Abstract
Skeletal muscle is a major organ for maintaining whole-body energy balance, yet how it adapts its transcriptional and metabolic programs to environmental cues remains unclear. Here, we report that histone mono-methyltransferase mixed lineage leukemia 4 (MLL4), a key enhancer regulator, directs muscle metabolic adaptation and systemic metabolism through AMPK signaling. Nutrient availability modulates MLL4 expression, and skeletal muscle-specific ablation of MLL4 in male mice protects against diet-induced obesity and improves glucose homeostasis despite reduced exercise endurance. These effects arise from enhanced fuel catabolism caused by marked activation of AMPK in MLL4-depleted muscles. Mechanistically, MLL4 cooperates with myocyte enhancer factor 2 to induce AMP-metabolizing enzymes cytosolic 5’-nucleotidase 1A and AMP-deaminase 3, which suppress AMPK activity. Pharmacologic inhibition of AMP-metabolizing pathway by Pentostatin activates muscle AMPK, confers resistance to obesity and improves metabolic health. These findings identify an enhancer regulator limiting AMPK-mediated muscle fuel catabolism, offering a potential strategy for treating obesity-related disorders.
Similar content being viewed by others
Introduction
Caloric intake outpaces caloric expenditure, causing obesity, a metabolic disorder that drives an array of metabolic pathologies1. As the largest organ and primary site of fuel substrate utilization (glucose and fatty acids) in the human body, skeletal muscle plays a central role in the control of whole-body metabolic homeostasis2,3,4,5. Skeletal muscle exhibits remarkable plasticity and undergoes extensive metabolic remodeling in response to various physiological and pathological challenges3,4,5. Numerous studies in animal models, and in humans, have supported that derangements in muscle fuel metabolism driven by chronic caloric excess is a major contributor to the onset of obesity and related metabolic disorders5,6,7,8. Conversely, significant evidence suggests that enhanced skeletal muscle fuel catabolism by exercise reduces the incidence of metabolic diseases4,5,9. Delineation of the molecular regulatory mechanisms involved in the adaptive and maladaptive remodeling of skeletal muscle fuel metabolism could yield therapeutic targets aimed at the prevention or treatment of the untoward effects of chronic caloric excess.
Skeletal muscle adapts to changes in nutrient availability and physiological demands by fine-tuning gene expression patterns and metabolism to maximize metabolic efficiency. However, the molecular mechanisms that enable and direct muscle transcriptional and metabolic responses to environmental cues are not well understood. Prior research has shown that various transcription factors, including the nuclear receptors PPARs and ERRs, as well as MEF2, in conjunction with coregulators such as PGC-1s, NCoR1, and HDACs, play key roles in the regulation of metabolic gene expression in skeletal muscle under physiological conditions10,11,12,13,14,15,16,17,18. Another metabolic regulator with strong impact on skeletal muscle transcriptional and metabolic responses is the AMP-activated protein kinase (AMPK), a serine/threonine kinase that senses cellular AMP/ATP ratios19,20,21. The activation of AMPK by AMP binding turns on glucose and fatty acids catabolic pathways in muscle to generate ATP, while simultaneously turning off muscle fuel anabolic pathways that consume ATP. As the key step for gene activation and the primary site of the gene-environment interactions, epigenetic regulation of enhancer activation is increasingly recognized as a key component in gene transcription adaptation to physiological and pathological stimuli22,23. Active enhancers are characterized by distinct histone modifications, including mono-methylation of histone H3 lysine 4 (H3K4me1) and acetylation of histone H3 lysine 27 (H3K27ac)22,24. The role of epigenetic regulation in enhancer activation and how it orchestrates the remodeling of skeletal muscle fuel metabolism, whether adaptive or maladaptive, remain to be fully understood.
In mammals, Mixed-lineage leukemia 4 (MLL4/KMT2D) functions as a key histone H3 lysine 4 (H3K4) mono-methyltransferase necessary for enhancer activation25,26,27,28,29. MLL4 has been reported to interact with lineage-determining transcription factors to establish H3K4me1 and H3K27ac signatures on enhancer regions, leading to enhancer activation and consequently gene transcription25,26,30,31. Loss of MLL4 activity prevents enhancer activation, leading to significant defects in gene expression and cell differentiation in multiple cell types25,26,30. Recent studies have highlighted the diverse, context-specific roles of the MLL4 protein in various biological processes, such as embryonic development, metabolic regulation, and cancer progression25,32,33,34,35. Additionally, mutations in MLL4 have been linked to pathological conditions, including Kabuki syndrome, congenital heart defects, and multiple forms of cancer, emphasizing the critical role of MLL4-dependent enhancer activation in human diseases35,36,37. While we recently found that MLL4 is required to activate slow type I muscle fiber gene enhancers to ensure running endurance33, it has yet to be unraveled whether this MLL4-dependent enhancer activation is implicated in regulating skeletal muscle nutrient handling that controls systemic metabolism.
In this study, we explored the potential of MLL4 in controlling the fuel metabolism adaptation in skeletal muscle. We found that muscle MLL4 expression is regulated in response to nutrient availability and, that depletion of MLL4 specifically in skeletal muscle, protects mice from HFD-induced obesity and improves systemic glucose homeostasis despite reducing exercise endurance. Mechanistically, the metabolic actions of MLL4 resulted from enhanced muscle fuel catabolism, ensuing from the marked activation of AMPK in MLL4-depleted muscles. Further, we uncovered a mechanism for the regulation of muscle AMPK activation mediated by the concerted action of a pair of AMP metabolizing enzymes NT5C1A and AMPD3 that are activated by the MLL4-MEF2 transcriptional complex in skeletal muscle. Importantly, pharmacologic inhibition of the AMP metabolizing pathway by Pentostatin potently activates muscle cell AMPK, protects mice against diet-induced obesity and ameliorates obesity-related metabolic disorders. Together, our findings establish MLL4 as a key physiological regulator of muscle fuel metabolism central to systemic metabolic homeostasis, and strongly suggest that modulations of the unique MLL4 downstream effectors in skeletal muscle are potential therapeutic approaches to treat metabolic diseases.
Results
Muscle MLL4 is regulated in response to nutrient availability
To investigate whether MLL4 is implicated in regulating skeletal muscle nutrient handling, we examined MLL4 regulation in response to nutritional cues. Mice were fasted for 6 or 12 h during the light phase and then re-fed during the dark phase for 6 or 12 h. Analyses of MLL4 expression revealed that its protein amounts were significantly reduced after 6 and 12 h of fasting compared to ad libitum-fed controls (Fig. 1a and Supplementary Fig. 1a), and were restored upon refeeding (Fig. 1a and Supplementary Fig. 1a). Furthermore, MLL4 levels progressively declined with prolonged fasting (16 and 24 h), despite all samples being collected at the same circadian time (Supplementary Fig. 1b, c). These findings indicate that nutrient deprivation is associated with reduction of muscle MLL4. We then tested whether nutrient excess also modulates MLL4 protein levels in the muscles. This was indeed the case, as muscle MLL4 protein levels increased after 1 week high-fat diet (HFD) feeding (Fig. 1b). However, Mll4 mRNA levels remained unchanged under the same conditions (Supplementary Fig. 1d), indicating that the increase in MLL4 protein is likely regulated through post-transcriptional mechanisms. Together, these data demonstrate that muscle MLL4 levels are regulated in response to nutrient availability.
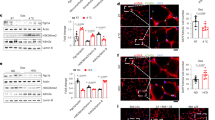
a Western blot of gastrocnemius (GC) muscle from WT mice under light-fasting/dark-refeeding. n = 4–6 mice per group. b Western blot and quantification of MLL4 protein in GC muscle from WT mice under chow diet (CD) or high-fat diet (HFD). n = 6 mice per group. P value: 0.0015. c, d Body weight (c) and weight gain (d). c CD-WT, n = 8; CD-MKO, n = 9; HFD-WT, n = 10; HFD-MKO, n = 6, P value: 0.0154, 0.0015, <0.0001, <0.0001, <0.0001, <0.0001. d CD-WT, n = 4; CD-MKO, n = 6; HFD-WT, n = 10; HFD-MKO, n = 6, P value: 0.0243, 0.0028, 0.001, 0.0007, 0.0007. e Schematic of treadmill endurance exercise test protocol. f, g Running time (f) and distance (g). CD-WT, n = 7; CD-MKO, n = 8; HFD-WT, n = 8; HFD-MKO, n = 8. P value: 0.0387, 0.0027, 0.0387, 0.0027. h Pictures of HFD-fed mice for 12 Weeks. i, j Oxygen consumption (VO2, i) and energy expenditure (EE, j) during the dark phase in HFD-fed mice. WT, n = 15; MKO, n = 11. j ANCOVA-predicted metabolic rate (MR) at 40 g body mass. WT, n = 15; MKO, n = 11. k Fasting glucose, WT, n = 10; MKO, n = 6 and insulin levels, WT, n = 9; MKO, n = 6. P value: 0.0087, 0.0347. l HOMA-IR of HFD-fed mice. WT, n = 9; MKO, n = 6. P value: 0.0279. m Glucose tolerance test (GTT) and area under the curve (AUC) for GTT. CD-WT, n = 7; CD-MKO, n = 7; HFD-WT, n = 10; HFD-MKO, n = 6. P value: 0.0135, 0.0037, 0.0356. n Insulin tolerance test (ITT) and AUC for ITT. CD-WT, n = 7; CD-MKO, n = 6; HFD-WT, n = 10; HFD-MKO, n = 6. P value: 0.0398, 0.0011, 0.0001, <0.0001, 0.0034. o Western blot and quantification of pAKT (S473, T308) in GC muscle after insulin stimulation. Representative Western blots are shown. n = 2 independent experiments. p Oil Red O staining of livers. Scale bar, 50 μm. n = 3–8 mice per group. q Liver triglyceride levels. CD-WT, n = 3; CD-MKO, n = 4; HFD-WT, n = 7; HFD-MKO, n = 6. P value: 0.0031. Data are mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 vs. corresponding WT by two-tailed Student’s t-test (b, f, g, k, l, q) or two-way ANOVA (c, d, m, n) with Fisher’s LSD. Source data are provided as a Source data file.
Muscle-specific deletion of MLL4 protects against high-fat diet-induced obesity and improves systemic glucose homeostasis despite reducing exercise endurance
We have previously generated skeletal muscle-specific Mll4-knockout (Mll4SETf/fHSA-Cre) (referred to as Mll4SET MKO) by crossing Mll4SETf/f mice with the human skeletal actin promoter-driven Cre mice (HSA-Cre)33. Mll4SET MKO mice did not exhibit an overt metabolic phenotype compared to wild-type (WT) littermates on normal chow diet (CD) (Supplementary Fig. 1e–g). We followed a cohort of Mll4SET MKO mice that show similar survival as respective WT littermates at least up to 1.5 years of age. To determine the importance of MLL4 in skeletal muscle nutrient handling, Mll4SET MKO and WT littermates were placed on HFD (60% kcal from fat). Mll4SET MKO mice showed no difference in body weight and energy expenditure when they were fed CD (Fig. 1c, d and Supplementary Fig. 1e–g). However, these mice were resistant to HFD-induced obesity, exhibiting much lower weight gain following HFD feeding despite consuming similar quantities of food (Fig. 1c, d and Supplementary Fig. 1h). These results indicate that Mll4SET MKO mice had a decreased efficiency in converting food into body mass. Interestingly, muscle-specific MLL4 deletion drives the development of exercise intolerance in the context of a “lean” phenotype (Fig. 1e–h). Similar to normally fed Mll4SET MKO mice33, HFD-fed Mll4SET MKO mice still run significantly shorter time and distance (~50%) as determined by a motorized treadmill test compared to WT littermates (Fig. 1f, g). Gene expression analysis reveals that MLL4 deficiency alters muscle regulatory programs involved in neuromuscular junction signaling and fiber-type specification (Supplementary Fig. 1i, j). MHC1 immunofluorescence staining confirmed a marked reduction in the number of type I fibers in muscles from HFD-fed Mll4SET MKO mice (Supplementary Fig. 1k), whereas the proportion of type 2b fibers remained unchanged (Supplementary Fig. 1l). Gene expression analysis of myosin heavy chain (MyHC) isoforms revealed a trend toward increased Myh1 expression, suggesting a possible shift toward type 2x fibers (Supplementary Fig. 1m). We also performed a maximal sprint capacity test and found that Mll4SET MKO mice achieved comparable peak running speeds to WT controls, indicating preserved high-intensity exercise performance (Supplementary Fig. 2a). Furthermore, following an acute treadmill exercise, while Mll4SET MKO muscles exhibited a reduction in mitochondrial respiratory capacity compared to WT controls, serum lactate concentrations were comparable between WT and Mll4SET MKO mice (Supplementary Fig. 2b, c). Together, these results demonstrate a uncoupling between HFD-induced obesity and exercise endurance by MLL4 ablation in skeletal muscle.
While Mll4SET MKO mice exhibited comparable fat mass and lean mass to WT controls under CD (Supplementary Fig. 2d), HFD-fed Mll4SET MKO mice had less fat mass (~25%) compared with HFD-fed WT controls (Supplementary Fig. 2e). Additionally, a modest but significant decrease in lean mass was also observed (Supplementary Fig. 2e). To explain the resistance to HFD-induced obesity, we performed indirect calorimetric analysis in HFD-fed mice using the Comprehensive Lab Animal Monitoring System (CLAMS). While no difference in locomotor activity was found (Supplementary Fig. 2f), Mll4SET MKO mice fed HFD have a significant increase in oxygen consumption, and energy expenditure during dark periods in comparison with WT controls (Fig. 1i, j). These data suggest that the observed reduction in adiposity resulted from an increase in systemic energy expenditure caused by MLL4 deficiency in skeletal muscle. We further determined glucose homeostasis in WT and Mll4SET MKO mice following HFD feeding. Both fasting glucose and insulin levels, as well as the homeostasis model assessment of insulin resistance (HOMA-IR), were significantly lower in Mll4SET MKO mice relative to WT controls (Fig. 1k, l), suggesting that Mll4SET MKO mice have an improved sensitivity to insulin. Indeed, glucose and insulin tolerance tests demonstrated that HFD-fed Mll4SET MKO mice were more tolerant to glucose challenge and more sensitive to insulin stimulation than HFD-fed WT controls (Fig. 1m, n). Moreover, insulin action, as indicated by insulin-stimulated AKT phosphorylation, in skeletal muscle, liver and adipose tissue was all enhanced in HFD-fed Mll4SET MKO mice (Fig. 1o and Supplementary Fig. 2g). Together, these results demonstrate that muscle MLL4 deficiency protects mice from HFD-induced insulin resistance despite reducing exercise endurance.
