, f). The hibernating state with Z-tRNA bound or empty E site has a small amount of 40S body rotation (states 7b; Fig. 3c; Supplementary Fig. 3), while most E-tRNA-bound states have larger 40S body rotation (states 7a-11a; Fig. 3c and Supplementary Fig. 3), suggesting the possibility that the rotation of the 40S body impacts binding affinity at the Z and E sites.
Ribosome-binding cofactors identified in situ
Our cryo-EM maps allowed us to identify many factors that bind transiently to ribosomes in situ, including endothelial differentiation-related factor 1 (EDF1), nascent polypeptide-associated complex β (NACβ), and eukaryotic initiation factor 6 (eIF6) (Fig. 4). EDF1 is involved in resolving ribosomal collisions9. In our cryo-EM maps, EDF1 is distinctly visible on approximately half of the ribosomes that are associated with a trailing ribosome (Fig. 5a,b). Surprisingly, in our EDF1-bound ribosome structures, helix 16 (h16) of the 18S rRNA displays a different conformation, marked by a repositioning of the solvent-exposed hairpin region of h16 and a substantial widening of the minor groove, compared to the EDF1-bound structure from purified samples9 (Fig. 4c). It appears to be “clamped” by the N- and C-terminal regions of EDF1 in a way that has not be observed before (Fig. 4c). These changes may be functionally important and again highlight the importance of conducting structural biology in situ.
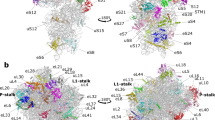
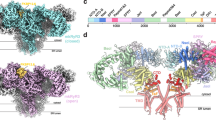
a Ribosome binding factors EDF1 (endothelial differentiation-related factor 1), EBP1 (ErbB3-binding protein 1), and NACβ (nascent polypeptide-associated complex β) were identified on ribosomes inside the cell. Representative cryo-EM maps (surface) and structural models (cartoon and sticks) are shown. Two patches of unknown density with fitted molecular model are shown in cyan (bottom). b eIF6 (eukaryotic initiation factor 6) (pink) and EBP1 densities in free 60S particles in the untreated cells. c Conformations of h16 of 18S rRNA and EDF1 observed in situ and in vitro.
a Structures of two types of di-ribosomes. i indicates the leading ribosome, i + 1 indicates the trailing ribosome. Class1 is a top-back (t-b) interaction and Class2 is a top-top (t-t) interaction. b Interaction interfaces of the two types of di-ribosomes. The t-b class has mainly inter-ribosomal 60S-40S subunit interactions, whereas the t-t class has mainly 40S-40S subunit interactions. c Comparison of the two classes of di-ribosomes. d Helical Class1 ribosome density appears when data processing was done with a large box. e The t-t interaction is incompatible with the helical structure shown in d, whereas the t-b interaction can form this high-order helical ribosome structure.
NACβ is involved in nascent peptide processing and trafficking. While it was previously primarily studied in in vitro reconstituted samples, recent structural evidence demonstrates its prevalent presence on ribosomes in cells10,28. Our maps show that the N-terminal anchor domain of NACβ is enriched on all translating ribosome states with occupancies of ~50% (Fig. 4a and Supplementary Fig. 8a). The C-terminal globular domain of NACβ, which we did not observe, has been reported to compete with the binding of EBP1, a protein associated with ribosomal translational regulation at the nascent peptide exit channel49. We observed that EBP1 and NACβ N-terminal occupations exhibit modest inverse correlation (Supplementary Fig. 8a). In hibernating 80S ribosomes and free 60S subunits, the occupancy of EBP1 increases, while the occupancy of NACβ drops. This correlation may be important for the nascent peptide exit and translation regulation. Furthermore, while a high-resolution cryo-ET study of human untreated ribosomes did not detect eIF6 bound to free 60S subunits12, our results and an independent cryo-ET work22 provide complementary evidence supporting eIF6-60S association in mammalian cells. Our 60S subunit structure revealed clear density for eIF6 bound at the inter-subunit interface (~60% occupancy), poised to prevent 40S ribosomal subunit joining (Fig. 4c).
Intriguingly, we observed two patches of previously unreported densities on the ribosome (Fig. 4a). The first patch of density is bound to RACK1 on the head of the 40S subunit, and the second patch of density is bound to ribosomal protein L30, L34, and L27 on the 60S subunit near the inter-subunit interface. We manually built the peptide backbone and placed potential sidechains that fit the density. While we cannot ascertain the identity of these unknown densities, our manually built model aligns closely with the AlphaFold2 prediction of various segments of SERBP1 bound to ribosome (Supplementary Fig. 8 b, c)50. Given that these densities appear on all ribosomal classes resolved in this study, they likely serve important regulatory functions.
