Abstract
Radiofrequency ablation (RFA) has been one of the most promising local treatments for colorectal liver metastases. However, tumor progression after RFA still severely affects patient prognosis, and its molecular mechanisms remain to be further explored. Here, we show that heat-stressed residual tumor cells following insufficient RFA (iRFA) can lead to lung metastases. Specific Cd177hiPad4hi neutrophils are found as the main pro-tumor phenotype in iRFA conditions, and neutrophil extracellular traps are produced to accelerate lung metastasis via the MEK/ERK signaling pathway. Mechanistically, PPARγ-triglyceride (TG) synthesis in residual tumor cells activates the P38 signaling pathway and CXCL5 secretion, contributing to Cd177hiPad4hi neutrophil infiltration. Furthermore, the specific CD36hi-EM (epithelial-mesenchymal) - circulating tumor cells (CTCs) with activated lipid metabolism characteristics are found to assist in early identification of residual tumor cells. This study not only provides specific targets for improving the efficacy of iRFA but also realizes its early identification and intervention.
Similar content being viewed by others
Introduction
Currently, over 50% of colorectal cancer (CRC) patients have colorectal liver metastases (CRLM) at diagnosis or develop CRLM during tumor progression1. Only 10–20% of patients are suitable for surgical resection2. To date, local ablative treatment has been the alternative measures for most CRLM patients who are not surgical candidates, especially radiofrequency ablation (RFA) considered the most promising technique for complete tumor destruction3. RFA relies on high-frequency electrical currents (400-500 kHz) to induce oscillation and friction among ions within the tissue, resulting in heat generation. This heat is conducted from the interior to the exterior of the tissue. When the temperature exceeds 50 °C, protein denaturation occurs; as the temperature continues to rise to between 70 °C and 100 °C, coagulative necrosis and dehydration of the tissue occur4.
However, 60–80% of recurrences are observed within the 5-year follow-up period after RFA5. Residual tumor cells in the periablational margin, defined as insufficient RFA (iRFA), are mainly responsible for tumor progression6. Influence from tumor microenvironment (TME), especially an inflammatory microenvironment, has also been implicated recently. iRFA-induced secretion of inflammatory factors, such as CCL2, TGFβ and IL-6, mediates the recruitment of tumor-associated macrophages and myeloid-derived suppressor cells (MDSCs) to form an immunosuppressive TME, thereby contributing to tumor progression7,8. A salvage alternative for residual tumors after iRFA is still challenging.
Some tumors secrete inflammatory cytokines, such as C-X-C motif chemokine ligand (CXCL)5, CXCL2, and CXCL1, promoting the recruitment of neutrophils9. Neutrophils inherently have environmentally driven plasticity and play a dual tumor-promoting and tumor-suppressing role10. It is necessary to further analyze the characteristic phenotypes and mechanisms of neutrophils in different disease processes, which still unclarified in iRFA.
How to distinguish iRFA immediately is another dilemma. Circulating tumor cells (CTCs) are the seeds of tumor metastasis and vital for prognostic and therapeutic monitoring11. Specific CTC subgroups are responsible for tumor metastasis12. However, no studies have been conducted on whether CTC subgroup identification can realize residual tumor cells monitoring in iRFA.
In this study, we discovered the residual tumor cells as the origin of lung metastases. The residual tumor cells with lipid metabolism adaption induced Cd177hiPad4hi neutrophils release neutrophil extracellular traps (NETs) and promote tumor metastasis. Meanwhile, CD36hi-EM-CTC cells were distinguished to assist in the early prediction of iRFA.
Results
Residual tumor cells after iRFA accelerate CRLM lung metastasis
One hundred and sixty-seven CRLM patients in Nanfang Hospital, Southern Medical University were enrolled in a retrospective case-controlled study. Among them, 41 patients were found to have iRFA, which was defined as observable residual tumor within 4 months after RFA according to previous studies7. Forty-six patients who achieved complete RFA (cRFA) without local residual tumor and eighty patients who underwent hepatic resection during the same study period were selected (Supplementary Data 1). A significantly shorter time to new extrahepatic metastasis was observed in the iRFA group compared with the cRFA and hepatic resection groups (Fig. 1A), indicating rapid extrahepatic progression after iRFA. Then, iRFA-CT26Luc/eGFP BALB/c and iRFA-MC38 C57BL/6 mouse models were established (Fig. 1B and Supplementary Fig. 1A, B). The in vivo imaging showed residual tumor fluorescence in the iRFA group and a loss of tumor fluorescence in the cRFA group, demonstrating the success of both iRFA and cRFA treatments (Supplementary Fig. 1C). A dynamic observation showed that mice treated with iRFA were more likely to develop lung metastasis (Fig. 1C). The increased number and area of lung metastases, as well as lung weight in the iRFA group, further confirmed that iRFA accelerated lung metastasis of CRLM, while cRFA could completely inhibit tumor progression (Fig. 1D, E and Supplementary Fig. 1D–F). Significantly shorter OS duration was also observed in iRFA-CT26Luc/eGFP BALB/c mice treated with iRFA compared to those treated with cRFA or subjected to sham operation (Fig. 1F). Similar results were obtained in C57BL/6 mice (Supplementary Fig. 1G–I).
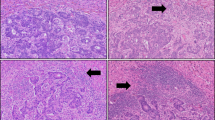
A Kaplan–Meier curves for time to new metastasis in CRLM patients (iRFA: n = 41; cRFA: n = 46; hepatic resection: n = 80). Log-rank test. B Study design of the iRFA-CT26Luc/eGFP BALB/c and iRFA-MC38 C57BL/6 mouse models. C Bioluminescence imaging of BALB/C mice treated with sham operation (Ctrl), iRFA, or cRFA of the implanted CT26Luc/eGFP tumors. Red and blue represent high and low fluorescence intensity, respectively. D Lung tissues from Ctrl and iRFA CT26Luc/eGFP BALB/c mice 14 days post-treatment (n = 8 mice). E (Left) HE staining of dissected lung tissues. Scale bars, 1 mm and 100 μm. (Right) The proportion of lung metastasis (n = 8 mice). Unpaired two-sided t-test. F Kaplan–Meier curves of OS (Ctrl: n = 27 mice, iRFA: n = 18 mice, cRFA: n = 23 mice). Log-rank test. Ctrl vs. iRFA: P = 0.001, Ctrl vs. cRFA: P = 0.029, and iRFA vs. cRFA: P = 0.001. G Study design of iRFA-CT26mCherry BALB/c mouse model (Triangles: sample collection time points). Representative microphotographs showed mCherry staining of lung tumors on days 0, 1, and 3 post-treatment (Arrows: mCherry+ tumor cells). Scale bars, 100 μm. H Lung tissues from CT26R-mCherry+L-GFP BALB/c mice on days 3 and 5 post-treatment (Left-iRFA or Right-iRFA). GFP (green) and mCherry (red) detected via immunofluorescence (Arrows: GFP+ or mCherry+ tumor cells). Scale bars, 100 or 10 μm. I Liver and lung tissues from the iRFA-resection or non-resection mice 14 days post-iRFA (n = 3 mice). J (Left) HE staining of dissected lung tissues. Scale bars, 1 mm. (Right) The proportion of lung metastasis area (n = 9 mice). P < 0.001, ANOVA. K Lung tissues from RFANormal and iRFATumor mice 14 days post-treatment. Lung weight shown on the right (Normal: n = 8 mice, Tumor: n = 7 mice), two-sided Mann-Whitney U test. L (Left) HE staining of lung tissues from the RFANormal and iRFATumor mice. Scale bars, 1 mm and 250 μm. (Right) The proportion of lung metastasis area (Normal: n = 8 mice, Tumor: n = 7 mice). Unpaired two-sided t-test. Data are presented as mean values ± SD. Source data are provided as a Source Data file.
Although promoted lung metastasis was observed after iRFA treatment, the origin remains unclear. By utilizing GFP and mCherry in tumor cells, we aimed to trace these cells through IF and differentiate the origins of lung metastasis. Firstly, the iRFA-CT26mCherry mice model was constructed, and mCherry-positive CT26 cells that formed lung metastases were only present on day 3 in the iRFA group, but not before RFA treatment (Fig. 1G and Supplementary Fig. 1J). Next, CT26mCherry cells were implanted into the right liver lobe, and CT26Luc/eGFP cells were implanted into the left lobe (CT26R-mCherry+L-GFP) to establish a bilateral liver lobe tumor-implanted mouse model (Supplementary Fig. 1K). iRFA was then performed for tumors in the left (Left-iRFA) or right (Right-iRFA) lobe. CT26Luc/eGFP cells were exclusively detected in lung tissues of the Left-iRFA group, whereas CT26mCherry cells were solely identified in lung tissues of the Right-iRFA group (Fig. 1H and Supplementary Fig. 1L), indicating that the pulmonary colonized cells originated from the tumor cells within the ablated regions. To minimize the influence of different ablated regions, iRFA-CT26L-mCherry+R-GFP and iRFA-CT26R-mCherry+L-GFP mouse models with Left-iRFA were constructed (Supplementary Fig. 1M). Notably, mCherry-positive lung metastases were only observed in the CT26L-mCherry+R-GFP mouse model subjected to Left-iRFA. These findings suggest that lung metastasis is related to the iRFA-influenced tumor cells.
Additionally, to verify whether the metastatic tumor cells originated from the escaped tumor cells during RFA treatment or from the residual tumor cells after iRFA treatment, we constructed the iRFA-Resection-CT26 BALB/c mice model (Supplementary Fig. 1N). Compared to the non-resection group, both day 1 and day 3 resection groups showed inhibited lung metastasis (Fig. 1I, J). Most significantly, lung metastases were almost completely suppressed in the day 1 resection group, emphasizing the residual tumor cells as the culprits of lung metastasis in iRFA. Finally, RFANormal and iRFATumor mouse models were established (Supplementary Fig. 1O). Lung metastasis occurred in iRFATumor but not in RFANormal (Fig. 1K, L), suggesting that the direct thermal stimulation by RFA or changes in the local microenvironment may result in lung metastasis. Overall, lung metastasis following iRFA originates from residual tumor cells.
Neutrophils act as “accomplices” in CRLM lung metastasis after iRFA
Bulk RNA-seq analysis was performed on the untreated and iRFA-treated CT26 tumors, showing 718 significantly changed genes (Supplementary Fig. 2A). Further Gene Ontology (GO) enrichment analysis revealed notably activated myeloid cell signatures, such as neutrophil migration, leukocyte migration, and neutrophil chemotaxis (Fig. 2A). Among the cellular composition, neutrophils were significantly increased in iRFA (Supplementary Fig. 2B).
A GO analysis of DEGs: iRFA-treated vs. untreated CT26 tumors (n = 3 mice/group). Hypergeometric tests. FDR correction. B Flow cytometric analysis and quantification of Cd11b+Ly6G+ cells in iRFA and Ctrl tumors on days 1 and 3 post-treatment (gated on CD45+ cells) (n = 5 mice). Unpaired two-sided t-test. C Ly6G detection in liver tumor tissues from Ctrl and iRFA mice 14 days post-treatment (T: tumor area, N: necrotic area). Scale bars, 20 μm. Ly6G+ cell counts shown on the right (n = 8 mice). Unpaired two-sided t-test. D Cd45, Cd11b, and S100a8 detection in liver tumor tissues from Ctrl and iRFA mice 14 days post-treatment. Scale bars, 50 μm and 10 μm. Quantification of S100a8+Cd11b+Cd45+ cells shown on the right (n = 10 mice). Unpaired two-sided t-test. E Study design of iRFA-CT26 mouse model treated with anti-Ly6G or IgG antibody (Circles: drug administration, Triangle: iRFA or Ctrl treatment). Cd45, Cd11b, and S100a8 detection in liver tumor tissues 14 days post-treatment. Scale bars, 50 μm. F Lung tissues from Ctrl and iRFA mice treated with anti-Ly6G or IgG antibody 14 days post-treatment (Circles: lung metastases). Lung weight shown on the right (n = 9 mice). P < 0.001, ANOVA. G scRNA-seq with uniform manifold approximation and projection (UMAP) plot of four cell clusters identified among neutrophils comparing the iRFA and Ctrl groups (n = 3 mice). H Expression of marker genes in the four clusters. I Bar chart showing identified neutrophil clusters in iRFA and Ctrl tumors. J Violin plots of identified neutrophil clusters characterized by functional gene expression. K Violin plot of gMDSC score in the clusters. N1: n = 593 cells, N2: n = 467 cells, N3: n = 273 cells, and N4: n = 125 cells. Box center line, median; box limits, upper and lower quartiles; whiskers, 1.5× interquartile range. L Visualization of neutrophil single-cell clades. Data are presented as mean values ± SD. Source data are provided as a Source Data file.
A significant increase in Cd45-positive cells was observed in the iRFA group (Supplementary Fig. 2C). There were no significant differences in the infiltration of lymphocytes, macrophages, and monocytes between groups (Supplementary Fig. 2D–F). Next, neutrophils were selectively separated, and a significant increase in neutrophils was verified in liver tumor tissues on days 1, 3, and 5 after iRFA treatment (Fig. 2B and Supplementary Fig. 2G, H). The increased accumulation of neutrophils in iRFA tumors was further confirmed by IF (Fig. 2C, D). Then, neutrophils in lung tissues, bone marrow, and peripheral blood were also analyzed. A slight increase of neutrophils was observed on day 5 in the lung tissue and on day 3 in bone marrow (Supplementary Fig. 2I, J). While neutrophils were increased on both day 1 and 3 in peripheral blood (Supplementary Fig. 2K). These data illustrate that neutrophil increases represent the most significant change in the TME treated with iRFA.
