Abstract
Cyclin-dependent kinase 4/6 inhibitors (CDK4/6i) combined with endocrine therapy are the standard first-line treatment for hormone receptor-positive, HER2−negative (HR+/HER2−) metastatic breast cancer, but resistance inevitably develops. In triple-negative breast cancer (TNBC), the efficacy of CDK4/6i remains uncertain. Our study shows that the selective CDK2 inhibitor BLU-222, while effective alone, enhances synergistic activity when combined with CDK4/6i in resistant HR+/HER2− and TNBC models, leading to increased apoptosis and cell cycle arrest. In vivo, combining BLU-222 with palbociclib or ribociclib produced significant antitumor activity across eight resistant models, driving durable tumor regression and prolonged survival. Mechanistically, BLU-222, alone or with palbociclib, upregulated p21 and p27 expression, enhanced p21 binding to CDK2 as well as p21 and p27 binding to CDK4. CRISPR knockout of p21 or p27 in palbociclib-resistant cells eliminated this synergy. Further, RNA sequencing revealed that combination treatment upregulated senescence and interferon pathways, providing mechanistic insight into the observed therapeutic synergy.
Similar content being viewed by others
Introduction
Cyclin-dependent kinases (CDKs) are essential regulators of the cell cycle, particularly in cell division and proliferation. CDK2, when bound to cyclin E or cyclin A, drives the G1-S phase transition and DNA replication1. Like CDK4/6, CDK2 phosphorylates the retinoblastoma protein (Rb) and targets other substrates necessary for DNA replication and S-phase progression2,3. CDK2 activity is tightly regulated by endogenous inhibitors, including p21 and p27, which ensure controlled cell cycle progression3. However, dysregulated CDK2 activity leads to uncontrolled cell proliferation, contributing to tumorigenesis in various cancers, including breast cancer.
The development of selective CDK2 inhibitors has historically faced challenges due to the high structural similarity among CDKs and the conserved nature of their ATP-binding sites. This has complicated the design of inhibitors with high specificity for CDK2. Early CDK2 inhibitors like flavopiridol and roscovitine lacked selectivity and had significant off-target effects, limiting their clinical utility4,5. Although dinaciclib demonstrated improved potency against CDK2, it also exhibited off-target toxicity, hindering broader applications6,7. However, recent advancements have led to the development of more selective CDK2 inhibitors such as BLU-222 (Blueprint Medicines), INX-315 (Incyclix Bio), INCB-123667 (Incyte), ARTS-021 (Allorion/Avenzo Therapeutics) and PF-07104091 (Pfizer), which show promise in preclinical models of cancers with aberrant CCNE1 and CDK4/6 inhibitor (CDK4/6i)-resistant breast cancers8,9.
CDK2 inhibition not only regulates cell cycle progression but can also induce cellular senescence, a state of irreversible growth arrest8. Senescence can be triggered by factors such as DNA damage or oncogene activation10, and the process involves the upregulation of p21 and p27, which inhibit CDK2 activity11. This, in turn, activates the p53 and Rb pathways, reinforcing the senescent state by blocking further cell cycle progression12. Additionally, senescence can promote the recruitment of immune cells via the senescence-associated secretory phenotype (SASP), aiding in the removal of senescent tumor cells13.
CDK2 inhibition has also emerged as a critical strategy for overcoming resistance to CDK4/6i in breast cancer. In hormone receptor-positive, HER2−negative (HR+/HER2−) breast cancers, upregulation of CDK2 activity, often driven by cyclin E overexpression14,15,16, enables tumor cells to bypass CDK4/6 inhibition, sustaining proliferation despite treatment17,18. This compensatory mechanism suggests that targeting CDK2 could provide a therapeutic approach to overcome resistance in these cancers19. Additionally, CDK2 inhibition holds promise in triple-negative breast cancer (TNBC), an aggressive subtype where certain tumors rely on CDK2/cyclin E-driven proliferation14,15,16. CDK2 inhibitors are also being explored in other cancer types, including CCNE1-abarrent ovarian and small cell lung cancers20,21.
BLU-222 represents a significant advancement in selective CDK2 inhibition, demonstrating strong antitumor activity in preclinical models of CCNE1-abarrent ovarian cancer and HR+/HER2− breast cancers that have progressed on CDK4/6i9. These findings paved the way for the VELA trial (NCT05252416), which evaluated BLU-222 as a monotherapy in patients with CCNE1-amplified solid tumors or in combination with ribociclib and fulvestrant in CDK4/6i-resistant breast cancers22,23. Previously presented results from the trial indicated that BLU-222 was generally well tolerated in patients and accompanied by markers of cell cycle modulation and anti-tumor activity observed22. Combining BLU-222 with ribociclib has shown no additional safety concerns and has led to reductions in thymidine kinase 1 and Rb activity, supporting its potential in combination therapies24.
In this study, we evaluate the efficacy of BLU-222 in combination with CDK4/6i in both in vitro and in vivo models of HR+/HER2− and TNBC subtypes. We hypothesize that dual inhibition of CDK4/6 and CDK2 can overcome resistance mechanisms, induce apoptosis, and more effectively halt tumor progression compared to single-agent treatments. Our study integrates transcriptomic and functional analyses to uncover the molecular mechanisms of p21 and p27 underlying the synergistic effects of BLU-222 and CDK4/6i, identifies biomarkers of response, and establishes a preclinical foundation for future clinical trials. These findings offer a promising therapeutic strategy for managing CDK4/6i-resistant breast cancers and improving patient outcomes.
Results
High levels of full-length or low molecular weight cyclin E sensitize CDK4/6 inhibitor-resistant breast cancer cells to BLU-222 across different subtypes
To examine the biological selectivity of BLU-222, we performed knockdown experiments in HCC1806, a TNBC cell line with CCNE1 amplification, targeting cyclin E (shCCNE1) and CDK2 (shCDK2) (Fig. 1A). Knocking down CDK2 and CCNE1 caused a 4-fold and 2-fold reduction in sensitivity to BLU-222 treatment compared to scrambled control cells. The IC50 values were 0.3 µM in control cells, compared to 1.1 µM in shCDK2 and 0.6 µM in shCCNE1 cells (Fig. 1B). Additionally, BLU-222 induced significant apoptosis and G1 cell cycle arrest in control cells, but these effects were attenuated in the shCCNE1 and shCDK2 cells (Fig. 1C, D, Supplemental Fig. 1A, B). These findings support the dependency of BLU-222’s antitumor activity on cyclin E and CDK2.
A Western blots of cyclin E, CDK2, pCDK2 (Thr160), Rb, and pRb (Ser807/811) in HCC1806 parental cells, scramble control, and CCNE1 or CDK2 knockdown cells. B Representative dose-response curves and IC50 values for BLU-222-treated HCC1806 scramble control and CCNE1 and CDK2 knockdown cell lines. Error bars represent standard error of the mean (SEM) from n = 3 experimental replicates. C Annexin V (+) quantification in HCC1806 control and CCNE1 and CDK2 knockdown cells treated with BLU-222 at 0.5 µM in for 0, 24, and 48 h. The experiments were done as two biological replicates. D Cell cycle distribution of HCC1806 control and CCNE1 and CDK2 knockdown cells treated with BLU-222 at 0.5 µM, measured by flow cytometry using propidium iodide (PI) staining. The experiments were done as two biological replicates. E Western blots of cyclin E1, CDK2, pCDK2 (Thr160), CDK4, CDK6, cyclin D1, and pRb (Ser807/811) in HCC1806 parental and CCNE1 or CDK2 knockdown cells, treated with BLU-222 at 0.3 µM, 0.6 µM, and 1.2 µM for 3 days. F Representative dose-response curves and IC50 values for palbociclib-treated HCC1806 parental and CCNE1 and CDK2 knockdown cells. Error bars represent SEM from n = 3 experimental replicates. G Western blots of cyclin E, CDK2, and pCDK2 (Thr160) in T47D parental cells, empty vector (EV), full-length cyclin E (EL), or low-molecular-weight truncated cyclin E (LMW-E) cells. H Representative dose-response curves and IC50 values for BLU-222-treated T47D parental cells, EL, and LMW-E. Error bars represent SEM from n = 3 experimental replicates. I Annexin V (+) quantification in T47D parental, EL, and LMW-E cells treated with palbociclib and/or BLU-222 (0.5 µM each) for 3 days. The experiments were done as 3 biological replicates. J-M IC50 values distribution for 15 breast cancer cell lines treated with single agents of J palbociclib, K abemaciclib, L ribociclib, or M BLU-222. HR+/HER2− cell lines include MCF7, T47D, and CAMA1. The HER2-enriched cell lines include BT474, SKBR3, MDA-MB-361, UACC812, HCC1569, and HCC1954. The triple-negative (TNBC) cell lines include MDA-MB-231, BT20, HCC38, MDA-MB-157, MDA-MB-468, and HCC1806. Each box plot displays the median (center line), the 25 and 75th percentiles (bounds of the box), and the minimum and maximum values (whiskers). Individual data points represent independent cell lines. N-P Correlation between BLU-222 IC50 values and those of N palbociclib, O abemaciclib, and P ribociclib. Differences between two groups were analyzed using a two-tailed unpaired Student’s t-test, and for multiple-group comparisons, a one-way ANOVA with Tukey’s multiple comparisons test was applied. The Pearson correlation coefficient (r) and associated p value were calculated for correlation analyses (ns, not significant).
Western blot analysis revealed that BLU-222 treatment elevated cyclin E1 levels in control and shCDK2 cells, while increasing CDK2 expression in shCCNE1 cells (Fig. 1E). Additionally, CDK4, CDK6, and cyclin D1 were upregulated in control and shCCNE1 cells, and while CDK6 and cyclin D1 induction was not observed in shCDK2 cells, CDK4 levels did increase following BLU-222 treatment (Fig. 1E, Supplemental Fig. 1C–E). Notably, pRb (Ser807/811)/Rb levels were significantly increased in all BLU-222-treated cells compared to untreated controls (Supplemental Fig. 1F). These findings indicate that BLU-222 affects both the CDK2 and CDK4/6 pathways by inducing compensatory upregulation of key cyclins and kinases in each pathway when the other is inhibited. This crosstalk suggests that BLU-222 modulates CDK2 and CDK4/6 activity in a coordinated manner, allowing breast cancer cells to adapt by shifting dependence between these pathways in response to targeted inhibition.
Further investigations revealed that knockdown of CDK2 or CCNE1 in HCC1806 cells sensitized them to palbociclib. Dose-response analysis showed that shCDK2 and shCCNE1 cells were 12-fold and 10-fold more sensitive to palbociclib than control cells, with IC50 values of 0.15 µM and 0.11 µM, respectively, compared to 1.8 µM in control cells (Fig. 1F). Similarly, CDK2 knockdown in MDA-MB-468, another TNBC cell line with CCNE1 amplification, increased sensitivity to palbociclib by 3-fold (Supplemental Fig. 1G). Conversely, CDK2 knockdown reduced BLU-222 sensitivity by approximately 2-fold in MDA-MB-468 cells (Supplemental Fig. 1H). These findings further support the notion of crosstalk between CDK2 and CDK4/6 pathways, suggesting that combined targeting of these pathways offers a therapeutic advantage in CDK4/6i-resistant breast cancers.
To assess how full-length cyclin E (EL) or low-molecular weight cyclin E (LMW-E) influences sensitivity to BLU-222, we treated T47D (HR+/HER2−) and MDA-MB-231 (TNBC) cells expressing either EL or LMW-E with BLU-222. LMW-E forms, generated by proteolytic cleavage, are known hyperactive forms of cyclin E that exhibit increased oncogenic activity14,25. Western blot analysis confirmed elevated expression levels of CDK2 and pCDK2 in LMW-E cells in T47D (Fig. 1G). Dose-response analysis revealed that LMW-E cells were more sensitive to BLU-222, with an IC50 value of 0.8 µM, compared to 6.3 µM in EL-expressing cells (Fig. 1H). BLU-222 treatment increased apoptosis in LMW-E-high cells and caused G2/M phase accumulation in EL and LMW-E cells (Fig. 1I, Supplemental Fig. 1I–K). Similar results were observed in LMW-E-overexpressing MDA-MB-231 cells [29], with a 2.5- to 4.5-fold decrease in BLU-222 IC50 values in LMW-E cells compared to control cells (Supplemental Fig. 1L). Interestingly, the impact of EL expression differed by breast cancer subtype. In T47D cells, a luminal cell line, EL overexpression conferred relative resistance to BLU-222 compared with parental cells, consistent with prior evidence that EL can delay cell cycle progression and checkpoint activation26. In contrast, EL-overexpressing MDA-MB-231 cells, a basal cell line, were ~10-fold more resistant to palbociclib but showed increased sensitivity to CDK2 inhibition, with a 3.8-fold reduction in BLU-222 IC50 and a 6-fold reduction with INX-315 compared to parental cells (Supplemental Fig. 1M). To directly validate these isoform-specific effects in a controlled background, we used CCNE1-knockout 76NE6 human immortalized mammary epithelial cells reconstituted with doxycycline-inducible EL or LMW-E (76NE6-EKO)25. Both isoforms increased CDK2 activation and Rb phosphorylation, but LMW-E induction produced robust sensitization to CDK2 inhibition, with an IC50 of 0.97 µM, approximately 16.6-fold lower than empty vector controls and 10-fold lower than EL-expressing cells (Supplemental Fig. 1N-O). In contrast, EL induction yielded only modest sensitization or, in some contexts, relative protection against BLU-222. Together, these findings demonstrate that LMW-E consistently drives CDK2 hyperactivation and therapeutic vulnerability across subtypes, whereas the effects of EL are context dependent, attenuating sensitivity in luminal T47D cells but enhancing it in basal MDA-MB-231 cells. Collectively, these results highlight the oncogenic role of LMW-E as a biomarker of CDK2 inhibitor sensitivity and underscore the importance of tumor subtype and isoform context in determining therapeutic response. Importantly, they also suggest an inverse relationship between palbociclib and BLU-222 sensitivity, where LMW-E expression confers resistance to CDK4/6 inhibition but enhances susceptibility to CDK2 inhibition, providing a mechanistic rationale for dual targeting in cyclin E–driven tumors.
To further test this hypothesis, we screened 15 breast cancer cell lines, comprising 3 HR+/HER2−, 6 HER2−enriched, and 6 TNBC subtypes, using three FDA-approved CDK4/6i (palbociclib, abemaciclib, and ribociclib) and BLU-222 (Supplemental Table S1). HR+HER2− cell lines were the most sensitive to CDK4/6i, followed by HER2−enriched and TNBC subtypes (Fig. 1J–L). Conversely, TNBC cell lines, were the most sensitive to BLU-222, followed by HER2−enriched and HR+/HER2− subtypes (Fig. 1M). Correlation analysis revealed a significant negative correlation between the IC50 values for the CDK4/6i and BLU-222 across all subtypes (Fig. 1N–P). This inverse correlation supports the complementary nature of responses to CDK4/6i and BLU-222, highlighting the therapeutic potential of dual targeting across different breast cancer subtypes.
Palbociclib resistance sensitizes HR+HER2− and TNBC cells to BLU-222, and combining BLU-222 with palbociclib results in synergistic tumor cell killing
Previous studies have shown that elevated cyclin E-CDK2 activity is a key driver of both intrinsic and acquired resistance to CDK4/6i in HR+/HER2− breast cancer27,28,29,30,31,32,33. Consistent with these reports, our data demonstrate significant upregulation of cyclin E (both full-length and low molecular-weight-forms) and CDK2 in palbociclib-resistant HR+/HER2− cells, accompanied by increased E2F signature expression28. These findings suggest that activation of cyclin E-CDK2 axis bypasses CDK4/6 inhibition, sustains E2F-driven transcription, and promotes drug resistance and cancer progression.
To evaluate whether BLU-222 could overcome palbociclib resistance, we tested BLU-222 in acquired palbociclib-resistant (PR) MCF7 and T47D cells, which were developed28 through gradually exposure to increasing concentrations of palbociclib, starting at 1.2 µM and escalating to 4.8 µM (Fig. 2A). Parental MCF7 and T47D cells and their PR counterparts, were treated with escalating doses of palbociclib or BLU-222 for 6 consecutive days, followed by 6 days drug-free period, after which cell viability was assessed. As expected, MCF7 PR cells exhibited 3–10 fold increased resistance to palbociclib relative to parental cells but were 5–6 fold more sensitive to BLU-222 (Fig. 2B, Supplemental Fig. 2A–C). Similarly, T47D PR cells showed a 4- to 8-fold increase in palbociclib resistance, while displaying 2-17-fold greater sensitivity to BLU-222 (Fig. 2C, Supplemental Fig. 2D–F). Together, these results highlight an inverse relationship between sensitivity to palbociclib and BLU-222 (Fig. 1N–P), supporting the ability of BLU-222 to overcome palbociclib resistance. Cell cycle and apoptosis analyses further revealed that, BLU-222 induced G2/M phase arrest and apoptosis in PR1.2 cells, whereas no significant changes were observed in parental cells (Supplemental Fig. 2G-J). In contrast, palbociclib treatment induced G1 phase arrest in parental cells, but had minimal effects in PR1.2 cells (Supplemental Fig. 2G-J). Based on the inverse sensitivities observed between palbociclib and BLU-222 in MCF7- and T47D PR cells, we hypothesized that combined BLU-222 and palbociclib treatment could enhance therapeutic efficacy in CDK4/6i-resistant breast cancer. To test this hypothesis, we performed combination treatments in parental and PR MCF7 and T47D cells using a high-throughput survival assay and evaluated drug interactions using the highest single agent (HSA) model, which quantifies combination efficacy relative to the most active single agen34.