High-resolution views of interacting interfaces between adjacent translating ribosomes
Multi-ribosome assemblies known as polysomes were identified in the mid-1960s51 and were often observed to form helical assemblies in previous low-resolution images52,53. Short segments with two or three colliding ribosomes, known as disomes and trisomes, respectively, were also frequently observed54,55,56, and one disome interface was resolved to high resolution using purified short polysomes57. 3D structures of helical polysomes in both prokaryotic and eukaryotic species have been studied by cryo-ET at moderate resolutions22,58,59,60. Here, we analyzed the polysome densities in different ribosome states. All translating states show some polysome density at the mRNA entry and exit positions (Supplementary Fig. 9a). Interestingly, the A/P-P/E states show strong polysome densities at the mRNA entry position, indicating that these states are enriched in trailing ribosomes. Additional classification of di-ribosomal particles resolved two main classes of di-ribosomes with different interaction interfaces, at resolutions between 3 to 4 Å (Fig. 5a, b, Supplementary Fig. 10 and Supplementary Table 6). One class (class 2, “top-top” interaction) is similar to previously published disome interfaces and has mainly 40S-40S subunit interactions12,23,57 (Fig. 5a, b). The other class (class 1, “top-back” interaction) is distinguished by a large rotation of the leading ribosome relative to the trailing ribosome (Fig. 5a–c). This interface is mainly formed by 60S-40S subunit interactions and is similar to the helical arrangement revealed by previous cryo-ET studies at low resolution (~25 Å)22,60. When the cryo-EM data were analyzed using a large box size, we indeed observed helical polysome density (Fig. 5d). Furthermore, fitting the class 1 di-ribosome structures indeed yielded a model for the helical polysome structure consistent with previous analysis22 (Fig. 5e). The leading ribosomes with the class 2 interaction consist of a mixture of both unrotated and rotated translation states, suggesting that the leading ribosome is actively translating (Fig. 5a), similar to that recently visualized in E. coli polysome61. By contrast, the compact class 1 helical polysomes consist of only specific A/P-P/E states for both leading and trailing ribosomes (Fig. 5a), suggesting that those ribosomes are stalled in that state, possibly because collision sterically hinders translocation. This is consistent with previous reports that protein synthesis inhibition and viral infection can induce helical polysomes53,62.
Native drug binding in the human ribosome
We further demonstrated that in situ single-particle cryo-EM can be applied to study the effects of pharmaceutical agents in living cells. We treated HEK293A cells with two ribosome inhibitors, cycloheximide (CHX), which competes with tRNA for binding at the E site14, and the anticancer drug homoharringtonine (HHT), which prevents peptide bond formation after aminoacyl-tRNA binding at the A site14,34,63. With our pipeline, we obtained a 2.4 Å 80S ribosome consensus map with clear densities for both compounds, allowing for atomic-level model building for the drug-ribosome interactions (Fig. 6a and Supplementary Table 1).
a Consensus map from the HHT + CHX-treated cells resolved at 2.4 Å resolution. The density (semitransparent surfaces) for CHX (top inset, red sticks), HHT (bottom inset, blue sticks), as well as their surrounding residues or nucleotides (modeled as grey sticks) are well resolved. b Comparison of the spermidine molecule contacting CHX with compositions of the same binding pocket observed in in vitro studies. c The translation states obtained from the HHT + CHX-treated cells, displayed as in Fig. 2a. CHX and HHT are highlighted in red and blue, respectively. d Only one hibernating state was observed from the CHX + HHT-treated cells.
Our in situ cryo-EM maps permitted us to visualize how the binding of the two inhibitors alters the dynamic landscape of protein synthesis. Of particular note, previous literature contains inconsistency regarding whether a spermidine molecule is present at the CHX binding pocket45,46. Here, we show clear density of the spermidine molecule contacting CHX and altering the chemical environment of the drug’s binding pocket (Fig. 6b). This exemplifies the importance of in situ ligand characterization for drug design and screening, as well as for understanding the role of ligands in translation regulation.
By employing focused classification and refinement, we resolved seven ribosome states in the CHX + HHT-treated cells at 3–4 Å resolutions, each with clear drug densities (Fig. 6c, d). Six of the observed states corresponded to translating ribosomes (states 1b, 1c, 2b, 2c, 3b, 3c; Fig. 6c and Supplementary Table 5) and accounted for 58% of 80S ribosomes. A single hibernating state (state 7c, Fig. 6d) accounted for the remaining 42% of 80S, which was an increase from 30% for untreated cells. As anticipated, no state harbored a tRNA in the E or P/E sites, consistent with the inhibition of tRNA translocation from the P to the P/E site by these two inhibitors. Our observations from the drug-treated cells also provided insight into the mechanistic role of the Z-tRNA. Nearly 50% of translating ribosomes (28.7% of all ribosomes) in the drug-treated cells contained Z-tRNA. This is higher than the proportion of Z-tRNA-containing states in the untreated cells (27.3% of translating ribosomes, 14.5% of all ribosomes) (Figs. 3a, 6c). This is possibly because CHX and HHT treatment abrogate tRNA binding at the E site, which competes with tRNA binding at the Z site. Importantly, the CHX and HHT treatment potently inhibited the P-to-E translocation of tRNA and thereby inhibited translation. Our observation is consistent with the previously reported enrichment of Z-tRNA on stalled ribosomes22. An accumulation of Z-tRNA in these translocation-stalled ribosome states could suggest that these stalled ribosomes picked up tRNA from solution for potential reverse translocation, which would need future studies to clarify.
