Abstract
Tumor-associated macrophages (TAMs) constitute a pivotal cellular component within the intricate tumor microenvironment (TME). However, the relationship between the N6-methyladenosine (m6A) and metabolic pattern of TAMs remains poorly understood. Here we show that the m6A methyltransferase METTL3 is conspicuously downregulated in monocyte-derived macrophages from glioblastoma (GBM) patients. Conditional knockout of Mettl3 in myeloid cells augments lipid metabolism and accelerates glioma progression. Furthermore, we found that METTL3 deficiency facilitates fatty acid synthase (FASN) expression, while compromising CD8+ T cell response. Mechanistically, ISG15 is highly expressed in METTL3-deficient macrophages. ISG15 interacts with FASN and regulates FASN ISGylation and its stability through impeding FASN ubiquitination. Notably, the suppression of ISG15 in METTL3-deficient macrophages reverses the enhanced FASN expression and restores CD8+ T cell functionality. Furthermore, we demonstrate that FASN knockout or FASN inhibitor treatment in myeloid cells abrogates the exaggerated tumor progression triggered by METTL3 knockout. Collectively, this study highlights the pivotal role of m6A in regulating macrophage metabolism and identifies potential targets in controlling GBM progression.
Similar content being viewed by others
Introduction
Glioblastoma (GBM) is the most common and aggressive type of primary adult malignant tumor in the central nervous system1. Owing to tumor microenvironment (TME) heterogeneity, GBM is a challenging cancer to treat2. Among the TME components, tumor-associated macrophages are the most abundant population of immune cells, comprising a staggering 50% of all viable cells in the whole GBM tumor mass3. Emerging evidence demonstrates that TAMs are critical for promoting tumor progression and orchestrating an immunosuppressive landscape in GBM4. However, therapeutic strategies for depleting TAMs have not been well translated into the clinic5, suggesting that our understanding of this cell population is still limited.
The TAM compartment is highly dynamic and heterogeneous, both within individual tumors and across distinct tumor types6,7. In recent years, cellular metabolism has emerged as a key regulator for macrophage activation. Notably, an elevated degree of TAM metabolism heterogeneity has indeed been revealed not merely among diverse cancer patients but also within the various malignant lesions of a single patient, as well as within a specific neoplastic lesion8,9. The TAM compartment evolves over time as well as in space through an extensive remodeling of core energy metabolism10. Once macrophages are recruited into the TME, they have to be reprogrammed in the metabolic pathways in order to adapt to the special environment11. Among the metabolic alterations in TAMs, lipid metabolism has been associated with the tumoricidal capability12. Noteworthily, an increased lipid accumulation in TAMs is positively and significantly associated with the identification of hypoxic macrophages in GBM with therapeutic potential for vasculature normalization9,13. The reprogrammed lipid metabolism not only determines their secretory profile, but also further influences the immune status of TME and ultimately the progression of tumors. Thus, elucidating lipid metabolic reprogramming-associated signaling pathways in TAMs holds significant promise for the development of other therapeutic targets in cancer treatment.
In recent years, N6-methyladenosine (m6A), the most diverse and reversible post-transcriptional modification of eukaryotic messenger RNAs (mRNAs), has been recognized as a strong regulator of mRNA splicing, stability, localization and translation14,15. This intricate process is orchestrated by the MTases complex (writer), demethylases (erasers), and mRNA-binding proteins (readers). The writer proteins act as methyltransferases for m6A formation in cells and mainly include METTL316, METTL1417, METTL1618, WTAP19, VIRMA20, RBM15/15B21, ZC3H1322,23, and HAKAI20. Conversely, the erasers (demethylases: FTO and ALKBH5) remove m6A modification from RNA molecules24,25. Meanwhile, the readers (including YTHDF1/2/3, YTHDC1/2, IGF2BPs, FMR1, and HNRNPA2B1) specifically recognize m6A-modified RNA25, influencing its fate through mechanisms such as degradation, stabilization, or modulation of translation26. m6A has been regarded as a major regulator on various tumor-related processes, including proliferation, apoptosis, migration, invasion, energy processing, and response to radiotherapy. However, research on m6A modification in tumors is still in its infancy, the unknown participants still need to be further explored and identified.
In the study, we demonstrated that Mettl3 knockout in myeloid cells significantly accelerates glioma progression. Further analysis implied that METTL3 deficiency facilitates ISG15-mediated upregulation of FASN expression and subsequent alterations in lipid metabolism. Mechanistically, we identified an interaction between ISG15 and FASN, which modulates the ISGylation status and stability of FASN. Furthermore, we showed that either the knockout of FASN or the treatment of myeloid cells with a FASN inhibitor effectively abolishes the enhanced tumor progression phenotype observed in mice with METTL3 depletion, underscoring the vital role of this pathway in glioma progression.
Results
Lower expression of METTL3 in TAMs derived from GBM
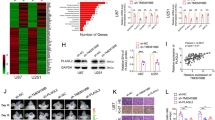
The GBM microenvironment is highly heterogeneity and employ huge of TAMs. To investigate the m6A modification in TAMs involved during GBM progression, we explored the TCGA datasets and discovered that the expression of METTL3 was diminished in mesenchymal subtype with a poorer prognosis and strong radiation resistance than other glioma subtypes (Supplementary Fig. 1a). Analysis of public single-cell RNA-seq (scRNA-seq) data indicated that the expression of METTL3 in TAMs was lower in high-grade gliomas than that in low-grade gliomas (Fig. 1a). Using expression data (ESTIMATE) algorithm27, which calculates the abundance of stromal and immune cells in malignant tumor tissues based on expression data, we found a negative correlation between the expression of METTL3 mRNA and immune scores in GBM (Supplementary Fig. 1b). By leveraging the TIMER 2.0 website, we found that the expression of METTL3 was negatively correlated with the infiltration of TAMs (Supplementary Fig. 1c). We then investigated the relationship of METTL3 in TAMs with tumor progression in mice. The results also showed that METTL3 expression was suppressed in TAMs from GL261 tumor-beared mice compared with PMs from healthy mice by immunofluorescence staining (Fig. 1b). Moreover, western blot analysis also confirmed the downregulation of METTL3 protein expression in TAMs compared with PMs from healthy mice (Fig. 1c). Meanwhile, we also analyzed and demonstrated that the expression of METTL3 in TAMs was markedly decreased in GBM patients compared with microglia from healthy control (Fig. 1d). Meanwhile, we observed a generalized reduction in METTL3 expression among myeloid populations within tumors relative to their circulating counterparts, and a similar down-regulation in tumor-infiltrating microglia compared with microglia in healthy brain tissue (Supplementary Fig. 1d). In summary, our comprehensive data underscore a consistent pattern of inhibited METTL3 expression in TAMs within the context of GBM, highlighting its potential importance in tumor progression.
