Abstract
Colonic smooth muscle cell (CSMC) hypertrophy and hyperplasia have been described in both human ulcerative colitis (UC) and animal models. The deletion of smooth muscle (SM) 22α induces the phenotypic switching of SMCs. Here, we report that Sm22α-deficient mice develop spontaneous colitis, which is characterized by radical S-adenosyl-methionine domain-containing 2 (RSAD2)-driven mitochondrial dysfunction and inflammation in CSMCs and ferroptosis in the colonic mucosa. Mechanistically, RSAD2 mediates YTH m6A RNA-binding protein 1 (YTHDF1) methylation and activation, thereby increasing the mRNA N6-methyladenosine (m6A) modification and translation of dynamin-related protein 1 (DRP1), resulting in mitochondrial fragmentation in CSMCs. Inflammatory CSMC-derived mitochondrial extracellular vesicles trigger intestinal epithelial ferroptosis by inducing ROS production. The ablation of RSAD2 in mice with SMC-specific Sm22α knockout alleviates colitis severity in this experimental model. Importantly, increases in both RSAD2 expression and the ferroptotic signature are observed in serum and/or colonic samples from UC patients. Overall, this study shows a mitochondrial mechanism underlying the ability of dysfunctional smooth muscle to drive colitis and highlights the potential of targeting the RSAD2-YTHDF1 axis as an innovative therapeutic strategy for colitis.
Similar content being viewed by others
Introduction
Inflammatory bowel disease (IBD) is a chronic, relapsing, and remitting inflammatory disorder of the gut that includes two main phenotypes, Crohn’s disease (CD) and ulcerative colitis (UC), and significantly disrupts bowel function and patient quality of life1. Known disease risk alleles highlight key pathways in the pathogenesis of IBD, including innate and adaptive immunity, gut barrier function, and pathogen sensing and responses. Epithelial cells, goblet cells, stroma and immune cells in the intestinal mucosa interact with each other or with the intestinal microbiota and the immune system, sending and receiving signals2. Furthermore, an inappropriate and continuing mucosal immune response to commensal microbes in a genetically susceptible host is also involved in intestinal inflammation in the gut and IBD2. However, the underlying genes at risk loci have not been mapped to their cells or pathways of action.
The gastrointestinal tract comprises an epithelium surrounded by layers of smooth muscle cells (SMCs). Fully mature SMCs have a unique ability to reversibly alter their phenotype from a contractile state to a synthetic state to respond to environmental cues3. Smooth muscle (SM) 22α, a marker of mature SMCs, is involved in actin filament assembly and stress fiber formation4, which are required for maintaining the differentiated phenotype of SMCs5,6. Our group and others have demonstrated that disruption of SM22α in vascular SMCs enhances the vascular inflammatory response by inducing the phenotypic switching of SMCs7,8. SMC hypertrophy (enlarged muscle cells) and hyperplasia (increased numbers of muscle fibers) have been described in the inflamed intestine in both human IBD and animal disease models and are being increasingly recognized as the main contributors to increased wall thickness in the fibrotic process of IBD9. Furthermore, aberrant SMC phenotypes, as environmental variables, may contribute to dysregulation of the mucosal immune and commensal ecosystem in the gut, ultimately initiating colitis. However, the importance of the smooth muscle component in IBD has been less investigated. This prompted us to ask whether and how SM22α disruption promotes intestinal inflammatory damage.
Radical S-adenosyl-L-methionine domain-containing 2 (RSAD2) is an interferon-stimulated gene that is renowned for its antiviral properties. RSAD2 is involved in complex interplay with mitochondria, and binding of the iron–sulfur (Fe–S) cluster is necessary for the involvement of RSAD2 in mitochondrial metabolism. RSAD2 influences the proliferation and development of immune cells and inflammation via different signaling pathways10. However, the function of RSAD2 has been shown to vary across studies, and knowledge of this topic is lacking. Although a previous study demonstrated that increasing RSAD2 expression in neutrophils suppressed intestinal inflammation in a mouse model of DSS-induced colitis11,12, very little is known about the crucial role of RSAD2 in the pro-inflammatory phenotypic switching of SMCs.
An increasing number of studies have underscored the new roles of YTH N6-methyladenosine (m6A) RNA-binding protein 1 (YTHDF1) in the gut13. Although YTHDF1 is largely dispensable for intestinal homeostasis under physiological conditions, YTHDF1 is required for intestinal epithelial regeneration and tumorigenesis because it maintains stem cell traits by amplifying Wnt/β-catenin signaling at the translational level14. Ythdf1 mRNA is upregulated in ulcerative colitis (UC) patients15 and downregulated in both LPS-induced macrophages and the colons of IBD model mice, whereas the upregulation of YTHDF1 alleviates the inflammatory response16, suggesting that YTHDF1-mediated RNA methylation events are involved in IBD pathogenesis17. In the current study, we demonstrate that SM22α deletion activates the RSAD2-YTHDF1 axis, leading to mitochondrial fragmentation and a proinflammatory phenotype in colonic SMCs (CSMCs). Mitochondrial extracellular vesicles (MitoEVs) from CSMCs induce intestinal epithelial cell ferroptosis and colitis development.
Results
SM22α deletion leads to spontaneous colitis
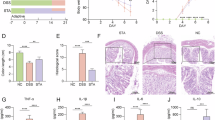
We first confirmed that the expression of SM22α was restricted to the muscular layer of the intestinal wall of wild-type (WT) mice (Supplementary Fig. 1a). Three-month-old Sm22α knockout (Sm22α−/−) mice exhibited signs of colitis. Weight loss was evident in these mice (Fig. 1a), which passed paste-like stools with high water contents (Supplementary Fig. 1b–d). Gross and microscopic examinations of Sm22α−/− mice revealed shorter colons (Fig. 1b) and increased histological activity index (HAI) scores, indicating architectural distortion and variation in the size and shape of crypts (Fig. 1c). Moreover, compared with their WT littermates, goblet cell loss, inflammatory cell infiltration and intestinal epithelial apoptosis, which are hallmarks of IBD18, were obvious in Sm22α-/− mice (Fig. 1c–e). Disrupted epithelial hypoxia (Fig. 1f) and increased plasma levels of FITC-conjugated dextran (Fig. 1g) indicated an increase in intestinal barrier permeability in Sm22α−/− mice, which was accompanied by impaired peristaltic contractions (Fig. 1h and Supplementary Fig. 1e, f). At the molecular level, the expression of the proinflammatory cytokines Tnf and Il-1β significantly increased in the colonic tissues of Sm22α-deficient mice, whereas the level of the anti-inflammatory cytokine Il-10 decreased (Fig. 1i), indicating an inflammatory phenotype. In addition, Sm22α−/− mice exhibited decreased proliferation and differentiation of intestinal epithelial cells without evidence of epithelial metaplasia (Fig. 1f and Supplementary Fig. 1g).
a Body weight of wild-type (WT) and SM22α-knockout (Sm22α−/−) mice from 8 to 16 weeks, n = 10 mice per group. b Representative images of colons and colon length quantification, n = 10 mice per group. c Representative haematoxylin and eosin (H&E)-stained colonic sections and corresponding histological activity index (HAI) scores, n = 10 mice per group. Arrows indicate infiltrating immune cells. Scale bar, 20 μm. d Alcian blue-Periodic acid Schiff (AB-PAS) staining of goblet cells and quantification, n = 10 mice per group. Scale bar, 20 μm. e TUNEL staining with haematoxylin counterstaining and quantification of TUNEL positive cells, n = 10 mice per group. Arrows indicate apoptotic epithelial cells. Scale bar, 50 μm. f Colonic sections stained for pimonidazole (green), Villin (orange) and Ki-67 (red), with quantification of Ki-67 positive cells. Nuclei were counterstained with DAPI (blue). n = 10 mice per group. Scale bar, 100 μm. g Serum fluorescein isothiocyanate (FITC)-dextran concentration as a measure of intestinal permeability, n = 10 mice per group. h Colonic transit time assessed by bead expulsion assay, n = 10 mice per group. i Cytokine mRNA expression in colonic tissue, n = 6 mice per group. j Body weight of floxed controls (Sm22αfl/fl) and smooth muscle-specific SM22α-knockout (smSm22α−/−) mice from 8 to 16 weeks, n = 10 mice per group. k Representative colon images and colon length measurements, n = 10 mice per group. l H&E-stained colonic sections and HAI scores, n = 10 mice per group. Arrows indicate infiltrating immune cells. Scale bar, 20 μm. m AB-PAS staining and quantification of goblet cells, n = 10 mice per group. Scale bar, 20 μm. n FITC-dextran assay for intestinal permeability, n = 10 mice per group. o Colonic transit time by bead expulsion assay, n = 10 mice per group. p Cytokine mRNA expression in colonic tissue, n = 6 mice per group. Data are presented as mean ± SEM of three independent experiments. P values were determined by 2-tailed Student’s t test. Source data are provided as a Source data file.
