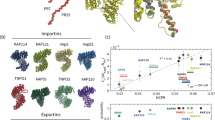
Abstract
A cellular gradient of the GTPase Ran orchestrates the movement of import and export complexes through the Nuclear Pore Complex (NPC). Ran-GTP modulates two essential activities of importin β during nuclear import. On the one hand, it reduces the avidity of importin β for phenylalanine-glycine-rich nucleoporins (FG-nups), thereby facilitating the passage of import complexes through the permeability barrier. On the other hand, it disassembles import complexes, releasing the import cargo into the nucleus. The precise mechanisms by which Ran-GTP modulates importin β activities have remained hypothetical. Leveraging cryogenic electron microscopy (cryo-EM) single-particle analysis, in this paper, we describe five distinct conformational states of importin β in complex with various effectors encountered during an import reaction, specifically IBB-cargos, FG-repeats, Ran-GTP, Ran-GTP:RanBP1, and Ran-GDP:RanBP1. Comparing these states allows us to decipher the conformational landscape of importin β without interference from crystallization agents and lattice forces. By correlating structural data with biochemical activities, we find that Ran-GTP, but not Ran-GDP, constrains the solenoid structure of importin β, closing high-affinity FG-binding pockets and displacing import cargos through allosteric crosstalk between the concave and convex surfaces. We propose that this allosteric mechanism is relevant to other β-karyopherins involved in nuclear import.
Similar content being viewed by others
Introduction
Importin β is the prototypical nuclear import receptor1,2,3,4 and the founding member of a family of transport receptors known as β-karyopherins5. Importin β binds import cargos directly or through the adapter protein importin α, which exists in seven isoforms in humans6. In the classical nuclear import pathway, cytoplasmic cargos containing a nuclear localization signal (NLS) assemble into a trimeric importin β:α1:NLS-cargo complex, which docks to the nuclear pore complex (NPC) and moves through the NPC channel due to importin β avidity for phenylalanine-glycine-rich nucleoporins (FG-nups)7,8. The chemical structure of the NPC inner channel is unknown, but ample evidence suggests FG-nups form a hydrogel that constitutes a selective permeability barrier9. The small GTPase Ran, enriched as Ran-GTP in the nucleus and Ran-GDP in the cytoplasm10,11, facilitates the import reaction by regulating the interaction between transport receptors (like importin β) and cargo, controlling the directionality of transport12. Ran-GTP binds importin β with high affinity, releasing the NLS-cargo in the nucleus. It also decreases the affinity of importin β for FG-nups, preventing importin β from stalling and thus clogging the NPC12,13,14,15. Importin β fragments that bind to the NPC but do not associate with Ran-GTP irreversibly clog the pore, inhibiting NLS-dependent protein import16.
Importin β can shuttle through the NPC on its own, without cargo or an adapter. In digitonin-permeabilized cells, purified importin β accumulates at NPCs and can enter the nucleus independently of importin α or cargo12,16. Single molecule/FRAP experiments17 directly observed importin β shuttling through NPCs, revealing transient interactions with FG repeats and a characteristic transport dwell time. A large portion of the entire cellular pool of importin β resides inside the NPC12,18,19, possibly divided into two pools within the NPC20: one pool is stably bound to the NPC, with a slow off-rate, regardless of the presence of Ran-GTP; the other pool is stably bound to the NPC only in the absence of Ran-GTP, which actively disengages importin β from the NPC21, ensuring receptor turnover and sustaining directional nuclear import.
Ran-GTP influences the permeability of the importin β:α1:NLS-cargo complex as it moves through the NPC22. Typically, large cargos require higher levels of Ran-GTP to navigate the NPC23,24, and Ran-GTP primarily regulates the cargo’s exit from the NPC22. Without Ran-GTP, cargos can enter and explore the entire NPC, but they are roughly 100 times more likely to exit the NPC on the cytoplasmic side, thus failing to reach the nucleus22. NLS-cargos bound to importin α1 require Ran-GTP and the export factor CAS17 to dissociate on the nucleoplasmic side of the NPC. In contrast, importin α1:NLS-cargo complexes that do not dissociate at the NPC are returned to the cytoplasm17. Recent evidence found that import and export complexes share the same central NPC path, rather than distinct lanes. Actively translocating complexes move within a confined ~40–50 nm annulus, while stalled complexes accumulate at the pore periphery21.
Ran binding protein 1, or RanBP1, is another key player in the import reaction. It exists as a soluble, 23 kDa protein located in the cytoplasm due to a strong nuclear export signal25, although some reports suggest that the protein may shuttle between the cytosol and the nucleus26,27,28. Four conserved RanBP1-like domains also exist in the filamentous nucleoporin Nup35829,30,31 at the cytoplasmic periphery of the NPC. A crystal structure revealed that RanBP1 binds Ran-GTP in a tight embrace32. The primary function of RanBP1 is to facilitate GTP hydrolysis by RanGAP1, countering the inhibitory activity of transport factors. Without RanBP1, the nanomolar binding affinity of Ran-GTP with importin β inhibits RanGAP1-dependent GTP hydrolysis. However, in the presence of RanBP1, the trimeric importin β:Ran-GTP: RanBP1 complex becomes susceptible to GTP hydrolysis, which additionally requires the importin β binding (IBB) domain of importin α1 in vitro33,34.
Structural and computational studies have been crucial in deciphering the organization, binding interactions, and conformational dynamics of importin β. Several crystal structures have revealed that importin β adopts a solenoid architecture formed by 19 HEAT repeats35, organized into two functionally polarized surfaces. Importin β inner (concave) surface, formed by HEAT helix B, binds Ran-GTP36 and the IBB domain37 of the adapter importin α, but can also associate directly with nonclassical cargos38,39,40. Importin β outer (convex) surface, formed by A helices, binds FG-nups41. A crystal structure of a fragment of importin β (res 1–442) bound to an FG-rich fragment of nucleoporin Nsp1p reveals two binding sites, one primary site on the A helix of HEAT 5 and another on HEAT 642. This FG-binding pattern was also observed in a structure of Kap95p, the yeast homolog of human importin β, bound to Nup1p43. Furthermore, a crystal structure of Kap95p bound to human Ran-GTP44 revealed an open conformation of Kap95p, in which the C-terminal domain moves away from the closed solenoid seen in the complex with the IBB domain. Based on the conformation of Kap95p observed in this crystal structure, it was proposed that Ran-GTP displaces cargo by inducing a change in the helicoidal pitch of importin β, which opens its concave surface and disassembles the import complex. However, it remains unclear, and is not explained by the above model, how Ran-GTP releases FG-nups from importin β during active transport.
Using cryogenic electron microscopy (cryo-EM) single-particle analysis (SPA), we determine six structures of importin β in a frozen-hydrated state bound to IBB-cargos, FG-nups, Ran-GTP, Ran-GTP: RanBP1, and Ran-GDP: RanBP1. This comparative structural analysis enables us to decipher the conformational plasticity of importin β, overcoming the limitations of previous crystal structures. Our work clarifies how the GTPase Ran-GTP alters the structure of importin β, regulating its binding affinities for cargos and FG-nups.
Results
Cryo-EM structure of importin β bound to IBBs and FG-nup
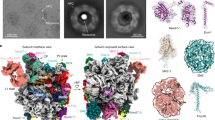
Molecular dynamics simulations41,45,46,47,48, Förster resonance energy transfer48, small-angle X-ray scattering46, crystallographic35,39,40,49, and biochemical50 data provided compelling evidence that importin β undergoes large structural fluctuations during nuclear transport. However, most importin β structures have been obtained in the crystalline state in the presence of dehydrating agents that can distort importin β flexible solenoid structure and potentially introduce artifacts40,49,51. This study aimed to investigate the structure and conformational changes that importin β undergoes during nuclear import under quasi-physiological conditions without potential crystallization-induced artifacts. We formed six complexes of importin β, corresponding to five discrete states observed during the nuclear import reaction. First (Fig. 1A), we assembled importin β with importin α1 IBB (αIBB) and XRIPα IBB (xIBB), an adapter protein involved in the nuclear import of replication protein A (RPA)52. Importin βːαIBB and importin βːxIBB are proxies for cytoplasmic importin β assemblies found in both classical and nonclassical import complexes. Second (Fig. 1B), we added an excess of the FxFG-rich nucleoporin Nsp142 to importin βːαIBB to mimic a classical import complex bound to high-affinity FG sites within the NPC central channel. Third (Fig. 1C), we assembled the heterodimeric importin β:Ran-GTP complex, which represents the final state of an import reaction presumably formed at the NPC basket, as importin β gets closer to the nucleus. Fourth (Fig. 1D), we assembled a trimeric importin β:Ran-GTP:RanBP1 complex that was identified nearly three decades ago33,53,54. And, finally, (Fig. 1E), we reconstituted the heterotrimeric importin β:Ran-GDP:RanBP1 complex53,54,55,56,57, which has also been known for decades but whose function remains unclear.
A Classical (importin β:α1:NLS-cargo) and nonclassical (importin β:XRP1α:RPA) import complexes assemble in the cytoplasm. B A classical import complex engages high-affinity FG-nups within the NPC (importin β:αIBB:FG). C An importin β:RanGTP:RanBP1 complex may form inside the NPC, whereas (D) the importin β:RanGDP:RanBP1 complex is thought to assemble at the cytoplasmic periphery. E At the nucleoplasmic periphery, where RanGTP concentration is highest, the terminal importin β:RanGTP complex is generated. The background image of the NPC was created in BioRender. Ko, T. (2025) https://BioRender.com/esum6zk.
All importin β complexes were vitrified and imaged using a 300 kV Krios microscope. A total of six cryo-EM reconstructions were determined using SPA, achieving maximal resolutions between 2.6 and 3.4 Å (Supplementary Figs. 1 and 2; Supplementary Table 1). The corresponding density maps had sufficiently high-resolution features to visualize all side chains (Supplementary Fig. 3), enabling us to build accurate atomic models of importin β bound to different effectors, which were refined to map-to-model correlation coefficients greater than 0.80 (Supplementary Table 1).
αIBB and xIBB stabilize distinct importin β conformations
We reconstructed importin β bound to the αIBB domain at 3.25 Å resolution (Supplementary Fig. 2A and Supplementary Table 1). This structure (Supplementary Fig. 4A), determined in a frozen-hydrated state, revealed significant variations compared to the crystal structure of the importin β:αIBB complex (RMSD ~ 4.7 Å)35, especially in the C-terminal HEATs 11–19 (Supplementary Fig. 4B). The N-to-C distance of the X-ray model is 11.5 Å shorter than the cryo-EM structure, suggesting that the constrained, snail-like conformation of importin β:αIBB crystallized at neutral pH35 represents a nonphysiological state. After establishing that cryo-EM is the preferred methodology for studying importin β curvature, we determined a cryo-EM reconstruction of importin β in complex with xIBB, which we refined to a resolution of 3.41 Å (Supplementary Fig. 2B and Supplementary Table 1). This structure revealed an elongated solenoid tightly bound to a ~60 Å long peptide, with importin β spanning ~110 Å (Fig. 2A). The xIBB exhibited strong density for residues 1–45 that contact the concave surface of importin β between HEATs 6–19. A comparison of the two cryo-EM structures of importin β in complex with xIBB and αIBB (solved at comparable resolutions) revealed systematic variations in the solenoid conformation (RMSD ~ 6.1 Å). The N-to-C distance of importin β bound to xIBB is 9 Å longer than that in complex with αIBB (Fig. 2B). The two IBBs are highly basic (isoelectric point of ~11) and exhibit a similar topology when bound to importin β (Fig. 2C), consisting of a long C-terminal α-helix and an unstructured N-terminal moiety. However, xIBB is slightly shorter: its C-terminal α-helix spans six turns of a helix (res. 20–45), compared to seven turns of the αIBB. Both IBBs contain an N-terminal 3/10 helix spanning residues R13–F17 and P8–Y13 in αIBB and xIBB, respectively. The bonding pattern is similar but not identical, primarily driven by electrostatic interactions. xIBB forms five salt bridges, seven hydrogen bonds, and 123 nonbonded contacts with importin β, which is significantly fewer than αIBB, where the binding interface with importin β is stabilized by eight salt bridges, ten hydrogen bonds, and 112 nonbonded contacts. Three tryptophan residues in importin β, W342, W430, and W86458 (Fig. 2C), play a crucial role in IBB recognition by making cation-π interactions with IBB arginine and lysine residues. Thus, cryo-EM reconstructions of importin β bound to IBBs reveal that the protein can extend its solenoid structure by up to 10%, repositioning the C-terminal α-helix of different IBBs relative to the HEATs 11–19.