a The METTL3 expression in TAMs from single cell RNA-seq datasets of GBM. The left panel shows per-cell METTL3 expression and the right panel shows per-sample averages. b F4/80 and METTL3 staining and corresponding quantification in PMs from healthy mice or tumor tissues from GL261-tumor bearing mice on day 21 post-xenograft. Scale bars: 30 μm. (n = 6 independent experiments). c Immunoblotting of METTL3 in PMs from healthy mice or TAMs isolated from tumor tissues of GL261-tumor bearing mice on day 21 post-xenograft. (n = 3 independent experiments). d Representative fluorescence micrographs of brains from the normal and glioma patient, immunolabeled with CD68 and METTL3 antibody. The intensity of the fluorescence signal is quantified. Scale bars: 10 μm. (n = 6 independent experiments). e–g Representative bioluminescence images from Mettl3fl/fl or Mettl3fl/flLyz2Cre mice implanted with GL261 cells on day 21. The intensity of the bioluminescent signal is quantified (e). Representative HE staining images of brain sections (f) are shown. Quantification of the tumor area was performed (g). (n = 6 mice per group). h The survival of mice injected with GL261 tumor cells was documented (n = 8 mice per group). i WT mice were injected with 4 × 104 GL261 cells mixed with 1 × 104 DMSO- or STM2457-pretreated BMDMs. Bioluminescence was measured at the indicated time points and quantified. (n = 10 mice per group). j A schematic presentation of the experiment design for bone marrow transplantation. k, l WT mice were irradiated and reconstituted for 4 weeks with 1 × 107 Mettl3fl/fl or Mettl3fl/flLyz2Cre bone marrow cells and then implanted with 3 × 105 GL261 cells. Mice were sacrificed on day 21 post-tumor implantation. Images of HE-stained brain sections are shown (k). The tumor area was quantified (l). (n = 7 mice per group). m, n 3 × 105 U87 GBM cells were implanted into the right striatum of nude mice for 3 weeks. Mettl3fl/fl or Mettl3fl/flLyz2Cre myeloid cells mixed with lymphocytes were injected into the mice via tail vein every week. The mice were sacrificed for HE staining to monitor the tumor growth after 3 weeks (m). Quantification of the tumor area was performed (n). (n = 7 mice per group). o, p Mettl3fl/fl or Mettl3fl/flLyz2Cre mice implanted with GL261 cells were injected with control or clodronate liposomes. Bioluminescence was measured at the indicated time points and quantified (o). Mice were sacrificed on day 21 post-tumor implantation. Representative HE staining images of brain sections are shown and the tumor area was quantified (p). (n = 7 mice per group). Data are expressed as the mean ± SD. P values were determined by a two-tailed t-test (b, d, e, g, i, l, n), one-way ANOVA with Tukey’s multiple comparisons (o, p) and the Gehan-Breslow-Wilcoxon test (h). The source data are provided as a Source data file.
Mettl3 deficiency in myeloid cells enhances glioma progression
To elucidate the functional role of METTL3 in TAMs involved in glioma progression, we employed Mettl3fl/flLyz2Cre conditional knockout mice and their littermates Mettl3fl/fl in the C57BL/6 genetic background for tumor growth experiments. We firstly confirmed the depletion of METTL3 in BMDMs by western blotting and m6A dot blot (Supplementary Fig. 2a, b). In the syngeneic murine GL261 tumor model, we observed a significant increase in tumor volume in Mettl3fl/flLyz2Cre mice compared to that in Mettl3fl/fl mice (Fig. 1e and Supplementary Fig. 2c). HE and ki67 staining also demonstrated that GL261 tumor cell proliferated more rapidly in Mettl3fl/flLyz2Cre mice than that in Mettl3fl/fl mice (Fig. 1f, g and Supplementary Fig. 2d). The survival of these conditional Mettl3 knockout mice with GL261 tumor was shortened (Fig. 1h). Further analysis showed that co-injection of GL261 cells with STM2457-treated BMDMs resulted in significantly accelerated tumor growth (Fig. 1i). To further validate the pathological role of METTL3 in GBM, we reconstituted irradiated Wild Type (WT) mice with total bone marrow-derived cells from Mettl3fl/flLyz2Cre or Mettl3fl/fl mice, and analyzed the contributions of bone marrow-derived macrophages in the chimaeras after reconstitution. The results showed that chimeric mice lacking Mettl3 exhibited an elevated tumor burden (Fig. 1j–l). Moreover, we employed a patient-derived xenograft model of GBM and discovered that Mettl3fl/flLyz2Cre myeloid cells could enhance U87 tumor growth (Fig. 1m, n). To further confirm whether macrophages with METTL3 depletion are required for the increased tumor burden in Mettl3fl/flLyz2Cre mice, the mice were administered an intravenous injection of clodronate liposome 24 h prior to tumor induction to destroy microglia and macrophages (Supplementary Fig. 2e, f). The results showed that clodronate-treated mice were unresponsive to Mettl3 depletion, relative to control mice, as characterized by an indistinguishable influence on tumor growth (Fig. 1o, p). In addition, we found that treatment with the CCR2 antagonist RS504393 abrogated the difference in tumor growth between Mettl3fl/fl or Mettl3fl/flLyz2Cre mice (Supplementary Fig. 3), indicating that the phenotype depends on monocyte-derived macrophages. Collectively, these results imply that Mettl3 ablation in TAMs promotes glioma progression, thereby emphasizing the importance of METTL3 in modulating macrophage-mediated tumor suppression.
Knockout of Mettl3 in myeloid cells inhibited CD8+IFN-γ+ cell formation
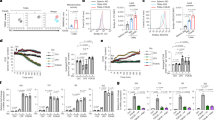
To determine whether the myeloid deficiency of METTL3 reshapes tumor immune microenvironment, we conducted a thorough analysis of the frequency of tumor-infiltrating immune cells. The results showed no apparent differences in the total number of macrophage between Mettl3fl/flLyz2Cre and Mettl3fl/fl mice (Fig. 2a and Supplementary Fig. 4a, b). However, we observed that the frequency of M1 macrophages (F4/80+NOS2+) and M2 macrophage (F4/80+CD206+ or F4/80+ARG1+), was increased in METTL3 deficient mice (Fig. 2a and Supplementary Fig. 4a, b). Moreover, immunofluorescence staining also demonstrated that CD206 and ARG1 exhibited increased expression in METTL3 knockout macrophages compared with that in wild-type cells (Supplementary Fig. 4c). We further investigated the effects of METTL3 deficient macrophage on other tumor infiltrating immune cells. Most of the immune-suppressive cell populations, including myeloid-derived suppressor cells (MDSCs) and neutrophils (PMN-MDSCs) exhibited increased cell numbers in tumors from METTL3-deficient mice relative to wild-type mice (Fig. 2b and Supplementary Fig. 4d). In contrast, the presence of anti-tumor CD8+ T cells was diminished in these tumors (Fig. 2c and Supplementary Fig. 4e). In addition, we examined the CD8+ cell functionality and observed the decreased numbers of CD8+IFN-γ+ and CD8+Granzyme B+ cells and the increased number of CD8+PD-1+, CD8+LAG-3+, CD8+TIM-3+ and CD8+CD39+ cells (Fig. 2d, e). In vitro coculture experiments also exhibited that METTL3-deficient BMDMs could inhibit the function of CD8+ tumor killer cells as demonstrated by reduced expression of IFN-γ (Fig. 2f, g). These results indicate that METTL3 knockout substantially remodels the TME from anti-tumor to pro-tumor by altering the composition and function of key immune cell populations.
a Quantification of total, M1 or M2 TAMs in tumor from Mettl3fl/fl or Mettl3fl/flLyz2Cre mice implanted with GL261 cells on day 21. (n = 5 mice per group). b Quantification of M-MDSCs and PMN-MDSCs in tumor from Mettl3fl/fl or Mettl3fl/flLyz2Cre mice implanted with GL261 cells on day 21. (n = 5 mice per group). c Quantification of T cells and NKT cells in tumor from Mettl3fl/fl or Mettl3fl/flLyz2Cre mice implanted with GL261 cells on day 21. (n = 5 mice per group). d, e Quantification of CD8+IFN-γ+, CD8+Granzyme B+, CD8+PD1+, CD8+LAG3+, CD8+TIM-3+ and CD8+CD39+ T cells in tumor from Mettl3fl/fl or Mettl3fl/flLyz2Cre mice implanted with GL261 cells on day 21. (n = 5 mice per group). f A schematic presentation of the experiment design for co-cultivation system. g T cell suppression by Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs co-cultured with lymphocytes at a ratio of 1:1 or 1:3. (n = 3 independent experiments). Data are expressed as the mean ± SD. P values were determined by a two-tailed t-test (a, b, c, d, e) and one-way ANOVA with Tukey’s multiple comparisons (g). The source data are provided as a Source data file.