A Generation of palbociclib-resistant (PR) cell lines by stepwise dose escalation from 1.2 to 4.8 µM. B,C IC50 values for MCF7 and T47D panels treated with single-agent palbociclib or BLU-222. Error bars represent SEM from n = 3 experimental replicates. D Representative 3D synergy plots for MCF7 parental and PR1.2 cells treated with different doses of palbociclib and BLU-222. E Representative 3D synergy plots for T47D parental and PR1.2 cells treated with different doses of palbociclib and BLU-222. F Annexin V (+) quantification in MCF7 parental and PR1.2 cells treated with single agents or combination. Error bars represent SEM from n = 3 experimental replicates. G Cell cycle distribution of MCF7 parental and PR1.2 cells treated with palbociclib and/or BLU-222 (0.3 µM, 3 days), measured by flow cytometry using PI staining. H Annexin V (+) quantification in T47D parental and PR1.2 cells treated with single agents or combination. The experiments were done as two biological replicates. I Cell cycle distribution of T47D parental and PR1.2 cells treated with palbociclib and/or BLU-222 (0.3 µM, 3 days). J Representative 3D synergy plots for HCC1806 and MDA-MB-157 cells treated with different doses of palbociclib and BLU-222. K Annexin V (+) quantification in HCC1806 and MDA-MB-157 cells treated with single agents or combination. The experiments were done as two biological replicates. L Cell cycle distribution of HCC1806 and MDA-MB-157 cells treated with palbociclib and/or BLU-222 (0.3 µM, 3 days), measured by flow cytometry using PI staining. The experiments were done as two biological replicates. M Representative 3D synergy plots for MDA-MB-231 parental and PR cells treated with different doses of palbociclib and BLU-222. N Annexin V (+) quantification in MDA-MB-231 parental and PR cells treated with single agents or combination. Error bars represent SEM from n = 3 experimental replicates. O Cell cycle distribution of MDA-MB-231 parental and PR cells under combination and single-agent treatment (0.3 µM, 3 days, n = 3). P Heatmap of highest single agent (HSA) synergic scores across HR+ , TNBC and acquired palbo-resistant breast cancer cell lines. Error bars represent SEM in at least three independent experiments. Comparisons between two treatment groups were analyzed using a two-tailed unpaired Student’s t-test; for comparisons among multiple groups, a one-way ANOVA with Tukey’s multiple comparisons test was applied.
Drug combination assays demonstrated strong synergy between BLU-222 (0.3 µM) and palbociclib (0.3 µM) in MCF7 PR1.2 cells, but not in parental or PR4.8 cells (Fig. 2D, Supplemental Fig. 2K). The lack of synergy in parental and PR4.8 cells is due to their inherent sensitivity to CDK4/6 inhibition (in parental cells) or BLU-222 (in PR4.8 cells) as single agents, resulting in high monotherapy activity that limits additional benefit from combination treatment. Consistently, HSA scores showed a 3-fold increase in synergy in PR1.2 cells compared to parental cells (HSA = 23 in PR1.2 vs. HSA = 7.4 in parental cells; Supplemental Fig. 2I). Similarly, T47D PR1.2 cells displayed significant synergy, while parental cells exhibited additive effects (Fig. 2E, Supplemental Fig. 2L). The HSA score in T47D PR1.2 cells was approximately 3.5-fold higher than in parental cells (HSA = 18.7 in PR1.2 vs. HSA = 5.3 in parental cells, Supplemental Fig. 2L), supporting combined CDK2 and CDK4/6 inhibition as a strategy to overcome resistance in these cell lines.
Previous studies have established Rb as a critical determinant of sensitivity to CDK4/6 inhibitor35. We assessed RB status across all cell lines and PDX models, including parental, resistant, and engineered lines (Supplemental Tables S1 and S2). RB-proficient models included HR+/HER2− MCF7 and T47D and TNBC MDA-MB-231; RB-deficient models included TNBC lines MDA-MB-468, HCC1806, and MDA-MB-157, as well as CDK4/6i-resistant clones T47D TCL and MCF7 MCL, where RB loss arose during long-term selection (Supplemental Fig. 2M and N). We also generated an RB knockdown MCF7 model (MCF7 shRB). Among the resistant pools, PR1.2 cells retained RB but exhibited high cyclin E expression, whereas PR4.8 cells showed complete RB loss, consistent with our prior reports28. As expected, RB-deficient TNBC lines, resistant pools (MCF7 PR4.8, T47D 4.8), resistant clones (TCL, MCL), and MCF7 shRB were refractory to CDK4/6i monotherapy but remained highly sensitive to BLU-222 (Supplemental Fig. 2M-P). Notably, BLU-222 monotherapy was particularly effective in RB-deficient PR4.8 cells, underscoring that CDK2 activity can drive proliferation independently of RB. BLU-222 plus palbociclib also produced strong synergy in RB-deficient TNBC models (HCC1806 and MDA-MB-157) and resistant clones (MDA-MB-231PR and BT20 PR), and showed additive effect in MCF7 shRB cells (Fig. 2J, M, Supplemental Fig. 2Q and R), with marked apoptosis and G2/M arrest (Fig. 2K, L, N, O, Supplemental Fig. 2S, T). In RB-proficient MCF7 and T47D parental cells, the combination was additive regardless of RB status. Collectively, these findings establish that while RB loss confers resistance to CDK4/6i, it does not diminish sensitivity to CDK2 inhibition; rather, BLU-222 effectively exploits CDK2/cyclin E–driven replication stress vulnerabilities in both RB-intact (PR1.2) and RB-deficient (PR4.8) contexts, providing a strong rationale for therapeutic targeting of this subset.
Apoptosis analysis showed that in MCF7 parental cells, palbociclib or combination treatment moderately increased apoptotic annexin V(+) cells ( ~ 6%), whereas BLU-222 alone did not increase apoptosis. In contrast, MCF7 PR1.2 cells treated with BLU-222 alone ( ~ 6%) or in combination with palbociclib ( ~ 10%) exhibited enhanced apoptosis compared to palbociclib alone or vehicle control (1–2%) (Fig. 2F, Supplemental Fig. 2K). Cell cycle analysis revealed G1 phase arrest with palbociclib or the combination in parental MCF7 cells, whereas MCF7 PR1.2 cells showed G2/M phase arrest with BLU-222 or the combination treatment (Fig. 2G, Supplemental Fig. 2P). T47D PR1.2 cells mirrored these results, with the combination treatment significantly increasing apoptosis and inducing G2/M phase arrest (Fig. 2H–I, Supplemental Fig. 2Q, R). These results suggest that BLU-222 effectively targets resistant cells by inducing apoptosis and cell cycle arrest through different mechanisms than in parental cells.
Next, we explored the combination effects of BLU-222 and palbociclib in TNBC cell lines in the context of both intrinsic and acquired resistance to CDK4/6i. In HCC1806 and MDA-MB-157 cells, which are intrinsically resistant due to cyclin E amplification, combination treatment with BLU-222 (0.3 µM) and palbociclib (0.3 µM) showed strong synergistic effects with HSA values of 23–30 (Fig. 2J). Apoptosis analysis confirmed a significant increase in Annexin V(+) apoptotic cells with BLU-222 alone, with further enhancement observed in combination-treated cells (Fig. 2K). Cell cycle analysis showed G2/M arrest in response to BLU-222 in both cell lines, with combination treatment amplifying this effect (Fig. 2L).
To evaluate the effects of BLU-222 and palbociclib in TNBC with acquired resistance, we generated palbociclib-resistant MDA-MB-231 and BT20 cells by gradually increasing palbociclib concentrations. Combination treatment of these PR cells revealed strong synergy (HSA scores 13.3–14.8) at low doses (0.3 µM each), while parental counterparts displayed only minimal synergy (HSA scores 2.4-4.1) (Fig. 2M and Supplemental Fig. 2L). Functional assays supported these findings: in PR cells, the combination increased apoptosis and promoted G2/M arrest, whereas parental cells primarily arrested in G1 without induction of apoptosis (Fig. 2N, O and Supplemental Fig. 2S, T). These data indicate that CDK2 inhibition preferentially enhances therapeutic efficacy in cells with acquired resistance to CDK4/6 inhibition.
To determine whether the observed synergy was subtype-specific, we compared responses across HR+ and TNBC parental and resistant models using HSA modeling (Fig. 2P). This analysis revealed a consistent trend: all parental cells, irrespective of subtype, exhibited additive effects, whereas all resistant cells displayed strong synergy, establishing BLU-222/palbociclib synergy as a hallmark of acquired resistance rather than lineage. In HR+ (MCF7 and T47D) and TNBC (MDA-MB-231 and BT20) parental cells, additive responses likely reflect their inherent sensitivity to CDK4/6 inhibition, while both HR+ (MCF7 PR1.2 and T47D PR1.2) and TNBC (MDA-MB-231 PR and BT20 PR) resistant cells showed enhanced synergy driven by compensatory CDK2 activation during CDK4/6i escape. The strongest effects were observed in TNBC models with intrinsic resistance due to cyclin E amplification, where BLU-222 effectively suppressed CDK2-driven cell cycle progression and yielded HSA scores approaching 30. Together, these findings demonstrate that BLU-222 and palbociclib combination therapy is effective across breast cancer subtypes, with activity in HR+/HER2− and TNBC and is particularly potent in tumors with intrinsic (through cyclin E amplification) or acquired resistance to CDK4/6 inhibition.
The induction of p21 and/or p27 in PR cells by BLU-222 mediates synergistic response to CDK4/6 inhibitors in combination treatment
Mechanisms of resistance to CDK4/6i often involve the overexpression of cyclin D1, CDK4, and CDK6, driving persistent cell cycle progression despite inhibition. Additionally, deregulation of cyclin E1 and CDK2 allows cells to bypass CDK4/6 inhibition, further promoting resistance18,32. To elucidate the mechanisms underlying the observed synergy between BLU-222 and CDK4/6 inhibition, we examined cell cycle protein expression in MCF7 and T47D parental cells, their palbociclib-resistant (PR) derivatives, and RB-deficient variants following treatment with BLU-222 (0.3 µM), palbociclib (0.3 µM), or the combination for three days (Fig. 3A, B). Western blot analysis confirmed elevated cyclin E1 and phosphorylated CDK2 in HR + PR cells, consistent with their CDK4/6i-resistant phenotype28. Cyclin D1 was decreased in T47D but remained stable in MCF7, while CDK4 expression was stable or modestly increased in both lines. In contrast, TNBC PR models (MDA-MB-231-PR and BT20-PR; Supplemental Fig. 3A–B) showed no consistent elevation of cyclin E1 or CDK2, and cyclin D1/CDK4 levels varied, underscoring the heterogeneity of resistance mechanisms across subtypes. Despite these differences, a unifying feature across resistant models was the consistent elevated levels of p21 and/or p27 following BLU-222 treatment, either alone or in combination with palbociclib. In parental HR+ cells, p21 levels rose only with the combination, while p27 induction was diminished by palbociclib alone. Notably, in RB-deficient resistant cells, p21 was strongly induced by the combination, whereas p27 expression remained largely unchanged under all conditions. These findings indicate that RB is partially required for p27 induction in response to CDK4/6 blockade, while p21 induction can be triggered by CDK2 inhibition independently of RB. Thus, across HR+ and TNBC contexts, and irrespective of RB status, BLU-222 drives Cip/Kip upregulation, highlighting p21 and p27 as central mediators of the synergy between CDK2 and CDK4/6 inhibition.
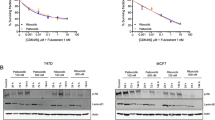
A,B Representative Western blots showing expression levels of cyclin E1, E2, A, B1, D1, CDK2, CDK4, phosphorylated and total Rb, p21, p27 and actin in (A) MCF7 parental, PR1.2, and shRB1 knockdown cell lines and B T47D parental and PR1.2 cell lines treated with 0.3 µM of single agents or a combination of palbociclib and BLU-222 for 3 days. Blots are representative of 3 biologically independent experiments performed on separate cell preparations with similar results. The samples were derived from the same experiment, and all gels/blots were processed in parallel under identical conditions. C Western blots showing successfully CRISPR p21 knockout (KO) in MCF7 parental and PR1.2 cells. Bar graph showing the HSA synergistic scores in MCF7 parental (n = 5) and PR1.2 cells (n = 4) treated with the combination of BLU-222 and palbociclib, as well as pooled (n = 4) and single clones of cells with p21 KO (A4, n = 4; C12, n = 4; CL1, n = 4; CL3, n = 3). Error bars represent SEM from n = 3 experimental replicates. D Representative 3D synergy plots illustrating the response of (D) MCF7 parental cells and p21 KO single clone (A4) and E MCF7 PR1.2 and p21 KO single clone (CL1) to combination treatment with different doses of palbociclib and BLU-222. F Western blots showing successfully CRISPR p27 KO in MCF7 parental (n = 5) and PR1.2 cells (n = 4). Bar graph showing the HSA synergistic scores in MCF7 and PR1.2 cells treated with the combination of BLU-222 and palbociclib, as well as pooled (n = 3) and single clones of cells with p27 KO (A4, n = 2; E9, n = 4; CL1, n = 4; CL3, n = 4). Error bars represent SEM from n = 3 experimental replicates. G Representative 3D synergy plots illustrating the response of MCF7 p27 KO single clone (A4) and PR1.2 p27 KO single clone (CL1) to combination treatment with different doses of palbociclib and BLU-222. H Heatmap showing HSA scores across MCF7 parental (par) and PR1.2 CRISPR KO pools and single clones for p21 or p27. I Bar graph showing the IC50 of MCF7 PR1.2 parental and CRISPR KO p21 pools or single clones treated with palbociclib (PR1.2, n = 4; pools, n = 4; CL1, n = 3; CL3, n = 5) or BLU-222 as single agents (PR1.2, n = 4; pools, n = 3; CL1, n = 4; CL3, n = 4). Box plots show the median (center line), 25th–75th percentiles (bounds of the box), and minimum–maximum values (whiskers). Error bars represent SEM from n = 3 experimental replicates. Comparisons between two treatment groups were analyzed using a two-tailed unpaired Student’s t-test; for comparisons among multiple groups, a one-way ANOVA with Tukey’s multiple comparisons test was applied.
p21, also known as CDK-interacting protein 1 (CIP1), directly interacts with CDKs, particularly CDK2, to halt cell cycle progression in response to DNA damage36,37,38. p27, also known as kinase inhibitory protein 1 (KIP1), primarily inhibits CDK2 and CDK4 complexes, controlling the transition from the G1 to the S phase39,40. CRISPR knockout (KO) of p21 in MCF7 parental and PR1.2 cells eliminated the synergistic effect of BLU-222 and palbociclib combination, shifting the response to antagonism (Fig. 3C–E). A similar shift to antagonism was observed when p27 was knocked out in MCF7 and PR1.2 cells (Fig. 3F–G). The heatmap of HSA modeling for the MCF7 parental and PR1.2 cells, and their respective p21 and p27 CRISPR KO pools and clones (n = 13 KO cell lines), reveals a marked shift in the drug response from synergism in PR1.2 to antagonism in all the p21 KO and p27 KO cells (Fig. 3H).
Consistent results were observed in T47D PR1.2 cells, where knockout of either p21 or p27 also led to antagonistic responses to the combination treatment of BLU-222 and palbociclib (Supplemental Fig. 3C–F). The disruption of this synergy upon knockout of either protein, suggests that BLU-222 relies on inducing endogenous inhibitors p21 and/or p27 to effectively suppress CDK2.
Additionally, knocking out of p21 or p27 rendered MCF7 PR1.2 cells more resistant to palbociclib while increasing their sensitivity to BLU-222 (Fig. 3I and Supplemental Fig. 3H), consistent with the results observed in T47D PR1.2 cells (Supplemental Fig. 3I, J). Collectively, these results highlight the pivotal roles of p21 and p27 as CIP/KIP proteins in coordinating the inhibition of both CDK4/6 and CDK2, which is essential for maximizing the efficacy of the combination treatment.
