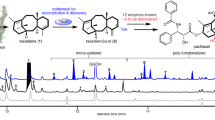
Abstract
Paclitaxel is one of the most prominent drugs for cancer therapy, and its synthesis has long been pursued by scientific community. Although the biosynthetic pathway has been gradually unraveled, its synthetic efficiency remains severely constrained by the low activity and low product selectivity of the hydrolases involved. In this study, an artificial toolbox of oxidases derived from mutants of TteUPO (unspecific peroxygenase from Thielavia terrestris) is developed for site-directed oxidation of taxanes at the C-4, C-6, C-10, C-11, C-12, and C-13 positions. Furthermore, de novo biosynthesis of oxidized taxadiene products is achieved in an engineered E. coli chassis. These findings offer an unconventional strategy for developing an efficient route toward paclitaxel and constructing paclitaxel analogs with different structures for medicinal evaluation.
Similar content being viewed by others
Introduction
Paclitaxel (Taxol®), a tetracyclic and functionally condensed diterpenoid, is renowned for its extensive anticancer efficacy. Its structural derivatives are widely utilized in the treatment of various malignancies (Fig. 1)1. Docetaxel and cabazitaxel, both classified alongside paclitaxel as first-generation taxane anticancer drugs, have therapeutic applications distinct from those of paclitaxel2,3. Next-generation taxoids feature modifications to the critical sites of paclitaxel, including the functional groups at C-10 and C-13. Notably, compounds such as SB-T-1214 have demonstrated remarkable efficacy against drug-resistant tumors4,5,6.
Structures of paclitaxel, docetaxel, cabazitaxel, and SBT-1214 are shown.
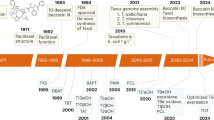
Paclitaxel is produced primarily via a semisynthetic method. Initial, 10-deacetyl-baclitatin III (10-DAB) is extracted from the needles of Taxus baccata L., after which its side chain is assembled through a 4-step chemical process7,8. Although this method has significantly improved the yield of paclitaxel compared with direct extraction from the bark of Taxus brevifolia Nutt.9,10,11, the clinical demand remains unfulfilled due to the long growth cycle and limited bioresources. The chemical synthesis of paclitaxel and its analogs has witnessed rapid advancements, yet challenges such as lengthy synthetic steps and low overall yields persist12,13,14,15,16,17.
In recent years, substantial progress has been made in elucidating and optimizing the natural biosynthetic pathway of paclitaxel, and all enzymes involved have been identified18,19,20,21,22,23,24,25,26,27,28. In 2010, the yield of taxadiene, the crucial paclitaxel precursor, produced by designed Escherichia coli cells was increased to 1 g/L18. The downstream biosynthetic of paclitaxel is severely limited by the low activity and low product selectivity of the cytochrome P450 monooxygenases (P450s) involved in native taxadiene oxidative derivation steps29,30,31. Despite the progress in elucidating the mechanism of taxadiene-5α-hydroxylase (T5αH), as the native enzyme responsible for the initial step in taxadiene oxidation, efforts to substantially enhance its catalytic efficiency and specificity via protein engineering have thus far proven unsuccessful32,33,34. To overcome the inherent limitations of T5αH, Xiao et al. investigated alternative taxadiene-oxidizing enzymes, finding that although CYP701A8 mediates C11-position oxidation, its application is severely constrained by pronounced allosteric effects35. Notably, Zhang et al. established the complete gene set requisite for heterologous paclitaxel biosynthesis and identified a taxadiene-epoxidizing enzyme that synergizes with T5αH to facilitate oxetane ring formation19. McClune et al. discovered FoTO1, an NTF2-like protein, can resolved the low product selectivity of T5αH, suppressing byproduct formation (e.g., OCT) and enabling the selective production of taxadiene-5α-ol (T5OH). This approach resulted in the isolation of baccatin III at ~8.7 μg/g DW from N. benthamiana36.
In the present study, we focus on identifying non-native taxadiene oxidases as alternatives to the native P450 enzymes. TteUPO (unspecific peroxygenase from Thielavia terrestris)37,38 is selected from 29 fungal-originated oxidases because of its highest activity toward taxadiene and excellent protein expression level in Pichia pastoris. We construct an enzyme toolbox comprising various TteUPO mutants, which exhibit distinct regioselectivities toward the taxadiene skeleton. Moreover, we engineer an E. coli BL21 (DE3) strain for the de novo biosynthesis of oxidized taxadiene products by introducing the taxadiene biosynthetic pathway and enabling the heterologous expression of a TteUPO mutant. These results lay a solid foundation for future development of artificial synthetic routes to paclitaxel and its analogs.
Results
Mining and characterization of taxane oxidases
Peroxidases and peroxygenases, both featuring heme coordinated by a cysteine ligand similar to that of P450s39,40,41,42, have been successfully utilized for the oxy-functionalization of carbon, sulfur, silicon, and nitrogen43,44,45,46,47. However, there are no reports of oxidases other than P450s that are capable of oxidizing taxanes. We initially screened 29 reported peroxidases and peroxygenases of fungal origin37,38,48,49,50,51,52,53,54, which were expressed in Pichia pastoris (syn. Komagataella phaffii). Among these, 13 oxidases exhibited activity toward taxadiene, producing identical oxidation products. TteUPO (unspecific peroxygenase from Thielavia terrestris)37,38 was chosen due to its highest activity and excellent protein expression level (Supplementary Note 1, Supplementary Tables 1, 2 and Supplementary Fig. 1, 2). TteUPO catalyzes the formation of hydroxylation products through the C-H activation of C-4 and C-6 positions in the taxadiene skeleton (Fig. 2A). Compounds 3 and 4 were identified as stereoisomers of the C-4 hydroxylation product ((S)-taxa-5(6),11(12)-dien-4-ol (3) and (R)-taxa-5(6),11(12)-diene-4-ol (4)), whereas 5 and 6 were found to be stereoisomers of the C-6 hydroxylation product ((R)-taxa-4(5),11(12)-diene-6-ol (5) and (S)-taxa-4(5),11(12)-dien-6-ol (6)). Additionally, during analysis of the given products by gas chromatography (GC), elimination of the allylic products resulted in a diene compound, namely, taxa-4(20),5(6),11(12)-triene (2) (Fig. 2B and Supplementary Fig. 3, 4, 5). This phenomenon was also observed when compounds 3 ~ 6 were dissolved in CDCl3 and CD2Cl2 for NMR experiments, indicating a facile elimination process that is enhanced by heat and acidic conditions.
A GC analysis of the reaction catalyzed by TteUPO. Compared to controls, five peaks (2-6) appeared in the TteUPO reaction; B The enzymatic catalytic reaction formula and the chemical structures of all products generated via TteUPO-catalyzed oxidation of taxadiene using ¹⁸O-labeled H₂O₂. The structure of 4 was confirmed by X-ray crystallographic analysis (deposition number: 2374527) and drawn via ORTEP (Supplementary Fig. 5); C Comparative oxidation of taxadiene by T5αH, DMDO, and TteUPO. Reaction pathway for T5αH proceeded as reported71. TteUPO neither oxidized taxadiene to form compound 7 nor catalyzed the conversion of compound 7 (synthesized via DMDO) to downstream products.