Knockout of Mettl3 in myeloid cells enhances lipid metabolism and fatty acid synthase (FASN) expression
TAMs and their impact on the overall metabolic profile of the TME have a major influence on tumor progression8. To explore whether METTL3 ablation influences the metabolic profile of macrophage, we conducted an unbiased metabolomic analysis comparing Mettl3fl/flLyz2Cre with their Mettl3fl/fl BMDMs counterparts. Metabolite analysis revealed 254 metabolites that were differentially regulated (125 upregulated, 129 downregulated) between Mettl3fl/flLyz2Cre and Mettl3fl/fl BMDMs (Fig. 3a). Enrichment analysis revealed that the changed metabolites were predominantly lipid and lipid like moleculars (Fig. 3b and Supplementary Fig. 5), suggesting that METTL3 participated in lipid metabolism pathway. So we tested the protein expression of key proteins related to lipid metabolism in BMDMs, and confirmed that FASN expression was significantly increased in Mettl3fl/flLyz2Cre BMDMs (Fig. 3c), accompanied with elevated lipid accumulation and expression of lipid laden macrophage-related gene Gpnmb (Fig. 3d, e). The gene expression analysis clearly revealed that FASN downstream genes involved in fatty acid elongation were significantly increased in Mettl3fl/flLyz2Cre BMDMs (Fig. 3f). However, qRT-PCR analysis showed that the expression of Fasn mRNA was unchanged (Fig. 3g). The mRNA decay assays also demonstrated that METTL3 had no effect on the stability of Fasn mRNA (Fig. 3h). Integrative genomics viewer (IGV) analysis showed that m6A peaks on the transcript of Fasn mRNAs remained unchanged in Mettl3fl/flLyz2Cre BMDMs (Supplementary Fig. 6a). These results implied that FASN was not directly regulated by METTL3-mediated modification. Previous studies have shown that METTL3 can bind to the gene promoters and influence their expression28. Our results indicated an absence of direct METTL3 binding to the Fasn gene promoter region (Supplementary Fig. 6b), further supporting the notion that METTL3 does not directly regulate FASN expression. Next, we reanalyzed the genes expression profiles in the single cell sequencing data GSE15915629 and discovered that 3 genes (ISG15, Lyz1, Plac8) were obviously upregulated in Mettl3fl/flLyz2Cre TAMs compared with Mettl3fl/fl TAMs (Fig. 3i). qRT-PCR results demonstrated that the expression of Isg15 was noticeably increased in the Mettl3fl/flLyz2Cre BMDMs, while the expression of Lyz1 and Plac8 remained unaffected (Fig. 3j). Western blot analysis also verified the increased expression of ISG15 in Mettl3fl/flLyz2Cre BMDMs (Fig. 3k). We next investigated the potential direct regulation of ISG15 by METTL3. However, IGV analysis showed that m6A peaks on the transcript of Isg15 remained unaltered in Mettl3fl/flLyz2Cre BMDMs (Supplementary Fig. 6c). In addition, METTL3 could not directly bind to the promoter region of Isg15 (Supplementary Fig. 6d). Given that ISG15 is an IFN-inducible protein30, we explored this avenue and found that IFN-β was upregulated in Mettl3fl/flLyz2Cre BMDMs and TAMs (Fig. 3l), aligning with previous reports of enhanced type I IFN expression following METTL3 ablation in monocytes31. In addition, we found that Mettl14 knockdown in BMDMs also robustly increased Ifnb1 expression (Fig. 3m). meRIP-qPCR assays further confirmed that Ifnb1 mRNA is indeed m⁶A-modified (Fig. 3n, o). Absence of Mettl3 enhanced the stability of Ifnb1 mRNAs (Fig. 3p). Furthermore, we demonstrated that IFN-β directly stimulated the expression of both ISG15 and FASN (Fig. 3q), while blocking IFN-β could abrogate the effects of METTL3 on FASN expression (Fig. 3r). In addition, treatment with IFN-β robustly elevated FASN levels in Mettl3fl/fl and Mettl3fl/flLyz2Cre BMDMs (Fig. 3s). These results suggested that METTL3 may regulate FASN expression and lipid metabolism through IFN-β/ISG15 axis.
a Scatter plots analysis of differential metabolites in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. b Heatmap analysis of differential metabolite in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. c Western blot analysis of the indicated proteins in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. (n = 3 replicates, each replicate includes 3 mice). d Nile Red staining and quantification were performed in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. Scale bars: 30 μm. (n = 4 independent experiments). e qRT-PCR analysis of Gpnmb expression in Mettl3fl/fl or Mettl3fl/flLyz2Cre TAMs isolated from tumor-bearing mice. (n = 3 biological replicates). f qRT-PCR analysis of lipogenic gene expression in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. (n = 3 biological replicates). g qRT-PCR analysis of Fasn expression in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. (n = 3 biological replicates). h Fasn mRNA stability in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs treated with actinomycin D at the indicated times was determined by qRT-PCR. (n = 3 biological replicates). i Scatter plots showing genes expression changes in cellular transcript levels in Mettl3fl/flLyz2Cre TAMs versus Mettl3fl/fl TAMs. j qRT-PCR analysis of genes expression in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. (n = 3 biological replicates). k Western blot analysis of the indicated proteins in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs. (n = 3 independent experiments). l qRT-PCR analysis of genes expression in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs and TAMs. (n = 3 biological replicates). m qRT-PCR analysis of genes expression in Mettl14 knockdown BMDMs. (n = 3 biological replicates). n m6A enrichment of Ifnb1 mRNA in BMDMs by m6A-RIP-qPCR. (n = 3 biological replicates). o m6A enrichment of Ifnb1 mRNA in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs by m6A-RIP-qPCR. (n = 3 biological replicates). p Ifnb1 mRNA stability in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs treated with actinomycin D at the indicated times determined by qRT-PCR. (n = 3 biological replicates). q Western blot analysis of the indicated proteins in BMDMs and HEK293T cells. (n = 3 independent experiments). r Immunoblotting analysis of the indicated proteins in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs transfected with NC siRNA or Ifnb siRNA. (n = 3 independent experiments). s Immunoblotting analysis of the indicated proteins in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs treated with IFN-β protein. (n = 3 independent experiments). Data are expressed as the mean ± SD. P values were determined by a two-tailed t-test (a, d, e, f, g, h, j, l, n, o, p) and one-way ANOVA with Tukey’s multiple comparisons (m). The source data are provided as a Source data file.
ISG15 interacts with FASN and regulates FASN ISGylation and stability through inhibiting FASN ubiquitination
To explore intricate regulatory mechanisms by which ISG15 modulates FASN expression. we searched the BioGRID (https://thebiogrid.org/) websites and found that ISG15 is a potential interacted protein of FASN32,33. To test the interactions between FASN and ISG15, reciprocal coimmunoprecipitation assays were performed. The results showed that endogenous FASN co-immunoprecipitated with ISG15, which was verified by coimmunoprecipitation of Myc-tagged ISG15 with endogenous FASN from transiently transfected HEK293T cells (Fig. 4a). Moreover, ISG15 and FASN that were epitope tagged and expressed in HEK293T cells were also able to interact with each other in co-immunoprecipitation experiments (Fig. 4b). Co-immunoprecipitation assays further confirmed endogenous ISG15 and FASN interaction in BMDMs (Fig. 4c). In vitro His pull-down assay also further definitively confirmed the direct interaction between FASN and ISG15 (Fig. 4d). To further corroborate this result, we performed immunofluorescence staining and observed strong colocalization of endogenous ISG15 and FASN in BMDMs (Fig. 4e). Ectopic expression of FLAG-FASN was also associated with Myc-ISG15 in 293T cells, providing compelling evidence that ISG15 is indeed in a protein complex with FASN (Fig. 4e). These data demonstrated that ISG15 indeed interacted with FASN. We then investigated the mechanism governing how ISG15 regulates FASN. We tested the dependence of FASN expression on ISG15. The results showed that ISG15 knockdown reduced FASN expression (Fig. 4f). Reversely, ISG15 overexpression enhanced FASN expression (Fig. 4g). Re-expression of ISG15 in ISG15 knockdown BMDMs reversed the reduced expression of FASN (Fig. 4f). Furthermore, we assessed the stability of FASN and observed the accelerated decay of FASN in BMDMs lacking ISG15 after cycloheximide (CHX) treatment (Fig. 4h). Re-expression of ISG15 restored the FASN protein stability in ISG15 knockdown BMDMs (Fig. 4h). Given that ISG15 is known to modify cellular proteins through a process called ISGylation30, we next searched the potential ISGylation sites of FASN protein by analyzing the ISGylome data previously published34. The results showed that there are 4 potential ISGylation sites in FASN protein (Fig. 4i). We then mutated these Lys (K) residues to Arg (R) residues and examined the mutant FASN expression. The results demonstrated that mutations at Lys 1582, Lys 1591 and Lys 1866 inhibited FASN protein expression (Fig. 4j), whereas the mRNA levels of FASN mutants were unchanged significantly (Fig. 4k). Co-immunoprecipitation assays further demonstrated that mutations at Lys 1582, Lys 1591 and Lys 1866 inhibited FASN and ISG15 interaction (Fig. 4l). The results indicated that Lys 1582, Lys 1591 and Lys 1866 truly served for FASN protein expression. We then investigated whether ISGylation affected the ubiquitination status of FASN. As shown in Fig. 5a, mutation at Lys 1582, Lys 1591 and Lys 1866 inhibited FASN ISGylation, but enhanced its ubiquitination. Furthermore, we demonstrated that inhibition of ISG15 led to a marked increase in FASN ubiquitination (Fig. 5b), while ISG15 overexpression impeded FASN ubiquitination (Fig. 5c). Notably, the ISG15 mutant lacking the C-terminal glycine residues (ISG15 △GG) could not inhibit FASN ubiquitination effectively (Fig. 5c). In addition, we demonstrated that the K48-linked ubiquitination of FASN was obviously downregulated upon ISG15 overexpression (Fig. 5d). These findings collectively suggest that ISG15 and ISGylation play a crucial role in regulating the ubiquitination-mediated degradation of FASN, thereby modulating its expression and stability.