Muscle MLL4 deficiency prevents HFD-induced hepatic steatosis
Evidence is emerging that skeletal muscle metabolic remodeling could be a modifier of hepatic steatosis4,38,39. Gross anatomical analysis of livers revealed markedly prevented hepatic steatosis in HFD-fed Mll4SET MKO mice compared to WT controls (Supplementary Fig. 3a). Hematoxylin and eosin (H&E) and Oil red O staining of histological sections provided further evidence of reduced liver lipid accumulation in HFD-fed Mll4SET MKO mice (Fig. 1p and Supplementary Fig. 3b). Biochemical measurements confirmed a pronounced (~50%) decrease in TG levels in HFD-fed Mll4SET MKO livers (Fig. 1q). The reduced hepatic lipid and TG accumulation in Mll4SET MKO mice on HFD suggested that MLL4 acts within skeletal muscle to modulate hepatic lipid handling. To gain further insight into the liver metabolic changes in Mll4SET MKO mice, we performed RNA-seq analysis on mRNA isolated from liver of the Mll4SET MKO mice and littermate controls (Supplementary Fig. 3c–f). Gene ontology (GO) analysis of downregulated genes revealed significant enrichment in lipid metabolic process, inflammatory response as well as collagen fibril organization (Supplementary Fig. 3e, f). Conversely, pathways of oxidation-reduction and epoxygenase P450 were significantly enriched in the upregulated gene set (Supplementary Fig. 3e, f). Gene expression validation studies demonstrated that the expression of the genes related to lipid uptake and de novo lipogenesis was reduced in HFD-fed Mll4SET MKO liver (Supplementary Fig. 3g, h), concomitant with an increased in the expression of the oxidation-reduction genes (Supplementary Fig. 3g). We also sought to determine whether major thermogenic pathways are affected in brown adipose tissue (BAT) from Mll4SET MKO mice. Consistent with the muscle-specific activity of HSA-Cre, Mll4 mRNA expression in BAT was unchanged in Mll4SET MKO mice compared to WT littermate controls (Supplementary Fig. 3i). We found that the expression of the genes involved in both UCP1-dependent and -independent thermogenesis were not difference in BAT from WT control and Mll4SET MKO mice (Supplementary Fig. 3j), suggesting that fuel catabolism is likely not upregulated in BAT of Mll4SET MKO mice. Together, these findings suggest that MLL4 deletion in skeletal muscle leads to protection from HFD-induced hepatic steatosis.
Loss of MLL4 promotes fuel catabolism in skeletal muscle
On the basis of the above data, we reasoned that loss of MLL4 might induce favorable metabolic remodeling within skeletal muscle. No significant differences were observed in either muscle triglyceride (TG) or glycogen levels between WT and Mll4SET MKO mice (Supplementary Fig. 4a, b). We also quantified mitochondrial DNA (mtDNA) and found no significant difference between WT and Mll4SET MKO muscles (Supplementary Fig. 4c). We next conducted high profiling metabolomic analysis on skeletal muscles from Mll4SET MKO mice. A large panel of organic acids (Krebs cycle intermediates), amino acids, acyl-carnitines and FFAs were measured in skeletal muscles from Mll4SET MKO mice and littermate controls under both CD and HFD conditions. More than 200~ metabolites were measured (Fig. 2, and Supplementary Fig. 4). These differential metabolites were classified into four different clusters (Fig. 2a and Supplementary Fig. 4d). The K-means clustering analysis (K = 4) was performed on the differential metabolites, grouping them into Cluster I (30 metabolites), Cluster II (23 metabolites), Cluster III (13 metabolites), and Cluster IV (11 metabolites) (Fig. 2a and Supplementary Fig. 4d). Cluster I and II showed prominently upregulated metabolites by MLL4 deficiency (Fig. 2a). While the metabolic differences between WT and Mll4SET MKO mice in Cluster I were more pronounced than under CD vs. HFD conditions (Fig. 2a), Cluster II metabolites showed significantly greater divergence in Mll4SET MKO mice upon HFD feeding (Supplementary Fig. 4d). Metabolite set enrichment analysis (MSEA) of MLL4-regulated metabolites in cluster I and II revealed significant enrichment in the Warburg effect and citric acid cycle (TCA) pathway as well as metabolism of amino acids (Supplementary Fig. 4e). We also observed an enhanced mitochondrial respiration in Mll4SET MKO muscles after HFD feeding (Supplementary Fig. 4f). Moreover, multiple fatty acids and FFA acyl-carnitines, which are products of mitochondrial fatty acid oxidative flux, increased in MLL4-deficient muscles compared to WT controls (Fig. 2b and Supplementary Fig. 4g–i), a phenotype that is more common in mice under CD conditions, suggesting that lipid overload masked the suppression effects of MLL4 on fatty acid catabolism. Specifically, MLL4-deficient muscles showed elevated levels of TCA cycle intermediates (citric acid, fumaric acid and malic acid) and acyl-carnitine species (C14:0, C14:1, C16:0, C16:1, C18:0 and C18:1) in comparison with WT muscles (Fig. 2c, d). To further explore lipid metabolic pathways, we analyzed the expression of genes involved in de novo lipogenesis, lipolysis, and fatty acid oxidation. De novo lipogenesis genes (e.g., Fasn, Scd1 and Pparg) were downregulated in Mll4SET MKO muscle, particularly under HFD (Supplementary Fig. 4j). Lipolytic genes (e.g., Lipe/HSL and Lpl) remained unchanged, while Pnpla2/ATGL showed modest upregulation and Pnpla7 was decreased (Supplementary Fig. 4k). Notably, there was an induction of fatty acid β-oxidation genes, including Cpt2, Acadm, Acadl, Acadvl, Hadhb, and Acaa2 (Supplementary Fig. 4l). Together, these data suggest that muscle MLL4 ablation promotes muscle fuel catabolism.
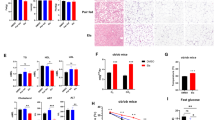
a, b Standardized heatmap showing unsupervised clustering of organic acids, amino acid (a) and acyl-carnitines (b) metabolite content in skeletal muscles from Mll4SET MKO mice and littermate controls under both CD and HFD conditions. n = 6 mice per group. c, d Quantitative data for Krebs cycle intermediates (c) and acyl-carnitines (d) metabolite content in skeletal muscles from indicated mice. CD-WT, n = 6; CD-MKO, n = 6; HFD-WT, n = 6; HFD-MKO, n = 6 mice per group. c P value: 0.0338, 0.0407, 0.0216. d P value: 0.0412, 0.026, 0.0126, 0.0078, 0.0238. Internal standards (decanoylcarnitine-d3, palmitoylcarnitine-d3, d3-FFA C16:0, and d3-FFA C18:0 for LC-MS; methionine sulfone and camphorsulfonic acid for CE-MS) were employed for analysis. Acquired signal intensity (peak area) of metabolites were corrected to an internal standard then normalized to the unit weight of each tissue specimen, relative abundance normalized to tissue weight ×105 for CE-MS and relative abundance normalized to tissue weight ×103 for LC-MS. e Diagram showing the metabolism of glucose and FFAs, the chief fuels for muscle cells. Red and green labels represent up- and down-regulation induced by MLL4 deficiency, respectively. f Schematic illustration of isotopic [13C]-glucose tracing experiments in C2C12 myotubes transduced with lentiviruses expressing scramble (shCtrl) or MLL4 shRNA (shMLL4). g, h Kinetic accumulation of uniformly labeled 3-phosphoglycerate (3-PG) (m + 3) and pyruvate (m + 3), citrate (m + 2 and m + 4), a-ketoglutarate (m + 2 and m + 4) and malate (m + 2) in MLL4 knock-down myotubes at 10 and 40 min after the addition of 25 mM 13C-glucose into the medium. n = 5 per group. P value: 0.0322, 0.0006, <0.0001, 0.0217, 0.0205, 0.0339, 0.0149, 0.0255. All data are shown as the mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 vs. corresponding WT or shCtrl controls determined by two-tailed unpaired Student’s t-test. Source data are provided as a Source data file.
To further investigate the above findings in a cell-autonomous setting, we performed comparative metabolome analysis in C2C12 cells transduced with lentiviruses expressing scramble or MLL4 shRNA. Consistent with the observations in vivo, numerous metabolites were altered in MLL4 knock down (KD) myotubes (Supplementary Fig. 5a–c). MSEA analysis of these upregulated metabolites confirmed significantly enriched in the citric acid cycle and pyruvate metabolism (Supplementary Fig. 5d). Indeed, the metabolites of glycolysis and the TCA cycle were most prominently upregulated (Supplementary Fig. 5e, f), and most of the glycolytic and TCA cycle intermediates were significantly increased (from ~1.5 to 4-fold) in MLL4 KD myotubes compared to control cells (Fig. 2e and Supplementary Fig. 5c, e, f), suggesting that MLL4 deficiency promotes the influx of muscle fuel to the TCA cycle. In fact, knockdown of MLL4 increased the rate of glucose consumption (Supplementary Fig. 5g). Thus, in vitro manipulation of MLL4 in myocytes provided further evidence that MLL4 exerts control on muscle fuel metabolism.
To further confirm the enhanced fuel utilization in MLL4-deficient muscle cells, we assessed the metabolic flux of glycolysis and the TCA cycle by conducting isotopic [13C]-glucose tracing experiments. Compared to control cells, the [13C]-glucose traced 3-phosphoglycerate (3-PG) (m + 3) and pyruvate (m + 3), were significantly increased in MLL4 KD myotubes (Fig. 2f–h). Moreover, we also observed elevated 13C incorporated TCA cycle intermediates, such as citrate (m + 2 and m + 4), α-ketoglutarate (m + 2 and m + 4) and malate (m + 2) in MLL4 KD myotubes (Fig. 2f–h). We also observed an increased trends in the labeled fractions of citrate (m + 4, m + 5), α-ketoglutarate (m + 2, m + 3), and malate (m + 2, m + 3) in MLL4 KD myotubes (Supplementary Fig. 5h–l). These results indicate that KD of MLL4 exhibits enhanced glucose utilization, which is consistent with the steady-state results in MLL4 KO muscles. Together, these results demonstrate that loss of MLL4 promotes fuel catabolism in muscle cells in a cell-autonomous manner.
Overall, these results suggest that MLL4 deletion in skeletal muscle enhances fuel catabolism. This muscle metabolic remodeling improves systemic glucose homeostasis and renders the liver protected from hepatic steatosis.
Muscle-specific deletion of MLL4 enhances glucose tolerance, insulin sensitivity, and reduces liver lipid accumulation primarily through AMPK activation in skeletal muscle
The activation of AMPK turns on fuel catabolism, and skeletal muscle AMPK activation alone has been shown to improve insulin sensitivity and glucose homeostasis. Phosphorylation of AMPK was significantly activated in muscles from Mll4SET MKO mice compared to WT controls under both CD and HFD conditions (Fig. 3a). In WT muscle, AMPK phosphorylation was increased during fasting and appropriately suppressed upon refeeding, consistent with the known energy-sensing role of AMPK (Supplementary Fig. 6a). In Mll4SET MKO mice, AMPK activation was not further enhanced during fasting, and notably, the suppression of AMPK phosphorylation upon refeeding was significantly blunted (Supplementary Fig. 6b). Consistent with the increase in muscle AMPK activation, the average percentage of SDH-positive myofibers was significantly increased in the muscle of HFD-fed Mll4SET MKO mice compared to WT controls (Supplementary Fig. 6c–e).
a Western blot and quantification of AMPK activation in GC muscles. n = 3 mice per group. P value: 0.0129, 0.0133. b Schematic of MyoAAV-mediated AMPK dominant-negative (DN) overexpression and HFD experiment. c, d Western blot and quantification of p-ACC, t-ACC, p-AMPK, and t-AMPK in GC muscles from MKO mice normalized to WT controls, n = 6 mice per group. P value: <0.0001, 0.0002. e Mouse images after 11 weeks of HFD. f, g Body weight and weight gain, WT-GFP, n = 13; MKO-GFP, n = 12; MKO-AMPK DN, n = 11. f *P value: 0.0003, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001. #P value: 0.013. g *P value: 0.0006, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001. #P value: 0.0248. h iWAT weight, WT-GFP, n = 13; MKO-GFP, n = 12; MKO-AMPK DN, n = 11. *P value: 0.003. #P value: <0.0001. i, j GTT and ITT with AUC analysis, WT-GFP, n = 13; MKO-GFP, n = 12; MKO-AMPK DN, n = 11. i *P value: 0.0003, 0.0011, <0.0001, 0.0001, 0.0035, 0.001. #P value: 0.0226, 0.0306, 0.0182, 0.0119, 0.0331, 0.03. j *P value: 0.0197, 0.0044, 0.0067, 0.0278, 0.0253. #P value: 0.0241, 0.036, 0.0252, 0.0113, 0.0378. k Liver weight, WT-GFP, n = 13; MKO-GFP, n = 12; MKO-AMPK DN, n = 11. *P value: 0.001. #P value: 0.0043. l H&E and Oil Red O staining of liver sections, scale bar = 50 μm, n = 10–13 mice per group. m Liver triglyceride levels, WT-GFP, n = 12; MKO-GFP, n = 11; MKO-AMPK DN, n = 11. *P value: 0.0001. #P value: 0.0015. Data are mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 vs. WT controls, #P < 0.05, ##P < 0.01, ###P < 0.001 vs. GFP controls, n.s. no significance. Statistical analysis performed using two-tailed unpaired Student’s t-test (d), one-way ANOVA (a, h, k, m) with Fisher’s LSD post-hoc test, or two-way ANOVA (f, g, i, j) with a Tukey’s post-hoc test. Source data are provided as a Source data file.