In contrast to the oxidation pattern of T5αH17,18,19,20, TteUPO catalyzed the oxidation of taxadiene to generate various hydroxylation substances that were distinct from byproducts such as OCT, iso-OCT or other compounds that constituted a large proportion of the products generated by T5αH in previous investigations18,29,30 (Fig. 2C). The oxidation of taxadiene by TteUPO was localized to either the C4 = C5 double bond or the allyl position of ring C. To elucidate the possible reaction pathway, taxadiene-4(5)-epoxide (7) was prepared by oxidation of taxadiene with DMDO55,56 (Fig. 2C and Supplementary Fig. 6). This unstable epoxy intermediate undergoes spontaneous rearrangement to form the products T5OH, OCT, and iso-OCT20,22,33,34. In TteUPO-catalyzed oxidation of taxadiene, taxadiene-4(5)-epoxide was not detected. Furthermore, epoxide 7 was not converted to compounds 3 ~ 6 by TteUPO. These observations led us to conclude that TteUPO-catalyzed taxadiene oxidation circumvents the taxadiene-4(5)-epoxide intermediate, thereby preventing the accumulation of byproducts (e.g., OCT). This finding also offers an alternative strategy to FoTO1 for addressing the McClune et al. discovered FoTO1, an NTF2-like protein, can resolved the low product selectivity of T5αH.
Protein engineering of TteUPO
Next, the catalytic efficiency was enhanced by protein engineering. Interestingly, the regioselectivity of TteUPO was also modified to enable oxidation at distinct sites of taxadiene. The crystal structure of TteUPO (PDB 9IZ8) was analyzed at a high resolution of 1.54 Å (Supplementary Fig. 7 and Supplementary Table 3). The conformation of substrate channel has been found to substantially influence the activity and selectivity of UPOs57,58,59,60,61. Molecular docking was performed via AutoDock Vina62, the fining docking structure indicates that the substrate channel of TteUPO is narrowed due to excessive steric hindrance of amino acids within the channel, thereby restricting the entry and binding of taxadiene (Supplementary Fig. 8). Furthermore, given the minimal polarity and strong hydrophobicity of taxadiene, which prevents hydrogen bond formation with surrounding amino acids, amino acids lining the substrate channel were mutated to alanine (Ala)—a sterically less hindered nonpolar amino acid. These mutations widen the channel and identify sensitive residues for further precise regulation of the channel geometry (Supplementary Tables 4 and 5).
The mutation of Leu208, which is located at the entrance of the substrate channel, to Ala, which results in less steric hindrance, widened the “bottle mouth”. This modification facilitated the entry of taxadiene into the active center (Fig. 3C, D). The L208A mutant exhibited a significant improvement in catalytic activity, with no alteration in product diversity (Fig. 3A, Table 1 and Supplementary Fig. 9A, 9C). The total turnover number (TON) increased from 82 to 4355, a marked 53-fold increase.
A GC analysis of taxadiene oxidation catalyzed by mutants of TteUPO. * Under GC conditions, compounds 11 and 12 partially degrade, yielding products with retention times identical to taxadiene (1); B Structures and MS (EI) spectra of the identified products (9, 11-13, 15, and 16). Compound 8 is the product generated by the thermal transformation of 15 during GC detection (Supplementary Fig. 15), upon scale-up, compounds 10 and 14 exhibited significantly diminished yields (Supplementary Fig. 9D); C Cross-sectional view of the TteUPO (WT) structure, highlighting Leu208 (red); D Cross-sectional view of the L208A mutant structure, highlighting Ala208 (red); E Cross-sectional view of the TteUPO (WT) structure, highlighting Leu60 (red); F Cross-sectional view of the L60A mutant structure, highlighting Ala60 (red); G Structure of the L60A/F156L mutant, highlighting Leu156 (red); H Structure of the L60A/G159L mutant, highlighting Leu159 (red). Color representation of taxadiene: blue for ring A, green for ring B and yellow for ring C.
Notably, the L60A mutant exhibited enhanced activity and demonstrated low product selectivity compared with the WT (Fig. 3A, B, Table 1 and Supplementary Fig. 9B, 9C). Leu60, which is positioned above the heme porphyrin ring, was substituted with alanine, thereby enhancing substrate accessibility to the catalytic site (Fig. 3E, F) and thus increasing its oxidative activity (Table 1). Moreover, expansion of the inner side of the substrate channel increases the diversity of the substrate binding conformations, resulting in the formation of more than 6 oxidation products (Fig. 3A, B). In contrast to the WT, the L60A mutant tended to attack the C11 = C12 double bond in ring A. In addition to the generation of hydroxylation products at the allylic positions (i.e., C-10 and C-13), epoxidation of the double bond in ring A was also observed, along with further oxidation of C-6 in ring C (Fig. 3A, B). Notably, the oxidation sites and configurations at C-10 and C-13 were identical to those in paclitaxel.
The L60S mutant exhibited activity and selectivity profiles comparable to those of L60A (Supplementary Fig. 9B). Although serine possesses a polar hydroxyl group not present in alanine, both residues impose similar steric constraints. This result further confirms that steric hindrance, rather than side-chain polarity, is the primary determinant of TteUPO’s activity and selectivity in taxadiene oxidation.
Interestingly, the L60A mutant exhibits remarkable versatility, catalyzing oxidations at the C-4, C-6, C-10, C-11, C-12, and C-13 positions of taxadiene. This renders it an excellent starting point for constructing a mutant library tailored to the regioselective oxidation of taxadiene. Mutants generated through FLA scanning (replaced with Phe, Leu, or Ala) and STY scanning (replaced with Ser, Thr, or Tyr)63, along with site-directed saturation mutagenesis of amino acids within the substrate channel, were then specifically designed to enhance the regioselective oxidation capabilities of the mutant (Supplementary Fig. 10).
The combined mutant L60A/F156A enables catalysis of epoxidation at C-11 and C-12 and oxidation at C-6 on taxadiene, resulting in the formation of compound 16 (Table 1). Compared with the L60A/F156A mutant, the L60A/F156L mutant exhibits greater steric hindrance at position 156 due to the bulkier leucine residue replacing alanine. This increased steric clash further constrains the binding conformation of taxadiene and limits the positioning of ring C near heme-Fe. Additionally, the “concave surface” of taxadiene can only enter the active site with the concave side facing Leu156, positioning the double bond at ring A close to the heme-Fe (Fig. 3G and Supplementary Fig. 11C). As a result, the C-10 and C-13 hydroxylation products (compounds 11 and 12), along with the C-11 and C-12 epoxidation product (compound 9), dominate for over 90% of the products, while oxidation at C-6 occurs only rarely (Table 1).