The direct interaction of p21 with CDK2 and p21/p27 with CDK4 mediates synergistic response to combined treatment with CDK4/6 and CDK2 inhibitors
Our study demonstrated that the combination of BLU-222 and palbociclib exhibits strong synergistic effects in both HR+/HER2− and TNBC subtypes, driven by the upregulation of p21 and p27. In palbociclib-resistant models, where p21 and/or p27 levels are downregulated, treatment with BLU-222 effectively restores their expression. To explore how the interactions of p21 and p27 with CDK2 and CDK4 contribute to these synergistic effects, we employed proximity ligation assay (PLA) (Supplemental Fig. 4A). This highly sensitive technique enables in situ visualization of endogenous protein-protein interactions, allowing us to quantify the interactions of p21 and p27 with CDK2 in both MCF7 parental and PR1.2 cells, on a per cell basis.
PLA results from MCF7 parental cells showed that the binding of p21 to CDK2 remained unchanged after palbociclib treatment. However, BLU-222 induced a slight increase in p21–CDK2 interactions. Notably, in cells treated with the combination of BLU-222 and palbociclib, a significant increase in p21–CDK2 binding was observed compared to all other treatments, including BLU-222 or palbociclib alone (Fig. 4A, B).
A Representative confocal images of proximity ligation assay (PLA) on MCF7 parental cells treated with DMSO, palbociclib, BLU-222, or the combination (0.3 µM each, 3 days) detecting proximity and binding between p21 and CDK2. Nuclei stained with DAPI (blue); PLA performed for p21 and CDK2 (red); and actin filaments stained with phalloidin (green). Scale bars: 15 μm. B Quantification of PLA signals per cell nucleus showing p21/CDK2 binding in MCF7 parental cells treated with DMSO, palbociclib, BLU-222, or the combination (0.3 µM each, 3 days). C Representative confocal images of PLA on MCF7 PR1.2 cells treated with DMSO, palbociclib, BLU-222, or the combination (0.3 µM, 3 days) detecting binding between p21 and CDK2. D Quantification of PLA signals per cell nucleus showing p21/CDK2 binding in MCF7 PR1.2 cells treated with DMSO, palbociclib, BLU-222, or the combination (0.3 µM each, 3 days). E Immunoprecipitation (IP)–Western blotting analysis of p21 and p27 interactions with CDK2 in MCF7 parental (left) and PR1.2 (right) cells treated with single agents or the combination of palbociclib and BLU-222 (0.3 µM, 3 days). D, DMSO; P, palbociclib; B, BLU-222. F IP–Western blotting analysis of p21 and p27 interactions with CDK4 in MCF7 parental (left) and PR1.2 (right) cells treated with single agents or the combination of palbociclib and BLU-222 (0.3 µM, 3 days). Blots are representative of 3 biologically independent experiments performed on separate cell preparations with similar results. The samples were derived from the same experiment, and all gels/blots were processed in parallel under identical conditions. Error bars represent SEM from n = 3 experimental replicates. Differences among groups were analyzed using a one-way ANOVA with Tukey’s multiple comparisons test.
In MCF-7 PR1.2 cells, the binding of p21 to CDK2 significantly increased with BLU-222 treatment alone, and this interaction was further enhanced significantly by the combination of BLU-222 and palbociclib, compared to either monotherapy treatment (Fig. 4C, D). These findings suggest that BLU-222 restores or enhances p21–CDK2 interactions in resistant cells, contributing to the observed synergy between CDK2 and CDK4/6 inhibition.
A similar PLA was performed to examine the binding of the p27 to CDK2. In parental cells, p27–CDK2 binding significantly increased following palbociclib treatment, while BLU-222 induced only a modest increase. The combination treatment resulted in levels of p27–CDK2 interaction similar to those seen with palbociclib alone (Supplemental Fig. 4B). However, in PR1.2 cells, treatment with either palbociclib, BLU-222, or the combination led to a pronounced increase in p27–CDK2 interactions compared to vehicle-treated cells, suggesting a broader role of p27 in mediating the response to CDK2 or CDK4/6 inhibition under resistant conditions (Supplemental Fig. 4C). Consistently, BLU-222 monotherapy significantly enhanced p21-CDK2 interactions in the TNBC cell lines MDA-MB-231 and BT-20, which were significantly enhanced by the combination with palbociclib in both MDA-MB-231 PR and BT-20 PR cells (Supplemental Fig. 4D-G).
To further explore the potential differential roles of p21 and p27 in mediating the response to combination treatment with BLU-222 and palbociclib, we performed immunoprecipitation (IP) and Western blot experiments. Unlike PLA, which allows for in situ visualization of protein interactions on a per cell basis, IP-Western blotting provides a biochemical approach to physically isolate and analyze protein complexes from cell lysates in bulk.
The results from the IP/Western blot analysis were consistent with the PLA findings, showing that p21 binding to CDK2 increased following BLU-222 as a single agent or following combination treatment in both parental and palbociclib-resistant cells (Fig. 4E). However, p27 binding to CDK2 appeared more ubiquitous, showing strong interactions not only in the BLU-222 and combination groups but also in the palbociclib treatment group, consistent with the previous PLA data (Fig. 4E and Supplemental Fig. 4H, I). We also examined p21 and p27 associations with CDK4 to assess whether these interactions contribute to the observed synergistic effects. Both p21 and p27 were found to bind to CDK4 when treated with BLU-222 alone or in combination with palbociclib (Fig. 4F and Supplemental Fig. 4H, I). This suggests that the interactions of p21 and p27 with CDK4 play a critical role in mediating the synergistic effects of dual treatment by enhancing CDK4/6 inhibition. Our studies in TNBC also suggest a broader role of p27 in MDA-MB-231 PR and BT-20 PR TNBC cells, where treatment with palbociclib, BLU-222, or their combination markedly increased p27-CDK2 binding. In contrast, this effect was absent in the corresponding parental cells, in which p27–CDK2 interactions remained unchanged following either monotherapy or combination therapy (Supplemental Fig. 4H, I).
Finally, to assess whether the p21/p27-mediated dual inhibition observed in resistant tumor cells is a general feature of cell cycle control, we tested the non-tumorigenic mammary epithelial line MCF-10A. Both palbociclib and BLU-222 reduced growth, but their effects were strictly additive, with no evidence of synergy by HSA analysis (Supplemental Fig. 4J). Mechanistically, PLA and co-immunoprecipitation assays showed no changes in p21 or p27 binding to CDK2 or CDK4 following treatment with either agent or the combination (Supplemental Fig. 4K, L). This implies that in normal epithelial cells, p21 and/or p27 may not be as dynamically regulated in response to CDK inhibitor treatment as observed in cancer cells, reflecting distinct mechanisms of cell cycle control between normal and transformed contexts.
In summary, the data demonstrate that the binding of p21 to CDK2 and the binding of both p21 and p27 to CDK4 are key components in the mechanism of synergy between BLU-222 and palbociclib in luminal and basal tumor cells. In both parental and resistant tumor cells lines, regardless of subtype, the restoration or enhancement of p21 and p27 interactions with CDK2 and CDK4 underscores their dual yet diverse roles in regulating CDK2 and CDK4/6 activity. These findings suggest that, beyond their canonical role as CDK inhibitors, p21 and p27 perform distinct, context-specific functions that contribute to the efficacy of dual inhibition. This mechanistic insight highlights their critical importance in overcoming resistance to CDK4/6i when combined with a CDK2 inhibitor in tumor cells.
Combination treatment with BLU-222 and CDK4/6 inhibitors is synergistic in HR+/HER2− PDX models that progressed on palbociclib
To evaluate the in vivo efficacy of BLU-222 in combination with CDK4/6i, we used four previously established ER+/HER2− PDX models (PR1–PR4), derived from patients whose tumors had progressed on palbociclib and endocrine therapy28,41. PR1 and PR2 represent acquired resistance, with progression after 16 and 6 months of treatment, respectively, whereas PR3 and PR4 represent intrinsic resistance, with progression within 3 months (Supplemental Table S2). Mechanistically, RNA-seq profiling revealed enrichment of IL-6/STAT3, EMT, and inflammatory signaling in the late progressors (PR1 and PR2), while the early progressors (PR3 and PR4) exhibited enrichment of cell cycle and DNA damage–associated programs, including E2F, G2/M checkpoint, and AURKA28,41. Genomic profiling further showed that all PR models harbor TP53 alterations, with additional RB1 loss in PR1, PR2, and PR4, and an ESR1 mutation in PR1 (Supplemental Tables S2). Together, these models capture clinically relevant mechanisms of acquired and intrinsic CDK4/6 inhibitor resistance and provide a robust platform to test whether CDK2 inhibition can overcome diverse resistance contexts.
Immunocompromised mice harboring these PDX models were randomized into 5-6 treatment groups: (i) vehicle, (ii) palbociclib (50 mg/kg once daily), (iii) BLU-222 (60 mg/kg twice daily), (iv) Palbo + fulvestrant (in PR-2) (2.5 mg/kg once weekly) (v) BLU-222 + palbociclib, and (v) BLU-222 + palbociclib + fulvestrant. In the PR1 and PR2 models, which represent acquired resistance, neither single-agent palbociclib or BLU-222, produced significant tumor growth inhibition. Notably, in PR2, even the standard-of-care (SOC) regimen of palbociclib plus fulvestrant failed to suppress tumor growth (Supplemental Fig. 5A). However, combination treatment with BLU-222 and palbociclib, as well as the triple combination with fulvestrant, led to significant (P < 0.0001) antitumor activity and durable tumor regression (Fig. 5A and Supplemental Fig. 5A). Waterfall plots further demonstrate the strong inhibitory effects of the combination treatments compared to single agents in individual mice (Fig. 5B and Supplemental Fig. 5B). Moreover, BLU-222 plus palbociclib significantly prolonged survival, with median survival not reached in the double and triple combination groups, while single-agent treatment resulted in median survival times of 17–22 days for PR1 (Fig. 5C) and 25.5–33 days for PR2 (Supplemental Fig. 5C).
A–I Tumor growth curves, waterfall plots showing tumor volume changes, and Kaplan-Meier survival curves of PDX palbociclib-resistant HR+/HER2− breast cancer models. A–C PR1 (vehicle, n = 5; BLU-222, n = 5; palbociclib, n = 4; BLU-222 + palbociclib, n = 4; BLU-222 + palbociclib + fulvestrant, n = 5). D–F PR3 (vehicle, n = 8; BLU-222, n = 3; palbociclib, n = 4; BLU-222 + palbociclib, n = 4; BLU-222 + palbociclib + fulvestrant, n = 4), and G–I PR4 (vehicle, n = 4; BLU-222, n = 4; palbociclib, n = 6; BLU-222 + palbociclib, n = 6; BLU-222 + palbociclib + fulvestrant, n = 4), treated with vehicle, palbociclib, BLU-222, BLU-222 plus palbociclib, or triple combination with fulvestrant. Tumor pieces from each model were xenograft implanted into the fat pad of NCR-NU-F sp/sp 10-week-old female mice. Drug treatment was initiated when tumors reached approximately 100 mm³. Palbociclib: 50 mg/kg, orally, once daily; BLU-222: 60 mg/kg, orally, twice daily; fulvestrant: 2.5 mg/kg, subcutaneously, once weekly. Error bars represent the SEM from n = biological replicates. J–W Immunohistochemistry (IHC) quantification of tumors collected from treated PDX mice, with percentage of cells with positive staining for Ki67, γH2AX, p21, and p27 in models. J–M PR1 (vehicle, n = 5; BLU-222, n = 5; palbociclib, n = 4; BLU-222 + palbociclib, n = 4; BLU-222 + palbociclib + fulvestrant, n = 5), N–Q PR3, (vehicle, n = 8; BLU-222, n = 3; palbociclib, n = 4; BLU-222 + palbociclib, n = 4; BLU-222 + palbociclib + fulvestrant, n = 4), and R–U PR4 (vehicle, n = 4; BLU-222, n = 4; palbociclib, n = 6; BLU-222 + palbociclib, n = 6; BLU-222 + palbociclib + fulvestrant, n = 4). V Representative IHC images of p21 and p27 expression in the PR1 model treated with vehicle, single agents, or a combination of palbociclib and BLU-222 +/- fulvestrant. Scale bar = 50 μm. Tumor growth curves were analyzed using a two-way ANOVA to compare treatment groups across time. Survival analyses were performed using the Kaplan–Meier method, and statistical significance among treatment groups was assessed with the log-rank (Mantel–Cox) test. Median survival times are reported in the figure. IHC quantification was analyzed using one-way ANOVA with Tukey’s multiple comparisons test. Data represent the mean ± SEM of the percentage of positively stained cells from the above-mentioned study animals per group (n), with each sample quantified from 2–3 randomly selected high-power fields.
Similarly, in the PR3 and PR4 models (both intrinsic resistance), palbociclib monotherapy did not result in significant tumor suppression (Fig. 5D–I). BLU-222 alone showed moderate efficacy, but the combination of BLU-222 and palbociclib, as well as the triple combination with fulvestrant, led to significant antitumor activity (Fig. 5D, G) and durable tumor regression (Fig. 5E, H). The combination treatments also significantly prolonged survival, with median survival not reached in the combination arms, while single-agent treatment resulted in median survival times of 15–39.5 days for PR3 and (Fig. 5F) and 26–28 days for PR4 (Fig. 5I). Additionally, the addition of fulvestrant in the combination of palbociclib and BLU-222 did not significantly enhance survival in any of the PR1–4 models (compare the green and yellow lines in Fig. 5A–I and Supplemental Fig. 5A–C).
Immunohistochemistry (IHC) analysis across PR1–PR4 tumors further supported the efficacy of combination treatments (Fig. 5J–V, Supplemental Fig. 5D–K). Ki67, a marker of cell proliferation, was significantly downregulated, while γH2AX, a marker of DNA damage, was upregulated in the combination arms compared to single agent-treated and vehicle-treated tumors Additionally, the expression levels of p21 and p27, key cell cycle regulators, were significantly elevated in the combination-treated tumors, contributing to the resensitization of palbociclib-resistant tumors to CDK4/6 inhibition.
Pharmacokinetic analysis of plasma concentration–time profiles, whether the compounds were administered individually or in combination, revealed substantial overlap (Supplemental Fig. 6A-E). This suggests there was no significant drug-drug interaction between palbociclib and BLU-222. Additionally, monitoring of mouse body weights (Supplemental Fig. 7A-J) and evaluation of blood cell populations (Supplemental Fig. 8A-D) demonstrated that BLU-222, whether administered alone or in combination twice daily in different models, was well tolerated.
We also assessed the combination of BLU-222 with ribociclib, another FDA-approved CDK4/6i, in the same PDX models. Similar to the palbociclib results, single-agent BLU-222 or ribociclib did not show significant tumor inhibition. The combination of BLU-222 with ribociclib, as well as the triple combination with fulvestrant, demonstrated strong synergistic tumor suppression and prolonged survival, where median survival was not reached (Fig. 6A–I). Compared to the SOC doublet of CDK4/6i plus fulvestrant, which produced only tumor growth delay without durable regression, addition of the CDK2 inhibitor BLU-222 significantly enhanced therapeutic efficacy across multiple resistant HR+/HER2− PDX models. The triplet regimen (BLU-222 + CDK4/6i + fulvestrant) induced robust, often complete or near-complete regressions, extended survival, and delayed recurrence (Fig. 6A–I; Supplemental Fig. 5A–C). These results highlight the potential of the triplet combination to overcome resistance to standard therapy and provide strong preclinical rationale for its clinical evaluation in HR+/HER2− breast cancer. IHC confirmed reduced Ki67 levels and increased γH2AX, p21, and p27 expression in combination-treated tumors (Fig. 6J–V, Supplemental Fig. 9A–C).