To evaluate if AMPK inhibition is able to reverse the metabolic improvements in Mll4SET MKO mice, we generated recombinant MyoAAV to express the dominant negative (DN) form (K45R) of AMPKα2. This MyoAAV approach allowed us to specifically suppress AMPK activation in the skeletal muscle of Mll4SET MKO mice and evaluate its effects on glucose tolerance, insulin sensitivity, body weight, and liver lipid accumulation upon HFD feeding (Fig. 3b). Western blot confirmed successful overexpression of AMPK DN specifically in skeletal muscle tissue (Supplementary Fig. 6f). Importantly, AMPK activity was suppressed in muscle as evident by reduced phosphorylation of ACC, a key downstream marker of AMPK activity (Fig. 3c, d). Whereas we observed an increase of iWAT weight in Mll4SET MKO-AMPK DN compared to Mll4SET MKO-GFP, there was no significant change in body weight (Fig. 3e–h), likely due to a concurrent reduction in lean tissue mass (Supplementary Fig. 6g). However, glucose tolerance and insulin tolerance tests revealed that the improvements in glucose tolerance observed in Mll4SET MKO mice were significantly diminished when muscle AMPK activation was suppressed (Fig. 3i, j). We further explored muscle AMPK activation in liver steatosis. Mll4SET MKO-AMPK DN mice showed significantly elevated liver weight compared to Mll4SET MKO-GFP mice (Fig. 3k). In addition, histological analysis demonstrated a marked reduction in hepatic steatosis in Mll4SET MKO mice but not in Mll4SET MKO-AMPK DN mice (Fig. 3l). Similarly, the reduction in liver triglyceride levels seen in Mll4SET MKO mice was also reversed in Mll4SET MKO-AMPK DN mice (Fig. 3m). Together, these findings suggest that skeletal muscle AMPK activation, driven by MLL4 deletion, plays a key role in regulating systemic glucose homeostasis and liver lipid metabolism.
MLL4 programs muscle AMPK activation
To dissect the mechanism through which MLL4 depletion activates muscle AMPK, we conducted a comprehensive study of the muscle transcriptome induced by MLL4 deletion, HFD feeding, or the combination. Comparison of gene expression profiles in Mll4SETf/f muscle revealed 638 genes in total whose expression was significantly regulated by HFD (Fig. 4a), and comparison between WT and Mll4SET MKO muscle showed 1430 and 1490 genes that were influenced by MLL4 abrogation on CD and HFD (Fig. 4a), respectively. Categorized into four subgroups (Fig. 4a), GO pathway analysis not only showed HFD-induced immune and inflammatory pathways that were suppressed by MLL4 deficiency, but also revealed muscle metabolic and contractile gene networks that were substantially altered by MLL4 abrogation (Fig. 4b and Supplementary Fig. 7a–e).
a Heatmap analysis of genes differentially regulated in muscles from indicated mice: WT or Mll4SET MKO mice were fed either a CD or HFD. Each group is represented by RNA-seq data from two independent samples generated from muscles from indicated mice. The differentially regulated genes were clustered into four groups, and a color scheme for fold change is provided. Red, relative increase in abundance, blue, relative decrease. b GO enrichment analysis of Cluster II (HFD-induced genes that were suppressed by MLL4 deficiency) and III (genes that were induced by MLL4 abrogation), with top five or ten terms shown. The P value is defined by one-sided Fisher’s exact test. c Analysis of the MLL4 ChIP-seq data in muscle combined with the mRNA-seq dataset upon muscle Mll4 deletion defines a set of genes directly regulated by MLL4. d Scatter plot of genes directly regulated by MLL4 under both CD and HFD conditions. A pair of genes (Nt5c1a and Ampd3) encoding AMP metabolizing enzymes are shown in blue. e A model for the regulation of muscle AMPK activation mediated by the concerted action of a pair of AMP metabolizing enzymes NT5C1A and AMPD3 that are transcriptionally activated by MLL4. f Heatmap showing AMP/ATP and ADP/ATP ratios in skeletal muscles from Mll4SET MKO mice and WT controls under both CD and HFD conditions. n = 6 mice per group. g AMP/ATP and ADP/ATP ratios in skeletal muscles from the indicated mouse genotypes under HFD conditions. n = 5 mice per group. P value: 0.0021, 0.0092. All data are shown as the mean ± SEM. **P < 0.05 vs. WT controls determined by two-tailed unpaired Student’s t-test (g). Source data are provided as a Source data file.
We next integrated transcriptomics and MLL4 ChIP-seq data to identify putative direct targets of MLL4 in skeletal muscle (Fig. 4c). Comparative analysis identified 213 direct targets that were commonly downregulated by MLL4 deletion under both CD and HFD conditions (Fig. 4d and Supplementary Table 1). Of particular interest were two genes implicated in the same AMP metabolizing pathway that regulates AMPK activation (Fig. 4e). Specifically, Nt5c1a, which is key for converting AMP into adenosine, and Ampd3, which catalyzes the hydrolytic deamination of AMP to inosine monophosphate (IMP), have been shown to negatively regulate AMPK activation40,41. The reduced expression of Nt5c1a and Ampd3 genes would be predicted to increase the intracellular availability of AMP, leading to AMPK activation and consequential fuel catabolism. Indeed, we confirmed that AMP/ATP and ADP/ATP ratios were increased in Mll4SET MKO muscle (Fig. 4f, g). Therefore, these results suggest that MLL4 stimulates the expression of a pair of AMP metabolizing enzymes NT5C1A and AMPD3, reducing the intracellular availability of AMP, thereby limiting AMPK activation and fuel catabolism in skeletal muscle.
Nt5c1a and Ampd3 mediate MLL4 regulation of muscle AMPK
Expression of NT5C1A and AMPD3 proteins was significantly decreased in Mll4SET MKO muscles compared to WT controls (Fig. 5a), concomitant with a marked increase in phosphorylation of AMPK. Consistent with increased MLL4 expression, both AMPD3 and NT5C1A protein levels were significantly elevated in the muscle of HFD-fed WT mice compared to CD conditions (Supplementary Fig. 8a, b). Interestingly, phosphorylation of AMPK remains unchanged under these conditions (Supplementary Fig. 8a, b). Notably, there are two isoforms of AMPD3 due to N-terminal truncation42,43. Both isoforms of AMPD3 were decreased in multiple muscle types from Mll4SET MKO mice compared to WT controls (Supplementary Fig. 8c, d). In contrast, AMPD1 remained unaffected by MLL4 deficiency (Supplementary Fig. 8c, d). Similar results were obtained in myotubes in culture, using lentiviruses-mediated MLL4 KD in C2C12 cells (Fig. 5b and Supplementary 8e, f). Furthermore, AMP/ATP ratio was increased in MLL4 KD myotubes (Fig. 5c). These results demonstrate that MLL4 regulates AMPK activation and fuel utilization in myotubes in a cell-autonomous manner. Additional studies were focused on Ampd3, given that Nt5c1a gene expression was undetectable in culture myotubes, as noted previously40 (Supplementary Fig. 8e). Similar to MLL4, lentiviral-mediated KD of Ampd3 resulted in a marked increase in AMPK phosphorylation (Fig. 5d and Supplementary Fig. 8g). To assess the relevance of this AMP metabolizing pathway in vivo, we performed Nt5c1a and Ampd3 gain- and loss-of-function studies in vivo in skeletal muscle. Adeno-associated virus (AAV)-mediated KD of both Nt5c1a and Ampd3 robustly stimulated AMPK phosphorylation in WT skeletal muscle (Fig. 5e, f). Conversely, AAV-mediated overexpression of Nt5c1a and Ampd3 resulted in attenuated AMPK phosphorylation in Mll4SET MKO muscles (Fig. 5g). Further, overexpression of Nt5c1a and Ampd3 markedly decreased muscle metabolites of glycolysis and acyl-carnitines in Mll4SET MKO mice to similar levels seen in WT control mice (Fig. 5h, i). To explore whether these changes could be attributed to alterations in fatty acid desaturation, we assessed the expression of key desaturase genes by qRT-PCR. As shown in Supplementary Fig. 8h–k, overexpression of Ampd3 and Nt5c1a in Mll4SET MKO muscle upregulated several desaturase transcripts, including Scd1 and Fads2 and fatty acid elongases Elovl1 and Elovl6, suggesting a possible role in maintaining unsaturated acyl-carnitine levels (Supplementary Fig. 8h, i). Interestingly, this overexpression did not significantly affect the expression of fatty acid oxidation genes but did induce de novo lipogenesis genes compared to Mll4SET MKO controls (Supplementary Fig. 8j, k). Together, these results suggest that MLL4 regulates muscle AMPK-mediated fuel catabolism in part through the regulation of Nt5c1a and Ampd3, although other MLL4-regulated genes may also contribute to this process.
a (Left) Representative Western blot of GC muscles lysates from WT and Mll4SET MKO mice under CD condition. (Right) Quantification of the NT5C1A/Tubulin, AMPD3/Tubulin, AMPD1/Tubulin, and p-AMPK/t-AMPK ratios were showed as arbitrary units (A. U.) and normalized (=1.0) to WT controls. n = 6 mice per group. P value: <0.0001, <0.0001, <0.0001. b Representative Western blot of AMPK activation in C2C12 myotubes with MLL4 knockdown. n = 4 independent experiments. c AMP/ATP ratio in MLL4 knock-down myotubes. n = 5 per group. P value: 0.0458. d Representative Western blot of AMPK activation in C2C12 myotubes with AMPD3 knockdown. n = 3 independent experiments. e Representative Western blot of AMPK activation in GC muscles of WT mice subjected to adeno-associated virus (AAV)-mediated knock-down of both Nt5c1a and Ampd3. f Quantification of Western blot in (e). n = 6 mice per group. P value: 0.0011, 0.0145. N.D. Not detected. g (Left) Representative Western blot of AMPK activation in GC muscles of indicated mice subjected to AAV-mediated overexpression of Nt5c1a and Ampd3. (Right) Quantification of the p-AMPK/t-AMPK signal ratios normalized (=1.0) to the WT-GFP control. WT-GFP, n = 7; MKO-GFP, n = 5; MKO-AMPK DN, n = 7. *P value: <0.0001. #P value: 0.0034. h Heatmap of acyl-carnitines and glycolysis metabolites in muscles of indicated mice. n = 5–8 mice per group. i Quantitative data for acyl-carnitines, WT-GFP, n = 6; MKO-GFP, n = 5; MKO-AD3 OE/NT5 OE, n = 8 and glycolysis metabolites, WT-GFP, n = 5; MKO-GFP, n = 5; MKO-AD3 OE/NT5 OE, n = 7 in muscles of indicated mice, internal standards and tissue weight were used to correct the areas of metabolites to obtain relative abundance. *P value: 0.0346, 0.0124. #P value: 0.0496, 0.0312, 0.0241. Data are mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 versus corresponding WT or shCtrl controls, #P < 0.05, ##P < 0.01 versus corresponding GFP controls, determined by two-tailed unpaired Student’s t-test (a, c, f) or one-way ANOVA (g, i) coupled to Fisher’s least significant difference (LSD) post-hoc test. Source data are provided as a Source data file.
Previous studies have identified futile cycle activation in skeletal muscle is crucial for systemic energy expenditure and body weight control44,45,46. We further analyzed our RNA-seq data and conducted gene expression validation studies. Whereas the expression of Pln and Atp2a1 was not induced in Mll4SET MKO muscles versus controls under HFD, Sln and Ucp3 gene level were significantly increased in Mll4SET MKO muscles (Supplementary Fig. 9a). These results were of interest because the Sln has been shown to be critical for the whole-body energy expenditure44,45. Western blotting confirmed the marked increased expression of SLN protein in Mll4SET MKO muscles under both CD and HFD (Supplementary Fig. 9b, c). The significant upregulation of Sln suggests that the calcium futile cycle is activated in Mll4SET MKO muscles, which likely also contributes to the protection from diet-induced obesity.