A Representative 2D class averages (top) and the 3D structure of the importin β:xIBB complex (bottom). Importin β is shown in blue, while xIBB is depicted in salmon. The final 3.41 Å cryo-EM density (shown in gray) for xIBB is overlaid on the refined model. B Secondary structure superimposition of importin β solved in complex with xIBB (blue) and αIBB (light gray) is shown as beads-on-a-string. Each bead represents a HEAT repeat. C Structural comparison of xBB to αIBB, colored salmon and yellow, respectively. The figure was generated by superimposing the importin β:xBB and importin β:αIBB complexes, displaying only the relative positions of the two IBBs. Importin β tryptophan residues (W342, W430, and W864) that bind to IBB residues are shown in green.
Seven FG repeats bind importin β outer surface
We reconstructed the structure of importin β:αIBB bound to the FG-rich nucleoporin Nsp1 at a maximum resolution of 3.21 Å (Supplementary Fig. 2C and Supplementary Table 1). The reconstruction revealed strong peaks of density on the convex surface of importin β (Fig. 3A). We identified five major FG-binding pockets on the convex surface of importin β, formed at the interface between HEAT repeats H4–H6 (pocket 1), H6–H7 (pocket 2), H7–H8 (pocket 3), H9–H10 (pocket 4), and H16–H17 (pocket 5) (Fig. 3B). Pocket 1 binds the PAFSFG motif, positioning the two phenylalanine side chains (FH5 and FH5’) within a bipartite FG-binding pocket between H5 and H6. This pocket is formed by L174, I178, N171, and F217 (Fig. 3C), and represents the previously identified major FG-binding site42. A proline residue N-terminal to the FSFG motif (PH4) contacts the interface between H4 and H5. Pocket 2 is also bipartite, and is filled by an AFSF motif, which includes phenylalanine FH6 and FH6’ (Fig. 3B). In contrast, all other pockets are filled with a small density (Fig. 3A) consistent with a single phenylalanine: FH7 for pocket 3 (Fig. 3B), and FH9 and FH16 for pockets 4 and 5, respectively (Fig. 3C). Like pocket 1, the two smaller pockets 4 and 5 exhibit both hydrophobic and polar residues that interact with the phenyl ring and mainchain atoms (Fig. 3C). For example, FH16 contacts one hydrophobic residue, L700, one uncharged polar residue, Q741, and one negatively charged glutamate, E703. The side chain oxygen of the glutamate forms a hydrogen bond with the glycine main chains and has a nonbonded contact with the FH16 side chain (Fig. 3C). Thus, the chemistry of FG-binding to importin β includes both hydrophobic and polar contacts, suggesting FG-binding pockets are hydrated when importin β is not associated with FG-nups.
A Representative 2D class averages (top) and 3D structure of the importin β:αIBB:FG complex (bottom). Importin β is colored blue, the αIBB is yellow, and FGs are red. The final 3.21 Å cryo-EM density (gray) is overlaid onto the refined model for FG-peptides in chicken-wire style, and around the αIBB as a solid surface. B Schematic diagram of the FG-binding pockets on the outer surface of importin β. FH5 and FH5’ create a bipartite binding site. C Schematic diagram of the importin β residues (colored blue) that surround FGs (colored red) in pocket 1 (left), pocket 4 (center), and pocket 5 (right).
The cryo-EM reconstruction presented here identified all major FG-binding major sites (FH5 and FH6) described crystallographically42 and probed biochemically (FH16)59,60. Additionally, we experimentally validated five of the eight FG sites identified in silico using molecular dynamics simulations41, all of which are located on the convex surface of importin β. Importantly, we found that the conformation of importin β observed in our cryo-EM reconstruction bound to FGs is similar to the importin:αIBB complex (Supplementary Fig. 4A), with an RMSD of approximately 1.9 Å. This indicates that FG-binding does not significantly alter the protein solenoid, at least in the αIBB-bound conformation61.
Ran-GTP closes the importin β solenoid
We elucidated the cryo-EM structure of the importin β:Ran-GTP complex at a resolution of 3.34 Å (Supplementary Fig. 2D and Supplementary Table 1). In this structure (Fig. 4A), importin β is significantly closed around Ran-GTP, making extensive contact with HEATs 1–13. Specifically, Ran-GTP engages at three contact points with the concave surface of importin β, at HEATs 1–4 (contact point a), HEATs 6–8 (contact point b), and HEAT 13 (contact point c) (Fig. 4B). Ala178 is the last residue of Ran-GTP with discernable density in our reconstruction, whereas C-terminal residues 179–216 are disordered. The association between importin β and Ran-GTP comprises 10 salt bridges and 11 hydrogen bonds, burying 2388 Å2 of the importin β concave surface. Perhaps the most striking finding is that the cryo-EM structure of the human importin β:Ran-GTP complex differs dramatically from the crystal structure of the Kap95p:Ran-GTP complex (PDB: 2BKU) with a staggering RMSD of 7.4 Å (Fig. 4C). In the cryo-EM structure, the C-terminus of importin β is completely closed relative to Ran-GTP (Fig. 4A), which is precisely the opposite of what is seen in the crystal structure of Kap95p:Ran-GTP44. The main difference between cryo-EM and crystal structures is that Kap95p:Ran-GTP crystallizes as a tight dimer stabilized by intermolecular dimerization of HEAT repeats 15–19 (Supplementary Fig. 5A, B), which forces the solenoid into an elongated conformation. In contrast, in vitreous ice, the importin β solenoid bound to Ran-GTP is under conformational strain, with N- and C-termini showing a maximum displacement of 19.4 Å compared to 40.4 Å in the Kap95p:Ran-GTP complex. Superimposing the cryo-EM structure of importin β with the Kap95p X-ray model reveals that the latter has open, relaxed, and untwisted C-terminal HEATs 17–19 with a displacement of approximately 25 Å relative to the equivalent HEAT repeats in the human homolog (Fig. 4C). Thus, the previous crystal structure of Kap95p bound to Ran-GTP44 adopts a tertiary structure that is drastically different from that of human importin β.
A Representative 2D class averages (top) and the 3D structure of the importin β:Ran-GTP complex (bottom). Importin β is colored blue, while Ran-GTP is colored magenta. The final 3.34 Å cryo-EM density (gray) for Ran-GTP is overlaid with the refined model. B A schematic diagram illustrates the contacts between importin β and Ran-GTP within a distance of 4.5 Å. C The superimposition of Kap95p:Ran-GTP (PDB: 2BKU) with the importin β:Ran-GTP solved in this study is represented as beads-on-a-string. Each bead represents a HEAT repeat, while Ran-GTP is depicted in ribbon representation. Kap95p is colored yellow, and importin β is colored blue.
RanBP1 intercalates between importin β and Ran-GTP
Importin β and RanBP1 do not bind to each other without Ran, but the three proteins form a complex together53,54, which accelerates Ran-GTP hydrolysis in the presence of importin α33,57.
We assembled importin β with RanQ69L, which is trapped in a GTP-bound state, along with RanBP1, and reconstructed this trimeric complex using cryo-EM SPA at 2.62 Å resolution. (Supplementary Fig. 2E and Supplementary Table 1). Unexpectedly, we found RanBP1 inserts itself at the interface between importin β and Ran-GTP, predominantly binding to the outer surface of Ran-GTP. The GTPase C-terminal moiety (res. 191–216) wraps around the RanBP1 core (Fig. 5A, B), recapitulating the tight embrace observed in the Ran-GTP:RanBP1 crystal structure32. Notably, RanBP1 makes limited contact with importin β in the ternary complex, limited to just one nonbonded interaction between R44 and H520 (Fig. 5B). Nonetheless, the association of RanBP1 with Ran-GTP profoundly influences the importin β solenoid, causing it to relax and extend, adopting a conformation that is even more elongated than when bound to xIBB (Fig. 2A).
A Representative 2D class averages (top) and 3D structure of the importin β:Ran-GTP:RanBP1 complex (bottom). Importin β is colored blue, Ran-GTP is magenta, and RanBP1 is colored orange. The final 2.62 Å cryo-EM density (gray) of Ran-GTP:RanBP1 is overlaid onto the refined model. B Schematic diagram of importin β contacts made with Ran-GTP:RanBP1 within a 4.5 Å distance. C Ribbon diagrams of importin β:Ran-GTP (left) and importin β:Ran-GTP:RanBP1 (right) visualized. Importin β is shown in light gray with HEAT 13 colored blue, while Ran-GTP and RanBP1 are represented as solvent surfaces colored magenta and orange, respectively. D An overlay of HEAT 13 residues 542–595 from the structure of importin β:Ran-GTP (gray) and importin β:Ran-GTP:RanBP1 (blue) reveals a dramatic structural change caused by the association with RanBP1. E The superimposition of importin β in the conformation bound to Ran-GTP:RanBP1 (blue) and Ran-GTP (gray) is shown as beads-on-a-string. Ran-GTP and RanBP1 are not displayed.
RanBP1 alters the association of importin β with Ran-GTP in three significant ways. First, the C-terminal helix of Ran-GTP, disordered in the importin β:Ran-GTP complex (Fig. 4A), becomes structured with RanBP1, disrupting a salt bridge between Ran-GTP R15 and importin β E46 in HEAT 1 (Fig. 5B, contact point a). Second, the binding interface between RanBP1 and Ran-GTP weakens the importin β:Ran-GTP binding interface at contact point c (red dashed line in Fig. 5B), resulting in the loss of two salt bridges and three hydrogen bonds. This is offset by 10 nonbonded interactions between RanBP1 R44 and HEAT 12 H520 (blue dashed lines in Fig. 5B). Third, the importin β:Ran-GTP association at HEAT 13 is partially disrupted due to a dramatic structural rearrangement of HEAT 13 (Fig. 5C). To accommodate both RanBP1 and Ran-GTP, HEAT 13 residues 542–595 bend by 90 degrees in the ternary complex (Fig. 5D), forming intramolecular contacts with the concave surface of importin β.
Thus, RanBP1’s association with importin β:Ran-GTP weakens two of the three bonding regions between the two proteins, specifically, points a and c, while keeping the bonding contacts between HEATs 2 and 8 seemingly unchanged (Fig. 5B). In the presence of RanBP1, the interaction between Ran-GTP and importin β decreases to six salt bridges and seven hydrogen bonds, with a total burial of 2038 Å2 of importin β‘s concave surface. Notably, the entire C-terminal cluster, consisting of HEATs 14–19, swings away from Ran-GTP in the presence of RanBP1, relaxing the solenoid in a way that contrasts significantly with the closed, snail-like conformation observed in the complex with Ran-GTP alone (Fig. 5E).