Notably, the L60A/G159L mutant exhibited a significant increase in selectivity, specifically increasing the proportion of the C-10 (compounds 11) and C-13 (compounds 12) hydroxylation products to 66% and 19%, respectively, with a conversion of 80.2% (Table 1). In the presence of Leu159, which exhibits greater steric hindrance and thus constrains other substrate binding conformations, the substrate preferentially enters the channel with a parallel conformation relative to the heme plane. Consequently, ring B of taxadiene comes closest to heme-Fe, thereby favoring the hydroxylation product at C-10 of ring B. However, as the distance increases, the oxidation activity slightly deteriorates (Fig. 3H, Supplementary Fig. 11A, 11B, and Supplementary Table 6). To restore activity, the L60A/G159L mutant was combined with the activity-enhancing L208A mutation to generate the triple mutant L60A/G159L/L208A. This modification led to a significant performance improvement, achieving a turnover number (TON) of 757. Additionally, the selectivity for the hydroxylation product at C-13 was increased to 33% (Table 1). The L60A/G159L/L208A mutant also exhibited excellent scalability. In a 100 mL reaction system, the yield of 0.5 mM taxadiene catalyzed by 4 mg of enzyme reached 75.3% within 2 h.
Development of a taxane oxidase toolbox
Seven TteUPO mutants were employed to construct an engineered oxidase toolbox, enabling site-specific oxidation of taxanes (Fig. 4A). Eleven distinct oxidized products were structurally characterized, revealing six oxidizable sites on the taxane scaffold (Fig. 4B and Supplementary Fig. 12). Compounds 11 and 12 exhibit regioselectivity and stereoselectivity consistent with those of paclitaxel, making them potential building blocks for reconstructing the paclitaxel synthesis pathway. Notably, the L60A/F156A and L60A mutants exhibit poor regioselectivity in catalyzing the oxidation of taxadiene. However, when additional steric hindrance is introduced at C-10 on ring B or C-4 on ring C of taxadiene (compound 11 and 4), these mutants exhibit high regioselectivity in oxidizing taxanes, yielding compounds 17 and 18. This highlights the versatility of the toolbox, which not only has been proven effective for taxadiene but also demonstrates even greater performance for taxadiene derivatives. The oxidation sites of compounds 5, 9, 13, 15, 16, 17, and 18 differ from those of paclitaxel, and these distinct chemical modifications may result in diverse pharmacological activities, while oxidation at the C-6 position is associated with paclitaxel metabolism in the human body64.
A Regioselectivity of mutants in the toolbox toward different taxanes. Definition of Regioselectivity: The peak area ratio between oxidation products at different positions on the taxane skeleton. Compound 15 or 16 is equally allocated between the C-6 position and combined C-11/C-12 positions. Unidentified products are excluded from the analysis; B Oxidation of Taxanes using TteUPO mutants in the constructed toolbox. The proportions of the depicted products relative to the total products were determined in a single-step enzymatic reaction. Y-axis Peak area ratio of illustrated compounds in the reaction mixture, with inclusion of unidentified products. Source data are provided as a Source Data file.
The biosynthesis of paclitaxel involves eight oxidative modifications of the taxadiene core structure (Fig. 1). The L208A mutant, which exhibits the highest activity, was employed to probe the oxidation capabilities of this engineered enzyme toolbox toward higher-oxidation-state taxane derivatives (compounds 19/21/23) (Fig. 4 and Supplementary Fig. 12). The introduction of a C2-carbonyl group in compound 19 (vs. taxadiene) dramatically enhances both regioselectivity and stereoselectivity in L208A catalysis, with compound 20 accounting for over 99% of the total products. L208A selectively oxidizes the configurationally inverted isomer 21 to compound 22, but shows no activity toward its stereoisomer 23. We ultimately achieved the synthesis of tri-oxygenated taxanes, demonstrating the potential of our established enzyme toolbox to generate high oxidation state taxane derivatives.
In vivo biosynthesis of oxidized taxadiene
Finally, we attempted to construct a biosynthetic pathway for in vivo production of oxidized taxadiene derivatives. Building upon a high-yielding E. coli taxadiene production chassis established in our previous work65, we successfully achieved functional expression of the L208A mutant in E. coli through N-terminal signal peptide truncation (Supplementary Fig. 14). Subsequent integration of this artificial oxidase module into the taxadiene-producing chassis, coupled with exogenous H2O2 supplementation, achieved de novo biosynthesis of taxane oxidation products at a total titer of 14.2 mg/L (Fig. 5).
A Schematic of the biosynthesis pathway of taxadiene oxidation products in engineered E. coli. GGPPS: geranylgeranyl pyrophosphate synthase; TDS: taxadiene synthase. H2O2 was exogenously supplemented; B GC analysis of products from the engineered E. coli chassis. Heterologous expression of L208A and exogenous supplementation of H2O2 enabled the detection of products 3, 4, 5, and 21. C Yields of taxane oxidation products in the engineered E. coli chassis. The bars are mean of n = 3 independent experiments and error bars indicate s.d. Source data are provided as a Source Data file.
Discussion
By screening of 29 oxidases, the ability of TteUPO to oxidize the skeleton of taxadiene was revealed. Based on the crystal structure of TteUPO, protein engineering was employed to remodel its substrate channel, enabling precise regulation of both activity and selectivity. Notably, the L208A mutant exhibited a 53-fold improvement in turnover number (TON), resolving the low catalytic activity intrinsic to the wild-type. Leveraging the catalytically promiscuous L60A variant through protein engineering enabled site-directed oxidation of taxadiene. An oxidase toolbox comprising TteUPO mutants was engineered to perform regio- and stereoselective oxidations at C-4, C-6, C-10, C-11, C-12, and C-13 positions of the skeleton of taxanes. This platform establishing a powerful strategy for diversifying oxidation patterns in structurally significant compounds. Finally, we validated the integration potential of this toolbox within the paclitaxel biosynthetic pathway. By truncating the N-terminal signal peptide, heterologous expression of the L208A mutant in E. coli was achieved, where implementation of a controlled H2O2 feeding protocol enabled de novo biosynthesis of oxidized taxadiene derivatives.
In summary, our research establishes steric hindrance within the substrate channel as the principal factor governing both activity and selectivity in TteUPO-catalyzed taxadiene oxidation. While numerous oxidation products remained unidentified due to low production titers, strategic remodeling of the channel of enzyme may enable engineered oxidation at additional taxane sites. Our research demonstrates significant potential as a foundation for the reconstruction of synthetic pathways for paclitaxel and its analogs.
Methods
Strains, genes and vectors
The gene sequences of oxidases were codon-optimized for expression in Pichia pastoris (syn. Komagataella phaffii) and cloned into the pPICZA vector by GenScript (Shanghai, China). Plasmid cloning was carried out using Escherichia coli TOP10 (Novagen, Darmstadt, Germany). Pichia pastoris X-33 was employed for the expression of oxidases.
A 6 × His tag was introduced after the signal peptide of TteUPO, with the specific primers detailed in Supplementary Table 1. All primers were synthesized by Tsingke Biotechnology Co., Ltd. (Shanghai, China).
All PCR reactions, including saturation mutation experiments, were conducted using PrimeSTAR® HS DNA polymerase (Takara, Shiga, Japan). The insertion of the 6 × His tag was achieved utilizing homologous recombination assay kits (ClonExpress Ultra One Step Cloning Kit, Vazyme, China).