Whether the lung metastasis of CRLM after iRFA treatment is associated with neutrophils still needed further verification. Anti-Ly6G antibody was administered to deplete neutrophils in vivo (Fig. 2E). As confirmed by IF and flow cytometry, neutrophil depletion was achieved using an anti-Ly6G antibody (Fig. 2E and Supplementary Fig. 3A–E). Lung metastasis was significantly inhibited when neutrophils were depleted (Fig. 2F and Supplementary Fig. 3F). The results also showed that the apoptosis of neutrophils infiltrating the tumor tissues in the iRFA group was indeed significantly reduced (Supplementary Fig. 3G). Therefore, neutrophils likely contribute to lung metastasis in CRLM following iRFA.
Single-cell RNA sequencing (scRNA-seq) was performed on Cd45-positive cells from the iRFA-related tumor tissue to further distinguish the neutrophil phenotype. Neutrophils and the other cell types (including fibroblasts, T cells, macrophages, B cells, dendritic cells, endothelial cells, and NK cells) were identified (Supplementary Fig. 3H–K). The findings further supported that neutrophils were increased in iRFA (4.47% vs. 12.89%, Supplementary Fig. 3I), and four main clusters of neutrophils were distinguished (Fig. 2G, H). Cd177 clearly separated two distinct neutrophil populations: clusters N1 and N2 demonstrated low Cd177 expression, while clusters N3 and N4 showed high Cd177 expression. CD177 is recognized as the second specific antigen of human neutrophils13. Previous study has demonstrated that neutrophils can be categorized into two subgroups based on CD177 expression14. N3 and N4 phenotypes can be further distinguished by Mki67. Cluster N3 experienced the most significant increase in iRFA (Fig. 2I). Regarding the functional gene characterization (Fig. 2J), cluster N3 expressed Padi4, while cluster N4 expressed Elane and Mpo, both of which are required for metastasis-related NET formation15,16. Gene signatures from granulocytic MDSCs17 were enriched in cluster N3 (Fig. 2K). Therefore, a cluster of neutrophils, Cd177hiPad4hi neutrophils (N3), was discovered in iRFA. These findings indicate a pro-tumor functionality of cluster N3 in iRFA. Developmental trajectory analysis revealed a sequential differentiation path from N4 to N3, then to N2, and finally to N1 (Fig. 2L).
NETs drive Cd177hiPad4hi neutrophil-facilitated CRLM lung metastasis
To further confirm the presence of N3 neutrophils in vivo, Cd177hi neutrophils were isolated by flow cytometry and were found to be significantly elevated in iRFA tumors (Fig. 3A). More infiltration of Pad4+Cd177+S100a8+ neutrophils was also confirmed (Fig. 3B). Cd177low neutrophils (N1 and N2), Cd177hiMki67- neutrophils (N3), and Cd177hiMki67+ neutrophils (N4) were distinguished by flow cytometry based on Cd177 and Mki67 expression. N3 neutrophils showed high Pad4 expression, suggesting the phenotypic characteristic of Cd177hiPad4hiMki67- neutrophils (P < 0.001; Fig. 3C). Meanwhile, the expression of NET marker protein citrullinated histone-H3 (cit-H3)18 was also significantly increased in N3 neutrophils (P = 0.003; Fig. 3C), indicating that Cd177hiPad4hi neutrophils might function through forming NETs.
A Flow cytometric analysis and quantification of Cd177hi neutrophils in liver tumor tissues dissected from iRFA and Ctrl mice (gated on Cd11b+Ly6G+ cells, n = 4 mice). Unpaired two-sided t-text. B Immunofluorescence detection of Cd177, Pad4, and S100a8 in liver tumors. Scale bars, 50 μm and 10 μm. Quantification of Pad4+Cd177+S100a8+ cells shown on the right (n = 15 mice). Unpaired two-sided t-test. C Western blotting analysis in Cd177-, Cd177+Mki67-, and Cd177+Mki67+ neutrophils. Quantification shown on the right (n = 3 biological replicates). ANOVA. D ELISA-detected MPO-dsDNA levels in neutrophils (n = 3 biological replicates). P < 0.001, ANOVA. E Transwell assay assessing CT26 cell invasion in co-culture with PBS- or G484-treated neutrophils. Scale bars, 200 μm. F dsDNA levels in peripheral blood from Ctrl and iRFA mice 3 days post-treatment (n = 5 mice). Unpaired two-sided t-test. G (Left) Immunofluorescence detection of Ly6G and cit-H3 in liver tumors. Scale bars, 20 μm. (Right) Quantification of the cit-H3+ area (n = 15 mice), two-sided Mann-Whitney U test. H Study design of iRFA-CT26 mice treated with G484 or PBS (Upper). Liver tumor tissues were dissected 14 days post-treatment (Lower). Circles: drug administration; triangle: iRFA or Ctrl treatments; asterisks: sample collection (peripheral blood was collected on days 3 and 5, while lung and liver tissues were collected on day 14). Circles and arrows: liver tumors. I, J MPO-dsDNA levels in peripheral blood on days (I) 3 and (J) 5 following G484 or PBS treatment (Day 3, G484+iRFA: n = 8 mice; other groups: n = 9 mice). P < 0.001, ANOVA. K, Lung tissues were dissected (Circles: lung tumors). HE staining of lung tissues. Scale bars, 1 mm. L Lung metastasis area proportion and (M) lung weight (n = 9 mice). P < 0.001, ANOVA. Data are presented as mean values ± SD. E, H, M G484: GSK484. Source data are provided as a Source Data file.
S100a6 was higher in N3 than in N4 (Figs. 2J and 3C; Fig. 3C, P = 0.002). As Pad4 and Mki67 are nuclear-expressed proteins, Cd177hiS100a6hi neutrophils were identified as N3 neutrophils to obtain living neutrophils. According to the MPO-DNA assay for NET detection, Cd177hiS100a6hi neutrophils showed the most NET formation in vitro (Fig. 3D). CT26 cells exhibited enhanced migration after coculture with Cd177hiS100a6hi neutrophils (P < 0.001), while this effect was inhibited when NETs were depleted using GSK484 (a Pad4 inhibitor) (P = 0.001) and DNase I (P < 0.001) (Fig. 3E and Supplementary Fig. 3L, M). These findings suggest the pro-metastatic contribution of N3 neutrophils mediated by NETs.
Furthermore, higher levels of dsDNA were detected in the peripheral blood of iRFA mice (Fig. 3F), and increased NETs were observed in iRFA tumors (Fig. 3G). To further examine the role of NETs in tumor metastasis, GSK484 was utilized in the mouse model to inhibit NET formation (Fig. 3H–J). Lung metastasis was significantly alleviated in NET-deficient mice compared with that in the controls (Fig. 3K–M). These findings confirm that Cd177hiPad4hi neutrophils facilitate CRLM lung metastasis in iRFA in a NET-mediated manner.
Lipid metabolism adaptation in residual tumor cells contribute to NET formation
The influence on NET formation of neutrophils after iRFA treatment still needs further study. Residual tumor tissues after iRFA (iRFA-TCS), necrotic tumor tissues (iRFA-NCS), normal liver tissues (NLCS), and control tumor tissues (Ctrl-TCS) were collected to prepare the culture supernatant for neutrophils, followed by NET detection. NETs were significantly more abundant in the iRFA-TCS group, suggesting that residual tumors promoted NET formation (Fig. 4A and Supplementary Fig. 4A). Based on the KEGG pathway enrichment analysis, upregulated genes were significantly enriched in the PPAR signaling pathway (Supplementary Fig. 4B). Fourteen PPAR-regulated genes19 (including Cyp4a10, Acox1, Cpt1a, Ehhadh, Pck1, Apoa5, Plin5, Etfdh, Slc22a5, Fabp4, Cd36, Fabp5, Angptl4, and Mmp9) were upregulated in iRFA vs. untreated tumors (Supplementary Fig. 4C), implicating activated PPAR signaling in iRFA tumors. Meanwhile, cytokine-cytokine receptor interaction was significantly enriched for both upregulated and downregulated genes, indicating stress-triggered self-regulatory feedback responses in cells. Tumor cells and the other cell types in the TME were distinguished through scRNA-seq analysis (Fig. 4B and Supplementary Fig. 4D). GSEA confirmed that PPAR-related genes were significantly enriched and upregulated in iRFA tumor cells (Fig. 4C).
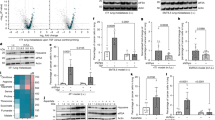
A Conditioned media from normal liver tissues (NLCS), residual tumors post-iRFA (iRFA-TCS), necrotic tissues post-iRFA (iRFA-NCS), and untreated tumors (Ctrl-TCS) were co-cultured with neutrophils. NETs were detected by immunofluorescence. Scale bars, 50 μm. B t-SNE plots comparing Ctrl versus iRFA groups. Ctrl group: 3561 cells; iRFA group: 3453 cells. C GSEA of the PPAR pathway in iRFA-treated vs. untreated CT26 tumors (n = 3 mice/group). Kolmogorov-Smirnov test. FDR correction. D Lipid metabolite differences between 37 °C-CT26 and 45 °C-CT26 cells (n = 6 technical replicates). E Tip47 immunofluorescence in liver tumors. Scale bars, 10 μm. F Oil Red O staining of liver tumors (T: tumor, L: liver, N: necrosis). Scale bars, 100 μm and 50 μm. G TG and FA levels in liver (Ctrl-L) and untreated tumors of Ctrl (Ctrl-T), and in the liver (iRFA-L), residual tumors (iRFA-T), and necrotic tissues (iRFA-N) of iRFA (TG: Ctrl-L/Ctrl-T: n = 8, iRFA-L/iRFA-T/iRFA-N: n = 9. FA: Ctrl-L/iRFA-N: n = 11, Ctrl-T: n = 10, iRFA-L: n = 8, iRFA-T: n = 13, mice). P < 0.001, ANOVA. H TG levels in CT26/DLD1 cells incubated at 37 °C/45 °C ± OA (n = 3 biological replicates). CT26: P < 0.001; DLD1: P = 0.01; ANOVA. I LD area and number in DLD1 cells at 37 °C/45 °C ± OA (n = 3 biological replicates). P < 0.001 and P = 0.001, respectively; ANOVA. J BODIPY 493 and (K) C16 immunofluorescence in cells at 37 °C/45 °C ± OA. Scale bars, 25 μm. L OCR measurements of Basal and Maximal respiration, spare respiration capacity (SRC), and ATP in CT26 cells (n = 3 biological replicates). Basal, Maximal, and ATP: P < 0.001; SRC: P = 0.001; ANOVA. M FAO rate and (N) ROS levels in CT26 cells (n = 3 biological replicates). M P < 0.001 and (N) P = 0.002, respectively; ANOVA. O, P Immunofluorescence in neutrophils co-cultured with (O) OA or (P) PF at 37 °C / 45 °C. Scale bars, 50 μm. Data are presented as mean values ± SD. (O and P) 37/45: 37 °C/45 °C; TCM: tumor-conditioned media. (M, N and P) PF: PF-04620110. Source data are provided as a Source Data file.
The PPAR signaling pathway, along with the above 14 upregulated genes in iRFA tumors, was mainly related to lipid metabolism, including fatty acid oxidation (Cyp4a10, Acox1, Cpt1a, Ehhadh, and Slc22a5), lipid storage (Apoa5, Plin5, and Etfdh), and fatty acid uptake (Fabp4, Cd36, and Fabp5)20. In vitro cell models (45 °C-CT26 and 37 °C-CT26) were established for lipid metabolomics analysis (Supplementary Fig. 5A). The 45 °C-CT26 cells simulate the residual tumor cells following iRFA, while the 37 °C-CT26 cells represent normal tumor cells that are unaffected by iRFA21. The increase in triglyceride (TG) levels in CT26 cells treated at 45 °C was the most significant (Fig. 4D). TG mainly comprises intracellular lipid droplets (LDs)22. The in vivo Tip47 and Oil Red O staining illustrated that LDs were formed in the residual tumor area in iRFA (Fig. 4E, F and Supplementary Fig. 5B). Liver tissues after iRFA (iRFA-L), residual tumor tissues after iRFA (iRFA-T), necrotic tumor tissues (iRFA-N), normal liver tissues (Ctrl-L), and the control tumor tissues (Ctrl-T) were collected for fatty acid (FA) and TG detection. TG was obviously increased in the iRFA-T group, while FA was increased in the iRFA-N group (Fig. 4G). Collectively, these findings suggest a high-FA TME and lipid accumulation in residual tumor cells following iRFA.
As the main components of TG, whether the free FAs in TME were responsible for TG formation in iRFA tumor cells needs further exploration. The lipid metabolomics analysis of the FA content in TG showed that oleic acid (C18:1) (OA) was the most abundant FA in TG (Supplementary Fig. 5C), and it was utilized in the following in vitro experiments. It was found that TG was increased more obviously after co-stimulation with 45 °C and OA both in CT26 and DLD1 cells (Fig. 4H). BODIPY 493 staining indicated that more LDs were deposited in cells treated with OA at 45 °C (Fig. 4I, J and Supplementary Fig. 5D). Additionally, the 45 °C + OA-treated group showed the highest FA level (Supplementary Fig. 5E). However, no difference was noticed in FA between the 37 °C and 45 °C groups, implying that FA uptake, rather than FA synthesis, contributed to the increased intracellular FA. BODIPY C16-stained FA further verified the increased FA uptake in tumor cells under heat stress (Fig. 4K and Supplementary Fig. 5F).