Discussion
We demonstrate here that the combination of automated cryo-FIB milling and single-particle cryo-EM holds the potential to transform the study of structural biology within native cellular contexts. This approach leverages advances in high-resolution template matching algorithms (GisSPA and 2DTM), allowing particles to be accurately identified and aligned from single-tilt micrographs, bypassing the need for tilt series collection. Compared to cryo-ET, this method is more user-friendly and integrates well with established data processing packages like CryoSPARC32 and RELION64. It also enables higher throughput data collection and achieves resolutions comparable to in vitro purified samples in an in situ setting. On the other hand, the in situ single-particle approach is inherently limited by the signal-to-noise ratio—particularly for smaller particles—and by the need for high particle abundance. Moreover, it does not provide an accurate representation of three-dimensional spatial relationships among particles or information about their surrounding cellular environment. We envision that combining the single-particle approach with cryo-ET will offer a more comprehensive and powerful framework for in situ structural biology.
Using this in situ cryo-EM method, we resolved human ribosomes in cells to 2.2 Å resolution. Our high-resolution consensus structures confirmed the presence of rRNA modifications previously identified through mass spectrometry and cryo-EM of purified ribosomes29,33,34,35. By comparing to recent in vitro studies, we observed numerous differences in local rRNA conformation, as well as native assignment of small molecules and ion occupancy. While our observations align with previously published in situ cryo-ET maps12,22, our higher resolution maps allowed us to identify these differences unambiguously. In addition to core ribosomal components, we identified additional protein factors, including NACβ and EDF1, many of which were previously captured on purified ribosomes with supplementation beyond physiological concentrations. Our findings suggest that some of these factors may be more commonly associated with ribosomes in vivo than previously thought. Additionally, we identified previously unobserved peptide densities—potentially corresponding to SERBP1—bound to the 60S and 40S ribosomal subunits in cells. Further studies will be required to confirm their identity and clarify their functional significance.
Beyond the high-resolution consensus structure, we obtained 23 distinct classes of ribosomes in HEK293A cells, creating a comprehensive translational landscape from a single dataset. Our dataset contains ~50 times more particles than previous cryo-ET studies12, allowing for more thorough and reliable classification. Unlike in vitro cryo-EM studies that use nucleoside triphosphate analogs or drugs to enrich for particular states, our translation landscape captures the translation cycle without such perturbations. The relative abundance of different classes sheds light on the kinetics of the state transitions. Notably, we identified tRNAs abundantly bound to the Z site. Previously, the Z site tRNA was discovered in purified hibernating ribosomes48, and was shown to have relatively low abundance in cells12 and to be enriched on stalled ribosomes22. Our structures show clear densities of mRNA, nascent peptides, and additional tRNAs, suggesting that some of these ribosomes might be actively translating, although we also observed increased Z-tRNA occupancy on inhibitor-stalled ribosomes. In addition, the 5% glycerol used as a cryoprotectant in our protocol, although applied for only a short duration (a few minutes), may induce cellular hibernation as a result of osmotic shock65. The observed enrichment of hibernating 80S ribosomes (Fig. 3) could reflect such stress. Consequently, the distribution of functional ribosome states (e.g., canonical elongating vs. hibernating), the prevalence of factors like SERBP1, and specific tRNA states (e.g., Z-tRNA) reported here may be influenced by the preparation conditions and may not solely represent the basal, unstressed cellular state. Future methodological improvements with faster freezing or gentler handling may help minimize these effects and more accurately preserve the native translational landscape.
Accurate and physiological models are crucial for molecular docking predictions, particularly in drug development. To test the potential of in situ cryo-EM in drug development, we investigated HEK293A cells treated with cycloheximide and homoharringtonine. We successfully identified drug densities and essential polyamine cofactors in our cryo-EM maps, as well as alterations in the translation landscape in the presence of these drugs. Our results suggest that future drug development efforts could benefit from this in situ approach.
Determining biological macromolecular structures at high resolution within their native cellular environment has long been a goal in structural biology. As highlighted in a recent review66 and exemplified in this study, in vitro structural biology does not always recapitulate what happens inside cells. A key feature of in situ data is that raw datasets contain the full repertoire of cellular factors and can be reanalyzed for entirely different macromolecular targets. This versatility opens exciting possibilities for structural biology to uncover the mechanisms of various cellular processes as they occur in their native context and to study differences across cell types or even among patient samples. The continued development of efficient, high-resolution in situ structural biology tools will undoubtedly accelerate this exploration.
Methods
Cell culture
The 293A cells (Thermo Fisher Scientific) were cultured in Dulbecco’s Modified Eagle Medium (DMEM, Gibco) supplemented with 10% fetal bovine serum (FBS, Sigma) and 1% Pen/Strep (Gibco). The cells were passaged every 2 days, following standard cell culture procedures. Cells were maintained at a temperature of 37 °C in a humidified chamber atmosphere containing 5% CO2.