Shake flask cultivation of the recombinant P. pastoris X-33
Shake flask cultivation of the recombinant Pichia pastoris was used for the activity screening.
The recombinant P. pastoris X-33 strains were cultured for 24 hours in YPD medium containing 100 μg/L zeocin (30 °C, 220 rpm). Subsequently, the strain was inoculated with 1% (vol/vol) of the inoculum into 50 mL MD medium (20 g/L peptone; 10 g/L yeast extract; 10 g/L glycerol) and incubated at 30 °C for 16–18 h. After allowing it to stand at room temperature for 4 hours to separate the supernatant and precipitate, 50 mL of induction medium (20 g/L peptone; 10 g/L yeast extract; 13.4 g/L YNB; 100 mM potassium phosphate buffer pH 6.0; 400 µg/L biotin; 200 µM ALA (5-aminolevulinic acid), 3 mM MgSO4) was added to the precipitation. The culture was then incubated with 1% methanol every 24 h for total 72 h.
To confirm that oxidases has been expressed, the activity of the expressed protein was assessed using the substrate ABTS (2,2′-Azino-bis (3-ethylbenzothiazoline-6-sulfonic acid) diammonium salt). The assay reaction mixture, containing 1 mM ABTS, 2 mM H2O2, and citrate buffer (pH 4.5), was incubated at 30 °C for 1 minute, and the absorbance change at 418 nm was measured (ε418 = 36,000 M−1 cm−1)48.
Screening of oxidases
29 reported oxidases, with protein sequences and signal peptides consistent with the published literatures, were screened. The expression vector was changed to pPICZA with the AOX1 promoter, and the enzymes were expressed in P. pastoris X-33. ABTS was utilized to confirm the correct expression of these oxidase. Subsequently, the expressed oxidases were employed to catalyze the oxidization of taxadiene, with 13 of them demonstrating activity against taxadiene and yielding the same oxidized products (Supplementary Fig. 1 and Supplementary Table 2). TteUPO was chosen due to its highest activity and excellent protein expression level.
The taxadiene oxidation reactions catalyzed by oxidases were conducted at 30 °C in a citrate buffer solution (CPBS) with a pH of 5.5. The reaction mixture contained 0.5 mM taxadiene, 2 mM H2O2, 30% (vol/vol) acetonitrile, and 50 μL/500 μL concentrated crude enzyme solution was incubated for 1 hour at 200 rpm. Crude enzyme solution was concentrated 50-fold from the fermentation supernatant.
Batch feeding cultivation of recombinant P. pastoris X-33
Batch feeding cultivation of TteUPO and its mutants are used for products preparing.
Initial Fermentation Medium: Dissolve 1.14 g/L of CaSO4·2H2O, 14.3 g/L of K2SO4, 12.2 g/L of MgSO4·7H2O, 3.3 g/L of KOH, 21 mL/L of phosphoric acid, and 40 g/L of glycerol in deionized water and subject the medium to autoclaving at 121 °C for 20 min. Prior toinoculation, adjust the fermentation pH to 6.0 using ammonia solution and supplement with 12 mL/L of PTM1.
Glycerol Feeding Medium: Dissolve 600 g/L of glycerol in deionized water to a total volume of 400 mL. And subject the medium to autoclaving at 121 °C for 20 min. After cooling, add 4.4 mL/L of PTM1 in a sterile environment.
Methanol Feeding Medium: Add 12 mL/L of PTM1 to pure methanol.
PTM1: Dissolve the following in deionized water in the specified order: 0.2 g/L biotin, 6 g/L CuSO4·5H2O, 0.09 g/L KI, 3 g/L MnSO4·H2O, 0.2 g/L Na2MoO4·2H2O, 0.02 g/L H3BO3, 0.9 g/L CoCl2·6H2O, 42.2 g/L ZnSO4·7H2O, 5 mL/L sulfuric acid, and 65 g/L FeSO4·7H2O. Filter the medium using 0.22 μm filtration to sterilize.
Batch feeding cultivation of TteUPO and its mutants were performed in the following five stage.
Stage 1: Activation of preserved recombinant P. pastoris X-33 strains by streaking on YPD plates. Pick single colonies and inoculate into 4 mL of YPD culture medium for 24 h. Inoculate 1% into 250 mL of MD medium and incubate overnight at 30 °C until OD600 = 2–6.
Stage 2: In a 5 L tank containing 2.5 L of liquid, use a 10% inoculation rate and incorporate 1 mM final concentration of Ampicillin, and culture at 28 °C. Set up a dissolved oxygen (DO)-stirring linkage, increasing stirring speed when DO < 30% in the range of 200-1200 rpm. Approximately 17–18 h later, there will be a notable increase in DO. Following manually supplementing glycerol, DO will decrease, marking the start of automatic feeding. The DO-STAT system is configured to feeding when DO exceeds 30%, with observed fluctuations maintained around 30% DO. The aeration rate is controlled at 1-2 vvm.
Stage 3: When the OD600 reaches a value between 150 and 200, stop feeding. At this point, DO will rapidly increase to a plateau at around 100% and remain steady for about 1 h, ensuring complete consumption of glycerol to prevent any interference with the subsequent methanol induction.
Stage 4: Add 200 μL/L of 1 M ALA, and infuse methanol at a rate not exceeding 7 mL/L/h for a duration of 2 hours to acclimatize the cells to methanol metabolism. Following this acclimation period, configure the DO-STAT to automatically supplement methanol when DO exceeds 30%. The induction culture is maintained for a total of 72 h.
Stage 5: Centrifuge the fermentation broth at 12,000 × g for 30 min and collect the supernatant from the fermentation.
Purification of TteUPO
Protein purification was carried out using Ni-NTA. Initially, impurities were eluted using buffer A (20 mM sodium dihydrogen phosphate, 500 mM sodium chloride, pH 7.4) with twice the column volume, followed by elution of TteUPO using buffer B (20 mM sodium dihydrogen phosphate, 500 mM sodium chloride, 50 mM imidazole, pH 8.0) with twice the column volume.
The protein content was determined using the Coomassie brilliant blue method.
Deglycosylation of TteUPO was performed with SpEndo H66 (mg TteUPO: mg SpEndo H = 5:1). The deglycosylated product was separated by Ni-NTA. Finally, the results of protein purification and deglycosylation were analyzed using SDS-PAGE (Supplementary Fig. 2).
Crystallization and structure determination
The crystallization was carried out at 18 °C in 96-well plates using sitting-drop vapor-diffusion with the commercial crystal condition (1.6% v/v Morpheus Ⅲ® Dipeptides Mix, 0.1 M Morpheus® -Buffer System 1 pH 6.5, 37.5% Precipitant Mix 4). The droplets consisted of 1 μL of 18 mg/mL purified TteUPO in buffer (10 mM potassium phosphate, pH 7.4) and 1 μL of the precipitant solution and were equilibrated against 50 μL of the precipitant solution. The crystals were flash-cooled in liquid nitrogen for data collection. Diffraction data were collected on beamlines BL10U2 and at Shanghai Synchrotron Radiation Facility. Data were processed with XDS and HKL2000 (Supplementary Fig. 7). The structure was solved by molecular replacement with the structure of TteUPO predicted by AF267. Iterative model building and refinement were performed using WINCOOT 0.9.8.168 and PHENIX 1.21.1.526869, respectively (Supplementary Table 3).