Importin β has a loose C-terminus in complex with Ran-GDP and RanBP1
Both in vitro53,54,55,56,57,62 and in live cells63, importin β forms a stable complex with Ran-GDP and RanBP1. The function of this complex is poorly understood, and the biochemical determinants responsible for its association are unclear, given that importin β binds Ran-GDP weakly (Kd ~2 μM), and RanBP1 exhibits undetectable binding to Ran-GDP but interacts strongly with Ran-GTP57. To decipher how importin β can associate with both Ran-GDP and RanBP1, we assembled a stoichiometric complex of the three proteins, which we subjected to cryo-EM SPA and used to reconstruct at 3.21 Å resolution (Supplementary Fig. 2F and Supplementary Table 1). Notably, the reconstruction of the trimeric complex shows density only for importin β residues 1–459, which include HEAT repeats 1–10, along with the full Ran-GDP:RanBP1 heterodimer (Fig. 6A, B). In the ternary complex, importin β has a flexible C-terminus, with HEATs 11–19 not visible in the reconstruction due to motion and lack of binding to Ran-GDP:RanBP1. Ran-GDP adopts a Ran-GTP-like conformation that makes nearly identical contacts with Ran-BP1 as seen in the importin β:Ran-GTP:RanBP1 complex (Fig. 5B). A similar GTP-like conformation of Ran-GDP was observed in a crystal structure of the yeast Kap95p:Ran-GDP complex64, suggesting that importin β can stabilize Ran’s conformation regardless of its nucleotide-bound state. In complex with RanBP1, Ran-GDP’s C-terminal moiety (res. 191–216) wraps around the RanBP1 core (Fig. 6A, B), embracing RanBP132. However, the Ran-GDP:RanBP1 heterodimer has only two contact points with importin β, at HEATs 1–4 (Fig. 6B, contact point a) and 8–9 (Fig. 6B, contact point b), but does not associate with HEAT 13 (contact point c in Fig. 5C). Because of this loss of binding, the C-terminus of importin β does not coil around the Ran-GDP:RanBP1 complex as it does with Ran-GTP, but instead remains flexible, likely resembling unliganded importin β45,46, which has proven intractable for cryo-EM single-particle analysis.
A Representative 2D class averages (top) and the 3D structure of the importin β:Ran-GDP:RanBP1 complex (bottom). Importin β is colored blue, Ran-GDP is light magenta, and RanBP1 is orange. The final 3.21 Å cryo-EM density (gray) of Ran-GDP:RanBP1 is overlaid onto the refined model. B Schematic diagram of importin β contacts with Ran-GDP:RanBP1 within a 4.5 Å distance. Notably, only HEAT repeats 1–10 are visible in the reconstruction. C Overlay of Ran-GDP:RanBP1 and Ran-GTP:RanBP1 achieved by superimposing importin β HEAT repeats 1–10 from the two experimentally determined ternary complexes, namely importin βːRan-GTP:RanBP1 (Fig. 5B) and importin βːRan-GDP:RanBP1 (Fig. 6B).
To rationalize the unexpected open structure of importin β in complex with Ran-GDP:RanBP1, we superimposed the Ran-GDP:RanBP1 and Ran-GTP:RanBP1 heterodimers bound to importin β (Fig. 6C). The global RMSD is only ~1.9 Å, suggesting that the overall conformation is highly similar. Subtle changes exist in switch loop I (res. 32–43) and switch loop II (res. 66–84) of Ran that surround the nucleotide. In Ran-GDP:RanBP1, the basic patch region (res. 122–138) is shifted outward by 5 Å relative to the GTP-bound state, toward the position occupied by HEAT 13 in the importin-β Ran-GDP:RanBP1 complex (Fig. 5D). This expansion of the Ran-GDP basic patch is driven by a series of small shifts triggered by the absence of the γ-phosphate in GDP, which causes the nucleotide to move downward between the switch loops, thereby pushing the basic patch outward.
Analysis of importin β domain motion
To rationalize the range of importin β conformational dynamics observed in our cryo-EM reconstructions, we performed a global superimposition of five cryo-EM structures of importin β determined in this study (Fig. 7A). Since all structures were obtained in a frozen-hydrated state, under physiological buffer, and refined at comparable resolution, it is reasonable to assume that the observed changes in importin β structure accurately reflect variations occurring during the nuclear import reaction. We found that importin β solenoid can stretch in response to binding effectors, ranging from approximately 85 to 110 Å, which is about one-quarter of the protein’s length. While the xIBB stabilizes the most extended conformation (around 110 Å), Ran-GTP instead strains the importin β solenoid, resulting in a globular assembly of roughly 85 Å (Fig. 7A). However, different binding effectors not only stretch the importin β solenoid but also induce local conformational changes at the N- and C-termini. The global RMSD among the different states described in this paper is significant, particularly for the Ran-GTP:RanBP1-bound complexes, namely, ~4.4 Å (xIBB vs. αIBB), ~7.3 Å (xIBB vs. Ran-GTP), ~9.3 Å (xIBB vs. Ran-GTP:RanBP1), ~3.6 Å (αIBB vs. Ran-GTP), ~8.2 Å (αIBB vs. Ran-GTP:RanBP1), and 7.2 Å (Ran-GTP vs. RanBP1). Because only the N-terminal 10 HEAT repeats are resolved in the reconstruction, we did not calculate a global RMSD for importin β:Ran-GDP:RanBP1 relative to the other states. The N-terminus of importin β, however, aligns closely with that in the Ran-GTP:RanBP1 complex (RMSD 1.5 Å), with only minor changes distributed anisotropically across different parts of the solenoid. This prompted us to analyze local domain movements using DynDom65.
A An overlay of all the importin β structures solved by cryo-EM in this study, represented as beads-on-a-string. Among the importin β complexes, the xIBB-bound form adopts the longest solenoid, whereas the RanGTP-bound form is the shortest. Color coding: salmon, for importin β:xIBB; gray, for importin β:αIBB:FG; blue, for importin β:Ran-GTP; and orange, for importin β:Ran-GTP:RanBP1. B Rationalization of importin β domain movement based on DynDom analysis. Importin β consists of three clusters of HEAT repeats. Moving clusters I (HEATs 1–4) and III (HEATs 11–19) are depicted in blue, while the fixed cluster II (HEATs 5–10) is shown in gray. Bending helices H5B and H10B are shown in green.
We compared IBBs and found that the xIBB-bound importin β structure is ~10 Å more open than the αIBB structure due to motion in the C-terminal HEATs 11–19, which rotates by about 8 degrees in the xIBB complex relative to αIBB (Fig. 7A). More surprising is the role of Ran-GTP. Instead of inducing global domain motion, such as the opening of the C-terminal arch, as previously reported44, Ran-GTP affects the local conformation of importin β HEAT repeats 1–4 and 11–19, which close toward each other and slightly rotate clockwise (Fig. 7A). As a result of this movement, the maximum length of importin β decreases by ~15 and ~25 Å relative to the αIBB and xIBB, respectively (Fig. 7A), highlighting significant closure of the protein. However, comparing importin β bound to Ran-GTP with the Ran-GTP:RanBP1, DynDom did not detect any domain movement in terms of the rotation of one domain relative to another. Importin β global stretching by approximately 20 Å occurs due to the insertion of RanBP1 between importin β and Ran-GTP, which is consistent with a rigid body movement.
Overall, DynDom analysis of importin β domain motion in response to binding effectors identified three clusters of HEAT repeats. Two moving clusters comprising N-terminal HEATs 1–4 (cluster I) and C-terminal HEATs 11–19 (cluster III) (blue in Fig. 7B), along with a fixed cluster II formed by HEATs 5–10 (gray in Fig. 7B), which is invariant in all our cryo-EM structures. Clusters I and III move relative to cluster II around the helices B of HEAT 5 (bending helix I) and HEAT 10 (bending helix III) (green in Fig. 7B). Ran-GTP binds to importin β, causing a 16-degree clockwise rotation in moving cluster I (H1–H4) and a 10-degree rotation in moving cluster III (H11–H19) while maintaining a fixed configuration in the region H5–H10. These rotations are linked to bending at the N- and C-termini around the helix B of H5 and H10, respectively (Fig. 7B). Because the N- and C-terminal clusters also twist around the two helices, the domain movement of clusters I and III is better described as bending rather than as hinge movement. Notably, importin β cluster III is fully mobile when complexed with Ran-GDP:RanBP1, where the rotation of cluster III relative to cluster II may resemble that of unbound importin β46. The two movements of H1–H4 and H11–H19 must account for the known activities of Ran-GTP, namely, reducing FG-nup binding avidity and displacing cargo from importin β.
Ran-GTP binding to importin β closes FG-binding pockets
Ran-GTP reduces the affinity of importin β for FG-nups12; however, none of the crystal structures determined to date have explained how this occurs. To address this question, we focused on the outer surface of frozen-hydrated importin β, which was solved in complex with αIBB:FG-repeats (Fig. 3A), Ran-GTP (Fig. 4A), Ran-GTP:RanBP1 (Fig. 5A), and Ran-GDP:RanBP1 (Fig. 6A). Guided by the atomic coordinates of FGs bound to importin β:αIBB (Fig. 3A), we measured the volume and solvent-accessible surface area (SASA) of each of the five pockets in importin β using FPocketWeb66,67. We found that the five pockets have volumes ranging from 502 to 154 Å3, displaying characteristics that lie between pocket-like invaginations and cavities (Supplementary Table 2). We then compared the volume and shape of these FG-binding pockets in importin β bound to Ran-GTP, Ran-GTP:RanBP1, and Ran-GDP:RanBP1. Strikingly, we observed a substantial size reduction in cavity volume and SASA in two pockets: pocket 1 (H4–H6) and pocket 4 (H9–H10) (Fig. 8A, B). Notably, the bipartite pocket between H5–H6 displayed a significant decrease in solvent-accessible volume at the first site (H5–H6) (Fig. 8B), diminishing from 502 to 122.0, 303, and 291 Å3, respectively. This site is likely vital, as a single point mutation at I178D abolishes importin β nuclear import in permeabilized cells42. Similarly, the pocket between H9–H10 (pocket 4) that binds FH9 exhibited a noteworthy reduction in solvent-accessible volume, with the FH9 pocket decreasing from 500 to 194, 325 and 346 Å3 (Fig. 8B). Notably, FH9 binds at the bending region identified when Ran-GTP or Ran-GDP, are associated with importin β (green in Fig. 6B). Additionally, pocket 5 located between H16–H17 at the C-terminal end undergoes a remarkable decrease in solvent-accessible volume only in the importin β:Ran-GTP:RanBP1 complex, diminishing from 545 to 290 Å3 (Fig. 8B). This pocket is not visible in the complex with Ran-GDP:RanBP1 that has a disordered C-terminus. Finally, Ran-GTP, Ran-GTP:RanBP1, and Ran-GDP:RanBP1 binding to importin β did not perturb pockets 2 and 3, which are very shallow (Supplementary Fig. 6A, B).
A Importin β is shown as a gray solvent surface, while the bound FG-peptides are depicted as red and gray sticks. FG-binding pockets 1 (top), 4 (middle), and 5 (bottom) are outlined in blue. B Magnified view of the FG-binding pockets 1 (top) and 4 (bottom) shown as a solvent surface from three importin β complexes: bound to αIBB (left), Ran-GTP (center-left), Ran-GTP:RanBP1 (center-right), and Ran-GDP:RanBP1 (right). The FGs bound to the pockets are shown as red sticks. Pockets 1 and 4 notably shrink after binding to Ran-GTP, Ran-GTP:RanBP1, and Ran-GDP:RanBP1. The volume and solvent-accessible surface area (SASA) of each pocket are calculated using FPocketWeb66,67, are indicated for each of the four conformations of importin β.