To determine the role of CSMCs in colitis development, we generated SMC-specific Sm22α knockout (smSm22α−/−) mice by crossing mice withloxP sites flanking Sm22α (Sm22α fl/fl mice) with SMC-specific Cre-expressing transgenic mice (myh11Cre mice). Similar colitis-related changes, including weight loss (Fig. 1j), shorter colons (Fig. 1k), elevated HAI scores (Fig. 1l), and goblet cell loss (Fig. 1m), were observed in 3-month-old smSm22α−/− mice. Additionally, intestinal permeability and peristaltic contraction were impaired (Fig. 1n, o) in these mice compared with Sm22αfl/fl mice, which was accompanied by increased inflammation (Fig. 1p).
To further investigate whether SM22α deletion in CSMCs plays a unique role in DSS-induced inflammation in mice, we administered DSS to induce colitis. Compared with Sm22αfl/fl control mice, smSm22α−/− mice had significantly shorter survival times (Fig. 2a), lower body weights (Fig. 2b), higher disease activity index (DAI) scores (Fig. 2c), shorter colons (Fig. 2d, e) and more severe tissue damage (Fig. 2f, g) 7d after the administration of DSS. Consistently, higher intestinal permeability and more severe peristaltic dysfunction and inflammation were observed in DSS-treated smSm22α−/− mice than in control mice (Fig. 2h–j). Additionally, we found that DSS impaired the contractility of CSMCs cultured in vitro (Supplementary Fig. 1h, i). Notably, the length of the small intestines shortened and the intestinal transit times of the marker that was orally administered, extended in Sm22α−/− mice compared with WT mice at baseline (Supplementary Fig. 2a-c). The small intestinal wall, especially the ileum, was significantly damaged with higher histological score in Sm22α−/− mice (Supplementary Fig. 2d), suggesting the small intestinal dysfunction. Nevertheless, no difference in the colitis phenotype between male and female Sm22α−/− mice was observed (Supplementary Fig. 2e–j). The rescue of SM22α expression by the administration of an adeno-associated virus 2/9 (AAV) carrying SM22α markedly ameliorated colitis phenotype in Sm22α−/− mice, with less pronounced colon shortening, tissue damage, peristaltic dysfunction and inflammation (Supplementary Fig. 3a–g). Collectively, these results suggest that the loss of SM22α in CSMCs plays a crucial role in colitis development.
a Survival rate during DSS exposure, n = 10 mice per group. b Daily body weight change, n = 10 mice per group. c Disease activity index (DAI) scores, n = 10 mice per group. d, e Representative images of colons and colon length quantification from Sm22αfl/fl and smSm22α−/− mice, n = 10 mice per group. f Representative haematoxylin and eosin (H&E)-stained colonic sections and corresponding histological activity index (HAI) scores, n = 10 mice per group. Scale bar, 20 μm. g Alcian blue-Periodic acid Schiff (AB-PAS) staining of goblet cells and quantification, n = 10 mice per group. Scale bar, 20 μm. h Measurement of intestinal permeability using Serum fluorescein isothiocyanate (FITC)-dextran levels in serum, n = 10 mice per group. i Colonic transit time assessed by bead expulsion assay, n = 10 mice per group. j Colonic mRNA expression of pro-inflammatory cytokines (Tnf and Il-1β) and anti-inflammatory cytokine (Il-10), n = 6 mice per group. Data are presented as mean ± SEM of three independent experiments. ns, not significant, P values were determined by 2-tailed Student’s t-test. Source data are provided as a Source data file.
SM22α deficiency disrupts mitochondrial homeostasis in CSMCs
To elucidate the molecular mechanisms linking SM22α deficiency to colitis, we performed RNA-seq and microarray analyses on colonic smooth muscle tissues from WT and Sm22α−/− mice. Transcriptomic analysis revealed 2,028 differentially expressed genes (DEGs), including 898 upregulated genes and 1,130 downregulated genes, in Sm22α−/− tissues (Supplementary Fig. 4a), as validated by quantitative PCR (qPCR) (Supplementary Fig. 4b). Gene Ontology (GO) enrichment analysis of the differentially expressed genes revealed that the upregulated genes were significantly associated with immune activation, inflammatory responses, and osteogenic processes. In contrast, the downregulated genes were primarily enriched in negative regulation of ossification, glycolysis, and fatty acid transport, suggesting that the colonic SMCs in Sm22α−/− mice undergo phenotypic switching characterized by proinflammatory and osteoblast-like transdifferentiation (Supplementary Fig. 4c). Notably, mitochondrial material transport, amino acid metabolism, oxidative phosphorylation, and ATP production regulation were significantly suppressed in the CSMCs of Sm22α−/− mice (Fig. 3a).
a Gene ontology terms enriched in up- and down- regulated genes from transcriptomes of colonic smooth muscle tissue isolated from Sm22α−/− mice compared to WT mice, n = 4 mice per group. b Representative transmission electron microscopy images of primary CSMCs. Arrows indicate morphological abnormalities. Scale bar, 1 μm. n = 4 biological replicates. c Immunofluorescence staining of mitochondria in WT and Sm22α−/− CSMCs using MitoTracker Deep Red (green), with nuclei counterstained with DAPI (blue)(lanes1-2). Mitochondrial superoxide levels were detected by MitoSox (red)(lane 3). Scale bar, 25 μm. d Quantification analysis of mitochondrial morphology from high-resolution confocal microscopy images (MitoTracker staining in c), n = 15 biological replicates. e Relative mitochondrial DNA (mtDNA) copy number measured by qRT-PCR in WT and Sm22α−/− CSMCs, n = 6 biological replicates. f Intracellular ATP levels in WT and Sm22α−/− CSMCs assessed using an ATP assay, n = 6 biological replicates. g, h Western blot analysis of mitochondrial fission regulators (FIS1 and DRP1) and fusion regulators (MFN1, MFN2 and OPA1) in WT and Sm22α−/− CSMCs with quantification of protein expression, n = 6 biological replicates. Data are presented as mean ± SEM of three independent experiments. ns, not significant, P values were determined by 2-tailed Student’s t test. Source data are provided as a Source data file.
Our previous study demonstrated that SM22α deficiency impaired mitochondrial structure and function in uterine SMCs, leading to contractility defects and dystocia19. Consistent with these findings, electron microscopy revealed mitochondrial shrinkage, severe cristae loss, and membrane rupture in primary Sm22α−/− CSMCs (Fig. 3b). We next evaluated mitochondrial morphology and discovered that mitochondrial fragmentation was substantially increased in Sm22α−/− cells. The mitochondrial number, mean length, and network branch counts were significantly lower in Sm22α−/− CSMCs than in WT controls (Fig. 3c, d). Moreover, mitochondrial dysfunction was further corroborated by elevated mitochondrial reactive oxygen species (mtROS) levels (Fig. 3c) and reduced mitochondrial DNA (mtDNA) copy numbers and ATP production (Fig. 3e, f). Notably, among mitochondrial fusion and fission markers, only dynamin-related protein 1 (DRP1) was markedly upregulated in Sm22α−/− CSMCs (Fig. 3g, h), which is consistent with a significant increase in mitochondrial fragmentation. Treatment with Mdivi-1, a selective DRP1 inhibitor, mitigated mitochondrial fragmentation in Sm22α−/− CSMCs (Supplementary Fig. 5a, b), suggesting that DRP1 plays an essential role in mitochondrial dysfunction in Sm22α−/− CSMCs.
Our previous research demonstrated that TNF induced the epigenetic silencing of SM22α transcription via H3K27 methylation in the SM22α promoter region in vascular SMCs8 and that the disruption of SM22α further aggravated vascular inflammation20, prompting us to explore whether the synergistic interaction of TNF stimulation and SM22α ablation impaired mitochondrial function in Sm22α−/− CSMCs. In line with our hypothesis, stimulation of CSMCs with TNF induced mitochondrial fragmentation and mtROS production (Supplementary Fig. 6a, b) and led to mitochondrial dysfunction (Supplementary Fig. 6c, d), which were similar to the phenotypic outcomes of Sm22α−/− CSMCs. Furthermore, TNF treatment significantly increased DRP1 protein levels (Supplementary Fig. 6e, f). Notably, TNF treatment exacerbated mitochondrial fragmentation and dysfunction in Sm22α−/− CSMCs (Supplementary Fig. 6g–l). Overall, these data indicate that inflammation signaling contributes to mitochondrial damage through the upregulation of DRP1 in CSMCs.