Cryo-EM sample preparation
The R2/4 holey carbon gold grids (200 mesh, Quantifoil) were glow discharged for 25 seconds at 20 mA and placed in 3.5 cm cell culture dishes (Matsunami). The grids were incubated with 70% ethanol and treated with UV for 1 hour. The grids were then washed 6 times with sterile distilled water and soaked in 0.1 mg/ml polylysine (Gibco) for over 4 hours to assist with cell attachment. The polylysine was removed, and the grids were rinsed twice with sterile distilled water. Then, 2 mL of suspended 293 A cells (~500,000 cells/ml) were applied to the culture dishes. Inhibitors-untreated cells were cultivated for 5–6 hours before plunge freezing. Two datasets of inhibitors-untreated cells were collected. In the case of the untreated dataset 1, the medium was changed to live cell imaging buffer (Invitrogen) with 5% glycerol before freezing within 30 minutes or less. For the untreated dataset 2, the medium was supplemented with a final 5% glycerol prior to freezing. The class distribution was highly similar between the two datasets (Supplementary Fig. 12), although an increased occurrence of hibernating states was observed in dataset 1, suggesting that the nutrition-free imaging buffer might have induced a state of cellular starvation, leading to the higher prevalence of hibernating states. The two datasets were combined for structural determination, but only dataset 2 from cells under normal growth conditions was used for translation state distribution analysis. For the HHT + CHX-treated cells, cells were first cultured for 4 hours on grids, then supplemented with HHT and CHX at a final concentration of 20 μM and 100 μg/ml (100 mg/ml stock in DMSO), respectively for another 1.5 hours. Like the untreated dataset 2, the medium was supplemented with 5% glycerol before freezing.
The grids were plunge-frozen using Vitrobot Mark 4 (Thermo Fisher Scientific). The grids were manually blotted for 4 seconds from the back side in a chamber at room temperature with 100% humidity. The frozen grids were then milled using an Aquilos machine (Thermo Fisher Scientific)67. The Maps software (Thermo Fisher Scientific) was used to select cells as milling targets. The grids were coated with an organometallic platinum layer for 26 seconds using a gas injection system (GIS)67. The AutoTEM68 software was used to set up automatic milling procedures, employing decreasing gallium ion-beam currents ranging from 0.5 nA to 10 pA, with a milling angle of 9 degrees. For the untreated dataset 1, 120 lamellae were polished with a final polishing current of 30 or 10 pA to a thickness of 120 to 180 nm. For the untreated dataset 2, 75 lamellae were polished with a final polishing current of 30 or 10 pA to a thickness of 120 to 130 nm. For the HHT + CHX-treated dataset, 110 lamellae were polished at 50 pA to a thickness of 120 nm. All reported lamellae thicknesses refer to the targeted lamellae thickness parameters during the milling process in AutoTEM.
It is worth noting that during cryo-FIB milling for the HHT + CHX-treated cells, higher polishing currents were used and lamellae were cut thinner, yet the resolution of the consensus map for this dataset was lower than for the untreated datasets, potentially due to greater damage incurred during cryo-FIB milling69.
Cryo-EM data acquisition
Data acquisition was conducted on a Titan Krios G2 electron microscope (Thermo Fisher Scientific), operated at 300 kV with an energy filter (Gatan) and K3 direct electron detector (Gatan). The imaging was performed at 5760 × 4092 pixels dimensions, with a pixel size of 1.068 Å. For single-particle data collection, a multi-shot pattern (18 shots) was employed at a single tilt angle (−9 or 9 degrees) to compensate for the angle at which the lamellae were cut. The target defocus ranged from −1.2 to −2.5 μm, and a total dose of 50 e/Å2 was applied during each shot. The data collection process was completed using the SerialEM software70. For the untreated dataset 1, 12,826 movies were acquired. For the untreated dataset 2, 6479 movies were acquired. For the HHT + CHX-treated sample.
Cryo-EM data processing
The acquired movies were processed for motion correction71 and Contrast Transfer Function (CTF) correction72 using CryoSPARC32. The processed micrographs were then manually curated to remove movies that are blurred or do not capture the cellular interior, resulting in 10,691 good micrographs for the untreated dataset 1, 5954 for the untreated dataset 2, and 5731 for the HHT + CHX-treated dataset. These micrographs were imported to GisSPA for particle picking, using a published 60S ribosome map (EMD-15113), low-pass filtered to 8.5 Å, as the template. Particles picked by GisSPA, along with their Euler angle information, were extracted using RELION364 and imported into CryoSPARC for subsequent analysis. The data processing workflows are summarized in Supplementary Fig. 3 (untreated) and Supplementary Fig. 11 (HHT + CHX-treated).
For the focused classification process, a 40S mask with a dilation radius of 1 and soft padding width of 5 pixels was generated by CryoSPARC32,73 to segregate the 80S and 60S particles effectively, as well as to remove false positives in particle picking. A class of 1,001,533 particles gave rise to clear 40S density, which had not been included in the particle-picking reference, and were unbiasedly identified as the 80S particles. After this initial classification, a 60S mask was employed to further refine the 80S particles to exclude additional low-quality particles. Using a final set of 411,079 80S particles, a consensus structure of the human 80S ribosome at 2.6 Å resolution was obtained using local refinement in CryoSPARC. The 2.6 Å map was used to repick particles from all the micrographs using GisSPA and CisTEM and led to 1,868,414 final particles of 80S, which after 3D classification on 60S and reference-based motion correction74 resulted in a consensus cryo-EM map at 2.2 Å resolution with 1,272,627 particles showing in the FSC curve (Supplementary Fig. 1c)75,76. Focused classification using a mask on the 40S body and 40S head improved resolution in those regions, and was used to create the composite map, as well as to identify the unknown density bound to the head of the 40S subunit. 3D classification was then used to distinguish the different conformations of the 80S ribosomes. Firstly, 40S-mask was used to classify the particles based on different 40S rotations. Then, masks covering the known binding sites of tRNA or cofactors were used to perform further classifications32,73. Each resultant translation state was then locally refined to a resolution of below 3 Å, except for the P state at 3.26 Å resolution.