a Co-immunoprecipitation to detect the interaction of Myc-ISG15 with endogenous FASN in HEK293T cells. (n = 3 independent experiments). b Co-immunoprecipitation to detect the interaction of Myc-ISG15 with FLAG-FASN in HEK293T cells. (n = 3 independent experiments). c Co-immunoprecipitation to detect the interaction of endogenous ISG15 and FASN in BMDMs. (n = 3 independent experiments). d In vitro His-tag pull-down was used to test the interaction of FASN and ISG15. (n = 3 independent experiments). e Representative images of endogenous FASN and ISG15 or FLAG-FASN and Myc-ISG15 distribution in HEK293T cells. Colocalization was quantitatively assessed. Scale bars: 10 μm. (n = 3 independent experiments). f BMDMs transfected with NC siRNA, Isg15 siRNAs or Isg15 siRNAs and Myc-ISG15 for 48 h were lysed, followed by immunoblotting with the indicated antibodies. (n = 3 independent experiments). g FLAG-FASN was co-expressed with or without Myc-ISG15 in HEK293T cells. Cell lysates were subjected to immunoblotting with the indicated antibodies. (n = 3 independent experiments). h Western blot to analyze FASN expression after ISG15 silence with or without Myc-ISG15 overexpression combined with 100 ug/ml cycloheximide (CHX) at the 0, 3, 6, and 9 h in BMDMs. (n = 3 independent experiments). i Schematic diagram showing the identified ISGylated lysine residues of FASN. j Western blot analysis of protein levels of FLAG-FASN mutants in 293 T cells. (n = 3 independent experiments). k qRT-PCR analysis of protein level of FASN mutants in 293T cells. (n = 3 independent experiments). l Co-immunoprecipitation to detect the interaction of Myc-ISG15 with FLAG-FASN mutants in human macrophage cells THP1. (n = 3 independent experiments). Data are expressed as the mean ± SD. The source data are provided as a Source data file.
a FLAG-FASN mutants along with HA-ubiquitin were co-expressed in THP1 cells. Cell lysates and immunoprecipitates were detected with indicated antibodies. (n = 3 independent experiments). b FLAG-FASN mutants co-expressed with HA-ubiquitin in ISG15 silenced THP1 cells treated with MG132. Cell lysates and immunoprecipitates were examined with indicated antibodies. (n = 3 independent experiments). c FLAG-FASN mutants, HA-ubiquitin and Myc-ISG15 or Myc-ISG15ΔGG were co-expressed in THP1 cells treated with MG132. Cell lysates and immunoprecipitates were detected with indicated antibodies. (n = 3 independent experiments). d FLAG-FASN mutants, HA-ubiquitin and Myc-ISG15 or Myc-ISG15ΔGG were co-expressed in THP1 cells treated with MG132. Cell lysates and immunoprecipitates were examined with indicated antibodies. (n = 3 independent experiments). The source data are provided as a Source data file.
The aforementioned data convincingly indicated that FASN was precisely modulated by ISG15, and METTL3 could inhibit ISG15 expression. To investigate whether ISG15 was a critical regulator for increased expression of FASN induced by METTL3 deficiency, we performed a rescue experiment to test the effect of ISG15 inhibition on FASN expression in METTL3-deficient BMDMs. The results showed that ISG15 knockdown inhibited the increased expression of FASN protein and FASN ISGylation while concurrently intensifying its ubiquitination upon METTL3 ablation (Fig. 6a). CHX assay also demonstrated that ISG15 inhibition abrogated the effect of METTL3 deficiency on FASN protein stability (Fig. 6b). ISG15 knockdown inhibited the M1-associated genes and the M2-associated genes expression in Mettl3fl/flLyz2Cre BMDMs (Fig. 6c). Notably, lipid accumulation was also significantly inhibited by ISG15 knockdown in Mettl3fl/flLyz2Cre BMDMs (Fig. 6d). Our previous results demonstrated that METTL3 deficiency in macrophages inhibited CD8+IFN-γ+ cell differentiation (Fig. 2d, g). We also tested the role of ISG15 in CD8+IFN-γ+ cell differentiation regulated by METTL3 ablation. The results displayed that ISG15 knockdown abrogated the inhibitory effect of METTL3 ablation on CD8+IFN-γ+ cell differentiation (Fig. 6e). Moreover, tumors from mice receiving CpG-siRNAIsg15 exhibited a markedly increased fraction of IFN-γ-producing CD8+ T cells compared with those from control-treated animals (Fig. 6f). Silencing ISG15 by targeted delivery of CpG-siRNAIsg15 to TAMs also reduced the expression of GPNMB (Fig. 6g). In summary, these data suggested that METTL3 regulated FASN expression and lipid metabolism through ISG15 during macrophage activation.
a Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs treated with NC siRNA or Isg15 siRNAs for 48 h were lysed. Cell lysates and immunoprecipitates were examined with indicated antibodies. (n = 3 independent experiments). b Western blot to analyze FASN expression in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs after ISG15 silence combined with 100 ug/ml cycloheximide (CHX) at the 0, 3, 6 and 9 h. (n = 3 independent experiments). c qRT-PCR analysis of genes expression in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs transfected with Isg15 siRNAs. (n = 3 biological replicates). d Nile Red staining and quantification were performed in Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs transfected with siNC or siIsg15. Scale bars: 30 μm. (n = 6 independent experiments). e T cell suppression by Mettl3fl/fl or Mettl3fl/flLyz2Cre BMDMs with ISG15 knockdown co-cultured with Lymphocytes at a ratio of 1:1. (n = 3 independent experiments). f, g Mice were engrafted with GL261 tumor cells and then injected with 5 mg/kg CpG-siRNAIsg15 every other day beginning on day 7 post-inoculation. Percentages of tumor-infiltrating IFN-γ-producing CD8+ T cells on day 21 were calculated (f). The expression of Gpnmb expression in TAMs were examined by qRT-PCR (g). (n = 5 biological replicates). Data are expressed as the mean ± SD. P values were determined by one-way ANOVA with Tukey’s multiple comparisons (c, d, e, f, g). The source data are provided as a Source data file.