GC-MS analysis
The analyses were performed with GCMS-QP2010 SE (Shimadzu, Kyoto, Japan), using an HP-5MS fused silica capillary column (30 m, 0.25 mm i. d., 0.50 μm film; Agilent). The oven temperature was from 80 °C to 220 °C (15 °C min−1) and held for 3 min, then by 20 min−1 to 280 °C, and held for 5 min. The scan range was set to m/z 35-400 and the electron ionization energy was 70 ev.
The reaction mixture was extracted with an equal volume of ethyl acetate, which contained 50 mg/L of octadecane as the internal standard for GC-MS analysis. Standard concentration curve of taxadiene is showed in Supplementary Fig. 3.
The quantitative analysis was performed using GC peak area integration, with the sample size set to n = 3. MS was employed for compound identification.
Preparation of taxadiene
The preparation of taxadiene (1) follows the established method from our laboratory65. After 72 h of induction culture of recombinant Escherichia coli BL21 (DE3), the cells were harvested via centrifugation, separating the supernatant from the bacterial sediment. The supernatant was subjected to extraction with an equal volume of ethyl acetate, while the bacterial pellet was soaked in an appropriate volume of methanol overnight before being extracted with an equivalent volume of n-hexane. The resulting extract was then dried using an appropriate amount of anhydrous sodium sulfate, and the solvent was removed through rotary evaporation at −0.1 Mpa and 30 °C. Preliminary purification of taxadiene was performed via silica gel column chromatography, followed by preparative HPLC to isolate taxadiene and its isomers. For the silica gel column chromatography, taxadiene was eluted by n-hexane. Detection was performed by TLC, developed with n-hexane, and observed by staining with phosphomolybdic acid. The preparative HPLC was conducted using an UltiMate 3000 chrome system (Thermo Fisher Scientific, USA) equipped with an Ultimate ® XB-C18 column (Welch, China). The conditions were set as follows: 100% acetonitrile at a flow rate of 3 mL/min, with detection at 210 nm UV.
Preparation of taxanes
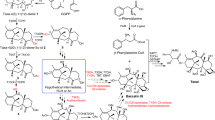
Compound 19, 21, and 23 were synthesized according to the protocol developed by the Baran’s group15,70.
The synthesis begins with the dibromocyclopropanation of a starting alkene using bromoform (CHBr₃) and potassium tert-butoxide (t-BuOK) as base. This resulting cyclopropane intermediate undergoes thermal rearrangement in N,N-dimethylaniline (PhNMe2) to form a brominated alkene. In a separate branch, a cyclohexanone derivative is subjected to nucleophilic addition with vinylmagnesium bromide to introduce a vinyl group. The two segments are then converged. The vinyl-functionalized cyclohexanone is deprotonated with n-butyllithium (n-BuLi) and protected with trimethylsilyl chloride (TMSCl). A key copper(I)-catalyzed conjugate addition is subsequently carried out using methylmagnesium bromide (MeMgBr) in the presence of a copper(II) triflate/N-heterocyclic carbene (NHC) catalyst system. The resulting enolate is trapped in an aldol reaction with acrolein, and the product is oxidized with Dess-Martin periodinane (DMP) to yield an advanced intermediate. This intermediate is then treated with boron trifluoride diethyl etherate (BF₃·OEt₂), which triggers Diels–Alder cyclization reaction to form the core ring structure, yielding compound 23 and 21. The final step is a palladium-catalyzed Negishi coupling between compound 23 and dimethylzinc (Me₂Zn), using tetrakis(triphenylphosphine)palladium(0) (Pd(PPh₃)₄) as the catalyst. This coupling installs the final methyl group, affording the desired product (compound 19) (Supplementary Fig. 13).
Biotransformations
Unless otherwise specified, the biotransformations of taxanes were conducted at 30 °C in a citrate buffer solution (CPBS) with a pH of 5.5. The reaction mixture contained 0.5 mM substrates, 2 mM H2O2, 30% (vol/vol) acetonitrile, and 40 μg/mL enzyme was incubated for one hour at 200 rpm. The addition of 5 mM 2-hydroxypropyl-β-cyclodextrin was found to be beneficial for the biotransformation of taxadiene.
The analytical-grade reaction is conducted in a 2 mL glass injection bottle for a 0.5 mL reaction, whereas the preparative-grade reaction takes place in a 2 L glass shake flask for a 500 mL reaction.
Isotope labeling
Utilizing H218O2 (Sigma-Aldrich®, Germany) as oxygen donor to oxidize taxadiene, under the conditions described in “Biotransformations” section, the molecular weight of compound 5 shifted from 288 to 290 (Supplementary Fig. 4). This suggests that H2O2 serves as the oxygen donor for unspecific peroxygenase during the oxidation of taxanes.
Product preparation and isolation
The reaction system was scaled up to 500 mL and conducted in a 2 L glass shake flask. The supernatant was extracted with ethyl acetate without an internal standard, dried with an appropriate amount of anhydrous sodium sulfate, and concentrated by rotary evaporation at −0.1 Mpa and 30 °C. Subsequently, the resulting product underwent rough separation using silica gel column chromatography. The target compounds were then collected and further purified through preparative HPLC using an UltiMate 3000 chrome system (Thermo Fisher Scientific, USA) equipped with an Ultimate ® XB-C18 column (Welch, China). For the silica gel column chromatography, compounds 2-16 were eluted by n-hexane: ethyl acetate = 30:1. Detection was performed by TLC, developed with a mixture of n-hexane: ethyl acetate in = 6:1, and observed by staining with phosphomolybdic acid. Compounds 17-22 were eluted by n-hexane: ethyl acetate = 10:1, and its detection was carried out by TLC with a development agent of n-hexane: ethyl acetate = 1:1, followed by staining with phosphomolybdic acid. The preparative HPLC conditions are as follows: 100% acetonitrile at a flow rate of 3 mL/min, with detection at 210 nm UV.
Analysis of catalytic mechanism
We synthesized taxadiene-4 (5)-epoxide (7) using DMDO55,56, and analyzed it by GC-MS and 1H NMR as shown in Supplementary Fig. 6.
