Abstract
O-GlcNAc modification is a key cellular signal, but its role in regulating senescence-associated transcription remains poorly understood. Here, we apply a time-resolved chemical genomics strategy to map dynamic O-GlcNAc chromatin-associated proteins (OCPs) during oncogene-induced senescence (OIS) in primary human fibroblasts. Chromatin O-GlcNAc modification continues to accumulate, while 1,987 senescence-associated OCPs undergo dynamic shifts in genomic occupancy across diverse epigenetic chromatin states and display bimodal regulatory activities within the 3,466-gene senescence transcriptome. O-GlcNAc facilitates the formation of dual-function complexes: TF–SWI/SNF activates senescence-associated secretory phenotype (SASP) genes at promoters, whereas NuRD enforces the repression of cell-cycle regulators at enhancers. Furthermore, we identify O-GlcNAc modified JUN and GATAD2A as key regulators of OIS phenotypes in both in vitro and in vivo models of senescence-driven tumorigenesis. These findings reveal dynamic regulation and chromatin organization principles of O-GlcNAc–related epigenetic factors, providing insights into cellular senescence and potential therapeutic strategies.
Similar content being viewed by others
Introduction
Cellular senescence, which involves the irreversible transition of cells from the cell cycle to a state of proliferative arrest due to internal and external stresses, is a primary promoter of age-related diseases such as cancer1,2. Cellular senescence is predominantly categorized into replicative senescence and premature senescence. An increase in cell division frequency and telomere shortening can induce replicative senescence, whereas diverse stimuli, such as oxidative stress, DNA damage, and oncogene activation, can trigger premature senescence3. Cellular senescence is a complex process that occurs over multiple stages and is characterized by dynamic shifts. During this process, senescent cells undergo a series of transformations that result in distinctive phenotypes. These transformations include the development of an enlarged flat morphology, cessation of the cell cycle, and an increase in senescence-associated β-galactosidase (SA-β-Gal) activity4,5. Concurrently, these cells also exhibit an altered secretion phenotype characteristic of the senescence-associated secretory phenotype (SASP), which has been associated with promoting neoplastic cell proliferation and metastasis in many cancers6,7,8,9. Further investigation revealed that these abnormal biological activities occurring during the senescence process are driven by transcriptional characteristics and temporal shifts in the gene expression profile, a phenomenon modulated predominantly through epigenetic regulation10,11,12. Therefore, clarifying the pivotal gene-regulatory networks governing senescent cell fate could identify potential targets for fighting aging-related diseases.
O-linked β-N-acetylglucosamine (O-GlcNAc) modification, a posttranslational modification (PTM) enriched in the nucleus and cytoplasm, contributes substantially to the regulation of chromatin assembly and gene expression13,14,15. O-GlcNAc is a ubiquitous and reversible monosaccharide modification that occurs on protein serine/threonine residues and is catalyzed by the corresponding enzymes: the glycosyltransferase OGT and the glycosidase OGA. Cells can efficaciously modulate the O-GlcNAc levels of an array of pivotal enzymes and signaling molecules, thereby altering the activity and biological functionalities of the substrate proteins, to respond adeptly to alterations in environmental conditions and signal stimulation16,17,18,19,20. We and others have substantiated that chromatin complexes commonly contain O-GlcNAc modifications, significantly impacting the activity of various epigenetic factors, such as transcription factors (TFs) and chromatin remodeling factors21,22,23,24,25,26,27. Dynamic alterations in O-GlcNAc levels, which occur by modulating the stability of these chromatin-binding proteins as well as protein–protein interactions and protein–DNA interactions, influence gene expression on a genomic scale, reconfigure cellular transcriptional patterns, and drive phenotypic manifestations in cells. Recent studies have indicated that O-GlcNAc modification plays an important role in the process of cellular senescence28,29. Moreover, O-GlcNAc can promote DNA damage repair in breast cancer cells and inhibit tumor therapy-induced cellular senescence (TIS)30. STAT3 also contains O-GlcNAc, and downregulating the glycosylation of this TF in neuronal cells can activate target gene expression and induce senescence phenotypes31. The repression complex TRIM28 needs OGT for its assembly on methylated promoters, with O-GlcNAc stimulating its functions and promoting cellular senescence phenotypes32. O-GlcNAc also stabilizes GATA4, promoting SASP acquisition and chronic inflammation; thus, O-GlcNAc plays a key role in osteoarthritis progression33. These findings suggest that the process of cellular senescence requires the involvement of various O-GlcNAc modified epigenetic factors, driving the formation of a characteristic senescence gene expression profile. However, revealing the integrated mechanism through which O-GlcNAc modification orchestrates multiple epigenetic factors and sequentially shapes the character of the senescent cell transcriptome remains challenging.
Herein, to elucidate the regulatory role of O-GlcNAc modification in the oncogene-induced senescence (OIS) process, we employed a chemical reporter-based multiomics strategy for time series proteomic profiling and genome-wide mapping of O-GlcNAc chromatin-associated proteins (OCPs) in primary human fibroblasts. Through dynamic analysis of the transcriptome and epigenome profiles, we demonstrated that OCPs undergo a genomic occupation switch across diverse epigenetic chromatin states and exhibit sequential regulatory activity in the senescence transcriptome. We demonstrate that O-GlcNAc modification facilitates the development of bimodal epigenetic regulatory complexes that control both the SASP and the cell cycle arrest-related genes during OIS. We further identified O-GlcNAcylated JUN and GATAD2A as regulatory epigenetic factors that influence OIS phenotypes using both in vitro and in vivo models of senescence and tumor growth. Collectively, our findings offer evidence for the dynamic behavior and chromatin organization principles of O-GlcNAc related epigenetic factors in driving the senescence process. Our findings also reveal potential avenues for therapeutic manipulation of aging-related diseases, including cancer.
Results
Dynamic accumulation of chromatin O-GlcNAc modifications is associated with the progression of premature senescence in human cells
Primary human lung and skin fibroblasts (derived from individuals; LU1015 and SK0304) and normal human epithelial cell lines (MCF10A, mammary origin; HaCaT, skin origin) were used to investigate the role of O-GlcNAc modification during cellular senescence. To avoid complications associated with the loss of telomeric sequences during premature senescence3, we immortalized the primary fibroblasts by transduction of a lentivirus expressing hTERT (generating monoclonal populations; Supplementary Fig. 1a). Oncogene-induced premature senescence (OIS) models were subsequently established using a doxycycline (Doxy)-responsive Tet-On human34 RASG12V expression system (abbreviated as LU-RAS, SK-RAS, MCF-RAS, and HaC-RAS cells), and the classic biomarkers12 of cellular senescence were assessed (Fig. 1a, b, Supplementary Fig. 1b–i). LU-RAS and SK-RAS fibroblasts undergoing OIS exhibited G2/M arrest, which is consistent with many established features of cellular senescence. In contrast, MCF-RAS epithelial cells were arrested in S phase, in agreement with prior reports35, and HaC-RAS epithelial cells exhibited a similar phenomenon. These differences may reflect cell type–specific programs and the distinct modalities by which senescence is induced.
a Representative SA-β-gal staining of LU-RAS and SK-RAS cells treated with Doxy to undergo OIS for the indicated times. Scale bars, 200 μm. b Flow cytometry was used to determine the percentages of cells in the G1, S, and G2 phases. LU-RAS and SK-RAS cells were treated with Doxy to undergo OIS for the indicated times. c Samples of LU-RAS and SK-RAS cells undergoing OIS were collected on D0, D2, D4, D6, D8, and D10 for time-course analysis. Total cell lysates and crosslinked chromatin complexes were isolated and subjected to immunoblot analysis. O-GlcNAc modification was detected using a CTD110.6 O-GlcNAc-specific antibody. d–g LU-RAS and SK-RAS cells were treated with shRNA targeting OGT, an OGT overexpression construct (pcDNA-OGT), OSMI-4, TMG, or an empty vector control and induced to undergo OIS for 4 days (D4) in the presence of Doxy. d Representative SA-β-Gal staining images are shown. Scale bars, 200 μm. e Representative fluorescence images of EdU incorporation and Hoechst staining in LU-RAS cells undergoing OIS at D4. Scale bars, 100 μm. Cells undergoing OIS and induced to progress to D4 were subsequently pulsed with EdU for 2 h, followed by Hoechst staining. f–g Total cell lysates were extracted and subjected to immunoblot analysis. O-GlcNAc modification was detected using a CTD110.6 O-GlcNAc-specific antibody. The data are presented as means ± SD. The p values were determined using a two-tailed unpaired Student’s t-test. For (a, b) and (d, e), n = 4 biologically independent experiments were performed with similar results. The immunoblots shown in (c, f–g) are representative of three biologically independent experiments with similar results. The source data for the graphs in (a, b, d, e) as well as the blots in (c, f, g) are provided in the Source Data File.
To elucidate the cellular O-GlcNAc modification signatures of OIS, we performed a time series analysis of these models at six time points: Days 0, 2, 4, 6, 8, and 10 (D0–D10). Notably, the formation of a senescence phenotype was accompanied by dynamically elevated levels of O-GlcNAc in both total cell lysates and chromatin precipitates (defined as O-GlcNAc chromatin-associated proteins, OCPs). Consistent with the findings of previous studies29, OGT expression increased with the induction of oncogenic RASG12V. Moreover, OGA expression was attenuated during OIS progression (Fig. 1c). We further revealed that chromatin-associated OGT is upregulated concomitant with OIS. These findings suggest that excess O-GlcNAc modification on chromatin, which is catalyzed by OGT, may be involved in regulating the senescence program.
Furthermore, we modulated cellular O‑GlcNAc levels by altering OGT expression using shRNA-mediated knockdown and an OGT overexpression construct (pcDNA‑OGT) and by pharmacologically inhibiting OGT and OGA with the potent inhibitors OSMI‑4 and Thiamet‑G (TMG), respectively. Indeed, SA-β-Gal activity and proliferation arrest assays revealed that the OIS process was markedly slowed when the O-GlcNAc level was decreased in both fibroblasts and epithelial cells (Fig. 1d, e, Supplementary Fig. 2). In addition, the expression of bona fide markers of senescence, such as p53, p21WAF1/Cip1, and p16INK4A, was decreased, whereas compared with that in control group fibroblasts undergoing OIS, the expression of LMNB1 and MKI67 increased in the absence of O-GlcNAc modification (Fig. 1f, g). In contrast, increasing O‑GlcNAc modification approximately doubled the number of SA‑β‑Gal–positive cells, reduced the number of EdU‑positive cells by approximately twofold, and effectively accelerated the acquisition of the senescence phenotype. Taken together, these results clearly show that O-GlcNAc modification facilitates the participation of human cells in senescence.
Time-resolved multiomics profiling reveals that OCPs impact transcriptional signatures in fibroblasts undergoing OIS
Accumulating evidence has demonstrated that the genomic binding of OCPs serves as a sensor for the rapid induction of transcriptome reprogramming in cell fate decisions18,36,37. We employed a metabolic chemical reporter-based multiomics strategy developed in a previous study18 to systematically elucidate the mechanisms by which OCPs orchestrate the OIS program. An optimal chemical reporter, 1,6-Pr2GalNAz, which is capable of efficient metabolic incorporation into O-GlcNAc modified proteins while preventing the nonspecific S-glyco-modification38, was added during the culture of senescent LU-RAS cells. We subsequently performed chemoselective conjugation using an alkyne–biotin-tag for specific isolation and hierarchical analysis of azide-labeled O-GlcNAc (O-GlcNAz) chromatin. We profiled the presence of nucleus-localized OCPs using quantitative proteomics and mapped the chromatin occupation dynamics of the interrelated OCPs using next-generation sequencing (chemoselective O-GlcNAc chromatin sequencing, COGC-seq) in a time series pattern during OIS progression. To further elucidate the transcriptomic and epigenomic dynamic signatures of OIS, RNA-seq, histone modification ChIP-seq, and whole-genome DNA methylation (WGBS) analyses were performed at the time points indicated in Fig. 2a. Crosslinked chromatin complexes were fractionated and extracted, with decrosslinking performed prior to proteomic analysis to minimize potential nonspecific carryover (Supplementary Fig. 3a). The successful incorporation of azides into OCPs was verified through immunoblotting of cell lysates, chromatin precipitates, and the biotin–streptavidin pulldown fraction from LU-RAS cells undergoing OIS and cultured with 1,6-Pr2GalNAz (Fig. 2b).