TG formation and FA uptake were strengthened in iRFA tumor. It is unclear whether fatty acid oxidation (FAO), another aspect of lipid metabolism, was influenced by iRFA. The oxygen consumption rate (OCR) was further examined (Fig. 4L and Supplementary Fig. 5G). Compared to the 37 °C-treated group, the 37 °C + OA group exhibited a higher basal and maximal respiration rate, and elevated intracellular ATP levels. However, these levels were significantly decreased in the 45 °C-treated group compared with 37 °C-treated group, and showed no difference between the 37 °C-treated group and the 45 °C + OA group. Similar results were seen in FAO rate detection (Fig. 4M and Supplementary Fig. 5H), indicating that FA uptake assisted in maintaining the homeostasis of oxidative phosphorylation under heat stress. A DGTA1 inhibitor (PF) was utilized to inhibit TG formation. Interestingly, the 37 °C + OA + PF-treated group exhibited a higher FAO rate than the 37 °C + OA-treated group, while PF significantly decreased the FAO rate in the 45 °C + OA-treated group, indicating that TG formation after 45 °C + OA treatment was conducive to increasing the FAO rate (Fig. 4M and Supplementary Fig. 5H). 45 °C + OA-treated tumor cells showed significantly lower levels of reactive oxygen species (ROS) when TG formation was inhibited (Fig. 4N). These results suggest that tumor cells co-stimulated by heat and FA have increased FA uptake, LD deposition, and FAO homeostasis; furthermore, LD formation is indispensable for maintaining FAO homeostasis.
To ensure whether these changes affect NET formation, conditioned media (CM) from the 37 °C, 37 °C + OA, 45 °C, and 45 °C + OA groups were collected and cocultured with neutrophils. The results showed the greatest NET formation in the 45 °C + OA-treated group (Fig. 4O and Supplementary Fig. 5I, J). In addition, we explored the impact of OA on NET formation. The results showed that the addition of OA alone did not increase NET formation (MPO-dsDNA) (Supplementary Fig. 5K). However, PF significantly inhibited NET formation in the 45 °C + OA-treated group (Fig. 4P and Supplementary Fig. 5L). Generally, the residual tumor cells in iRFA demonstrated lipid metabolism adaptation, maintaining homeostasis after heat stress and contributing to neutrophil NET formation.
Crosstalk between PPARγ and P38 MAPK in residual tumor cells regulates NET formation
It was found that the upregulation of the PPAR signaling pathway was most significant in the residual tumor cells. Among the PPARα, PPARβ/δ, and PPARγ subtypes, PPARα and PPARγ were notably increased and activated in CT26 and DLD1 cells treated with 45 °C + OA (Fig. 5A, CT26: Ppara and Pparg, P < 0.001; Ppard, P = 0.051; DLD1: PPARa and PPARg, P < 0.001; Fig. 5B, Pparα and Pparγ, P < 0.001; Pparβ/δ, P = 0.12). While PPARγ was primarily expressed in the nucleus, it exhibited the highest expression levels in iRFA (Fig. 5C, D, and Supplementary Fig. 6A, B). PPAR, as a nuclear transcription factor, can transcriptionally regulate the expression of downstream target genes. Among the genes targeted by PPAR, the protein and mRNA expression levels of CD36, FABP4, and FABP5 were significantly increased, as confirmed by both in vitro and in vivo studies (Fig. 5E, F, and Supplementary Fig. 6C-F). To further investigate the regulation of CD36, FABP4, and FABP5 expression by PPARγ, we constructed Pparg/PPARg-silenced (siPparg/PPARg) tumor cells (Supplementary Fig. 7A). The expression levels of CD36, FABP4, and FABP5 were significantly reduced in siPparg/PPARg tumor cells (Fig. 5G and Supplementary Fig. 7B–D). In addition, treatment with T0070907 (a PPARγ inhibitor) resulted in a significant decrease in CD36, FABP4, and FABP5 levels (Supplementary Fig. 7E). Generally, the expression of CD36, FABP4, and FABP5 in residual tumor cells was regulated by PPARγ.
A PPARa, PPARd, and PPARg mRNA detected via qPCR in cells (n = 7 biological replicates). ANOVA. B PPRE binding ability for PPAR (n = 8 biological replicates). ANOVA. C, D Nuclear and cytoplasmic PPARα, PPARβ/δ, and PPARγ proteins in (C) tumor cells and (D) tumor tissues detected via Western blotting. E, F CD36, FABP4, and FABP5 proteins in (E) tumor cells and (F) tumor tissues detected via Western blotting. G CD36, FABP4, and FABP5 proteins in CT26 and DLD1 cells treated with siCtrl or siPPARg at 37 °C and 45 °C + OA were detected via Western blotting. H Detection of FA and TG levels in CT26 cells treated with siCtrl or siPparg at 37 °C and 45 °C + OA (Upper). Detection of FA and TG levels in CT26 cells treated with DMSO or T0 at 37 °C and 45 °C + OA (Lower). FA: n = 3; TG: n = 5; biological replicates. P < 0.001, ANOVA. I Detection of FA and TG levels in DLD1 cells treated with DMSO or T0 at 37 °C and 45 °C + OA (FA: n = 3, TG: n = 5, biological replicates). P < 0.001, ANOVA. J BODIPY C16 was detected in CT26 cells treated with DMSO or T0 at 37 °C and 45 °C + OA via immunofluorescence. Scale bars, 25 μm. K The positive LD area in DLD1 cells treated with DMSO or T0 at 37 °C and 45 °C + OA (n = 3 biological replicates). P < 0.001, ANOVA. L BODIPY 493 was detected in DLD1 cells treated with DMSO or T0 at 37 °C and 45 °C + OA via immunofluorescence. Scale bars, 25 μm. M,Nile red staining of CT26 cells treated with siCtrl or siPparg at 37 °C and 45 °C + OA. Data are presented as mean values ± SD. C, E and G 37/45: 37 °C/45 °C. H–L T0: T0070907. Source data are provided as a Source Data file.
As CD36 is recognized as a membrane receptor associated with FA uptake, and FABP4/5 function as FA transporters, we hypothesized that PPARγ plays a role in FA uptake and lipid accumulation in residual tumor cells, mediated by CD36 and FABP4/5. Both FA and TG were significantly decreased by Pparg silencing and PPARγ inhibition (Fig. 5H, I). The reduced uptake of FA and LD synthesis was also observed in siPparg/PPARg tumor cells or as a result of PPARγ inhibition (Fig. 5J-M and Supplementary Fig. 7F, G). In conclusion, PPARγ accelerated FA uptake and lipid accumulation in residual tumor cells, mediated by CD36 and FABP4/5.
mTOR, MAPK (JNK, ERK, and P38), and PI3K/AKT signaling pathways are regulated by lipids23. Western blotting found that p-P38 was upregulated by 45 °C and OA treatment (Fig. 6A and Supplementary Fig. 8A-C). The GSEA of the scRNA-seq data also showed that the MAPK signaling pathway was enriched in iRFA tumor cells (P < 0.001, Fig. 6B). The same results were obtained in vivo (Fig. 6C; P38, P = 0.753; p-P38, P = 0.018), suggesting activation of the P38 MAPK in residual tumor cells.
A mTOR, p-mTOR, Akt, p-Akt, P38, and p-P38 proteins detected in CT26 cells at 37 °C/45 °C ± OA. B Representative GSEA analysis of the MAPK pathway in scRNA-seq data from untreated and iRFA-treated CT26 tumors (n = 3 mice). Kolmogorov-Smirnov test. FDR correction. C P38 and p-P38 in untreated and iRFA-treated tumor tissues. The quantification shown on the right (n = 9 biological replicates). ANOVA. D Nuclear PPARγ, and cytosol CD36, FABP4, and FABP5 proteins in CT26 cells treated with DMSO or SB202190 at 37 °C and 45 °C + OA. E FA levels in CT26/DLD1 cells treated with DMSO, SB190, or SB580 at 37 °C and 45 °C + OA (n = 3 biological replicates). P < 0.001, ANOVA. F Nile red staining of CT26 cells treated with DMSO or SB190 at 37 °C and 45 °C + OA. G p-P38 and P38 in CT26 cells treated with siCtrl or siPparg at 37 °C and 45 °C + OA (Upper), and treated with DMSO or T0070907 (Lower). H p-P38 and P38 proteins in CT26 and DLD1 cells treated with DMSO or PF at 37 °C and 45 °C + OA. I CD36, FABP4, FABP5, PPARγ, and p-P38 staining in liver tumors from iRFA and Ctrl mice. J Pearson correlation between P38 and neutrophil infiltration. K P38 score and neutrophil infiltration score of each patient obtained through ssGSEA. Survival analysis in NEUHp38H and NEULwp38L patients. L MPO-dsDNA levels in neutrophil co-cultured with conditioned media from CT26/DLD1 cells treated with DMSO or SB190 at 37 °C, 37 °C + OA, 45 °C, and 45 °C + OA (n = 3 biological replicates), P < 0.001, ANOVA. M Lung weight of iRFA and Ctrl CT26 mice treated with SB190 or PBS (n = 5 mice), P < 0.001, ANOVA. N HE staining of lung tumors. Scale bars, 1 mm and 50 μm. O Immunofluorescence detection of Ly6G and cit-H3 in liver tumors from iRFA-CT26 mice treated with SB190 or PBS. Scale bars, 50 μm. Data are presented as mean values ± SD. A, D, G, and H 37/45: 37 °C/45 °C. E, F, L, M, and O SB190: SB202190. E SB580: SB203580. H PF: PF-04620110. Source data are provided as a Source Data file.
The expression of Pparγ, Cd36, Fabp4, and Fabp5 was significantly inhibited by the P38 inhibitor, SB202190 (Fig. 6D and Supplementary Fig. 8D, E). Furthermore, the FA and LD levels in 45 °C + OA-treated tumor cells were significantly decreased by P38 inhibition (SB202190 and SB203580) (Fig. 6E, F and Supplementary Fig. 8F). These data suggest that PPARγ-mediated lipid formation is regulated by the P38 MAPK signaling pathway. Meanwhile, silencing Pparg/PPARg or inhibiting PPARγ significantly downregulated p-P38, as did inhibiting TG, suggesting PPARγ-mediated lipid formation activated the P38 MAPK signaling pathway (Fig. 6G, H and Supplementary Fig. 8G–J). Cd36, Fabp4, Fabp5, Pparγ, and p-P38 were all increased by iRFA in vivo (Fig. 6I and Supplementary Fig. 8K). Together, positive feedback was confirmed within the PPARγ-TG-P38 MAPK axis.
The TCGA-COREAD cohort includes mRNA data and clinical information from CRC patients, which was obtained from the UCSC Xena Browser. Based on the TCGA-COREAD cohort, activation of the P38 MAPK signaling pathway was positively associated with neutrophil infiltration (Fig. 6J). After further classifying patients into NEUHP38H and NEULP38L using the ssGSEA method, poorer survival outcomes were observed in NEUHP38H patients (Fig. 6K). In vitro inhibition of P38 significantly decreased NET formation, and in vivo inhibition suppressed lung metastasis and decreased NETs (Fig. 6L–O and Supplementary Fig. 8L, M). These data suggest that the P38 MAPK signaling pathway is closely related to neutrophil infiltration and NET formation.
The P38 MAPK-CXCL5-pERK axis mediates NET formation
According to bulk RNA-seq data, Cxcl5, Csf3, Cxcl2, Cxcl1, Il1b, and Tnf were significantly increased in iRFA tumors (Fig. 7A). Elevated mRNA expression levels of Cxcl5/CXCL5 and Il1b/IL1B were confirmed in cells treated with 45 °C + OA (Supplementary Fig. 9A). Regarding protein levels, the Cxcl5 protein level in the 45 °C + OA group was significantly elevated (Fig. 7B). With the P38 inhibitors, both mRNA and protein expression levels of Cxcl5 were decreased (Fig. 7C and Supplementary Fig. 9B), indicating that the secretion of Cxcl5 by residual tumor cells mediated by the P38 signaling pathway. NET formation were promoted in neutrophils in 37 °C-treated tumors with added Cxcl5, and these effects were inhibited in 45 °C + OA-treated tumor cells exposed to P38 or CXCR1/2 (Reparixin) inhibitors (Figs. 6L, 7D and Supplementary Fig. 9C, D). Thus, Cxcl5 was the crucial driving factor for NET formation in neutrophils, mediated via P38 MAPK activation.