Synthesis procedure of taxadiene-4(5)-epoxide (7): NaHCO3 (12.00 g, 142.85 mmol) was dissolved in a mixture of H2O (10 mL) and acetone (15 mL). The reaction vessel was placed in an ice-water bath with stirring at 200 rpm. Oxone® (12.5 g total) was added batchwise within 1 min. After stirring for 30 min, the mixture was extracted with diethyl ether (20 mL). The resulting ethereal extract containing DMDO was stored in an ice-water bath. Taxadiene was dissolved in the DMDO/ether solution to a final concentration of 2 mM and reacted with stirring (200 rpm) in an ice-water bath for 1 h. The solvents were removed by rotary evaporation under reduced pressure. Taxadiene-4(5)-epoxide (7) was purified using silica gel column chromatography, eluted by n-hexane/ethyl acetate (30:1, v/v). Reaction progress and product fractions were monitored by TLC (silica gel, n-hexane/ethyl acetate = 6:1, v/v), with visualization achieved by staining with phosphomolybdic acid. The target compound was then collected and further purified through preparative HPLC using an UltiMate 3000 chrome system (Thermo Fisher Scientific, USA) equipped with an Ultimate® XB-C18 column (Welch, China).
Selection of amino acids within the substrate channel
L56, L60, F63, F156, G159, and L208 were selected for the first round of alanine scanning (Supplementary Fig. 8A). It is worth noting that E160 is a conserved site thought to be involved in the oxidation reaction with H2O2, and saturation mutagenesis at this position has been significantly diminished the activity.
Building upon the L60A mutant, the following residues were selected for FLA and STY scanning: L56, L59, F63, A155, F156, G159, L208, A163, and the residues M20 and L21 located behind the heme (Supplementary Fig. 8B).
Construction of mutagenesis library
Site-directed mutagenesis was conducted by performing whole-plasmid PCR amplifications of the parental plasmids using PrimeSTAR HS (premix) from Takara along with the respective mutagenesis primers (Supplementary Table 4) carrying the designed mutations. The amplified product was then subjected to digestion with Dpn I (NEB) at 37 °C for 2 h to selectively degrade the parental methylated strands. Subsequently, the digested product was transformed into Escherichia coli TOP10 cells. For site-directed mutagenesis, an Escherichia coli TOP10 colony of the designed mutant was selected from an agar plate and inoculated into 4 mL of LLB medium (containing 5 g/L NaCl, 5 g/L yeast powder, and 10 g/L peptone) supplemented with 100 μg/mL zeocin in a glass tube, and cultivated at 37 °C and 200 rpm. After 12 h of cultivation, the recombinant plasmid was extracted. Linearization of the recombinant plasmid was performed using Sac Ⅰ for 3 h. Linearized fragments were then electro-transformed into P. pastoris X33 with a micropulser (BioRad, Hercules, CA, USA). The mutants were cultivated in 250 mL shake flasks containing 50 mL of media (as described in “Shake flask cultivation of the recombinant P. pastoris X-33” section).
Site-directed saturation mutagenesis involves mutating a specific site to each of the other 19 amino acids.
Screening of mutants
The mutants were cultivated in 250 mL shake flasks containing 50 mL of media and the secreted supernatant was subsequently purified using Ni-NTA (as described in “Purification of TteUPO” section). The purified enzymes were evaluated for their activity and selectivity through reactions with taxadiene, with the results analyzed via gas chromatography (GC) (as described in “GC-MS analysis” section) (Supplementary Tables 5, 6 and Supplementary Fig. 9-11).
MS, HR-MS and NMR spectroscopy
MS data were measured as described in “GC-MS analysis” section (Supplementary Fig. 15). HR-MS data were measured on a GCT Premier mass spectrometer (Waters Corporation, USA) (Supplementary Fig. 16). HR-MS analysis was performed using dried pure compounds, with a sample size of n = 1. NMR spectra were recorded on a Bruker (Billerica, MA, USA) AV Neo 400 MHz, AV Neo 500 MHz, or AV Neo 600 MHz at ambient temperature. Spectra were referenced against solvent signals (1H-NMR, residual proton signal of C6D6: δ = 7.16 ppm, CD2Cl2: δ = 5.32 ppm; 13C-NMR, signal of C6D6: δ = 128.06 ppm, CD2Cl2: δ = 53.84 ppm), with a sample size of n = 1. (Supplementary Tables 7–21 and Supplementary Figs. 17–121).
Construction and cultivation of recombinant E. coli
L208A codon optimization for E. coli was performed at GenScript (Shanghai, China), and the gene was synthesized into the plasmid pET 28a (+). After removal of the N-terminal signal peptide, L208A was successfully expressed in E. coli and exhibited activity against ABTS and taxadiene (Supplementary Fig. 14A–14F). By optimizing the concentration of the inducer IPTG to 0.1 mM during a 25-h induction, the expression level of L208A was enhanced by over two-fold (Supplementary Fig. 14G). H₂O₂ supplementation was initiated after the first 8 hours of induction and repeated every 8 h (total of 3 additions) When H₂O₂ concentration per addition was kept below 5 mM, bacterial growth was largely unaffected (Supplementary Fig. 14H). Based on these results, we transformed the recombinant plasmid carrying L208A into a taxadiene-producing E. coli chassis strain constructed previously in our laboratory (Supplementary Fig. 14I)65. The engineered E. coli chassis for taxadiene production was designed to overexpress two key upstream enzymes, 1-deoxy-D-xylulose-5-phosphate synthase (DXS) and isopentenyl diphosphate isomerase (IDI), to enhance the supply of the precursors, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). This elevated precursor pool then enabled the de novo biosynthesis of taxadiene through heterologous expression of geranylgeranyl pyrophosphate synthase (GGPPS) and taxadiene synthase (TS).
The recombinant E. coli strain was cultured under the following conditions: A 2% inoculum was transferred into 50 mL of Terrific Broth (TB) medium (12 g/L peptone, 24 g/L yeast extract, 5.2 g/L glycerol, 2.31 g/L KH₂PO₄, 12.54 g/L K₂HPO₄). Initial cultivation was carried out at 37 °C and 200 rpm for 3 h, followed by temperature reduction to 20 °C for 30 min. Subsequently, 0.1 mM IPTG was added, and cultivation continued for another 16 h. Then, 2.5 mM H₂O₂ was added, and cultivation proceeded for an additional 8 h. Finally, the fermentation broth was extracted with an equal volume of ethyl acetate for GC analysis.
Statistics and reproducibility
Unless otherwise specified, all statistical experiments were determined with n = 3. No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The Investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Bioinformatics data; Mutant screening data; Protein crystallization data; GC, MS (EI) and HR-MS chromatograms; and NMR spectra are included in the Supporting Information. Crystallographic data for the structures reported in this Article have been deposited at the Cambridge Crystallographic Data Center, which are available at the CCDC accession 2374527 code (https://www.ccdc.cam.ac.uk/structures/Search?Ccdcid=2374527). Diffraction data for protein reported in this Article have been deposited at Protein Data Bank, which are available at the PDB accession 9IZ8 code (https://doi.org/10.2210/pdb9iz8/pdb). Source data are provided with this paper.
References
Holmes, F. A. et al. Phase II trial of taxol, an active drug in the treatment of metastatic breast cancer. J. Natl. Cancer Inst. 69, 341–345 (1994).
Bissery, M.-C., Nohynek, G., Sanderink, G.-J. & Lavelie, F. Docetaxel (Taxotere®) a review of preclinical and clinical experience. Part I: preclinical experience. Anti-Cancer Drugs 6, 339–355 (1995).