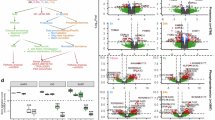
a Schematic overview of defining the gene-regulatory code of OIS in LU-RAS cells. b Immunoblotting and silver staining of 1,6-Pr2GalNAz–labeled LU-RAS cell lysates and chromatin collected during OIS at D0, D2, D4, D6, D8, and D10. β-actin, Histone 3 (H3), and SRSF2 mark cytoplasmic, chromatin, and nucleoplasmic fractions, respectively. c PCA plot showing distinct clustering of OCP quantitative proteomics data across the OIS time series. d Volcano plot of OCP quantitative proteomics comparing D0 and D10, analyzed by two-tailed unpaired Student’s t test. Representative OCPs are highlighted. e GO terms enriched among differentially quantified OCPs between D0 and D10. f Scatter plot of multidimensional scaling (MDS) based on normalized RNA-seq data across the OIS time series in LU-RAS cells. Sample distances represent leading log fold changes, calculated as the root-mean-square of log(FC) values for genes distinguishing each sample pair. g Heatmaps showing hierarchical clustering of normalized RNA-seq DEGs at the indicated time points. Cluster 1 genes are upregulated, Cluster 2 genes downregulated during OIS. Data are shown as row Z-scores. Representative genes are indicated. h Bubble plot of GO term enrichment for cluster genes in (g). Point size reflects the number of genes per GO term; color indicates significance. i Annotation of LU-RAS COGC-seq peaks (MACS2 callpeak) relative to genomic elements (top). Heatmap of COGC-seq signal enrichment within ±2 kb of peak centers (bottom; white: zero, red: low, blue: high). Blots are representative of three biologically independent experiments (b). Data in (c–e) are from n = 9 biologically independent experiments; data in (f–i) are from two biologically independent experiments. GO enrichment used a one-sided hypergeometric test with Benjamini-Hochberg correction where appropriate (e, h). The source data for the graphs in (c–h) as well as the blots in (b) are provided in the Source Data File.
Principal component analysis (PCA) and quantitative analysis of the proteomics data demonstrated both the reproducibility across biological replicates and the significant changes in OCPs over OIS time courses (Fig. 2c). Specifically, compared with that at D0, high-confidence OCPs (identified in at least six of nine biological replicates at any timepoint) exhibited increased chromatin binding and enrichment in cellular senescence-related functions at D10, with 436 upregulated and 93 downregulated OCPs meeting significance thresholds (fold change ≥ 2, p ≤ 0.05; Fig. 2d, e). Furthermore, 1987 OCPs (nucleus-localized) were identified, all of which exhibited a consistent quantitative increase in abundance during OIS progression (Supplementary Data 1). Ontological analyses of these OCPs revealed their enrichment in terms related to transcriptional regulation, chromatin organization, cellular senescence and epigenetic regulation of gene expression, and further protein–protein interaction (PPI) network analysis demonstrated interconnected nodes among these functional complexes, suggesting a role for OCPs in coordinating OIS transcriptional signatures (Supplementary Fig. 3b–d). Subsequently, global gene expression profiling by RNA-seq revealed a total of 10,776 differentially expressed genes (DEGs), including 5430 upregulated and 5346 downregulated genes, between at least two time points during OIS progression (fold change ≥ 2 and FDR ≤ 0.05; Fig. 2f; Supplementary Data 2). The transcriptome data revealed dynamic gene expression trajectories, with a notable shift at D4 (D4–D10 showing similar expression trends), supporting the selection of D0, D2, D4, and D10 as key time points for advanced COGC-seq and epigenome analyses (Fig. 2g and Supplementary Fig. 4a). Both gene ontology (GO) and gene set enrichment analysis (GSEA) of the DEGs revealed that cell cycle-related pathways were significantly downregulated in senescent cells, whereas SASP-associated pathways, such as the TNF-α and inflammatory response pathways, were upregulated (Fig. 2h and Supplementary Fig. 4b). COGC-seq peak calling analysis revealed a progressive increase in both the peak-centered signal intensity and the proportion of peaks mapping to promoters for OCPs during the OIS process, accompanied by distinct peak-associated genes and unique characteristics of OCP loci across the genome (Fig. 2i, Supplementary Fig. 5, and Supplementary Data 3). These data obtained from the chemoselective O-GlcNAz chromatin sequencing analysis suggest that the increasing O-GlcNAc modification during OIS in fibroblasts results in increased binding of OCPs to chromatin, potentially mediating the precise transcriptional regulation of senescence-related genes.
OCPs undergo a genomic occupation switch and exhibit bimodal epigenetic regulatory activity in the senescence transcriptome
To comprehensively investigate the role of O-GlcNAc modification in orchestrating gene expression during OIS cell fate transition, we examined multiple chromatin epigenetic marks and mapped a comprehensive landscape of the dynamic chromatin loci of OCPs. COGC-seq and histone 3 PTM ChIP-seq datasets were integrated, and unsupervised ChromHMM analysis was performed. This approach allowed us to estimate the genome-wide co-occurrence of O-GlcNAc with or without distinct chromatin state markers (H3K4me1, putative enhancers; H3K4me3, putative promoters; H3K27ac, active cis-elements; and H3K27me3, repressed chromatin; Supplementary Fig. 6). Across all four OIS time points (D0, D2, D4, and D10), we delineated 8 chromatin states, each characterized by a distinct enrichment pattern pertaining to one or more markers, including O-GlcNAc (Fig. 3a). The fraction of O-GlcNAc related chromatin states can be categorized into four functional classes: O-GlcNAc promoter, O-GlcNAc enhancer, O-GlcNAc occupied transcriptionally repressed chromatin (O-GlcNAc-repressed), and a class lacking any of the profiled histone modifications (O-GlcNAc-unmarked). Consistent with the genome-wide increase in OCP binding, the cumulative fraction of chromatin states occupied by O-GlcNAc (i.e., the sum of the O-GlcNAc promoter, O-GlcNAc-repressed, and O-GlcNAc-unmarked classes) was increased in OIS-affected cells, from a minimum of 64.1% at D2 to a maximum of 83.5% at D10 of the genomic analysis. Notably, there were two main extensive internal switch directions within the chromatin states of the entire class from D0 to D10, with a high frequency of chromatin state transitions detected at O-GlcNAc-enhancers and a moderate influence observed at O-GlcNAc-promoters (Fig. 3b and Supplementary Fig. 7). During the D0–D2 and D4–D10 intervals, many enhancer-marked regions underwent complete removal of active markers and became either O-GlcNAc-unmarked or O-GlcNAc-repressed chromatin, indicating the repression of these chromatin regions during OIS. Another category exemplified the transition from other chromatin states to the O-GlcNAc promoter state, which resulted in a gradual increase in the proportion of this specific promoter type, indicating the activation of these chromatin regions during OIS. These data indicate that OCPs participate in the bimodal dynamics of both sequential enhancer repression and promoter activation during the OIS process. These results were also confirmed by the increased degree of conservation with the distributions of H3K27me3, H3K4me1, and DNA methylation in COGC-seq–annotated intergenic/enhancer peaks (the subset of COGC-seq peaks annotated as intergenic/enhancer regions, from D0 to D10), while the COGC-seq–annotated promoter peaks (COGC-seq peaks annotated as promoter regions) exhibited increased intensity levels of activating histone markers (H3K4me3 and H3K27ac, Fig. 3c).
a Eight chromatin states were identified by ChromHMM using COGC-seq and histone mark ChIP-seq. Each column represents a chromatin state and each row an O-GlcNAc or histone mark (left). The genome coverage of each state at the indicated time points is shown (right). b Sankey diagram illustrating dynamic chromatin state transitions during OIS in LU-RAS cells. Edge width reflects the number of transitions, representing the average transition landscape. c Average ChIP-seq enrichment profiles for histone marks or DNA methylation at COGC-seq-annotated promoter (top, red) or enhancer/intergenic (bottom, green) peaks at the indicated OIS time points. d Heatmap of average read counts within COGC-seq peaks that increased or decreased relative to D0, shown as row Z-scores. e Bubble plot of GO term enrichment for genes linked to quantitatively increased or decreased peaks during OIS. Bubble size indicates gene number and color indicates enrichment significance. f Average ChIP-seq enrichment profiles for histone marks or DNA methylation at quantitatively increased promoter peaks (top, red) and quantitatively decreased enhancer/intergenic peaks (bottom, green) at D0 and D10. g Venn diagrams showing overlap between upregulated (top) or downregulated (bottom) RNA-seq DEGs and genes associated with quantitative COGC-seq peaks. h Box plots showing mRNA expression changes of overlapping genes in (g) at the indicated OIS time points (median, box: 25th–75th percentiles; whiskers: min–max). p values, two-tailed unpaired Student’s t test. i GO enrichment analysis of overlapping genes in (g). Data are from two biologically independent experiments (a–i). GO enrichment used a one-sided hypergeometric test with Benjamini–Hochberg correction where appropriate (e, h). Source data for (a, b, d, e, g–i) are provided in the Source Data file.
To further elucidate the characteristics of the genomic occupation switch of OCPs during the process of OIS, we quantitatively analyzed the peaks derived from COGC-seq datasets (Supplementary Fig. 8 and Supplementary Data 4). Given the increased genomic binding of OCPs, along with their occupancy at promoters, and the decreased signals at intergenic/enhancer regions (Fig. 2i), we further conducted a functional analysis of these quantitatively altered COGC-seq peaks (all MAnorm quantitatively differential peaks compared to D0 were included; Fig. 3d). Functional enrichment analysis revealed that genes associated with the quantitatively decreased subset of the above OCP chromatin-binding sites (COGC-seq peaks) were linked to cell cycle regulation, whereas numerous genes related to the quantitatively increased subset were annotated as being involved in SASP-related signaling (Fig. 3e). Despite the lack of discernible alterations in the distribution of DNA methylation throughout the entire genome (Supplementary Fig. 9), compared with the peaks at D0, the enhancer peaks quantitatively decreased at D2–D10 exhibited prominently higher levels of DNA methylation, H3K4me1 and the repressive marker H3K27me3 (Fig. 3f, Supplementary Fig. 10). On the other hand, at the quantitatively increased promoter peaks, the intensity of H3K4me3 and H3K27ac were clearly increased, whereas DNA methylation was slightly decreased. These findings underscore the dual regulatory role of OCPs in orchestrating the OIS program. To determine the correlations between OIS-induced OCP chromatin binding and gene expression, we compared differential COGC-seq peak-associated genes with DEGs identified by RNA-seq (Fig. 3g, Supplementary Data 5). Importantly, genes associated with COGC-seq–annotated decreased enhancer peaks (e.g., CCND1, STAG2, CDK6, and E2F8) displayed progressively decreased expression (2187 genes) and were enriched in processes related to the cell cycle and proliferative arrest. Conversely, genes linked to promoter peaks with increased OCP binding according to COGC-seq (e.g., IL6 and IGFBP2) showed temporally upregulated expression (1279 genes) and were associated with cellular stress response- and SASP-related functional terms (Fig. 3h, i), indicating that O-GlcNAc associated enhancer suppression and promoter activation were tightly coordinated with the temporal expression patterns of these genes. Collectively, our findings suggest that during OIS, OCPs exhibit an increase in genome-wide chromatin interactions, undergo a transcriptionally repressed enhancer and activated promoter bidirectional genome-wide binding switch, and thus participate in orchestrating the transcriptomic characteristics of OIS phenotypes.