RSAD2 expression is upregulated during inflammation
To further elucidate the molecular mechanisms underlying mitochondrial dysfunction in Sm22α−/− CSMCs, we compared our RNA-seq data with 1,218 mitochondria-related genes cataloged in the MitoCarta 3.0 database21. This analysis revealed 43 overlapping genes, 9 of which were upregulated and 34 of which were downregulated in Sm22α−/− smooth muscle tissues (Fig. 4a, b). Notably, RSAD2, a free radical S-adenosylmethionine (SAM) enzyme associated with immune, inflammatory, and mitochondrial processes22, was significantly upregulated (Fig. 4b), suggesting its potential role in mediating mitochondrial dysfunction linked to SM22α deletion. Therefore, we examined the expression and distribution of RSAD2 in WT and Sm22α−/− colonic tissues and found that the mRNA and protein levels of RSAD2 were markedly increased throughout the entire intestinal wall (Fig. 4c–g), especially in the smooth muscle layers of Sm22α−/− mice (Fig. 4f). Parallel observations in human samples revealed thickening of the colonic muscularis propria and increased RSAD2 expression in both smooth muscle tissues and serum from patients with UC compared with controls without colitis (Supplementary Tables 1 and 2 and Supplementary Fig. 7a–d). Overall, these results suggest that RSAD2 plays a critical role in colitis pathogenesis.
a Venn diagram showing the overlap between differentially expressed genes (DEGs) from RNA-seq and mitochondria-related proteins from MitoCarta 3.0. b Heatmap of 43 mitochondria-related genes in WT and Sm22α−/− mice, n = 4 mice per group. c Rsad2 mRNA levels in mouse colonic tissue, n = 7 mice per group. d, e Western blot analysis and quantification of RSAD2 protein levels in mouse colonic tissue, n = 4 mice per group. f Immunofluorescence staining of RSAD2 (red) with DAPI counterstaining (blue) in mouse colonic tissue. Scale bar, 100 μm. g Quantification of RSAD2 immunofluorescence intensity (f), n = 7 mice per group. h Rsad2 mRNA levels in primary CSMCs, n = 7 biological replicates. i, j Western blot analysis and quantification of RSAD2 protein in CSMCs, n = 6 biological replicates. k Immunofluorescence staining for RSAD2 (red), and mitochrondria (green) in primary CSMCs. Nuclei were counterstained with DAPI (blue). Scale bar, 25 μm. l Colocalization analysis using Pearson’s correlation coefficient from (k), n = 7 biological replicates. m, n siRNA-mediated knockdown of SM22α in WT CSMCs, followed by Western blot analysis and quantification of RSAD2 levels, n = 6 biological replicates. o, p Western blot analysis and quantification of RSAD2 levels in Sm22α−/− CSMCs infected with Ad-EGFP and Ad-SM22α-EGFP, n = 6 biological replicates. q, r Western blot analysis and quantification of RSAD2 levels in WT CSMCs with or without tumor necrosis factor-α (TNF) treatment, n = 6 biological replicates. Data are presented as mean ± SEM of three independent experiments. P values were determined by 2-tailed Student’s t test. Source data are provided as a Source data file.
Next, we extended these findings by performing complementary experiments in CSMCs. Consistent with the in vivo observations, the expression of RSAD2 at both the mRNA and protein levels was significantly elevated (Fig. 4h–j), and RSAD2 colocalized mainly with mitochondria in the cytoplasm of Sm22α−/− CSMCs (Fig. 4k, l). Sm22α knockdown by a specific siRNA increased the expression of the RSAD2 protein (Fig. 4m, n), whereas rescuing Sm22α expression in Sm22α−/− CSMCs abolished the increase in RSAD2 protein expression (Fig. 4o, p). Consistently, TNF treatment significantly increased RSAD2 protein levels in CSMCs (Fig. 4q, r). Together, these results indicated that inflammatory signaling contributes to RSAD2 upregulation.
RSAD2 leads to mitochondrial fragmentation and dysfunction in CSMCs
RSAD2 is an endoplasmic reticulum (ER)-localized IFN-inducible iron sulfur (Fe–S) cluster-binding antiviral protein22. We next investigated whether RSAD2 is involved in mitochondrial dysfunction in CSMCs. Analysis of mitochondrial morphology revealed that mitochondrial fragmentation was induced by the overexpression of RSAD2 and that the majority of mitochondria were relatively short and punctate (Fig. 5a, b). Furthermore, mtROS levels markedly increased (Fig. 5a), which was accompanied by decreases in mtDNA copy numbers (Fig. 5c) and ATP production (Fig. 5d). Given the significant upregulation of DRP1 in Sm22α−/− CSMCs or upon TNF treatment, we examined the effect of RSAD2 on DRP1 expression. RSAD2 overexpression markedly increased the protein level of DRP1 in CSMCs (Fig. 5e, f), indicating that RSAD2 strongly affects mitochondrial dysfunction.
a Immunofluorescence staining of mitochondria (green) in CSMCs with or without RSAD2 overexpression (OE or CON; lanes 1-2). Nuclei were counterstained with DAPI (blue). Mitochondrial superoxide levels were detected by MitoSox (red) in WT CSMCs with or without RSAD2 overexpression (lane 3). Scale bar, 25 μm. b Quantification analysis of mitochondrial morphology from confocal microscopy images (a), n = 15 biological replicates. c Relative mtDNA copy number in WT CSMCs with or without RSAD2 overexpression assessed by qRT-PCR, n = 6 biological replicates. d Intracellular ATP levels in CSMCs measured by ATP assay, n = 6 biological replicates. e, f Western blot analysis and quantification of DRP1 protein levels, n = 6 biological replicates. g Immunofluorescence staining of mitochondria (green) in TNF-treated WT and Rsad2−/− CSMCs, with DAPI (blue) counterstaining (lanes 1-2). Mitochondrial superoxide levels were detected by MitoSox (red) in TNF treated WT and Rsad2−/− CSMCs (lane 3). Scale bar, 25 μm. h Quantification analysis of mitochondrial morphology from confocal microscopy images (g), n = 15 biological replicates. i Relative mtDNA copy number in TNF-treated WT and Rsad2−/− CSMCs measured by qRT-PCR, n = 6 biological replicates. j Intracellular ATP levels in TNF-treated WT and Rsad2−/− CSMCs, n = 6 biological replicates. k, l Western blot analysis and quantification of DRP1 protein levels, n = 6 biological replicates. Data are presented as mean ± SEM of three independent experiments. P values were determined by 2-tailed Student’s t-test. Source data are provided as a Source data file.
To further confirm the role of RSAD2 in mitochondrial dysfunction under inflammatory conditions, WT and Rsad2−/− CSMCs were treated with TNF. We showed that RSAD2 deficiency mitigated TNF-induced mitochondrial dysfunction, which was characterized by reduced mitochondrial fragmentation (Fig. 5g, h) and decreased mtROS levels (Fig. 5g). Additionally, compared with those in WT cells, the mtDNA copy number and ATP production in Rsad2−/− CSMCs increased upon TNF treatment (Fig. 5i, j). Consistent with these findings, DRP1 expression was significantly downregulated in Rsad2−/− CSMCs compared with WT controls following TNF treatment (Fig. 5k, l), further indicating that RSAD2 plays a dominant role in DRP1-related mitochondrial dysfunction.
We next investigated whether RSAD2 deficiency plays a unique role in protecting mice against DSS-induced colitis. WT and Rsad2−/− mice were given 2.5% DSS in their drinking water for 7 days. Compared with WT mice, Rsad2−/− mice exhibited significantly less weight loss, lower DAIs, and less colon shortening (Supplementary Fig. 8a–d) after DSS administration, findings that were confirmed by histological analysis (Supplementary Fig. 8e–g). Furthermore, DSS-treated Rsad2−/− mice survived longer (Supplementary Fig. 8h). Overall, our data support the conclusion that RSAD2 plays a crucial role in intestinal inflammation.