A Differences in RNA-seq between Ctrl and iRFA on day 3 post-iRFA (n = 3 mice). Volcano plot of DEGs identified by DESeq2, based on a negative binomial model. B Cxcl5 (P < 0.001) and Il1b (P = 0.045) in CT26 cells ± OA treated at 37 °C/45 °C via ELISA (n = 3 biological replicates). ANOVA. C Cxcl5 in CT26 cells treated with DMSO, SB190, or SB580 at 37 °C/45 °C ± OA (n = 3 biological replicates). P < 0.001, ANOVA. #, P < 0.001 vs. 45 °C + OA + DMSO group. D MPO-dsDNA levels in neutrophils co-cultured with conditioned media from CT26 cells (37 °C/45 °C) ± OA, with Cxcl5 and / or CXCR1/2 inhibitor (Reparixin) treatment (n = 3 biological replicates). P < 0.001, ANOVA. # vs. 37 °C; ▼ vs. 37 °C + OA; ★ vs. 45 °C; P < 0.001. E (Left) Immunofluorescence in neutrophils co-cultured with 37 °C and 45 °C + OA CT26 / DLD1 TCM supplemented with DMSO / Trametinib (MEK/ERK inhibitor). (Right) cit-H3+ area quantification (n = 3 biological replicates). P < 0.001, ANOVA. F, Erk and pErk in the neutrophils from iRFA mice. G–I Proteins in neutrophils treated with 37 °C or 45 °C ± OA CT26 TCM supplemented with ± (G) Cxcl5, (H) SB190, and (I) Reparixin. J Lung tissues from the Ctrl and iRFA mice treated with DMSO, Trametinib and/or a P38 inhibitor (Ralimetinib). K HE staining of lung tissues (Circles: lung metastases). L The quantification of lung metastatic area (n = 9 mice). Ctrl groups: P = 0.059; iRFA groups: P < 0.001; ANOVA. M (Left) Ly6G and cit-H3 immunofluorescence in liver tumors. (Right) Quantification of the cit-H3+ area (n = 9 mice). P < 0.001, ANOVA. Scale bars, 200 μm. N Ly6G and pErk immunofluorescence in liver tumors. Scale bars, 50 μm. Data are presented as mean values ± SD. C, H SB190: SB202190. C SB580: SB203580. E 37/45: 37 °C/45 °C. E and G–I TCM: conditioned media from tumor cells. Source data are provided as a Source Data file.
NET formation is regulated through the STAT, PI3K/AKT, and MEK/ERK signaling pathways24. Therefore, STAT1/3/5, PI3K, and MEK inhibitors were combined with tumor-CM to determine their influence on NET formation. The most significant decrease was observed with MEK/ERK inhibitor (Trametinib) treatment (Fig. 7E and Supplementary Fig. 9E). Meanwhile, Erk and pErk were significantly increased in N3 neutrophils (Cd177hiMki67- neutrophils) (Fig. 7F and Supplementary Fig. 9F).
Exogenous Cxcl5 elevated the levels of pErk1/2 and cit-H3 in neutrophils (Fig. 7G and Supplementary Fig. 9G, H), whereas P38 and CXCR1/2 inhibitors reduced these levels in the same cell type (Fig. 7H, I and Supplementary Fig. 9I–K). P38 (Ralimetinib) plus MEK/ERK inhibitors treatment in vivo significantly inhibited lung metastasis and NET formation compared with monotherapy (Fig. 7J-M and Supplementary Fig. 10A). Further, pErk+ neutrophil levels were significantly decreased by the combined treatment of P38 and MEK/ERK inhibitor (Fig. 7N and Supplementary Fig. 10B, C). The P38 and MEK/ERK dual-targeted therapy is expected to be a salvage alternative for iRFA.
CD36hi-EM-CTC detection assists in the early identification of CRLM patients with iRFA
We next focused on paired samples from CRLM patients who underwent iRFA to validate the above findings (Supplementary Data 2). A representative CRLM patient who developed new intrahepatic metastasis (liver S7) and lung metastases after iRFA was shown in Fig. 8A. Liver S5 (iRFA-treated lesion) and S7 tissues were obtained through hepatic resection. Pathological examination showed residual tumors in liver S5 tissues after iRFA, and greater CD177hiPAD4hi neutrophil infiltration and higher NET formation were observed in S5 tissues compared with those in S7 tissues (Supplementary Fig. 11A, B). Bulk RNA-seq supported the previous conclusion that neutrophil levels were increased and the PPAR signaling pathway was activated after iRFA (Fig. 8A). Additionally, the expression of CD36 and FABP4 was increased.
A Treatment timeline for a representative CRLM patient. The patient underwent Dixon surgery following CRLM diagnosis. First tumor evaluation showed a new metastasis in liver S5, and subsequently, RFA was performed. Postoperative PET/CT showed no tumor viability. Intrahepatic S7 metastasis appeared 4 months later. S5 and S7 tissues were obtained through hepatic resection, and pathological examination as well as sequencing were performed. The second progression with new lung metastases occurred at 7 months. Triangles: liver or lung metastases. B Neutrophil infiltration levels in tumors pre- and post-iRFA analyzed by different datasets (Box center line: median; box limits: upper and lower quartiles; whiskers: 1.5× interquartile range). C Representative GSEA analysis of FA metabolism and the PPAR pathway, and the differences in scRNA-seq data between pre- and post-iRFA tumors. Kolmogorov-Smirnov test. FDR correction. D CD36 and CXCL5 staining of the tumors pre- and post-iRFA. E Density quantification of CD36+ and CXCL5+ areas in the tumors pre- and post-iRFA. F CD66b immunofluorescence in tumors pre- and post-iRFA. Scale bars, 50 μm and 10 μm. CD66b+ neutrophil counts shown on the left. G MPO and cit-H3 immunofluorescence in tumors pre- and post-iRFA. Scale bars, 50 μm. The cit-H3+ area shown on the left. H Mesenchymal markers, epithelial markers, and CD36 immunofluorescence on CTCs were analyzed. The data represented the proportions of CD36hi-CTCs, CD36hi-E-CTCs, and CD36hi-EM-CTCs in the tumors pre- and post-iRFA. Scale bars, 10 μm. I MPO-dsDNA levels in the peripheral blood pre- and post-iRFA. J The data represented the proportions of CD36hi-CTCs and CD36hi-EM-CTCs in the cRFA and iRFA tumors. K MPO-dsDNA levels in the peripheral blood of cRFA and iRFA patients. Data are presented as mean values ± SD; Statistical analysis was performed based on Paired t-test (B, E, F, and G, n = 8 patients; H and I, n = 13 patients) or Unpaired two-sided t-test (J, K); cRFA, n = 5 patients, iRFA, n = 13 patients). Source data are provided as a Source Data file.
Bulk RNA-seq was then performed on liver metastases from eight CRLM patients before and after iRFA. Neutrophil infiltration levels were significantly increased, and fatty acid metabolism and the PPAR signaling pathway were enriched in post-iRFA tumors (Fig. 8B, C). Further, CD36 and CXCL5 were significantly upregulated in post-iRFA tumors (Fig. 8D, E). Although there was no statistical difference, FABP4, FABP5, and PPARγ levels were increased in the residual tumor tissues of most patients with iRFA compared with pre-treatment tissues (Supplementary Fig. 11C). Moreover, neutrophil infiltration and NET formation were significantly increased after iRFA (Fig. 8F, G). These data further support our findings from the animal study.
It was concluded that iRFA is closely related to subsequent lung metastasis in CRLM patients; however, the timely and early identification of patients who have suffered from iRFA remains a challenging issue. PET/CT examination cannot accurately detect tumor residue after iRFA. CTCs are widely accepted for the early detection and monitoring of minimal residual disease11; in particular, CTC phenotyping enables the specific recognition of metastatic tumor cells. Nevertheless, there were no differences in the total epithelial, mesenchymal, and epithelial-mesenchymal (EM) CTC counts before and after iRFA treatment in 13 CRLM patients (Supplementary Fig. 11D, E). CTCs with epithelial and mesenchymal phenotypes and medium-to-high expression of CD36 were defined as CD36hi-EM-CTCs (Fig. 8H). Increased CD36hi-EM-CTCs and NET formation were observed in peripheral blood from CRLM patients with iRFA (Fig. 8H, I); however, there was no significant change in the 5 CRLM patients with cRFA (Supplementary Fig. 11F, G). Further analysis showed that CD36hi-CTCs, CD36hi-EM-CTCs, and NET formation were significantly higher in iRFA patients than in cRFA patients (Fig. 8J, K). CD36hi-EM-CTCs and NET detection are expected to realize the early identification of residual tumor cells.
Discussion
Metastasis and recurrence remain the main causes of reduced RFA efficacy. Residual tumor cells in the transitional zone undergo reversible hyperthermia and eventually acquire a more aggressive cancer phenotype. However, no direct evidence has confirmed the origin of the tumor cells that cause iRFA-related metastasis. In this study, we demonstrate that iRFA induces the emergence of residual tumor cells that act as lung metastasis seeds.
iRFA elicits complex physiological reactions in residual tumors, during which the local inflammatory TME releases proinflammatory cytokines and chemokines into the transitional zone7. Changes in the local TME following iRFA were also observed in our mouse models and tissue specimens from CRLM patients undergoing iRFA, characterized by substantial neutrophil accumulation. Lung metastasis after iRFA is notably regressed when neutrophils are depleted, supporting neutrophils as the central participant in tumor progression25. However, the therapeutic strategies to eliminate neutrophils have limited clinical value.
Neutrophil heterogeneity has been extensively investigated10,26. Different neutrophil subsets express different marker genes and function diversely26. Here, we employed scRNA-seq analysis to profile the neutrophils infiltrating into residual tumors. Cd177-positive neutrophils are functionally activated27. As a surface molecule, CD177 facilitates neutrophil migration, regulates neutrophil-mediated inflammatory responses, and demonstrates complex immunoregulatory functions through its expression in various tissues and organs28. However, the specific mechanisms of action of CD177-positive neutrophils in various tissues and diseases are not yet fully understood. We found that Cd177hiPad4hi neutrophils were enriched in the residual tumors and prominent in NET formation, promoting tumor invasion. After blocking NET formation with a specific Pad4 antagonist, lung metastasis was alleviated in iRFA. This finding provides a salvage alternative for treating residual tumors after iRFA.
Tumor lipid synthesis is considered a sign of invasiveness and adaptation to harmful stimuli29. We found that the residual tumor cells were in a high FA microenvironment, suggesting that the in vivo microenvironment of residual tumor is not merely thermal stimulation or a high-FA environment but rather a combination of both. Therefore, in our study, the 45 °C + OA group was chosen as the main in vitro experimental group, differing from the previous recommendation of using only the 45 °C group, which represents the biggest difference in experimental design. Under ablation-induced FA abundance conditions, excess FAs are incorporated into lipids to confer resistance to sublethal thermal stress and maintain basal FAO, which is favorable for malignant tumors. Previous studies have demonstrated that PPARγ functions as a tumor suppressor involved in regulating tumor proliferation, differentiation, apoptosis, and metastasis. In this study, the PPARγ-TG synthesis pathway in residual tumor cells is activated to maintain lipid uptake and FAO homeostasis.
Moreover, the metabolic adaption of cancer is associated with immune evasion and tumor-promoting inflammation. Increasing evidence have indicated that PPARγ can function as a cancer promoter, particularly in a lipid-rich environment. Our data further suggest that high lipid content activates the P38 signaling pathway, playing a role in the secretion of the inflammatory mediator CXCL5, neutrophil infiltration, activation of the MEK/ERK signaling pathway, and NET formation. The role of NETs in driving metastasis is well-established. In this study, we found that CXCL5 secreted by residual tumor cells mediates increased ERK phosphorylation in infiltrating neutrophils after iRFA, thereby promoting the release of NETs from CD177hiPAD4hi neutrophils. This contributes to a deeper understanding of the mechanisms by which neutrophils release NETs. The mechanisms by which residual tumors educate neutrophils provide opportunities for therapeutic intervention in CRLM patients undergoing iRFA. The positive feedback loop of the P38 signaling pathway through the PPARγ axis on TG synthesis was also confirmed in this study. These findings are consistent with previous reports that activation of the P38 pathway can upregulate PPARγ expression30,31, and P38 signaling is also a downstream signal of PPARγ30,32. Metabolic inflammatory crosstalk is considered the causal mechanistic link between the neutrophil-oriented inflammatory microenvironment and tumor metastasis.
P38 MAPK inhibitors have been evaluated in clinical trials involving advanced CRC patients, especially in combination with chemotherapy33. However, it has not been applied clinically at present. It is still challenging to determine how to select the effective population and optimize the combination therapy regimens. Our research has confirmed the potential application value of p38 inhibitors in combination regimens for iRFA patients. MEK inhibitor combined with BRAF + EGFR inhibition was demonstrated to be tolerable and showed promising activity in patients with BRAFV600E colorectal cancer34. However, the treatment strategies involving these three drug regimens are still not recommended by guidelines. Based on our results, targeting P38 and MEK/ERK jointly completely inhibits lung metastasis without any noticeable toxic effects in mice with iRFA. P38- and MEK/ERK-targeting therapy presents an opportunity for controlling tumor progression after iRFA and addresses many challenges in clinical efficacy and toxicity.
Identifying patients with postoperative residual tumor cells is another key issue in iRFA. Imaging examinations, such as PET/CT, often fail to detect tumor residue after iRFA. Our animal study revealed a higher incidence of lung metastasis in the day 3 resection group compared to that in the day 1 resection group, indicating that early detection and timely remedial measures are conducive to reducing the adverse effects of iRFA. Among various liquid biopsy markers, CTCs are widely accepted, and CTC counts have been used in the clinic11. As tumor heterogeneity is gradually realized in tumor metastasis35, phenotyping of CTCs is also emerging for prognostic monitoring. The development of single-cell profiling has resulted in the discovery of various CTC phenotypic markers. Fumagalli A et al. have found Lgr5-Negative CTCs to be the most disseminated CRC cells driving distant metastases12. In this study, metabolic adaptation was observed in residual tumor cells, and we identified CD36hi-EM-CTCs, which represent the disseminated residual tumor cells in iRFA, as having activated lipid metabolism. This finding further supports that CTC phenotyping is crucial for prognostic monitoring in addition to CTC counts. Regarding RFA treatment, CD36hi-EM-CTC detection is expected to enable early identification and intervention of residual tumor cells, and address the limitations of the previous epithelial and mesenchymal phenotype-based classification.