FASN knockout or FASN inhibitor treatment abrogates the enhanced tumor progression induced by METTL3 knockout
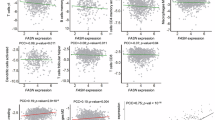
To investigate whether FASN was a key regulator for enhanced tumor progression induced by METTL3 deficiency, we generated double knockout mice for METTL3 and FASN specifically in myeloid cells. Utilizing immunoblotting, we confirmed the successful depletion of METTL3 and FASN in BMDMs in Mettl3fl/flFasnfl/flLyz2Cre mice (Fig. 7a). Then the GL261 cells were injected into the brain of WT, Mettl3fl/flLyz2Cre and Mettl3fl/flFasnfl/flLyz2Cre mice. The results displayed that METTL3 and FASN double knockout inhibited the increased tumor growth observed in METTL3-deficient mice (Fig. 7b, c). Immunofluorescence staining also showed that METTL3 and FASN double knockout hindered M2 macrophage polarization caused by METTL3 deficiency, as demonstrated by reduced expression of CD206 (Supplementary Fig. 7a). Moreover, Nile red staining revealed that the macrophage lipid content was also limited to a lower level by FASN knockout in Mettl3fl/flLyz2Cre BMDMs (Supplementary Fig. 7b). In vitro experiments exhibited that Mettl3fl/flFasnfl/flLyz2Cre BMDMs enhanced the CD8+IFN-γ+ cell differentiation compared with Mettl3fl/flLyz2Cre BMDMs (Fig. 7d). In addition, we observed that Mettl3fl/flLyz2Cre BMDMs with FASN knockdown increased IFN-γ production in CD8+ T cells, contrasting sharply with those co-cultured with METTL3 deficient BMDMs (Supplementary Fig. 7c, d). FASN knockdown inhibited the M1-associated genes and the M2-associated genes expression in Mettl3fl/flLyz2Cre BMDMs (Supplementary Fig. 7e). The lipid content analysis further confirmed the suppressive effect of FASN knockdown on lipid accumulation in Mettl3fl/flLyz2Cre BMDMs (Supplementary Fig. 7f). To confirm the role of FASN in tumor progression in Mettl3fl/flLyz2Cre mice, we also examined the tumor progression in Mettl3fl/fl and Mettl3fl/flLyz2Cre mice treated with a well-proved FASN inhibitor C75. The results showed that pharmacological inhibition of FASN with C75 largely abolished the aggravative effect of METTL3 depletion on tumor progression (Fig. 7e–g). Consistently, C75 treatment impeded F4/80+CD206+ M2 macrophage polarization and lipid accumulation associated with METTL3 depletion (Supplementary Fig. 8a, b). The reduced IFN-γ production in CD8+ T cells co-cultured with METTL3 deficient BMDMs was also significantly reversed upon FASN inhibition (Fig. 7h and Supplementary Fig. 8c). These data suggested that the effects of METTL3 on tumor growth are largely dependent on FASN expression. Clinical specimens analysis also showed that the glioma patients who had lower METTL3 expression in TAMs tended to have higher levels of ISG15 and FASN in tumor section, consistent with the results obtained in the study (Fig. 7i). To interrogate the prognostic impact of METTL3 expression within the tumor-associated macrophage compartment, we performed immunofluorescence co-staining for CD68 and METTL3 on tissue microarrays. Quantitative image analysis revealed that low METTL3 levels in CD68⁺ macrophages are strongly and independently associated with diminished overall survival (Fig. 7j). In summary, we found that inhibition of METTL3 regulates ISG15-FASN axis and lipid metabolism to promote glioma progression (Fig. 7k).
a The knockout efficiency of FASN and METTL3 in WT, Mettl3fl/flLyz2Cre and Mettl3fl/flFasnfl/flLyz2Cre BMDMs was analyzed by western blot. (n = 3 independent experiments). b, c Representative images of tumors in WT, Mettl3fl/flLyz2Cre and Mettl3fl/flFasnfl/flLyz2Cre mice implanted with GL261 cells (b) on day 21 and representative HE staining images of brain sections (c) are shown. The tumor area was quantified (c). (n = 6 mice per group). d Representative plots and percentages of IFN-γ-producing CD8+ T cells cocultured with WT, Mettl3fl/flLyz2Cre and Mettl3fl/flFasnfl/flLyz2Cre BMDMs. (n = 5 independent experiments). e, f Representative bioluminescent images of Mettl3fl/fl or Mettl3fl/flLyz2Cre mice bearing orthotopic murine glioma GL261 cell xenografts treated i.p. with vehicle or C75 (e) and bioluminescent signal was quantified (f). (n = 6 mice per group). g Representative HE staining images of brain sections from orthotopic GL261 murine glioma-bearing mice on day 21 post-xenograft, following intraperitoneal treatment with vehicle or C75. The tumor area was quantified. (n = 6 mice per group). h Lymphocytes were cocultured with WT and Mettl3fl/flLyz2Cre BMDMs treated with vehicle or C75 and percentages of IFN-γ-producing CD8+ T cells were quantified. Scale bars: 50 μm. (n = 4 independent experiments). i Immunofluorescence staining for METTL3, ISG15 and FASN in sections from patients with glioma tumors. Quantitative image analysis was performed. Scale bars: 30 μm. (n = 3 independent experiments). j Quantitative image analysis revealed that lower METTL3 levels in CD68⁺ macrophages are strongly and independently associated with diminished overall survival. k Schematic illustrating the mechanism by which METTL3-deficient macrophages promote glioma progression through ISG15-FASN axis-mediated reprogramming of lipid metabolism. Data are expressed as the mean ± SD. P values were determined by one-way ANOVA with Tukey’s multiple comparisons (c, d, f, g, h) and the Gehan-Breslow-Wilcoxon test (j). The source data are provided as a Source data file.
Discussion
TAMs are a major cellular component in the TME. Once macrophages are recruited into the TME, they have to be reprogrammed in the metabolic pathways in order to adapt to the the unique tumor niche11. It has been well-established that m6A was found to be closely related to macrophage phenotype and dysfunction in tumors. Our previous study revealed that Mettl3 deficiency reprograms macrophage and accelerates tumor progression29. This finding was supported by the work from Tong et al. who found that Mettl3-deficient macrophages exhibited reduced TNF-α production upon LPS stimulation and enhanced tumor growth35. Dong et al. also found that Macrophage-specific knockout of an m(6)A methyltransferase METTL14 skews CD8(+) T cell differentiation towards a dysfunctional state, impairing CD8(+) T cells to eliminate tumors36. By contrast, a separate study reported that TAM-specific Mettl3 deletion restrained MC38 tumor growth37. This apparent contradiction underscores the existence of a multifaceted, context-dependent m⁶A-regulatory network that governs myeloid cell behavior within the TME and warrants further dissection. In the study, we found that the expression of METTL3 in TAMs was markedly decreased in GBM patients compared to healthy control. Similarly, a reduction in METTL3 expression was observed in TAMs from mice harboring GL261 tumors. Mechanismly, we discovered that METTL3 deficiency in macrophages facilitates FASN expression, thereby enhancing lipid metabolism, subsequently accelerating glioma progression. Notably, genetic ablation of FASN or treatment with the specific inhibitor C75 effectively nullified the pro-tumorigenic and immunosuppressive effects triggered by METTL3 depletion.
A latest research revealed that METTL3 inhibits hepatic insulin sensitivity via N6-methyladenosine modification of Fasn mRNA and promoting fatty acid metabolism38. Intriguingly, the Fasn mRNA modification was unaltered in METTL3-deficient macrophages, suggesting an alternative mode of METTL3-mediated FASN regulation. Therefore, the molecular bases of FASN regulation in different cells would require future investigations. The expression of FASN is intricately orchestrated at both transcriptional and post-transcriptional levels. Key transcription factors, such as SREBP-139, USF40 and LXR41, have been found to control FASN expression. Nevertheless, transcription factors are difficult to target directly, rendering post-translational regulators as more attractive and feasible drug targets. Thus, the detailed regulatory mechanism of FASN needs to be revealed in order to further explore other promising therapeutic strategies aiming at specifically modulating FASN activity under pathological conditions.