MLL4 and MEF2 function cooperatively to control Nt5c1a and Ampd3 expression
To define the mechanism involved in the activation of Nt5c1a and Ampd3 gene expression by MLL4, we interrogated muscle RNA-seq dataset in conjunction with our muscle ChIP-seq data for MLL433. Genomic localization of MLL4 was confirmed on Nt5c1a and Ampd3 gene loci in muscles. Specifically, both Nt5c1a and Ampd3 genes showed strong MLL4 binding (Fig. 6a, b). Importantly, deletion of Mll4 decreased mRNA levels of both Nt5c1a and Ampd3 in Mll4SET MKO muscles (Fig. 6a, b). Real-time PCR further confirmed that the mRNA expression of Nt5c1a and Ampd3, but not other AMP metabolism genes (e.g., Nt5c2, Ak1, Adk, Ada), was reduced in the GC muscles of Mll4SET MKO mice compared to WT controls (Fig. 6c). Notably, Nt5c1a and Ampd3 mRNA levels were also markedly decreased in the soleus muscles of Mll4SET MKO mice (Supplementary Fig. 9d). Because we have recently uncovered that MLL4 directly binds to enhancers and functions as a coactivator of MEF2 to activate transcription of slow-oxidative myofiber genes33, we thus sought to determine whether MLL4 and MEF2 function cooperatively to regulate Nt5c1a and Ampd3 expression. Interestingly, we found that MLL4 and MEF2 could co-localize on both of Nt5c1a and Ampd3 gene loci (Fig. 6d). ChIP-qPCR analysis confirmed MEF2 binding at these loci, with a reduced enrichment trend observed in Mll4SET MKO muscles (Supplementary Fig. 9e). Moreover, within a MLL4/MEF2 co-bound region located approximately 4.6 kb downstream of the Ampd3 transcription start site (TSS), we identified a highly conserved DNA sequences conforming to the consensus binding sites for MEF2 [(C/T)TA(T/A)4TA(G/A)] (Fig. 6e). These results strongly suggest that MEF2 is involved in the MLL4-mediated regulation of Nt5c1a and Ampd3 gene transcription. Consistent with this conclusion, AAV-mediated overexpression of MEF2C in WT skeletal muscle resulted in markedly increased expression of AMPD3 (Fig. 6f, g). Notably, NT5C1A levels did not increase upon MEF2C overexpression, indicating that additional regulatory factors might require for induction of NT5C1A expression (Fig. 6g and Supplementary Fig. 9f). Interestingly, MEF2C-mediated induction of AMPD3 was significantly attenuated in the absence of MLL4 (Fig. 6g), demonstrating the requirement of MLL4 for the stimulatory effects of MEF2 on Ampd3 gene. Taken together, these results suggest that MLL4 cooperates with MEF2 to drive the expression of Ampd3 and Nt5c1a.
a, b MLL4-dependent regulation of Nt5c1a and Ampd3 gene expression. (Top) ChIP-seq binding profiles for MLL4 in WT and Mll4SET MKO muscle. mRNA-seq data from WT and Mll4SET MKO muscle is shown at the bottom, indicating a high correlation of the two datasets. Input, genomic DNA from myocytes. c Expression of genes (qRT–PCR) involved in AMP metabolism in GC muscles from the indicated genotypes. Ampd3, WT, n = 5; MKO, n = 5 mice per group. Nt5c1a, Nt5c2, Ak1, Adk, Ada, WT, n = 6; MKO, n = 6 mice per group. P value: <0.0001, 0.0067. d MLL4 and MEF2D ChIP-seq tracks from myocytes at the Ampd3 and N5c1a loci. Data obtained from published datasets GSE138994 and GSE43223 were analyzed. e Putative conserved MEF2-binding site within the intron 1 of mouse Ampd3 gene. f AAV-mediated overexpression of MEF2C in WT and Mll4SET MKO muscle. Ex1, exon 1. g (Left) Representative Western blot analysis of protein extracts prepared from GC muscles of indicated mice subjected to AAV-mediated overexpression of MEF2C. (Right) Quantification of the AMPD3/GAPDH signal ratios were normalized (=1.0) to WT GFP controls. n = 5 mice per group. *P value: <0.0001. #P value: <0.0001. All data are shown as the mean ± SEM. **P < 0.01, ***P < 0.001 vs. corresponding WT controls, ###P < 0.001 versus corresponding GFP controls, determined by two-tailed unpaired Student’s t-test (c) or one-way ANOVA (g) coupled to Fisher’s least significant difference (LSD) post-hoc test. Source data are provided as a Source data file.
Muscle MLL4-MEF2 regulatory axis is associated with fasting plasma glucose in humans
To determine the relevance of the muscle MLL4-MEF2 regulatory circuit in humans, muscle biopsies from 29 young and 23 old sedentary subjects were analyzed. Previous analyses demonstrated that older participants had higher fasting plasma glucose and BMI (Body Mass Index) than the young participants (Supplementary Table 2)47. As expected, muscle tissue from the older group exhibited higher MLL4, MEF2A, MEF2B and MEF2D gene expression compared to the young group (Fig. 7a). Expression levels of AMPD3 were also significantly elevated in the older group compared to the young controls (Fig. 7a). However, this up-regulation was not observed with MEF2C and NT5C1A levels (Fig. 7a and Supplementary Fig. 10a). In addition, there was a significant positive correlation between MLL4 vs. AMPD3 and NT5C1A mRNA levels (Fig. 7b). Moreover, MEF2 mRNA levels were also positively correlated with AMPD3 and NT5C1A levels (Fig. 7b and Supplementary Fig. 10b).
Samples are from 29 young and 23 old sedentary subjects. a mRNA expression levels of MLL4, MEF2D, MEF2A, MEF2B, AMPD3, NT5C1A were determined by RNA-seq. Data represent the mean ± SEM. Significant differences were analyzed using two-sided unpaired Mann-Whitney test. *P < 0.05 vs young controls. b Correlation between MLL4 and MEF2D gene expression and that of muscle metabolic genes, AMPD3, and NT5C1A. Spearman correlation analysis was used to determine the correlation. P value: 0.003, <0.001, <0.001, <0.001. c Correlation between MLL4 and MEF2 and fasting plasma glucose and BMI. Spearman correlation analysis was used to determine the correlation. d Western blot analysis of human skeletal muscle myotubes transduced with lentiviruses expressing scramble (shCtrl) or MLL4 shRNA (shMLL4), showing MLL4, AMPD3, p-AMPK, t-AMPK, and GAPDH levels. e Quantification of MLL4/GAPDH, AMPD3/GAPDH, and p-AMPK/t-AMPK signal ratios normalized to shCtrl (=1.0). n = 3 independent experiments. P value: 0.0056, 0.0013, 0.0053. f Mll4 and AMPD3 gene expression (qRT-PCR) in MLL4 knockdown myotubes. n = 3 independent experiments. P value: 0.0008, 0.0183. g Glucose consumption in LHCN-M2 myotubes after 8, 16, and 24 h. n = 6 per group. P value: 0.032. All data are presented as mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 vs. corresponding controls, determined by two-tailed unpaired Student’s t-test. Source data are provided as a Source data file.
The relationship between the muscle MLL4-MEF2 regulatory circuit with fasting plasma glucose and BMI was next assessed. As shown in Fig. 7c, a moderate positive correlation was observed between the expression of MLL4 and fasting plasma glucose levels. MEF2A, MEF2B and MEF2D expression also positively correlated with fasting plasma glucose levels (Fig. 7c and Supplementary Fig. 10c). MEF2A but not MLL4 mRNA levels were also positively correlated with human BMI (Fig. 7c). Notably, the levels of AMPD3 mRNA neither exhibited a significant correlation with fasting plasma glucose or human BMI (Supplementary Fig. 10c).
We further validated the MLL4 regulatory axis in human myotubes. The LHCN-M2 cell is a well-characterized human skeletal muscle cell widely used for human muscle metabolism and disease studies48,49,50,51. The LHCN-M2 myotubes were transduced with either a control scramble shRNA or MLL4 shRNA. Knockdown of MLL4 resulted in reduced protein levels of MLL4 and AMPD3, along with an increase in the phosphorylation of AMPK (Fig. 7d, e), indicating AMPK activation in response to MLL4 knockdown in human skeletal muscle cells. qRT-PCR confirmed that MLL4 knockdown decreased AMPD3 expression at the transcriptional level (Fig. 7f). Moreover, MLL4 knockdown led to a significant increase in glucose consumption (Fig. 7g), supporting the role of the MLL4/AMPD3/AMPK regulatory axis in enhancing glucose metabolism in human skeletal muscle cells. Together, these findings demonstrate the conserved role of the MLL4/AMPD3/AMPK axis in human skeletal muscle cells and underscore the relevance of MLL4-mediated pathways in human muscle metabolism.
Pharmacologic inhibition of the AMP metabolizing pathway activates muscle AMPK and protects against diet-induced obesity and improves glucose homeostasis
The MLL4 regulatory and target pathways unveiled in this study show tremendous promise for the development of strategies to enhance muscle fuel catabolism relevant to the ravages of caloric excess. Pentostatin is a United States Food and Drug Administration (FDA)-approved drug that dually inhibits ADA and AMPD enzyme activities52,53,54. This raises the interesting question as to whether pharmacologic inhibition of the AMP metabolizing pathway by Pentostatin is able to activate muscle cell AMPK and improve insulin sensitivity and glucose homeostasis. To test this idea, we treated C2C12 myotubes with Pentostatin, which resulted in elevated intracellular AMP/ATP and adenosine levels, accompanied by increased phosphorylation of AMPK α subunit in myotubes (Fig. 8a–c). Similarly, treatment of differentiated human LHCN-M2 myotubes with Pentostatin significantly enhanced AMPK phosphorylation at Thr172. Notably, phosphorylation of ACC, an established downstream target of AMPK, was also increased (Supplementary Fig. 11a, b).
a (Top) Schematic depicting the AMP metabolizing pathway that is inhibited by Pentostatin. (Bottom) Representative Western blot of AMPK activation in C2C12 myotubes treated with DMSO (vehicle) or Pentostatin (100 μM or 200 μM) for 4 h. n = 3 independent experiments. b Quantification of the p-AMPK/t-AMPK ratios normalized (=1.0) to the vehicle control. n = 3 independent experiments. P value: 0.0334, 0.0111. c Quantitative data for AMP/ATP in C2C12 myotubes treated with vehicle or Pentostatin (200 μM) for 4 h. n = 5 per group. P value: 0.038, 0.0021. d, e Body weight (d) and body weight gain (e). Six-week-old male C57BL/6 mice were fed a HFD for 4 months. During the HFD feeding, Pentostatin or vehicle control was administered intraperitoneally as once-every two days dose (0.6 mg/kg). n = 9 mice per group. d P value: 0.0156, 0.0149, 0.0007, <0.0001, <0.0001, <0.0001, <0.0001. e P value: 0.04, 0.0047, 0.0045, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001. f iWAT weight. n = 9 mice per group. P value: <0.0001. g Food consumption per day on HFD. n = 9 mice per group. h, i Oxygen consumption (h) and energy expenditure (i) during the dark period in mice treated with Pentostatin for the prevention of dietary obesity. i ANCOVA-predicted MR at a body mass of 40 g. n = 8 mice per group. j Locomotor activity over 12 h light/dark cycle. HFD 0.9% NaCl, n = 8; HFD 0.6 mg/kg Pento, n = 4. k, l Glucose tolerance test (GTT) and insulin tolerance test (ITT). AUC for GTT and ITT is shown. n = 9 mice per group. k P value: 0.041, 0.0116, 0.0006, <0.0001, 0.0052, 0.0008. l P value: <0.0001, <0.0001, 0.0204, 0.002, 0.0003, <0.0001, <0.0001. Data are mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 vs. corresponding vehicle controls determined by two-tailed unpaired Student’s t-test (c, f, g), one-way (b) or two-way ANOVA (d, e, k, l) coupled to Fisher’s least significant difference (LSD) post-hoc test. Source data are provided as a Source data file.
Next, we assessed the pharmacological potential of Pentostatin in the context of diet-induced obesity and insulin resistance. C57BL/6 mice were fed a HFD for 4 months. During the HFD feeding, Pentostatin or vehicle control was administered intraperitoneally as once-every two days dose (0.6 mg/kg) (Fig. 8d). Notably, Gdf15, a key stress marker, was not upregulated in muscle tissue of Pentostatin-treated mice (Supplementary Fig. 11c). Pentostatin administration significantly slowed weight gain and reduced iWAT weight in HFD-fed mice without affecting their food intake and locomotor activity (Fig. 8e–g, j). Pentostatin-treated mice displayed higher oxygen consumption and energy expenditure in comparison with control mice (Fig. 8h, i). Moreover, glucose and insulin tolerance tests revealed a significant improvement in glucose homeostasis in Pentostatin-treated mice relative to control mice (Fig. 8k, l). In parallel, Pentostatin also significantly enhanced AMPK phosphorylation in both the liver and white adipose tissue (Supplementary Fig. 11d, e). Together, these data demonstrate that pharmacological inhibition of the AMP metabolizing pathway by Pentostatin protects from diet-induced obesity and improves metabolic parameters in mice.
Discussion
Changes in nutrient availability and physiological demands evoke a metabolic re-programming in skeletal muscle. Our results demonstrate that skeletal muscle cells sense and transform the environmental signal into transcriptional and metabolic adaptation through an enhancer activation regulatory mechanism. We specifically identify the MLL4-dependent enhancer activation as a critical regulatory node for the governance of skeletal muscle metabolic efficiency and the control of systemic metabolic homeostasis (Fig. 9). We showed that MLL4 expression is regulated in response to nutrient availability in skeletal muscle. Muscle-specific deletion of MLL4 in mice confers resistance to dietary obesity with improved systemic insulin sensitivity and glucose tolerance despite reducing exercise endurance. These seemingly paradoxical phenotypes are due to lower endurance type I muscle fiber proportion but greater fuel catabolism in MLL4 deficient muscles. Mechanistically, MLL4 directly binds to and cooperates with MEF2 to activate expression of a pair of AMP metabolizing enzymes NT5C1A and AMPD3, reducing the intracellular availability of AMP, thereby limiting muscle AMPK activation and fuel catabolism. Moreover, pharmacologic targeting the AMP metabolizing pathway by Pentostatin activates muscle AMPK, protects mice from diet-induced obesity and ameliorates obesity-related morbidities. Thus, our results highlight a critical role of enhancer regulator MLL4 in the governance of muscle metabolic efficiency and systemic metabolism via the control of muscle AMPK activation. Our current study also offers potential strategies for the pharmacological treatment of obesity and related metabolic disorders.
The schematic depicts the proposed model for the governance of skeletal muscle metabolic efficiency and systemic metabolism by MLL4. MLL4 directs skeletal muscle metabolic programming by limiting AMPK-mediated fuel catabolism.
Our findings and conclusions of the current studies shed insight into a previously unrecognized role of enhancer regulator MLL4 in modulating muscle AMPK activity that is crucial for systemic glucose homeostasis, and underscore the point that links MLL4’s epigenetic control with AMPK’s metabolic actions. Our data suggest that AMPK activation in MLL4-deficient muscle is indeed critical for mediating many of the metabolic improvements observed in Mll4SET MKO mice. For instance, our finding indicates that the reductions in liver lipids observed in Mll4SET MKO mice are not merely secondary to reduced body weight, but instead rely on active AMPK signaling in skeletal muscle. Future studies are needed to directly test whether muscle-derived signals or altered substrate flux contribute to liver lipid metabolism. Our results also suggest that the SLN-mediated calcium futile cycle is activated in Mll4SET MKO muscles, leading to an increase in systemic energy expenditure, which likely contributes to the protection from diet-induced obesity. It would seem likely that muscle-specific deletion of MLL4 drives a concordant activation of muscle AMPK and SLN futile cycle to improve glucose homeostasis, liver lipid accumulation and protection from obesity.