Overall, we have demonstrated that Cd177hiPad4hi neutrophils in the local microenvironment drive the metastasis of residual tumor cells through NET formation. Targeting the P38 and MER/ERK signaling pathways blocks neutrophil infiltration and NET formation, significantly suppressing metastasis. This study provides insight into the mechanisms of the intricate interaction between cancer and inflammation, which should eventually lead to more innovative anti-inflammatory cancer therapies reaching the clinic. With CD36hi-EM-CTC detection, the early identification and intervention of iRFA can be realized, improving the effect of RFA in CRLM patients.
Methods
Patients and human tissue samples
To investigate the correlation between RFA and hepatectomy regarding the development of new metastases. This study included patients diagnosed with CRLM at Nanfang Hospital, Southern Medical University, from September 2009 to July 2024, who underwent either RFA or resection of liver metastases. We collected clinicopathological parameters and postoperative imaging data for the patients who were followed up until the occurrence of new extrahepatic metastases, death, or loss to follow-up. Exclusion criteria included patients who did not undergo imaging reexamination within four months after surgery, making it impossible to assess the residual status of postoperative liver metastases, as well as those who developed extrahepatic metastases within two months after surgery (to exclude patients with pre-existing metastases prior to surgery).
Eight paired pre- and post-iRFA liver tumor samples from iRFA-treated CRLM patients were obtained for bulk RNA-seq and immunohistochemistry. Among these, liver S5 (iRFA-treated lesion) and S7 (new intrahepatic metastasis) tumor specimens from a representative iRFA case were obtained through hepatic resection in our study. The pre-iRFA sample is the biopsy specimen taken at the time of definitive diagnosis, while the post-iRFA specimen is the biopsy-confirmed specimen or surgically obtained specimen after the suspected tumor residue was identified through PET/CT examination.
We collected peripheral blood samples from 18 CRLM patients who underwent RFA and conducted CTC and NET detection. Among these cases, 13 were classified as iRFA, which was defined as observable residual tumor within 4 months after RFA7, while 5 were classified as cRFA.
As the information on gender was not relevant in this study, no related analysis was conducted. The clinical characteristics of patients are shown in Supplementary Data 1 and 2, and we obtained consent to collect samples and publish relevant clinical information before enrollment. No statistical method was used to predetermine the sample size. No data were excluded from the analyses.
Cell culture and treatment
CT26 (Cat. TCM37), MC38 (Cat. TCM46), and DLD1 (Cat. TCHu134) colon cancer cells were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China) and maintained in RPMI-1640 medium (Hyclone, Logan, UT, USA) supplemented with 10% fetal bovine serum (FBS) (GIBCO BRL, Invitrogen, Grand Island, NY, USA) at 37 °C in 5% CO2.
For thermal stimulation, cell culture-medium containing 10% FBS was preheated in a 45 °C cell incubator and added to cells at 60-70% confluency. Cells were then transferred to 45 °C for 15 min and subsequently to 37 °C for culture.
After growing to 60-70% confluence, cells were incubated with OA (OA:10% FBS = 1:10000, 0.31509 mM) (112-80-1, Sigma-Aldrich) for 48 h. DGAT1 inhibtor (25 μM) (PF-04620110, S7192, Selleck), p38 inhibitor (20 μM) (SB202190/SB203580, T2301/T6335, TargetMol), and PPARγ inhibitor (20 μM) (T0070907, S2871, Selleck) were used to inhibit TG formation, p38 signaling pathway, and PPARγ, respectively. Tumor cell supernate supplemented with MEK inhibitor (Trametinib, GSK1120212, Selleck) or CXCR1/2 inhibitor (1 μM) (Repertaxin, SX-682, Selleck) was used to treated neutrophils.
Cell transfection
CT26Luc/eGFP and CT26mCherry cells were constructed by LV-Luc/eGFP and LV-mCherry transfection (FulenGen, Guangzhou, China). CT26 cells were seeded into the 12-well plates and transfected with LV-Luc/eGFP or LV-mCherry using polybrene (Obio technology, Shanghai, China). Successfully transfected cells were selected with puromycin (1 μg/mL) and confirmed by microscopy (Bx51, Olympus, Japan).
PPARγ-silenced tumor cells were constructed by transfecting siRNA against PPARγ or a negative control siRNA (NC) (RiboBio Co., Ltd., Guangdong, China) into the cells using Lipofectamine 2000 Transfection Reagent (Invitrogen, Carlsbad, CA, USA) according to manufacturer’s instructions. After incubated for 4-6 h, fresh media supplemented with 10% FBS were added for another 24-48 h incubation.
Animal models and treatment
All animal experiments were conducted in accordance with the ethical guidelines approved by the Nanfang Hospital Ethics Review Board. Six-week-old BALB/c and C57BL/6 mice were obtained from the Animal Center of Southern Medical University (Guangzhou, China) and housed under specific pathogen-free (SPF) conditions, maintained on a 12-hour light/dark cycle in an environment with an ambient temperature of 22 ± 2 °C and a relative humidity of 50 ± 10%. In accordance with institutional ethical thresholds, the maximal permitted tumor burden was defined as 1.5 cm in diameter or 10% of the animal’s body weight. Tumor dimensions and body weight were measured every 3 days using digital calipers and a precision scale, respectively. Significant differences in the sizes of lung metastases and liver lesions were observed on the 14th day after treatment between the iRFA group and the control group by using in vivo imaging. Therefore, the 14th day was selected as the endpoint of the animal experiment. No animals exceeded the approved tumor burden threshold during the study period. Both male and female mice were included; however, sex was not analyzed as a separate variable due to the nature of the research and the absence of prior evidence indicating sex differences in the outcome measures.
The specific cells were harvested through trypsinization, washed twice with PBS, and then resuspended in mixture of PBS at a concentration of 2 × 107 cells/mL. Six-week-old BALB/c mice were anesthetized with 1% pentobarbital sodium (0.15–0.2 mL/10 g body weight), and a small midline incision was made to expose the left liver lobe. Twenty-five microliters of cells (5 × 105) were injected into the left liver lobe beneath the capsule. Treatments were initiated 7 days after cell implantation. RFA was performed using a 17-gauge single ablation electrode (Medsphere International Holdings. Inc., CA, USA) with a 5 mm active tip inserted into the tumors. Complete RFA (cRFA): Electrodes were inserted along the long axis of the tumor, with the tips of the electrodes reaching the contralateral edge of the tumor. Treatments were administered for 3 to 5 min, at 2 watts. iRFA: The tips of the electrodes reached the middlepoint of the tumor’s long axis. Treatments were administered for 1 to 2 min, at 2 watts. In the sham-operated group, electrode puncture was performed for 1 min without conducting RFA.
For the CT26, CT26Luc/eGFP, and CT26mCherry mouse models, 5 × 105 CT26, CT26Luc/eGFP, and CT26mCherry cells were injected into the left liver lobe beneath the capsule in BALB/c mice. For MC38 mouse model, 5 × 105 MC38 cells were injected into the left liver lobe beneath the capsule in C57BL/6 mice. Subsequently, treatment (either iRFA, cRFA or Ctrl) were performed on tumors located in the left lobe of the liver.
For the CT26R-mCherry+L-GFP mouse model, 5 × 105 CT26Luc/eGFP cells were injected into the left liver lobe, and 5 × 105 CT26mCherry cells were injected into the right liver lobe of BALB/c mice. For the CT26L-mCherry+R-GFP mouse model, 5 × 105 CT26mCherry cells were injected into the left liver lobe, and 5 × 105 CT26Luc/eGFP cells were injected into the right liver lobe of BALB/c mice. Subsequently, treatment (either iRFA or control) was performed on tumors located in the left or right lobe of the liver.
For the iRFA-Resection-CT26 BALB/c mouse model, 5 × 105 CT26 cells were injected into the left liver lobe beneath the capsule in BALB/c mice. Subsequently, iRFA was performed on tumors located in the left lobe.of the liver. The left liver lobes were resected on day 1 or 3 following iRFA, respectively. The non-resection group was only incised and sutured.
For the RFANormal and iRFATumor mice models, 5 × 105 CT26 cells were injected into the left liver lobe beneath the capsule in BALB/c mice. Subsequently, RFA was performed on liver tissues on the right lobe (RFANormal), while iRFA was conducted on tumors located in the left lobe (iRFAtumor).
A variety of drug-treated mouse models were used in our studies. MEK inhibitor (1 mg/kg/day; Trametinib, GSK1120212, Selleck, USA)36, p38 inhibitor (5 mg/kg/day; Ralimetinib, LY2228820, Selleck, USA)37, and PAD4 inhibitor (4 mg/kg/day; GSK484, S7803, Selleck, USA)38 were administered orally. InVivoMAb anti-mouse Ly6G (initial dose 400 μg, and subsequently 100 μg; clone1A8, BE0075-1, BioXcell, USA)39 and CXCR1/2 inhibitor (initial dose 80 mg/kg, and subsequently 20 mg/kg/day; Repertaxin, SX-682, Selleck, USA)40 were administered intraperitoneally. All processes were according to the instructions provided by the manufacturer.
Bioluminescent in vivo imaging
Bioluminescent in vivo imaging was performed to evaluate CT26Luc/eGFP tumors41. Briefly, the VivoGlo™ Luciferin in vivo Grade solution (150 mg/kg) was injected intraperitoneally into the tumor-bearing mice. Within 10-12 minutes after injection, bioluminescence-positive signals were photographed using an in vivo imaging system (FX Pro, Bruker, USA).
HE staining41
Tumor tissues were fixed in 10% formalin for 1 day, then embedded in paraffin and cut into 4-μm thick sections. The tissue sections were dried at 62 °C for 30 minutes to 1 hours. They were then subjected to two rounds of dewaxing in xylene for 20 minutes each, followed by sequential hydration in decreasing concentrations of ethanol (100%, 95%, 90%, 80%, and 70%) for 3-5 minutes per concentration. Hematoxylin staining was performed for 5–15 minutes, after which the sections were washed with running water. Differentiation was carried out using 1% hydrochloric acid in alcohol for 2–5 seconds, followed by counterstaining to restore blue coloration. The sections were subsequently dehydrated in increasing concentrations of ethanol (70%, 80%, 95%), stained with eosin for 30 seconds to 2 minutes, and further dehydrated in absolute ethanol (I and II) for 5 minutes each. Clearing was achieved by immersing the sections in xylene (I and II) for 10 minutes each. Finally, the sections were mounted with neutral balsam and examined under a microscope (SQS-120P, TEKSQRAY, China), with images captured for documentation.
Immunohistochemistry
Paraffin-embedded slices were prepared for immunohistochemistry. Briefly, paraffin-embedded tissues were cut into 4-μm sections. After dewaxing, hydration, and antigen retrieval, the slices were incubated with the following primary antibodies: mCherry (1:200, ab167453, Abcam), Cd45 (1:200, GB11066, Servicebio), CD36 (1:200, 18836-1-AP, Proteintech), FABP4 (1:200, 12802-1-AP, Proteintech), FABP5 (1:200, 12348-1-AP, Proteintech), PPARγ (1:200, 10156-2-AP, Proteintech), p-P38 (1:200, 4511 T, Cell Signaling Technology), and CXCL5 (1:250, bs-2549R, Bioss). The slices were then incubated with the following secondary antibodies: enzyme-labeled polymer goat anti-rabbit (PV-6001, ZSGB-Bio) and enzyme-labeled polymer goat anti-mouse (PV-6002, ZSGB-Bio). After incubation with 3,3’-diaminobenzidine (DAB, ZLI-9019, ZSGB-Bio) solution for color developments, the stained slices were captured by a light microscope (SQS-120P, TEKSQRAY, China).
For the immunohistochemical scoring of protein expression, such as CD36 and FABP4, five visual fields were randomly selected from each stained tissue section. The staining intensity of each field was scored on a scale from 0 to 3 points, categorized as follows: colorless (0), light yellow (1), brownish-yellow (2), and yellow-brown (3). The number of positive cells was counted and classified into the following categories: ≤5% (0 points), 6%-25% (1 point), 26%-50% (2 points), 51%-74% (3 points), and ≥75% (4 points). The final score was determined by multiplying the scores from these two assessments. All results must be independently reviewed in a double-blind manner by two experienced pathologists to minimize errors.
Tissue immunofluorescence
Paraffin-embedded slices were prepared for immunofluorescence as in immunohistochemistry. Slices were incubated with primary antibodies against GFP (1:200, ab183734, Abcam), mCherry (1:200, ab167453, Abcam), Ly6G (1:200, #87048, Cell Signaling Technology), Cd177 (1:50, sc-376329, Santa Cruz), Pad4 (1:200, ab214810, Abcam), citrulline Histone H3 (1:200, ab5103, Abcam), Tip47 (1:200, ab47639, Abcam), pERK (1:200, #4370, Cell Signaling Technology), MPO (1:300, GB11224, Servicebio), CD66b (1:200, ab300122, Abcam), and p-P38 (1:200, #4511, Cell Signaling Technology). After incubation with the following fluorescent secondary antibodies: Cy3-labeled goat anti-Rabbit (1:500, A0516, Beyotime) and Alexa Fluor 488-labeled goat anti-mouse (1:500, A0428, Beyotime), the slices were sealed by anti-quenching agents and photographed under a microscope (Ti2-E, Nikon, Japan). Nuclei were stained with 4,6-diamidino-2-phenylindole (DAPI) (C1002, Beyotime) for 5 minutes at room temperature.