Previous studies have illuminated that FASN degradation is also regulated by the ubiquitin-proteasome pathway. The E3 ligase COP1 can bind FASN to form a complex that promotes FASN degradation42, while the deubiquitinating enzymes USP2a43 and USP1444 directly interact with FASN and increase its stability to promote triglyceride accumulation. SNX8 binds to FASN and promotes its protein degradation by recruiting an E3 ligase TRIM2845. In our study, we found that ISG15 acts as an interactor that regulates FASN ISGylation and stability through inhibiting its ubiquitination. Previous study have demonstrated that ISG15 is an IFN-inducible protein30. Our results also demonstrated that METTL3 knockout enhanced IFN-β expression, mirroring the heightened type I IFN expression observed upon genetic ablation of METTL3 in monocytes31. The METTL3 deficiency resulted in increased expression of ISG15, which contributed to enhanced stability of FASN. Knockdown of ISG15 in METTL3 deficient BMDMs attenuated the stability of FASN and inhibited FASN expression, which resulted in the recovery of CD8+ T cell activation. Similarly, knockdown of FASN in METTL3 deficient BMDMs also lead to the recovery of CD8+ T cell activation. Therefore, our results underscore that the ISG15-regulated FASN expression plays a vital role in accelerating tumor progression caused by METTL3-dificient macrophage.
In vivo experiments also indicated that METTL3 and FASN double knockout facilitated CD8+IFN-γ+ T cell differentiation and inhibited the increased tumor growth triggered by METTL3 depletion. Consistently, pharmacological inhibition of FASN with C75 largely abolished the aggravative effect of METTL3 depletion on tumor progression.
In conclusion, our findings reveal a regulatory mechanism by which METTL3 modulated FASN stability at posttranslational level via ISG15 and lipid metabolism. Specifically, ISG15 inhibited proteasomal degradation of FASN through enhancing FASN ISGylation and blocking FASN polyubiquitination. These findings propose a regulatory network for FASN expression and lipid metabolism in macrophage, supporting the METTL3-ISG15-FASN axis as a promising therapeutic target for managing GBM progression. In murine and human gliomas, the pool of TAMs is composed of brain-resident microglia, which develop from fetal yolk sac progenitors, and peripherally recruited monocyte derived macrophages46. In our study, conditional deletion of Mettl3 in the myeloid lineage reprogrammed recruited monocyte-derived macrophages from a pro-inflammatory M1 to an anti-inflammatory M2 phenotype, thereby fostering an immunosuppressive TME and potentially modulating the activation state of brain-resident microglia during glioma progression.
Method
Clinical specimens
Frozen and paraffin-embedded primary brain tumor tissues were obtained from Chinese patients at Xijing Hospital (Xi’an, China), a subsidiary hospital of the Fourth Military Medical University. The use of clinical specimens in this study was approved by the Animal Experiment Administration Commission in Fourth Military Medical University (KY20253615-1). All adult participants provided written informed consent. No participant compensation was provided. Sex and gender were not considered in the study, as the primary focus of this study was unrelated to sex or gender.
Mice
Lyz2Cre mice (catalog 004781, The Jackson Laboratory) were crossed with floxed Mettl3 mice (Beijing Biocytogen Co., Ltd) to generate Mettl3fl/flLyz2-/- and Mettl3fl/flLyz2cre/- mice. Mettl3fl/flLyz2cre/- mice were crossed with Fasnfl/fl mice (Gempharmatech Co., Ltd) to generate Mettl3fl/flFasnfl/flLyz2cre/- mice. All experimental mice were bred and maintained under specific pathogen-free conditions, fed standard laboratory chow, and kept on 12 h light to 12 h dark cycles. The temperature and humidity were set at 22 ± 1 °C and 55% ± 5%, respectively. Six- to twelve-week-old mice of both sexes were used for the experiments. Cohoused Cre-negative littermate mice were used as controls. All experimental procedures were approved by the Institutional Animal Care and Use Committee of Xinxiang Medical University (XYLL-20230041).
Cell lines
Mouse glioma cell line GL261 and human embryonic kidney cell line HEK293T were cultured in Dulbecco’s Modified Eagle’s Medium supplemented with 10% FBS, 1% penicillin/streptomycin, and 1% L-glutamine, and maintained at 37 °C and 5% CO2.
Plasmid construction
The FASN and ISG15 promoter was amplified and cloned into the PGL-3.0 vector. FLAG-FASN was constructed by cloning human full-length FASN cDNA into the p3×FLAG-CMV-10 vector. FLAG-FASN mutants were generated by homologous recombination method based on FLAG-FASN plasmid. Myc-ISG15 was generated by cloning full-length ISG15 cDNA into the pCMV-Myc vector. Myc-ISG15 ΔGG was generated by depleting the two C-terminal glycine residues.
Metabolite extraction and mass-spectrometry-based metabolomic analysis
The cell culture media samples were thawed on ice, and metabolites were extracted from 20 µL of each sample using 120 µL of precooled 50% methanol buffer. Then the mixture of metabolites was vortexed for 1 min and incubated for 10 min at room temperature, and stored at −20 °C overnight. The mixture was centrifugated at 4000 × g for 20 min, subsequently the supernatant was transferred to 96-well plates. The samples were stored at −80 °C prior to the LC-MS analysis. Pooled quality control sample were also prepared by combining 10 μl of each extraction mixture.
All samples were analyzed using a Triple TOF 5600 Plus high resolution tandem mass spectrometer (SCIEX, Warrington, UK) with both positive and negative ion modes. Chromatographic separation was performed using an ultra performance liquid chromatography (UPLC) system (SCIEX, UK). An ACQUITY UPLC T3 column (100 mm*2.1 mm, 1.8 µm, Waters, UK) was used for the reversed-phase separation. The TripleTOF 5600 Plus system was used to detect metabolites eluted from the column.
Tumor cell implantation
GL261-Luc cells were grown in Dulbecco’s modified eagle medium (DMEM) containing glucose, L-glutamine, and 10% FBS, at 37 °C with 5% CO2 and maintained under normal adherent culture conditions. Prior to tumor cell injection, use a sterile 25 gauge sharp needle to puncture the skull at 2 mm to the right of the bregma and 1 mm anterior to the coronal suture, thereby creating an opening for the injection of tumor cells. A Hamilton syringe (22 gauge needle) containing 300,000 GL261-Luc cells in 3 μl DMEM was inserted into the hole to a depth of 3 mm, and the cells were injected over 5 min. The needle was left in place for 2 min to reduce backflow before the wound was closed with staples.
GBM xenografts
3 × 10⁵ U87 GBM cells were prepared in PBS for stereotactic injection into 6–8 weeks old Nu/Nu mice. Then 5 × 106 Mettl3fl/fl or Mettl3fl/flLyz2Cre myeloid cells mixed with 5 × 106 lymphocytes were injected into Nu/Nu mice via tail vein every week. The mice were sacrificed for HE staining to monitor the tumor growth after 3 weeks. Mice were sacrificed at the indicated time points after tumor implantation and xenografts were harvested for further analyses.
In vivo bioluminescence imaging
For in vivo bioluminescence imaging, mice were weighted, injected intraperitoneally with D-luciferin at 150 mg/kg, and anesthetized with isoflurane. The mice were imaged using an IVIS® Spectrum In Vivo Imaging System (Perkin Elmer, Waltham, MA). The Living Image software was used to draw an ROI around the tumor signal and measure the size of each tumor.
Generation of bone marrow derived macrophages (BMDMs)
Bone marrow was isolated from femurs and tibia of mice, and red blood cells were lysed. The bone marrow cells were cultured in DMEM supplemented with 10% fetal bovine serum, penicillin (100 U/ml), streptomycin (100 μg/ml), and M-CSF (40 ng/ml). Cells were cultured for 7 days to generate BMDMs. Cultured BMDMs were used for qRT-PCR analysis, western blotting, and other experiments.
Transfection
For siRNA transfection, cells were seeded 1 day before transfection. Transfection was performed using Lipofectamine RNAiMAX (Thermo Fisher Scientific) according to the manufacturer’s instructions. The siRNA duplex oligonucleotides used are listed in S1 Table. The plasmids were transfected by Lipofectamine 2000 (Thermo Fisher Scientific) following the manufacturer’s instructions.
qRT-PCR
Total RNA was isolated from cells using TRIzol (Thermo Fisher Scientific). For qRT-PCR analysis of mRNAs, cDNA was generated using a PrimeScript RT reagent kit (TaKaRa). mRNA expression levels were analyzed using SYBR Green PCR master mix (TaKaRa). The primers used for qRT-PCR are listed in S2 Table.