Recently, we found that MLL4-dependent enhancer activation is required for establishing and maintaining slow type I muscle fiber program to ensure running endurance33. The results described herein raise the interesting question regarding what the normal biological role is of MLL4 signaling in skeletal muscle. Our results indicate that an intact MLL4 regulatory pathway is necessary for the coupling of exercise capacity and fuel anabolism to optimize muscle metabolic efficiency. Given that MLL4, a fasting-repressed protein, is nonetheless required for efficient transition from a catabolic to an anabolic state. We propose that transient downregulation of MLL4 during acute fasting may facilitate catabolism for survival, whereas chronic deletion of MLL4, as modeled in our Mll4SET MKO mice, creates a constitutive “pseudo-fasted” state, even under nutrient-rich conditions. In this state, energy utilization remains persistently high, and energy is dissipated, in part via activation of futile cycles (e.g., SLN-mediated calcium cycling). This may underlie the observed reductions in metabolic efficiency and the redirection of nutrients away from storage toward continuous fuel catabolism.
Enhancers play a pivotal role in controlling gene expression and are marked by modifications such as H3K4me1/222,24,55. While the transcriptional regulation of skeletal muscle metabolic gene expression through transcription factors like nuclear receptor PPARs, ERRs, MEF2, Tbx15, and NURR1, as well as their associated coregulators, has been well-established10,11,12,13,14,15,16,17,18,56,57, the mechanisms governing skeletal muscle metabolic reprogramming at the level of enhancer activation remain unclear. In this study, we identified a key role for enhancer regulator MLL4-directed gene programs in the control of muscle cell fuel metabolism both in vivo and in vitro. Our findings are consistent with results from an earlier study reporting that whole-body Mll4+/- protects against diet-induced obesity and fatty liver but suggest that muscle-specific deletion is sufficient to cause the phenotype58. Whereas our data suggest that MLL4 cooperates with MEF2 to drive the expression of metabolic genes in skeletal muscle, it remains to be tested whether MLL4 may also serves as a major coactivator for diverse nuclear receptors. Our findings have expanded our understanding of the physiological importance of MLL4-mediated enhancer activation. Conceivably, this muscle fuel metabolism regulatory action may reflect the multi-functions of MLL4 in utilizing its enhancer remodeling activity to direct gene transcription adaptation to various physiological and pathological stimuli. A critical next step in elucidating MLL4’s mechanism of action will be to distinguish between its enzymatic and non-enzymatic functions. A definitive approach for addressing this would be the generation of enzyme-dead MLL4 knock-in mice carrying SET domain-inactivating mutations.
Combining comprehensive metabolomics and genomic analyses, we have defined the unique MLL4 target metabolic pathway in skeletal muscle. Specifically, MLL4 coordinately stimulates the expression of a pair of AMP-metabolizing enzymes NT5C1A and AMPD3 to limit AMPK activation and muscle fuel catabolism in a cell-autonomous manner. We found that Nt5c1a and Ampd3 gene expression were suppressed in MLL4 MKO muscles, concomitant with a marked activation of AMPK. These results were intriguing because NT5C1A and AMPD3 are known to negatively affect AMPK activation40,41. AMPK is a master regulator of cellular and organismal metabolism that is implicated in controlling multiple aspects of glucose and lipid metabolism19,20,21. In addition to the well-known role in regulating cellular metabolism acutely through phosphorylation of multiple metabolic enzymes, AMPK also modulates cell metabolic adaptation via transcriptional regulation19,20,21,59,60. Notably, KD of Nt5c1a and Ampd3 in WT skeletal muscle reproduced the effects of MLL4 deficiency on AMPK activation, whereas overexpression of Nt5c1a and Ampd3 in MLL4-deficient muscles attenuated AMPK activation and fuel catabolism. Thus, we have uncovered an enhancer activation mechanism for the regulation of muscle AMPK mediated by the concerted action of a pair of Nt5c1a and Ampd3 genes that are activated by MLL4 in muscle. Our results suggest that although MLL4 activation by HFD enhances the expression of AMP-catabolizing enzymes, this may not be sufficient to significantly reduce the cellular AMP:ATP ratio to suppress AMPK activation in vivo. We speculate that in early HFD exposure, the induction of AMP-metabolizing enzymes may buffer AMP accumulation to some extent, but not sufficiently suppress AMPK activity due to ongoing metabolic stress. Notably, Kviklyte et al. reported findings that contradict earlier studies, showing no modulation of AMP levels or AMPK activity in NT5C1A and NT5C2 knockout muscles61. This discrepancy could be attributed to compensatory mechanisms or secondary effects in global knockout models. In addition, previous study also demonstrated that NT5C2 deletion improved insulin resistance on a HFD, likely independent of AMPK activation, though participation of AMPK could not be fully excluded62.
Our data did not suggest a detrimental impact on overall health or lifespan in muscle-specific Mll4SET MKO mice. Mll4SET MKO mice maintain normal food intake, and display improved glucose tolerance, enhanced insulin sensitivity, and reduced muscle inflammation. Although AMPK is activated in Mll4SET MKO muscle, it does not lead to abnormal glycogen accumulation. AMP and ADP are potent allosteric activators of phosphofructokinase-1 (PFK1), the rate-limiting enzyme in glycolysis, and this aligns with our observation that glycolytic pathway activity is increased in MLL4-deficient muscles. While PFK1 activation could contribute to the glycolytic phenotype, we also noted elevated levels of TCA intermediates and fatty acid β-oxidation metabolites. Therefore, these changes likely reflect the combined effects of enhanced AMPK signaling and nucleotide-driven metabolic reprogramming.
Our survey of the human muscle samples demonstrated that the MLL4-MEF2 circuit was activated in old sedentary subjects. Specifically, MLL4, MEF2A, MEF2B, MEF2D and AMPD3 levels were higher in muscle samples obtained from older participants group compared with that of the young group. We also found that the expression of MLL4 and MEF2 was correlated with expression of AMPD3 and NT5C1A genes, and fasting plasma glucose levels. However, we did not see similar correlations with their downstream targets AMPD3 or NT5C1A expression in humans. This could be related to species differences. Alternatively, the downstream programs of MLL4 directed by AMPD3 and NT5C1A in humans may not be manifested in the older versus young group comparison because the body composition differences were small as the participants were not obese. Given that our studies were performed while the participants were on similar diet, perhaps the response would have been different if fat intake were different. Notably, our findings demonstrated the conserved role of the MLL4/AMPD3/AMPK axis in human skeletal muscle cells, and thus strengthened the human relevance of our findings.
Activation of skeletal muscle AMPK to enhance fuel catabolism has been a powerful therapeutic strategy to treat metabolic disorders20,21,63. However, due to the broad family of AMPK heterotrimers, pharmacologically targeting the AMPK heterotrimers has been a challenge. Pentostatin is a FDA-approved drug that dually inhibits ADA and AMPD52,53,54. We found that Pentostatin is capable of eliciting AMPK activation in cultured myotubes. Our findings offer a pharmacologic strategy to activate the muscle AMPK via altering the expression or activity of AMP-metabolizing enzymes. The weight loss by Pentostatin treatment was not due to reduced food intake and physical activity, making a generalized toxic effect of the drug unlikely. It should be noted that our use of Pentostatin served to mechanistically validate the role of AMP-metabolizing enzymes, rather than to propose it as a clinical anti-obesity agent. Because Pentostatin was administered systemically, our data support the idea that the metabolic benefits of Pentostatin are the result of combined AMPK activation across multiple organs. Future studies using tissue-targeted delivery or skeletal muscle–selective AMPK activators will be valuable to further dissect the contribution of muscle-specific AMPK activation.
In summary, our results identify a pivotal role for enhancer regulator MLL4 in controlling skeletal muscle fuel metabolism and metabolic efficiency that governs systemic metabolism. Our study links MLL4’s epigenetic control to AMPK’s metabolic actions and suggests a potential therapeutic strategy for metabolic diseases.
Methods
Animal studies
All animal studies were conducted in strict accordance with the institutional guidelines for the humane treatment of animals and were approved by the Institutional Animal Care and Use Committee at the Model Animal Research Center (MARC) of Nanjing University (Approval No. GZJ07). Wild-type male C57BL/6 J wild-type mice were from GemPharmatech Co., Ltd (Jiangsu, China). Generation of muscle-specific Mll4-knockout (Mll4SETf/fHSA-Cre) mice has been described33. All mice were back-crossed into C57BL/6 J background for more than 6 generations before being used in experiments. Male mice were allowed ad libitum access to standard laboratory rodent chow or high fat diet (60% of calories from fat, Research Diets no. D12492) as indicated. Pentostatin was purchased from Selleckchem (CAS.53910-25-1), and Pentostatin was dissolved in 0.9% NaCl for intraperitoneal injection. For preventing obesity, 6-week-old male C57BL/6 mice were fed a HFD for 4 months and were intraperitoneally injected with Pentostatin (0.6 mg/kg mouse body weight) as once-every two days dose during the HFD feeding. Male mice of the indicated genotypes and ages (2 months to 5 months) were used. The mouse numbers for each experiment is provided in the legend of each Figure and Supplementary Figs. The animals were maintained with free access to pellet food (XieTong Biology, 1010082) and water in plastic cages at 21 ± 2 °C, relative humidity of 55 ± 10% and kept on a 12 h light-dark cycle. All mice are harbored in the specific pathogen-free facility in Nanjing University. Littermate controls were used in all cases.
Human studies
Details on subject characteristics and procedures have been described47 and are provided in Supplementary Table 2. The study included 18–30 years old sedentary younger adults (16 male, 13 female) and 65–80 years old sedentary older adults (12 male, 11 female). Briefly, the study design was approved by the Mayo Clinic Institutional Review Board and registered under Clinical Trials #NCT01477164 and # NCT01738568 (clinicaltrials.gov). All participants were informed of study procedures and provided written consent. Body composition was measured after an overnight fast with dual-energy X-ray absorptiometry. Metabolic measurements were performed after 3 days of weight-maintenance meals (20% protein, 50% carbohydrates, and 30% fat). Participants refrained from exercise for 72 h prior to metabolic study days. Participants repeated the 3-days of weighed meals and were admitted to the Clinical Research Unit of the evening of the third day. A light snack was provided at 2100 then participants remained fasting overnight. At 1000 h, biopsies (~350 mg) were collected from the vastus lateralis for RNA sequencing.
Metabolite extraction
Metabolite extraction of skeletal muscle tissues and C2C12 myotubes was performed through a chloroform/methanol/water system or a methanol/methyl tert-butyl ether/water system64,65. In brief, we followed a rapid dissection process, ensuring that muscles were snap-frozen within seconds of excision to preserve the native nucleotide levels. The muscle tissues were dissected immediately and quickly immersed in liquid nitrogen. All experiments included a minimum of five biological replicates per group. We employed capillary electrophoresis-mass spectrometry (CE-MS), which is a well-established and precise method for detecting polar metabolites such as adenine nucleotides (AMP, ADP, and ATP)65,66. We implemented a relative quantification strategy enhanced by multiple safeguards against technical variation. Specifically, we used a panel of isotope-labeled internal standards—including decanoylcarnitine-d3, palmitoylcarnitine-d3, FFA C16:0-d3, and FFA C18:0-d3 (for liquid chromatography-mass spectrometry, LC-MS), and methionine sulfone and camphorsulfonic acid (for CE-MS)—to adjust for extraction efficiency, matrix interference, and instrument drift. Furthermore, all metabolite abundances were normalized to tissue weight (mg) to control for inter-sample loading differences, providing a consistent basis for biological comparison. 500 μL ice-cold methanol with internal standard 1 (IS1, including L-methionine sulfone and D-camphor-10-sulfonic acid, Human Metabolome Technologies, H3304-1002, 1:200, used to standardize the metabolite intensity and to adjust the migration time) were added to samples, and then submitted to mixed grinding apparatus (Scientz-48) for homogenization (35 Hz, 1 min) followed by addition of 500 μL chloroform and vortex for 30 s. After phase breaking using 200 μL water and centrifugation (13,000 × g, 4 °C, 15 min), 420 μL hydrophilic layer was transferred for ultrafiltration through a 5-kDa cutoff filter (Millipore, cat. UFC3LCCNB-HMT). Simultaneously, the quality control (QC) sample was prepared by combining the aqueous phase from each sample and then filtered. Samples were then vacuum dried and stored at −80 °C until CE-MS analysis. For acyl-carnitines and fatty acids analysis using LC-MS, isotope-labeled internal standards as stated above were used to improve the precision of quantitative analysis and monitor errors arising from the sample pretreatment and analysis procedures. 300 μL ice-cold methanol with internal standards were added to samples, and then submitted to mixed grinding apparatus for homogenization (35 Hz, 1 min) followed by addition of 1 mL methyl tert-butyl ether and vortex for 1 min. After phase breaking using 300 μL water and centrifugation (13,000 × g, 4 °C, 15 min), 600 μL hydrophobic layer was collected and freeze-dried for fatty acids analysis. At the same time, the QC sample was prepared by combining the hydrophobic layer from each sample and then vacuum dried. For acyl-carnitines analysis, 300 μL hydrophilic layer and 300 μL hydrophobic layer were freeze-dried. QC sample was also prepared by combining the aqueous phase and then vacuum dried to evaluate the analytical quality.
Myotubes were rinsed with 25 ml of 5% mannitol solution (water dissolved) and instantly frozen in liquid nitrogen. 1 mL methanol containing IS1 were added before the cells are scraped off from the dish, mixed with 1 mL chloroform and vortexed for 20 s. After phase breaking using 400 μL water and centrifugation at 15,000 × g for 15 min at 4 °C, 400 μL of aqueous phase was collected and was then filtrated through a 5-kDa cutoff filter by centrifuging at 12,000 × g for 3 h at 4 °C. The filtered aqueous phase was then freeze-dried in a vacuum concentrator and stored at −80 °C until CE-MS based metabolite analysis.