De Bono, J. S. et al. Prednisone plus cabazitaxel or mitoxantrone for metastatic castration-resistant prostate cancer progressing after docetaxel treatment: a randomised open-label trial. Lancet 376, 1147–1154 (2010).
Jayanetti, K., Takemura, K., Bendale, H., Garg, A. & Ojima, I. Recent advances in the strategic incorporation of fluorine into new-generation taxoid anticancer agents. J. Fluor. Chem. 267, 110106 (2023).
Mohelnikova-Duchonova, B. et al. Hedgehog pathway overexpression in pancreatic cancer is abrogated by new-generation taxoid SB-T-1216. Pharmacogenomics J 17, 452–460 (2017).
Seitz, J. D., Wang, T., Vineberg, J. G., Honda, T. & Ojima, I. Synthesis of a next-generation taxoid by rapid methylation amenable for 11C-labeling. J. Org. Chem. 83, 2847–2857 (2018).
Denis, J. N. et al. Highly efficient, practical approach to natural taxol. J. Am. Chem. Soc. 110, 5917–5919 (1988).
Holton, R. A. Method for preparation of Taxol. EP Patent, EP0400971A2 (1990).
Malik, S. et al. Production of the anticancer drug Taxol in Taxus baccata suspension cultures: a review. Process Biochem 46, 23–34 (2011).
Nadeem, M., Rikhari, H. C., Kumar, A., Palni, L. M. S. & Nandi, S. K. Taxol content in the bark of Himalayan yew in relation to tree age and sex. Phytochemistry 60, 627–631 (2002).
Wheeler, N. C. et al. Effects of genetic, epigenetic, and environmental factors on Taxol content in Taxus brevifolia and related species. J. Nat. Prod. 55, 432–440 (1992).
Holton, R. A. et al. First total synthesis of Taxol. 1. Functionalization of the B ring. J. Am. Chem. Soc. 116, 1597–1598 (1994).
Holton, R. A. et al. First total synthesis of taxol. 2. Completion of the C and D rings. J. Am. Chem. Soc. 116, 1599–1600 (1994).
Nicolaou, K. C. et al. Total synthesis of taxol. Nature 367, 630–634 (1994).
Kanda, Y. et al. Two-phase synthesis of taxol. J. Am. Chem. Soc. 142, 10526–10533 (2020).
Hu, Y. J., Gu, C. C., Wang, X. F., Min, L. & Li, C. C. Asymmetric total synthesis of taxol. J. Am. Chem. Soc. 143, 17862–17870 (2021).
Min, L. et al. Strategies and lessons learned from total synthesis of taxol. Chem. Rev. 123, 4934–4971 (2023).
Ajikumar, P. K. et al. Isoprenoid pathway optimization for taxol precursor overproduction in Escherichia coli. Science 330, 70–74 (2010).
Zhang, Y. et al. R. Synthetic biology identifies the minimal gene set required for paclitaxel biosynthesis in a plant chassis. Mol. Plant. 16, 1951–1961 (2023).
Zhao, Y. et al. Oxetane ring formation in taxol biosynthesis is catalyzed by a bifunctional cytochrome P450 enzyme. J. Am. Chem. Soc. 146, 801–810 (2023).
Jiang, B. et al. Characterization and heterologous reconstitution of Taxus biosynthetic enzymes leading to baccatin III. Science 383, 622–629 (2024).
Liu, J. C.-T., De La Peña, R., Tocol, C. & Sattely, E. S. Reconstitution of early paclitaxel biosynthetic network. Nat. Commun. 15, 1419 (2024).
Yang, C. et al. Biosynthesis of the highly oxygenated tetracyclic core skeleton of taxol. Nat. Comm. 15, 2339 (2024).
Croteau, R., Ketchum, R. E., Long, R. M., Kaspera, R. & Wildung, M. R. Taxol biosynthesis and molecular genetics. Phytochem. Rev. 5, 75–97 (2006).
Jennewein, S., Wildung, M. R., Chau, M., Walker, K. & Croteau, R. Random sequencing of an induced Taxus cell cDNA library for identification of clones involved in Taxol biosynthesis. Proc. Natl. Acad. Sci. USA 101, 9149–9154 (2004).
Dejong, J. M. et al. Genetic engineering of taxol biosynthetic genes in Saccharomyces cerevisiae. Biotechnol. Bioeng 93, 212–224 (2006).
Guerra-Bubb, J., Croteau, R. & Williams, R. M. The early stages of taxol biosynthesis: an interim report on the synthesis and identification of early pathway metabolites. Nat. Prod. Rep. 29, 683–696 (2012).
Kaspera, R. & Croteau, R. Cytochrome P450 oxygenases of taxol biosynthesis. Phytochem. Rev. 5, 433–444 (2006).
Biggs, B. W. et al. Overcoming heterologous protein interdependency to optimize P450-mediated taxol precursor synthesis in Escherichia coli. Proc. Natl. Acad. Sci. USA 113, 3209–3214 (2016).
Li, J. et al. Chloroplastic metabolic engineering coupled with isoprenoid pool enhancement for committed taxanes biosynthesis in Nicotiana benthamiana. Nat. Commun. 10, 4850 (2019).
Mutanda, I., Li, J., Xu, F. & Wang, Y. Recent advances in metabolic engineering, protein engineering, and transcriptome-guided insights toward synthetic production of taxol. Front. Bioeng. Biotechnol. 9, 632269 (2021).
Edgar, S., Li, F. S., Qiao, K., Weng, J. K. & Stephanopoulos, G. Engineering of taxadiene synthase for improved selectivity and yield of a key taxol biosynthetic intermediate. ACS Synth. Biol. 6, 201–205 (2017).
Yadav, V. G. Unraveling the multispecificity and catalytic promiscuity of taxadiene monooxygenase. J. Mol. Catal. B-Enzym. 110, 154–164 (2014).
Song, X. et al. Unraveling the catalytic mechanism of taxadiene-5α-hydroxylase from crystallography and computational analyses. ACS Catal 14, 3912–3925 (2024).
Xiao, Z. et al. Cytochrome P450-mediated skeleton rearrangement of taxadiene in an engineered Escherichia coli system. Org. Lett. 26, 1640–1644 (2024).
McClune, C. J. et al. Discovery of FoTO1 and taxol genes enables biosynthesis of baccatin III. Nature 643, 582–592 (2025).
Püllmann, P. et al. A modular two yeast species secretion system for the production and preparative application of unspecific peroxygenases. Commun. Biol. 4, 562 (2021).
Püllmann, P. & Weissenborn, M. J. Improving the heterologous production of fungal peroxygenases through an episomal Pichia pastoris promoter and signal peptide shuffling system. ACS Synth. Biol. 10, 1360–1372 (2021).
Grogan, G. Hemoprotein catalyzed oxygenations: P450s, UPOs, and progress toward scalable reactions. JACS Au 1, 1312–1329 (2021).
Hofrichter, M. & Ullrich, R. Oxidations catalyzed by fungal peroxygenases. Curr. Opin. Chem. Biol. 19, 116–125 (2014).