The 60S dataset was also analyzed. This dataset contains both free 60S particles and noise particles. To remove the noise particles, the 60S particles were first 3D classified into two classes without using a local focus mask. One class had significantly less contrast, could not be refined above 6 Å, and did not contain additional factor densities. This class was discarded and not used in any further analysis. The other class, containing 520,484 particles, was locally further classified to remove low-quality particles. 220,458 particles of 60S can be refined to 2.9 Å. These 60S particles were further 3D classified using a focus mask covering the location of eIF6, resulting in one class (132,487 particles) with eIF6, and one without (88,131 particles). Each class was locally refined to resolutions of 3.0 and 3.1 Å, respectively. When repicking 80S particles, 225,80 particles of 60S can also be picked. Combining these particles and removing duplicate and bad quality particles, A final class of 248,791 particles was refined to 2.5 Å resolution after reference-based motion correction. Besides the high-resolution features, EBP1 density was present in this consensus map, but not in the original template that had been used for particle picking, suggesting that this particle set was largely free of template bias.
The same procedure was applied to the HHT + CHX-treated dataset. We obtained an initial 2.9 Å resolution 80S ribosome consensus map with 168,292 particles. Further classification with 40S, tRNAs, and cofactors mask yielded 6 translation states and 1 hibernating state. All states were refined to resolutions of 3–4 Å. For the 60S particles, a 4.1 Å resolution consensus map was made using 65,012 particles. Of these, 27,744 particles contain eIF6 and were refined to a 4.5 Å resolution map. The 80S particles were repicked as described for the WT dataset, which resulted in a 2.4 Å consensus 80S map with 463,982 particles.
Model building and refinement
Reference structures (6Z6M, 4V6X, 6ZM7) were used as starting models to generate the 80S ribosome model for each conformational class. For the HHT + CHX-treated ribosome models, structures from 7Q08 and 8A3D were used as references. Additionally, 7OSM and 7QWR were used as references for building the mRNA and nascent peptide models. 7QW7 was used for building eIF6 densities. These initial models were fitted into the experimental maps and refined using Phenix44. The refined models were adjusted manually using Coot45 and then further refined in Phenix for the final models. All the ions were identified by comparing to ions in previously reported high-resolution structures and based on geometry35.
Model building on the 2 patches of unknown densities on ribosomes
For unknown density 2, the C-alpha positions were manually traced in Coot, and the most likely side chains were assigned based on the observed high-resolution densities. BLASTp77 was then used—with uncertain positions marked as “x”—to identify candidate proteins. SERBP1 aa. 1–37 was determined to be the most likely match, and a preliminary model was built using its sequence. To further validate this assignment, AlphaFold 250 was employed to predict the structure of SERBP1 aa. 1–37 in the context of neighboring proteins (see AlphaFold Prediction section). The independent prediction matched the experimental density map and the built model with exceptional agreement, with high pLDDT scores and low PAE (Supplementary Fig. 8b-c). Subsequently, a BLASTp search using a side-chain–weighted scoring matrix (as described in cryoID78) was performed against the human reference proteome (UP000005640_9606, UniProt), which further confirmed SERBP1 aa. 1–37 as the overwhelming top hit.
For unknown density 1, limited resolution allowed a single aromatic density to be visually identified. AlphaFold 2 was used to predict structures for various SERBP1 fragments containing aromatic residues, and it was found that the fragment corresponding to aa. 389–402 matched the experimental density with exceptional agreement, again with excellent pLDDT scores and low PAE (Supplementary Fig. 8b, c). A BLASTp search using the cryoID scoring matrix further confirmed SERBP1 residues 389–402 as the top hit—three orders of magnitude more significant than the next best match.