RSAD2 facilitates DRP1 translation through m6A deposition
To elucidate the molecular mechanism through which RSAD2 affects DRP1-related mitochondrial dysfunction, we examined the regulatory pathways involved in DRP1 protein expression, including mRNA transcription, stability, and translation, as well as ubiquitination and degradation, in CSMCs from WT and Rsad2−/− mice. We showed that the mRNA level of Drp1 did not change upon TNF treatment (Fig. 6a), and its decay kinetics did not significantly differ between WT and Rsad2−/− CSMCs upon actinomycin D treatment (Fig. 6b), suggesting that RSAD2 does not affect Drp1 mRNA transcription or stability. We then assessed the role of RSAD2 in DRP1 protein stability using ubiquitin immunoprecipitation assays. No significant differences in DRP1 ubiquitination or degradation were detected between WT and Rsad2−/− CSMCs (Fig. 6c), effectively ruling out protein stability as a regulatory mechanism. Taken together, these results indicated that RSAD2 may increase DRP1 expression at the translational level.
a Drp1 mRNA expression in TNF-treated WT and Rsad2−/− CSMCs, n = 6 biological replicates. b Drp1 mRNA stability in TNF-treated WT and Rsad2−/− CSMCs following actinomycin D treatment for the indicated times, n = 5 biological replicates. c Immunoblot analysis of ubiquitinated RSAD2 in CSMCs treated with 5 μM MG132 for 6 hours; IgG served as a control. d Total m6A RNA levels in TNF-treated WT and Rsad2−/− CSMCs, n = 6 biological replicates. e MeRIP-qPCR analysis of Drp1 m6A modification in TNF-treated WT and Rsad2−/− CSMCs, n = 6 biological replicates. f RNA pull-down assay followed by immunoblot demonstrating RSAD2 binding to Drp1 mRNA. g MeRIP-qPCR analysis of Drp1 m6A levels, n = 5 biological replicates. h MeRIP-qPCR analysis of Drp1 m6A levels in WT CSMCs transducted with siNC or siYTHDF1, n = 6 biological replicates. i Western blot analysis of RSAD2 levels in WT CSMCs transducted with siNC or siYTHDF1, n = 5 biological replicates. j RIP-qPCR analysis of YTHDF1 binding to Drp1 mRNA, n = 6 biological replicates. k RNA pull-down assay followed by immunoblot demonstrating YTHDF1 binding to Drp1 mRNA, n = 5 biological replicates. Data are presented as mean ± SEM of three independent experiments. P values were determined by 2-tailed Student’s t test (a, b, h and i) or one-way ANOVA followed by multiple comparisons test (d, e, g, j and k). Source data are provided as a Source data file.
The N6-methyladenosine (m6A) RNA modification plays a crucial role in regulating RNA metabolism, including its transport, decay, and translation. Given that RNA methylation activity is associated with RSAD223, we hypothesized that RSAD2 influences Drp1 mRNA translation via m6A deposition. Under inflammatory conditions, global m6A levels were elevated in CSMCs, whereas RSAD2 deficiency significantly attenuated m6A deposition (Fig. 6d). Methylated RNA immunoprecipitation (MeRIP)–qPCR analysis revealed significant enrichment of m6A on Drp1 mRNA in CSMCs following TNF treatment (Fig. 6e). In contrast, Rsad2−/− CSMCs exhibited markedly reduced m6A deposition under the same conditions, suggesting that RSAD2 modulates m6A deposition. However, unexpectedly, we did not detect an interaction between RSAD2 and Drp1 mRNA (Fig. 6f), which excluded the possibility that the enrichment of m6A on Drp1 mRNA was due to RSAD2-associated RNA methylation activity.
Next, we conducted RNA interference (RNAi) targeting key m6A regulatory proteins, including writers (METTL3 and METTL14), erasers (ALKBH5 and FTO), and readers (YTHDF1 and YTHDF3). Among these, knockdown of YTH domain family protein 1 (YTHDF1) significantly reduced m6A enrichment on the mRNA of Drp1 upon TNF treatment, as indicated by MeRIP–qPCR (Fig. 6g, h). m6A was recently shown to mediate the regulation of gene expression at the mRNA level through the direct recruitment of reader proteins to the transcript24. We showed that the expression of the DRP1 protein decreased after YTHDF1 knockdown (Fig. 6i), suggesting that YTHDF1, as a reader, may increase the translation of Drp1 mRNA by recognizing and binding to its m6A modifications.
To verify our speculation, we investigated the binding of YTHDF1 to Drp1 mRNA via RNA immunoprecipitation (RIP) assays with an anti-YTHDF1 antibody. TNF treatment increased the interaction of YTHDF1 with Drp1 mRNA. Drp1 mRNA was strongly enriched in the immunoprecipitated complexes (Fig. 6j). RNA pull-down assays confirmed that YTHDF1 binds to Drp1 mRNA (Fig. 6k), which is consistent with the increase in DRP1 protein expression. Importantly, the binding of YTHDF1 to Drp1 mRNA was markedly suppressed in Rsad2−/− CSMCs (Fig. 6j, k), indicating that YTHDF1 reader activity depends on RSAD2. Together, these data suggest that RSAD2 increases the translation of Drp1 mRNA through YTHDF1-mediated recognition of m6A modification.
RSAD2 mediates YTHDF1 methylation and activation
To determine how RSAD2 modulates YTHDF1 activity, we conducted computational docking analyses using the HDOCK server (Supplementary Fig. 9a). The top-scoring model predicted an interaction between RSAD2 and YTHDF1 (Supplementary Data 1). This interaction was validated by coimmunoprecipitation (CoIP) assays; furthermore, the binding interaction was enhanced following TNF treatment (Fig. 7a–c). Notably, the protein level of YTHDF1 did not change upon TNF treatment or RSAD2 deletion (Fig. 7d).
a, b Co-immunoprecipitation (CoIP) demonstrating RSAD2-YTHDF1 interaction, n = 5 biological replicates. c Western blot analysis showing enhanced RSAD2-YTHDF1 interaction upon TNF stimulation, n = 6 biological replicates. d TNF stimulation or RSAD2 deficiency did not alter YTHDF1 protein abundance, n = 6 biological replicates. e Immunoprecipitation of YTHDF1 followed by detection of its methylation using Pan me1/me2, n = 5 biological replicates. f WT and Rsad2−/− CSMCs, with or without TNF treatment, were subjected to YTHDF1 immunoprecipitation or IgG control, followed by Pan me1/me2 detection, n = 5 biological replicates. g Colonic tissues from DSS-treated Sm22αfl/fl and smSm22α−/− mice were analyzed as in f, n = 5 mice per group. h Colonic tissues from DSS-treated WT and Rsad2−/− mice were analyzed similarly, n = 5 mice per group. i Schematic of full-length and truncated RSAD2 constructs. j Proximity ligation assay (PLA) showing RSAD2-YTHDF1 interactions in HEK293A cells co-transfected with HA-YTHDF1 and the indicated RSAD2 constructs. PLA signals are shown in red and nuclei in blue (DAPI). n = 5 biological replicates. k COIP of HEK293A cells expressing HA-YTHDF1 and the indicated RSAD2 constructs, followed by HA immunoblotting, n = 5 biological replicates. l YTHDF1 methylation in cells treated as in (k), detected using the Pan me1/me2 antibody, n = 5 biological replicates. m CoIP of HEK293A cells expressing HA-YTHDF1 and either the SAM domain or GFP control, followed by HA detection, n = 5 biological replicates. n YTHDF1 methylation in cells treated as in (m), n = 5 biological replicates. o WT CSMCs transfected with the SAM domain or GFP were subjected to YTHDF1 RIP followed by qPCR quantification of Drp1 mRNA enrichment, n = 6 biological replicates. p DRP1 protein levels in WT CSMCs expressing the SAM domain or GFP. Data are presented as mean ± SEM from three independent experiments. ns, not significant. P values were determined by 2-tailed Student’s t test (c, g, h, o and p) or one-way ANOVA followed by multiple comparisons (d, f). Source data are provided as a Source data file.
Posttranslational modifications of readers may also alter their m6A binding affinity, cellular localization and interactions with protein partners25. Although RSAD2 is a radical SAM enzyme, neither the reaction it catalyzes nor its substrate has been identified23. We hypothesized that RSAD2 may mediate YTHDF1 methylation. To validate the physical and functional interaction of RSAD2 with YTHDF1, we performed immunoprecipitation (IP) assays with anti-α-mono- and anti-dimethyl lysine antibodies. Western blot analysis revealed that YTHDF1 was indeed methylated in CSMCs (Fig. 7e). Furthermore, TNF treatment increased YTHDF1 methylation, which was abolished by RSAD2 deletion (Fig. 7f). These results have been also validated in vivo. In DSS-induced colitis models, YTHDF1 methylation was increased in smSm22α−/− mice but reduced in Rsad2−/− mice (Fig. 7g, h). Overall, these data indicated that RSAD2 plays a dominant role in YTHDF1 methylation.