We also examined the FG-binding pockets of Kap95p identified in the crystal structure with Nup143 (PDB: 5OWU) and compared them to those in the crystal structure with Ran-GTP44 (PDB: 2BKU) to detect changes in volume and SASA. Notably, Nup1 contacts three FG-pockets in Kap95p outer surface (Supplementary Fig. 7A), which are analogous to pockets #1, #2, and #4 in importin β (Fig. 3B). The volume of these pockets was surprisingly reduced in complex with Ran-GTP (Supplementary Fig. 7B). Especially, the pocket #1 (H5–H6) and #3 (H7–H8) display a significant decrease in solvent-accessible volume, which drops from 459 to 228.0 Å3 and 463 to 221 Å3, respectively, when Ran-GTP is bound (Supplementary Fig. 7B). The pocket #2 instead did not have significant changes in solvent-accessible volume and SASA. Therefore, in solution as well as in the crystalline state, Ran-GTP binding to the inner surface of importin β and Kap95p induces an allosteric change on the outer surface that closes the primary FG-binding pockets, located between HEAT repeats H4–H6, H9–H10, and H16–H17 in importin β, and between H5–H6 and H7–H8 in Kap95p.
Ran-GTP, alone or in complex with RanBP1, reduces importin β‘s avidity for FG-nups
We experimentally validated the prediction that importin β bound to RanGTP:RanBP1 exhibits reduced binding affinity for FG-nucleoporins, as expected for the RanGTP-bound state12. To this end, we immobilized GST-tagged Nup358, Nup62, and Nup153 on glutathione beads, representing FG-nups localized to the cytoplasmic face, central channel, and nuclear basket of the NPC, respectively (Fig. 9A–C, lane 2, and Supplementary Fig. 8A–C). In an initial set of pulldown assays, all three FG-nups efficiently captured free importin β as well as importin β in complex with αIBB (Fig. 9A–C, lanes 3–4). Notably, Nup62 bound free importin β slightly more strongly than the importin β:αIBB complex (Fig. 9B, lane 4), consistent with a previous report showing that importin β affinity for Nup62 decreases by up to 40-fold when bound to αIBB37. By contrast, no statistically significant difference was observed between importin β alone and αIBB–IBB to Nup358 or Nup153 (Fig. 9A, C, lane 4). In the presence of Ran-GTP alone or Ran-GTP plus RanBP1, importin β exhibited dramatically reduced binding to all tested FG-nups (Fig. 9A–C, lanes 5–6, see relative quantification). Ran-GTP and Ran-GTP: RanBP1 dissociated ~90–95% of all importin β from GST-Nup62 and -Nup358 (Fig. 9A, B), whereas about 20–25% of importin β remained bound to GST-Nup153 (Fig. 9C). No statistically significant difference was observed between Ran-GTP alone and Ran-GTP-RanBP1, both of which equally displaced importin β from FG-nups (Fig. 9A–C). Therefore, Ran-GTP binding to the concave surface of importin β induces a long-distance conformational change, which reduces the solvent-accessible surface of at least two FG-binding pockets on the outer surface, thereby disrupting FG association in vitro. The ternary importin β:Ran-GTP: RanBP1 complex exhibits a low affinity for FG-nups due to the closure of three FG-binding pockets 1, 4, and 5 (Fig. 8B).
Each panel shows (top) representative cropped Coomassie-stained SDS-PAGE gels from pulldown assays and (bottom) the corresponding quantification obtained by densitometric analysis of bands from four independent experiments (n = 4). A GST-Nup358 (res. 2503–2893), B GST-Nup62, and C GST-Nup153 (res. 946–1472) were coupled to glutathione agarose beads and incubated with equal amounts of purified importin β alone (lane 3), or importin β with αIBB (lane 4), Ran-GTP (lane 5), or Ran-GTP plus RanBP1 (lane 6). The intensity of importin β bands bound to GST-FG-nups was quantified using ImageJ78 across four independent SDS-PAGE experiments (n = 4). Histogram columns represent the ratio of importin β band intensity (black circles) normalized by the intensity of the GST-FG-nup band, with relative standard deviation indicated by error bars. Histograms are presented as mean values +/− standard deviations. A Shows that 45.6 ± 6.9% of importin β was pulled down by GST-Nup358 beads. This value dropped to 36.0 ± 6.6% in the presence of MBP-αIBB, 14.8 ± 4.9% with Ran-GTP, and 7.9 ± 4.2% with Ran-GTP:RanBP1. B Shows that 70.9 ± 11.4% of importin β was pulled down by GST-Nup62, decreasing to 40.4 ± 12.3% with MBP-αIBB, 5.3 ± 0.9% with Ran-GTP, and 4.0 ± 0.9% with Ran-GTP:RanBP1. C Shows that 83.1 ± 11.2% of importin β was pulled down by GST-Nup153 beads, and slightly more, 88.5 ± 13.6%, in the presence of MBP-αIBB. Importin β binding to Nup153 dropped to 23.7 ± 6.1% with Ran-GTP and 17.6 ± 5.1% with Ran-GTP:RanBP1. D–F Show comparable pulldown experiments as in A–C but using Ran-GDP. Specifically, D GST-Nup358 (res. 2503–2893), E GST-Nup62, and F GST-Nup153 (res. 946–1472) coupled to glutathione agarose beads were incubated with equal amounts of purified importin β (lane 3) alone, or importin β together with αIBB (lane 4), Ran-GDP (lane 5) or Ran-GDP plus RanBP1 (lane 6). D Shows that 62.9 ± 8.4% of importin β was pulled down by GST-Nup358 beads; 56.8 ± 4.6% in the presence of MBP-IBB, 55.2 ± 5.6% with Ran-GDP, and 57.2 ± 19.6% with Ran-GDP:RanBP1. E Shows that 107.0 ± 16.7% of importin β was pulled down by GST-Nup62, 78.2 ± 8.2% with MBP-αIBB, 104.7 ± 17.7% with Ran-GDP, and 56.9 ± 11.4% with Ran-GDP + RanBP1. F shows that 88.4 ± 10.8% of importin β was pulled down by GST-Nup153 beads; 83.8 ± 25.3% with MBP-αIBB, 78.1 ± 10.2% with Ran-GDP, and 55.8 ± 14.6% with Ran-GDP + RanBP1. For all pulldown gels A–F, only fractions bound to each FG-nup and recovered on beads are shown. M.W. molecular weight markers. See Supplementary Fig. 8 for the uncropped gels. A significant (p < 0.01) deviation from the importin β:GST-FG-nup ratio was assessed using a two-tailed Student’s t test, assuming unequal variance. In this analysis, *p < 0.05 and ***p < 0.001 are considered statistically significant, while n.s. indicates not significant. Source data, including all SDS-PAGE gels, are provided as a Source data file.
We then repeated the pull-down assay described above using Ran-GDP, both alone and together with RanBP1 (Fig. 9D–F and Supplementary Fig. 8D–F). As expected12, Ran-GDP alone did not significantly alter the association of importin β with FG-nups (Fig. 9D–F, lane 5). Notably, the addition of RanBP1 with Ran-GDP did not affect the binding of importin β to Nup358 (Fig. 9D, lane 6). Still, it produced a very small yet statistically significant reduction in binding to Nup62 (Fig. 9E, lane 6) and Nup153 (Fig. 9F, lane 6). This modest decrease in affinity may be explained by the partial closure of the FG-binding pockets #1 and #4 observed in the structural model displayed in Fig. 8B.
IBB and Ran-GTP bind to partially different regions of importin β
We next investigated the IBB interface for IBB and Ran-GTP and identified three binding regions: HEATs 1–6 (region I), HEATs 7–13 (region II), and HEATs 14–19 (region III) (Fig. 10A). These three regions are similar but not identical to the three moving clusters identified through local domain motion analysis (Fig. 7B). Both αIBB and xIBB domains do not contact region I, form only two salt bridges with region II, and create over ten hydrogen bonds with importin β C-terminal HEATs 14–19, which is part of region III (Supplementary Fig. 9A). In contrast, Ran-GTP primarily binds to importin β through the N-terminal regions I and II (Supplementary Fig. 9B). The interaction with importin β involves five hydrogen bonds in region I and five salt bridges along with six hydrogen bonds in region II. Residues E281 and D288 in region II are the only residues in importin β bound by either IBB or Ran-GTP, both engaging in electrostatic interactions (Supplementary Fig. 9B). Thus, the binding sites for IBBs and Ran-GTP in importin β are mainly distinct. IBB associates primarily with the C-terminal region III (HEATs 7–19), whereas Ran-GTP binds to the N-terminal arch comprising HEATs 1–13, validating previous mutagenesis studies16, with some overlap in HEAT 13. Strikingly, in the presence of Ran-GTP:RanBP1 (Supplementary Fig. 9C), three salt bridges and three hydrogen bonds between Ran-GTP and importin β in region II are disrupted by RanBP1, resulting in an overall opening of HEATs 14–19, potentially creating space to accommodate the IBB.
A Solvent surface representations of importin β bound to αIBB (yellow ribbon, left), Ran-GTP alone (magenta, center), and Ran-GTP:RanBP1 (magenta and orange, right). Semitransparent blue circles labeled I, II, and III highlight three binding regions in importin β. B Native gel electrophoresis shows a sequential addition of Ran-GTP (lane 3) and Ran-GTP:RanBP1 (lane 4) to purified importin β (lane 2) along with a titration of αIBB (lanes 5–12) against a preformed importin β:Ran-GTP:RanBP1 complex. Lane 1 has molecular weight (M.W.) native gel markers. The MBP-αIBB control is in lane 13, although this fusion protein is basic and does not migrate well on a native acrylamide gel. The quantification of the native gel in B is presented in Supplementary Fig. 10A. Source data, including all native gels, are provided as a Source data file.
We then asked whether RanBP1 would facilitate the formation of a larger assembly comprising both αIBB and Ran-GTP. We titrated an equimolar quantity of purified Ran-GTP against 5 μg of importin β and visualized the relative association using native PAGE (Fig. 10B, lanes 2–3). We assembled the importin β:Ran-GTP:RanBP1 complex by adding a stoichiometric amount of RanBP1 (Fig. 10B, lane 4), resulting in a ternary complex that shifted upward compared to importin β:Ran-GTP (Fig. 10B, lane 3). Finally, we titrated a 0–10-fold excess of αIBB fused to the maltose binding protein (MBP-αIBB) against the importin β:Ran-GTP:RanBP1 ternary complex (Fig. 10B, lanes 5–12). Although the MBP-αIBB chimera is basic and does not migrate well on a native gel (Fig. 10B, lane 13), it significantly shifted the migration of the trimeric importin β:Ran-GTP:RanBP1 complex. In the presence of MBP-αIBB, a new band shifted upwards was observed, corresponding to a tetrameric complex formed by importin β:Ran-GTP:RanBP1:MBP-αIBB (Fig. 10B, lanes 7–12). Quantification of native gel bands indicated that about half (e.g., 52.1 ± 12.8%) of the trimeric importin β:Ran-GTP:RanBP1 complex was converted into a tetrameric complex upon addition of a 4-fold molar excess of αIBB (Supplementary Fig. 10A). This tetrameric importin β:Ran-GTP:RanBP1:αIBB complex was not observed when the αIBB was titrated against a preformed importin β:Ran-GTP without RanBP1 (Supplementary Fig. 10B). Therefore, the binding of RanBP1 to importin β and Ran-GTP stabilizes a conformation of the importin β C-terminus that is capable of binding αIBB, which in this study serves as a proxy for the classical import complex (e.g., importin α1:NLS-cargo).