AlphaFold Prediction
AlphaFold2 multimer50 was used to confirm the binding between SERBP1 and various ribosomal proteins. The default setting was used in all cases, and the model with the highest overall pLDDT was used for downstream analysis. For predicting SERBP1 N-terminal segment binding to ribosome (Fig. 3b.ii), we used the SERBP1 sequence aa.1–49 (Uniprot accession number Q8NC51 isoform 2), eL30 aa.9–106 (Uniprot accession number P62888), eL34 aa.2–115 (Uniprot accession number P49207), and eL27 aa. 2–136 (Uniprot accession number P61353). For predicting SERBP1 C-terminal segment binding to ribosome (Fig. 3b.iv), we used the human SERBP1 sequence aa. 380–402 (Uniprot accession number Q8NC51 isoform 2) and the full-length RACK1 sequence (Uniprot accession number P63244).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic models of the untreated composite, the consensus with EBP1, 60S, 60S with eIF6, state 1a, 1b, 1c, 2a, 2b, 2c, 2a’, 2b’, 2c’, 3a, 3b, 3c, 4, 5, 6a, 6b, 7a, 7b, 7c, 8a, 9a, 10a, 11a, the HHT- and CHX-treated consensus, state 1b, 1c, 2b, 2c, 3b, 3c, 6 h, and the t-t and t-b disome consensus composite have been deposited in the Protein Data Bank (PDB) under accession codes: 9PBE, 9PA7, 9P9J, 9P9K, 9P72, 9P6Z, 9PKG, 9P8B, 9P7O, 9P76, 9P7K, 9P7N, 9P7L, 9P7Y, 9P7X, 9P7W, 9P8H, 9P73, 9P9I, 9P9H, 9P78, 9P79, 9P8C, 9P7A, 9P7C, 9P7D, 9P7E, 9PKD, 9P8I, 9P7J, 9P7G, 9P7F, 9PKB, 9B0S, 9B0Q. The globally and locally refined cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes: EMD-71477, EMD-71440, EMD-71441, EMD-71442, EMD-71443, EMD-71331, EMD-71329, EMD-71696, EMD-71371, EMD-71349, EMD-71333, EMD-71345, EMD-71348, EMD-71346, EMD-71357, EMD-71356, EMD-71355, EMD-71377, EMD-71332, EMD-71334, EMD-71335, EMD-71372, EMD-71336, EMD-71337, EMD-71338, EMD-71339, EMD-71694, EMD-71378, EMD-71344, EMD-71341, EMD-71340, EMD-71690, EMD-44052, EMD-44057, EMD-44056, EMD-44050, EMD-44054, EMD-44053. Also see Supplementary Table 1 for a complete list of accession codes.
References
Korostelev, A. A. The structural dynamics of translation. Annu. Rev. Biochem. 91, 245–267 (2022).
Voorhees, R. M. & Ramakrishnan, V. Structural basis of the translational elongation cycle. Annu. Rev. Biochem. 82, 203–236 (2013).
Ramakrishnan, V. The ribosome emerges from a black box. Cell 159, 979–984 (2014).
Schuller, A. P. & Green, R. Roadblocks and resolutions in eukaryotic translation. Nat. Rev. Mol. Cell Biol. 19, 526–541 (2018).
Anger, A. M. et al. Structures of the human and Drosophila 80S ribosome. Nature 497, 80–85 (2013).
Beckmann, R. et al. Architecture of the protein-conducting channel associated with the translating 80S ribosome. Cell 107, 361–372 (2001).
Lyumkis, D. et al. Structural basis for translational surveillance by the large ribosomal subunit-associated protein quality control complex. Proc. Natl. Acad. Sci. USA 111, 15981–15986 (2014).
Shen, P. S. et al. Protein synthesis. Rqc2p and 60S ribosomal subunits mediate mRNA-independent elongation of nascent chains. Science 347, 75–78 (2015).
Sinha, N. K. et al. EDF1 coordinates cellular responses to ribosome collisions. Elife 9 https://doi.org/10.7554/eLife.58828 (2020).
Jomaa, A. et al. Mechanism of signal sequence handover from NAC to SRP on ribosomes during ER-protein targeting. Science 375, 839–844 (2022).
Brown, A. & Shao, S. Ribosomes and cryo-EM: a duet. Curr. Opin. Struct. Biol. 52, 1–7 (2018).
Xing, H. et al. Translation dynamics in human cells visualized at high resolution reveal cancer drug action. Science 381, 70–75 (2023).
Brown, A., Shao, S., Murray, J., Hegde, R. S. & Ramakrishnan, V. Structural basis for stop codon recognition in eukaryotes. Nature 524, 493–496 (2015).
Garreau de Loubresse, N. et al. Structural basis for the inhibition of the eukaryotic ribosome. Nature 513, 517–522 (2014).
Wong, W. et al. Cryo-EM structure of the Plasmodium falciparum 80S ribosome bound to the anti-protozoan drug emetine. Elife 3 https://doi.org/10.7554/eLife.03080 (2014).
Wan, W. & Briggs, J. A. G. The resolution revolution: recent advances in cryoEM. Methods in Enzymology. Vol. 579 (ed. Crowther R. A.) 329–367 (2016).
Scheres, S. H. W. In Methods in Enzymology 579 (The Resolution Revolution: Recent Advances in cryoEM) (ed. Crowther, R. A.), 125–157 (Academic Press, 2016).
Zhang, P. Advances in cryo-electron tomography and subtomogram averaging and classification. Curr. Opin. Struct. Biol. 58, 249–258 (2019).
Marko, M., Hsieh, C., Schalek, R., Frank, J. & Mannella, C. Focused-ion-beam thinning of frozen-hydrated biological specimens for cryo-electron microscopy. Nat. Methods 4, 215–217 (2007).
Hylton, R. K. & Swulius, M. T. Challenges and triumphs in cryo-electron tomography. Iscience 24, 102959 (2021).
Young, L. N. & Villa, E. Bringing Structure to Cell Biology with Cryo-Electron Tomography. Annu. Rev. Biophys. 52, 573–595 (2023).
Fedry, J. et al. Visualization of translation reorganization upon persistent ribosome collision stress in mammalian cells. Mol. Cell 84, 1078–1089.e1074 (2024).
Xue, L. et al. Visualizing translation dynamics at atomic detail inside a bacterial cell. Nature 610, 205–211 (2022).
Hoffmann, P. C. et al. Structures of the eukaryotic ribosome and its translational states in situ. Nat. Commun. 13, 7435 (2022).