To further define the role of RSAD2 in YTHDF1 methylation, we generated a series of RSAD2 truncation mutants, including the SAM domain (GFP-SAM) and a truncated SAM domain (GFP-ΔSAM) (Fig. 7i). Proximity ligation assays (PLAs) in HEK293A cells co-transfected with the HA-YTHDF1 and RSAD2 constructs revealed that both full-length (FL) RSAD2 and its SAM domain interacted with YTHDF1, whereas the SAM domain deletion construct (GFP-ΔSAM) failed to bind (Fig. 7j). CoIP assays further confirmed that the SAM domain mediated the binding of RSAD2 to YTHDF1 (Fig. 7k). Consistently, both full-length RSAD2 and the SAM domain increased YTHDF1 methylation (Fig. 7l). We validated these results in HEK293A cells co-transfected with the HA-YTHDF1 and GFP-SAM constructs, which exhibited strong interactions between the SAM domain and YTHDF1 (Fig. 7m), accompanied by increased YTHDF1 methylation (Fig. 7n). Moreover, overexpression of GFP-SAM markedly increased the effect of YTHDF1 on Drp1 mRNA expression (Fig. 7o), which was accompanied by an increase in DRP1 protein expression in CSMCs (Fig. 7p), indicating that the RSAD2-YTHDF1 axis positively affects DRP1 expression. Together, these results suggest that YTHDF1 is an endogenous substrate of RSAD2.
In addition to recognizing m6A modifications, reader proteins may indirectly regulate m6A deposition. For example, m6A readers can compete with m6A demethylases for binding to m6A sites, resulting in varying degrees of m6A deposition26. As mentioned above, upon TNF treatment, Drp1 mRNA m6A deposition increased in CSMCs but decreased in Rsad2−/− cells (Fig. 6e). The results of the RNA pull-down experiments confirmed that the binding of ALKBH5, an m6A demethylase, to Drp1 mRNA markedly decreased with increased YTHDF1 expression in TNF-treated cells compared with control cells (Supplementary Fig. 10a, b). This decrease was reversed in Rsad2−/− CSMCs, which exhibited increased enrichment of ALKBH5 on Drp1 mRNA, coincident with the reduced m6A levels of Drp1 mRNA, suggesting that YTHDF1 occupancy at the m6A site of Drp1 mRNA suppresses ALKBH5 demethylase activity. Taken together, these findings suggest that RSAD2 promotes the YTHDF1-driven translational upregulation of Drp1 mRNA by mediating YTHDF1 methylation.
Inflammatory CSMC-derived mitochondrial extracellular vesicles induce ferroptosis in intestinal epithelial cells, contributing to colitis pathology
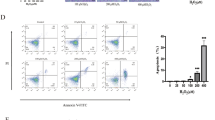
Recent advances have illustrated the functional importance of mitochondrial extracellular vesicle (MitoEV) release in an increasing number of tissue contexts27. Specific mitochondrial stresses or inflammatory signals trigger MitoEV release28. To verify the causal relationship between CSMC dysfunction and colitis, we first identified CD63-marked MitoEVs using immunofluorescence analysis in CSMCs (Supplementary Fig. 11a). We then isolated the MitoEVs from the conditioned medium of the CSMCs using differential centrifugation (Fig. 8a). Western blot analysis confirmed the presence of the mitochondrial outer membrane proteins TOM20 and TOM40 as well as the EV marker CD63 in CSMCs and in their released MitoEVs (Fig. 8b). Using FACS, we revealed that TNF treatment increased MitoEV release from CSMCs, which was significantly reduced in Rsad2−/− cells (Supplementary Fig. 11b). To trace MitoEV-mediated mitochondrial transfer, we first labeled the mitochondria in CSMCs with MitoTracker Deep Red and the mitochondria in intestinal epithelial cells (IECs) with MitoTracker Green. MitoEVs isolated from the conditioned medium of TNF-treated CSMCs were then used to treat IECs (HCT116 cells). Immunofluorescence analysis confirmed that the CSMC-derived MitoEVs were taken up by IECs, a process inhibited by dynasore, a clathrin-mediated endocytosis inhibitor (Fig. 8c), suggesting that intercellular mitochondrial transfer occurred between CSMCs and IECs.
a Workflow for isolation of mitoEVs secreted by CSMCs. Arrows indicate MitoTracker-positive staining (green). Nuclei were counterstained with DAPI (blue). Scale bar, 25 μm. b Western blot analysis of CSMC lysates and secreted mitochondria, n = 5 biological replicates. c Representative immunofluorescence co-staining of MitoTracker Deep Red (red)(mitochondria secreted by CSMCs) and MitoTracker Green (green)(mitochondria in IECs). Nuclei were counterstained with DAPI (blue). n = 5 biological replicates. Scale bar, 25 μm. d-e Confocal microscopy and quantification of lipid peroxidation in IECs following C11-BODIPY probe staining. Scale bar, 25 μm, n = 6 biological replicates. f–h Relative levels of malondialdehyde (MDA), glutathione (GSH), and total iron in IECs transfected with mitoEVs from distinct CSMCs, n = 6 biological replicates. i, j Confocal microscopy and quantification of intracellular free Fe2+ using FerroOrange staining (red). Scale bar, 25 μm, n = 6 biological replicates. k Western blot analysis and quantification of ferroptosis marker proteins, n = 6 biological replicates. l Representative immunofluorescence staining of COX2 (green) and GPX4 (red) in mouse colon tissue. Scale bar, 100 μm. m Quantification of (l) n = 6 mice per group. Data are presented as mean ± SEM of three independent experiments. ns, not significant. P values were determined by 2-tailed Student’s t test (m) or one-way ANOVA followed by multiple comparisons test (e–h, and j, k). Source data are provided as a Source data file.
Intercellular mitochondrial transfer not only rescues tissue damage but also contributes to multifunctional cellular activity and the regulation of inflammation29. An elegant study demonstrated that MitoEV-mediated mitochondrial transfer between cells in vivo resulted in increased mitochondrial ROS production in recipient cells30. To examine this possibility, we treated IECs with MitoEVs derived from different cellular contexts, including WT and Rsad2−/− CSMCs with or without TNF treatment. We observed increased lipid peroxidation (Fig. 8d, e) and elevated MDA levels (Fig. 8f) in IECs treated with MitoEVs of TNF-induced CSMCs, accompanied by a decreased GSH/GSSG ratio (Fig. 8g), suggesting increased oxidative stress. Similar results were observed in other colonic epithelial cell lines, including FHC and Caco-2 (Supplementary Fig. 11c, d). In contrast, Rsad2−/− MitoEVs seemed to have a smaller effect on lipid ROS levels (Fig. 8d–g). Together, these results suggested that MitoEVs from inflammatory CSMCs induced lipid peroxidation in the intestinal epithelium.
Lipid peroxidation products are the main drivers of ferroptosis, an iron-dependent form of necrotic cell death31. We showed that TNF-induced MitoEVs increased the levels of total ferrous iron and free Fe2+ in IECs (Fig. 8h–j). Moreover, expression of the ferroptosis marker COX2 significantly increased while GPX4 expression was decreased (Fig. 8k). In contrast, inflammatory Rsad2−/− MitoEVs had a mild effect on ferroptosis in IECs. In vivo, DSS-treated Rsad2−/− mice displayed less epithelial cell ferroptosis than the WT controls did (Fig. 8l, m and Supplementary Fig. 8i–k).
Similar to the findings in IECs, mouse intestinal organoids were shown to internalize CSMC-derived MitoEVs (Supplementary Fig. 12a, b). Exposure to inflammatory MitoEVs resulted in reduced expression of the barrier-associated protein Claudin-1 in colonic organoids, which was accompanied by elevated oxidative stress and the pronounced downregulation of the ferroptosis marker protein GPX4 and the upregulation of COX2 (Supplementary Fig. 12c–e). In contrast, inflammatory Rsad2−/− MitoEVs failed to induce substantial changes in ROS levels or ferroptosis markers in intestinal organoids (Supplementary Fig. 12c–e).