We also repeated the titration in Fig. 10B using Ran-GDP, which, as expected, bound importin β and RanBP1, forming a trimeric importin β:Ran-GDP:RanBP1 complex (Supplementary Fig. 11A, B, lanes 4). However, adding up to a 10-fold molar excess of MBP-αIBB to a pre-formed importin β:Ran-GDP:RanBP1 complex did not cause a further upward shift on the native gel (Supplementary Fig. 11A, B, lanes 5–12), indicating that the importin β:Ran-GDP:RanBP1 complex cannot bind cargo. Overall, this data suggests that the importin β:Ran-GDP:RanBP1 complex is not directly involved in mediating the nuclear import of NLS cargos through the NPC.
Discussion
This study aimed to elucidate two key mechanisms of nuclear import that have remained unclear despite decades of extensive research. On one hand, we sought to determine how Ran-GTP dissociates importin β from high-affinity FG-binding sites lining the NPC, allowing import complexes to move through the selective barrier instead of clogging the NPC16. On the other hand, we asked how Ran-GTP disassembles the import complex, releasing the cargo into the nucleus and preventing futile rounds of stochastic, Ran-GTP-induced dissociation and reassociation within the NPC68. To address these questions, we determined six unbiased structures of importin β bound to various effectors encountered in a typical import reaction and complemented the structural data with biochemical measurements of importin β activity. Overall, we shed light on three aspects of importin β biology relevant to answering the two questions above.
First, we present a comprehensive description of the importin β solenoid conformational landscape through a comparative analysis of the structure of importin β obtained in vitreous ice. Our analysis suggests that importin β resembles an intrinsically disordered protein, in which the nineteen HEAT repeats maintain their secondary structures but lack a tertiary structure, as the solenoid fluctuates among various conformations, sampling different states that are possibly separated by low-energy barriers49. We were unable to reconstruct the structure of unliganded importin-β because its high conformational heterogeneity precludes single-particle analysis. In the absence of binding partners, importin β likely interconverts between multiple, substantially different conformations, preventing current algorithms from aligning particles and yielding stable 2D class averages. Stabilizing the solenoid with binding partners enabled us to reconstruct five conformations of importin β bound to its cellular partners. We show evidence that distinct importin β conformers bind to and are stabilized by αIBB, xIBB, Ran-GTP, Ran-GTP:RanBP1, and Ran-GDP:RanBP1, leading to structurally different solenoids. We suggest that the states reconstructed via cryo-EM in this study represent the major conformations that human importin β adopts in solution. However, we cannot rule out the possibility that importin β solenoid also exists in other conformations not immediately visible in this study due to our inability to determine particle orientation and position computationally. We identified three HEAT repeat clusters in importin β, two of which (HEATs 1–4 and 11–19) move relative to an invariant core (HEATs 5–10) by bending around helices H5B and H10B (Fig. 7B). This parameterization of importin β conformational dynamics could not have been achieved using X-ray crystallography. All previously determined crystal structures of importin β exhibit significant fluctuations primarily due to the type of crystallization agents, pH, and lattice forces, unlike cryo-EM structures obtained under quasi-physiological conditions. We provide experimental visualization of five FG-binding pockets on the outer surface of importin β. At the current resolution (~3.2 Å), we did not observe significant conformational changes in the importin β solenoid upon FG-binding, suggesting that FG-repeats function like molecular Velcro69, decorating the outside of importin β without inducing substantial fluctuations in the import receptor.
Second, we analyzed how Ran-GTP binds to importin β, regulating its activities essential for nuclear transport. The GTPase stabilizes a highly restrained conformation of the importin β solenoid that wraps around Ran-GTP by making three contact points (Fig. 4B). This tripartite recognition forces importin β to adopt a globular conformation that alters the protein curvature, closing two of the five FG-binding pockets on its outer surface. Consequently, the association of Ran-GTP with the concave surface of importin β triggers an allosteric change on the convex surface, affecting both the volume and the solvent-accessible surface of the FG pockets. By doing this, Ran-GTP reduces the affinity for FG-nups lining the NPC but does not eliminate it entirely, enabling the import complex to continue moving through the NPC and bypassing the permeability barrier. Remarkably, Ran-GTP does not open the C-terminus of importin β, as previously reported. The cryo-EM structure of human importin β bound to Ran-GTP presented in this study significantly differs from a previous crystal structure of yeast Kap95p bound to Ran-GTP44, demonstrating that Kap95p is structurally unrepresentative of human importin β. As a corollary, the proposed model for Ran-GTP-mediated cargo displacement based on a change in the helicoidal pitch of the importin β C-terminus is unlikely. Our cryo-EM reconstructions suggest that Ran-GTP and IBBs partially overlap in their binding to importin β region II (Fig. 10A), where the two binding effectors compete for a limited number of residues. The IBB domain primarily associates with the C-terminal region III (HEATs 7–19), while Ran-GTP binds to HEATs 1–13 at three contact points (Fig. 4B). We envision a sequential mechanism for cargo release that begins with the association of Ran-GTP with importin β at contact point a (Fig. 4B). This interaction causes a 16-degree clockwise rotation of the moving cluster I (HEATs 1–4) relative to the fixed cluster II (HEATs 5–10) and a 10-degree counterclockwise rotation of cluster III (H11–H19) (Fig. 7B). The globularization of importin β around Ran-GTP reduced the N-to-C distance by ~15 Å relative to the αIBB-bound conformation, breaking the bonds between αIBB and HEATs 13–19 in region III (Supplementary Figs. 4A and 9A). Thus, Ran-GTP allosterically displaces the import complex, triggering long-range conformational changes in the importin β solenoid that displace the IBB domain.
Third, our study provides a molecular explanation of the role of RanBP1 in the import reaction. RanBP1 promotes the translocation of the classical nuclear import complex through the NPC55,56, a function also conserved in yeast70. Earlier research found that RanBP1 increases the affinity of both Ran-GDP and Ran-GTP for importin β to the same level, suggesting that the nucleotide-bound state of Ran does not affect the stability of the importin β:Ran:RanBP1 complex53. We validated this idea by visualizing frozen-hydrated structures of importin β bound to Ran-GTP:RanBP1 and Ran-GDP:RanBP1, which revealed that the Ran:RanBP1 heterodimer maintains the same conformation in both GDP- and GTP-bound states when bound to importin β. This is because importin β N-terminus and RanBP1 can force Ran-GDP to adopt a GTP-like conformation, as previously described for Kap95p64. However, we found that Ran’s nucleotide state influences importin β‘s structural flexibility and, consequently, its biochemical activities. The importin β:Ran-GTP:RanBP1 complex displays a stretched conformation of the importin β solenoid, which biochemically has reduced affinity for FG-nups but retains the ability to associate with the IBB-domain. In contrast, the importin β:Ran-GDP:RanBP1 complex lacks density for the entire C-terminus of importin β (HEATs 11–19), which are too flexible and dynamic to be aligned during SPA. This results in an importin β that is unable to associate with the cargo and retains avidity for FG-nups.
How do these structural and biochemical findings help us understand the role of Ran and RanBP1 in the nuclear translocation of importin β? In the literature, RanBP1 is primarily implicated in the disassembly of the import complex33, serving as the key player that removes importin β’s inhibition of GTP hydrolysis, directly promoting the dissociation of the import complex at Nup358, where RanGAP1 is cross-linked through sumoylation. Interestingly, this process also requires the IBB domain of importin α in vitro33. Our study finds that in the presence of Ran-GTP and RanBP1, importin β retains a low affinity for FG-nups and can hold onto the cargo due to the extended position of the C-terminal HEATs 11–19, suggesting the possible formation of a larger assembly consisting of the importin complex (importin β:α1:NLS-cargo) associated with Ran-GTP:RanBP1. Thus, we hypothesize that a transient assembly of the import complex bound to both Ran-GTP and RanBP1 may exhibit sufficient stability and decreased avidity for FG-nups, allowing movement across the NPC. This would enable the import complex to traverse the permeability barrier and pass through the central channel as a complete entity, circumventing unproductive cycles of cargo release and reassociation. This transient assembly would possess the properties of importin β bound to Ran-GTP in terms of affinity for FGs, yet it would remain attached to the NLS-cargo, similar to an import complex formed in the cytoplasm. The final disassembly of the import complex would take place at the NPC basket, where import complexes encounter a higher concentration of nuclear Ran-GTP. This model agrees with a recent report21 that RanGTP facilitates the release of importin β from the NPC. In experiments with permeabilized cells, NPCs decorated with fluorescently tagged importin β were liberated from NPC-bound importin β upon washing with a Ran mix containing Ran-GDP, NTF2, GTP, RanBP1, and RanGAP, which caused a significant loss of importin β. This mix summarizes both the Ran-GTP:RanBP1 and Ran-GDP:RanBP1 complexes described in this paper, which we nonetheless place at different stages of the import reaction.
Unlike the importin β:RanGTP:RanBP1 complex, which we implicate in translocating the import complex, the role of the importin β:RanGDP:RanBP1 complex in nuclear import is more difficult to decipher. The evidence presented here—that this trimeric species cannot associate with αIBB yet retains binding to FG-nups—suggests it does not contribute to the translocation of import complexes through the NPC. As suggested by Plafker and Macara63, this ternary complex of importin β may represent an intermediate that helps terminate/resolve transport complexes in the cytosol, at the cytoplasmic surface of the NPC, where Nup358 is located8, and feed Ran back into the Ran-GTP/Ran-GDP cycle. The trimeric importin β:Ran-GDP:RanBP1 complex could form a pool of importin β (alongside Ran-GDP) at the RanBP1-like domains of RanBP2, which function similarly to RanBP1 studied here57. However, we cannot rule out that this ternary species represents an off-pathway complex formed by importin β in the cytoplasm: a nonproductive side complex that can accumulate in vitro and in cells when Ran-GDP and RanBP1 are available but does not contribute to productive import cycles.
In summary, we have elucidated the complete conformational dynamics of importin β and provided insight into the function of RanGTP. This GTPase has evolved to regulate the affinity of β-karyopherins for FG-nups by allosterically altering their curvature and, as demonstrated here, modulating the volume and solvent-accessible surface of FG-binding pockets. Our data support a model in which Ran-GTP and RanBP1 facilitate cargo passage through the NPC and its controlled release at the NPC basket, while Ran-GDP, in conjunction with RanBP1, can bind importin β and potentially sequester it in a manner that does not promote transport. We propose that the structural principles elucidated in this work for importin β are conserved and applicable to members of the β-karyopherin family involved in nuclear import and export. It is possible that β-karyopherins and Ran-GTP coevolved under selective pressure to achieve efficient and rapid nuclear import.
Methods
Cloning, expression, and purification of recombinant proteins
The genes encoding the importin β:α1 heterodimer were cloned into the vector pACYCDuet-1 (EMD Millipore) with importin α1 fused to an N-terminal 6× His-tag and importin β as untagged71. The proteins were expressed in BL21-DE3 Escherichia coli (E. coli) cells. Protein expression was induced with 500 µM IPTG for 3 h at 30 oC, and the heterodimer was purified to homogeneity using metal affinity chromatography and size-exclusion chromatography (SEC) using a Superdex 200 16/600 preparative column (Cytiva)71,72. Untagged importin β was separated from the importin β:α1 complex bound to the nickel agarose beads (GenScript) using a Separation Buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 250 mM MgCl2, and 3 mM 2-mercaptoethanol). Importin β was further purified by SEC on a Superdex 200 26/600 preparative column (Cytiva) equilibrated in β SEC Buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1 mM PMSF, and 3 mM 2-mercaptoethanol).