A–I Tumor growth curves and waterfall plots showing tumor volume changes and Kaplan-Meier survival curves of PDX PR HR+/HER2− breast cancer models. A–C PR1, treated with vehicle (n = 5), ribociclib (n = 6), fulvestrant (n = 4), ribociclib + fulvestrant (n = 5), palbociclib + fulvestrant (n = 6), BLU-222 + ribociclib (n = 5), BLU-222 + ribociclib + fulvestrant (n = 5). D–F PR3, treated with vehicle (n = 5), ribociclib (n = 5), fulvestrant (n = 5), palbociclib + fulvestrant (n = 5), BLU-222 + ribociclib + fulvestrant (n = 4), and G–I PR4, treated with vehicle (n = 4), ribociclib (n = 5), fulvestrant (n = 4), palbociclib + fulvestrant (n = 5), BLU-222 + ribociclib (n = 5), BLU-222 + ribociclib + fulvestrant (n = 5). Ribociclib: 50 mg/kg, orally, once daily; Palbociclib: 50 mg/kg, orally, once daily; BLU-222: 60 mg/kg, orally, twice daily; Fulvestrant: 2.5 mg/kg, subcutaneously, once weekly. Error bars represent the SEM from n = biological replicates. J–U IHC quantification of tumors collected from treated PDX mice, with percentage of cells with positive staining for Ki67, γH2AX, p21, and p27 in models. J-M PR1 (vehicle, n = 5; ribociclib, n = 6; fulvestrant, n = 4; ribociclib + fulvestrant, n = 5; palbociclib + fulvestrant, n = 6); BLU-222 + ribociclib, n = 5; BLU-222 + ribociclib + fulvestrant, n = 5). N-Q PR3 (vehicle, n = 5; ribociclib, n = 5; fulvestrant, n = 5; palbociclib + fulvestrant, n = 5; BLU-222 + ribociclib + fulvestrant, n = 4), and R-U PR4 (vehicle, n = 4; ribociclib, n = 5; fulvestrant, n = 4; palbociclib + fulvestrant, n = 5; BLU-222 + ribociclib, n = 5; BLU-222 + ribociclib + fulvestrant, n = 5). V Representative IHC images of p21 and p27 expression in the PR1 model treated with vehicle, single agents, or a combination of ribociclib and BLU-222 +/- fulvestrant. Scale bar = 50 μm. Tumor growth curves were analyzed using a two-way ANOVA test to compare treatment groups across time. Survival analyses were performed using the Kaplan–Meier method, and statistical significance among treatment groups was assessed with the log-rank (Mantel–Cox) test. Median survival times are reported in the figure. IHC quantification was analyzed using one-way ANOVA with Tukey’s multiple comparisons test. Data represent the mean ± SEM of the percentage of positively stained cells from the above-mentioned study animals per group (n), with each sample quantified from 2–3 randomly selected high-power fields.
Overall, the results consistently demonstrated strong synergistic effects of combining BLU-222 with either palbociclib or ribociclib in CDK4/6i-resistant models. In the PR1–4 models, the combination treatments significantly inhibited cell proliferation, increased DNA damage as indicated by γH2AX levels, and robust expression of p21 and p27, underscoring their role as key mediators of the response. These effects were observed with both CDK4/6 inhibitors, highlighting the broad potential of BLU-222 to overcome both acquired and intrinsic resistance to CDK4/6 inhibition in HR+/HER2− breast cancer. Importantly, BLU-222 retained efficacy across all PR models despite diverse genomic alterations, including RB1 loss, ESR1 mutations, and PIK3CA mutations, demonstrating that its therapeutic benefit is not limited by these resistance-associated lesions (Supplemental Table S2). Notably, strong p21 induction was detected even in the TP53-deficient or mutated PR1-4 models, indicating that CDK2 inhibition activates p21 expression through p53-independent mechanisms. Together, these findings establish that PR1–4 not only recapitulate clinically relevant resistance mechanisms but also provide a robust platform showing that dual CDK2/CDK4 inhibition can overcome diverse contexts of resistance.
Combination treatment with BLU-222 and palbociclib is synergistic in TNBC models mirroring the results observed in HR+/HER2− models
In addition to the HR+/HER2− breast cancer subtype, our previous in vitro data also showed the synergistic effects of BLU-222 and palbociclib in the TNBC subtype. To validate these findings in vivo, we treated 3 TNBC PDX models, BCX070, BCX017 and XC517201342,43 (Supplemental Table S2), with single-agent BLU-222, palbociclib, or the combination. All three models were generated from patients with TNBC who had received chemotherapy. The XC5172013 model, which lacks CCNE1 amplification or cyclin E1 overexpression, did not show tumor growth inhibition with either monotherapy, but the combination treatment consistently inhibited tumor growth, maintaining this effect even after treatment withdrawal and reinitiation (Fig. 7A). The waterfall plot demonstrates a strong synergistic inhibition of tumor growth with the combination treatment (Fig. 7A), and survival analysis revealed a significant increase in overall survival with median survival not reached in the combination arm, while single-agent treatment resulted in median survival times of 24.5–39 days in the single-agent and vehicle groups (Fig. 7A). This highlights the ability of the combination to overcome resistance in TNBC models lacking CCNE1 amplification.
A–D Tumor growth curves and waterfall plots showing tumor volume changes and Kaplan-Meier survival curves of PDX PR TNBC breast cancer models. A XC5172013 (vehicle, n = 7; palbociclib, n = 2; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 8; BLU-222 + palbociclib, n = 5), B BCX070 (vehicle, n = 5; palbociclib, n = 3; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 5; BLU-222 + palbociclib, n = 4), C BCX017 (vehicle, n = 6; palbociclib, n = 5; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 5; BLU-222 + palbociclib, n = 5), and D MMTV-T1p, a transgenic model overexpressing LMW-E (vehicle, n = 5; palbociclib, n = 5; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 5; BLU-222 + palbociclib, n = 4). Palbociclib: 50 mg/kg, orally, once daily; BLU-222: low-dose 50 mg/kg and high-dose 100 mg/kg, orally, twice daily. Error bars represent the SEM from n = biological replicates. E-T IHC quantification of tumors collected from treated mice, with percentage of cells with positive staining for Ki67, γH2AX, p21, and p27 in models. E, I, M, Q XC5172013 (vehicle, n = 7; palbociclib, n = 2; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 8; BLU-222 + palbociclib, n = 5), (F, J, N, R) BCX070 (vehicle, n = 5; palbociclib, n = 3; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 5; BLU-222 + palbociclib, n = 4), (G, K, O, S) BCX017 (vehicle, n = 6; palbociclib, n = 5; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 5; BLU-222 + palbociclib, n = 5), and (H, L, P, T) transgenic T1p (vehicle, n = 5; palbociclib, n = 5; BLU-222 50 mg/kg, n = 5; BLU-222 100 mg/kg, n = 5; BLU-222 + palbociclib, n = 4). Each tumor piece was quantified by the average percentage of positivity of cells in 2-3 different areas. Tumor growth curves were analyzed using a two-way ANOVA with Tukey’s multiple comparisons test to compare treatment groups across time. Survival analyses were performed using the Kaplan–Meier method, and statistical significance among treatment groups was assessed with the log-rank (Mantel–Cox) test. Median survival times are reported in the figure. IHC quantification was analyzed using one-way ANOVA with Tukey’s multiple comparisons test. Data represent the mean ± SEM of the percentage of positively stained cells from the above-mentioned study animals per group (n), with each sample quantified from 2–3 randomly selected high-power fields.
In contrast, the BCX070 and BCX017 models, derived from TNBC patients with CCNE1 amplification (Supplemental Table S2), responded moderately to 50 mg/kg BID BLU-222 and exhibited complete tumor growth inhibition with 100 mg/kg BID BLU-222 (Fig. 7B, C). These findings suggest that CCNE1 amplification could be a biomarker for response to BLU-222 treatment. The waterfall plots for the BCX070 (Fig. 7B) and BCX017 (Fig. 7C) models confirm the strong inhibitory effects of 100 mg/kg of BLU-222 and a combination treatment (50 mg/kg BLU-222 + palbociclib). Survival analysis further demonstrated a significant extension in survival for both the 100 mg/kg BLU-222 and combination treatment groups. In these groups, the median survival was not reached, contrasting with the vehicle and single-agent treatment groups, where median survival times ranged from 24 to 42 days in both CCNE1 high models (Fig. 7B, C). This suggests that TNBC tumors with cyclin E1 overexpression are particularly sensitive to BLU-222.
We further examined the role of LMW-E in driving sensitivity to BLU-222 using the MMTV-T1p mouse model44,45, which overexpresses LMW-E with heterozygous p53 to mimic the TNBC subtype. As previously shown in vitro, LMW-E–expressing cells were more sensitive to BLU-222 (Fig. 1H, Supplemental Fig. 1L, N, O). In vivo, palbociclib or 50 mg/kg BID BLU-222 monotherapy only moderately inhibited T1p tumor growth. However, 100 mg/kg BID BLU-222 or the combination treatment (50 mg/kg BLU-222 + palbociclib) produced a strong synergistic effect, as shown in the tumor growth curve, waterfall plot, and survival analysis, where the median survival was not reached (Fig. 7D). These findings provide in vivo validation of the heightened sensitivity of LMW-E–expressing tumors to BLU-222, especially in combination with palbociclib.
IHC analysis across the TNBC models provided further insights into the mechanisms underlying these effects. In all 4 models (XC5172013, BCX070, BCX017 and T1p), combination treatment significantly reduced cell proliferation, as indicated by decreased Ki67 levels (Fig. 7E–H and Supplemental Fig. 10A–D). BLU-222, both as a monotherapy and in the combination arm, induced DNA damage, evidenced by increased γH2AX levels (Fig. 7I–L, Supplemental Fig. 10A–D). Importantly, BLU-222 also upregulated p21 and p27 in the combination-treated tumors across all models (Fig. 7M–T, Supplemental Fig. 10E–H), suggesting that the observed anti-tumor activity may be mediated through cell cycle regulation.
To evaluate the translational relevance of BLU-222 in TNBC, we tested its efficacy in combination with both SOC chemotherapies and CDK4/6 inhibition across multiple preclinical models (Supplemental Fig. 11A–F). In the TNBC PDX model BCX070 and the TNBC CDX model HCC1569, both harboring CCNE1 amplification, BLU-222 was combined with carboplatin (BCX070) or paclitaxel (HCC1569). In both settings, the combination achieved equal or slightly more durable responses than either monotherapy (Supplemental Fig. 11A–D). These findings parallel results from a recently published study demonstrating that BLU-222 synergizes with chemotherapy in CCNE1-abarrent ovarian cancer models, further reinforcing the dependence of this effect on CCNE1 amplification and its translational significance.
In contrast, in the XC5172013 TNBC PDX model lacking CCNE1 amplification, BLU-222 combined with carboplatin produced only modest improvements compared to monotherapies (Supplemental Fig. 11E, F). By comparison, BLU-222 combined with palbociclib resulted in complete tumor eradication, whereas neither palbociclib nor BLU-222 monotherapy had therapeutic impact. Importantly, none of the treatment regimens, including the BLU-222 plus carboplatin combination, caused measurable body weight loss, indicating that the therapies were well tolerated (Supplemental Fig. 7I, J). These results highlight a critical distinction: while BLU-222 synergizes with chemotherapy in CCNE1-amplified tumors, its most profound therapeutic effect in non-amplified TNBC emerges from dual CDK2 and CDK4/6 inhibition, which can overcome resistance where both SOC chemotherapy and CDK4/6 inhibitors fail.
In summary, the combination of BLU-222 and palbociclib produced robust synergistic effects in TNBC models, characterized by reduced cell proliferation, increased DNA damage, and upregulation of key cell cycle inhibitors p21 and p27. These findings are consistent with the in vitro data and further support the therapeutic potential of combining BLU-222 with CDK4/6i in TNBC, particularly in tumors with CCNE1 amplification or LMW-E expression.
Senescence and interferon pathways were significantly upregulated after combination treatment with BLU-222 plus CDK4/6 inhibitors
To build on the observation of strong synergistic effects of BLU-222 combined with CDK4/6i in both HR+/HER2− and TNBC models with intrinsic and acquired resistance, as well as the upregulation of p21 and p27, we sought to further explore the mechanisms involved. To achieve this, we performed RNA sequencing analysis on 2 HR+/HER2− PDX models (PR1, representing acquired resistance, and PR4, representing intrinsic resistance) and 3 TNBC PDX models (BCX070, BCX 017 which have CCNE1 amplification, and XC5172013, without CCNE1 amplification). The differential expression and functional analyses were focused on comparing combination-treated tumors to vehicle-treated controls to identify the pathways contributing to the observed synergy.
Gene set enrichment analysis (GSEA) revealed that the most significantly upregulated pathways in the combination-treated group included interferon-γ and α responses, TNF-α signaling, and inflammatory responses. On the other hand, the MYC target gene set was downregulated in the XC5172013 model (Fig. 8A), suggesting that the combination treatment influences immune pathways while suppressing oncogenic signals. The Venn diagram further illustrates the overlapping or unique upregulated genes associated with pathways across models, with the interferon-γ pathway consistently activated in all groups (Fig. 8B).
A Dot plot showing normalized enrichment scores (NES) of Hallmark gene sets from gene set enrichment analysis (GSEA) for the combination BLU-222 and palbociclib treatment arm across HR+/HER2− (PR1, n = 3 and PR4, n = 4) and TNBC (BCX070, n = 5 and XC5172013, n = 4) PDX models. B Venn diagram showing common and unique upregulated genes associated with pathways upon combination treatment with BLU-222 and palbociclib across 4 PDX models. C–F Heatmaps showing the standardized single-sample GSEA (ssGSEA) scores for each sample of interferon-α and -γ pathways, TNF-α, and inflammation response pathways across 4 PDX models. Bar plots on top of each heatmap indicate the average value of the standardized ssGSEA scores for each sample. G,H Enrichment plots of the interferon-γ gene set from GSEA performed on XC5172013 and PR1 tumors for the comparison of the treatment combination of BLU-222 and palbociclib versus vehicle treatment. I,J Heatmaps of fold-change values of genes associated with interferon-α and -γ pathways in treated tumors compared to vehicle-treated tumors across all PDX models. K–N Heatmaps showing the standardized ssGSEA scores for each sample of gene sets related to senescence across 4 PDX models. O,P Representative images of Sudan Black B (SBB)-stained lipofuscin (far-red, Cy5), indicative of therapy-induced senescence, in O BCX070 and P PR4 tumors from vehicle-treated and combination-treated (BLU-222+palbociclib) mice. Scale bars, 100 μm. Q Quantification of SBB-positive lipofuscin area (%) in BCX070 (vehicle, n = 5; palbociclib, n = 3; BLU-222 100 mg/kg, n = 5; BLU-222 + palbociclib, n = 4), XC5172013 (vehicle, n = 7; palbociclib, n = 2; BLU-222 100 mg/kg, n = 8; BLU-222 + palbociclib, n = 5), PR1 (vehicle, n = 5; BLU-222, n = 5; palbociclib, n = 4; BLU-222 + palbociclib + fulvestrant, n = 5), and PR4 tumors (vehicle, n = 4; BLU-222, n = 4; palbociclib, n = 6; BLU-222 + palbociclib + fulvestrant, n = 4) across indicated treatment groups. Data represent the mean ± SEM of the percentage of positively stained cells from the above-mentioned study animals per group (n), with each sample quantified from 2–3 randomly selected high-power fields. Statistical significance was determined by one-way ANOVA with Tukey’s multiple comparisons test.
To gain further insights into the molecular mechanisms, we performed single-sample GSEA (ssGSEA) in addition to traditional GSEA. While GSEA identifies pathway enrichment between groups of samples (treated vs. vehicle), ssGSEA calculates enrichment scores for each predefined gene set on a per-sample basis. By using ssGSEA, we were able to explore individual variability in pathway activation, particularly in hallmark gene sets related to interferon-α, interferon-γ, TNF-α signaling, and inflammatory response across HR+ and TNBC models. In the TNBC BCX017 and XC5172013 models, both 100 mg/kg BLU-222 and the combination treatment significantly increased the expression of these immune response pathways (Fig. 8C, D), with similar findings observed in the HR+ PR1 and PR4 models (Fig. 8E, F). The activation of these immune-related pathways across HR+ and TNBC models supports the hypothesis that the combination treatment induces a broad immune response, contributing to the therapeutic effect. In addition to ssGSEA, GSEA enrichment plots for the interferon-γ pathway further confirmed its significant activation in the XC5172013 and PR4 models (Fig. 8G, H), highlighting the consistent role of this immune pathway across models. These data suggest that interferon signaling may be a key mechanism by which BLU-222 enhances the efficacy of CDK4/6i.
Next, to examine specific genes associated with interferon responses, we analyzed interferon-related gene sets across all models. A heatmap revealed consistent upregulation of genes such as SAMD9, CD47, TRIM26, and several interferon-α induced genes (IFI44L, IFI35, IFI27) across all models (Fig. 8I). Similarly, interferon-γ pathway genes, including OAS2, HLA-A, and PSMB8, were upregulated in the combination-treated group (Fig. 8J). This consistent upregulation across models underscores the involvement of interferon pathways in mediating the anti-tumor response. Volcano plots from individual models, such as PR1, confirmed the significant upregulation or induction of these interferon-related genes following combination treatment (Supplemental Fig. 12A).
To investigate whether interferon signaling contributes to the cytotoxic effects of CDK2 and CDK4/6 co-inhibition, we treated HR+ and TNBC parental and PR cells with BLU-222 and palbociclib in the presence or absence of JAK inhibitors (Upadacitinib or Abrocitinib), which block downstream signaling from both IFNAR (type I) and IFNGR (type II) interferon receptors. In resistant lines, including HR+ MCF7 and T47D cells and TNBC MDA-MB-231 cells, JAK inhibition with Upadacitinib markedly rescued cell viability (Supplemental Fig. 12B–D), indicating that JAK/STAT signaling is required for the full cytotoxic effect of combined CDK2 and CDK4/6 inhibition. Similar results were obtained with the JAK inhibitor Abrocitinib (Supplemental Fig. 12E–G), further confirming that interferon signaling is necessary for the observed therapeutic activity. Collectively, these findings support a model in which activation of interferon signaling via JAK/STAT contributes to the anti-tumor effects of CDK2/4/6 co-inhibition specifically in resistant cells and provide functional validation of the interferon pathway enrichment identified in our RNA-seq analyses.