Given that RSAD2 expression is increased in Sm22α−/− mice with colitis, we investigated whether ferroptosis occurs in the intestinal mucosa in vivo. Western blot analysis revealed significant upregulation of COX2, which was accompanied by the downregulation of GPX4 (Supplementary Fig. 13a, b). Immunofluorescence analysis corroborated the increase in COX2 expression and the reduction in GPX4 expression (Supplementary Fig. 13c). Moreover, ferrous ion and MDA assays confirmed the occurrence of ferroptosis in the intestinal mucosa of Sm22α−/− mice (Supplementary Fig. 13d, e). Intriguingly, genetic deletion of RSAD2 in SMC-specific Sm22α knockout mice alleviated colitis severity (Supplementary Fig. 13f–h) and reduced epithelial ferroptosis (Supplementary Fig. 13i–m). Importantly, compared with those from healthy controls, the colonic mucosa from patients with UC exhibited a similar ferroptotic signature, with COX2 upregulation and GPX4 downregulation (Supplementary Fig. 7e). Together, these results further demonstrated that CSMC dysfunction caused intestinal epithelial damage.
Discussion
This study reveals the pivotal role of CSMC dysfunction in colonic inflammation through intercellular mitochondrial transfer and intestinal epithelial cell ferroptosis. We showed that SM22α deficiency-induced inflammation markedly increased the expression of RSAD2 in CSMCs, which increased the m6A modification of Drp1 mRNA via YTHDF1 methylation. This modification facilitates the translation of Drp1 mRNA, resulting in excessive mitochondrial fragmentation and CSMC functional impairment. Dysfunctional mitochondria are subsequently released from CSMCs via MitoEV and engulfed by IECs, triggering oxidative stress, ferroptosis, and epithelial injury and culminating in colitis.
In addition to being involved in motor function, SMCs that undergo phenotypic switching can synthesize and secrete proinflammatory cytokines, chemotaxis-associated molecules, and growth factors, which have been studied in detail in vascular muscle32. SMC plasticity likely also occurs in the gut33 and could have important implications for the pathogenesis of different gut motor disorders. The smooth muscle hypertrophy and hyperplasia observed in both human IBD and animal disease models are generally considered to result from intestinal inflammation, but little is known about the ability of SMCs to modulate intestinal epithelial function. In the current study, we showed that the colonic SMCs of smSm22α−/− mice underwent phenotypic switching are proinflammatory and exhibit osteoblast-like transdifferentiation. DSS-treated smSm22α−/− mice displayed more tissue damage and a higher pathological score, further suggesting the involvement of SMC phenotypic switching in colitis pathogenesis.
RSAD2 (also known as Viperin, cig5, or vig1) is localized to the ER and is an interferon (IFN)-inducible multifunctional protein in various cell types34. RSAD2 is constitutively expressed at high levels in the liver, heart, adipose tissue, and some immune cells. Research has shown that RSAD2 expression in the immune system can be directly or indirectly induced by various stimuli, including viral infection, interferons, synthetic analogs of double-stranded RNA or B-form DNA, and LPS, via IFN-dependent and IFN-independent pathways35. The basal and IFN-induced expression of RSAD2 is negatively regulated by a multiexonic nucleus-localized long noncoding RNA known as lncRNA-CMPK2 (cytidylate monophosphate kinase 2)36. IFN-independent induction of RSAD2 expression can occur through interferon regulatory factor 3 (IRF3), activator protein 1 (AP-1), and IRF135. In addition to its broad-spectrum antiviral activity37, RSAD2 also regulates cellular lipid metabolism38, mediates Toll-like receptor signaling pathways39, and plays broader roles in cellular function and proliferation40. Additionally, RSAD2 is localized to mitochondria in brown adipose tissue cells41 and transiently in human foreskin fibroblasts, where it regulates the β-oxidation of fatty acids and energy production38. A recent study demonstrated that IFNλ can modulate the function of neutrophils by upregulating RSAD2, leading to the suppression of intestinal inflammation in a mouse model of DSS-induced colitis, which was recapitulated in human neutrophils11,12. Here, we provide contradictory evidence concerning the role of RSAD2 in DSS-induced colitis. Increased expression of RSAD2 was observed in the colonic muscle layer of mice with colitis, colonic samples from UC patients, and CSMCs that had undergone proinflammatory phenotypic switching. RSAD2 contributed to mitochondrial fragmentation and dysfunction in CSMCs by increasing DRP1 expression and activity in a YTHDF1-dependent manner. Furthermore, after treatment with DSS, the colon shortening, tissue damage, weight loss, and disease activity index scores of Rsad2−/− mice significantly decreased. We speculated that the positive or negative effects of RSAD2 on colitis may be associated with cell type. Together, our findings provide evidence for the cell type-specific function of RSAD2 and show that increased expression of RSAD2 in CSMCs plays a dominant role in the development of colitis.
YTHDF1, a critical m6A reader protein, plays a key role in the increased translation of m6A-modified mRNAs. YTHDF1 promotes the translation of target transcripts, and this activity is regulated by its glycan modification and modification of its partner RBP Fragile X mental retardation protein (FMRP)42. YTHDF1 activity is also regulated by other posttranslational modifications, such as O-GlcNAc glycosylation43, ubiquitination44, and lactylation45. Furthermore, increased YTHDF1 expression has been demonstrated to be associated with disruption of intestinal barrier integrity in CD patients and colitis mice46. Deletion of YTHDF1 induces CD8+ T cells to stimulate the production of high levels of programmed cell death 1 ligand 1, ultimately exacerbating IBD47. YTHDF1 overexpression increases the mRNA expression of immunosuppressive molecules in an m6A-dependent manner and restructures the environment of inflammation in colitis48. Additionally, the increase in DNA copy number and the overexpression of YTHDF1 are considered frequent events in UC and inflammatory colorectal cancer49,50. Our study demonstrated a methylation-dependent mechanism of YTHDF1 activity in IBD pathogenesis. We revealed that RSAD2 directly interacted with YTHDF1 via its SAM domain, promoting the lysine methylation of YTHDF1, which was accompanied by increased YTHDF1-driven translation efficiency of m6A-modified Drp1 mRNA, contributing to SMC-driven intestinal epithelial inflammation. Further investigations are needed to determine whether targeting YTHDF1 may increase the efficacy of current IBD therapeutic regimens.
Given that the substrate of RSAD2 is not a protein but a common metabolite23, we speculated that YTHDF1 methylation may not be directly catalyzed by the radical SAM activity of RSAD2 but may indirectly result from SAM accumulation. The binding of YTHDF1 to RSAD2 might lead to the inability of RSAD2 to catalyze the reductive cleavage of SAM51, resulting in SAM accumulation. In fact, both the accumulation of SAM and the exhaustion of the cellular pool of this metabolite affect the SAM-dependent methyl transferases of DNA, protein, and phospholipids, which are involved in different cellular processes52. This discovery provides insights into the regulatory landscape of m6A readers and their posttranslational regulation under inflammatory conditions, offering promising avenues for therapeutic intervention.
The functions of mitochondria are not only associated with their metabolic roles in producing energy and generating essential metabolites but also linked to diverse processes, including cell signaling, the initiation of apoptosis, immune responses, and the regulation of cell proliferation and development53. Emerging studies have revealed that cells can also export some of their mitochondria for delivery to recipient cells, contributing to both normal physiological processes and disease pathogenesis. The release and capture of extracellular MitoEVs as cargo constitute common mechanisms of intercellular mitochondrial transfer. These MitoEVs are marked with the tetraspanins CD63, CD9, and CD81, which are used to eliminate damaged mitochondrial components. MitoEVs enable intercellular communication with either beneficial or harmful effects on recipient cells, depending on the cellular source as well as the nature of the cargo and the originating stimulus54. Brown adipocytes undergoing thermogenic stress eliminate damaged mitochondrial components via the formation of MitoEVs27. Osteoblasts release MitoEVs containing fragmented mitochondria to promote osteoprogenitor cell differentiation into osteoblasts55. Our current study revealed that TNF treatment increased MitoEV release from CSMCs, which was significantly reduced in Rsad2−/− cells. Furthermore, CSMC-derived MitoEVs were taken up by IECs. The TNF-induced increase in the number of MitoEVs resulted in an increase in the total ferrous iron content; moreover, the expression of the ferroptosis marker COX2 increased in IECs while the expression of GPX4 decreased, and these effects were reduced in Rsad2−/− MitoEV-treated IECs. Moreover, the induction of ferroptosis by inflammatory CSMC-derived MitoEVs was also observed in intestinal crypt organoids. In line with this finding, ferroptosis increased in cells in the intestinal mucosa of Sm22α−/− mice but decreased in Rsad2−/− mice. Overall, our data suggest that CSMCs with an inflammatory phenotype exacerbate intestinal inflammation via the induction of ferroptosis by MitoEVs.