The importin α1 IBB domain (res 1–65) C-terminally fused to a 6xHis Maltose Binding Protein (MBP-αIBB) was cloned by PCR into an engineered pET28a vector73 that also contains the MBP gene downstream of the 6xHis tag, using the following forward (5′-CAGCCGGATCCATGTCCACCAACGAGAATGCTA-3′) and reverse (5′- CAGCCAAGCTTCTACTGCAGCGGAGAAGTAGCATC-3′) oligos. MBP-αIBB was expressed in BL21-DE3 E. coli cells. Protein expression was induced with 500 µM IPTG for 3 h at 30 °C. Cells were resuspended in IBB Lysis Buffer (20 mM Tris-HCl, pH 8.0, 200 mM NaCl, 1 mM PMSF, 3 mM 2-mercaptoethanol, 1 mM EDTA, and 0.1% Triton X-100, DNase, and RNase) and after sonication, the soluble fraction was bound to amylose resin (New England Biolabs). The protein was washed using a lysis buffer and then eluted with a lysis buffer with 15 mM maltose. MBP-αIBB was further purified by SEC using a Superdex 200 16/600 preparative column (Cytiva) equilibrated with IBB SEC Buffer (20 mM Tris-HCl, pH 8.0, 200 mM NaCl).
Ran-GTP was expressed and purified using a Ran-Q69L mutant that remains in a GTP-bound conformation. Ran-Q69L was cloned into a pET28a-PPase construct with an N-terminal 6xHis tag and expressed in BL21-DE3 E. coli strain supplemented with kanamycin37. Protein was expressed overnight at 18 °C with 500 µM IPTG. Cells were resuspended in Ran Lysis Buffer (50 mM KPO4, pH 7.0, 250 mM NaCl, 2 mM MgCl2, 1 mM PMSF, and 3 mM 2-mercaptoethanol) and, after sonication, the soluble fraction was bound to low-density nickel agarose beads (GoldBio). The protein was washed using a lysis buffer containing 10 mM imidazole and then eluted with a lysis buffer with Ran Elution Buffer (50 mM KPO4, pH 7.0, 250 mM NaCl, 2 mM MgCl2, 1 mM PMSF, 3 mM 2-mercaptoethanol, and 150 mM imidazole). Ran-GTP was further purified by SEC using a Superdex 200 16/600 preparative column (Cytiva) equilibrated with Ran SEC Buffer (50 mM KPO4, pH 7.0, 250 mM NaCl, 2 mM MgCl2, 1 mM PMSF, and 3 mM 2-mercaptoethanol). The importin β:Ran-GTP complex was formed by incubating importin β and Ran-GTP at 1–4 molar ratio at 4 °C for 30 min.
To produce Ran-GDP, we expressed the full-length human Ran cloned into a pET28a-PPase vector with an N-terminal 6xHis tag in BL21-DE3 E. coli40. Protein was expressed overnight at 18 °C with 500 µM IPTG. Cells were resuspended in Ran-GDP Lysis Buffer (20 mM Tris-HCl, pH 8.0, 250 mM NaCl, 2 mM MgCl2, 1 mM PMSF, and 3 mM 2-mercaptoethanol) and, after sonication, the soluble fraction was bound to low-density nickel agarose beads (GoldBio). The protein was washed using a lysis buffer containing 10 mM imidazole and then eluted with a lysis buffer with Ran-GDP Elution Buffer (20 mM Tris-HCl, pH 8.0, 250 mM NaCl, 2 mM MgCl2, 1 mM PMSF, and 3 mM 2-mercaptoethanol, 150 mM imidazole). The Ran protein was incubated with 2 mM GDP on ice for 30 min before SEC. Ran-GDP was further purified by SEC using a Superdex 200 16/600 preparative column (Cytiva) equilibrated with Ran-GDP SEC Buffer (20 mM Tris-HCl, pH 8.0, 250 mM NaCl, 2 mM MgCl2, 1 mM PMSF, and 3 mM 2-mercaptoethanol). The importin β:Ran-GDP:RanBP1 complex was formed by pre-incubating importin β and Ran-GDP and RanBP1 at 4 °C for 30 min and purified by SEC using a Superose 6 increase 10/300 GL column (Cytiva) equilibrated with Ran-GDP SEC Buffer.
RanBP1 was cloned into a pET28a-PPase vector with an N-terminal 6xHis tag and expressed in BL21-DE3 E. coli cells37. Protein expression was induced with 500 µM IPTG for 3 h at 30 °C. Cells were resuspended in a Low Salt Lysis Buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1 mM PMSF, and 3 mM 2-mercaptoethanol), lysed by sonication, and the soluble fraction isolated by centrifugation and incubated with low-density nickel agarose beads (GoldBio). The protein was washed using a low salt lysis buffer containing 5–20 mM imidazole, 50 mL for each wash, then eluted with Elution Buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1 mM PMSF, and 3 mM 2-mercaptoethanol, 150 mM imidazole). RanBP1 was further purified by SEC using a Superdex 200 16/600 preparative column (Cytiva) equilibrated with PBS plus 0.1 mM PMSF.
The FxFG-rich fragment (residues 497–608) of yeast Nsp174 was cloned into expression vector pMW172 with an N-terminal 6xHis tag75. The protein was expressed for 3 h at 30 °C after adding 400 µM IPTG to cells with an optical density at 600 nm of ~0.6. The Nsp1 fragment was purified in a High Salt Lysis Buffer (20 mM Tris-HCl, pH 8.0, 600 mM NaCl, 1 mM PMSF, and 3 mM 2-mercaptoethanol and 5% glycerol) and bound to low-density nickel agarose beads (GoldBio). The protein was washed with a high salt buffer containing 5/10/20 mM imidazole, 10 mL for each wash, then eluted with Elution Buffer (20 mM Tris-HCl pH 8.0, 150 mM NaCl, 1 mM PMSF, and 3 mM 2-mercaptoethanol, 150 mM imidazole) supplemented with 1.25% glycerol. The importin β:α1:FG complex was formed by adding a 2-fold molar excess of Nsp1 (497–608) to purified importin α1/β and used immediately for vitrification.
GST-tagged constructs of Nup358 (res. 2503–2893)76, full-length Nup6276, and Nup153 (res. 946–1472)76 were cloned into the vector pGEX-4T-1 (Cytiva) and expressed in BL21-DE3 E. coli cells. Protein expression was induced with 500 µM IPTG for 3 h at 30 oC. Cells were resuspended in a Medium Salt Lysis Buffer (20 mM Tris-HCl pH 8.0, 300 mM NaCl, 0.2% Triton X-100, 1 mM PMSF, and 3 mM 2-mercaptoethanol) plus protease inhibitor, and after sonication, the soluble fraction was bound to glutathione beads (GenScript). The protein was washed using a lysis buffer containing 0.2% Triton X and then buffer-exchanged to Protein Binding Buffer (20 mM HEPES pH 7.0, 150 mM KOAc, 2 mM Mg(OAc)2, 1 mM dithiothreitol, and 0.1% Tween) plus protease inhibitors. The immobilized protein was used immediately to avoid degradation. A peptide comprising XRIP1α residues 6–47 (xIBB) was synthesized by GenScript.
Pulldown assays and native gel electrophoresis
GST-tagged fragments of Nup358 (res. 2503–2893) and Nup153 (res. 946–1472), as well as full-length p62 (res. 1–522) were immobilized on glutathione beads (GoldBio) and resuspended in Pulldown Binding Buffer (20 mM HEPES pH 7.0, 150 mM KOAc, 2 mm Mg(OAc)2, 1 mM dithiothreitol, and 0.1% Tween 20). Pulldown assays were carried out as described37,77. Briefly, batches of 50 μL of glutathione beads coupled with GST-Nups were incubated with either 3 μM importin β alone, or (3 μM importin β + 3 μM MBP-αIBB), or (3 μM importin β + 3 μM Ran-GTP), or (3 μM importin β + 3 μM Ran-GTP + 3 μM RanBP1). All reactions were incubated for 1 h at 4 oC. Unbound fractions were washed away, and beads were washed three times with 150 μL of Pulldown Wash Buffer (20 mM HEPES pH 7.0, 150 mM KOAc, 2 mM Mg(OAc)2, 1 mm dithiothreitol, and 0.1% Tween 20). The entire pool of 50 μL of washed beads that contains the bound fractions was resuspended in 25 μL of 4× SDS-sample buffer and 5 μL was analyzed on an 8–16% Tris-Glycine mini protein gel (Invitrogen, XP08162BOX). SDS-PAGE gels were run across four independent experiments (n = 4), and the average ratio of importin β band intensity normalized to each GST-FG-nup was determined through densitometry using ImageJ78. We employed a two-tailed Student’s t test, assuming unequal variances, to determine statistical significance between the means of the control and experimental pulldowns. The alpha parameter had to be outside the 99th percentile for statistical significance (p < 0.01). Error bars indicate the standard deviation of the mean.
Native Gel electrophoresis was performed using NativePAGETM Bis-Tris Mini Protein Gels, 3–12% (Invitrogen, BN1001). Briefly, 5 μg of importin β or importin β complexes were incubated on ice for one hour, with increasing molar excesses (typically between 0.5 and 10×) of Ran-GTP (or Ran-GDP), RanBP1, and/or αIBB. After 1 h, 3.75 μL of 4× Loading buffer (Invitrogen, BN2003) was added to the sample to reach a final volume of 15 μL. Samples were then loaded onto a 3–12 % Bis-Tris native gel, run the gel with Bis-Tris non-denaturing running buffer (Invitrogen, BN2007) at 150 V for 90 min, at 4°C. The gel was stained with Coomassie Brilliant Blue-G-250 and destained overnight in 50% (v/v) methanol in water with 10% (v/v) acetic acid.
Vitrification and data collection
For all importin β complexes studied in this paper, typically 2.0 µl of importin β pre-bound to αIBB, xIBB, αIBB:Nsp1, Ran-GTP, Ran-GTP:RanBP1, or Ran-GDP:RanBP1 at 2 mg/ml final concentration were applied to a 300-mesh copper Quantifoil R 1.2/1.3 holey carbon grid (EMS) previously glow-discharged negatively for 60 s at 15 mA using an easiGlow (PELCO). Only for the importin β:αIBB:Nsp1 complex (referred to herein as importin β:αIBB:FG) and importin β:Ran-GTP:RanBP1, we used 200-mesh UltraAUfoil R 1.2/1.3 grid (EMS) instead of standard carbon grids. All grids were blotted for 5 s at blot force 5 and frozen in liquid ethane using Vitrobot Mark IV (FEI). All micrographs were pre-screened in-house on a 200 kV Glacios2 equipped with a Falcon 4i detector at the UAB Cryo-EM Facility. EPU (v3.10)79 software was used for data screening and collection using the accurate positioning mode. All high-resolution datasets were collected at Titan Krios microscopes available at three federal facilities equipped with either a K3 direct electron detector camera (e.g., NCEF and NCCAT) or a Falcon 4i (e.g., S2C2 and NCCAT). The software used for data collection was Serial EM (v4.2.10)80 for Gatan K3 and EPU (v3.10)79 for Falcon 4i. 9497 micrographs were collected for importin β:xIBB in super-resolution mode plus energy filter at 20 eV with an image pixel size of 0.436 Å at ×105,000 magnification, a nominal total dose of 50 e/Å2, 40 frames, and defocus range −1.0 to −2.5 μm. 27,987 micrographs were collected for importin β:αIBB in super-resolution mode plus energy filter at 20 eV with an image pixel size of 0.371 Å at ×165,000 magnification, a nominal total dose of 50 e/Å2, 40 frames, and defocus range −0.8 to −2.5 mm. 18,016 micrographs were collected for importin β:αIBB:FG plus energy filter at 20 eV with an image pixel size of 0.959 Å at ×130,000 magnification, a nominal total dose of 50 e/Å2, 40 frames, and defocus range −0.8 to −2.5 μm. 13,468 micrographs were collected for importin β:Ran-GTP in super-resolution mode with an energy filter at 20 eV, using an image pixel size of 0.436 Å at ×105,000 magnification, a nominal total dose of 50 e/Å2, 40 frames, and a defocus range of −1.0 to −2.5 μm. 11,100 micrographs were collected for importin β:Ran-GTP:RanBP1 plus energy filter at 20 eV with an image pixel size of 0.371 Å at ×165,000 magnification, a nominal total dose of 50 e/Å2, 40 frames, and defocus range −0.8 to −2.5 μm. 22,555 micrographs were collected for importin β:Ran-GDP:RanBP1 plus energy filter at 20 eV with an image pixel size of 0.371 Å at ×165,000 magnification, a nominal total dose of 50 e/Å2, 40 frames, and defocus range −0.6 to −3.7 μm. All collection parameters are in Supplementary Table 1.