OCPs assemble distinct epigenetic transcriptional complexes during the progression of OIS
To further elucidate the mechanism by which OCPs regulate the OIS transcriptome, we leveraged quantitative proteomics data to construct an OCP interaction network (Supplementary Fig. 3e), revealing two principal groups of epigenetic transcriptional OCP complexes. PPI analysis demonstrated that the transcriptional repression group contained nodes such as chromatin remodeling proteins and DNA methylation-dependent regulatory factors, which centrally engage the NuRD repressor complex as the core. Another group was centered on the SWI/SNF complex, encompassing multiple TFs, which play an important role in transcriptional activation (Fig. 4a). Furthermore, we confirmed that multiple members of the NuRD and SWI/SNF complexes, along with TFs and OGT, exhibited stronger interactions with chromatin at D10 than at D0 in OIS-affected cells (Fig. 4b). To identify the TFs regulated by O-GlcNAc and involved in gene expression regulation during the OIS process, we subsequently scanned the quantitatively increased COGC-seq–annotated promoter peaks (from D0 to D10) using de novo motif discovery algorithms. After evaluating the potential TF-binding sites and motif enrichment values, we identified a variety of TF motifs that were also identified by OCP proteomics (Fig. 2d), with distinct distribution patterns varying from D0 to D10 (Fig. 4c). bZip TFs, particularly AP-1 superfamily members (e.g., JUN, ATF7, and FOSL2), were found to exhibit increased activity and a continuously increase in chromatin binding during OIS, corroborating previous reports39,40. In addition, substantial changes in the chromatin-binding parameter were identified for the fork head helix (FOX) family (FOXK1, RFX5) and C2H2 zinc finger family of TFs, implying that O-GlcNAc modification of these epigenetic factors could have an appreciable effect on OIS phenotypes.
a STRING PPI network and functional enrichment analysis of epigenetic transcriptional OCPs in LU-RAS cells during OIS progression. The node fill colors represent the proteomic quantification of OCPs (fold change at D10 relative to D0; red: high, blue: low). b Immunoblot analysis of chromatin complexes isolated from LU-RAS cells at OIS time points D0 and D10. c Heatmap showing the chromatin binding sites and TF enrichment derived from motif analysis of upregulated COGC-seq-annotated promoter peaks in LU-RAS cells during OIS. The dot color represents the degree of TF motif enrichment, and the dot size reflects the percentage of target sequences with TF-binding sites. P values were calculated using a hypergeometric test. Multiple testing correction was performed using the Benjamini–Hochberg method. d Whole-cell lysates of LU-RAS cells at specified OIS time points were immunoprecipitated with an anti-O-GlcNAc antibody (EPR19847), and the levels of NuRD and SWI/SNF components were analyzed by immunoblotting. e LU-RAS cells at OIS time points D0 and D10 were treated with OSMI-4 or vehicle. Whole-cell lysates were subjected to immunoprecipitation with an anti-GATAD2A or anti-JUN antibody. Immunoprecipitates were analyzed by immunoblotting to detect O-GlcNAc (CTD110.6) and the presence of NuRD or SWI/SNF complex components. The immunoblots are representative of three biologically independent experiments that yielded similar results (b, d, e). The source data for the graphs in (c) and the blots in (b, d, e) are provided in the Source Data File.
We then examined the O-GlcNAc state of the NuRD and SWI/SNF components during the OIS process (Fig. 4d). Consistent with the OCP proteomics results (Fig. 2d), nearly all components of the NuRD repressor complex (MTA1, MBD2, CHD3, and GATAD2A) were identified as being O‑GlcNAcylated, with the extent of glycosylation increasing as OIS progressed. The results of pulldown assays using succinylated wheat germ agglutinin (sWGA), a lectin that specifically recognizes the O‑GlcNAc moiety41, corroborated these findings (Supplementary Fig. 11). Similar results were obtained for the SWI/SNF activator complex (including SMCA4, SMRC1, and SNF5) and TFs (including JUN and FOXK1), indicating the important role of O-GlcNAc modification in the activity of these diverse epigenetic complexes. Co-IP assays further demonstrated that the interactions between NuRD and TF-SWI/SNF complexes were amplified in OIS-affected cells (Fig. 4e). However, inhibiting O‑GlcNAcylation with the small molecule OGT inhibitor OSMI‑4 markedly disrupted the assembly of these complexes, indicating the important role of O-GlcNAc in the interactions among diverse epigenetic complex components during OIS.
O-GlcNAc modification facilitates the formation of TF-SWI/SNF and NuRD complexes that perform diverse functions in senescence reprogramming
To elucidate the role of O-GlcNAc modification in the assembly of epigenetic complexes and the impact of these complexes on senescence reprogramming, a comprehensive analysis was conducted to identify O-GlcNAc sites in OCPs (Fig. 5a). By utilizing an alkyne–biotin tag with a UV-cleavable linker, we concentrated and examined the O-GlcNAz modified peptides from OIS chromatin through higher-energy collision dissociation product-dependent electron transfer/higher-energy collisional dissociation (HCD pdEThcD)-tandem mass spectrometry (MS). In total, 128, 526, 657, and 688 high-confidence O-GlcNAc sites were identified (present in at least two of three biological replicates at any given time point) in OIS-affected cells at D0, D2, D4, and D10, respectively. Altogether, 969 sites were identified across all the samples, indicating dynamic changes in O-GlcNAc modifications during OIS (Supplementary Data 6). These sites were distributed across 405 proteins, which exhibited significant enrichment in biological processes such as chromatin organization and epigenetic regulation of gene expression (Fig. 5b and Supplementary Fig. 12). Although several O-GlcNAc sites have been reported in the O-GlcNAc Database v2.042, our mass spectrometry data acquired during OIS in LU-RAS cells enabled the pinpointing of potential O-GlcNAc sites on key components of the NuRD and TF–SWI/SNF complexes—specifically, S182 and S625 of GATAD2A, S84 of JUN, T679 of FOXK1, and S1494 of ARID2 (Fig. 5c, and Supplementary Fig. 13). Moreover, a subset of these sites was well conserved across species (e.g., S182/S625 of GATAD2A and S84 of JUN), implying a pivotal functional role. However, previously reported sites near GATAD2A S182 (T175, T180, S182, S185, and T188) and in other proteins were not detected in our MS analysis. A plausible explanation for this discrepancy is the source difference, since many database entries are derived from tumors with elevated O-GlcNAcylation, whereas our normal human embryonic lung fibroblasts have lower O-GlcNAcylation levels, which reduces their detectability even with high-sensitivity MS.
a Workflow for quantitative and site-specific analysis of O-GlcNAc modifications on OCPs during OIS in LU-RAS cells. b Venn diagram showing overlap of high-confidence O-GlcNAc sites across OIS time points. c Mapped O-GlcNAc sites in GATAD2A and JUN. d, e LU-RAS/KD cells stably expressing empty vector (vehicle), wild-type (Flag-GATAD2AWT and HA-JUNWT; LU-RAS/KD-WT cells) or mutant (Flag-GATAD2AS182A and HA-JUNS84A; LU-RAS/KD-AA cells) constructs were analyzed at D0 and D10. Proteins were immunoprecipitated with anti-Flag or anti-HA magnetic beads (d) or pulled down using sWGA lectin (e). f, h Immunoprecipitation of cells in (d) at D0 and D10 followed by immunoblot analysis of NuRD complex components (f) and TF-SWI/SNF complex components (h). g Size exclusion chromatography (Superose 6) of nuclear extracts from the indicated cells, followed by immunoblot analysis of NuRD and SWI/SNF subunits. Fractions 4–13 represent holo-NuRD and SWI/SNF complexes; fractions 13–25 contain unassembled components. i Immunofluorescence images showing nuclear colocalization of Flag-GATAD2A with MBD2 and HA-JUN with SNF5 in LU-RAS/KD-WT and LU-RAS/KD-AA cells at D10. Nuclei were stained with DAPI (blue). Scale bars, 10 μm. A Pearson coefficient of 1 indicates perfect colocalization, 0 indicates none. j Truncated variants of vehicle (vehicle-1 to vehicle-3), Flag-GATAD2A (ΔGATAD2A-1 to ΔGATAD2A-3) and HA-JUN (ΔJUN-1 to ΔJUN-2) were designed (top), and co-IP was performed using Flag and HA tags (bottom). k Simulation of GATAD2A interaction with the NuRD complex (replica 1), with GATAD2A in wheat, MTA1 blue, MBD2 gray, HDAC1 green. Zoomed view shows the O-GlcNAc-S182GATAD2A (red sticks) interaction network, with the P157MBD2–S85HDAC1 distance used to characterize binding. Distance I curves for indicated systems are shown. l Overall model of JUN within SWI/SNF component SNF5 and nucleosome (replica 1), with JUN in wheat, SNF5 sky blue, other proteins gray, DNA green. Zoomed view shows O-GlcNAc-S84JUN interaction with SNF5 and the O-GlcNAc JUN-E59SNF5 distance. Distance curves for indicated systems are shown. Immunoblots (d–h, j) are representative of three independent experiments. Source data for (b) and for immunoblots (d–h, j) are in the Source Data File.
GATAD2A plays a crucial role in the assembly of the integrated NuRD complex, significantly contributing to methylated DNA binding and transcriptional repression43,44,45. Moreover, the AP-1 family of TFs, typified by JUN, plays an important role in cellular senescence39,46. Hence, in this study, their functionalities related to O-GlcNAc modifications were analyzed in depth. LU-RAS cells were stably transduced with lentiviruses containing shRNAs targeting GATAD2A and JUN (shGATAD2A and shJUN, respectively; referred to as LU-RAS/KD cells), and recombinant wild-type constructs (tandem expression of Flag-GATAD2AWT and HA-JUNWT; LU-RAS/KD-WT cells) or mutant constructs (tandem expression of Flag-GATAD2AS182A and HA-JUNS84A; LU-RAS/KD-AA cells) with expression at comparable levels were then reintroduced (Supplementary Fig. 14a, b). The co-IP and sWGA pulldown results demonstrated that mutations at these sites led to a significant decrease in the O-GlcNAc signal on both GATAD2A and JUN (Fig. 5d, e). Similar results were obtained in HEK‑293 T cells. Residual O‑GlcNAc signals were observed in the S182A and S84A mutants, indicating that GATAD2A and JUN are O‑GlcNAcylated predominantly at S182 and S84, respectively (Supplementary Fig. 14c).
Previous research has indicated that the bridging of repression components to NuRD core subunits by GATAD2A also relies on the N-terminal coiled-coil domain45,47, which contains the O-GlcNAc site. Our findings showed that GATAD2AWT coimmunoprecipitated multiple NuRD components in OIS-affected cells at D10. Mutation of the glycosylation site in GATAD2A moderately reduces its interaction with MTA1 and HDAC1 but markedly diminished its interaction with the DNA methylation reader MBD248 and the transcriptional repressor CHD449, suggesting that O-GlcNAc regulates the recruitment of repressors to the core subunits of NuRD during the OIS process (Fig. 5f). Furthermore, we performed a fractionation assay on the nuclear extract from OIS-affected cells at D10 using size exclusion chromatography and immunoblot analysis for CHD4, MBD2, MTA1, and the Flag tag (GATAD2AWT or GATAD2AS182A). In the GATAD2AWT group, all four proteins were detected in fractions of very high molecular weight (fractions 4–13), signifying their participation in the complete NuRD repressor complex (Fig. 5g). In contrast, GATAD2AS182A was found in a lower molecular weight complex in fractions containing MTA1 (fractions 13-19) but had minimal overlap with CHD4 (fractions 16-22) and MBD2 (fractions 19-25). This pattern suggests that the lack of O-GlcNAc modification inhibited the formation of the MBD2/CHD4-containing NuRD repressor complex. In the same vein, we observed a comparable scenario in the assembly of JUN and the SWI/SNF transcriptional activator complex, wherein O-GlcNAc is capable of increasing the interaction of JUN with multiple components of the complex (Fig. 5h). Notably, immunofluorescence staining revealed colocalization of Flag-GATAD2AWT:MBD2 and HA-JUNWT:SNF5 in the nucleus in OIS-affected cells. However, the absence of O-GlcNAc resulted in the dissociation of these complexes (Fig. 5i). By expressing truncated forms of GATAD2A (Fig. 5j), we deduced that a region in the N-terminal coiled-coil domain (ΔGATAD2A-1, aa 1–240) is necessary for facilitating the interaction between MTA1 and MBD2. This region lacks a clear domain structure but contains a coiled-coil, which was reported to be O-GlcNAcylated at the terminus with multiple potential modification sites50,51. In the interaction between SNF5 and JUN, the loosely organized N-terminal domain (ΔJUN-1) containing the O-GlcNAc site is crucial. This finding implies the important role of O-GlcNAc modification in the assembly of TF-SWI/SNF and NuRD complexes that perform different functions.
Using all-atom molecular dynamics simulations, we investigated how O-GlcNAc modulates the binding of GATAD2A and JUN with their partners. O-GlcNAcylated and non-glycosylated GATAD2A-CR1 (aa 137–190) and the JUN N-terminus (aa 69–91) exhibited distinct binding interactions with MBD2 and SNF5, respectively, with O-GlcNAcylation strengthening these interactions (Fig. 5k, l). Specifically, O-GlcNAc on GATAD2A promoted its interaction with MBD2, stabilizing the MBD2–HDAC1/MTA1 assembly (Supplementary Fig. 15a–e), as indicated by the decreases in Distance I (S85HDAC1 − P157MBD2, Fig. 5k and Supplementary Fig. 15g) and Distance II (K302MTA1 − E162MBD2, Supplementary Fig. 15f, g), the more gradual time evolution plots of the C-α root mean square deviation (RMSD) for MBD2 and HDAC1/MTA1, and the increased MBD2–HDAC1/MTA1 contact area, including the changes at key interfacial residues (Supplementary Fig. 15h–k). Similarly, in the JUN system, O-GlcNAcylation stabilized JUN–SNF5 binding and thereby affected SNF5–DNA stability (Supplementary Fig. 16a–f), as supported by the decreased S84JUN–E59SNF5 distance (Fig. 5l; Supplementary Fig. 16h), increased contact area (Supplementary Fig. 16g, h), and altered interactions at key interfacial residues (Supplementary Fig. 16i, j). Across the systems, the three replicates yielded consistent results (Supplementary Fig. 15l and m; Supplementary Fig. 16k, l). Consequently, these findings clearly demonstrate that O-GlcNAc facilitates the formation of TF-SWI/SNF and NuRD complexes, which carry out dual transcriptional regulatory functions at promoters and enhancers, respectively, during senescence reprogramming.