Interestingly, in the CCNE1-amplified BCX070 and BCX017 models, which displayed unique sensitivity to BLU-222 monotherapy, we observed a marked induction of interferon signaling following CDK2 inhibition (Fig. 8A-J). To further evaluate the immunomodulatory consequences of this treatment, we assessed tumor-infiltrating lymphocytes (TILs) in the immune-competent T1p model, which overexpresses low-molecular-weight cyclin E. Histological analysis and quantitative assessment of TILs revealed that BLU-222, administered either as a single agent or in combination with palbociclib, significantly increased TIL density compared to vehicle or palbociclib alone (p < 0.0001; Supplemental Fig. 12H). These findings suggest that CDK2 inhibition, in addition to disrupting tumor cell cycle progression, promotes remodeling of the tumor immune microenvironment. Importantly, this effect appears particularly pronounced in cyclin E-driven tumors, highlighting a potential link between CDK2 inhibition, interferon pathway activation, and enhanced antitumor immunity. Together, these results raise the possibility that CDK2 inhibition could potentiate immune responsiveness and improve outcomes when integrated with immunotherapeutic strategies.
Given that p21 and p27 are known markers of senescence, we next investigated whether the combination treatment also induces senescence-associated gene sets across all models46,47,48. Using ssGSEA, we found that combination treatments upregulated senescence-related gene sets in both TNBC (BCX070, XC5172013) and HR+ (PR1 and PR4) models (Fig. 8K–N). These data suggest that the combination treatment promotes tumor cell senescence, contributing to tumor growth inhibition.
To further investigate the cellular consequences of combined CDK2 and CDK4/6 inhibition, we analyzed senescence induction using Sudan Black B (SBB) staining for lipofuscin accumulation, a hallmark of senescent cells49. SBB staining revealed a significant increase in lipofuscin-positive cells in tumors treated with the combination of BLU-222 and CDK4/6 inhibitors across multiple models, including two HR+ (PR1 and PR4) and two TNBC (BCX070 and XC5172013) PDX models (Fig. 8O–Q, Supplemental Fig. 13A, B). These results indicate robust induction of cellular senescence, suggesting that combination of BLU-222 and CDK4/6 inhibition contributes to therapeutic efficacy not only through cell cycle suppression but also by promoting a senescent phenotype in tumor cells.
To functionally validate transcriptomic findings, we assessed senescence induction following single agents and combination treatment with BLU-222 and palbociclib in vitro. β-galactosidase staining revealed robust induction of cellular senescence following BLU-222 and CDK4/6 inhibitor combination treatment across multiple models, including Palbociclib-resistant MCF7, T47D, and MDA-MB-231 derivatives (Supplemental Fig. 13C–H). These results further support senescence as a key mechanism contributing to the therapeutic efficacy of CDK2 and CDK4/6 co-inhibition. These results, observed consistently across HR+ and TNBC in both in vitro and in vivo models, highlight senescence as a conserved and biologically relevant mechanism that contributes to the therapeutic efficacy of CDK2 and CDK4/6 co-inhibition in resistant tumors.
In addition to the observed senescence, we examined the DREAM pathway, which has been shown to mediate p21-induced senescence and tumor suppression50. ssGSEA analysis showed an upregulation of DREAM pathway–associated gene sets in either single-agent 100 mg/kg BLU-222 (XC5172013 and PR4) or combination treatments (BCX070 and PR1), suggesting that p21-driven senescence may be a crucial mechanism of BLU-222’s anti-tumor effects (Supplemental Fig. 14A–D).
Further investigation of other hallmark gene sets related to p21 and p27 (apoptosis, PI3K/AKT/mTOR, hypoxia, E2F targets) revealed model-specific effects. In the TNBC BCX070 model, all 4 gene sets were upregulated in the combination group, whereas in the TNBC XC5172013 model, only apoptosis was upregulated, while the E2F targets and PI3K/AKT/mTOR pathway were downregulated with combination treatment (Supplemental Fig. 14E, F). In HR+ models, apoptosis and E2F targets were consistently upregulated, while the hypoxia and PI3K/AKT/mTOR pathways exhibited mixed responses based on resistance type (Supplemental Fig. 14G, H). These findings suggest that BLU-222, in combination with CDK4/6i, induces cell death and modulates cellular stress responses differently depending on the resistance mechanisms in place.
To functionally validate apoptotic induction, we assessed caspase-3 activation following single-agent and combination treatment with BLU-222 and CDK4/6i across five PDX models. IHC staining revealed modest caspase-3 positivity with BLU-222 monotherapy, minimal staining with CDK4/6i, and robust induction of apoptosis with the combination in PR1, PR2, PR4, XC5172013, and BCX070 (Supplemental Fig. 14N–R). While these findings establish apoptosis as an important contributor to therapeutic efficacy, they complement rather than replace the broader biological outcomes observed. In particular, earlier analyses demonstrated that CDK2/CDK4/6 co-inhibition triggers replication stress marked by γH2AX and drives durable growth arrest through senescence, as evidenced by β-galactosidase and Sudan Black B staining (Fig. 8O, P). Together, these data suggest that apoptosis occurs in selected contexts, but that DNA damage–associated senescence represents the predominant and durable therapeutic mechanism, with apoptosis acting in parallel to enhance overall tumor suppression.
Lastly, the Venn diagram identified common and unique downregulated genes, with the common genes associated with apoptosis and hypoxia pathways across all models (Supplemental Fig. 14I). In particular, hypoxia gene sets showed significant downregulation in both the XC5172013 and PR1 models, as confirmed by negative enrichment scores in preranked GSEA plots (Supplemental Fig. 14J, K). Volcano plots highlighted the downregulation of specific hypoxia-related genes in the combination-treated groups in both XC5172013 and PR1 models (Supplemental Fig. 14L–M). These findings indicate that the combination treatment may reduce the hypoxic environment within tumors, further enhancing the therapeutic effects by disrupting tumor cell survival mechanisms. Overall, these RNA sequencing results demonstrate that the combination of BLU-222 and CDK4/6i consistently upregulates p21- and p27-associated senescence and interferon response pathways across HR+ and TNBC models, while hypoxia pathways are downregulated. These findings underscore the critical role of p21 and p27 in mediating the synergistic effects of the combination treatment and provide insight into the molecular mechanisms driving tumor regression in both acquired and intrinsic resistance settings.
Discussion
This study provides robust in vitro and in vivo evidence that combining the selective CDK2 inhibitor BLU-222 with CDK4/6i effectively overcomes resistance in breast cancer, particularly in HR+/HER2− and TNBC subtypes. Both HR+ and TNBC models demonstrated strong synergistic effects when BLU-222 was combined with CDK4/6 inhibitors such as palbociclib and ribociclib, underscoring the broad potential of this combination strategy. This is particularly relevant for TNBC, which lacks targeted therapies and remains aggressive, highlighting the ability of BLU-222 to target resistant phenotypes across diverse disease settings.
A central finding of this work is that BLU-222 or combination treatment induces de novo expression of the endogenous CDK inhibitors p21 and p27, which act as obligate mediators of cell cycle arrest and senescence particularly in CDK4/6i-resistant cells where p21 and p27 are often downregulated51,52. Functional CRISPR knockout confirmed that loss of either protein abolished therapeutic synergy, establishing them as active mediators rather than passive biomarkers. Mechanistically, BLU-222 enhanced binding of p21 to both CDK2 and CDK4, and p27 to CDK4, imposing dual inhibitory control over CDK complexes. This reprogramming contrasts with structural studies showing that p27 can assemble CDK4/6–cyclin D complexes and antagonize palbociclib inhibition53 emphasizing that BLU-222 fundamentally shifts CIP/KIP function from potential activators to obligate inhibitors in resistant tumor contexts. By restoring this inhibitory control, BLU-222 not only enhances antitumor efficacy but also introduces a mechanistic framework for overcoming drug resistance.
This mechanism is distinct from prior redistribution models. Previous studies have shown that CDK4/6 inhibitors can decrease p21 in resistant cells54,55,56 and dissociate it from CDK4, leading to redistribution toward CDK257. In contrast, our data show that BLU-222 leads to de novo induction of p21 and p27, whose dual engagement with CDK2 and CDK4 is essential for therapeutic synergy. These effects were consistent across HR+ and TNBC models, including both intrinsic and acquired resistance, highlighting their broad relevance. Notably, p21 and p27 induction coincided with senescence and interferon pathway activation, suggesting multilayered tumor-suppressive responses.
Transcriptomic analyses provided further mechanistic clarity. Downregulation of the G2/M checkpoint and E2F targets was most evident in XC5172013, reflecting treatment timing and model-specific kinetics. Because RNA-seq was performed at study endpoints, after regression in most tumors, the dominant transcriptional signatures reflected interferon signaling, TNF-α, and inflammatory pathways consistent with senescence and immune activation. The retreatment schedule in XC5172013 likely captured an intermediate stage where cell cycle suppression was still detectable. Together, these findings suggest that suppression of G2/M and E2F programs are early events, whereas senescence and immune remodeling dominate at later stages.
Importantly, these mechanisms were restricted to tumor contexts. In the non-tumorigenic MCF-10A line, palbociclib and BLU-222 yielded only additive inhibition with no changes in p21/p27 binding, confirming that dual engagement is tumor-specific and not a universal feature of CDK inhibition. Moreover, BLU-222 activated the p21–DREAM pathway58,59,60, providing transcriptional repression of cell cycle genes and reinforcing long-term arrest61. The centrality of this axis highlights how replication stress, p21 induction, and DREAM-mediated repression converge to sustain durable responses.
RB status, a known determinant of CDK4/6 inhibitor response [37], did not diminish BLU-222 activity. Resistant PR1.2 cells retained RB but displayed elevated cyclin E, while PR4.8 cells lost RB entirely. Both remained highly sensitive to BLU-222. Across RB-proficient and RB-deficient contexts, including TNBC lines, resistant clones, and MCF7 shRB, BLU-222 retained potency and synergized with CDK4/6 inhibition, producing apoptosis and G2/M arrest. These findings underscore that CDK2 dependence persists even in RB-null settings, consistent with reports that CDK2 inhibition is synthetically lethal in RB1-deficient cancers62,63.
Cyclin E isoforms further shaped therapeutic response. Full-length cyclin E (EL) delayed cell cycle progression in luminal T47D cells, attenuating BLU-222 sensitivity, whereas LMW-E64 consistently drove hyperactive CDK2 activity and hypersensitivity to BLU-22264. This dichotomy, validated in inducible 76NE6 models65, established an inverse relationship between CDK4/6 and CDK2 inhibitor sensitivity: LMW-E drove palbociclib resistance but enhanced BLU-222 efficacy. Mechanistically, LMW-E upregulation allows cells to bypass the G1 checkpoint by hyperactivating CDK2, sustaining proliferation despite CDK4/6 inhibition. Although prior studies suggest that CDK2 inhibition can initially suppress this activity before adaptive entry into S phase via E2F and cyclin A219, our findings demonstrate that dual inhibition is required to sustain durable arrest.
In vivo, BLU-222 combined with CDK4/6 inhibition effectively overcame cyclin E–mediated resistance, producing tumor regressions in CCNE1-amplified and LMW-E–overexpressing PDX models (BCX017, BCX070, T1p). This dual blockade not only halted CDK2-driven proliferation but also sustained cell cycle arrest, providing a potent therapeutic strategy to counteract resistance in both breast66,67 and ovarian cancer24,68 models. Collectively, these data establish a mechanistic rationale for dual CDK2/CDK4/6 targeting in cyclin E–driven tumors and position LMW-E as a clinically actionable biomarker for patient stratification, enabling the identification of patients most likely to benefit from this therapeutic strategy.
The translational impact of these findings is highlighted by direct comparisons against SOC regimens. Benchmarking showed that the triplet therapy (BLU-222 + CDK4/6 inhibitor + endocrine therapy) consistently outperformed palbociclib + fulvestrant, producing more durable tumor regressions and survival benefit in resistant PDX models. In TNBC, therapeutic partners differed by genomic context: BLU-222 synergized with chemotherapy in CCNE1-amplified tumors, but in non-amplified models, only BLU-222 plus CDK4/6 inhibition achieved durable control. These results argue for tailoring CDK2-based combinations to molecular subsets of breast cancer.
Beyond cell cycle arrest, BLU-222 induced senescence and immune remodeling. Upregulation of interferon pathways suggested immune-mediated elimination of senescent cells, consistent with prior reports linking CDK4/6 inhibition to cGAS-STING activation69. Importantly, these effects were not uniform across all tumors but appeared to be shaped by cyclin E status. In CCNE1-amplified BCX070, BLU-222 monotherapy induced a particularly strong interferon response, highlighting that cyclin E amplification confers an immune-primed phenotype. This aligns with prior clinical evidence linking low-molecular-weight cyclin E to heightened tumor-infiltrating lymphocyte (TIL) presence and immune activity in TNBC70. Supporting this concept, in the immune-competent T1p model engineered to overexpress LMW-E, CDK2 inhibition significantly increased TIL density, with even greater effects when combined with palbociclib. Together, these findings support a dual role for CDK2 inhibition: directly suppressing proliferation while indirectly remodeling the tumor immune microenvironment. Cyclin E–driven tumors may therefore be uniquely poised to couple CDK2 blockade with immune remodeling. More broadly, the convergence of interferon signaling and enhanced immune infiltration positions CDK2 inhibition as a potential strategy to augment antitumor immunity and strengthens the rationale for its integration with immunotherapies in defined breast cancer subsets.
Several limitations warrant consideration. Although we interrogated a broad panel of models, including parental, resistant, engineered, PDX systems, and one immune-competent model (T1p), our studies cannot fully capture the heterogeneity of clinical resistance. The inclusion of T1p provided important validation of our findings, as it demonstrated that CDK2/CDK4/6 co-inhibition synergistically suppressed tumor growth while also remodeling the immune microenvironment, consistent with interferon activation observed in PDX models. However, because most PDX models lack functional human immune compartments, they remain limited in fully recapitulating immune-tumor interactions, and thus the generalizability of immune-related findings requires further evaluation. Mechanistically, while γH2AX induction, p21/p27 upregulation, and DREAM pathway activation provide insight into the cellular consequences of CDK2 inhibition, these endpoints remain correlative and require time-resolved or genetic validation to establish causality. In addition, while we accounted for recurrent genomic alterations such as RB1, TP53, PIK3CA, and ESR1, other genomic drivers not represented in our cohort may influence therapeutic response. Finally, although our preclinical data demonstrate consistent efficacy and tolerability of BLU-222–based combinations, the long-term safety, feasibility, and optimal scheduling of triplet regimens will need to be carefully tested in clinical trials.
Recent advances with other CDK2 inhibitors, such as Pfizer’s PF-07104091, Incyclix’s INX-315 and Incyte’s INCB-123667, which have shown efficacy in HR+ and CCNE1-amplified models8,71,72, further highlight the promise of this therapeutic strategy and support continued development of CDK2-based combinations. These complementary efforts underscore CDK2 as a clinically relevant target, strengthening the translational potential of BLU-222.
In conclusion, our study demonstrates that BLU-222, when combined with CDK4/6 inhibitors, effectively overcomes resistance in both HR+ and TNBC models, including challenging contexts such as RB1-deficient and LMW-E–overexpressing tumors where CDK4/6i monotherapy is typically ineffective. We uncover a mechanistic framework in which upregulation of p21 and p27, coupled with their enhanced binding to both CDK2 and CDK4 complexes, drives therapeutic synergy. Functional CRISPR knockout experiments confirm these proteins as essential mediators rather than passive biomarkers. This effect is further reinforced by increased cellular senescence, validated in vitro and in vivo through biomarker analysis, activation of the p21–DREAM complex, and induction of type I and II interferon signaling, contributing to immune pathway engagement. We also show remodeling of the tumor immune microenvironment, including tumor-infiltrating lymphocyte enrichment in immune-competent models, underscoring a potential immunomodulatory component to the response. Importantly, systematic in vivo benchmarking against SOC regimens across multiple independent PDX and syngeneic models establishes the translational superiority of dual CDK2/CDK4/6 inhibition. Together, these findings advance our biological understanding of resistance mechanisms and provide a strong rationale for clinical testing of CDK2i combinations, as a promising therapeutic strategy for patients with CDK4/6i-resistant breast cancer across diverse molecular subtypes.