Cryo-EM single particle analysis (SPA)
Micrographs were subjected to patched motion-correction using MotionCor2 (v1.6.4)81 and patched contrast transfer function (CTF) estimation using CTFFIND4 (v4)82. All subsequent steps of SPA were carried out using cryoSPARC version 4.2.183. Particle picking was done using one round of template picker and at least three rounds of Topaz training and extraction84, followed by extracting particles from micrographs. In total, we picked 4.3 million for importin β:xIBB, 11.9 million for importin β:αIBB, 5.8 million particles for importin β:αIBB:FG, 6.5 million for importin β:Ran-GTP, 9.6 million for importin β:Ran-GTP:RanBP1, and 1.81 million for importin β:Ran-GDP:RanBP1. The initial maps were reconstructed by using a combination of selected 2D classes and ab initio reconstruction. In total, 293,677, 157,234, 593,354, 1,400,279, 721,883, and 241,237 particles were used to calculate 3D densities for importin β:xIBB, importin β:αIBB, importin β:αIBB:FG, importin β:Ran-GTP, importin β:Ran-GTP:RanBP1, and importin β:Ran-GDP:RanBP1, respectively. The 3D maps were further refined using heterogeneous 3D classification, homogeneous refinement with CTF and defocus refinement options, and post-processing on a 10-GPU cluster. The final densities were sharpened using phenix.auto_sharpen85. A flowchart of cryo-EM SPA methods used in this paper is displayed in Supplementary Fig. 1.
Model building, refinement, and structure analysis
All atomic models were built using Coot (v0.9.8.7)86 and ChimeraX (v1.9)87 followed by several rounds of rigid-body, real-space, and B-factor refinement using phenix.real_space_refinement in Phenix (v1.21)88. All final models were validated using MolProbity89 (Supplementary Table 1). All images of models and cryo-EM maps were generated using ChimeraX (v1.9)87 and PyMol (v2.3)90. RMSD between superimposed PDBs, structural comparison, and domain motion analysis were done using the DynDom server65 and SuperPose 1.091. FPocketWeb66,67 was used to identify and characterize binding pockets within importin β. Binding interfaces were analyzed using PISA92 and PDBsum93.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic coordinates generated in this study have been deposited in the Protein Data Bank database under accession codes 9N86 (Importin β:IBB complex), 9N87 (Importin β:xIBB complex), 9BFC (Importin β:xIBB:FG complex), 9BAW (Importin β:Ran-GTP complex), 9N85 (Importin β:Ran-GTP:RanBP1 complex), and 9YB5 (Importin β:Ran-GDP:RanBP1 complex). The cryo-EM density maps generated in this study have been deposited in the Electron Microscopy Data Bank under accession codes EMD-49116 (Importin β:IBB reconstruction), EMD-49117 (Importin β:xIBB reconstruction), EMD-44492 (Importin β:xIBB:FG reconstruction), EMD-44412 (Importin β:Ran-GTP reconstruction), EMD-49114. (Importin β:Ran-GTP:RanBP1 reconstruction), and EMD-72749 (Importin β:Ran-GDP:RanBP1 reconstruction). Previously published PDB codes referred to in this paper include: 2BKU (Kap95:Ran-GTP complex), 5OWU (Kap95:Nup1 complex),1QGK (Importin β:IBB complex). The minimum datasets (e.g., motion corrected micrographs) necessary to interpret and verify all six cryo-EM reconstructions presented in this paper can be downloaded from the University of Alabama at Birmingham (UAB) Cheaha Supercomputer (https://www.uab.edu/it/home/research-computing/cheaha) upon request to the authors. The source data underlying Figs. 9A–F, 10B; Supplementary Fig. 10 and 11 are provided as a Source data file.
References
Bednenko, J., Cingolani, G. & Gerace, L. Nucleocytoplasmic transport: navigating the channel. Traffic 4, 127–135 (2003).
Macara, I. G. Transport into and out of the nucleus. Microbiol. Mol. Biol. Rev. 65, 570–594 (2001).
Stewart, M. Molecular mechanism of the nuclear protein import cycle. Nat. Rev. Mol. Cell Biol. 8, 195–208 (2007).
Wing, C. E., Fung, H. Y. J. & Chook, Y. M. Karyopherin-mediated nucleocytoplasmic transport. Nat. Rev. Mol. Cell Biol. 23, 307–328 (2022).
Strom, A. C. & Weis, K. Importin-beta-like nuclear transport receptors. Genome Biol. 2, REVIEWS3008 (2001).
Pumroy, R. A. & Cingolani, G. Diversification of importin-alpha isoforms in cellular trafficking and disease states. Biochem. J. 466, 13–28 (2015).
Aramburu, I. V. & Lemke, E. A. Floppy but not sloppy: interaction mechanism of FG-nucleoporins and nuclear transport receptors. Semin. Cell Dev. Biol. 68, 34–41 (2017).
Lin, D. H. & Hoelz, A. The structure of the nuclear pore complex (an update). Annu. Rev. Biochem. 88, 725–783 (2019).
Frey, S., Richter, R. P. & Gorlich, D. FG-rich repeats of nuclear pore proteins form a three-dimensional meshwork with hydrogel-like properties. Science 314, 815–817 (2006).
Kalab, P., Weis, K. & Heald, R. Visualization of a Ran-GTP gradient in interphase and mitotic Xenopus egg extracts. Science 295, 2452–2456 (2002).
Izaurralde, E., Kutay, U., von Kobbe, C., Mattaj, I. W. & Gorlich, D. The asymmetric distribution of the constituents of the Ran system is essential for transport into and out of the nucleus. EMBO J. 16, 6535–6547 (1997).
Gorlich, D., Pante, N., Kutay, U., Aebi, U. & Bischoff, F. R. Identification of different roles for RanGDP and RanGTP in nuclear protein import. EMBO J. 15, 5584–5594 (1996).
Moore, M. S. & Blobel, G. The GTP-binding protein Ran/TC4 is required for protein import into the nucleus. Nature 365, 661–663 (1993).
Gorlich, D., Prehn, S., Laskey, R. A. & Hartmann, E. Isolation of a protein that is essential for the first step of nuclear protein import. Cell 79, 767–778 (1994).
Moore, M. S. & Blobel, G. Purification of a Ran-interacting protein that is required for protein import into the nucleus. Proc. Natl. Acad. Sci. USA 91, 10212–10216 (1994).
Kutay, U., Izaurralde, E., Bischoff, F. R., Mattaj, I. W. & Gorlich, D. Dominant-negative mutants of importin-beta block multiple pathways of import and export through the nuclear pore complex. EMBO J. 16, 1153–1163 (1997).
Sun, C., Yang, W., Tu, L. C. & Musser, S. M. Single-molecule measurements of importin alpha/cargo complex dissociation at the nuclear pore. Proc. Natl. Acad. Sci. USA 105, 8613–8618 (2008).
Kose, S., Imamoto, N., Tachibana, T., Shimamoto, T. & Yoneda, Y. Ran-unassisted nuclear migration of a 97-kD component of nuclear pore-targeting complex. J. Cell Biol. 139, 841–849 (1997).
Rabut, G., Doye, V. & Ellenberg, J. Mapping the dynamic organization of the nuclear pore complex inside single living cells. Nat. Cell Biol. 6, 1114–1121 (2004).
Lowe, A. R. et al. Importin-beta modulates the permeability of the nuclear pore complex in a Ran-dependent manner. Elife 4, e04052 (2015).
Sau, A. et al. Overlapping nuclear import and export paths unveiled by two-colour MINFLUX. Nature 640, 821–827 (2025).
Lowe, A. R. et al. Selectivity mechanism of the nuclear pore complex characterized by single cargo tracking. Nature 467, 600–603 (2010).
Lyman, S. K., Guan, T., Bednenko, J., Wodrich, H. & Gerace, L. Influence of cargo size on Ran and energy requirements for nuclear protein import. J. Cell Biol. 159, 55–67 (2002).
Snow, C. J., Dar, A., Dutta, A., Kehlenbach, R. H. & Paschal, B. M. Defective nuclear import of Tpr in progeria reflects the Ran sensitivity of large cargo transport. J. Cell Biol. 201, 541–557 (2013).
Plafker, K. & Macara, I. G. Facilitated nucleocytoplasmic shuttling of the Ran binding protein RanBP1. Mol. Cell Biol. 20, 3510–3521 (2000).
Zolotukhin, A. S. & Felber, B. K. Mutations in the nuclear export signal of human Ran-binding protein RanBP1 block the Rev-mediated posttranscriptional regulation of human immunodeficiency virus type 1. J. Biol. Chem. 272, 11356–11360 (1997).
Schlenstedt, G. et al. Yrb4p, a yeast ran-GTP-binding protein involved in import of ribosomal protein L25 into the nucleus. EMBO J. 16, 6237–6249 (1997).
Pasquinelli, A. E., Powers, M. A., Lund, E., Forbes, D. & Dahlberg, J. E. Inhibition of mRNA export in vertebrate cells by nuclear export signal conjugates. Proc. Natl. Acad. Sci. USA 94, 14394–14399 (1997).
Wu, J., Matunis, M. J., Kraemer, D., Blobel, G. & Coutavas, E. Nup358, a cytoplasmically exposed nucleoporin with peptide repeats, Ran-GTP binding sites, zinc fingers, a cyclophilin A homologous domain, and a leucine-rich region. J. Biol. Chem. 270, 14209–14213 (1995).
Yokoyama, N. et al. A giant nucleopore protein that binds Ran/TC4. Nature 376, 184–188 (1995).
Yaseen, N. R. & Blobel, G. Two distinct classes of Ran-binding sites on the nucleoporin Nup-358. Proc. Natl. Acad. Sci. USA 96, 5516–5521 (1999).
Vetter, I. R., Nowak, C., Nishimoto, T., Kuhlmann, J. & Wittinghofer, A. Structure of a Ran-binding domain complexed with Ran bound to a GTP analogue: implications for nuclear transport. Nature 398, 39–46 (1999).
Bischoff, F. R. & Gorlich, D. RanBP1 is crucial for the release of RanGTP from importin beta-related nuclear transport factors. FEBS Lett. 419, 249–254 (1997).
Floer, M., Blobel, G. & Rexach, M. Disassembly of RanGTP-karyopherin beta complex, an intermediate in nuclear protein import. J. Biol. Chem. 272, 19538–19546 (1997).