Bulk RNA-sequencing and further analysis
Total RNA was isolated from the indicated tissue samples using Trizol reagent according to the manufacturer’s protocol. Ribosomal RNA was removed from the total RNA using EpicentreRibo-Zero rRNA Removal Kit (Illumina, USA), and the remaining RNA was fragmented to approximately 200 bp. Subsequently, RNA-seq libraries were generated according to the instructions of the NEBNext® Ultra™ RNA Library Prep Kit for Illumina (NEB, USA). The purified library products were evaluated using the Agilent 2200 TapeStation and Qubit®2.0 (Life Technologies, USA), and then paired-end sequencing (2 × 150 bp) was performed on the HiSeq 3000. The clean reads were aligned to the mouse reference genome mm10 with HISAT2 (version 2.0.5). HTSeq v0.6.0 was employed to count the reads numbers mapped to each gene.
DEGs were identified by the “DEseq2” package (version 1.34.0) and were declared statistically significant according to the criteria of | log2 (fold change) | > 1 and Benjamini-Hochberg (BH) adjusted P-value < 0.05. Heatmaps of DEGs were plotted using the “pheatmap” package (version 1.0.12). Meanwhile, the “ggplot2” package (version 3.4.4) was used to draw a volcano plot of DEGs.
GO and KEGG enrichment analyses were assessed using the “clusterProfiler” package (version 4.10). A BH-adjusted P-value < 0.05 was used as the threshold to determine significant enrichment of the gene sets.
To estimate the immune infiltration in mouse tissues, the seq-ImmuCC method [https://github.com/wuaipinglab/ImmuCC] was used to assess the relative proportions of 10 major immune cell types from RNA-Seq data of mouse tissues. To quantify neutrophil infiltration, we collected signature genes from Bindea et al. MCPCOUNTER, and QUANTISEQ to calculate the neutrophil infiltration score.
Single-Cell RNA sequencing and data preprocessing
According to the research purpose and animal ethical management standards, fresh liver tumor samples meeting the requirements of the project were obtained from a mouse animal model to prepare a single-cell suspension according to reference paper26. CD45+ cells were then obtained using CD45 microbeads (Miltenyi Biotec, 130-052-301). Precisely, 6 × 103 cells were loaded onto the 10 × Genomics single cell A chip. After droplet generation, samples were transferred onto a pre-chilled 8-well tube, heat-sealed, and reverse transcription was performed using a Veriti 96-well thermal cycler (Thermo Fisher), followed by Dynabeads cDNA cleanup and amplification (12 cycles). cDNA libraries were prepared as outlined by the Single Cell 3’ Reagent Kits v3 user guide with appropriate modifications to the PCR cycles based on the calculated cDNA concentration (as recommended by 10× Genomics). The molarity of each library was calculated based on library size as measured using a Bioanalyzer (Agilent Technologies) and qPCR amplification data. Samples were pooled and normalized to 10 nM, then diluted to 2 nM using elution buffer with 0.1% Tween20 (Sigma). Samples were sequenced on a Novaseq 6000 with the following run parameters: Read1 26 cycles, Read2 98 cycles, index 1-8 cycles.
Raw sequencing data (bcl files) were converted to fastq files using Illumina bcl2fastq, version 2.19.1, and aligned to the human genome reference sequence (http://cf.10xgenomics.com/supp/cell-exp/refdata-cellranger-GRCh38-1.2.0.tar.gz). The CellRanger (10× Genomics) analysis pipeline was used to generate a digital gene expression matrix from this data. The raw digital gene expression matrix (UMI counts per gene per cell) was filtered, normalized, and clustered using R [https://www.R project.org/]. Cell and gene filtering was performed as follows: cells with a very small library size (<1500) and a very high (> 0.5) mitochondrial genome transcript ratio were removed. To ensure robust gene expression signals, genes detected in fewer than three cells (with a UMI count > 0) were excluded. Data normalization was subsequently performed to reduce technical biases, employing the scran R package through its default pooled deconvolution method. Following normalization, cell clustering was conducted using standardized protocols from established software packages. The enrichment of cellular gMDSC was quantified using the AddModuleScore function from the Seurat package (version 5.0.1)42.
To reconstruct potential differentiation trajectories, we utilized Monocle 3 (version 0.2.3.0)43. Cell ordering along inferred pseudotemporal paths was performed by first applying dimensionality reduction using the DDRTree method (reduceDimension), followed by trajectory inference (orderCells).
For GSEA, mouse gene symbols were firstly converted to human gene symbols using the “biomaRt” package (version 2.58.1). Gene sets were downloaded from the MSigDB database. Data were analyzed using GSEAv4.0.2 software or the “clusterProfiler” package (version 4.10) in R 4.3.2.
Flow cytometry
Single-cell suspensions were prepared from liver tissues using the Tumor Dissociation Kit (130-096-730, Miltenyi Biotec), from lung tissues using the Lung Dissociation Kit (130-095-927, Miltenyi Biotec), and from bone marrow and peripheral blood using a dissociation kit (R1010, Solarbio), according to the manufacturer’s protocol. The cells were incubated with antibodies against Cd45 (CD45 PE-CY5, 1:100, 15-0451-81, Invitrogen), Cd11b (CD11b FITC, 1:100, AH011B01, Multi sciences; 1:100, 17-0112-82, Invitrogen), and Ly6G (Ly6G APC-CY7, 1:100, 127623, Biolegend; 1:100, 127606, Biolegend) to identify neutrophils.
For the identification of Cd177+Ly6G+ neutrophils, the cells were incubated with antibodies against Cd45 (CD45 PE-CY5, 1:100, 15045-81, Invitrogen), Cdd11b (CD11b FITC, 1:100, AH011B01, Multi sciences), Ly6G (Ly6G APC-CY7, 1:100, 127623, Biolegend), and Cd177 (CD177 PE, 1:100, fab8186p, R&D). For the pErk+Ly6G+ neutrophils test, the cells were incubated with Cd45 (CD45 PE-CY5, 1:100, 15045-81, Invitrogen), Cd11b (CD11b FITC, 1:100, AH011B01, Multi sciences), Ly6G (Ly6G APC-CY7, 1:100, 127623, Biolegend), and pErk (pERK Alexa Fluor 647, 1:100, 369504, Biolegend) antibodies. Incubation was performed for 30 minutes at 4 °C in the dark. Neutrophils were then measured by flow cytometry (FACSMelody, BD, USA) and analyzed using Flowjo software.
To assess neutrophil apoptosis in tumor tissues, single-cell suspensions were prepared from mouse tumor tissues as described above. Antibodies against Cd45 (1:100, 103133, Biolegend), Cd11b (1:100, 101245, Biolegend) and Ly6G (1:100, 70-F21LY6G03-100, Multi Sciences) were added according to the flow cytometry antibody instructions and incubated on ice for 30 minutes. Following a wash with pre-cooled PBS, the cells were resuspended in 1 × Binding Buffer from the apoptosis detection kit (70-AP101-100, Multi Sciences). Subsequently, 5 µL of Annexin V-FITC and 10 µL of PI were added to each tube. After gentle mixing, the samples were incubated at room temperature in the dark for five minutes before proceeding to flow cytometric analysis.
All flow cytometry gating strategies have been illustrated in Supplementary Fig. 2G. In Fig. 2B, Fig. 3A, Supplementary Fig. 2H, Supplementary Fig. 2I, Supplementary Fig. 3G, and Supplementary Fig. 10C, similar to Supplementary Fig. 2G, we first circled all cells morphologically similar to immune cells based on FSC and SSC, selected all individual cell populations among these cells, labeled all Cd45-positive immune cells, and finally identified CD11b- and Ly6G-positive neutrophils.
Neutrophil subtype isolation
Single cell suspension was prepared from liver tissue using the Tumor Dissociation Kit (130-096-730, Miltenyi Biotec), and then, a neutrophil suspension was obtained by neutrophil isolation liquid (P2430, Solarbio). Neutrophils were maintained in RPMI-1640 medium (Hyclone, Logan, UT, USA) with 10% fetal bovine serum (FBS) (GIBCO BRL, Invitrogen, Grand Island, NY, USA) at 37 °C in 5% CO2. For western blotting, after incubation with Cd45 (CD45 PE-CY5, 15-0451-81, Invitrogen) and Cd177 (CD177 PE, fab8186p, R&D) antibodies, the cell membrane was lysed using the eBioscienceTM Foxp3/Transcription factor staining buffer set (00-5523-00, Invitrogen). Then, the Mki67 antibody (151211, Biolegend) was used to isolate Cd45+Cd177-, Cd45+Cd177+Mki67+, and Cd45+Cd177+Mki67- neutrophils by flow cytometry (FACSMelody, BD, USA).
For co-culture with tumor cells, the neutrophils were successively incubated with anti-S100a6 (1:100, ab181975, Abcam) and Alexa Fluor 488 anti-rabbit (1:500, ab150077, Abcam) for 30 min at room temperature while protected from light. Then, Cd45 (15-0451-81, Invitrogen) and Cd177 (fab8186p, BD) antibodies were used to isolate Cd45+Cd177-, Cd45+Cd177+S100a6+, and Cd45+Cd177+S100a6- neutrophils by flow cytometry (FACSMelody).
Western blotting
Briefly, cultured cells and tumor tissues were collected and lysed in lysis buffer mixture (FD009, Fdbio science). Then tissues were ground and centrifuged for supernate. Protein concentration was measured by BCA Protein Assay Kit (KeyGEN BioTECH). 20–30 μg total protein was separated by 8–10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), transferred to polyvinyl difluoride (PVDF) membranes, and blocked in 5% BSA (diluted by TBS supplemented with 0.1% Tween-20) for 1 h at room temperature. After incubation with the following primary antibodies: Pad4 (1:800, ab96758, Abcam), Histone H3 (citrulline R2 + R8 + R17) (1:800, ab5103, Abcam), S100a6 (1:800, ab181975, Abcam), PPARα (1:800, YT3835, Immunoway), PPARβ/δ (1:800, 16643-1-AP, Proteintech), PPARγ (1:800, 10156-2-AP, Proteintech), CD36 (1:1000, 18836-1-AP, Proteintech), FABP4 (1:1000, 12802-1-AP, Proteintech), FABP5 (1:1000, 12348-1-AP, Proteintech), p-PI3K (1:1000, #11889, Cell Signaling Technology), p-AKT (1:1000, #4060, Cell Signaling Technology), p-mTOR (1:1000, #5536, Cell Signaling Technology), p-ERK1/2 (Thr202/Tyr204) (1:1000, #4370, Cell Signaling Technology), p-SAPK/JNK (Thr183/Tyr185) (1:1000, #4668, Cell Signaling Technology), and p-P38 MAPK (1:1000, Thr180/Tyr182) (#4511, Cell Signaling Technology), at 4 °C overnight, the membranes were subsequently incubated with IRDye® 680RD goat anti-mouse (1:10000, P/N 926-68070, LI-COR) or IRDye® 800CW goat anti-rabbit (1:10000, P/N 926-32211, LI-COR) for 1 h at room temperature. The signals were captured by Odyssey Infrared Imaging System (Li-Cor Bioscience, USA). The antibody against Tubb (1:5000, FD0064, Fdbio science), β-actin (1:5000, 60009-1-Ig, Proteintech), and Histone H3 (1:5000, A01502, GenScript) were used as the loading control.
The samples derive from the same experiment, but different gels were processed in parallel. Figure 3C: Different gels for Pad4, cit-H3, Tubb, one for S100a6 and Actb were processed in parallel. Figures 5C, D: Different gels for PPARα and Histone H3, one for PPARγ, and another for PPARβ/δ and Actb were processed in parallel. Figure 5E–G and Supplementary Fig. 7E: Different gels for CD36 and Actb, one for FABP4, and another for FABP5 were processed in parallel. Figure 6A: Different gels for p-mTOR, p-Akt, p-P38, and Actb, and another for mTOR, Akt, P38, and Actb were processed in parallel. Supplementary Fig. 8A: Different gels for Pi3k, Jun, Erk1/2, and Actb, and another for p-Pi3k, p-Jun, and p-Erk1/2 were processed in parallel. Supplementary Fig. 8C: Different gels for mTOR, p-mTOR, P38, p-P38, JUN, ERK1/2, AKT, and ACTB, and another for p-ERK1/2, PI3K, p-PI3K, p-JUN, p-AKT, and ACTB were processed in parallel. Figures 6C, G, H, and Supplementary Fig. 8G: Different gels for p-P38 and another for P38 and Tubb were processed in parallel. Figure 7F: Different gels for Erk1/2 and Tubb, and another for pErk1/2 and Tubb were processed in parallel. Figure 7G–I and Supplementary Fig. 9G, J, and K: Different gels for Mek1/2, Erk1/2, cit-H3, and another for pMek1/2, pErk1/2, and Gapdh were processed in parallel.
Transwell
Neutrophils were isolated and placed (2 × 104) in the upper chambers of the 0.8 μm Transwell 24-well culture inserts (Corning Costar, Corning, NY, USA). Then, CT26 cells (10 × 104) were added to the lower compartment. After 24 h, neutrophils adhered to the low surface were fixed, washed, and stained with 0.1% crystal violet for 30 min. Representative images were obtained by microscopy (Bx51, Olympus, Japan).