Methods
Cell lines and culture conditions
All cell lines used in this study were obtained from the American Type Culture Collection (ATCC). The following authenticated human breast cancer cell lines were used: MCF7 (ATCC HTB-22), T47D (ATCC HTB-133), HCC1806 (ATCC CRL-2335), MDA-MB-157 (ATCC HTB-24), MDA-MB-231 (ATCC HTB-26), and BT20 (ATCC HTB-19). MCF7 and T47D palbociclib-resistant (PR) cells were generated by continuously culturing parental pools in complete medium with stepwise increases in palbociclib concentration (1.2, 2.4, 3.6, and 4.8 μM) over a 6–7-month period28. Similarly, the MDAMB231 PR and BT20 PR cells were generated in medium supplemented with increasing concentrations of palbociclib in a stepwise manner. All PR cells were maintained with corresponding dose of palbociclib used to develop resistance. The cells were maintained in Eagle minimum essential medium, alpha modification, supplemented with 10% fetal bovine serum, 10 mM HEPES, 0.1 mM nonessential amino acids, 1 mM sodium pyruvate, 2 mM L-glutamine, 1 mg/mL insulin, 25 mg/mL epidermal growth factor, and 1 mg/mL hydrocortisone. All cell lines were maintained in a humidified tissue culture incubator at 37 °C and in a 5% CO2 atmosphere. All cell lines were routinely tested for mycoplasma contamination using the MycoAlert Mycoplasma Detection Kit (Lonza) and were authenticated every 6 months by short tandem repeat (STR) profiling and karyotype analysis at the MD Anderson CCSG Cell Line Authentication Core. No cell lines were used beyond 20 passages after authentication.
BLU-222
BLU-222 was provided by Blueprint Medicines (Cambridge, MA). For in vitro studies the drug was prepared in dimethyl sulfoxide (DMSO) at a concentration of 10 mM as stock solution and diluted in full media for cell treatment. For in vivo use, BLU-222 was prepared for 50–100 mg/kg administration in 0.5% (w/v) sodium carboxymethyl cellulose (Na-CMC), 1% (v/v) Tween 80 in water and administered via oral gavage every 12 hours (BID). Stocks for in vivo use were freshly prepared every day and vortexed at 4 °C after preparation.
Dose-response assay
Cells were seeded at a density of 500–3000 cells per well in a 96-well plate overnight and treated with a single drug at the indicated concentrations25,28. In vitro treatments with single agents palbociclib (HY-50767, MCE), abemaciclib (HY-16297A, MCE), ribociclib (HY-15777, MCE), upadacitinib (HY-19569, MCE), abrocitinib (HY-107429, MCE), or BLU-222 were administered for 6 consecutive days, followed by 6 days without treatment. The medium was replaced with or without the drug every other day. At the end of the experiment, cells were incubated in 100 µL of 2.5 mg/mL MTT (RPI) in serum-free media for 4 h at 37 °C, then solubilized in 100 µL of DMSO for 30 minutes at room temperature on a horizontal shaker. Absorbance was measured at an optical density of 560 nm using the Epoch Microplate Spectrophotometer (BioTek Instruments, Inc.). The readings were normalized to those of the untreated controls and analyzed by nonlinear regression using GraphPad Prism 10 software to obtain half-maximal inhibitory concentration (IC50) values. Each condition included eight technical replicates per concentration and was repeated in at 3 three independent experiments.
Flow cytometry
Cell cycle analysis: Cells were plated at 3–5 × 105 cells on 10-cm petri dish and grown in medium containing 0.3 μM BLU-222 and/or 0.3 μM of palbociclib for 3 days. The medium was replaced every other day with or without drug. At the end of the treatment, cells were trypsinized, washed with 1× phosphate-buffered saline (PBS), and fixed with 3.5 mL of ice-cold 1× PBS and 1.5 mL of 95% ethanol at 4 °C overnight. Cells were then stained with propidium iodide (1 mg/mL) and RNaseA (1 mg/mL) in 1× PBS without Ca/Mg containing 0.5% (v/v) Tween 20 (Sigma-Aldrich) and 0.5% bovine serum albumin at 4 °C overnight protected from light. Samples were read using an Attune Flow Cytometer (Thermo Fisher Scientific), and data were analyzed with FlowJo v10.10.0 software. Each condition included 2 technical replicates and was repeated in at least 3 independent experiments.
Apoptosis analysis: Cells were plated at 3–5 × 105 cells on 10-cm petri dish and grown in medium containing 0.3 μM BLU-222 and/or 0.3 μM of palbociclib for 3 days. At the end of the treatment, cells were trypsinized, centrifuged and washed with 1× PBS. The apoptotic cells were labeled with Alexa Fluor 488 Annexin V according to the manufacturer’s protocol (Invitrogen, #V13241). Samples were read using an Attune Flow Cytometer, and data were analyzed with FlowJo v10.10.0 software. Each condition included 2 technical replicates and was repeated in at least 3 independent experiments.
Synergistic assay
For combination treatments, drug 1 was used at fixed concentrations of 0, IC25, IC50, and IC75, while drug 2 was applied in a serial dilution ranging from 0 to 10 mM. All cells were treated with the drugs for 6 consecutive days (drug-on) followed by 6 days without treatment (drug-off), with the medium (with or without drugs) replaced every other day. At the end of the experiment, cells were stained with 0.5% crystal violet solution and then solubilized with 0.1% sodium citrate in 50% ethanol. Absorbance was measured at 570 nm using the Epoch Microplate Spectrophotometer. Each condition included 4 technical replicates and was repeated in at least three independent experiments.
Cell viability values for single agents and combination treatments were normalized to their respective no-treatment controls. The cell viability data were averaged for each condition and used to generate a matrix table, which was processed using SynergyFinder version 3.0, a web application with R package-implemented algorithms to calculate synergy scores for dose–response matrix data (https://synergyfinder.fimm.fi). The highest single agent (HSA) model was applied to calculate the synergy score. A synergy score less than 0 indicates that the interaction between the two drugs is likely antagonistic, a score between 0 and 10 suggests an additive interaction, and a score greater than 10 indicates a synergistic interaction.
Western blot and immunoprecipitation analysis
Protein expression was analyzed by Western blotting using whole cell lysates lysed in radioimmunoprecipitation assay buffer with a cocktail of protease/phosphatase inhibitors (25 g/mL leupeptin, 25 g/mL aprotinin, 10 g/mL pepstatin, 1 mmol/L benzamidine, 10 g/mL soybean trypsin inhibitor, 0.5 mmol/L phenylmethylsulfonyl fluoride, 50 mmol/L sodium fluoride, and 0.5 mmol/L sodium orthovanadate)73,74. Antibodies as follows: cyclin E1 (HE12; Santa Cruz Biotechnology #sc-247, RRID: AB_627357; 1:1000), cyclin D1 (H-295; Santa Cruz Biotechnology #sc-753, RRID: AB_2070433; 1:1000), Rb (4H1; Cell Signaling Technology #9309, RRID: AB_823629; 1:1000), phospho-Rb (Ser807/811; Cell Signaling Technology #9308, RRID: AB_331472; 1:1000), CDK4 (C-22; Santa Cruz Biotechnology #sc-260-G, RRID:AB_631219; 1:1000), CDK6 (C-21; Santa Cruz Biotechnology #sc-177-G, RRID: AB_631226; 1:1000), CDK2 (D-12; Santa Cruz Biotechnology #sc-6248, RRID: AB_627238; 1:1000), phospho-CDK2 (Thr160; Cell Signaling Technology #2561, RRID:AB_2078685; 1:1000), phospho-histone H2AX (Ser139, γ-H2AX; Millipore, #05-636, 1:1000, RRID:AB_309864; 1:1000), p21 (12D1; Cell Signaling Technology, #2947S, RRID:AB_823586; 1:1000), p27 (D69C12; Cell Signaling Technology, #3686S, RRID:AB_2077850; 1:1000), vinculin (hVIN-1; Sigma-Aldrich, #V9131, RRID: AB_477629; 1:2000) and actin (C4; Millipore #MAB1501, RRID: AB_2223041; 1:5000).
For immunoprecipitation, 1000 µg of protein extracted from total-cell lysates for each immunoprecipitation was used with polyclonal antibodies against CDK2, CDK4, or normal mouse immunoglobulin G (IgG) (SC-2025; Santa Cruz Biotechnology) and incubated with protein A–coated or protein G–coated beads (Calbiochem)75. Beads were washed three times with lysis buffer, resuspended in SDS sample buffer, and heated at 95 °C for 5 min. Immunoprecipitated proteins were resolved by SDS–PAGE and analyzed by immunoblotting with the antibodies listed above. Uncropped and unprocessed scans of the Western blots are provided in the Source Data file. Antibody validation was based on expected molecular weight, published validation data, and manufacturer-provided validation resources.
Short hairpin RNA knockdown and cyclin E overexpression
To establishment of the CDK2 or CCNE1-knockdown HCC1806 and MDA-MB-468 cell lines, and Rb1-knockdown MCF7 cells, lentiviral constructs expressing shRNA targeting the CDK2 gene (Clone ID: V3LHS_637573), CCNE1 gene (Clone ID: V2LHS_642332), Rb1#1 (Clone ID: V2LHS_130609), Rb1#2 (Clone ID: V2LHS_130608) were obtained from the MD Anderson Functional Genomics Core. ShRNA constructs and the packaging vectors pMDG.2 and pCMV deltaR8.2 (obtained from the laboratory of Didier Trono, École Polytechnique Fédérale de Lausanne) were co-transfected into HEK293T cells for lentiviral packaging. After 48 h of transfection, the virus-containing medium was collected, filtered through a 0.45-μm filter and added to the cells of interest in the presence of 8 μg/mL of polybrene (Millipore). GFP expression was confirmed and the lentivirus-infected cells were selected with 1 μg/mL puromycin (InvivoGen, San Diego, CA), which was followed by confirmation of successful knockdown via Western blotting42.
The T47D, MDA-MB-231 and 76NE6-EKO cells were generated by stable transfection with either full-length cyclin E (FL-cycEL) or low-molecular-weight cyclin E (LMW-E). MCF-7 cells were transfected with empty vector pcDNA4.0 (Invitrogen), cyclin EL1-FLAG, or cyclin E-FLAG LMW constructs using FuGENE (Invitrogen). Transfected cells were selected in 80 μg/mL zeocin, and individual colonies were isolated and screened by Western blotting with anti-FLAG (M2; Sigma) and anti–cyclin E (HE-12; Santa Cruz Biotechnology)18,75.
CRISPR-Cas9 genome editing
Waf1/Cip1/CDKN1A p21 CRISPR/Cas9 KO plasmids (sc-400013, Santa Cruz Biotechnology) or CDKN1B/Kip1 p27 CRISPR/Cas9 KO plasmids (sc-400074-KO-2, Santa Cruz Biotechnology) were transfected into early-passage MCF7 and T47D parental and PR cells using Lipofectamine 3000 according to the manufacturer’s instructions (L300001; Invitrogen). The p21 CRISPR/Cas9 KO plasmid is a pool of 3 different gRNA plasmids:
-
1.
sc-400013 A (CCGCGACTGTGATGCGCTAA)
-
2.
sc-400013 B (CCATTAGCGCATCACAGTCG)
-
3.
sc-400013 C (AGATCAGCCGGCGTTTGGAG)
The p27 Kip1 CRISPR/Cas9 KO plasmid is a pool of 3 different gRNA plasmids:
-
1.
sc-400074-KO-2 A (TCAAACGTGCGAGTGTCTAA)
-
2.
sc-400074-KO-2 B (TCAGACGGTTCCCCAAATGC)
-
3.
sc-400074-KO-2 C (TTCCCCAAATGCCGGTTCTG)
After 96 hours of transfection, single-cell sorting was performed based on green fluorescent protein (GFP) expression to select successfully transfected cells. The top 5% of GFP-positive cells were gated and sorted into a 96-well plate for culturing as single clones. Colonies derived from single cells were expanded, and successful knockouts were confirmed using Western blotting.
Proximity ligation assay
Cells were seeded in an 8-well chamber slide (Nunc Lab-Tek Chamber Slide, #177445) at 20% confluence 1 day before treatment. Palbociclib and BLU-222 were dissolved in DMSO at a stock concentration of 10 mM. These inhibitors were then freshly diluted to a final concentration of 0.3 μM in cell culture media at the time of the experiment. Cells were incubated with DMSO, palbociclib, BLU-222, or a combination of palbociclib and BLU-222 for 48 h. The inhibitor-containing media were refreshed, and cells were further incubated for another 24 h, for a total treatment duration of 72 h. The cells were fixed with freshly prepared 4% paraformaldehyde on ice for 10 min, then permeabilized with 0.2% Triton X-100 for 5 min at room temperature. The Duolink PLA assay (Sigma-Aldrich, #DUO92008) was performed according to the manufacturer’s protocol using the following primary antibodies: mouse anti-CDK2 (Abnova, #00001017-D01P, 1:80), rabbit anti-p21 (Cell Signaling Technology, #2947, 1:100), and rabbit anti-p27 (Cell Signaling Technology, #3686, 1:100). After amplification and final washing steps, the slides were stained with Alexa Fluor 488 phalloidin (Invitrogen, #A12379, 1:1000) for 30 minutes at room temperature before applying DAPI-containing Duolink In Situ Mounting media. The fluorescent signals were examined and recorded using a Zeiss Confocal 810, and puncta were counted using BlobFinder76.
Senescence-associated β-galactosidase (SA-β-gal) staining
We used a commercially available SA-β-Gal staining kit (Cell Signaling Technology, Catalog No. 9860S) according to the manufacturer’s protocol. Cells were seeded in 24-well plates at a density of 5 ×103 cells per well and treated the following day with vehicle, 0.3 μM palbociclib, 0.3 μM BLU-222, or their combination for 4 days. At endpoint, cells were gently rinsed twice with PBS (3 min each), fixed with 500 μL of fixative solution for 20 min at room temperature, and washed once with PBS. Subsequently, 1 mL of staining working solution was added per well, and plates were sealed with Parafilm and wrapped in aluminum foil to protect from light and evaporation. Samples were incubated at 37 °C in a CO2-free incubator for 24 h. Images were acquired on an inverted microscope at 40x magnification. For quantification, five randomly selected non-overlapping fields per well were analyzed, and the percentage of SA-β-Gal-positive (blue-green) cells was calculated as (number of positive cells/total cells) × 100%. Each experiment was repeated 3 times independently.
PDX model generation and maintenance
The TNBC PDX models BCX017, BCX070 and XC5172013 were previously developed42,43. Briefly, fresh tumor fragments ( ~ 3 mm³) were implanted into immunodeficient mice within 1 h of resection. When outgrowths reached ~10 mm, fragments were passage into new hosts (n = 3). Stable growth after two passages was considered successful PDX establishment. The HR+ breast cancer PDX model PR4 (originally named T141-004), which progressed on palbociclib, was provided by Dr. Funda Meric-Bernstam’s laboratory at MD Anderson. The generation of the 3 PDX models PR1, PR2, and PR3, originally named CTG-3277, CTG-3266, and CTG-2308, respectively, were obtained from Champions Oncology, Inc. through a Material Transfer Agreement with MDACC28.
In this study, the PDX models were re-established from the liquid nitrogen tumor bank and transplanted into the T4 mammary fat pad of 8-week-old female NSG mice from Jackson Lab (strain #005557). After transplantation, tumor size was monitored, and when the primary tumor outgrowths reached 1000–1500 mm³, 4×4-mm fragments of the outgrowths were implanted in new hosts (n = 3 per tumor) as a secondary passage. As the tumor tissues can stably grow after two passages with our protocol, we considered the PDX line to have been successfully established at that point. During maintenance, HR+ breast cancer PDX models were given drinking water supplemented with 8 μg/mL estradiol (E2), except for the PR4 and the two TNBC models, which are independent of E2. Only female mice were used, consistent with standard breast cancer tumor implantation models. Sex was therefore not analyzed as an independent variable. Sex-disaggregated data are provided as female-only, as all mice used in the study were female. The number of animals used in each experiment is provided in the corresponding figure legends.
In vivo CDX model establishment
In vivo HCC1569 CDX study was performed at Shanghai ChemPartner (RRID:SCR_026139) according to the protocols approved by the IACUC. Female NOD-SCID mice (6–8 weeks old) were implanted subcutaneously in the flanks with 1 × 10⁷ HCC1569 cells mixed with Matrigel. Tumor growth was monitored twice weekly with digital calipers. When mean tumor volume reached 150-250 mm³, mice were randomized into four treatment groups (n = 4 per group): vehicle, BLU-222 (100 mg/kg BID, oral gavage), paclitaxel (10 mg/kg QW, intraperitoneal), or BLU-222 + paclitaxel combination (100 mg/kg BID + 10 mg/kg QW). BLU-222 was formulated in 0.5% (w/v) CMC-Na with 1% (v/v) Tween-80, and paclitaxel was prepared in 5% Cremophor EL + 5% ethanol + 90% saline. Dosing was performed for 28 consecutive days, after which treatments were discontinued and tumors monitored for regrowth until day 49.