Co-Immunoprecipitation
HEK293T cells were lysed in NP-40 buffer containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 0.3% NP-40, and protease inhibitor cocktail. Cell lysates were incubated with specific antibodies or control IgG for 3 h at 4 °C. The dynabeads protein G (10004D, Invitrogen) were washed three times with NP-40 buffer and mixed with the cell lysates, followed by incubation at 4˚C overnight. Beads were then washed four times using NP-40 buffer. The precipitate samples were subjected to immunoblotting analysis.
Immunoblotting
For immunoblotting, cells were lysed in RIPA buffer and protein concentrations were determined with a BCA assay. Proteins were separated by SDS–PAGE, electrotransferred to PVDF membranes (Millipore), and probed with the indicated antibodies. The primary antibody information is as follows: rabbit anti-FASN (1:1000, 3180S, Cell Signaling Technology), rabbit anti-ACLY (1:1000, 4332S, Cell Signaling Technology), rabbit anti-Phospho-ACLY (1:1000, 4331T, Cell Signaling Technology), rabbit anti-Lipin 1 (1:1000, 14906S, Cell Signaling Technology), rabbit anti-ACSL1 (1:1000, 9189S, Cell Signaling Technology), rabbit anti-C/EBPα (1:1000, 8178S, Cell Signaling Technology), rabbit anti-Perilipin-1 (1:1000, 9349S, Cell Signaling Technology), rabbit anti-PPARγ (1:1000, 2435S, Cell Signaling Technology), rabbit anti-AceCS1 (1:1000, 3658S, Cell Signaling Technology), Mouse anti-ISG15 (1:1000, sc-166755, Santa Cruz Biotechnology), mouse anti-FLAG (1:2000, Sigma), rabbit anti-FLAG (1:1000, 20543-1-AP, Proteintech), mouse anti-Myc (1:2000, M192-3, MBL), rabbit anti-METTL3 (1:1000, ab18810, Abcam), and mouse anti-β-actin (1:5000, Sigma).
mRNA stability analysis
BMDMs were treated with actinomycin D (Sigma) at a final concentration of 5 μg/ml and collected at the indicated time. The total RNA was extracted, then the mRNA transcript levels of interest were determined by qRT-PCR.
Bone marrow chimeras
The bone marrow of recipient mice was ablated with 9.5 Gy irradiation. Meanwhile, donor bone marrow cells were collected from the femurs and tibias of mice by flushing with cold DMEM. The bone marrow was filtered through a 70 µm filter to produce single cell suspensions. Cells were counted and re-suspended in cold PBS. After irradiation, 1 × 107 cells of donor bone marrow cells in 200 µl PBS were injected into recipient mice through the tail vein for reconstruction.
Depletion of macrophages
Macrophages were depleted using Clodronate Liposomes. Briefly, 200 μL Clodronate Liposomes were intraperitoneally injected into mice on days −1, 1, 3, and 5 during tumor growth. Immunofluorescence staining for F4/80+ cells was used to confirm macrophages depletion.
Flow cytometry
To obtain a single cell suspension, brain tumors were finely chopped and digested for 30 min using dispase and collagenase Ⅰ in 1640 medium with intermittent shaking at 37 °C, and then passed through a 70 μm filter. Cell pellets were washed twice in PBS and resuspended in FACS buffer. For surface staining, cells were incubated with the antibodies diluted in FACS buffer on ice for 30 min. To perform intracellular immuno-staining, cells were fixed and permeabilized with a Staining Buffer (ThermoFisher Scientific) according to the manufacturer’s protocol. Samples were run on a BD Caliper (BD) or a CytoFLEX S (Beckman Coulter), and all subsequent compensation and gating design was performed using FlowJo software. The following antibodies were used for flow cytometry: CD45 (30-F11, Biolegend), CD11b (M1/70, Biolegend), Gr1 (RB6-8C5, Biolegend), CD3 (145-2C11, eBioscience), CD8 (clone 53-6.7, Biolegend), IFN-γ (XMG1.2, Biolegend), CD4 (GK1.5, Biolegend), CD49b (HMα2, Biolegend), F4/80 (BM8, Biolegend), Ly6G (1A8, Biolegend), CD11b (clone M1/70, Biolegend), LAG-3 (C9B7W, Biolegend), CD39 (Duha59, Biolegend), TIM-3 (B8.2C12, Biolegend), CX3CR1 (SA011F11, Biolegend), ARG1 (A1exF5, eBioscience), NOS2 (CXNFT, eBioscience).
Immunofluorescence staining
Immunofluorescence was performed using the standard protocol in cells and tissues. Antibodies specific to F4/80 (1:500, ab6640, Abcam), CD68 (1:2000, 66231-2-Ig, Proteintech), FLAG (1:1000, 20543-1-AP, Proteintech), Myc (1:2000, M192-3, MBL), METTL3 (1:1000, ab18810, Abcam), ISG15 (1:1000, sc-166755, Santa Cruz Biotechnology), NOS2 (1:100, PA1-036, ThermoFisher Scientific) and CD206 (1:100, 64693, Abcam) were used. Images were acquired using a microscope (Olympus).
In vitro T cell activation and co-culture with BMDMs
Spleens were collected under sterile conditions. The spleen was smashed on a 70 μm cell strainer and single cells were collected by centrifugation. Red blood cells were lysed by Red Blood Cell Lysis Buffer. Lymphocytes were activated with anti-CD3 (eBioscience, 5 mg/ml) and anti-CD28 (eBioscience, 2.5 mg/ml) for 5 days47. The activated Lymphocytes were cocultured with BMDMs for the indicated time. The cells were then subjected to surface marker and intracellular cytokine staining and analyzed by FACS.
Oligonucleotide design treatment
The phosphothioate ODN48,49 and sense strands (SS) of siRNAs were joined using six units of C3 carbon chain linker, (CH2)3 (GlenResearch). The resulting constructs were annealed to complementary siRNA antisense (AS) strands to generate CpG–siRNA conjugates (asterisks indicate phosphothioation sites). Sequences of single-stranded constructs are listed below: CpG-siRNA Isg15: 5′-T*C*C*A*T*G*A*C*G*T*T*C*C*T*G*A*T*G*C*T/linker-G*C*GGGACCUAAAGGUGAAGAU −3′; In vivo delivery of CpG-siRNAIsg15 for treatment began every other day on day 7 post-inoculation. Mice were sacrificed at the indicated time points after tumor implantation and xenografts were harvested for further analyses.
Statistical analysis
Statistical significance was determined by two-paired t test or ANOVA, and differences were considered statistically significant when P < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data featured in this paper and is provided as part of the paper or as part of the source data file. Source data are provided with this paper.
References
Roda, E. & Bottone, M. G. Editorial: brain cancers: new perspectives and therapies. Front. Neurosci. 16, 857408 (2022).
Qazi, M. A. et al. Intratumoral heterogeneity: pathways to treatment resistance and relapse in human glioblastoma. Ann. Oncol. 28, 1448–1456 (2017).
Xuan, W., Lesniak, M. S., James, C. D., Heimberger, A. B. & Chen, P. Context-dependent glioblastoma-macrophage/microglia symbiosis and associated mechanisms. Trends Immunol. 42, 280–292 (2021).
Simonds, E. F. et al. Deep immune profiling reveals targetable mechanisms of immune evasion in immune checkpoint inhibitor-refractory glioblastoma. J. Immunother. Cancer 9, e002181 (2021).
Butowski, N. et al. Orally administered colony stimulating factor 1 receptor inhibitor PLX3397 in recurrent glioblastoma: an Ivy Foundation Early Phase Clinical Trials Consortium phase II study. Neuro Oncol. 18, 557–564 (2016).
Khan, F. et al Macrophages and microglia in glioblastoma: heterogeneity, plasticity, and therapy. J. Clin. Investig. 133, e163446 (2023).
Zhang, L., Jiang, Y., Zhang, G. & Wei, S. The diversity and dynamics of tumor-associated macrophages in recurrent glioblastoma. Front. Immunol. 14, 1238233 (2023).
Vitale, I., Manic, G., Coussens, L. M., Kroemer, G. & Galluzzi, L. Macrophages and metabolism in the tumor microenvironment. Cell Metab. 30, 36–50 (2019).