CT26 cells (2 × 104) were placed in the upper chambers of 0.8 μm Transwell 24-well culture inserts. Then, separated neutrophil cells (1 × 105) were added to the lower compartment. After 24 h, CT26 cells adhered to the low surface were fixed, washed, and stained with 0.1% crystal violet for 30 minutes to assess their invasion ability.
MPO-DNA ELISA for NET detection
In brief, 50 μl of plasma samples collected from BALB/c mice and patients, and cell supernate were added to the 96-well plate coated with anti-MPO antibody (Human: JL11580-96T, Jianglaibio; Mouse: JL10367-96T, Jianglaibio)44. After incubated for 1 h at room temperature, the plate was washed five times with wash buffer. Next, 100 μl of PicoGreen reagent from Quant-iTTM PicoGreen dsDNA Assay Kit (#P7589, InvitrogenTM) was added to the wells for 5 min at room temperature with protecting from light. The PicoGreen dye that was bound to dsDNA was measured using a fluorescence microplate reader (SpectraMax M5, Molecular Devices) and a multimode microplate reader (SYNERGY H1, BioTek). The following settings were used to read the PicoGreen results: Excitation wavelength and bandwidth were 485 nm and 14 nm, respectively. Emission wavelength and bandwidth were 535 nm and 25 nm, respectively.
To evaluate the effect of oleic acid on the formation of NETs, the following method for cell treatment was employed: the isolated human peripheral blood neutrophils or mouse bone marrow neutrophils were first evenly seeded at a density of 2.5 × 105 cells per well in a 24-well cell culture plate. Subsequently, the culture plate was supplemented with either RPMI-1640 medium, 3 µM oleic acid (oleic acid-BSA aqueous solution, Pythonbio, AAPR-K5), or 200 nM PMA (PMA, Selleck, S7791) to stimulate the neutrophils for 12 hours. After stimulation, the supernatant was collected from the cell cultures and analyzed for MPO-dsDNA content.
NET immunofluorescence
Before seeding neutrophils, round glass coverslips (13 mm) were plated into the 24-well plate. 5 × 105 neutrophils were then seeded in each well in RPMI-1640 medium supplemented with tumor cell supernate (1:1), tumor tissue supernate (1:1), DGAT1 inhibitor (PF-04260110, S719, Selleck, USA) or MEK inhibitor (Trametinib, GSK1120212, Selleck, USA) for 12 h at 37 °C. Tumor cells were centrifuged at 1000 g for 10 min after 24 h culture for tumor cell supernate. Tumor tissues were collected and cultured with RPMI-1640 medium for 24 h, and then tumor tissue supernate was obtained by centrifuged at 1000 g for 10 min. After fixed with 4% paraformaldehyde, the glass coverslips with attached neutrophils were carefully removed and put upside down on a drop of PBS for thrice washing. Incubate the coverslips with primary antibodies: anti-Histone H3 (citrulline R2 + R8 + R17) (1:200, ab5103, Abcam) and anti-Ly6G (1:200, ab9535, Abcam), or anti-MPO (1:200, ab300650, Abcam) at 4 °C overnight. Next day, the coverslips were incubated with fluorescent secondary antibodies: Alexa Fluor 488-labeled goat anti-Rabbit (1:500, ab150073, Abcam) and Cy3-labeled goat anti-Rat (1:500, ab6953, Abcam) for 1 h following washing with PBS. Set a 10 μl drop of anti-quenching agent onto a glass slide and mount coverslips with cells upside down. Representative images were acquired with microscopy (Bx63, Olympus, Japan). Nuclei were stained with DAPI for 5 min at room temperature.
Lipidomics
Freshly collected 37 °C-CT26 and 45 °C-CT26 cells (1 × 107 cells/per sample, n = 6) were resuspended in 1 mL of PBS, snapped frozen and saved at −80 °C until processing. To the sample 1 mL of lipid extract was added, followed by votexing and centrifugation at 13 000 × g for 10 min at 4 °C. The organic fraction was transferred to a clean vial and dried under N2, followed by reconstitution in 100 mL acetonitrile (ACN) / isopropanol (IPA). For Untargeted Lipidomics LC-MS, reconstituted extracts were injected into a Ultra Performance Liquid Chromatography (UPLC) (Shim-pack UFLC SHIMADZU CBM30A) coupled to a Tandem mass spectrometry (QTRAP® 6500 + ). Lipids were separated using a Thermo C30 column (2.1 × 100 mm) with the following solvents A: 5 mmol/L ammonium formate with 0.04% acetic acid in 60:40 ACN: water and B: 5 mmol/L ammonium formate with 0.04% acetic acid in 10:90 ACN: IPA at a flow rate of 0.35 mL/min. The gradient program was 0 min A/B (80:20 V/V), 3 min 50:50 V/V, 5 min 35:65 V/V, 9 min 25:75 V/V, and 15.5 min 10:90 V/V. The mass spectrometer was equipped with an ESI source. Statistically significant was defined according to the criteria of | log2(fold change) | > 1.
Oil Red O staining
OCT-embedded frozen tissues were prepared for Oil Red O staining45. The slices were washed with distilled water several times, preincubated in 60% isopropanol, and finally stained with filtered Oil Red O working solution [60% Oil Red O stock solution (Beijing Solarbio Science & Technology Co., Ltd., China) and 40% deionized water]. After several washing in 60% isopropanol, the nuclei were counterstained with hematoxylin, and the slices were differentiated in 1% hydrochloric acid in alcohol. Finally, after sealed with glycerin-gelatin, representative images were captured using a microscope (Bx51, Olympus, Japan).
Collection of supernatants
CT26-iRFA and CT26-Ctrl mouse models were established. Twenty-four hours after treatment, normal liver tissues following iRFA (iRFA-L), residual tumors after iRFA (iRFA-TCS or iRFA-T), necrotic tissues after iRFA (iRFA-NCS or iRFA-N), as well as untreated liver tissues (Ctrl-L) and tumors (Ctrl-TCS or Ctrl-T) were dissected and cocultured with 10% fetal bovine serum for another 24 h. Conditioned media were then collected for co-culturing with neutrophils for 12 h or for the detection of FA and TG.
CT26 and DLD1 cancer cells were maintained in RPMI-1640 medium (Hyclone, Logan, UT, USA) with 10% fetal bovine serum (FBS) (GIBCO BRL, Invitrogen, Grand Island, NY, USA) at 37 °C in 5% CO2. After growing to 60-70% confluence, cells were incubated with preheated OA (OA:10% FBS = 1:10000, 0.31509 mM) (112-80-1, Sigma-Aldrich) or 10% FBS, and then transferred to 45 °C for 15 min stimulation and subsequently to 37 °C for 48 h culture. The supernatant was collected as 45 °C + OA or 45 °C TCM. For 37 °C and 37 °C + OA TCM, cancer cells were treated without thermal stimulation.
Free fatty acid and triglyceride test
Before each test, the tissue and cell samples were prepared. For tissues samples, the indicated tissues were removed and weighted. Normal saline was added (9 mL/g) to make homogenate. After centrifuged at 1250 × g for 10 min, the supernate was collected. For cell samples, CT26 and DLD1 cells were harvested by trypsinization after indicated treatment, and resuspended (1 × 106 cells/mL) in a mixture of PBS. After centrifuging at 200 × g for 10 min, the supernate was discarded and 100 μl 2% TritonX-100 was added. Cell homogenate was made using Ultrasonic Cell Disruptor (3000 Walt, 3-5 s). Then gently rock the tube at 4 °C for another 30-40 min using a shaking platform. Keep a small aliquot of the extract to quantitate the protein concentration.
Next, the content of free fatty acid and triglycerid in tissue and cell samples were tested using a Nonesterified Free fatty acids assay kit and Triglyceride assay kit (Nanjing Jiancheng Bioengineering Institute, Jiangsu, China) following the manufacturer’s instructions. The OD values were recorded at 546 nm and 510 nm using a microplate reader (SpectraMax M5).
Intracellular fatty acid determination
For intracellular FA content, cells were pretreated with BODIPY™ 493/503 (D3922, Invitrogen). Images were taken under confocal microscopy (Ti2-E, Nikon, Japan) using an excitation wavelength of 493 nm and an emission light at 504 nm. For FA uptake in tumor cells, CT26 and DLD1 cells were pretreated with BODIPY™ FL C16 (D3821, Invitrogen). Images were taken under confocal microscopy (Ti2-E, Nikon, Japan) using an excitation wavelength of 488 nm with the same laser intensity.
Seahorse metabolic profiling experiments
The metabolic profiling was performed following the instruction of Seahorse mitochondrial stress kits. CT26 or DLD1 cells (2 × 104 cells/per well or 3 × 104 cells/per well) were seeded in triplicate in an XF96-well culture plate, and the growth medium was supplemented with or without oleic acid (0.3 mM). One hour after cell incubation, the cells were exposed to heat shock at 45 °C for 15 min in the heated group. After 24 h incubation at 37 °C in 5% CO2, the medium from each well was carefully removed and replaced with 180 μl of assay medium. The assay medium was freshly prepared, and the pH was adjusted to 7.4. The oxygen consumption rate (OCR) was measured using Seahorse XFp analyzer (Seahorse Bioscience, USA), and oligomycin (1 μM), carbonyl cyanide-4-trifluoro methoxy phenyl hydrazone (FCCP) (1 μM), and rotenone/antimycin A (0.5 μM) were sequentially added to each well. The data were analyzed by Wave software.
Fatty acid oxidation assay
According to the instructions provided by FAOBlue (Fatty Acid Oxidation Detection Reagent, FDV-0033, Funakoshi), CT26 or DLD1 cells were uniformly planted on 24-well plates at a density of 2 × 105 cells per well. Corresponding stimulation was added during the logarithmic growth phase of the cells. After 24 hours, the cells were washed twice with serum-free RPMI-1640 medium. Subsequently, RPMI-1640 medium containing 20 µM FAOBlue was added; after an additional 24-hour incubation period, the cells were washed again with serum-free RPMI-1640 medium and observed using fluorescence microscopy. The mean fluorescence intensity was quantified for analysis.
Measurement of ROS
Cells were seeded on 6-well plates at a density of 2 × 105 cells per well. ROS was measured through flow cytometry by fluorescent 2’,7’-dichlorofluorescin diacetate (DCF-DA) as described in the manufacturer’s protocol in the commercial kit (S0033S, Beyontime, China). Flow cytometry was performed at 488 mm excitation wavelength and 525 mm emission wavelength.
Quantitative real-time PCR
RNAiso Plus reagent (TaKaRa, Japan) was used to extract total RNA from cultured CT26 and DLD1 cells according to manufacturer’s instruction, and RNA (2 μg) was transcribed to cDNA using PrimeScriptTM RT Master Mix (TaKaRa, Japan). Quantitative real-time PCR was performed with LightCycler 480 system (Roche Diagnostics, USA) using SYBR Premix Ex Taq II (Tli RNaseH Plus) (TaKaRa, Japan). The primers used in our experiments are listed in the Table 1. β-actin was used as the reference gene for normalization. The relative gene expression was calculated according to the 2-ΔΔCt method.
PPAR binding assay
Nucleus fractions was isolated from cultured cells using Nuclear and Cytoplasmic Protein Extraction Kit (KeyGEN BioTECH), and used in PPARα, β/δ, γ Complete Transcription Factor Assay (Abnova). The procedures were performed following the manufacturer’s protocols. The OD values were recorded at 450 nm using a microplate reader (SpectraMax M5).
Nile red staining
CT26 cells were seeded into the 6-well plate with 30% confluence, followed by indicated treatment. Next day, the cells were incubated with Nile red working solution (Beijing Solarbio Science & Technology Co., Ltd., China) for 10 min at 37 °C protecting from light. Representative images were captured using a microscope (Ti2-E, Nikon, Japan).
ELISA
The levels of Cxcl5 and Il1b in the supernatant of CT26, MC38 and DLD1 cells (1 × 106 cells/mL) were quantified by a commercially purchased enzyme-linked immunoabsorbent assay (ELISA) kit used according to the manufacturer’ s instructions (Jianglaibio, China). Briefly, 100 μl supernate were added to the 96-well plate. After incubated for 90 min at room temperature, the plate was washed five times with wash buffer. Next, 100 μl of biotinylation antibody was added for 60 min incubation. Finally, the OD values were recorded at 450 nm using a microplate reader (SpectraMax M5).
Data acquisition and analysis of TCGA
Fragments Per Kilobase per Million (FPKM) data and clinical information of Colorectal adenocarcinoma (TCGA-COREAD cohort) were acquired from the UCSC Xena website (https://xenabrowser.net/). FPKM data were converted to Transcripts PerKilobase Million (TPM) values and used to calculate enrichment score by the single-sample GSEA (ssGSEA). Gene sets of neutrophils46 and BIOCARTA_P38MAPK_PATHWAY downloaded from MSigDB were used to compute neutrophils infiltration score and P38 MAPK pathway enrichment score separately. Scatter plot was created by package “ggpubr” (version 0.6.0), and Pearson’s correlation coefficients r was calculated to characterize a linear relationship between the two variables. Kaplan-Meier plots were generated using R packages “survival” (version 3.5) and “survminer” (version 0.4.9).