In vivo PDX efficacy studies
PDX tumor fragments (4×4 mm) were transplanted into the T4 mammary fat pad of female NCR-NU-F sp/sp nude mice (10-week-old) from Taconic Biosciences. Tumors were allowed to grow until they reached 150–200 mm³. HR+ breast cancer PDX models (except PR4 and the TNBC models) were maintained with drinking water supplemented with E2 (8 μg/mL). Once tumors reached an average volume of 200 mm³, mice were randomized to the indicated treatment arms and treated with vehicle, palbociclib (50 mg/kg once daily, oral gavage), BLU-222 (60 mg/kg twice daily, oral), or a combination of palbociclib (50 mg/kg) and BLU-222 (60 mg/kg) plus fulvestrant (2.5 mg/kg, once weekly, subcutaneous). The vehicle for palbociclib and ribociclib is 0.5 w/v methylcellulose (Fujifilm, #133-17815), the vehicle for BLU-222 is 0.5% Na-CMC/1% TW80, and the vehicle for fulvestrant is 10% DMSO diluted in 90% corn oil. The E2 water was continued throughout the treatment except PR4 and TNBC models. The length and width of tumor xenografts were measured by caliper 3 times per week, and tumor volume was calculated using the following formula: volume = (length × (width)²)/2. All animal procedures complied with the tumor size and humane endpoint limits approved by the MD Anderson Institutional Animal Care and Use Committee. The maximal tumor burden permitted under our protocol was a volume of 1500 mm³ or a diameter not exceeding 20 mm. No tumor exceeded these IACUC-approved limits during the study. Following euthanasia, all tumors were harvested, random fragments were frozen and stored at −80 °C for RNA/protein assessment, and one piece was fixed in formalin for histopathological and IHC analysis. Palbociclib isethionate and fulvestrant were purchased from MedChemExpress (palbociclib #HY-A0065; fulvestrant #HY-13636).
Pharmacokinetics
The pharmacokinetic (PK) profiles of palbociclib and BLU-222 were evaluated in 10-week-old female FBV (The Jackson Laboratory) and NCR-NU-F nude mice (Taconic Biosciences) using single-agent and combination dosing. Mice were randomly divided into 4 treatment groups: single-agent palbociclib (50 mg/kg), single-agent BLU-222 (60 mg/kg and 100 mg/kg), and a combination of palbociclib and BLU-222 (50 mg/kg palbociclib and 60 mg/kg BLU-222). BLU-222 was formulated in 0.5% (w/v) CMC-Na with 1% (v/v) Tween-80, and palbociclib was prepared in 0.5% (w/v) methylcellulose (Fujifilm, #133-17815). and administered via oral gavage, either alone or simultaneously for the combination group. Blood samples (approximately 50 µL) were collected from the retro-orbital sinus at predetermined time points (2, 11.5, 14, and 24 hours post-dosing) following the Retro-Orbital Bleed Standard Operating Procedure approved by IACUC. Mice were anesthetized prior to sampling to minimize discomfort. Collected blood samples were immediately transferred into EDTA-coated tubes and centrifuged for 5 minutes at 1500 x g without using the brake to prevent sample disturbance. The separated plasma was carefully transferred into pre-labeled tubes and stored at −80 °C until further analysis. Plasma concentrations of palbociclib and BLU-222 were quantified using a validated high-performance liquid chromatography–tandem mass spectrometry method.
Complete Blood Count Analysis for Blood Toxicity in Treated Mice
At the end of the treatment period, female FBV (The Jackson Laboratory) and NCR-NU-F nude mice (Taconic Biosciences) bearing tumors were euthanized, and blood samples were collected via cardiac puncture. Briefly, a needle was carefully inserted into the thoracic cavity, and blood was slowly withdrawn upon confirmation of successful entry into the heart chamber. Approximately 500 µL of blood was collected and transferred into EDTA-coated capillary tubes. The samples were immediately submitted to the Animal Pathology Core at MD Anderson Cancer Center and processed for complete blood count analysis using an automated hematology analyzer (Hemavet 950FS). The parameters measured included white blood cell count, red blood cell count, hemoglobin, platelet count, and other relevant indices. Statistical significance was determined using 1-way analysis of variance (ANOVA) followed by post hoc analysis, with a p-value < 0.05 considered indicative of significant blood toxicity.
IHC and analysis of tumor tissues
Tumor tissues were formalin-fixed and embedded into paraffin blocks and sectioned for IHC. The slides were deparaffinized and rehydrated, and heat-induced epitope retrieval was performed for 20 minutes in a citrate- or Tris-based antigen unmasking solution (H-3300 or H-3301, Vector Labs) and subsequently cooled at room temperature for 30 minutes. The slides were then incubated with 3% hydrogen peroxide in methanol for 10 min and 1.5% normal goat or horse serum for 1 hour. The sections were incubated overnight at 4 °C in a humid chamber with primary antibodies at indicated dilutions (Supplemental Table S3). For some antibodies, steps were followed from the manual of VECTASTAIN Elite ABC kit (rabbit IgG) (#6101, Vector Labs), and the signals were detected by DAB substrate (#SK-4100, Vector Labs). For others, the slides were incubated with a secondary antibody horse anti-mouse IgG H&L (HRP) (#PI-2000, Vector Labs) at room temperature for 2 hours after washing, and the signals were detected by ImmPACT DAB substrate (#SK-4105, Vector Labs). After that, the sections were counterstained with hematoxylin (Dako) and mounted with Permount Mounting Medium (#SP15-100, Fisher Chemical). All washing steps were performed in PBS alone or PBS with 0.1% Tween. The IHC conditions for the markers were listed in Supplemental Table S3. Stained slides were thereafter scanned on the Aperio CS2 slide scanner (20× magnification).
For the analysis of IHC antibody expression, tumor slides were scored for the percentage of positive cells. Three areas per tumor were randomly selected and calculated for positive cell detection. The intensity threshold parameter was set to 1+ with a cutoff of 0.2. Positivity percentage was calculated by the mean value of quantification from three separate areas per sample. The data were analyzed using QuPath v0.5.1 (https://qupath.github.io).
Sudan Black B (SBB) staining for senescence detection
Senescence-associated lipofuscin accumulation was detected using a Sudan Black B (SBB) staining protocol adapted from Davan-Wetton and Montero-Melendez (PLoS One, 2024)77. Briefly, tumor tissue sections were fixed in 4% paraformaldehyde for 15 min at room temperature, washed in PBS, and permeabilized with 70% ethanol for 2 min. Samples were incubated with freshly prepared, triple-filtered saturated SBB solution (1.2 g in 80 mL 70% ethanol, stirred overnight and filtered through 70, 0.45, and 0.22 μm filters) for 8 min on a plate shaker, followed by washing in distilled water. Nuclei were counterstained with Nuclear Fast Red (200 mL of 0.5% solution in 5% aluminum sulfate) for 10 min, or with DAPI (1 μg/mL, 10 min) for fluorescent quantification. Slides were washed in PBS and mounted in aqueous medium. Brightfield imaging was performed for Nuclear Fast Red visualization, while Sudan Black B autofluorescence was quantified using a Cy5 filter set (far-red, excitation 628/40 nm, emission 692/40 nm). Negative controls without SBB were included to confirm specificity. For quantification, three randomly selected non-overlapping fields per sample were analyzed, and the percentage of far-red positive cells was quantified using QuPath.
Tumor-infiltrating Lymphocytes (TILs) analysis
FFPE samples from T1P tumors were stained with hematoxylin and eosin (H&E). TIL scoring was performed according to the Immuno-Oncology International TILs Working Group guidelines, in which stromal TILs are defined as the proportion of the stromal area occupied by lymphocytes without direct contact with invasive tumor cells. Microscopic assessment was carried out in three randomly selected non-overlapping fields per sample. Detailed guidance for TIL quantification in breast tissue is available at https://www.tilsinbreastcancer.org.
RNA sequencing and gene set enrichment analysis
Total RNA was extracted with TRIzol reagent (Ambion Life Technologies 15596026) following the manufacturer’s instructions. RNA purity and concentration were measured on a NanoDROP 2000c spectrophotometer (Thermo Fisher Scientific). Preparation of RNA library and transcriptome sequencing was conducted by Novogene Corporation Inc. Briefly, messenger RNA was purified from total RNA using poly-T oligo-attached magnetic beads. After fragmentation, the first-strand cDNA was synthesized using random hexamer primers, and then the second-strand cDNA was synthesized using deoxythymidine triphosphate for a nondirectional library. The library was checked with Qubit and real-time polymerase chain reaction for quantification and bioanalyzer for size distribution detection. Quantified libraries were pooled and sequenced on Illumina platforms, according to effective library concentration and data amount. The clustering of the index-coded samples was performed according to the manufacturer’s instructions. After cluster generation, the library preparations were sequenced on an Illumina platform, and paired-end reads were generated.
The data processing and bioinformatics analyses were performed by the Department of Bioinformatics and Computational Biology at MD Anderson Cancer Center. Briefly, paired-end reads in FASTQ format were checked for read quality using FastQC/v0.11.8 and then aligned to the combined reference genome (GENCODE human GRCh38 and mouse GRCm38 concatenated) using TopHat2/v2.1.178. FeatureCounts79 (Subread 1.6.3) was used to count reads that are mapped to each feature. The features were annotated using the combined gene transfer format (GTF) file using GENCODE human and mouse reference (gencode.vM20.annotation.gtf for mouse and gencode.v29.annotation.gtf for human)80. Only the human gene count matrix was retrieved for downstream analysis. The read counts for all samples were then normalized using the trimmed mean of M method implemented in the R Bioconductor package (Bioconductor, RRID:SCR_006442) edgeR (edgeR, RRID:SCR_012802) to generate the abundance for each feature81. We applied the edgeR pipeline with default filtering, TMM normalization, dispersion estimation, and quasi-likelihood F-test based on the negative binomial generalized linear model to determine differentially expressed genes (controlled by false discovery rate (FDR) cutoff 5%) between groups. The Benjamini-Hochberg correction was applied to the resulting p-values for multiple testing adjustment82.
Functional analysis of the differentially expressed gene sets or pathways was performed using gene set enrichment analysis (GSEA)83. We conducted pre-ranked GSEA for the comparisons of each resistant cell line vs. the parental cell line to explore enriched pathways. The Hallmark and Reactome gene sets from the Molecular Signatures Database (MSigDB) were used. All the genes tested were annotated in HUGO gene symbols and ranked based on the significance (p-values) in combination with the sign of log2 fold change values resulting from 2-group comparisons. We used the FDR cutoff 5% to select significant gene sets for each comparison.
Single-sample GSEA (ssGSEA) was performed using the GSVA package84. This analysis was performed using the log2-transformed data in transcripts per million data, which was generated using RSEM/v1.3.3 based on the FASTQ files with human reads and GENCODE human reference GRCh38. For each individual sample, an enrichment score was calculated by applying a weighted running sum to the normalized ranking scores of genes within the gene set. For generating senescence scores, gene sets were obtained from Reactome_Cellular-Senescence85 and from Casella86, Fridman87, Hernandez-Segura88, and Purcell and colleagues89. For p21/DREAM pathway scores, gene sets were obtained from MSigDB database v7.5.1: FISCHER_DREAM_TARGETS (DREAM-F), FISCHER_DIRECT_P53_TARGETS_META_ANALYSIS, FISCHER_G1_S_CELL_CYCLE, FISCHER_G2_M_CELL_CYCLE, SHEPARD_BMYB_TARGETS. The FOXM1 and E2F-RB target genes were obtained from Fischer et al.50. A second set of DREAM targets (DREAM-E) was obtained from Engeland90. A set of DREAM-dependent p53-targets (DREAM-U) and a set of DREAM-independent p53 target genes were obtained from Uxa et al.91.
Statistical analyses
GraphPad Prism (v10), IBM SPSS Statistics (v24; RRID: SCR_002865) software, and R (v4.2.2; https://www.r-project.org/), a publicly available statistics analysis tool, were used for data analysis. Statistical analyses were performed using the Student t-test for comparisons between 2 groups, and 1-way or 2-way ANOVA for comparisons among multiple groups. All data represent the mean ( ± standard error of the mean) from at least 3 independent experiments or samples unless otherwise specified. For survival studies, Kaplan-Meier curves were used to compare cohorts, and the log-rank (Mantel-Cox) test was applied to determine differences. Band intensity from western blotting was quantified using ImageJ (RRID: SCR_003070). Specific statistical methods are outlined in the respective figure legends where applicable.
Ethics Statement
All animal experiments were conducted in compliance with institutional and national guidelines and were approved by the MD Anderson Institutional Animal Care and Use Committee. Procedures adhered to the standards specified by the Association for Assessment and Accreditation of Laboratory Animal Care (AAALAC International).
Human breast tumor tissues used to establish patient-derived xenograft (PDX) models were collected during surgery after written informed consent, in accordance with protocols approved by the MD Anderson Cancer Center Institutional Review Board and all relevant ethical regulations.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The publicly available data used in this study are available in the Gene Expression Omnibus (GEO) under accession code GSE282705. The dataset is publicly available without restrictions. All data supporting the findings of this study are available within the article, the Supplemental Information, and the Source Data file. No restricted or confidential datasets were generated for this study. Source data are provided with this paper.
References
Malumbres, M. & Barbacid, M. Cell cycle, CDKs and cancer: a changing paradigm. Nat. Rev. Cancer 9, 153–166 (2009).
Fagundes, R. & Teixeira, L. K. Cyclin E/CDK2: DNA replication, replication stress and genomic instability. Front Cell Dev. Biol. 9, 774845 (2021).
Tadesse, S. et al. Targeting CDK2 in cancer: challenges and opportunities for therapy. Drug Discov. Today 25, 406–413 (2020).
Senderowicz, A. M. Flavopiridol: the first cyclin-dependent kinase inhibitor in human clinical trials. Invest N. Drugs 17, 313–320 (1999).
Meijer, L. & Raymond, E. Roscovitine and other purines as kinase inhibitors. From starfish oocytes to clinical trials. Acc. Chem. Res 36, 417–425 (2003).
Parry, D. et al. Dinaciclib (SCH 727965), a novel and potent cyclin-dependent kinase inhibitor. Mol. Cancer Ther. 9, 2344–2353 (2010).
Mita, M. M. et al. Randomized phase II trial of the cyclin-dependent kinase inhibitor dinaciclib (MK-7965) versus capecitabine in patients with advanced breast cancer. Clin. Breast Cancer 14, 169–176 (2014).
Dietrich, C. et al. INX-315, a selective CDK2 inhibitor, induces cell cycle arrest and senescence in solid tumors. Cancer Discov. 14, 446–467 (2024).
House N. C. et al. Profiling the Activity of the Potent and Highly Selective CDK2 Inhibitor BLU-222 Reveals Determinants of Response in CCNE1-Aberrant Ovarian and Endometrial Tumors. Cancer 85, 1297–1309 (2025).
Hydbring, P. et al. Phosphorylation by Cdk2 is required for Myc to repress Ras-induced senescence in cotransformation. Proc. Natl. Acad. Sci. USA 107, 58–63 (2010).
Martín, A. et al. Cdk2 is dispensable for cell cycle inhibition and tumor suppression mediated by p27Kip1 and p21Cip1. Cancer Cell 7, 591–598 (2005).
Zalzali, H. et al. CDK2 transcriptional repression is an essential effector in p53-dependent cellular senescence—implications for therapeutic intervention. Mol. Cancer Res. 13, 29–40 (2015).
Zingoni, A. et al. The senescence journey in cancer immunoediting. Mol. Cancer 23, 68 (2024).
Caruso, J. A., Duong, M. T., Carey, J. P. W., Hunt, K. K. & Keyomarsi, K. Low-molecular-weight cyclin E in human cancer: cellular consequences and opportunities for targeted therapies. Cancer Res. 78, 5481–5491 (2018).
Karakas, C. et al. Cytoplasmic cyclin E and phospho-cyclin-dependent kinase 2 are biomarkers of aggressive breast cancer. Am. J. Pathol. 186, 1900–1912 (2016).
Hunt, K. K. et al. Cytoplasmic cyclin E predicts recurrence in patients with breast cancer. Clin. Cancer Res. 10, 2217 (2016).
Vijayaraghavan, S., Moulder, S., Keyomarsi, K. & Layman, R. M. Inhibiting CDK in cancer therapy: current evidence and future directions. Target Oncol. 13, 21–38 (2017).
Vijayaraghavan, S. et al. CDK4/6 and autophagy inhibitors synergistically induce senescence in Rb positive cytoplasmic cyclin E negative cancers. Nat. Commun. 8, 15916 (2017).
Arora, M. et al. Rapid adaptation to CDK2 inhibition exposes intrinsic cell-cycle plasticity. Cell 186, 2628–2643 e2621 (2023).
Kawakami, M. et al. Next-generation CDK2/9 inhibitors and anaphase catastrophe in lung cancer. J. Natl. Cancer Inst. 109 https://doi.org/10.1093/jnci/djw297 (2017).