Kloosterman, D. J. et al. Macrophage-mediated myelin recycling fuels brain cancer malignancy. Cell 187, 5336–5356.e5330 (2024).
Mazzone, M., Menga, A. & Castegna, A. Metabolism and TAM functions-it takes two to tango. FEBS J. 285, 700–716 (2018).
Ley, K. M1 means kill; M2 means heal. J. Immunol. 199, 2191–2193 (2017).
Li, J., DeNicola, G. M. & Ruffell, B. Metabolism in tumor-associated macrophages. Int. Rev. Cell Mol. Biol. 367, 65–100 (2022).
Masetti, M. et al. Lipid-loaded tumor-associated macrophages sustain tumor growth and invasiveness in prostate cancer. J. Exp. Med. 219, e20210564 (2022).
Schwartz, S. Cracking the epitranscriptome. RNA 22, 169–174 (2016).
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Bokar, J. A., Shambaugh, M. E., Polayes, D., Matera, A. G. & Rottman, F. M. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3, 1233–1247 (1997).
Liu, J. et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 10, 93–95 (2014).
Pendleton, K. E. et al. The U6 snRNA m(6)A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell 169, 824–835.e814 (2017).
Ping, X. L. et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014).
Yue, Y. et al. VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discov. 4, 10 (2018).
Patil, D. P. et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016).
Wen, J. et al. Zc3h13 regulates nuclear RNA m(6)A methylation and mouse embryonic stem cell self-renewal. Mol. Cell 69, 1028–1038.e1026 (2018).
Knuckles, P. et al. Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m(6)A machinery component Wtap/Fl(2)d. Genes Dev. 32, 415–429 (2018).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Chen, X. Y., Zhang, J. & Zhu, J. S. The role of m(6)A RNA methylation in human cancer. Mol. Cancer 18, 103 (2019).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Barbieri, I. et al. Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature 552, 126–131 (2017).
Yin, H. et al. RNA m6A methylation orchestrates cancer growth and metastasis via macrophage reprogramming. Nat. Commun. 12, 1394 (2021).
Perng, Y. C. & Lenschow, D. J. ISG15 in antiviral immunity and beyond. Nat. Rev. Microbiol. 16, 423–439 (2018).
Qiu, W. et al. N(6)-methyladenosine RNA modification suppresses antiviral innate sensing pathways via reshaping double-stranded RNA. Nat. Commun. 12, 1582 (2021).
Bhushan, J. et al. ISG15 connects autophagy and IFN-γ-dependent control of toxoplasma gondii infection in human cells. mBio 11, e00852-20 (2020).
Zhao, C., Denison, C., Huibregtse, J. M., Gygi, S. & Krug, R. M. Human ISG15 conjugation targets both IFN-induced and constitutively expressed proteins functioning in diverse cellular pathways. Proc. Natl. Acad. Sci. USA 102, 10200–10205 (2005).
Munnur, D. et al. Altered ISGylation drives aberrant macrophage-dependent immune responses during SARS-CoV-2 infection. Nat. Immunol. 22, 1416–1427 (2021).
Tong, J. et al. Pooled CRISPR screening identifies m(6)A as a positive regulator of macrophage activation. Sci. Adv. 7, eabd4742 (2021).
Dong, L. et al. The loss of RNA N(6)-adenosine methyltransferase Mettl14 in tumor-associated macrophages promotes CD8(+) T cell dysfunction and tumor growth. Cancer Cell 39, 945–957.e910 (2021).
Xiong, J. et al. Lactylation-driven METTL3-mediated RNA m(6)A modification promotes immunosuppression of tumor-infiltrating myeloid cells. Mol. Cell 82, 1660–1677.e1610 (2022).
Xie, W., Ma, L. L., Xu, Y. Q., Wang, B. H. & Li, S. M. METTL3 inhibits hepatic insulin sensitivity via N6-methyladenosine modification of Fasn mRNA and promoting fatty acid metabolism. Biochem. Biophys. Res. Commun. 518, 120–126 (2019).
Choi, W. I. et al. Proto-oncogene FBI-1 (Pokemon) and SREBP-1 synergistically activate transcription of fatty-acid synthase gene (FASN). J. Biol. Chem. 283, 29341–29354 (2008).
Wong, R. H. et al. A role of DNA-PK for the metabolic gene regulation in response to insulin. Cell 136, 1056–1072 (2009).
Na, T. Y. et al. Liver X receptor mediates hepatitis B virus X protein-induced lipogenesis in hepatitis B virus-associated hepatocellular carcinoma. Hepatology 49, 1122–1131 (2009).
Yu, J. et al. Modulation of fatty acid synthase degradation by concerted action of p38 MAP kinase, E3 ligase COP1, and SH2-tyrosine phosphatase Shp2. J. Biol. Chem. 288, 3823–3830 (2013).
Graner, E. et al. The isopeptidase USP2a regulates the stability of fatty acid synthase in prostate cancer. Cancer Cell 5, 253–261 (2004).
Liu, B. et al. Proteome-wide analysis of USP14 substrates revealed its role in hepatosteatosis via stabilization of FASN. Nat. Commun. 9, 4770 (2018).
Hu, Y. et al. Fatty acid synthase-suppressor screening identifies sorting nexin 8 as a therapeutic target for NAFLD. Hepatology 74, 2508–2525 (2021).
Akkari, L. A.-O. X. et al. Dynamic changes in glioma macrophage populations after radiotherapy reveal CSF-1R inhibition as a strategy to overcome resistance. Sci. Transl. Med. 12, eaaw7843 (2020).
Liao, P. et al. CD8(+) T cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell 40, 365–378.e366 (2022).
Kortylewski, M. et al. In vivo delivery of siRNA to immune cells by conjugation to a TLR9 agonist enhances antitumor immune responses. Nat. Biotechnol. 27, 925–932 (2009).
Ma, S. et al. YTHDF2 orchestrates tumor-associated macrophage reprogramming and controls antitumor immunity through CD8(+) T cells. Nat. Immunol. 24, 255–266 (2023).
Acknowledgements
We are grateful to Mrs. Qingqing Wang from Xinxiang Medical University for carefully breeding and reproduction of mice. We appreciate the contribution of in vivo IVIS Spectrum imaging team, especially Zhao Shan in Equipment Sharing Service Platform in Xinxiang Medical University. This study was supported by the Natural Science Foundation of Henan Province (Grant No. 242300421093), the National Natural Science Foundation of China (Grant No. 82173046, 82472702, 32330047), Shaanxi Province Funds for Distinguished Young Youths (Grant No. 2023-JC-JQ-66), Shaanxi Province key industrial innovation chain project (Grant No. 2024SF ZDCYL-04-04), Rapid response project of AMU (Grant No. 2023KXKT035), the Zhongyuan Academician Fund from Henan Province (No. C24185), 111 Project (No. D20036), Postgraduate Education Reform and Quality Improvement Project of Henan Province (No. YJS2026YBGZZ29), the Fund of State Key Laboratory of Holistic Integrative Management of Gastrointestinal Cancers (Grants No. CBSKL2022ZZ04 and CBSKL2022KF03), the Program for Ph.D. Starting Research Funding from Xinxiang Medical University (Grant No. 505249), the Natural Science Foundation of Henan Province (Grant No. 252300421680). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
H.Y., F.W., R.Z., and A.Y. conceived the research, designed the study, interpreted data and wrote the manuscript; H.Y., X.Y., C.H., Y.Y., M.W., Y.X., B.Z., C.Z., W.Y., Y.L., Z.Y., L.Z., H.M., Y.C., C.W., Z.J., B.L., M.S., and X.C. performed experiments; H.Y., Y.Y., and B.Z. analyzed data; F.W., R.Z., and A.Y. are co-senior authors who directed the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jo Van Ginderachter, Juyeun Lee, and Severa Bunda for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yin, H., Yu, X., Hu, C. et al. METTL3 abrogation promotes glioma progression through regulating the ISG15-FASN axis-mediated lipid metabolism in macrophages. Nat Commun 17, 1326 (2026). https://doi.org/10.1038/s41467-025-68079-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68079-4