Cingolani, G., Petosa, C., Weis, K. & Muller, C. W. Structure of importin-beta bound to the IBB domain of importin-alpha. Nature 399, 221–229 (1999).
Vetter, I. R., Arndt, A., Kutay, U., Gorlich, D. & Wittinghofer, A. Structural view of the Ran-Importin beta interaction at 2.3 A resolution. Cell 97, 635–646 (1999).
Lott, K., Bhardwaj, A., Mitrousis, G., Pante, N. & Cingolani, G. The importin beta binding domain modulates the avidity of importin beta for the nuclear pore complex. J. Biol. Chem. 285, 13769–13780 (2010).
Cingolani, G., Bednenko, J., Gillespie, M. T. & Gerace, L. Molecular basis for the recognition of a nonclassical nuclear localization signal by importin beta. Mol. Cell 10, 1345–1353 (2002).
Lee, S. J. et al. The structure of importin-beta bound to SREBP-2: nuclear import of a transcription factor. Science 302, 1571–1575 (2003).
Mitrousis, G., Olia, A. S., Walker-Kopp, N. & Cingolani, G. Molecular basis for the recognition of snurportin 1 by importin beta. J. Biol. Chem. 283, 7877–7884 (2008).
Isgro, T. A. & Schulten, K. Binding dynamics of isolated nucleoporin repeat regions to importin-beta. Structure 13, 1869–1879 (2005).
Bayliss, R., Littlewood, T. & Stewart, M. Structural basis for the interaction between FxFG nucleoporin repeats and importin-beta in nuclear trafficking. Cell 102, 99–108 (2000).
Liu, S. M. & Stewart, M. Structural basis for the high-affinity binding of nucleoporin Nup1p to the Saccharomyces cerevisiae importin-beta homologue, Kap95p. J. Mol. Biol. 349, 515–525 (2005).
Lee, S. J., Matsuura, Y., Liu, S. M. & Stewart, M. Structural basis for nuclear import complex dissociation by RanGTP. Nature 435, 693–696 (2005).
Zachariae, U. & Grubmuller, H. Importin-beta: structural and dynamic determinants of a molecular spring. Structure 16, 906–915 (2008).
Forwood, J. K. et al. Quantitative structural analysis of importin-beta flexibility: paradigm for solenoid protein structures. Structure 18, 1171–1183 (2010).
Kappel, C., Zachariae, U., Dolker, N. & Grubmuller, H. An unusual hydrophobic core confers extreme flexibility to HEAT repeat proteins. Biophys. J. 99, 1596–1603 (2010).
Halder, K. et al. MD simulations and FRET reveal an environment-sensitive conformational plasticity of importin-b. Biophys. J. 109, 277–286 (2015).
Bhardwaj, A. & Cingolani, G. Conformational selection in the recognition of the snurportin importin beta binding domain by importin beta. Biochemistry 49, 5042–5047 (2010).
Cingolani, G., Lashuel, H. A., Gerace, L. & Muller, C. W. Nuclear import factors importin alpha and importin beta undergo mutually induced conformational changes upon association. FEBS Lett. 484, 291–298 (2000).
Tauchert, M. J. et al. Impact of the crystallization condition on importin-beta conformation. Acta Crystallogr. D Struct. Biol. 72, 705–717 (2016).
Jullien, D., Gorlich, D., Laemmli, U. K. & Adachi, Y. Nuclear import of RPA in Xenopus egg extracts requires a novel protein XRIPalpha but not importin alpha. EMBO J. 18, 4348–4358 (1999).
Chi, N. C., Adam, E. J. & Adam, S. A. Different binding domains for Ran-GTP and Ran-GDP/RanBP1 on nuclear import factor p97. J. Biol. Chem. 272, 6818–6822 (1997).
Lounsbury, K. M. & Macara, I. G. Ran-binding protein 1 (RanBP1) forms a ternary complex with Ran and karyopherin beta and reduces Ran GTPase-activating protein (RanGAP) inhibition by karyopherin beta. J. Biol. Chem. 272, 551–555 (1997).
Chi, N. C., Adam, E. J., Visser, G. D. & Adam, S. A. RanBP1 stabilizes the interaction of Ran with p97 nuclear protein import. J. Cell Biol. 135, 559–569 (1996).
Lounsbury, K. M., Richards, S. A., Perlungher, R. R. & Macara, I. G. Ran binding domains promote the interaction of Ran with p97/beta-karyopherin, linking the docking and translocation steps of nuclear import. J. Biol. Chem. 271, 2357–2360 (1996).
Villa Braslavsky, C. I., Nowak, C., Gorlich, D., Wittinghofer, A. & Kuhlmann, J. Different structural and kinetic requirements for the interaction of Ran with the Ran-binding domains from RanBP2 and importin-beta. Biochemistry 39, 11629–11639 (2000).
Koerner, C., Guan, T., Gerace, L. & Cingolani, G. Synergy of silent and hot spot mutations in importin beta reveals a dynamic mechanism for recognition of a nuclear localization signal. J. Biol. Chem. 278, 16216–16221 (2003).
Bednenko, J., Cingolani, G. & Gerace, L. Importin beta contains a COOH-terminal nucleoporin binding region important for nuclear transport. J. Cell Biol. 162, 391–401 (2003).
Otsuka, S., Iwasaka, S., Yoneda, Y., Takeyasu, K. & Yoshimura, S. H. Individual binding pockets of importin-beta for FG-nucleoporins have different binding properties and different sensitivities to RanGTP. Proc. Natl. Acad. Sci. USA 105, 16101–16106 (2008).
Lott, K. & Cingolani, G. The importin beta binding domain as a master regulator of nucleocytoplasmic transport. Biochim. Biophys. Acta 1813, 1578–1592 (2011).
Pu, R. T. & Dasso, M. The balance of RanBP1 and RCC1 is critical for nuclear assembly and nuclear transport. Mol. Biol. Cell 8, 1955–1970 (1997).
Plafker, K. & Macara, I. G. Fluorescence resonance energy transfer biosensors that detect Ran conformational changes and a Ran x GDP-importin-beta -RanBP1 complex in vitro and in intact cells. J. Biol. Chem. 277, 30121–30127 (2002).
Forwood, J. K. et al. Kap95p binding induces the switch loops of RanGDP to adopt the GTP-bound conformation: implications for nuclear import complex assembly dynamics. J. Mol. Biol. 383, 772–782 (2008).
Girdlestone, C. & Hayward, S. The DynDom3D webserver for the analysis of domain movements in multimeric proteins. J. Comput. Biol. 23, 21–26 (2016).
Kochnev, Y. & Durrant, J. D. FPocketWeb: protein pocket hunting in a web browser. J. Cheminform. 14, 58 (2022).
Le Guilloux, V., Schmidtke, P. & Tuffery, P. Fpocket: an open source platform for ligand pocket detection. BMC Bioinform. 10, 168 (2009).
Rexach, M. & Blobel, G. Protein import into nuclei: association and dissociation reactions involving transport substrate, transport factors, and nucleoporins. Cell 83, 683–692 (1995).
Onischenko, E. et al. Natively unfolded FG repeats stabilize the structure of the nuclear pore complex. Cell 171, 904–917.e19 (2017).
Schlenstedt, G., Wong, D. H., Koepp, D. M. & Silver, P. A. Mutants in a yeast Ran binding protein are defective in nuclear transport. EMBO J. 14, 5367–5378 (1995).
Doll, S. G. et al. Recognition of the TDP-43 nuclear localization signal by importin α1/β. Cell Rep. 39, 111007 (2022).
Yang, R. et al. Structural basis for nuclear import of hepatitis B virus (HBV) nucleocapsid core. Sci. Adv. 10, eadi7606 (2024).
Pumroy, R. A., Nardozzi, J. D., Hart, D. J., Root, M. J. & Cingolani, G. Nucleoporin Nup50 stabilizes closed conformation of armadillo repeat 10 in importin alpha5. J. Biol. Chem. 287, 2022–2031 (2012).
Nehrbass, U. et al. NSP1: a yeast nuclear envelope protein localized at the nuclear pores exerts its essential function by its carboxy-terminal domain. Cell 61, 979–989 (1990).
Clarkson, W. D., Kent, H. M. & Stewart, M. Separate binding sites on nuclear transport factor 2 (NTF2) for GDP-Ran and the phenylalanine-rich repeat regions of nucleoporins p62 and Nsp1p. J. Mol. Biol. 263, 517–524 (1996).
Ben-Efraim, I. & Gerace, L. Gradient of increasing affinity of importin beta for nucleoporins along the pathway of nuclear import. J. Cell Biol. 152, 411–417 (2001).
Pumroy, R. A., Ke, S., Hart, D. J., Zachariae, U. & Cingolani, G. Molecular determinants for nuclear import of influenza A PB2 by importin alpha isoforms 3 and 7. Structure 23, 374–384 (2015).
Abramoff, M. D., Magelhaes, P. J. & Ram, S. J. Image processing with ImageJ. Biophotonics Int. 11, 36–42 (2004).
Thermo Fisher Scientific. EPU (version 3.10). Thermo Fisher Scientific, Waltham, MA, USA (2024).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36-51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T., Kelley, K., Noble, A. J. & Berger, B. Topaz-Denoise: general deep denoising models for cryoEM and cryoET. Nat. Commun. 11, 5208 (2020).
Terwilliger, T. C., Sobolev, O. V., Afonine, P. V. & Adams, P. D. Automated map sharpening by maximization of detail and connectivity. Acta Crystallogr. D Struct. Biol. 74, 545–559 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Schrödinger, LLC. The PyMOL Molecular Graphics System, Version 2.3 (Schrödinger, LLC, New York, NY, 2020).
Maiti, R., Van Domselaar, G. H., Zhang, H. & Wishart, D. S. SuperPose: a simple server for sophisticated structural superposition. Nucleic Acids Res. 32, W590–W594 (2004).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Laskowski, R. A. PDBsum new things. Nucleic Acids Res. 37, D355–D359 (2009).
Acknowledgements
This work was supported by NIH grants R01 GM100888 and R35 GM140733 to G.C. Electron microscopy was carried out in the UAB cryo-EM Facility (RRID: SCR_025450), supported by the Institutional Research Core Program and O’Neal Comprehensive Cancer Center (NIH grant P30 CA013148), with additional funding from NIH grant S10 OD024978. We thank Dr. James Kizziah for assisting with the in-house data collection. We also thank three federal cryo-EM facilities for access to 300 kV Krios microscopes and assistance with data collection: the National Cancer Institute’s National cryo-EM Facility at the Frederick National Laboratory for Cancer Research (contract 75N91019D00024); the National Center for CryoEM Access and Training (NCCAT) and the Simons Electron Microscopy Center (supported by the NIH U24 GM129539, the Simons Foundation SF349247, and NY State Assembly); Stanford-SLAC cryo-EM Center (S2C2), supported by NIH U24 GM129541.
Author information
Authors and Affiliations
Contributions
Y-H.K. and G.C. conceived the study. Y-H.K., F.L., S.S.S., and R.K.L. performed biochemical experiments. Y-H.K., F.L., S.S.S., J.L., and C-F.D.H. performed cryo-EM studies. Y-H.K., F.L., S.S.S., and G.C. built and refined structural models. Y-H.K. and G.C. wrote the manuscript with feedback from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kuo-Chiang Hsia and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ko, YH., Li, F., Suinn, S.S. et al. Ran modulates allosteric crosstalk between importin β surfaces. Nat Commun 16, 11425 (2025). https://doi.org/10.1038/s41467-025-66255-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66255-0