O-GlcNAc modification of JUN and GATAD2A plays an important role in intensive chromatin occupation correlated with the SASP and cell cycle arrest
To analyze the influence of O-GlcNAc modification on the dynamic targeting of the SWI/SNF and NuRD complexes in OIS gene reprogramming, we evaluated the chromatin binding patterns of JUN and GATAD2A in OIS-affected cells with varying O-GlcNAc levels. The levels of chromatin-bound JUNWT and GATAD2AWT in LU-RAS/KD-WT cells were markedly greater than those in JUNS84A and GATAD2AS182A-mutant LU-RAS/KD-AA cells at both D0 and D10, respectively (Fig. 6a). The levels of JUNS84A and GATAD2AS182A were increased in the nuclear soluble and cytoplasmic fractions, corroborating the finding that glycosylation promotes the chromatin association of JUN and GATAD2A. Moreover, suppressing the glycosylation of JUN and GATAD2A diminished the chromatin accumulation of SNF5 and MBD2 at both D0 and D10, indicating a role for O-GlcNAcylation in mediating the interactions and colocalization among these factors. We next performed ChIP-seq analysis of these variants in OIS-affected cells to precisely determine the target genes of the O-GlcNAc modified SWI/SNF and NuRD complexes (Fig. 6b). Upon comparing the ChIP-seq signals of the JUN and GATAD2A variants, we observed a notable reduction in the intensity of JUNS84A at the COGC-seq–annotated promoter peaks in OIS-affected cells (Fig. 6c). In addition, the removal of O-GlcNAc interfered with the presence of SNF5 (as indicated by SNF5 ChIP-seq) at JUN-bound regions, suggesting that O-GlcNAc is necessary for maintaining the presence of SWI/SNF chromatin and the colocalization of the complex with JUN (Fig. 6d). Moreover, the accumulation of GATAD2AS182A was also decreased at the COGC-seq–annotated intergenic/enhancer peaks, resulting in decreased colocalization of GATAD2A with MBD2. We further revealed greater saturation of active promoter marks (H3K4me3 and H3K27Ac) in JUNWT chromatin-anchored areas than in JUNS84A-bound ChIP-seq peaks. O-GlcNAc-induced GATAD2A chromatin binding was also linked to a notably increased degree of DNA methylation compared with that in GATAD2AS182A-binding regions (Supplementary Fig. 17), consistent with the finding that O-GlcNAc promotes the interaction between GATAD2A and the NuRD complex. Illustrative results from the visual representation of the distinct genomic locations occupied by JUN and GATAD2A variants, including SASP-related genes such as IL6 and IGFBP2 and cell cycle-related genes such as CCND1 and STAG2, show O-GlcNAc-induced alterations in chromatin binding at the individual gene level (Fig. 6e). These findings point to a crucial link between O-GlcNAc and epigenetic complexes, where O-GlcNAc could function as a navigational factor for genomic targeting of SWI/SNF and NuRD.
a Immunoblot analysis of chromatin extracted from LU-RAS/KD-WT and LU-RAS/KD-AA cells at OIS time points D0 and D10, showing chromatin-bound Flag-GATAD2A, HA-JUN, their variants, MBD2, and SNF5. Whole‑cell lysates, cytoplasmic fractions, and soluble nuclear fractions served as controls. n = 3 biologically independent samples were analyzed, with similar results obtained across replicates. b Average enrichment profiles of ChIP-seq reads (n = 2 biological replicates) for Flag-GATAD2AWT, Flag-GATAD2AS182A, HA-JUNWT, and HA-JUNS84A at COGC-seq peaks in LU-RAS/KD-WT and LU-RAS/KD-AA cells at the D0 and D10 OIS time points. c Heatmap showing ChIP-seq signal enrichment for Flag-GATAD2AWT and Flag-GATAD2AS182A within ±2 kb of COGC-seq-annotated enhancer/intergenic peak centers and for HA-JUNWT and HA-JUNS84A within ±2 kb of COGC-seq-annotated promoter peak centers. The color indicates the enrichment level (white: low; green/red: high). d Average ChIP-seq enrichment profiles of MBD2 at the Flag-GATAD2AWT and Flag-GATAD2AS182A ChIP-seq peaks and of SNF5 at the HA-JUNWT and HA-JUNS84A ChIP-seq peaks in LU-RAS cells at the D10 OIS time point. e Integrative Genomics Viewer (IGV) tracks displaying ChIP-seq signals for Flag-GATAD2AWT, Flag-GATAD2AS182A, HA-JUNWT, and HA-JUNS84A at the D0 and D10 OIS time points across representative SASP- and cell cycle-related genes. f Heatmaps showing RNA-seq data for representative SASP- and cell cycle-related genes in LU-RAS/KD-WT and LU-RAS/KD-AA cells at the D10 OIS time point. The data are presented as row Z-scores based on two biologically independent experiments. g, h Validation of O-GlcNAc GATAD2A and JUN binding peaks at the indicated gene loci by ChIP–qPCR (g). The bars show the % of total input chromatin, with IgG used as a negative control. Quantitative PCR (qPCR) analysis of the mRNA levels of the indicated genes in LU-RAS, LU-RAS/KD (vehicle), LU-RAS/KD-WT, LU-RAS/KD-AA and LU-RAS/KD-WT cells treated with OSMI‑4 (5 μM) for 48 h at the D10 OIS time points (h). The data are presented as mean ± SD from four biological replicates with similar results. The p values were calculated using a two-tailed unpaired Student’s t test. The source data for the graphs in (f–h) and immunoblots in (a) are provided in the Source Data File.
To further investigate the regulatory functions of O-GlcNAc modification in OIS, we carried out RNA-seq analysis on cells expressing JUN and GATAD2A variants (Supplementary Data 7). The results of the transcriptomic analyses revealed that at D10, overexpression of JUNWT and GATAD2AWT—relative to their O-GlcNAc–deficient mutants—significantly decreased the expression of cell-cycle–related genes and increased the expression of SASP genes (Fig. 6f and Supplementary Fig. 18). Consistent with this finding, depletion of O-GlcNAc resulted in reductions in JUN and GATAD2A ChIP-seq signals at senescence-associated loci, including key cell cycle genes (CCND1, STAG2, CDK6, and E2F5) and SASP genes (IL6, IGFBP2, MIF, and CXCL1) (Fig. 6g and Supplementary Fig. 19a). Moreover, compared with that in LU-RAS control cells, knockdown of JUN and GATAD2A increased the expression of these cell cycle genes and decreased the expression of the aforementioned SASP genes. Pharmacologic reduction of O-GlcNAc modification with OSMI-4 similarly increased the abundance of cell cycle gene transcripts while decreasing SASP mRNA levels (Fig. 6h and Supplementary Fig. 19b), closely mirroring the effects of JUN and GATAD2A knockdown in LU-RAS cells. Collectively, these findings indicate that O-GlcNAc modification of JUN and GATAD2A is important for their chromatin occupancy at SASP- and cell cycle arrest-related loci and that the O-GlcNAc modified forms rather than the non-glycosylated proteins modulate the transcriptional regulation of senescence-associated gene expression.
Attenuation of O-GlcNAc modification on JUN and GATAD2A impairs cellular senescence progression and SASP-associated tumor growth
Considering that the O-GlcNAc modification of JUN and GATAD2A affects the expression of senescence-associated genes, we investigated whether O-GlcNAc influences cellular functionality via the aforementioned mechanism. Compared with LU-RAS/KD-WT cells, LU-RAS/KD-AA cells, which expressed similar levels of JUNS84A and GATAD2AS182A, exhibited significantly decreased SA-β-Gal activity and increased capacities for cell cycle progression and proliferation at D10 of the OIS process (Fig. 7a, b). Consistent results were observed in other OIS cell models (Supplementary Fig. 20). One of the principal effects of OIS in cells is the promotion of tumor expansion and spread. Consequently, we examined whether the lack of O-GlcNAc modification on JUN and GATAD2A in OIS-affected fibroblasts could induce an increase in cancer cell proliferation, enlargement, and mobility in a Transwell coculture system. This system physically partitions senescent cells and cancer cells but permits them to share media and secrete soluble components such as SASP components (Fig. 7c). The reduction in O‑GlcNAc on JUN and GATAD2A attenuated certain senescence markers, including SASP production, in OIS-affected LU‑RAS cells (Fig. 7d, e). OIS-affected LU-RAS/KD-WT cells significantly stimulated the expansion, migration and invasion of A549 lung cancer cells through the secretion of SASP components. However, LU-RAS/KD-AA cells did not considerably influence these malignant phenotypes of cancer cells compared to those of non-cocultured A549 cells (Fig. 7f, g).
a Representative SA-β-Gal staining of LU-RAS (vehicle), LU-RAS/KD-WT, LU-RAS/KD-AA, SK-RAS (vehicle), SK-RAS/KD-WT, and SK-RAS/KD-AA cells at D10 of OIS. SK-RAS cells and either wild-type (Flag-GATAD2AWT, HA-JUNWT) or mutant (Flag-GATAD2AS182A and HA-JUNS84A) constructs were then reintroduced. Scale bars, 200 μm. b Flow cytometric analysis of the cell cycle distribution (G1, S, and G2 phases) in LU-RAS (vehicle), LU-RAS/KD-WT, LU-RAS/KD-AA, SK-RAS (vehicle), SK-RAS/KD-WT, and SK-RAS/KD-AA cells at D10 of OIS. c Schematic of transwell coculture experiments assessing the effects of SASP factors derived from LU-RAS, LU-RAS/KD-WT, and LU-RAS/KD-AA cells at D10 of OIS on A549 cell growth. d The relative levels of senescence conditioned medium multiple SASP factors at the OIS D10 time point in LU-RAS and its variant cell lines were quantified by ELISA, with the level in vehicle control cells set as 1 (fold change). e LU-RAS and its variant cells were induced to undergo OIS at D10 in the presence of doxycycline (2 μg/mL) under transwell coculture conditions. Whole-cell lysates were extracted and subjected to immunoblot analysis. The immunoblots are representative of three biologically independent experiments with similar results. f Counts of A549 cells cultured alone or cocultured with the indicated cells at D10 of OIS. Scale bars, 200 μm. g Images from the scratch assay of A549 cells at 0 h and 96 h; the cells were cultured alone or cocultured with the indicated cells at D10 of OIS. Scale bars, 200 μm. The data are presented as mean ± SD from four biological replicates with similar results. The p values were calculated using a two-tailed unpaired Student’s t test (a, b, d, f, g). The source data for the values and the graphs in (a, b, d, f, g) and immunoblots in (e) are provided in the Source Data File.
We next used immunocompromised (nude) mice to determine whether subcutaneous coinjection of A549 cancer cells (carrying the GFP gene) and OIS-affected LU-RAS cells expressing JUN and GATAD2A variants influenced tumor progression in vivo. All LU-RAS and its variant cells were marked with the cytoplasmic membrane dye NIR790, allowing us to visualize both the tumor cells and OIS-affected cells using an in vivo imaging system (Fig. 8a). The NIR790 images revealed that OIS-affected cells remained at the injection site within the growing tumor throughout the period during which we monitored tumor development (Fig. 8b, c). As predicted, compared with injection of only A549 cells, coinjection of A549 and LU-RAS cells led to increased tumor volume. In contrast, coinjection of LU-RAS/KD-AA cells did not increase tumor growth. Moreover, we found a substantial increase in tumor size when mice were coinjected with OIS-affected cells expressing JUNWT and GATAD2AWT, unlike the findings after coinjection of LU-RAS/KD-AA cells, as determined by the integrated density of GFP fluorescence and visual inspection of the extracted tumors (Fig. 8d, e). Immunoblot and IHC analyses revealed that compared with tumors from mice injected with LU‑RAS/KD‑AA cells, tumors from mice injected with LU‑RAS/KD‑WT cells exhibited increased expression of senescence markers (p16INK4A and p21WAF1/Cip1) and increased SA‑β‑Gal activity (Fig. 8f, g). Additionally, in the subcutaneous coinjection model—in which A549 tumor cells and OIS-affected LU‑RAS cells were present within the same lesions and could not be unequivocally distinguished by routine histology—we observed noticeable increases in MKI67 and LMNB1 (Fig. 8f), as well as IL6 and IGFBP2, in tissues from mice injected with LU‑RAS and LU‑RAS/KD‑WT cells, compared with those injected with LU‑RAS/KD‑AA cells. Meanwhile, in these tissues, the A549 tumor cells—exposed to SASP factors secreted by the senescent LU‑RAS cells—exhibited enhanced upregulation of the cell cycle–related factors CCND1 and STAG2, which constitute major components of the tumor cell population (Fig. 8g, h).
a Schematic of the coinjection model of D10 LU-RAS (vehicle), LU-RAS/KD-WT, or LU-RAS/KD-AA cells with A549 cells in nude mice. b, c Fluorescence images of GFP-expressing A549 cells injected alone or coinjected with LU-RAS (vehicle), LU-RAS/KD-WT, or LU-RAS/KD-AA cells (NIR790-stained) at D10 of OIS (n = 6 per group); images were acquired on days 5, 15, and 28. One representative image is shown (b). The relative fluorescence intensity was quantified (c). d Tumors were excised after 28 days. e Tumor volumes were measured. f Immunoblot analysis of senescence markers in xenograft tumor samples. n = 3 biologically independent samples were analyzed, and similar results were obtained across replicates. g, h Xenograft tumor samples were subjected to H&E, SA-β-Gal, IL6, IGFBP2, CCND1, and STAG2 staining. Scale bars, 100 μm. Representative images (g); quantitative analysis of mean IHC staining (h). The data are presented as mean ± SD from six biological replicates with similar results. p values were calculated using a two-tailed unpaired Student’s t-test (c, e, h). The source data for the graphs in (c, e, h) and immunoblots in (f) are provided in the Source Data File.