13C6-glucose flux
For metabolic flux analysis, [U-13C6] glucose (from Cambridge Isotope Laboratories) was added to C2C12 myotubes for the indicated times at a final concentration of 25 mM following 30 min of starvation. CE-MS-based metabolic flux analysis was performed, followed by metabolite identification and quantitation. In-house developed software (SIAM) was used to calculate mass isotopologue distribution profiles65,67. Relative abundance of isotopologues was analyzed through peak area normalization method. Cells were washed and then quenched in liquid nitrogen and stored at −80 °C until MS analysis was performed.
Metabolomics analyses
CE-MS-based metabolomics and metabolic flux analysis were conducted on CE (G7100A, Agilent) coupled to the time of flight (TOF) mass spectrometry (G6224A, Agilent). The fused silica capillary (50 μm i.d. × 80 cm, HMT, Japan) was used for sample separation. Two analysis modes were performed and the temperature of the capillary was set at 20 °C68. For cation mode, 1 M formic acid was used as the background electrolyte and a positive voltage of 27 kV was applied during the CE separation. 50 mM ammonium acetate (pH = 8.5) was prepared as the running electrolyte for anion mode and a voltage of 30 kV was used during the separation. To assist the electroosmotic flow, an internal pressure of 17 mbar was also applied to the inlet capillary. For TOF/MS, the electrospray ionization was performed. Mass spectrometry parameters of positive ion mode were set as follows: dry gas temperature, 300 °C; nitrogen flow, 7 L/min; nebulizer pressure, 5 psig; capillary voltage, 4 kV; fragmentor, 105 V; skimmer, 50 V; Oct RFV, 650 V; acquisition rate, 1.5 spectra/s; mass range, 60–1000 Da. For negative ion mode, most parameters were identical to those used in the cation mode, except that the scanning range of 50–1000 Da. Besides, the voltage of the capillary and fragmentor were reset at 3.5 kV and 125 V, respectively. The qualitative analysis of metabolites was preceded based on the pre-analyzed metabolite standard library (HMT) and peak extraction and identification were carried out with Quantitative Analysis Software (B.04.00, Agilent). While we did not convert these signals to absolute concentrations (e.g., nmol/g tissue), the combination of IS-based correction and tissue normalization provides high-quality, biologically interpretable data.
LC-MS analysis was performed by an ACQUITY UPLC system (Waters) coupled with a tripleTOF™ 5600 plus mass spectrometer (AB SCIEX). The C8 ACQUITY column (2.1 mm × 50 mm × 1.7 µm) was used for acyl-carnitines separation in positive ion mode. The mobile phase A consisted of 0.1% formic acid in water and phase B consisted of 0.1% formic acid in acetonitrile69. The flow rate was set as 0.4 mL/min and the column temperature was 60 °C. Gradient elution started at 5% B, was held at this concentration for 0.5 min, was linearly increased to 40% B at 2 min, reached 100% B at 8 min and was held at this concentration for 2 min. Finally, the elution gradient was returned to 5% B within 0.1 min and held at this concentration for 1.9 min for equilibration. The complete running time of the program was 12 min. The C8 ACQUITY column (2.1 mm × 100 mm × 1.7 µm) was used for fatty acids separation in negative ion mode70. The mobile phase A consisted of acetonitrile/water (60:40, v/v) containing 10 mM ammonium acetate, and mobile phase B was isopropanol/acetonitrile (90:10, v/v) with the same additives. A 13-min gradient program was employed, starting at 50% B, was held at this concentration for 1.5 min, was linearly increased to 85% B at 9.0 min, reached 100% B at 9.1 min, and was held at this concentration for 1.9 min. Finally, the elution gradient was returned to 50% B within 0.1 min and held at this concentration for 1.9 min for equilibration. The flow rate was set as 0.3 mL/min and the column temperature was 60 °C. The ion spray voltage for MS was set at 5500 V and 4500 V in positive and negative ion modes, respectively. The interface heater temperature was 500 °C and 550 °C in positive and negative ion modes, respectively. Ion source gas 1, ion source gas 2, and curtain gas were set at 50, 50, and 35 psi in positive ion mode and 55, 55, and 35 psi in negative ion mode, respectively. The MS/MS fragmentation patterns were acquired using an information-dependent analysis; the collision energy was set to 30 V (positive mode) and −30 V (negative mode) with a collision energy spread of 15 V. The MS scan range was 80–1000 Da for acyl-carnitines analysis in positive ion mode and 150–1250 Da for fatty acids in negative mode. All samples were randomly analyzed to decrease errors derived from preparation and instrument analysis. Additionally, the QC samples were identically inserted into the analytical sequence to monitor the reproducibility of the analytical method. Acyl-carnitines and fatty acids identification was based on exact mass, retention time and MS/MS pattern. Peakview workstation (V1.2, AB SCIEX, USA) was used to check MS/MS information of metabolites and Multiquant (V2.1, AB SCIEX, USA) was used to obtain the area of identified metabolites. The applied database search engines were HMDB, Metlin (https://metlin.scripps.edu) and LIPID MAPS. The metabolomic datasets generated in this study are provided as Supplementary Data 1–5.
For statistical analysis, first, the raw data from CE-MS and LC-MS was normalized by peak area of internal standard and the weight of tissues. For cell’s metabolomics analysis, the quantity of metabolites was normalized by total peak area of whole metabolites from a cell sample. All metabolomic studies were analyzed by Student’s t test (2-tailed) or one-way ANOVA coupled to a Fisher’s least significant difference (LSD) post-hoc test when more than two groups were compared, setting p < 0.05 as the significant difference levels. K-means cluster analysis (K = 4) was performed with the Cluster 3.0 for the normalized metabolite data (p < 0.05) from CE-MS. Metabolite-associated pathways were analyzed using MetaboAnalyst 5.0 (Xia Lab at McGill University, Montreal, Canada; metaboanalyst.ca). Principal component analysis (PCA) and heatmap analyses were generated by using R software, version 4.0.3 with the factoextra/FactoMineR/ggpubr and gplots packages, respectively.
AAV injection
MyoAAVs expressing GFP or the dominant-negative (DN) form (K45R) of AMPKα271 were generated and provided by OBiO Technology Corp., Ltd. (Shanghai, China). These MyoAAVs were packaged into the MyoAAV-1A vector. MyoAAV-GFP and MyoAAV-AMPK DN were diluted in 0.9% NaCl to a concentration of 5 × 1012 VG/mL and administered via tail-vein injection (100 μL per mouse) to 6-week-old male WT and Mll4SET MKO mice (n = 11–13 mice per group). AAVs for in vivo expression of GFP, AMPD3, NT5C1A or MEF2C were generated and provided by the Rongsen Gene Technology Co., Ltd (Jiangsu, China). Briefly, the PCR products were then sub-cloned into a pAAV-CAG plasmid to produce the pAAV-CAG-AMPD3, pAAV-CAG-NT5C1A and pAAV-CAG-MEF2C plasmid. AAVs were subsequently generated using packaging plasmids pAAV-helper and pAAV2/9 together with pAAV-CAG- AMPD3 and pAAV-CAG-NT5C1A by Rongsen Gene Technology Co., Ltd (Jiangsu, China). AAV-AMPD3 and AAV-NT5C1A were diluted in 0.9% NaCl at 1 × 1013 virus particles/mL, and injected 30 μl intraperitoneally at postnatal day 3 (P3) and day 5 (P5) male WT and Mll4SET MKO mice (n = 5–8 mice per group). AAV-MEF2C was diluted in 0.9% NaCl at 1 × 1013 virus particles/mL and injected into 6-week-old male WT and Mll4SET MKO mice muscles (50 μl per GC muscle, n = 5 mice per group). AAV-GFP was used as a control. AAVs for in vivo knockdown of AMPD3 and NT5C1A were generated and provided by the Rongsen Gene Technology Co., Ltd (Jiangsu, China). The knockdown sequence: shAMPD3: 5’CCCTATGCTATGCCTGAGTAT-3’, 5’GCCCTATAGTTACTACCTGTA-3’, shNT5C1A: 5’-GTGCCCGATTCGCACGTATTT-3’, 5’-TCAACAGTATCAACCATTATG-3’, 5’-GTACCCTGATAGCGAAGATAT-3’. AAVs were diluted in 0.9% NaCl at 1 × 1013 virus particles/mL, and injected into 6-week-old male WT and Mll4SET MKO mice muscles (50 μl per GC muscle, n = 6 mice per group). AAV-shCtrl was used as a control.
Exercise stress test
Mice were acclimated (run for 9 min at 10 meters (m)/min followed by 1 min at 20 m/min) to the treadmill for 2 consecutive days prior to the experimental protocol. Four-month-old male WT and Mll4SET MKO mice under CD and HFD condition were run for 10 min at 10 m/min followed by a constant speed of 20 m/min until exhaustion33. Tail blood was taken after exercise and measured for lactate (Lactate Scout, Senelab, Germany) according to manufacturer’s instruction. To determine maximal sprint capacity, 2-month-old male WT and Mll4SET MKO mice were subjected to a high intensity exercise test consisting of an increasing speed every 2 min at 5° inclination until exhaustion. The increasing speeds used in the protocol were 10, 14, 18, 22, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, and 48 m/min.
Metabolic measurements in vivo
Mice were housed individually in metabolic cages at a 12-h light and dark cycle with free access to food and water using the CLAMS (Columbus Instruments). Four-month-old male WT and Mll4SET MKO mice (n = 11–15 mice per group) under HFD condition were acclimated in the metabolic cage for 1 day prior to the recording according to the instructions of the manufacturer. Food, energy expenditure, physical activity, VO2 and VCO2 were assessed simultaneously72.
Body composition analyses
Two-month-old and 4-month-old male WT and Mll4SET MKO mice (n = 6–10 mice per group) under CD and HFD condition body composition parameters including fat mass and lean tissue mass were determined via dual-energy X-ray absorptiometry (DEXA) using a Lunar PIXImus II densitometer (GE Healthcare) or nuclear magnetic resonance (NMR) with a MiniSpec LF50 (Bruker Optics), according to the manufacturer’s instructions73.
Glucose and insulin tolerance testing
Prior to studies, 4-month-old male WT and Mll4SET MKO mice (n = 6–10 mice per group) mice under CD and HFD condition were fasted overnight (GTT) or for 4 h (ITT). For GTT studies, mice were injected with 1.5 g/kg of D-glucose. For ITT, mice received an intraperitoneal injection of human regular insulin (Sigma-Aldrich) at a dose of 1 U/kg body weight. Blood glucose levels were determined at 0, 15, 30, 60, 90, and 120 min after challenge using a OneTouch ultramini glucose meter (OneTouch). Area under the curve (AUC) was defined as the difference between baseline glucose levels and the deflection caused by the glucose or insulin challenge. Total AUC was calculated using the trapezoidal rule. For assessment of insulin signaling, mice received an intraperitoneal injection of human regular insulin (Sigma-Aldrich) at a dose of 2 U/kg body weight.
Blood and tissue chemistry
After a 12-week high fat diet regimen, 4-month-old male WT and Mll4SET MKO mice under HFD condition were fasted overnight (16 h) beginning at 5 p.m. Blood samples were obtained for glucose and insulin measurements. Serum insulin levels were determined using an insulin ELISA kit (Millipore). Liver tissue (50 mg) was homogenized, and centrifuged supernatants were harvested. For muscle TG analysis, triglycerides were extracted from freeze-dried muscle in 2:1 chloroform:methanol and dried in air. The triglyceride was then saponified in 4% ethanol-KOH solution at 60 °C, and the glycerol concentration was measured fluorometrically. The TGs levels were determined with the Free Glycerol Reagent (F6428, Sigma-Aldrich) using glycerol (G7793, Sigma-Aldrich) as standard for calculation. For measurement of muscle glycogen, muscle tissue was acid hydrolyzed in 2 M HCl at 95 °C for 2 h, neutralized with an equal volume of 2 M NaOH, and centrifuged at 12,000 × g for 10 min. The liberated free-glycosyl units of the supernatant were determined using the glucose hexokinase kit (Wako, 298-65701) according to the manufacturer’s instructions.
Mitochondrial respiration studies
Mitochondrial respiration rates were measured in saponin-permeablized extensor digital longus muscle fibers with palmitoyl-L-carnitine chloride (PC) as a substrate74,75. In brief, the muscle fibers were separated and transferred to BIOPS buffer (7.23 mM K2EGTA, 2.77 mM CaK2EGTA, 20 mM imidazole, 20 mM taurine, 50 mM potassium 2-[N-morpholino]-ethanesulfonic acid, 0.5 mM dithiothreitol, 6.56 mM MgCl2, 5.7 mM ATP, and 14.3 mM phosphocreatine [PCr], pH 7.1). The muscle fibers bundles were then permeabilized with 50 μg/ml saponin in BIOPS solution. Measurement of oxygen consumption in permeabilized muscle fibers was performed in buffer Z (105 mM potassium 2-[N-morpholino]-ethanesulfonic acid, 30 mM KCl, 10 mM KH2PO4, 5 mM MgCl2, 5 mg/ml BSA, 1 mM EGTA, pH 7.4) at 37 °C and in the oxygen concentration range 220–150 nmol O2/ml in the respiration chambers of an Oxygraph 2 K (Oroboros Inc., Innsbruck, Austria). Following measurement of basal, palmitoyl-L-carnitine chloride (PC) (40 μM)/malate (5 mM) respiration, maximal (ADP-stimulated) respiration was determined by exposing the mitochondria to 4 mM ADP. Uncoupled respiration was evaluated following addition of oligomycin (1 μg/mL). Respiration rates were determined and normalized to tissue wet weight using Datlab 5 software (Oroboros Inc., Innsbruck, Austria), the data were expressed as “pmol O2 s−1mg wet weight−1”.