Schomer, N. et al. CDK2 inhibition demonstrates synthetic lethality in SCLC through apoptotic induction. Mol. Cancer Therapeutics 22 https://doi.org/10.1158/1535-7163.Targ-23-A022 (2023).
Patel, M. R. et al. BLU-222, an oral, potent, and selective CDK2 inhibitor, in patients with advanced solid tumors: phase 1 monotherapy dose escalation. J. Clin. Oncol. 41 https://doi.org/10.1200/JCO.2023.41.16_suppl.3095 (2023).
Yap, T. A. et al. in Annals of Oncology 33, S765-S765 (2022).
Juric, D. et al. BLU-222, an investigational, oral, potent, and highly selective CDK2 inhibitor (CDK2i), as monotherapy in patients (pts) with advanced solid tumors and in combination with ribociclib (RIBO) and fulvestrant (FUL) in HR+/HER2-breast cancer (BC). J. Clin. Oncol. 42, 1056 (2024).
Li, M. et al. Low molecular weight cyclin E confers a vulnerability to PKMYT1 inhibition in triple-negative breast cancer. Cancer Res. 84, 3864–3880 (2024).
Bagheri-Yarmand, R., Biernacka, A., Hunt, K. K. & Keyomarsi, K. Low molecular weight cyclin E overexpression shortens mitosis, leading to chromosome missegregation and centrosome amplification. Cancer Res. 70, 5074–5084 (2010).
Wingate, H. F. & Keyomarsi, K. Distinct mechanisms of resistance to CDK4/6 inhibitors require specific subsequent treatment strategies: one size does not fit all. Cancer Res. 83, 3165–3167 (2023).
Navarro-Yepes, J. et al. Abemaciclib is effective in palbociclib-resistant hormone receptor-positive metastatic breast cancers. Cancer Res 83, 3264–3283 (2023).
Kettner, N. M. et al. Combined inhibition of STAT3 and DNA repair in palbociclib-resistant ER-positive breast cancer. Clin. Cancer Res. 25, 3996–4013 (2019).
Portman, N. et al. Overcoming CDK4/6 inhibitor resistance in ER-positive breast cancer. Endocr. Relat. Cancer 26, R15–R30 (2019).
Herrera-Abreu, M. T. et al. Early adaptation and acquired resistance to CDK4/6 inhibition in estrogen receptor-positive breast cancer. Cancer Res 76, 2301–2313 (2016).
Turner, N. C. et al. Cyclin E1 expression and palbociclib efficacy in previously treated hormone receptor-positive metastatic breast cancer. J. Clin. Oncol. 37, 1169–1178 (2019).
Guarducci, C. et al. Cyclin E1 and Rb modulation as common events at time of resistance to palbociclib in hormone receptor-positive breast cancer. npj Breast Cancer 4, 38 (2018).
Duarte, D. & Vale, N. Evaluation of synergism in drug combinations and reference models for future orientations in oncology. Curr. Res Pharm. Drug Discov. 3, 100110 (2022).
Finn, R. S. et al. PD 0332991, a selective cyclin D kinase 4/6 inhibitor, preferentially inhibits proliferation of luminal estrogen receptor-positive human breast cancer cell lines in vitro. Breast Cancer Res. 11, 1–13 (2009).
Harper, J. W., Adami, G. R., Wei, N., Keyomarsi, K. & Elledge, S. J. The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 75, 805–816 (1993).
el-Deiry, W. S. et al. WAF1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993).
Abbas, T. & Dutta, A. p21 in cancer: intricate networks and multiple activities. Nat. Rev. Cancer 9, 400–414 (2009).
Polyak, K. et al. p27KIP1, a cyclin-cdk inhibitor, links transforming growth factorß and contact inhibition to cell cycle arrest. Genes Dev. 8, 9–22 (1994).
Slingerland, J. & Pagano, M. Regulation of the cdk inhibitor p27 and its deregulation in cancer. J. Cell Physiol. 183, 10–17 (2000).
Kettner, N. M. et al. IL-6 predicts CDK4/6 inhibitor resistance, identifying STAT3 as a target in HR + /HER2-negative metastatic breast cancer. NPJ Precis Oncol. 9, 260 (2025).
Chen, X. et al. Cyclin E overexpression sensitizes triple-negative breast cancer to wee1 kinase inhibition. Clin. Cancer Res. 24, 6594–6610 (2018).
Evans, K. W. et al. A population of heterogeneous breast cancer patient-derived xenografts demonstrate broad activity of PARP inhibitor in BRCA1/2 wild-type tumors. Clin. Cancer Res. 23, 6468–6477 (2017).
Akli, S., Van Pelt, C. S., Bui, T., Meijer, L. & Keyomarsi, K. Cdk2 is required for breast cancer mediated by the low-molecular-weight isoform of cyclin E. Cancer Res. 71, 3377–3386 (2011).
Akli, S. et al. Overexpression of the low molecular weight cyclin E in transgenic mice induces metastatic mammary carcinomas through the disruption of the ARF-p53 pathway. Cancer Res. 67, 7212–7222 (2007).
Mandal, M., Bandyopadhyay, D., Goepfert, T. M. & Kumar, R. Interferon-induces expression of cyclin-dependent kinase-inhibitors p21WAF1 and p27Kip1 that prevent activation of cyclin-dependent kinase by CDK-activating kinase (CAK). Oncogene 16, 217–225 (1998).
Xiao, S. et al. RIG-G as a key mediator of the antiproliferative activity of interferon-related pathways through enhancing p21 and p27 proteins. Proc. Natl. Acad. Sci. USA 103, 16448–16453 (2006).
Huang, W., Hickson, L. J., Eirin, A., Kirkland, J. L. & Lerman, L. O. Cellular senescence: the good, the bad and the unknown. Nat. Rev. Nephrol. 18, 611–627 (2022).
Georgakopoulou, E. A. et al. Specific lipofuscin staining as a novel biomarker to detect replicative and stress-induced senescence. A method applicable in cryo-preserved and archival tissues. Aging 5, 37–50 (2013).
Fischer, M., Grossmann, P., Padi, M. & DeCaprio, J. A. Integration of TP53, DREAM, MMB-FOXM1 and RB-E2F target gene analyses identifies cell cycle gene regulatory networks. Nucleic Acids Res. 44, 6070–6086 (2016).
Orlando, S. et al. p27Kip1 and p21Cip1 collaborate in the regulation of transcription by recruiting cyclin-Cdk complexes on the promoters of target genes. Nucleic Acids Res. 43, 6860–6873 (2015).
Alvarez-Fernandez, M. & Malumbres, M. Mechanisms of sensitivity and resistance to CDK4/6 inhibition. Cancer Cell 37, 514–529 (2020).
Guiley, K. Z. et al. p27 allosterically activates cyclin-dependent kinase 4 and antagonizes palbociclib inhibition. Science 366 https://doi.org/10.1126/science.aaw2106 (2019).
Zhang, C. et al. Signalling involving MET and FAK supports cell division independent of the activity of the cell cycle-regulating CDK4/6 kinases. Oncogene 38, 5905–5920 (2019).
Iida, M. et al. The p21 levels have the potential to be a monitoring marker for ribociclib in breast cancer. Oncotarget 10, 4907–4918 (2019).
Poulet, S. et al. Genome-wide in vivo CRISPR screen identifies TGFβ3 as actionable biomarker of palbociclib resistance in triple negative breast cancer. Mol. Cancer 23, 118 (2024).
Pack, L. R., Daigh, L. H., Chung, M. & Meyer, T. Clinical CDK4/6 inhibitors induce selective and immediate dissociation of p21 from cyclin D-CDK4 to inhibit CDK2. Nat. Commun. 12, 3356 (2021).
Gartel, A. L. et al. p21 expression is induced in newly nondividing cells in diverse epithelial and during differentiation of the Caco-2 intestinal cell line. Exp. Cell Res. 227, 171–181 (1996).
Rossiello, F., Herbig, U., Longhese, M. P., Fumagalli, M. & d’Adda di Fagagna, F. Irreparable telomeric DNA damage and persistent DDR signalling as a shared causative mechanism of cellular senescence and ageing. Curr. Opin. Genet Dev. 26, 89–95 (2014).
Kumari, R. & Jat, P. Mechanisms of cellular senescence: cell cycle arrest and senescence associated secretory phenotype. Front Cell Dev. Biol. 9, 645593 (2021).
Quaas, M., Muller, G. A. & Engeland, K. p53 can repress transcription of cell cycle genes through a p21(WAF1/CIP1)-dependent switch from MMB to DREAM protein complex binding at CHR promoter elements. Cell Cycle 11, 4661–4672 (2012).
Dommer, A. P. et al. Tumor Suppressors Condition Differential Responses to the Selective CDK2 Inhibitor BLU-222. Cancer Res. 85, 1310–1326 (2025).
Lyu, J. et al. Synthetic lethality of RB1 and aurora A is driven by stathmin-mediated disruption of microtubule dynamics. Nat. Commun. 11, 5105 (2020).
Spencer, S. L. et al. The proliferation-quiescence decision is controlled by a bifurcation in CDK2 activity at mitotic exit. Cell 155, 369–383 (2013).
Li, M. et al. Low–molecular weight cyclin E confers a vulnerability to PKMYT1 inhibition in triple-negative breast cancer. Cancer Res. 84, 3864–3880 (2024).
Luo, L. J. et al. Combination treatment with CDK2 inhibitor (BLU-222) and either palbociclib or ribociclib is synergistic in pre-clinical models of CDK4/6 inhibitor-resistant breast cancer. Cancer Res. 84 https://doi.org/10.1158/1538-7445.Sabcs23-Po1-18-05 (2024).
Luo, L. J. et al. Anti-tumor activity of CDK2 inhibitor BLU-222 in combination with CDK4/6 inhibitors for overcoming resistance in HR positive and triple negative metastatic breast cancers models. Cancer Res. 84 https://doi.org/10.1158/1538-7445.Am2024-4622 (2023).
Brown, V. et al. CDK2 inhibition with BLU-222 in combination with ribociclib demonstrates robust antitumor activity in pre-clinical models of CDK4/6 inhibitor-naive and -resistant HR+/HER2-breast cancer. Cancer Res. 83 https://doi.org/10.1158/1538-7445.Sabcs22-P6-10-07 (2023).
Fan, H. et al. DNA damage induced by CDK4 and CDK6 blockade triggers anti-tumor immune responses through cGAS-STING pathway. Commun. Biol. 6, 1041 (2023).
Karakas, C. et al. Cytoplasmic cyclin E expression predicts for response to neoadjuvant chemotherapy in breast cancer. Ann. Surg. 274, e150–e159 (2021).
Yap, T. A. et al. First-in-human phase 1/2a study of a potent and novel CDK2-selective inhibitor PF-07104091 in patients (pts) with advanced solid tumors, enriched for CDK4/6 inhibitor resistant HR+/HER2-breast cancer. J. Clin. Oncol. 41, 3010 (2023).
Shen, C. et al. PF-07104091, a first-in-class CDK2-selective inhibitor for the treatment of HR+/HER2-breast cancer and CCNE1high ovarian cancer. Cancer Res. 84 https://doi.org/10.1158/1538-7445.Am2024-5709 (2023).
Nguyen, T. D. T. et al. Sequential targeting of retinoblastoma and DNA synthesis pathways is a therapeutic strategy for sarcomas that can be monitored in real time. Cancer Res 83, 939–955 (2023).
Li, M. et al. Low-molecular-weight cyclin E deregulates DNA replication and damage repair to promote genomic instability in breast cancer. Oncogene 41, 5331–5346 (2022).
Akli, S. et al. Tumor-specific low molecular weight forms of cyclin E induce genomic instability and resistance to p21, p27, and antiestrogens in breast cancer. Cancer Res. 64, 3198–3208 (2004).
Allalou, A. & Wahlby, C. BlobFinder, a tool for fluorescence microscopy image cytometry. Comput Methods Prog. Biomed. 94, 58–65 (2009).
Davan-Wetton, C. S., & Montero-Melendez, T. An optimised protocol for the detection of lipofuscin, a versatile and quantifiable marker of cellular senescence. Plos one 19, e0306275 (2024).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Frankish, A. et al. Gencode 2021. Nucleic Acids Res. 49, D916–D923 (2021).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 14, 7 (2013).
Fabregat, A. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 46, D649–D655 (2018).
Casella, G. et al. Transcriptome signature of cellular senescence. Nucleic Acids Res. 47, 11476 (2019).
Fridman, A. L. & Tainsky, M. A. Critical pathways in cellular senescence and immortalization revealed by gene expression profiling. Oncogene 27, 5975–5987 (2008).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652–2660.e2654 (2017).
Purcell, M., Kruger, A. & Tainsky, M. A. Gene expression profiling of replicative and induced senescence. Cell Cycle 13, 3927–3937 (2014).
Engeland, K. Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ. 25, 114–132 (2018).
Uxa, S. et al. DREAM and RB cooperate to induce gene repression and cell-cycle arrest in response to p53 activation. Nucleic Acids Res 47, 9087–9103 (2019).
Acknowledgements
We sincerely thank Dr. Joseph Marszalek, director of Translational Research to Advance Therapeutics and Innovation in Oncology (TRACTION) at MDACC, for supporting the project; and Dr. Scott Ribich, Dr. Nealia House, Dr. Lakshmi Muthuswamy, and Dr. Mikael Rinne from Blueprint Medicines for their critical review of the manuscript. This work was supported by NIH grant P30CA016672 to The University of Texas MD Anderson Cancer Center, NIH/NCI grants CA255960 and CA223772 (K.K.), Cancer Prevention and Research Institute of Texas (CPRIT) Multi-Investigator grant RP180712 (K.K.H. and K.K.), and Blueprint Medicines (K.K.). C.W.U. is supported by the NIH 1R25CA240137-01A1 UPWARDS Training Program (Undergraduate Students Working Towards Research in Science) and by the CPRIT Research Training Award CPRIT Training Program (RP210028).
Author information
Authors and Affiliations
Contributions
L.L. and K.K. conceived the study, designed the experiments, and wrote the manuscript with input from all authors. L.L. performed the majority of the experiments and data analysis, with the following contributions: Y.W. assisted with IHC and SBB assays; T.B. assisted with sample processing, western blotting, and co-IP assays; X.J. assisted with in vivo treatment studies; M.-K.C. assisted with PLA assays; S.M. assisted with co-IP assays; M.L. assisted with in vitro treatments and western blots; and S.K., R.Y.K., S.K.A., and C.W.U. assisted with cell culture and combination in vitro assays. X.R. and S.T. performed bioinformatic analyses and assisted with statistical modeling and data interpretation. J.N.-Y. and N.M.K. provided technical assistance and experimental support. F.M.-B., K.K.H., J.W., H.W., and K.F. provided critical resources, clinical expertise, and guidance on study design. K.K. and K.K.H. acquired funding and supervised all aspects of the project. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The following authors have relationships with the companies listed: F Meric-Bernstam: AbbVie, Aduro BioTech, Aileron Therapeutics, Alkermes, AstraZeneca, Bayer Healthcare Pharmaceutical, Black Diamond, Biovica, Calithera Biosciences, Chugai Pharmaceutical, Curis, CytomX Therapeutics, Daiichi Sankyo, Debiopharm International, Ecor1 Capital, eFFECTOR Therapeutics, Eisai, European Organization for Research and Treatment of Cancer (EORTC), European Society for Medical Oncology (ESMO), F. Hoffmann-La Roche, FogPharma, Genentech, GT Apeiron, Guardant Health, Harbinger Health, IBM Watson, Immunomedics, Infinity Pharmaceuticals, Inflection Biosciences, Jackson Laboratory, Karyopharm Therapeutics, Klus Pharma, Kolon Life Science, Lengo Therapeutics, Loxo Oncology, Menarini Group, Mersana Therapeutics, Novartis, OnCusp Therapeutics, OrigiMed, PACT Pharma, Parexel International, Pfizer, Protai Bio, Puma Biotechnology, Samsung Bioepis, Sanofi, Seattle Genetics, Silverback Therapeutics, Spectrum Pharmaceuticals, Taiho Pharmaceutical, Takeda Pharmaceutical, Tallac Therapeutics, Tyra Biosciences, Xencor, Zentalis, Zymeworks. KK Hunt: Armada Health, AstraZeneca, Cairn Surgical, Eli Lilly & Co, Lumicell. S Damodaran: EMD Serono, Guardant Health, Novartis, Pfizer, Sermonix, Taiho. K Faia: Blueprint Medicines. K Keyomarsi: Apeiron, Blueprint Medicines, REPARE, Schrodinger, Novartis. All other authors declare no conflicts of interest.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, L., Wang, Y., Bui, T. et al. CDK2 inhibitor BLU-222 synergizes with CDK4/6 inhibitors in drug resistant breast cancers through p21/p27 induction. Nat Commun 17, 619 (2026). https://doi.org/10.1038/s41467-025-67865-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67865-4