These results suggest that O-GlcNAc deficiency in these two key epigenetic factors can prevent the acquisition of the OIS phenotype, thereby inhibiting the stimulation of tumorigenesis both in vitro and in vivo. Hence, O-GlcNAc plays a specific role in controlling the detrimental behaviors of senescent cells, and targeting O-GlcNAc modified JUN and GATAD2A could constitute an alternative strategy for combating senescence-induced tumor development.
Discussion
Perturbations in cellular homeostasis, such as DNA damage, oncogene activation, oxidative stress, and organelle stress, can trigger premature senescence3,12. This state entails sustained and often irreversible growth arrest and serves as a key mechanism in tumorigenesis6. Tumor cells typically gain proliferative capacity by evading senescence, particularly OIS, which underpins immortalization and malignant progression52. Notably, senescence and cancer exhibit a bidirectional relationship: senescence-associated microenvironments can promote tumor initiation and progression, and OIS, while tumor suppressive, is frequently detected in premalignant and malignant lesions53,54,55. Therefore, identifying the regulatory mechanisms of senescence is essential for understanding tumor progression, identifying therapeutic targets and predicting outcomes in aging‑related cancers.
O-GlcNAc may act as both a sensor and a cytoprotective mechanism of senescence, enabling rapid modulation of key fate determinants to confer resistance to environmental fluctuations and exogenous stimuli56,57,58. Using a human RASG12V-induced in vitro senescence model39, we investigated the role of O-GlcNAc in OIS and tumorigenesis. In primary human fibroblasts, O‑GlcNAc levels increased along with the emergence of senescence phenotypes. During the OIS process, OGT and OGA expression fluctuated markedly, which may have contributed to this increase. However, direct suppression of O‑GlcNAc impeded OIS progression and attenuated the phenotype. This finding is in accordance with our results in prior chemoresistance models, in which OGT levels remained constant59. These data support a governing role for protein O-GlcNAc modification in OIS and highlight this modification as a potential antisenescence target with implications for aging-related diseases, including cancer. Notably, the level of OGT mRNA decreased by approximately twofold during OIS progression (Supplementary Data 2), but the level of the OGT protein increased. The results of cycloheximide chase and MG132 assays revealed that the half-life of OGT was prolonged during the OIS process and that the protein was further stabilized by proteasome inhibition (Supplementary Fig. 4c). Moreover, pharmacologic modulation of global O-GlcNAc did not appreciably alter OGT abundance (Fig. 1c, d). This may reflect compensatory regulation elicited by enzyme inhibition that is highly dependent on cell type and stress context and may be manifested primarily at the level of UDP-GlcNAc substrate flux rather than as a change in the OGT protein level. Conversely, although OGT itself can be O-GlcNAc modified60,61, our data indicate that OIS-associated OGT stabilization and the resulting increase in protein abundance are largely independent of O-GlcNAc levels. Future work should address the contrasting trajectories of OGT and OGA levels during OIS progression to clarify how O-GlcNAc levels are specifically regulated in this process.
Approximately half of the >7000 intracellular O-GlcNAcylated proteins localize to the nucleus, where they contribute to chromatin function and gene regulation62,63. Chromatin-binding factors (TFs/cofactors and remodeling complexes) frequently interact with OGT and are O-GlcNAcylated57,64,65. However, the mechanisms linking chromatin O-GlcNAc to transcriptional reprogramming in cell fate decisions remain elusive. We employed a chemical genomics approach56,66 to enrich O‑GlcNAc associated chromatin complexes and, using a dynamic multiomics approach, mapped how O‑GlcNAc–regulated gene expression drives OIS phenotypes. We established a time-resolved model of chromatin O-GlcNAc regulated transcriptional reprogramming, revealing an unrecognized mechanism of O-GlcNAc mediated transcriptional control during senescence. A central finding was that during OIS, O-GlcNAc chromatin proteins (OCPs) assemble distinct epigenetic regulatory complexes. Two major classes were enriched: a TF-SWI/SNF complex that binds promoters (including those of SASP genes) to activate transcription and a NuRD complex that associates with enhancers to repress the transcription of cell cycle and proliferation genes. Global or site-specific suppression of O-GlcNAc impaired the assembly of these complexes during OIS. Together with the results of molecular dynamics simulations showing that O‑GlcNAc facilitates component interactions, these results establish O‑GlcNAc as being integral to both complex assembly and transcriptional activity. Under O-GlcNAc mediated control, these complexes coordinate the time-dependent expression of OIS-related genes, revealing a more intimate mechanism by which O-GlcNAc regulates senescence and the acquisition of the SASP18,67. Similar O-GlcNAc based mechanisms may operate in other physiological and pathological contexts to shape cell fate.
We further observed temporal redistribution of OCPs across the genome during the OIS process, coordinated with classical epigenetic marks (histone modifications and DNA methylation). A subset of OCPs transitioned among distinct promoters, thereby potentially activating target genes; another subset was redistributed from enhancer-like regions bearing transcriptionally active histone marks to a state characterized by transcriptionally repressive DNA hypermethylation. This switch appears to be linked to O-GlcNAc modified GATAD2A-mediated recruitment of MBD2/CHD4 to a sub-NuRD complex. The distributional switch suppresses cell cycle and proliferation genes, thereby inducing proliferative arrest, whereas O-GlcNAc modified JUN-mediated recruitment of a sub-SWI/SNF complex undergoes transitions among promoters, with its targets evolving to drive SASP gene activation. Consequently, OIS affected cells exhibit increased secretion of IL6 and IGFBP2. Moreover, we observed that different cell types enter distinct phases of cell cycle arrest following OIS induction. Regardless of which phase of the cell cycle is arrested, the core molecular mechanism is attributed to the transcriptional repression of various cell cycle factors mediated by O-GlcNAc modification. In addition to supporting the O-GlcNAc facilitated assembly of functionally distinct transcriptional complexes, these findings underscore the presence of synergistic crosstalk between chromatin O-GlcNAc and other epigenetic marks. We propose that chromatin O-GlcNAc represents an epigenetic code that acts via the glycosylation of chromatin-bound regulators and TFs; by modifying readers of DNA methylation and histone marks, O-GlcNAc alters their activity and interactions, thereby facilitating or impeding transcriptional regulation. Thus, O-GlcNAc may serve as an adapter of epigenetic signals and function as a downstream layer of these classical epigenetic codes.
Finally, we constructed a time-resolved, highly dynamic model of the OIS gene expression network controlled by OCPs, emphasizing the combinatorial O-GlcNAc modified TF interactome. In addition to recognized senescence genes, we identified candidates not previously linked to OIS. Among the SASP-related inflammatory factors that strongly influence tumorigenesis8,9, O-GlcNAc modified JUN is a prominent contributor. Concurrently, glycosylated GATAD2A, a nexus for NuRD-mediated repression43,44,45, plays a key role in cell cycle arrest. Although we mapped temporal O-GlcNAc landscapes on OCPs during OIS, the glycosylation of two factors, JUN and GATAD2A, was found to be able to partially reset the senescence clock. As an AP-1 family TF, JUN is essential for senescence and proliferation39; here, we showed that O-GlcNAc modified JUN rather than unmodified JUN orchestrates OIS and SASP acquisition, a process critical for OIS-driven malignant progression in vitro and in vivo. Accordingly, glycosylated factors such as JUN and GATAD2A are promising antisenescence targets; selectively inhibiting the O-GlcNAc modification of these factors—by developing substrate-specific O-GlcNAc inhibition techniques—may offer a feasible strategy to modulate senescence phenotypes.
In conclusion, in the present multiomics study, we elucidated how O-GlcNAc cooperates with other epigenetic components to regulate gene expression during cellular senescence, revealing previously unrecognized epigenetic functions of O-GlcNAc and its control over the temporal dynamics of gene expression networks. The temporal increase in chromatin O-GlcNAc and the glycosylation of key OCPs play important roles in the acquisition of OIS phenotypes (Fig. 9). These findings provide a rationale for exploring pharmacological targets for cancer therapy within a senescence-based paradigm.
Cellular senescence triggers an increase in O-GlcNAc levels and alters OCP genomic binding dynamics. O-GlcNAc modifications on JUN and GATAD2A support the assembly of distinct epigenetic complexes (TF-SWI/SNF and NuRD), which regulate SASP acquisition and cell cycle arrest, ultimately driving tumor progression both in vitro and in vivo.
Methods
This research was conducted in accordance with all applicable ethical guidelines and regulations and was approved by the Ethics Committee of Dalian University of Technology under approval number DUTSCEOLS240228-05.
Cell culture and reagents
LU1015 human primary lung fibroblasts (#T1015) and SK0304 human primary benign skin fibroblasts (#T0304) were purchased from Applied Biological Materials Inc. (abm) (Richmond, Canada). Normal human epithelial cell lines (MCF10A and HaCaT) and HEK-293T (293T) cells and A549 lung cancer cells were obtained from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China) and were used within 6 months of resuscitation. LU1015 and their corresponding cells were cultured in medium PriGrow VII (abm, #TM017), 10% fetal bovine serum (FBS) and 1% Penicillin/Streptomycin Solution (Gibco). SK0304 and their corresponding cells were cultured in medium PriGrow III (abm, #TM003), 10% FBS and 1% Penicillin/Streptomycin Solution (Gibco). MCF10A and their corresponding cells were cultured in Complete Medium (Procell, #CM-0525) and 100 ng/mL cholera toxin (Sigma, #C8052). 293T and HaCaT cell lines were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% FBS and 1% penicillin/streptomycin (Gibco). A549 lung cancer cells were cultured in RPMI-1640, 10% FBS and 1% Penicillin/Streptomycin Solution (Gibco). All Cells were maintained at 37 °C in a humidified atmosphere with 5% CO2 and were routinely tested for Mycoplasma contamination on a monthly basis using the PlasmoTestTM Mycoplasma Detection Kit (InvivoGen). OSMI-4 (#S8910) and Thiamet-G (#S7213, TMG) were purchased from Selleck, and doxycycline were purchased from Sigma-Aldrich. Other reagents were used as analytic grade or better.
Plasmids, lentiviral production, and transfection
LU1015 and SK0304 fibroblasts were transduced with a recombinant lentiviral vector carrying the hTERT gene (Applied Biological Materials Inc.) to achieve cellular immortalization. At 72 h post-transduction, cells were cultured in growth medium supplemented with puromycin to select for stable hTERT-transduced clones. To generate a doxycycline-inducible Tet-ON human RASG12V expression system, a synthetic gene encoding the human RASG12V coding sequence was cloned into the lentiviral pLVX-Tet3G vector34 (Addgene #28061), which was modified to introduce a G418 resistance selection marker, resulting in the pLVX-Tet3G-RASG12V-G418 plasmid. Lentiviral particles were produced by transient co-transfection of 293T cells with pLVX-Tet3G-RASG12V-G418 and appropriate packaging plasmids (psPAX2 and pMD2.G) using the calcium phosphate transfection method. After 48–72 h, viral supernatants were collected, filtered, and used for cell infection. LU-RAS, SK-RAS, MCF-RAS, and HaC-RAS cells were infected with the pLVX-Tet3G-RASG12V-G418 lentivirus. Following infection, stable monoclonal populations of infected cells were generated by limiting dilution and subsequent selection in G418-containing medium. Senescence was induced by treating cells with 2 μg/mL doxycycline, and samples were collected at the specified time points for further analysis.