Mitochondrial DNA analyses
Mitochondrial DNA content was determined by SYBR Green analysis (Vazyme Biotech). The levels of NADH dehydrogenase subunit 1(mitochondrial DNA) were normalized to the levels of lipoprotein lipase (genomic DNA). The primer sequences are noted in Supplementary Table 3. All primers are ordered from Tsingke Biotechnology Co.,Ltd.
Histological analyses
Mouse muscle tissues were frozen in isopentane that had been cooled in liquid nitrogen. Notably, the gastrocnemius (GC) and plantaris of 5-month-old C57BL/6J male mice of the indicated genotypes under HFD condition were dissected intact and processed into 10-μm serial sections. GC muscles cross-sections were cut from the knee cut side in a Leica CM1850 cryostat at −20 °C and mounted on positively charged glass slides. Transverse sections collected from the widest part (mid-belly) of the GC muscle were used for histological comparison to keep consistent between different mice. For IF stains, the muscle fibers were stained with antibodies directed against MHC1 (BA-D5) and MHC2b (BF-F3) (MHC1, green; MHC2, red). For succinate dehydrogenase (SDH) histochemical staining, the frozen sections were thawed to room temperature and then incubated in a staining solution consisted of 100 mM sodium phosphate buffer (pH 7.6), 48 mM disodium succinate, 1 mM l-methoxyphenazine methosulfate (MPMS), 1.5 mM nitro blue tetrazolium (NBT), 5 mM EDTA, and 1 mM sodium azide for 10 min at 37 °C in the dark76. Livers of mice were embedded in Tissue-Tek OCT cryostat molds (Leica) and frozen at −80 °C. 10-μm-thick sections of livers were generated in a cryostat. Tissue sections were stained in 0.5% oil red O and counterstained with hematoxylin and eosin (Sigma-Aldrich) according to the standard protocol. For fiber type and SDH staining quantification in GC muscle, data were presented as percentage of positive fiber type/SDH staining myofibers per lateral head of GC to keep consistent between different mice.
RNA-seq studies
Transcriptomics analyses were performed using RNA-sequencing. Total RNA was isolated from the gastrocnemius muscle or liver of 5-month-old male Mll4SET MKO and WT control mice fed either CD or HFD using RNAiso Plus (Takara Bio). RNA-Seq using Illumina HiSeq 4000 was performed by the Beijing Novogene Bioinformatics Technology Co. Two independent samples per group were analyzed. Paired-end 150 nt reads were obtained from the same sequencing lane. The sequencing reads were then aligned to the UCSC mm10 genome assembly using TopHat, version 2.0.14, with the default parameters. Fragments per Kb of exon per million mapped reads (FPKM) were calculated using Cufflinks, version 2.2.1. The criteria for a regulated gene was a fold change greater than 1.5 (either direction) and a significant P value (<0.05). K-means cluster analysis (K = 4) was used to group differentially regulated genes in muscle. The numbers of regulated genes in muscle analysis are 252 (Cluster I), 480 (Cluster II), 784 (Cluster III) and 917 (Cluster IV), respectively (Fig. 4a). A total of 1534 genes were regulated in livers from HFD-fed Mll4SET MKO mice, with 573 up- and 961 down-regulated, respectively (Supplementary Fig. 3d). For pathway analysis, the filtered datasets were uploaded into DAVID Bioinformatics Resources 6.8 to review the biological pathways using the Functional Categories database. The GO analysis was used to interpret data, and the regulated terms ranked by P value are shown in Fig. 4b and Supplementary Fig. 7a, b. The volcano plot and heatmap analysis of regulated genes were generated by using R software, version 4.0.3, and the ggplot2/gplots package. The muscle and liver RNA-seq data have been deposited in the NCBI Gene Expression Omnibus and are accessible through GEO Series accession numbers GSE206506 and GSE206416.
RNA analyses
Total RNA was extracted from mouse skeletal muscle using Trizol reagent77. Briefly, total RNA was extracted from entire gastrocnemius muscle using RNAiso Plus (Takara Bio). Isolated total RNA integrity was electrophoretically verified by ethidium bromide staining. 1 μg total RNA samples were then reverse transcribed using the PrimeScript RT Reagent Kit with gDNA Eraser (Takara Bio). Real-time quantitative RT-PCR was performed using the ABI Prism Step-One system with Reagent Kit from Takara Bio. Specific oligonucleotide primers for target gene sequences are listed in Supplementary Table 3. All primers are ordered from Tsingke Biotechnology Co.,Ltd. Arbitrary units of target mRNA were corrected to the expression of 36b4.
ChIP, ChIP-qPCR, and ChIP-seq
Mouse gastrocnemius muscles from 2-month-old WT and Mll4SET MKO male mice were harvested, minced, and cross-linked in 2.5% formaldehyde for 15 min followed by quenching with 1/20 vol of 2.5 M glycine solution for 5 min and two washes with 1× PBS wash buffer78. Nuclear extracts were prepared by a Dounce homogenizer in ice-cold hypotonic buffer supplemented with protease inhibitors. Chromatin fragmentation was performed by sonication in lysis buffer (50 mM Tris-HCL at pH 8.0, 1%SDS, 10 mM EDTA) using a Bioruptor (Diagenode). An aliquot of chromatin was pre-cleared with protein G and immunoprecipitated with anti-MLL4 (ABE1867, Sigma-Aldrich), anti-MEF2C (ab211493, Abcam) antibodies. Following reversal of cross-linking, DNA was isolated using the standard Phenol-chloroform method. Immunoprecipitated DNA was analyzed by quantitative PCR using the Roche LightCycler 96 Instrument with Taq Pro Universal SYBR qPCR Master Mix from Vazyme Bio. The enrichment of genomic regions was calculated relative to input DNA using the standard curve method. Specific oligonucleotide primers for target regions are listed in Supplementary Table 3. For ChIP-seq, the precipitated DNA samples were then amplified according to ChIP Sequencing Sample Preparation Guide provided by Illumina. Deep sequencing using Illumina NovaSeq 6000 was performed by Beijing Novogene Bioinformatics Technology Co., Ltd.
ChIP-seq data processing
ChIP-seq data were processed by aligning the sequencing reads to the mm10 genome and subsequent peak calling33,78. Briefly, single end 50 nt reads were mapped to the mouse genome (UCSC mm10) using Bowtie2 (Version 2.2.5). Only the sequences uniquely mapped with no more than 1 mismatch were kept and used as valid reads. The peak caller program MACS2 (Version 2.1.1) was used to identify peaks with the following parameter settings: --keep-dup = 1, -B, --SPMR to generate signal pileup tracks in bedGraph format on a per million reads basis. Genome regions were associated to the gene with the nearest transcription start sites (TSS) from the UCSC genome browser. Differentially expressed genes identified from CD- and HFD-muscle RNA-seq data, respectively, were integrated with our previously published MLL4 ChIP-seq data (GSE138994)33 to define a set of genes (298 genes) directly regulated by MLL4. The MEF2 ChIP-seq dataset (GSE43223)79. Genome browser tracks of ChIP-seq data were visualized in IGV (Version 2.3.70).
Antibodies and immunoblotting studies
Antibodies directed against MHC1 (BA-D5, 1:200 dilution) and MHC2b (BF-F3, 1:200 dilution) were purchased from the Developmental Studies Hybridoma Bank; anti-MEF2C (ab211493, 1:500 dilution) was from Abcam; anti-Flag (#F1804, 1:1000 dilution) was from Sigma; AMPKα Thr172 (#2535, 1:1000 dilution), AMPKα (#5831, 1:1000 dilution), p-AKT Ser473 (#4060, 1:1000 dilution), p-AKT Thr308 (#13038, 1:1000 dilution), AKT (#9272, 1:1000 dilution) p-ACC Ser79 (#11818, 1:1000 dilution), ACC (#3676, 1:1000 dilution) were from Cell Signaling Technology; AMPD3 (#23997-1-AP, 1:1000 dilution), AMPD1 (#19780-1-AP, 1:1000 dilution), GAPDH (#60004-1-Ig, 1:1000 dilution), HSP90 (#13171-1-AP, 1:1000 dilution) were all from Proteintech; NT5C1A (#C15296, 1:1000 dilution) was from AssaybioTech; MLL4 (#ABE1867, 1:1000 dilution) and Sarcolipin (#ABT13, 1:1000 dilution) was from Millipore. Western blotting studies were performed74. The total protein concentration was measured by BCA assay using Pierce BCA Assay Kit Protocol (ThermoFischer Scientific). Equal total protein was loaded to each lane.
Cell culture and lentivirus infection
HEK293T (#CRL-3216) and C2C12 (#CRL-1772) Cells were obtained from the American Type Culture Collection and were cultured at 37 °C and 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS, 1000 U/ml penicillin and 100 mg/mL streptomycin. LHCN-M2 human skeletal muscle cells (T0756, abm) were obtained from Applied Biological Materials Inc. (BC, Canada) and cultured in DMEM/M199 medium (4:1) supplemented with 1000 U/mL penicillin, 100 mg/mL streptomycin, 15% (v/v) FBS, 20 mM HEPES, 30 ng/mL zinc sulfate, 1.4 µg/mL vitamin B12, 55 ng/mL dexamethasone, 2.5 ng/mL recombinant human hepatocyte growth factor, and 10 ng/mL basic FGF. For myocyte differentiation, LHCN-M2 and C2C12 cells were cultured with 2% horse-serum/DMEM differentiation medium. Lenti-shMLL4 (shMLL4: 5’TTCATCGAGTTGCGACATA) was used for mouse muscle cells, while lenti-shKMT2D/MLL4 (sh-KMT2D/MLL4: 5’CCTGAATTGAACAACAGTCTT) was used for human muscle cells, and lenti-shAMPD3 (shAMPD3: 5’GCCCTATAGTTACTACCTGTA) viruses were generated by co-transfection of 293T cells with the lentiviral vector and two packaging plasmids (psPAX2 and pMD2.G) at a ratio of 2:1.5:1. The virus-containing supernatants were collected at 48 h post-transfection. The filtered (0.45 μm) supernatants were then used to infect the C2C12 cells. Cells were then incubated in growth media with puromycin, 48–72 h post-selection, cells were induced to differentiation for 5 days.
Glucose consumption assay
Cells grown in 6-well plates were differentiated into myotubes for 5 days. Cells were then treated with fresh differentiation medium. At 0 h, 8 h, 16 h and 24 h post incubation, culture medium was collected from each well and frozen in −80 °C for future glucose measurements using Glucose assay kit (298-65701, Wako).
Statistical and reproducibility
All mouse and cell studies were analyzed by Student’s t test when two groups were compared. One-way ANOVA or two-way ANOVA coupled to a Fisher’s least significant difference (LSD) post-hoc test or a Tukey’s post-hoc test was used when more than two groups were compared. Statistical analyses in human studies were performed using Graphpad Prism 8.0 or SPSS 25.0 software. Gene expression levels in human studies were analyzed using Mann-Whitney test or Spearman correlation test. Significant differences were defined as P < 0.05. No statistical methods were used to predetermine sample sizes, and sample sizes are explicitly stated in the figure legends. No data were excluded from the analyses. The experiments were not randomized. The Investigators were not blinded to allocation during experiments and outcome assessment. All data points were used in statistical analyses. Data represent the mean ± SEM, with a statistically significant difference defined as a value of P < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq and ChIP-seq data reported in this paper have been deposited in the NCBI Gene Expression Omnibus. The human muscle RNA-seq data could be accessible through accession number GSE97084. The WT and Mll4SET MKO mice muscle RNA-seq data could be accessible through accession number GSE206506. The WT and Mll4SET MKO mice liver RNA-seq data could be accessible through accession number GSE206416. The MLL4 ChIP-seq data could be accessible through accession number GSE138994. The MEF2 ChIP-seq data could be accessible through accession number GSE43223. All other data supporting the findings of this study are available with the article. Source data are provided with this paper.
Change history
04 February 2026
In the version of Supplementary Information initially published alongside this article, Supplementary Fig. 3a contained an incorrect representative gross liver image panel (“CD WT” and “CD Mll4SET MKO”) introduced during figure assembly. The panel has been replaced with the correct representative images from the Mll4SET MKO cohort used in this study. This update is limited to the Supplementary Information figure presentation and does not affect any underlying data, analyses, or the conclusions of the article.
References
Blüher, M. Obesity: global epidemiology and pathogenesis. Nat. Rev. Endocrinol. 15, 288–298 (2019).
Baskin, K. K., Winders, B. R. & Olson, E. N. Muscle as a “mediator” of systemic metabolism. Cell Metab. 21, 237–248 (2015).
Goodpaster, B. H. & Sparks, L. M. Metabolic flexibility in health and disease. Cell Metab. 25, 1027–1036 (2017).
Egan, B. & Zierath, J. R. Exercise metabolism and the molecular regulation of skeletal muscle adaptation. Cell Metab. 17, 162–184 (2013).
Gan, Z., Fu, T., Kelly, D. P. & Vega, R. B. Skeletal muscle mitochondrial remodeling in exercise and diseases. Cell Res. 28, 969–980 (2018).
Lowell, B. B. & Shulman, G. I. Mitochondrial dysfunction and type 2 diabetes. Science 307, 384–387 (2005).
Muoio, D. M. & Neufer, P. D. Lipid-induced mitochondrial stress and insulin action in muscle. Cell Metab. 15, 595–605 (2012).
Hesselink, M. K., Schrauwen-Hinderling, V. & Schrauwen, P. Skeletal muscle mitochondria as a target to prevent or treat type 2 diabetes mellitus. Nat. Rev. Endocrinol. 12, 633–645 (2016).
Neufer, P. D. et al. Understanding the cellular and molecular mechanisms of physical activity-induced health benefits. Cell Metab. 22, 4–11 (2015).
Wang, Y. X. et al. Regulation of muscle fiber type and running endurance by PPARdelta. PLoS Biol. 2, e294 (2004).
Gan, Z. et al. The nuclear receptor PPARβ/δ programs muscle glucose metabolism in cooperation with AMPK and MEF2. Genes Dev. 25, 2619–2630 (2011).
Gan, Z. et al. Nuclear receptor/microRNA circuitry links muscle fiber type to energy metabolism. J. Clin. Investig. 123, 2564–2575 (2013).