CTC detection
Through the CanPatrolTM CTC detection technology, a filter-based method combined with RNA in situ hybridization (RNA-ISH) method that was based on the branched DNA signal amplification technology, CTCs were isolated for categorization47. The methods are summarized as follows: before enriching CTCs, red blood cell lyses buffer (Surexam Biotech, Guangzhou, China) was used to lyse the erythrocytes in the blood specimens followed by centrifugation to remove the supernatant and addition of 4% formaldehyde (Sigma, St. Louis, USA) to fix the remaining cells. Subsequently, the cell suspension of each blood specimen was transferred to a filtration tube that which contained a calibrated membrane with 8-μm diameter pores (Millipore, Billerica, USA) to remove the eluent from the filtration tube using a vacuum pump.
The membrane was treated with a protease (Qiagen, Hilden, Germany) and then hybridized with three groups of capture probes. Group 1 probes were specific for the epithelial biomarkers, epithelial cell adhesion molecules (EpCAM) and cytokeratins (CK) 8/18/19. Group 2 probes had two mesenchymal biomarkers, vimentin and twist. Group 3 probes had the lipid transporter biomarker, CD36. Hybridization was performed at 42 °C for 2 h, followed by incubating with preamplifier solution (Surexam Biotech, Guangzhou, China), amplifier solution (Surexam Biotech, Guangzhou, China), and fluorescently-labeled probes. Isolated CTCs were stained with 4’, 6-diamidino-2-phenylindole (DAPI) (Sigma, St. Louis, USA) and counted under fluorescence microscope (Olympus BX53, Tokyo, Japan).
Results interpretation: The epithelial CTCs were displayed in the red fluorescent channel, the mesenchymal CTCs in the green fluorescent channel, while a mixture of two types was displayed in both red and green fluorescent. The CTCs isoforms with lipid metabolism showed purple fluorescence.
Statistic and reproducibility
The experiments were analyzed according to the instructions provided in the Figure legend, and the data were presented as mean ± SD. The sample sizes are denoted in the figure legends. Each experiments was performed in triplicate. The images represent n = 3 independent replicates of experiments with similar results.
The Kaplan-Meier method was used to measure survival. All statistical analyses were evaluated using Student’s t-test, Paired t-test, Mann-Whitney U test, one-way ANOVA with LSD or Tamhane/Dunnett post-hoc test, depending on the case, conducting with SPSS 22.0 for Windows (SPSS Inc., Chicago, IL, USA) or Prism 9 software (GraphPad). P < 0.05 was considered statistically significant.
Ethics statement
This study adheres to all pertinent ethical guidelines and regulations. This study received approval from the Ethics Committee of NangFang Hospital, Southern Medical University (NFEC-2017-206). For the utilization of human tissue and blood specimens in this research, informed consent was obtained from all participants. The animal care protocols and experimental procedures were reviewed and approved by the Animal Ethics Committee of Nanfang Hospital, Southern Medical University.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq raw data from colon cancer tumor tissue in the livers of iRFA-treated mice, as well as those from human samples collected before and after iRFA, have been deposited in the Genome Sequence Archive (GSA) at the National Genomics Data Center (NGDC), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences. These data are accessible under project PRJCA033119. This study complies with the “Guidance of the Ministry of Science and Technology (MOST) for the Review and Approval of Human Genetic Resources”. All required approvals from the MOST of China related to this data have been secured (Approval No. 2025BAT00980). TCGA Level 3 RNA-seq data and clinical information for CRC patients in the TCGA-COREAD dataset were obtained from the UCSC Xena platform (https://xenabrowser.net/datapages/?cohort=TCGA%20Colon%20and%20Rectal%20Cancer%20(COADREAD)&removeHub=https%3 A%2 F%2Fxena.treehouse.gi.ucsc.edu%3A443]. The lipidomics data generated in this study have been deposited in the Mendeley database (https://data.mendeley.com/datasets/jdw9tbv9jd/1). Source data are provided with this paper.
Code availability
The analyses in this study were performed using standard and publicly available R packages (Table 2), as detailed in the Methods section. No custom code essential for reproducing the results was developed. Therefore, no additional code is available for public sharing.
References
Manfredi, S. et al. Epidemiology and management of liver metastases from colorectal cancer. Ann. Surg. 244, 254–259 (2006).
Goldberg, S. N. et al. Image-guided tumor ablation: proposal for standardization of terms and reporting criteria. Radiology 228, 335–345 (2003).
Ruers, T. et al. Local Treatment of Unresectable Colorectal Liver Metastases: Results of a Randomized Phase II Trial. J Natl Cancer Inst. 109, (2017).
Wu, J. et al. Radiofrequency ablation: mechanisms and clinical applications. MedComm (2020) 5, e746 (2024).
Sucandy, I. et al. Longterm survival outcomes of patients undergoing treatment with radiofrequency ablation for hepatocellular carcinoma and metastatic colorectal cancer liver tumors. HPB (Oxf.) 18, 756–763 (2016).
Tanis, E. et al. Local recurrence rates after radiofrequency ablation or resection of colorectal liver metastases. Analysis of the European Organisation for Research and Treatment of Cancer #40004 and #40983. Eur. J. Cancer 50, 912–919 (2014).
Shi, L. et al. Inflammation induced by incomplete radiofrequency ablation accelerates tumor progression and hinders PD-1 immunotherapy. Nat. Commun. 10, 5421 (2019).
Wu, H. et al. Palliative radiofrequency ablation accelerates the residual tumor progression through increasing tumor-Infiltrating MDSCs and reducing T-cell-mediated anti-tumor immune responses in animal model. Front Oncol. 10, 1308 (2020).
Zhang, W. et al. CXCL5/CXCR2 axis in tumor microenvironment as potential diagnostic biomarker and therapeutic target. Cancer Commun. (Lond.) 40, 69–80 (2020).
Hedrick, C. C. & Malanchi, I. Neutrophils in cancer: heterogeneous and multifaceted. Nat. Rev. Immunol. 22, 173–187 (2022).
Raza, A. et al. Dynamic liquid biopsy components as predictive and prognostic biomarkers in colorectal cancer. J. Exp. Clin. Cancer Res 41, 99 (2022).
Fumagalli, A. et al. Plasticity of Lgr5-negative cancer cells drives metastasis in colorectal cancer. Cell Stem Cell 26, 569–578 (2020).
Lalezari, P., Murphy, G. B. & Allen, F. J. NB1, a new neutrophil-specific antigen involved in the pathogenesis of neonatal neutropenia. J. Clin. Invest. 50, 1108–1115 (1971).
Dahlstrand, R. A. et al. The neutrophil subset defined by CD177 expression is preferentially recruited to gingival crevicular fluid in periodontitis. J. Leukoc. Biol. 109, 349–362 (2021).
Lewis, H. D. et al. Inhibition of PAD4 activity is sufficient to disrupt mouse and human NET formation. Nat. Chem. Biol. 11, 189–191 (2015).
Adrover, J. M., McDowell, S., He, X. Y., Quail, D. F. & Egeblad, M. NETworking with cancer: The bidirectional interplay between cancer and neutrophil extracellular traps. Cancer Cell 41, 505–526 (2023).
Bayik, D. et al. Myeloid-derived suppressor cell subsets drive glioblastoma growth in a sex-specific manner. Cancer Discov. 10, 1210–1225 (2020).
Wang, Y. et al. Histone hypercitrullination mediates chromatin decondensation and neutrophil extracellular trap formation. J. Cell Biol. 184, 205–213 (2009).
Costantino, S. et al. Obesity-induced activation of JunD promotes myocardial lipid accumulation and metabolic cardiomyopathy. Eur. Heart J. 40, 997–1008 (2019).
Peters, J. M., Shah, Y. M. & Gonzalez, F. J. The role of peroxisome proliferator-activated receptors in carcinogenesis and chemoprevention. Nat. Rev. Cancer 12, 181–195 (2012).
Yoshida, S. et al. Sublethal heat treatment promotes epithelial-mesenchymal transition and enhances the malignant potential of hepatocellular carcinoma. Hepatology 58, 1667–1680 (2013).
Cruz, A., Barreto, E. A., Fazolini, N., Viola, J. & Bozza, P. T. Lipid droplets: platforms with multiple functions in cancer hallmarks. Cell Death Dis. 11, 105 (2020).
Martin-Perez, M., Urdiroz-Urricelqui, U., Bigas, C. & Benitah, S. A. The role of lipids in cancer progression and metastasis. Cell Metab. 34, 1675–1699 (2022).
Papayannopoulos, V. Neutrophil extracellular traps in immunity and disease. Nat. Rev. Immunol. 18, 134–147 (2017).
Xiong, S., Dong, L. & Cheng, L. Neutrophils in cancer carcinogenesis and metastasis. J. Hematol. Oncol. 14, 173 (2021).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Engblom, C. et al. Osteoblasts remotely supply lung tumors with cancer-promoting SiglecF(high) neutrophils. Science. 358, eaal5081 (2017).
Zheng, C. et al. Dual role of CD177 + neutrophils in inflammatory bowel disease: a review. J. Transl. Med. 22, 813 (2024).
Luo, X. et al. Emerging roles of lipid metabolism in cancer metastasis. Mol. Cancer 16, 76 (2017).
Min, K. J., Um, H. J., Cho, K. H. & Kwon, T. K. Curcumin inhibits oxLDL-induced CD36 expression and foam cell formation through the inhibition of p38 MAPK phosphorylation. Food Chem. Toxicol. 58, 77–85 (2013).
Lu, Y. et al. p38 MAPK-inhibited dendritic cells induce superior antitumour immune responses and overcome regulatory T-cell-mediated immunosuppression. Nat. Commun. 5, 4229 (2014).
Liu, Y. et al. Telmisartan inhibits oxalate and calcium oxalate crystal-induced epithelial-mesenchymal transformation via PPAR-γ-AKT/STAT3/p38 MAPK-Snail pathway. Life Sci. 241, 117108 (2020).
Pranteda, A., Piastra, V., Stramucci, L., Fratantonio, D. & Bossi, G. The p38 MAPK signaling activation in colorectal cancer upon therapeutic treatments. Int. J. Mol. Sci. 21, 2773 (2020).
Corcoran, R. B. et al. Combined BRAF, EGFR, and MEK Inhibition in patients with BRAF(V600E)-mutant colorectal cancer. Cancer Discov. 8, 428–443 (2018).
Cañellas-Socias, A. et al. Metastatic recurrence in colorectal cancer arises from residual EMP1(+) cells. Nature 611, 603–613 (2022).
Neuhöfer, P. et al. Acinar cell clonal expansion in pancreas homeostasis and carcinogenesis. Nature 597, 715–719 (2021).
Alicea-Torres, K. et al. Immune suppressive activity of myeloid-derived suppressor cells in cancer requires inactivation of the type I interferon pathway. Nat. Commun. 12, 1717 (2021).
Cedervall, J. et al. Pharmacological targeting of peptidylarginine deiminase 4 prevents cancer-associated kidney injury in mice. OncoImmunology 6, e1320009 (2017).
Coffelt, S. B. et al. IL-17-producing gammadelta T cells and neutrophils conspire to promote breast cancer metastasis. Nature 522, 345–348 (2015).
Teijeira, Á et al. CXCR1 and CXCR2 chemokine receptor agonists produced by tumors induce neutrophil extracellular traps that interfere with immune cytotoxicity. Immunity 52, 856–871 (2020).
Huang, N. et al. Gold nanoparticles induce tumor vessel normalization and impair metastasis by inhibiting endothelial Smad2/3 signaling. ACS Nano 14, 7940–7958 (2020).
Wilk, A. J. et al. A single-cell atlas of the peripheral immune response in patients with severe COVID-19. Nat. Med. 26, 1070–1076 (2020).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Vong, L., Sherman, P. M. & Glogauer, M. Quantification and visualization of neutrophil extracellular traps (NETs) from murine bone marrow-derived neutrophils. Methods Mol. Biol. 1960, 63–73 (2019).
Marfella, R. et al. Lipid accumulation in hearts transplanted from nondiabetic donors to diabetic recipients. J. Am. Coll. Cardiol. 75, 1249–1262 (2020).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795 (2013).
Wu, S. et al. Classification of circulating tumor cells by epithelial-mesenchymal transition markers. PLoS One 10, e123976 (2015).
Acknowledgements
This study was supported by grants from the National Natural Science Foundation of China (82173187 to N.H., 82002555 to J.W., and 82272635 to W.L.), the Science and Technology Projects in Guangzhou (2023A04J2360 to Y.F.), the Guangdong Basic and Applied Basic Research Foundation (2023A1515110893 to Y.F.), the Natural Science Foundation of Guangdong Province (2021A1515011705 to N.H.), the Outstanding Youths Development Scheme of Nanfang Hospital, Southern Medical University (2022J005 to N.H.), and the Science and Technology Projects in Guangzhou (202201011433 to S.Z.). We appreciate the sequencing service provided by GenePlus.
Author information
Authors and Affiliations
Contributions
W.L., M.S., N.H., and Z.W. designed the study. W.L., M.S., N.H., Y.F., S.Z., and J.W. interpreted results and drafted the paper. N.H., Y.F., S.Z., G.H., Y.G., B.X., Y.H., and X.D. prepared all figures. N.H., Y.F., S.Z., J.W., G.H., Y.G., B.L., and D.Z. performed experiments and/or analyzed data. X.R. and C.W. collected and analyzed clinical data. W.L., M.S., J.B., and Y.L. supervised the study. All authors reviewed, edited, or commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, N., Fang, Y., Zheng, S. et al. Residual tumor cells after insufficient radiofrequency ablation promote lung metastasis by educating CD177hiPAD4hi neutrophils. Nat Commun 17, 208 (2026). https://doi.org/10.1038/s41467-025-66897-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66897-0