Lentiviral shRNA plasmids targeting GATAD2A and JUN were constructed using the pLKO.1-Hygro vector (Addgene, #24150), which was modified to enable simultaneous knockdown of these two genes. Specifically, an additional U6 promoter was introduced into the vector downstream of the GATAD2A shRNA cassette, and the JUN shRNA sequence was cloned under this promoter. This tandem shRNA expression system ensures that each shRNA is expressed independently. Two distinct shRNAs (shRNA-1, shRNA-2) were designed for both GATAD2A and JUN, and the sequence information is provided in the Supplementary Data 8. Hygromycin was used in the subsequent lentiviral transduction selection process to obtain monoclonal cells (LU-RAS/KD). Lentiviral shRNA plasmids for OGT (#sc-40780- V, Santa Cruz) were used for gene knockdown. Scrambled shRNA plasmid (#1864) was purchased from Addgene.
We constructed a lentiviral vector based on the pLenti-CMV-MCS-blank backbone (Addgene, #17448) to co-express full-length wild-type human GATAD2A, JUN, and O-GlcNAcylation site mutants (S182 → A or S625 → A for GATAD2A, and S84 → A for JUN), with Zeocin as the selection marker. The original antibiotic resistance gene was replaced by the Zeocin resistance gene (ble). To enable the simultaneous expression of the two genes, 3×Flag-GATAD2A and HA-JUN were linked via a self-cleaving P2A peptide under the control of the CMV promoter. The 3×Flag and HA tags were placed at the C-terminus of GATAD2A and JUN, respectively. LU-RAS/KD-WT (stably tandem expression of Flag-GATAD2AWT and HA-JUNWT) and LU-RAS/KD-AA (stably tandem expression of Flag-GATAD2AS182A and HA-JUNS84A) cells were generated via lentiviral transduction, followed by Zeocin selection. The truncated mutants of Flag-GATAD2A and HA-JUN were subcloned into pCMV vector. The construction of pcDNA-OGT was reported previously59.
Immunoblot/lectin blot and co-immunoprecipitation (co-IP)
For immunoblot/lectin blot, cells were collected and lysed using Western/IP lysis buffer (Beyotime, #P0013) supplemented with protease and phosphatase inhibitor cocktails (Roche) for 20 min on ice. Protein concentrations were determined using the Bradford assay (Bio-Rad). Equivalent protein amounts were prepared by mixing the lysates with SDS-PAGE loading buffer, followed by boiling at 95 °C for 5 min. Proteins were separated by SDS-PAGE and subsequently transferred onto a polyvinylidene fluoride (PVDF) membrane (Immobilon-P, Millipore-Sigma). The membrane was then blocked with 5% non-fat milk for 1 h to prevent non-specific binding. Primary antibodies were applied overnight at 4 °C. After thorough washing to remove unbound antibody, the membrane was incubated with HRP-conjugated secondary antibodies at for 1 h.
The primary anti-bodies used were anti-O-GlcNAc CTD110.6 (CST, #9875, 1:1000), RasG12V (CST, #14412, 1:1000), anti-JUN (Abcam, #ab40766, 1:1000), anti-GATAD2A (CST, #17705, 1:1000), anti-OGT (Abcam, #ab96718, 1:1000), anti-OGA (CST, #60406, 1:1000), anti-GAPDH (CST, #5174, 1:4000), anti-Flag (CST, #14793, 1:1000), anti-HA (CST, #3724, 1:1000), anti-Histone 3 (CST, #4499, 1:4000), MBD2 (Abcam, #ab188474, 1:1000), MTA1 (Proteintech, #30545-1-AP, 1:1000), CHD4 (Proteintech, #14173-1-AP, 1:1000), SMCA4 (Proteintech, #21634-1-AP, 1:1000), SMRC1 (Proteintech, # 17722-1-AP, 1:3000), SNF5 (Abcam, #ab307985, 1:1000), p53 (Proteintech, #10442-1-AP, 1:4000), p21WAF1/Cip1 (Proteintech, #10355-1-AP, 1:1000), p16INK4A (Proteintech, # 10883-1-AP, 1:1000), LMNB1 (Proteintech, #12987-1-AP, 1:5000), MKI67 (Proteintech, #84192-4-RR, 1:5000), SP1 (Proteintech, #21962-1-AP, 1:5000), SRSF2 (Proteintech, #20371-1-AP, 1:5000), β-actin (Proteintech, #20536-1-AP, 1:5000), TERT (Proteintech, #27586-1-AP, 1:500), FOXK1 (Proteintech, # 29338-1-AP, 1:2000). Lectin sWGA (Vector Laboratories, #B-1025S, 1:2000) was used for lectin blotting. The appropriate secondary antibody used were anti-mouse IgG-HRP (CST, #7076, 1:20,000), anti-rabbit IgG-HRP (CST, #7074, 1:20,000), anti-mouse IgM-HRP (Abcam, #ab97230, 1:20,000), Streptavidin-HRP (CST, #3999, 1:50,000), and the signals were detected by the ECL Plus kit (GE Health-care). All immunoblot experiments were conducted in three biological replicates.
For co-IP, cells were collected and lysed in Western/IP lysis buffer supplemented with protease and phosphatase inhibitor cocktails (Roche) on ice for 20 min. The lysates were then centrifuged at 20,000 × g for 10 min at 4 °C to remove cellular debris. The resulting supernatant was incubated with primary antibodies and protein A/G magnetic beads (Bimake, #B23201) overnight at 4 °C under gentle rotation. After incubation, the immunoprecipitates were washed five times with ice-cold Western/IP lysis buffer to minimize non-specific interactions. The samples were then prepared for immunoblot analysis. The antibodies and beads used for IP were anti-JUN (Abcam, #ab40766, 1:50), anti-GATAD2A (CST, #17705, 1:50), anti-O-GlcNAc CTD110.6 (CST, #9875, 1:1000), anti-O-GlcNAc EPR19847 (Abcam, #ab202665, 1:50), anti-Flag-magnetic beads (Bimake, #B26102), and anti-HA-magnetic beads (Bimake, #B26202). For lectin pull down assay, each cell lysate was incubated with sWGA bound agarose beads (Vector Laboratories, #AL-1023S). All samples utilized the same volume (60 μL) of magnetic beads to ensure consistency in experimental conditions, thereby guaranteeing the reliability of IP results. To avoid interference from IgG, antibodies from different species were used in the immunoblotting detection following the co-IP experiments.
To confirm the O-GlcNAc modification of OCPs and eliminate the possibility of contamination from O-GlcNAc interacting proteins in the immunoprecipitated samples, cells were lysed in 1× SDS lysis buffer (Beyotime, #P0013G) by heating at 95 °C for 15 min. The denatured samples were centrifuged, and the supernatant was diluted 15-fold with Western/IP lysis buffer. The diluted lysates were then subjected to immunoprecipitation using specific antibodies or anti-Flag/anti-HA-magnetic beads. Immunoprecipitates were subsequently analyzed using the anti-O-GlcNAc antibody.
Chromatin complexes capture
Cells (1 × 107) were washed with PBS and crosslinked with 1% formaldehyde for 15 min to stabilize proteins interacting with chromatin for subsequent immunoblot, proteomic, and DNA cis-element analyses. The crosslinking reaction was quenched with 2.5 M glycine. Cells were scraped from culture plates, pelleted by centrifugation, and resuspended in 1 mL hypotonic buffer (10 mM HEPES, pH 7.9; 10 mM KCl; 1.5 mM MgCl2, 0.34 M sucrose; 10% glycerol (v/v); 0.4% Igepal CA-630 (v/v); 1 mM DTT), supplemented with protease and phosphatase inhibitor cocktails, to disrupt the plasma membrane. Nuclei were isolated by centrifugation (use the supernatant as the cytoplasm fraction), gently washed with nuclear extraction buffer (20 mM HEPES, pH 7.9; 150 mM NaCl; 1.5 mM MgCl2; 0.2 mM EDTA; 10% glycerol; 0.4% Igepal CA-630 (v/v); protease and phosphatase inhibitors), incubated for 5 min, and centrifuged at 1000 × g. The intermediate salt strength (150 mM NaCl) at this step is intended to increase ionic strength sufficiently to remove soluble nuclear constituents and weakly associated proteins, thereby partitioning soluble nuclear proteins and weakly bound complexes into the supernatant (nucleoplasm) while retaining chromatin-bound material in the pellet. Subsequently, nuclei were lysed in nuclear lysis buffer (50 mM HEPES, pH 7.9; 150 mM NaCl; 1.5 mM MgCl2; 1% Igepal CA-630 (v/v); 0.1% SDS (w/v)) containing protease and phosphatase inhibitors, and the lysates were incubated on a rotating platform at 4 °C for 1 h. Chromatin was then fragmented by sonication (Sonics VCX130pb) until DNA was sheared to ~200–500 bp, followed by centrifugation at 20,000 × g at 4 °C for 15 min to remove nuclear membranes and scaffolding components. The crosslinked chromatin complexes were collected from the resulting supernatant and used for downstream assays. For detection of chromatin-associated proteins by immunoblotting, the isolated chromatin complexes were treated with benzonase nuclease (5 µg/mL; HaiGene) and RNase A (10 µg/mL; Sigma) prior to protein analysis.
Cell proliferation assay
Approximately 2000 cells per well were seeded into a 96-well plate and treated with either the vehicle control or 2 μg/mL doxycycline for 2–10 days, with or without OSMI-4 (5 μM) or TMG (50 μM), for 48 h. Cell proliferation was assessed using the Enhanced Cell Counting Kit-8 (CCK-8, Beyotime, #C0041) following the manufacturer’s protocol. Absorbance at 450 nm was recorded for each well and normalized to the vehicle-treated control. All experiments were conducted in four biological replicates.
Transwell cancer growth and scratch experiments
LU-RAS and its derivative cells were grown to 70–80% confluency on Transwell permeable supports (Costar). Ten days after senescence induction, 5 × 104 A549 cells were seeded in 6-well plates, and Transwell supports containing no cells or LU-RAS and its derivative cells were placed above the A549 cells. After incubation, cell numbers were counted under a microscope. For Transwell in vitro scratch assays, A549 cells were plated in 6-well plates and cultured to confluency. A scratch was created using a P200 pipette tip, and cell debris was removed by washing with fresh medium. The medium was replaced, and the initial scratch area was imaged. LU-RAS and its derivative cells medium were then added to the culture plates. The scratches were re-imaged 96 h later using a phase-contrast microscope.
EdU staining and senescence-associated beta galactosidase activity
For EdU staining, representative samples from the senescence time series were analyzed for EdU incorporation using the Click-iT EdU Alexa Fluor Imaging Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions. Cells were seeded in 6-well plates and cultured under senescence-inducing conditions to the indicated time points (D0, D2, D4, D10). Immediately prior to fixation, cells were pulsed with EdU for 2 h. After incubation, the medium was aspirated, and cells were washed twice with PBS and fixed with paraformaldehyde. Cells were then washed three times with washing buffer (PBS containing 3% BSA) and permeabilized with 0.3% Triton X-100 for 15 min. Following three additional washes with washing buffer, cells were incubated with the Click Reaction Mixture. For nuclear counterstaining, Hoechst dye was added at a final concentration of 1 μg/mL and incubated for 10 min at 25°C in the dark. Excess dye was removed by washing with PBS prior to imaging. Cells exhibiting both green (EdU) and blue (Hoechst) signals were scored as EdU-positive. Regardless of the specific phase in which cells are arrested, once they enter cell‑cycle arrest they lose proliferative capacity and no longer incorporate EdU. Thus, a lack of EdU incorporation (negative EdU staining) is indicative of senescent cells.
For SA-β-Gal activity assays, cells were washed with PBS and fixed in a solution containing 2% formaldehyde and 0.2% glutaraldehyde for 10 min at 25 °C. After fixation, cells were incubated with freshly prepared SA-β-Gal staining solution (40 mM citric acid/sodium phosphate buffer, pH 6.0, 1 mg/mL X-gal, 150 mM NaCl, 2 mM MgCl2, 5 mM potassium ferrocyanide, and 5 mM potassium ferricyanide) at 37 °C without CO2 until visible blue staining was detected. Imaging was performed using a Leica confocal fluorescence microscope.