Narkar, V. A. et al. Exercise and PGC-1α-independent synchronization of type I muscle metabolism and vasculature by ERRγ. Cell Metab. 13, 283–293 (2011).
Potthoff, M. J. et al. Histone deacetylase degradation and MEF2 activation promote the formation of slow-twitch myofibers. J. Clin. Investig. 117, 2459–2467 (2007).
Lin, J. et al. Transcriptional co-activator PGC-1 alpha drives the formation of slow-twitch muscle fibres. Nature 418, 797–801 (2002).
Arany, Z. et al. The transcriptional coactivator PGC-1beta drives the formation of oxidative type IIX fibers in skeletal muscle. Cell Metab. 5, 35–46 (2007).
Yamamoto, H. et al. NCoR1 is a conserved physiological modulator of muscle mass and oxidative function. Cell 147, 827–839 (2011).
Hong, S. et al. Dissociation of muscle insulin sensitivity from exercise endurance in mice by HDAC3 depletion. Nat. Med. 23, 223–234 (2017).
Hardie, D. G. AMP-activated/SNF1 protein kinases: conserved guardians of cellular energy. Nat. Rev. Mol. Cell Biol. 8, 774–785 (2007).
Steinberg, G. R. & Kemp, B. E. AMPK in health and disease. Physiol. Rev. 89, 1025–1078 (2009).
Hardie, D. G., Schaffer, B. E. & Brunet, A. AMPK: an energy-sensing pathway with multiple inputs and outputs. Trends Cell Biol. 26, 190–201 (2016).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why?. Mol. Cell 49, 825–837 (2013).
Kang, S., Tsai, L. T. & Rosen, E. D. Nuclear mechanisms of insulin resistance. Trends Cell Biol. 26, 341–351 (2016).
Heinz, S., Romanoski, C. E., Benner, C. & Glass, C. K. The selection and function of cell type-specific enhancers. Nat. Rev. Mol. Cell Biol. 16, 144–154 (2015).
Lee, J. E. et al. H3K4 mono- and di-methyltransferase MLL4 is required for enhancer activation during cell differentiation. eLife 2, e01503 (2013).
Wang, C. et al. Enhancer priming by H3K4 methyltransferase MLL4 controls cell fate transition. Proc. Natl. Acad. Sci. USA 113, 11871–11876 (2016).
Park, Y.-K. et al. Interplay of BAF and MLL4 promotes cell type-specific enhancer activation. Nat. Commun. 12, 1630 (2021).
Cao, K. et al. An Mll4/COMPASS-Lsd1 epigenetic axis governs enhancer function and pluripotency transition in embryonic stem cells. Sci. Adv. 4, eaap8747 (2018).
Hu, D. et al. The MLL3/MLL4 branches of the COMPASS family function as major histone H3K4 monomethylases at enhancers. Mol. Cell. Biol. 33, 4745–4754 (2013).
Lai, B. et al. MLL3/MLL4 are required for CBP/p300 binding on enhancers and super-enhancer formation in brown adipogenesis. Nucleic Acids Res. 45, 6388–6403 (2017).
Wang, S. P. et al. A UTX-MLL4-p300 transcriptional regulatory network coordinately shapes active enhancer landscapes for eliciting transcription. Mol. Cell 67, 308–321.e306 (2017).
Ang, S. Y. et al. KMT2D regulates specific programs in heart development via histone H3 lysine 4 di-methylation. Developement 143, 810–821 (2016).
Liu, L. et al. Histone methyltransferase MLL4 controls myofiber identity and muscle performance through MEF2 interaction. J. Clin. Investig. 130, 4710–4725 (2020).
Kim, D. H. et al. Critical roles of the histone Methyltransferase MLL4/KMT2D in murine hepatic steatosis directed by ABL1 and PPARγ2. Cell Rep. 17, 1671–1682 (2016).
Rao, R. C. & Dou, Y. Hijacked in cancer: the KMT2 (MLL) family of methyltransferases. Nat. Rev. Cancer 15, 334–346 (2015).
Ng, S. B. et al. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat. Genet. 42, 790–793 (2010).
Zaidi, S. et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 498, 220–223 (2013).
Amoasii, L. et al. A MED13-dependent skeletal muscle gene program controls systemic glucose homeostasis and hepatic metabolism. Genes Dev. 30, 434–446 (2016).
Izumiya, Y. et al. Fast/Glycolytic muscle fiber growth reduces fat mass and improves metabolic parameters in obese mice. Cell Metab. 7, 159–172 (2008).
Kulkarni, S. S. et al. Suppression of 5’-nucleotidase enzymes promotes AMP-activated protein kinase (AMPK) phosphorylation and metabolism in human and mouse skeletal muscle. J. Biol. Chem. 286, 34567–34574 (2011).
Mi, Z. et al. Super-enhancer-driven metabolic reprogramming promotes cystogenesis in autosomal dominant polycystic kidney disease. Nat. Metab. 2, 717–731 (2020).
Mahnke-Zizelman, D. K., Tullson, P. C. & Sabina, R. L. Novel aspects of tetramer assembly and N-terminal domain structure and function are revealed by recombinant expression of human AMP deaminase isoforms. J. Biol. Chem. 273, 35118–35125 (1998).
Tatekoshi, Y. et al. Translational regulation by miR-301b upregulates AMP deaminase in diabetic hearts. J. Mol. Cell. Cardiol. 119, 138–146 (2018).
Bal, N. C. et al. Sarcolipin is a newly identified regulator of muscle-based thermogenesis in mammals. Nat. Med. 18, 1575–1579 (2012).
Wang, D. et al. GDF15 promotes weight loss by enhancing energy expenditure in muscle. Nature 619, 143–150 (2023).
Sharma, A. K., Khandelwal, R. & Wolfrum, C. Futile lipid cycling: from biochemistry to physiology. Nat. Metab. 6, 808–824 (2024).
Robinson, M. M. et al. Enhanced protein translation underlies improved metabolic and physical adaptations to different exercise training modes in young and old humans. Cell Metab. 25, 581–592 (2017).
Zhu, C.-H. et al. Cellular senescence in human myoblasts is overcome by human telomerase reverse transcriptase and cyclin-dependent kinase 4: consequences in aging muscle and therapeutic strategies for muscular dystrophies. Aging Cell 6, 515–523 (2007).
Williamson, A. et al. Genome-wide association study and functional characterization identifies candidate genes for insulin-stimulated glucose uptake. Nat. Genet. 55, 973–983 (2023).
Singh, S. et al. Targeting KDM4 for treating PAX3-FOXO1-driven alveolar rhabdomyosarcoma. Sci. Transl. Med. 14, eabq2096 (2022).
Frampton, J., Murphy, K. G., Frost, G. & Chambers, E. S. Short-chain fatty acids as potential regulators of skeletal muscle metabolism and function. Nat. Metab. 2, 840–848 (2020).
Agarwal, R. P. & Parks, R. E. Potent inhibition of muscle 5’-AMP deaminase by the nucleoside antibiotics coformycin and deoxycoformycin. Biochem. Pharmacol. 26, 663–666 (1977).
Grever, M. R. & Lozanski, G. Modern strategies for hairy cell leukemia. J. Clin. Oncol. 29, 583–590 (2011).
Saven, A. & Piro, L. Newer purine analogues for the treatment of hairy-cell leukemia. N. Engl. J. Med. 330, 691–697 (1994).
Piunti, A. & Shilatifard, A. Epigenetic balance of gene expression by Polycomb and COMPASS families. Science 352, aad9780 (2016).
Lee, K. Y. et al. Tbx15 controls skeletal muscle fibre-type determination and muscle metabolism. Nat. Commun. 6, 8054 (2015).
Amoasii, L. et al. NURR1 activation in skeletal muscle controls systemic energy homeostasis. Proc. Natl. Acad. Sci. USA 116, 11299–11308 (2019).
Kim, J. et al. UBE3A suppresses overnutrition-induced expression of the steatosis target genes of MLL4 by degrading MLL4. Hepatology 69, 1122–1134 (2019).
Bergeron, R. et al. Chronic activation of AMP kinase results in NRF-1 activation and mitochondrial biogenesis. Am. J. Physiol. Endocrinol. Metab. 281, E1340–E1346 (2001).
Jäger, S., Handschin, C., St-Pierre, J. & Spiegelman, B. M. AMP-activated protein kinase (AMPK) action in skeletal muscle via direct phosphorylation of PGC-1alpha. Proc. Natl. Acad. Sci. USA 104, 12017–12022 (2007).
Kviklyte, S. et al. Effects of genetic deletion of soluble 5′-nucleotidases NT5C1A and NT5C2 on AMPK activation and nucleotide levels in contracting mouse skeletal muscles. Am. J. Physiol.-Endocrinol. Metab. 313, E48–E62 (2017).
Johanns, M. et al. Genetic deletion of soluble 5′-nucleotidase II reduces body weight gain and insulin resistance induced by a high-fat diet. Mol. Genet. Metab. 126, 377–387 (2019).
Cokorinos, E. C. et al. Activation of skeletal muscle AMPK promotes glucose disposal and glucose lowering in non-human primates and mice. Cell Metab. 25, 1147–1159.e1110 (2017).
Li, J. et al. Integration of lipidomics and transcriptomics unravels aberrant lipid metabolism and defines cholesteryl oleate as potential biomarker of prostate cancer. Sci. Rep. 6, 20984 (2016).
Yan, M. et al. Metabolomics profiling of metformin-mediated metabolic reprogramming bypassing AMPKα. Metab.: Clin. Exp. 91, 18–29 (2019).
Zhang, C.-S. et al. Fructose-1,6-bisphosphate and aldolase mediate glucose sensing by AMPK. Nature 548, 112–116 (2017).
Zeng, J. et al. Comprehensive profiling by non-targeted stable isotope tracing capillary electrophoresis-mass spectrometry: a new tool complementing metabolomic analyses of polar metabolites. Chem. – A Eur. J. 25, 5427–5432 (2019).
Zeng, J. et al. Metabolomics study of hepatocellular carcinoma: discovery and validation of serum potential biomarkers by using capillary electrophoresis-mass spectrometry. J. Proteome Res. 13, 3420–3431 (2014).
Ouyang, Y. et al. A high throughput metabolomics method and its application in female serum samples in a normal menstrual cycle based on liquid chromatography-mass spectrometry. Talanta 185, 483–490 (2018).
Xuan, Q. et al. Development of a high coverage pseudotargeted lipidomics method based on ultra-high performance liquid chromatography–mass spectrometry. Anal. Chem. 90, 7608–7616 (2018).
Mu, J., Brozinick, J. T. Jr., Valladares, O., Bucan, M. & Birnbaum, M. J. A role for AMP-activated protein kinase in contraction- and hypoxia-regulated glucose transport in skeletal muscle. Mol. Cell 7, 1085–1094 (2001).
Yin, Y. et al. FNIP1 regulates adipocyte browning and systemic glucose homeostasis in mice by shaping intracellular calcium dynamics. J. Exp. Med. 219, e20212491 (2022).
Fu, T. et al. Mitophagy directs muscle-adipose crosstalk to alleviate dietary obesity. Cell Rep. 23, 1357–1372 (2018).
Xu, Z. et al. Disuse-associated loss of the protease LONP1 in muscle impairs mitochondrial function and causes reduced skeletal muscle mass and strength. Nat. Commun. 13, 894 (2022).
Guo, Q. et al. Mitochondrial proteostasis stress in muscle drives a long-range protective response to alleviate dietary obesity independently of ATF4. Sci. Adv. 8, eabo0340 (2022).
Liu, J. et al. Coupling of mitochondrial function and skeletal muscle fiber type by a miR-499/Fnip1/AMPK circuit. EMBO Mol. Med. 8, 1212–1228 (2016).
Xiao, L. et al. AMPK-dependent and -independent coordination of mitochondrial function and muscle fiber type by FNIP1. PLoS Genet. 17, e1009488 (2021).
Liu, L. et al. Coupling of COPII vesicle trafficking to nutrient availability by the IRE1α-XBP1s axis. Proc. Natl. Acad. Sci. USA 116, 11776–11785 (2019).
Sebastian, S. et al. Tissue-specific splicing of a ubiquitously expressed transcription factor is essential for muscle differentiation. Genes Dev. 27, 1247–1259 (2013).
Acknowledgements
We thank Dr. Yong Liu (Wuhan University) for critical reading the manuscript. This work was supported by grants from National Natural Science Foundation of China (No. 32425019, 92457302, 31922033, 91857105, 32471226, 32071136, and 32300970) to Z.G., T.F., and Y.Y., the Ministry of Science and Technology of China (National Key R&D Program of China 2022YFA0806000) to Z.G., Noncommunicable Chronic Diseases-National Science and Technology Major Project 2024ZD0530100 to T.F., Fundamental Research Funds for the Central Universities Grant No. KG202503, 021414380533, 021414380517, 021414380511, 021414380529, and 021414380524 (to T.F. and Z.G.). This project was also funded by the China Postdoctoral Science Foundation Grant No. 2023 M731633 (to Y.Y.).
Author information
Authors and Affiliations
Contributions
L.Y., L.L., and W.W. contributed equally to this work. L.Y., L.L., and W.W. designed and performed most of the experiments, analyzed the data, and wrote the manuscript. C.D., A.A., T.F., D.Z., Z.X., T.F., Q.G., Z.Z., P.S, G.S., L.X., Y.Y., Z.S., Y.M., W.S., J.L., and J.X. participated in the collection and analysis of the data. M.Z., K.G., and KS.N. contributed reagents, samples and provided scientific insight and discussion. P.H. and Z.G. conceptualized, interpreted the experiments, and wrote the manuscript. Z.G. supervised the work. All authors reviewed and contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, L., Liu, L., Wang, W. et al. Enhancer regulator MLL4 controls skeletal muscle metabolic efficiency by limiting AMPK-mediated fuel catabolism. Nat Commun 16, 11644 (2025). https://doi.org/10.1038/s41467-025-66684-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66684-x