Wang, X., Peter, S., Kinne, M., Hofrichter, M. & Groves, J. T. Detection and kinetic characterization of a highly reactive heme-thiolate peroxygenase compound I. J. Am. Chem. Soc. 134, 12897–12900 (2012).
Sigmund, M.-C. & Poelarends, G. J. Current state and future perspectives of engineered and artificial peroxygenases for the oxyfunctionalization of organic molecules. Nat. Catal. 3, 690–702 (2020).
Ullrich, R., Nüske, J. R., Scheibner, K., Spantzel, J. & Hofrichter, M. Novel haloperoxidase from the agaric basidiomycete Agrocybe aegerita oxidizes aryl alcohols and aldehydes. Appl. Environ. Microbiol. 70, 4575–4581 (2004).
Peter, S., Kinne, M., Ullrich, R., Kayser, G. & Hofrichter, M. Epoxidation of linear, branched and cyclic alkenes catalyzed by unspecific peroxygenase. Enzyme Microb. Technol. 52, 370–376 (2013).
Scheibner, K., Ullrich, R., Kiebist, J., Kellner, H. & Hofrichter, M. Unspezifische Peroxygenasen — Oxyfunktionalisierung außerhalb der Pilzhyphe. Biospektrum 26, 103–106 (2020).
Beltrán-Nogal, A. et al. Surfing the wave of oxyfunctionalization chemistry by engineering fungal unspecific peroxygenases. Curr. Opin. Struct. Biol. 73, 102342 (2022).
Xu, X. et al. Peroxygenase-catalysed selective oxidation of silanes to silanols. Angew. Chem. Int. Ed. 62, e202302844 (2023).
Molina-Espeja, P. et al. Directed evolution of unspecific peroxygenase from Agrocybe aegerita. Appl. Environ. Microbiol. 80, 3496–3507 (2014).
Rotilio, L. et al. Structural and biochemical studies enlighten the unspecific peroxygenase from Hypoxylon sp. EC38 as an efficient oxidative biocatalyst. ACS Catal 11, 11511–11525 (2021).
König, R. et al. Novel unspecific peroxygenase from Truncatella angustata catalyzes the synthesis of bioactive lipid mediators. Microorganisms 10, 1267 (2022).
Gomez de Santos, P. et al. Functional expression of two unusual acidic peroxygenases from Candolleomyces aberdarensis in yeasts by adopting evolved secretion mutations. Appl. Environ. Microbiol. 87, e0087821 (2021).
Li, T. et al. A novel unspecific peroxygenase from Agaricus bisporus var. bisporus for biocatalytic oxyfunctionalisation reactions. Mol. Catal. 546, 113275 (2023).
Bormann, S. et al. Broadening the biocatalytic toolbox—screening and expression of new unspecific peroxygenases. Antioxidants (Basel) 11, 223 (2022).
Domenico, N. K. et al. Recombinant heme thiolate oxygenases. US Patent, US20220282227A1 (2022).
Barton, N. A. et al. Accessing low-oxidation state taxanes: is taxadiene-4(5)-epoxide on the taxol biosynthetic pathway?. Chem. Sci. 7, 3102–3107 (2016).
Hébert, M., Bellavance, G. & Barriault, L. Total synthesis of ginkgolide C and formal syntheses of ginkgolides A and B. J. Am. Chem. Soc. 144, 17792–17796 (2022).
Carro, J. et al. Modulating fatty acid epoxidation vs hydroxylation in a fungal peroxygenase. ACS Catal 9, 6234–6242 (2019).
González Benjumea, A. et al. Fatty acid epoxidation by Collariella virescens peroxygenase and heme-channel variants. Catal. Sci. Technol. 10, 717–725 (2020).
Knorrscheidt, A. et al. Accessing chemo- and regioselective benzylic and aromatic oxidations by protein engineering of an unspecific peroxygenase. ACS Catal 11, 7327–7338 (2021).
Gomez de Santos, P. et al. Engineering a highly regioselective fungal peroxygenase for the synthesis of hydroxy fatty acids. Angew. Chem. Int. Ed. 62, e202217372 (2023).
Münch, J. et al. Computational-aided engineering of a selective unspecific peroxygenase toward enantiodivergent β-ionone hydroxylation. ACS Catal 13, 8963–8972 (2023).
Jayadeepa, R. M. & Niveditha, M. S. Computational approaches to screen candidate ligands with anti- Parkinson’s activity using R programming. Curr. Top. Med. Chem. 12, 1807–1814 (2012).
Wu, T. et al. Reshaping substrate-binding pocket of leucine dehydrogenase for bidirectionally accessing structurally diverse substrates. ACS Catal 13, 158–168 (2023).
Yue, D., Ng, E. W. H. & Hirao, H. Hydrogen-bond-assisted catalysis: hydroxylation of paclitaxel by human CYP2C8. J. Am. Chem. Soc. 146, 30117–30125 (2024).
Wang, J.-Y. et al. Facile biosynthesis of taxadiene by a newly constructed Escherichia coli strain fusing enzymes taxadiene synthase and geranylgeranyl pyrophosphate synthase. Proc. Biochem. 122, 129–136 (2022).
Trumbly, R. J. et al. Amplified expression of streptomyces endo-beta-N-acetylglucosaminidase H in Escherichia coli and characterization of the enzyme product. J. Biol. Chem. 260, 5683–5690 (1985).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta. Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta. Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Mendoza, A., Ishihara, Y. & Baran, P. S. Scalable enantioselective total synthesis of taxanes. Nat. Chem. 4, 21–25 (2012).
Rontein, D. et al. CYP725A4 from yew catalyzes complex structural rearrangement of taxa-4(5),11(12)-diene into the cyclic ether 5(12)-oxa-3(11)-cyclotaxane. J. Biol. Chem. 283, 6067–6075 (2008).
Acknowledgements
We thank Xudong Kong at Shanghai Jiao Tong University for his valuable advice on protein crystallization and structure determination. This work was financially supported by the National Key Research and Development Program of China (2019YFA0905000, Jian He Xu), the Shanghai Commission of Science and Technology (23HC1400200, Hui-Lei Yu) and the National Natural Science Foundation of China (32271540, Hui-Lei Yu and U24A20484, Ran Hong), Shanghai Municipal Science and Technology Major Project (Jian He Xu). Diffraction data were collected at Shanghai Synchrotron Radiation Facility.
Author information
Authors and Affiliations
Contributions
Hui-Lei Yu designed the experiments. Mingyuan Lai played a central role in all experiments and completed the majority of the work. Yu Weng, Zhengyu Huang, and Longhao Huang contributed to the preparation and structural analysis of several compounds. Jian Wei, Xin-Yi Lu, and Hangcheng Li assisted with the construction and cultivation of various mutants. Zhijun Zhang, Lili Zhu, Jian-He Xu, and Ran Hong provided valuable support in designing the experiments and offered technical guidance.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Zhihua Zhou, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lai, M., Weng, Y., Wei, J. et al. Designing an oxidase toolbox for site-directed oxidation of taxanes. Nat Commun 17, 841 (2026). https://doi.org/10.1038/s41467-025-67544-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67544-4