Flow cytometry
Cells were seeded in 6-well plates and cultured under conditions to induce senescence as previously described, with or without the addition of OSMI-4 (5 μM) or TMG (50 μM) for 48 h. The cells were then fixed in 70% (v/v) ethanol overnight at −20 °C. Following fixation, cells were washed with PBS and incubated with PBS containing RNase A (Sigma) for 30 min at 37 °C in the dark. Subsequently, cells were stained with 5 μL of propidium iodide (PI, Sigma) for 30 min at 25 °C in the dark. Flow cytometry analysis was then performed using a BD flow cytometer and the ModFit LT (Version4.1) was used to analyze Flow Cytometry Data. The flow cytometry gating strategy is presented in Supplementary Fig. 21. All experiments were conducted in four biological replicates.
Gel filtration-based fractionation of nuclear extract
To analyze the co-migration of NuRD or TF-SWI/SNF components, nuclear extracts were prepared from LU-RAS/KD-WT and LU-RAS/KD-AA cells. A total of 0.3 mg of nuclear extract was treated with Benzonase for 1 h followed by centrifugation at maximum speed for 10 min at 4 °C. The supernatant was then loaded onto a Superose 6 10/300 column pre-equilibrated with a buffer containing 20 mM Tris (pH 7.5), 300 mM NaCl, 2 mM MgCl2, 5% glycerol, 1 mM PMSF, and complete protease inhibitors. Fractions of 0.25 mL were collected and precipitated using trichloroacetic acid. The resulting protein pellets were resuspended in SDS-PAGE loading buffer, boiled, and analyzed by western blot, with every other fraction being loaded for electrophoresis. All experiments were conducted in four biological replicates.
Metabolic labeling and click reaction
OIS cells at different senescence time points in 10-cm dishes were cultured in medium supplemented with 100 μM 1,6-Pr2GalNAz (generously provided by Prof. Bo Chen, Peking University) for 48 h. Approximately 3 × 107 cells were harvested by trypsinization and washed three times with PBS. Crosslinked chromatin was prepared from metabolically labeled cells as described above. To specifically biotinylate O-GlcNAz modified proteins, the chromatin was incubated with 100 μM alkyne-biotin (Click Chemistry Tools, #1266-25), 300 μM of a premixed CuSO4-BTTAA (Click Chemistry Tools, #1236-500) complex (molar ratio 1:2), and 2.5 mM freshly prepared sodium ascorbate at 25 °C for 2 h. Methanol (10 mL) was added to the reaction and the mixture was stored at −80 °C overnight. Precipitated proteins were pelleted by centrifugation at 10,000 × g for 15 min at 4 °C, washed twice with ice-cold methanol, and resuspended in 1 mL of recovery buffer. To ensure thorough decrosslinking of remote PPI, samples were heated at 100 °C for 20 min followed by 60°C for 2 h, minimizing crosslink‑driven co‑capture and nonspecific recovery of non‑O‑GlcNAz proteins68. The resulting solution was diluted 15-fold with Western/IP lysis buffer, pre-cleared with 100 μL of vehicle-magnetic beads (BEAVER Life Science, #70301), and incubated with 200 μL of streptavidin-magnetic beads (BEAVER Life Science, #22308) at 25 °C for 4 h on a rotating platform to capture OCPs. To minimize nonspecific binding, the beads were stringently washed five times with low-salt buffer (20 mM Tris-HCl, pH 8.1, 0.1% SDS, 2 mM EDTA, 1% Triton X-100, 150 mM NaCl), five times with high-salt buffer (buffer composition identical to low-salt but containing 500 mM NaCl), and twice with LiCl buffer (10 mM Tris-HCl, pH 8.0, 0.25 M LiCl, 0.5% NP-40, 1% sodium deoxycholate, 1 mM EDTA). Finally, the beads were resuspended in SDS-PAGE loading buffer for immunoblot analysis of OCPs. For LC-MS/MS analysis, the beads were resuspended in 500 μL of 6 M urea in PBS. Protein samples were reduced with 10 mM DTT in water at 65 °C for 15 min and alkylated with 20 mM iodoacetamide in water at 35 °C in the dark for 30 min. The buffer was subsequently exchanged with 200 μL of 2 M urea in PBS, after which 4 μL of trypsin (Promega, 0.5 μg/μL) and 2 μL of 100 mM CaCl2 were added to digest the proteins. Digestion was performed at 37 °C for 16 h, and the resulting peptides were used for LC-MS/MS analysis.
For omics-level mapping of O-GlcNAc sites, OIS cells at various senescence time points were metabolically labeled with 100 μM 1,6-Pr2GalNAz (generously provided by Prof. Bo Chen, Peking University). After lysis, 1 mL of crosslinked chromatin (2 mg/mL) prepared from the labeled cells was incubated at 25 °C for 2 h with 300 μM of a premixed CuSO4-BTTAA complex (molar ratio 1:2), 100 μM alkyne-PC-biotin (Vector Laboratories, #CCT-1118), and 2.5 mM freshly prepared sodium ascorbate. The protein pellets obtained by centrifugation were resuspended in 300 μL of 8 M urea in water. The suspensions were then diluted to 4 M urea with 100 mM ammonium bicarbonate (ABC) and reduced with 10 mM DTT at 65 °C for 15 min, followed by alkylation with 20 mM iodoacetamide at 35 °C in the dark for 30 min. The solution was further diluted to 0.8 M urea using 50 mM ABC and digested with trypsin at a 1:50 enzyme-to-substrate ratio (w/w) at 37 °C for 16 h. After digestion, the samples were diluted to 0.5 M urea with 50 mM ABC and incubated with 100 μL of streptavidin-magnetic beads at 25 °C for 4 h. The beads were washed six times with PBS (pH 7.4) and six times with Milli-Q water. Subsequently, the beads were resuspended in 200 μL of 0.1% formic acid (v/v) in water and irradiated with UV light (9 J/cm2, 365 nm) using a UV cross-linker (#CL-1000, UVP). After removing the streptavidin beads, the samples were evaporated to dryness and analyzed by LC-MS/MS.
LC-MS/MS analysis and label-free quantification
The digested peptides were analyzed using a timsTOF Pro2 mass spectrometer (Bruker Daltonics) coupled to a nanoElute HPLC system. Peptides were first loaded onto a 300 μm i.d. ×5 mm C18 trap column (μ-Precolumn, Thermo Scientific) and subsequently separated on a 75 μm i.d. ×25 cm analytical column packed in-house with 1.8 μm C18 reverse-phase particles. Separation was performed using a 60-min gradient at a flow rate of 300 nL/min, with buffer A (0.1% formic acid in water) and buffer B (0.1% formic acid in acetonitrile): 2% B to 22% B in 45 min, followed by an increase to 35% B in 10 min, then to 80% B in 5 min, held at 80% B for another 5 min. Each sample was analyzed in nine replicates for robust peptide identification. Data acquisition was performed in DIA (data-independent acquisition) mode using the dia-PASEF method. The method consisted of a single MS1 scan followed by twelve dia-PASEF scans, utilizing 28 variable mass windows of 25 Da each, covering an m/z range of 452-1152. Each mass window was further split into two ion mobility windows, resulting in 56 total acquisition windows. The ion mobility range spanned 0.75–1.4 Vs/cm2. The ion source voltage was set at 1700 V, and ions were fragmented via CID with collision energy ranging from 20 eV at 1/K0 of 0.6 Vs/cm2 to 59 eV at 1/K0 of 1.6 Vs/cm2. DIA files were processed using Spectronaut software with the direct DIA workflow and default settings. Parameters included specifying Trypsin/P as the digestion enzyme, permitting peptide lengths between 7 and 52 amino acids with up to two missed cleavages, and incorporating fixed carbamidomethylation of cysteine along with variable modifications, such as methionine oxidation and protein N-terminal acetylation. A 1% FDR was set for both peptide-spectrum match (PSM) and protein identifications.
For the qualitative analysis of OCPs, representative proteins were determined if they were identified in at least six out of nine replicates analyzed, and only proteins annotated as nuclear were considered OCPs. For quantitative analysis based on DIA data, label-free quantification (LFQ) was performed using non-labeled peptide intensities. Protein quantification was based on the summed intensities of unique and razor peptides with a minimum of two quantified peptides per protein group. Missing values were imputed using a normal distribution with a downshift of 1.8 standard deviations and a width of 0.3 standard deviations to account for low-intensity features. Proteins identified in at least six replicates across nine samples in at least one experimental group were included for further statistical processing. Differentially abundant proteins were identified based on a fold change ≥2 and a p ≤ 0.05 (two-sided unpaired Student’s t test) after correcting for multiple hypothesis testing using the Benjamini-Hochberg false discovery rate (FDR) set at 0.05. To assess the reproducibility of biological replicates, PCA was performed. Functional annotation and GO enrichment analyses were conducted using Metascape software (v3.5)69. Protein-protein interaction networks were constructed and visualized using STRING (v11.0)70 and Cytoscape software (v3.6.1)71. Additional bioinformatic analyses were carried out using the statistical software R (v3.4.3) with several packages, including beanplot (v1.2), ggplot2 (v3.0.0), igraph (v1.2.1), and venneuler (v1.1-0).
Protein O-GlcNAc sites were acquired from an Orbitrap Fusion Lumos mass spectrometer (Thermo). The resuspended samples were separated with a 75 µm × 50 cm EasySpray reversed-phase LC column (Thermo) packed by 100 Å, 2 µm, PepMap C18 particles. A Dionex Ultimate 3000 RPLC nano system (Thermo) was employed to deliver the following LC gradient: 1–7% B in 9 min, 7-35% B in 302 min, 35–44% B in 42 min, 44–99% B in 3 min (A is water with 0.1% formic acid and B is 80% acetonitrile with 0.1% formic acid). A Full MS scan (350–2000 Th) was obtained with a maximum injection time of 50 ms, a resolution of 120,000, and RF lens at 60%. Monoisotopic precursor selection was enabled, and the multi-charged precursors with z = 2–8 were selected for data-dependent MS/MS scans. HCD product-dependent electron-transfer/higher-energy collisional dissociation (HCD pd EThcD) was used for MS/MS fragmentation. If a HCD scan produced one of the following glycan oxonium ions (m/z 168.0655, 186.0761, 204.0865, 274.0921, 292.1027, 300.1302, 329.1455, 366.1395, 388.1463, 399.1992, and 405.213), a separated EThcD scan was performed on the captured precursor. The identification and quantification of O-GlcNAc glycopeptides were performed using MaxQuant software integrated with the Andromeda search engine (version 1.5.3.28). MS/MS spectra were searched against the SwissProt human database (20,379 sequences), which was concatenated with a ReverseDecoy database and a list of common contaminants. The search parameters for O-GlcNAc site identification followed previously established criteria64. Specifically, UV-HexNAc modified sites (resulting from UV cleavage of alkyne-PC-biotin fragments, m/z 299.12) with a localization probability above 0.75 on confidently identified UV-HexNAc modified peptides (FDR of 1%, Andromeda score > 40, and delta score >8) were classified as unambiguous O-GlcNAc sites. Extracellular sites with UV-HexNAc modifications were excluded from the final O-GlcNAc site list. For quantification of O-GlcNAc sites, LFQ was applied to measure ion intensities of peptides containing UV-HexNAc modifications. Only unambiguous O-GlcNAc sites that were quantified in at least two out of three biological replicates were selected for downstream quantitative analyses, as described previously. The raw mass spectral data obtained in our study are available via iProX with the identifier PXD063140.
Immunofluorescence
Cells grown on coverslips were fixed with 4% paraformaldehyde for 10 min at 25 °C, followed by permeabilization with 0.1% Triton X-100. After washing with PBS, the cells were blocked with 5% goat serum for 30 min to prevent nonspecific binding. Primary antibodies were then applied and incubated overnight at 4 °C. The following antibodies were used: anti-Flag (CST, #8146, 1:500), anti-HA (CST, #2367, 1:500), anti-MBD2 (Abcam, #ab188474, 1:500), and anti-SNF5 (Abcam, #ab192864, 1:100). Then the cells were washed and incubated with fluorophore-conjugated secondary antibodies, including both anti-mouse IgG H&L (Alexa Fluor 488, CST, #4408) and goat anti-rabbit IgG H&L (Alexa Fluor 647, CST, #4414), at 25 °C for 2 h. Nuclear DNA was stained with 4′,6-diamidino-2-phenylindole (DAPI). Images were acquired using a Nikon Eclipse Ti confocal microscope with Nikon Eclipse C2 software (v5.21.00). Image resolution is 1024 × 1024 pixels. Images were captured using a 100× oil immersion objective. All experiments were conducted in four biological replicates.