Xue, L., Spahn, C. M. T., Schacherl, M. & Mahamid, J. Structural insights into context-dependent inhibitory mechanisms of chloramphenicol in cells. Nat. Struct. Mol. Biol. 32, 257–267 (2025).
You, X. et al. In situ structure of the red algal phycobilisome-PSII-PSI-LHC megacomplex. Nature 616, 199–206 (2023).
Bronwyn, A. L., Benjamin, A. H. & Nikolaus, G. Baited reconstruction with 2D template matching for high-resolution structure determination in vitro and in vivo without template bias. bioRxiv, 2023.2007.2003.547552 https://doi.org/10.1101/2023.07.03.547552 (2023).
Rickgauer, J. P., Choi, H., Moore, A. S., Denk, W. & Lippincott-Schwartz, J. Structural dynamics of human ribosomes in situ reconstructed by exhaustive high-resolution template matching. Mol. Cell 84, 4912–4928.e4917 (2024).
Faille, A., Dent, K. C., Pellegrino, S., Jaako, P. & Warren, A. J. The chemical landscape of the human ribosome at 1.67 A resolution. bioRxiv, 2023.2002.2028.530191 https://doi.org/10.1101/2023.02.28.530191 (2023).
Cheng, J. et al. Determining protein structures in cellular lamella at pseudo-atomic resolution by GisSPA. Nat. Commun. 14, 1282 (2023).
Lucas, B. A. et al. Locating macromolecular assemblies in cells by 2D template matching with cisTEM. Elife 10 https://doi.org/10.7554/eLife.68946 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Taoka, M. et al. Landscape of the complete RNA chemical modifications in the human 80S ribosome. Nucleic Acids Res 46, 9289–9298 (2018).
Natchiar, S. K., Myasnikov, A. G., Kratzat, H., Hazemann, I. & Klaholz, B. P. Visualization of chemical modifications in the human 80S ribosome structure. Nature 551, 472–477 (2017).
Holvec, S. et al. The structure of the human 80S ribosome at 1.9 Å resolution reveals the molecular role of chemical modifications and ions in RNA. Nat. Struct. Mol. Biol. 1–14 (2024).
Nierhaus, K. H. Mg2+, K+, and the ribosome. J. Bacteriol. 196, 3817–3819 (2014).
Cohen, S. S. & Lichtenstein, J. Polyamines and ribosome structure. J. Biol. Chem. 235, 2112–2116 (1960).
Konecki, D., Kramer, G., Pinphanichakarn, P. & Hardesty, B. Polyamines are necessary for maximum in vitro synthesis of globin peptides and play a role in chain initiation. Arch. Biochem. Biophys. 169, 192–198 (1975).
Bartetzko, A. & Nierhaus, K. H. In Methods in Enzymology 164 650-658 (Elsevier, 1988).
McCarthy, B. The effects of magnesium starvation on the ribosome content of Escherichia coli. Biochim. Biophys. Acta (BBA)-Spec. Sect. Nucleic Acids Relat. Subj. 55, 880–889 (1962).
Gesteland, R. F. Unfolding of Escherichia coli ribosomes by removal of magnesium. J. Mol. Biol. 18, 356–IN314 (1966).
Weiss, R. L. & Morris, D. R. Cations and ribosome structure. I. Effects of the 30S subunit of substituting polyamines for magnesium ions. Biochemistry 12, 435–441 (1973).
Kimes, B. W. & Morris, D. R. Cations and ribosome structure. II. Effect of the 50S subunit of substituting polyamines for magnesium ions. Biochemistry 12, 442–449 (1973).
Tal, M. Metal ions and ribosomal conformation. Biochim. Biophys. Acta (BBA)-Nucleic Acids Protein Synth. 195, 76–86 (1969).
Holm, M. et al. mRNA decoding in human is kinetically and structurally distinct from bacteria. Nature 617, 200–207 (2023).
Myasnikov, A. G. et al. Structure–function insights reveal the human ribosome as a cancer target for antibiotics. Nat. Commun. 7, 12856 (2016).
Gemmer, M. et al. Visualization of translation and protein biogenesis at the ER membrane. Nature 614, 160–167 (2023).
Brown, A., Baird, M. R., Yip, M. C., Murray, J. & Shao, S. Structures of translationally inactive mammalian ribosomes. Elife 7, e40486 (2018).
Wild, K. et al. MetAP-like Ebp1 occupies the human ribosomal tunnel exit and recruits flexible rRNA expansion segments. Nat. Commun. 11, 776 (2020).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. bioRxiv, 2021.2010.2004.463034 https://doi.org/10.1101/2021.10.04.463034 (2022).
Warner, J. R., Rich, A. & Hall, C. E. Electron microscope studies of ribosomal clusters synthesizing hemoglobin. Science 138, 1399–1403 (1962).
Weiss, P. & Grover, N. Helical array of polyribosomes. Proc. Natl. Acad. Sci. 59, 763–768 (1968).
Djaczenko, W., Benedetto, A. & Pezzi, R. Formation of helical polyribosomes in poliovirus-infected cells of the 37 RC line. J. Cell Biol. 45, 173 (1970).
Juszkiewicz, S. et al. ZNF598 is a quality control sensor of collided ribosomes. Mol. Cell 72, 469–481. e467 (2018).