In the present study, we cannot exclude the potential effects of vascular smooth muscle on the intestinal mucosa and disease phenotype via changes in gut vascular barrier function. Given the importance of the muscular component in IBD, a mouse model of IBD induced by colonic smooth muscle-specific inactivation of the Sm22α gene is necessary in future studies. How RSAD2 expression is induced in the CSMCs of the inflamed intestine in vivo or in vitro, and what intermediate molecule mediates RSAD2-driven YTHDF1 methylation, are unclear. Additionally, a more in-depth characterization of inflammatory CSMC-derived MitoEV biology is needed to enable the use of these cells as biomarkers in clinical studies.
In conclusion, this study reveals a pathophysiological mechanism linking smooth muscle dysfunction and intestinal epithelial damage, emphasizing the importance of mitochondrial dynamics-related CSMC dysfunction in the development of colitis. RSAD2 and its downstream effectors, including YTHDF1 and DRP1, may serve as biomarkers for colitis activity and severity, enabling more precise disease monitoring and facilitating personalized treatment.
Methods
Study approval
Approval for all the experimental procedures was granted by the Institutional Laboratory Animal Care and Use Committee of Hebei Medical University, Permit No. IACUC-Hebmu-P2023006.
Human samples
The study involving humans received approval from the Medical Ethics Committee of Hebei Medical University, and informed consent was obtained from all participants (2023-001). Between May 2023 and May 2024, colonic tissue and blood samples were collected via endoscopic biopsies at the Second Hospital of Hebei Medical University. UC was diagnosed by colonoscopy and pathological examination of colon tissue samples. This research was conducted with the voluntary participation of individuals who were not offered any compensation. Sex or gender information was not collected, and no sex or gender analysis was carried out.
Animals
All animal studies were carried out in accordance with the guidelines sanctioned by the Ethics Committee for Animal Experiments and the Institutional Animal Care and Use Committee at Hebei Medical University. The experimental procedures followed the guidelines set by Directive 2010/63/EU of the European Parliament regarding the protection of animals used in scientific research. Cyagen Biosciences (S-KO-11081) provided C57BL/6 J wild-type (WT) mice and Rsad2−/− mice. Sm22α−/− mice were acquired from the Jackson Laboratory (006878), and Myh11-cre/ERT2 mice were obtained from Youdu Bio (YDS0815). All the mice were kept in a pathogen-free facility under controlled conditions, including a temperature ranging from 21 to 23 °C, a humidity ranging from 50 to 60%, and a 12-h light/dark cycle. Mice aged 12–16 weeks were used for the experiments. Tamoxifen was administered intraperitoneally (i.p.) at 100 mg kg−1 in corn oil, either as a single dose or every other day for 14 days, starting at 5–8 weeks of age.
Culture and treatment of CSMCs
After being isolated from the colonic muscular layer of young mice, the CSMCs were cultured in high-glucose Dulbecco’s modified Eagle’s medium (DMEM; Invitrogen) supplemented with 20% fetal bovine serum (FBS; Gibco) and 1% penicillin‒streptomycin (PS; Biological Industries). The cells were incubated in a humidified environment at 37 °C with 5% CO2. Only cells from passages 4 through 10 were used in the experiments. The cells were serum-starved by incubation in 0.5% FBS for 24 h before being stimulated with TNF (10 ng mL−1) and transfected with siRNA or plasmids.
Culture and treatment of intestinal organoids
Organoid cultures and crypt cells were isolated using methods detailed in earlier studies56. The procedures involved dissecting the colon lengthwise, washing it with cold DPBS, and incubating the crypts in Gentle Cell Dissociation Reagent (STEMCELL Technologies) at room temperature on a rocking platform at 20 rpm for 20 min. A 70-μm cell strainer was used to filter the crypt suspension, and the crypt fraction was collected by centrifugation at 290 × g for 5 minutes at 4 °C. The crypt pellets were then embedded in Matrigel (BD Biosciences) on ice and subsequently plated in 24-well plates. Once Matrigel polymerization was complete, 750 μL of complete IntestiCult Organoid Growth Medium (STEMCELL Technologies) was added to each well at room temperature. The culture medium was changed every 2 days. The mitoEVs from the CSMCs were used to treat the organoids for 48 h prior to additional experiments.
Methylated RNA immunoprecipitation (MeRIP) assay
In accordance with the manufacturer’s protocol, the m6A modification level of the Drp1 mRNA was determined using an EpiQuik CUT & RUN m6A RNA Enrichment Kit (P-9018; Epigentek). Briefly, m6A-containing RNA fragments were selectively isolated using beads conjugated with an anti-m6A capture antibody. The alterations in m6A methylation of Drp1 mRNA were quantified by qPCR.
Plasmid or siRNA transfection
The cultured CSMCs were grown to 50–60% cofluence and then transfected with overexpression plasmid (General Biosystems, Anhui, China) or siRNA (Gene Pharma, Suzhou, China) using HighGene transfection reagent (ABclonal, Wuhan, China) according to the manufacturer’s instructions. All plasmids were constructed using standard methods and confirmed by DNA sequencing. The siRNA sequences used for siRNA transfection are listed in Supplementary Table.
Statistical analysis
Unless specified in the figure legends, the data are displayed as the means ± SEMs. Statistical analyses were performed using GraphPad Prism (version 10.1.2; GraphPad Software, La Jolla, CA, USA). Comparisons between two groups were evaluated using two-tailed Student’s t tests. For analyses involving more than two groups, one-way ANOVA followed by Tukey’s post hoc multiple comparisons test was applied. The data and images represent at least three independent experiments. Statistical significance was defined as P < 0.05.
The Online Data Supplement contains an expanded methods section.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data generated in this study have been deposited in the NCBI Sequence Read Archive (SRA) under accession code PRJNA1312247(ID 1312247 - BioProject - NCBI). All other data are available in the article and its Supplementary files. Source data are provided with this paper.
References
Ramos, G. P. & Papadakis, K. A. Mechanisms of disease: inflammatory bowel diseases. Mayo Clin. Proc. 94, 155–165 (2019).
Khor, B., Gardet, A. & Xavier, R. J. Genetics and pathogenesis of inflammatory bowel disease. Nature 474, 307–317 (2011).
Scirocco, A. et al. Cellular and molecular mechanisms of phenotypic switch in gastrointestinal smooth muscle. J. Cell Physiol. 231, 295–302 (2016).
Zhang, J. C. et al. Analysis of SM22alpha-deficient mice reveals unanticipated insights into smooth muscle cell differentiation and function. Mol. Cell. Biol. 21, 1336–1344 (2001).
Fu, Y. et al. Mutagenesis analysis of human SM22: characterization of actin binding. J. Appl. Physiol. 89, 1985–1990 (1985).
Han, M. et al. Smooth muscle 22 alpha maintains the differentiated phenotype of vascular smooth muscle cells by inducing filamentous actin bundling. Life Sci. 84, 394–401 (2009).
Shen, J. et al. Disruption of SM22 promotes inflammation after artery injury via nuclear factor kappaB activation. Circ. Res. 106, 1351–1362 (2010).
Shu, Y. N. et al. CKII-SIRT1-SM22α loop evokes a self-limited inflammatory response in vascular smooth muscle cells. Cardiovasc. Res. 113, 1198–1207 (2017).
Speca, S., Giusti, I., Rieder, F. & Latella, G. Cellular and molecular mechanisms of intestinal fibrosis. World J. Gastroenterol. 18, 3635–3661 (2012).
Chen, S. et al. Crucial roles of RSAD2/viperin in immunomodulation, mitochondrial metabolism and autoimmune diseases. Inflammation 48, 520–540 (2025).
Broggi, A., Tan, Y., Granucci, F. & Zanoni, I. IFN-λ suppresses intestinal inflammation by non-translational regulation of neutrophil function. Nat. Immunol. 18, 1084–1093 (2017).
Starling, S. Neutrophils: interfering with intestinal inflammation. Nat. Rev. Immunol. 17, 594 (2017).
Wang, H., Han, J. & Zhang, X. A. Interplay of m6A RNA methylation and gut microbiota in modulating gut injury. Gut Microbes 17, 2467213 (2025).
Han, B. et al. YTHDF1-mediated translation amplifies Wnt-driven intestinal stemness. EMBO Rep. 21, e49229 (2020).