Ikeuchi, K. et al. Collided ribosomes form a unique structural interface to induce Hel2-driven quality control pathways. EMBO J. 38, e100276 (2019).
Matsuo, Y. et al. RQT complex dissociates ribosomes collided on endogenous RQC substrate SDD1. Nat. Struct. Mol. Biol. 27, 323–332 (2020).
Narita, M. et al. A distinct mammalian disome collision interface harbors K63-linked polyubiquitination of uS10 to trigger hRQT-mediated subunit dissociation. Nat. Commun. 13, 6411 (2022).
Brandt, F. et al. The native 3D organization of bacterial polysomes. Cell 136, 261–271 (2009).
Brandt, F., Carlson, L. A., Hartl, F. U., Baumeister, W. & Grunewald, K. The three-dimensional organization of polyribosomes in intact human cells. Mol. Cell 39, 560–569 (2010).
Myasnikov, A. G. et al. The molecular structure of the left-handed supra-molecular helix of eukaryotic polyribosomes. Nat. Commun. 5, 5294 (2014).
Flügel, T. et al. Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM. Nat. Commun. 15, 1756 (2024).
Popp, J. A. & Hisashi, S. Inhibition of protein synthesis and induction of helical polysomes in rat liver by N-hydroxy-2-fluorenylacetamide. Chem. -Biol. Interact. 9, 37–43 (1974).
Gürel, G., Blaha, G., Moore, P. B. & Steitz, T. A. U2504 Determines the Species Specificity of the A-Site Cleft Antibiotics: The Structures of Tiamulin, Homoharringtonine, and Bruceantin Bound to the Ribosome. J. Mol. Biol. 389, 146–156 (2009).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Mazur, P. Freezing of living cells: mechanisms and implications. Am. J. Physiol. -Cell Physiol. 247, C125–C142 (1984).
Beck, M., Covino, R., Hänelt, I. & Müller-McNicoll, M. Understanding the cell: Future views of structural biology. Cell 187, 545–562 (2024).
Wagner, F. R. et al. Preparing samples from whole cells using focused-ion-beam milling for cryo-electron tomography. Nat. Protoc. 15, 2041–2070 (2020).
Kuba, J. et al. Advanced cryo-tomography workflow developments – correlative microscopy, milling automation and cryo-lift-out. J. Microsc. 281, 112–124 (2021).
Lucas, B. A. & Grigorieff, N. Quantification of gallium cryo-FIB milling damage in biological lamellae. Proc. Natl. Acad. Sci. USA 120, e2301852120 (2023).
Mastronarde, D. N. SerialEM: A program for automated Tilt series acquisition on Tecnai microscopes using prediction of specimen position. Microsc. Microanal. 9, 1182–1183 (2003).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Barchet, C. et al. Focused classifications and refinements in high-resolution single particle cryo-EM analysis. J. Struct. Biol. 215, 108015 (2023).
Rubinstein, J. L. & Brubaker, M. A. Alignment of cryo-EM movies of individual particles by optimization of image translations. J. Struct. Biol. 192, 188–195 (2015).
Saxton, W. & Baumeister, W. The correlation averaging of a regularly arranged bacterial cell envelope protein. J. Microsc. 127, 127–138 (1982).
Van Heel, M., Keegstra, W., Schutter, W. & Van Bruggen, E. Arthropod hemocyanin structures studied by image analysis. Life Chem. Rep. Suppl. 1, 5 (1982).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Ho, C.-M. et al. Bottom-up structural proteomics: cryoEM of protein complexes enriched from the cellular milieu. Nat. Methods 17, 79–85 (2020).
Acknowledgements
The project is funded by Yale's discretionary funds to Y.X. The Aquilos Cryo-FIB is funded by R01AI152421 (J.L.). We thank Yale CryoEM Resource for maintaining the Krios Cryo-TEM used for data collection. We thank the Yale Center for Research Computing for support with computing resources. We thank colleagues at the Department of Molecular Biophysics and Biochemistry at Yale University for suggestions and discussions on this study.
Author information
Authors and Affiliations
Contributions
Y.X. supervised the project. W.Z. prepared the sample and performed FIB milling with the help of J.F.L., J.L., and W.G. W.Z., S.P.W., and Y.X. collected the data. W.Z., P.C., S.H.W., Y.Z., K.Z., and Y.X. processed the data. W.Z., J.W., Y.Z., S.H.W., S.C.D., E.J.B., I.L., and Y.X. interpreted the data. W.Z., J.W., Y.Z., and C.Z. built and refined the atomic models. W.Z., Y.Z., S.H.W., E.J.B., S.C.D., and Y.X. prepared the Figures. W.Z., E.J.B., Y.Z., I.L., and J.W. wrote the manuscript with input from all authors. J.W., Y.Z., E.J.B., I.L., S.H.W., W.Z., and Y.X. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, W., Zhang, Y., Wang, J. et al. Visualizing the translation landscape in human cells at high resolution. Nat Commun 16, 10757 (2025). https://doi.org/10.1038/s41467-025-65795-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65795-9
This article is cited by
-
NatA engages in multi-factor complexes at the ribosomal polypeptide tunnel exit
Nature Communications (2026)
-
Structural basis of TACO1-mediated efficient mitochondrial translation
Nature Communications (2026)