Motawi, T. K., Shaker, O. G., Amr, G. & Senousy, M. A. RNA methylation machinery and m(6)A target genes as circulating biomarkers of ulcerative colitis and Crohn’s disease: correlation with disease activity, location, and inflammatory cytokines. Clin. Chim. Acta 561, 119831 (2024).
Xu, X. et al. hucMSC-Ex alleviates inflammatory bowel disease in mice by enhancing M2-type macrophage polarization via the METTL3-Slc37a2-YTHDF1 axis. Cell Biol. Toxicol. 40, 74 (2024).
Sebastian-delaCruz, M. et al. Implication of m6A mRNA methylation in susceptibility to inflammatory bowel disease. Epigenomes 4, 16 (2020).
Kurashima, Y. & Kiyono, H. Mucosal ecological network of epithelium and immune cells for gut homeostasis and tissue healing. Annu. Rev. Immunol. 35, 119–147 (2017).
Gao, Y. et al. Smooth muscle 22α deficiency impairs oxytocin-induced uterine contractility in mice at full-term pregnancy. Biochem. Biophys. Res. Commun. 529, 884–889 (2020).
Shu, Y. N. et al. SM22α inhibits vascular inflammation via stabilization of IκBα in vascular smooth muscle cells. J. Mol. Cell Cardiol. 84, 191–199 (2015).
Rath, S. et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 49, D1541–D1547 (2021).
Seo, J. Y., Yaneva, R., Hinson, E. R. & Cresswell, P. Human cytomegalovirus directly induces the antiviral protein viperin to enhance infectivity. Science 332, 1093–1097 (2011).
Fenwick, M. K., Li, Y., Cresswell, P., Modis, Y. & Ealick, S. E. Structural studies of viperin, an antiviral radical SAM enzyme. Proc. Natl. Acad. Sci. USA 114, 6806–6811 (2017).
Liu, Y. et al. N (6)-methyladenosine RNA modification-mediated cellular metabolism rewiring inhibits viral replication. Science 365, 1171–1176 (2019).
He, P. C. & He, C. m(6) A RNA methylation: from mechanisms to therapeutic potential. EMBO J. 40, e105977 (2021).
Wang, J. et al. ALKBH5 attenuates mitochondrial fission and ameliorates liver fibrosis by reducing Drp1 methylation. Pharmacol. Res. 187, 106608 (2023).
Rosina, M. et al. Ejection of damaged mitochondria and their removal by macrophages ensure efficient thermogenesis in brown adipose tissue. Cell Metab. 34, 533–548.e12 (2022).
König, T. & McBride, H. M. Mitochondrial-derived vesicles in metabolism, disease, and aging. Cell Metab. 36, 21–35 (2024).
Liu, D. et al. Intercellular mitochondrial transfer as a means of tissue revitalization. Signal Transduct. Target Ther. 6, 65 (2021).
Brestoff, J. R. et al. Intercellular mitochondria transfer to macrophages regulates white adipose tissue homeostasis and is impaired in obesity. Cell Metab. 33, 270–282.e8 (2021).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Owens, G. K., Kumar, M. S. & Wamhoff, B. R. Molecular regulation of vascular smooth muscle cell differentiation in development and disease. Physiol. Rev. 84, 767–801 (2004).
Nair, D. G., Han, T. Y., Lourenssen, S. & Blennerhassett, M. G. Proliferation modulates intestinal smooth muscle phenotype in vitro and in colitis in vivo. Am. J. Physiol. Gastrointest. Liver Physiol. 300, G903–G913 (2011).
Seo, J. Y., Yaneva, R. & Cresswell, P. Viperin: a multifunctional, interferon-inducible protein that regulates virus replication. Cell Host Microbe 10, 534–539 (2011).
Rivera-Serrano, E. E. et al. Viperin reveals its true function. Annu. Rev. Virol. 7, 421–446 (2020).
Kambara, H. et al. Negative regulation of the interferon response by an interferon-induced long non-coding RNA. Nucleic Acids Res. 42, 10668–10680 (2014).
Rivieccio, M. A. et al. TLR3 ligation activates an antiviral response in human fetal astrocytes: a role for viperin/cig5. J. Immunol. 177, 4735–4741 (2006).
Seo, J. Y. & Cresswell, P. Viperin regulates cellular lipid metabolism during human cytomegalovirus infection. PLoS Pathog. 9, e1003497 (2013).
Saitoh, T. et al. Antiviral protein Viperin promotes Toll-like receptor 7- and Toll-like receptor 9-mediated type I interferon production in plasmacytoid dendritic cells. Immunity 34, 352–363 (2011).
Aguilar, H. et al. Biological reprogramming in acquired resistance to endocrine therapy of breast cancer. Oncogene 29, 6071–6083 (2010).
Eom, J. et al. Intrinsic expression of viperin regulates thermogenesis in adipose tissues. Proc. Natl. Acad. Sci. USA 116, 17419–17428 (2019).
Zou, Z. & He, C. The YTHDF proteins display distinct cellular functions on m(6)A-modified RNA. Trends Biochem. Sci. 49, 611–621 (2024).
Chen, Y. et al. O-GlcNAcylation determines the translational regulation and phase separation of YTHDF proteins. Nat. Cell Biol. 25, 1676–1690 (2023).
Wang, A. et al. USP47 inhibits m6A-dependent c-Myc translation to maintain regulatory T cell metabolic and functional homeostasis. J. Clin. Investig. 133, e169365 (2023).
Liu, B. et al. Severe fever with thrombocytopenia syndrome virus induces lactylation of m6A reader protein YTHDF1 to facilitate viral replication. EMBO Rep. 25, 5599–5619 (2024).
Wang, J. et al. Elevated SPARC disrupts the intestinal barrier integrity in Crohn’s disease by interacting with OTUD4 and activating the MYD88/NF-κB pathway. Adv. Sci. 12, e2409419 (2025).
Du, Y. et al. Role of epigenetic modifications and aging in inflammatory bowel disease. Medcomm Future Med. 2, e63 (2023).
Chen, Q. et al. WTAP/YTHDF1-mediated m6A modification amplifies IFN-γ-induced immunosuppressive properties of human MSCs. J. Adv. Res. 71, 441–455 (2025).
Gu, C. et al. Immune infiltration of Ulcerative Colitis and detection of the m6A subtype. J. Immunol. Res. 2022, 7280977 (2022).
Wang, S. et al. N6-methyladenosine reader YTHDF1 promotes ARHGEF2 translation and RhoA signaling in colorectal cancer. Gastroenterology 162, 1183–1196 (2022).
Nelp, M. T., Young, A. P., Stepanski, B. M. & Bandarian, V. Human viperin causes radical SAM-dependent elongation of escherichia coli, hinting at its physiological role. Biochemistry 56, 3874–3876 (2017).
Honarmand Ebrahimi, K. A unifying view of the broad-spectrum antiviral activity of RSAD2 (viperin) based on its radical-SAM chemistry. Metallomics 10, 539–552 (2018).
Borcherding, N. & Brestoff, J. R. The power and potential of mitochondria transfer. Nature 623, 283–291 (2023).
Picca, A. et al. Circulating extracellular vesicles: friends and foes in neurodegeneration. Neural Regen. Res. 17, 534–542 (2022).
Suh, J. et al. Mitochondrial fragmentation and donut formation enhance mitochondrial secretion to promote osteogenesis. Cell Metab. 35, 345–360.e7 (2023).
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32371161 to M.H., 82370474 to P.K., and 82400508 to D.D.Z.), the S&T Program of Hebei (22377759D to Y.B.L.), and the Graduate Student Innovation Fund of Hebei Province (CXZZBS2023101 to W.D.Z).
Author information
Authors and Affiliations
Contributions
M.H. designed the study. W.D.Z., X.W., D.D.Z., X.L.L., S.Z.Z., W.S., X.Q.L., L.J.L., H.B.W., and D.M.L. performed the animal study and analyzed the data. W.D.Z. and P.K. performed the sequencing experiments, bioinformatics analyses, and statistical analyses. X.L.Z., Y.B.L., and C.Y.L. collected the human samples and provided material support. D.D.Z. and F.Q.L. were responsible for visualization. W.D.Z. assisted with the drafting of the manuscript. M.H. is the principal investigator who obtained funding and drafted the manuscript. All the authors reviewed, contributed to, and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, WD., Zhang, DD., Wang, X. et al. Activation of the RSAD2-YTHDF1 axis in smooth muscle causes inflammatory bowel disease via intercellular mitochondrial transfer. Nat Commun 17, 976 (2026). https://doi.org/10.1038/s41467-025-67707-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67707-3