COGC-seq/ChIP-seq and bioinformatics
COGC-seq assays were carried out as previously described56, using ~3 × 107 cells per experiment. Crosslinked chromatin complexes were isolated from 1,6-Pr2GalNAz metabolically labeled cells and biotinylated as detailed above. Biotinylated O-GlcNAz modified chromatin complexes were pre-cleared and then captured by incubating with 200 μL of streptavidin magnetic beads at 25 °C for 4 h. Stringent washes were conducted using low-salt, high-salt, and LiCl wash buffers, following the protocol for OCP enrichment. The beads were resuspended in 100 μL of TE buffer, followed by digestion with RNase (10 μg/mL, Sigma) at 37 °C for 30 min. SDS was added to a final concentration of 0.5%, and the complexes were de-crosslinked overnight at 65 °C with Proteinase K (0.2 mg/mL). DNA was then purified using the MiniBEST DNA Fragment Purification Kit (Takara, #9761) and sequenced using the Illumina NovaSeq 6000 platform (Illumina, at Novogene Technology, China) and/or a Salus Progene sequencer (Shenzhen Salus BioMed Co., Ltd, China). Two biological replicates were performed for the sequencing.
ChIP-seq assays for Flag-GATAD2AWT, Flag-GATAD2AS182A, HA-JUNWT, HA-JUNS84A, H3K27ac, H3K27me3, H3K4me1, and H3K4me3 were performed as described26. In brief, crosslinked chromatin complexes were isolated from ~3 × 107 cells and sonicated using a Sonics Vcx130pb. After pre-clearing with vehicle-coated magnetic beads, chromatin complexes were immunoprecipitated with either anti-Flag magnetic beads, anti-HA magnetic beads, or antibodies against histone modifications, including H3K27me3 (CST, #9733, 1:50), H3K27ac (CST, #8173, 1:50), H3K4me1 (CST, #5326, 1:50), or H3K4me3 (CST, #9751, 1:50). Protein A/G magnetic beads were added, and the samples were incubated at 4 °C for 4 h. Chromatin-bead complexes were then washed extensively, five times each with low-salt and high-salt buffers, followed by two washes with LiCl buffer at 4 °C. De-crosslinking was performed using Proteinase K (Sigma), and the resulting DNA was purified. Purified DNA was utilized for either qPCR or next-generation sequencing, which was carried out using the Illumina NovaSeq 6000 platform (Illumina) at Novogene Technology (Beijing, China). Each experiment was conducted in two biological replicates.
The raw data for COGC-seq and ChIP-seq are available in the GEO database under the accession number GSE295087. Fastp (v0.23.1) was used to trim reads with the parameters set as follows: length_required = 50, n_base_limit = 6, poly_x_min_len = 10. Quality control of the clean reads was performed using FastQC (v0.11.8), and the quality of the filtered data was assessed. Clean sequencing datasets were aligned to the human reference genome (hg19) using Bowtie2 (v2.3.4.3) and processed with Samtools (v1.2). Picard was used to remove duplicate reads with default parameters. Next, Macs2 (v2.2.9.1) was used for peak calling with the parameters set, COGC-seq as follows: -f BAMPE -g hs -B -p 0.01. H3K27me3 and H3K4me1 ChIP-seq as follows: -f BAMPE -g hs -B -broad -broad-cutoff 0.05 -qvalue 0.05. H3K27ac and H3K4me3 ChIP-seq as follows: -f BAMPE -g hs -B -q 0.05. The resulting BAM files were converted into BigWig format using the bamCoverage tool from deepTools (v3.3.2.0.0) and normalized by the reads per kilobase per million mapped reads (RPKM) method. Genomic annotation, distribution, and comparisons of COGC-seq/ChIP-seq peaks were analyzed with ChIPseeker (v1.18.0). Then, deepTools computeMatrix was used to analyze COGC-seq peaks intensity and Histone ChIP-seq signal distribution in reference-point mode. The specific command was as follows: ComputeMatrix reference-point --beforeRegionStartLength 2000 --afterRegionStartLength 2000 –skipZeros. The output of ComputeMatrix was further visualized as heatmaps using deeptools plotHeatmap or plotProfile. Differential binding regions were identified using MAnorm (v1.2.0) based on normalized read counts, with criteria set at |M value| ≥ 1 and p ≤ 0.05. Visualization of sequencing tracks was performed using the Integrative Genomics Viewer (IGV, v2.5.1), while heatmaps and signal density profiles were generated using R package ClusterGVis (v0.99.1) and deepTools. ChromHMM (v1.22) was used to identify and characterize chromatin states based on five epigenetic modifications, including O-GlcNAc. Deduplicated BAM files for histone and O-GlcNAc samples, along with their corresponding input controls, were processed using the BinarizeBam function in ChromHMM with default settings: LearnModel -paired -b 200 8 hg19. An 8-state chromatin model was generated, with each state annotated as promoter, enhancer, repressed, O-GlcNAc promoter, O-GlcNAc enhancer, O-GlcNAc repressed, or O-GlcNAc-unmarked. The model was manually curated to achieve a biologically relevant balance between unique and overlapping combinatorial features. Chromatin state transitions were visualized using the geom_alluvium function from the R package ggalluvial (v3.6.1). Motif enrichment and discovery were carried out with Homer (v4.11) (-size 200 -nomotif -float -mask) and the MEME suite (v5.1.1) applying sequences for analysis. GO enrichment analysis was conducted through Metascape (v3.5). All bubble plot were generated using the R package ggplot2 (v3.3.3). Additional bioinformatics analyses, including statistical testing, were performed using the phyper function in the R software package.
DNA methylation
Whole-genome bisulfite sequencing (WGBS) was performed to assess DNA methylation at single-base resolution. Genomic DNA was extracted from ~1 × 10⁶ cells and treated with bisulfite using the EZ DNA Methylation-Gold Kit (Zymo Research), ensuring the conversion of unmethylated cytosines to uracil, while preserving methylated cytosines. After bisulfite treatment, library preparation was completed according to standard protocols, followed by sequencing on an Illumina platform. The raw data for bisulfite sequencing are available in the GEO database under the accession number GSE295084. Fastp (v0.23.1) was used to trim reads with the parameters set as follows: --cut_right_window_size = 4 --cut_right_mean_quality = 15 --length_required = 36. Sequencing data were aligned to the reference genome (hg19) using Bismark (v0.22.3): -N 0 -L 20 --quiet --un --ambiguous. Duplicate reads were removed: deduplicate_bismark -p. DNA methylation levels were determined as the ratio of methylated reads to total reads at each CpG site: bismark_methylation_extractor --comprehensive --no_overlap -p --parallel 30 --split_by_chromosome --bedGraph --counts --CX_context --cytosine_report --report --buffer_size 90G. The genome-wide distribution of DNA methylation was analyzed using the scale-regions mode of DeepTools computeMatrix with the following parameters: ComputeMatrix scale-regions -S -R --beforeRegionStartLength 2000 --regionBodyLength 5000 --afterRegionStartLength 2000 --binSize 100. The output of ComputeMatrix was further visualized as heatmaps using deeptools plotProfile. R packages circlize (v0.4.15) and R were used for creating circular genome plots and karyotype-based visualizations, respectively.
RNA sequencing
Total RNA was extracted from 5 × 104 cells and subsequently fragmented before being reverse-transcribed to construct cDNA libraries. Sequencing of the libraries was performed using the Illumina NovaSeq 6000 platform (Illumina) at Novogene Technology (Beijing, China). Each RNA-seq experiment included two biological replicates. The raw sequencing data have been deposited in the GEO database under the accession number GSE295083. Fastp (v0.23.1) was used to trim reads with the parameters set as follows: -g -q 5 -u 50 -n 15 -l 150 --overlap_diff_limit 1 --overlap_diff_percent_limit 10. Clean reads were aligned to the human reference genome (hg19) using Hisat2 (v2.0.5): hisat2 --dta -x −1 −2 -S. After alignment, the results were sorted and converted to BAM files. Alignment quality was assessed using samtools flagstat, and gene expression counts were obtained using featureCounts (v1.5.0-p3). The specific commands were as follows: featureCounts -T 8 -p -B -C -a -o. DEGs were identified using R-R- with thresholds set at counts ≥10, fold change ≥2 and p ≤ 0.05. GO functional enrichment analysis was performed using Metascape (v3.5). GSEA (v4.3.2) was carried out with the R package clusterProfiler (v4.6.0) and the MsigDB (v7.0) GMTs gene set collection, with gene sets considered significantly enriched at FDR < 0.05 after 1000 permutations. Additional bioinformatics analyses were conducted with the phyper function in R.
Quantitative real-time PCR analysis (qPCR), ChIP-qPCR
Total RNA was extracted using the Trizol reagent (Invitrogen) following the manufacturer’s protocol. RNA (5 μg) was reverse transcribed and amplified using the One-Step SYBR PrimeScript PLUS RT-PCR Kit (Takara) on a Thermal Cycler Dice system (TaKaRa), according to the kit instructions. The relative enrichment of various subunits at each site was quantified using the 2-ΔΔCt method. For ChIP-qPCR, ChIP assays were conducted as described above. DNA bound by HA-JUN, Flag-GATAD2A or IgG control was de-crosslinked and purified for subsequent qPCR analysis. Enrichment was determined by qPCR using primers targeting the promoters of the indicated genes. IgG was used as a negative control. The sequences of all primers used in this study were designed to target specific genes using SnapGene software (v 5.1.7). All primers are listed in Supplementary Data 8. All experiments were conducted in four biological replicates.
Molecular simulation
For GATAD2A systems modeling, the MBD2-MTA1-HDAC1 complex system was obtained from the crystal structure (PDB ID: 7AO9) from the Protein Data Bank (PDB) and complemented gaps in the structure with MODELLER (v10.4). GATAD2ACR1 structure (residues 137–190) was built using MODELLER with crystal structure (PDB ID: 2L2L) as the template. The O-GlcNAc modified GADAT2ACR1 structure at S182 was built based on the GADAT2ACR1 model structure with the online platform GLYCAM. ZDOCK (v3.0.2) was utilized to predict the interactions between the non-glycosylated GATAD2ACR1 and O-GlcNAc modified GATAD2ACR1 structures with the MBD2-MTA1-HDAC1 complex, forming two distinct systems to investigate the impact of O-GlcNAc on the recruitment of MBD2-MTA1-HDAC1 and GATAD2ACR1. These two GATAD2ACR1-bound systems and MBD2-MTA1-HDAC1 system were subjected to run 500 ns molecular dynamics (MD) simulations using Amber23 software.
For JUN systems modeling, we took the DNA-interacting domain (residues 9-106) of SNF5 and a 14 base-pair DNA helical structure from one cryo-EM structure of SWI/SNF complex (PDB ID: 7Y8R), and the AlphaFold2 modeling structure of the JUN N-terminal domain (residues 69–91) of JUN (UniProt ID: P05412), which includes the glycosylation site S84JUN, to predict their interactions with HDOCKlite (v1.1). The O-GlcNAc site S84JUN was specified as an interface residue, and AI-based molecular surface interaction fingerprinting (MaSIF) algorithm was used to detect the potential interface and to analysis the docking results. Several potential interfaces for JUN N-terminal domain were identified, most are located at the opposite of the DNA binding pocket of SNF5 and are formed by Q59, L54, R53, and Q20.
Supplementary Table 1 presents the simulation setup for each system. All MD simulations were performed using Amber23 with the Amber14 force field. The structure was solvated in a cubic TIP3P water box with a 1 nm distance from the edge, and the system was neutralized by sodium ions. After four steps of energy minimization, the temperature of the system was gradually increased to 300 K over 100 ps to perform 5 ns NVT equilibration followed by 5 ns NPT equilibration. Finally, production MD simulations at 300 K and 1 atm were carried out with the LINCS algorithm to restrain the hydrogen positions at their equilibrium distances, allowing for an integration time step of 2 fs. Energies and coordinates were saved every 10 ps for postproduction analysis of the MD simulations. All MD simulations were performed on a high-performance computer cluster running the Linux operating system. Cpptraj and GetContacts packages were used to analysis the MD simulation results.
Xenograft mouse model
Ethical approval for all animal experiments was granted by the Ethics Committee of Dalian University of Technology (DUTSCEOLS240228-05). The maximal tumor size or burden permitted by the committee was 1.5 cm in diameter, and this limit was not exceeded in any of the experiments. GFP-stably expressing A549 lung cancer cells were established in our laboratory. To create tumor xenografts, 5-week-old nude mice with equal numbers of males and females (Changsheng Biotechnology Co., Ltd., Benxi, China) were used. All nude mice were housed in an SPF-level barrier system. Housing conditions: 12-h light/dark cycle (08:00–20:00), temperature 24 °C ± 1 °C, relative humidity 40–70%, with free access to water and food. For tumor formation, 2.5 × 106 A549-GFP cells were injected subcutaneously, alone or combined with 5 × 106 LU-RAS or its derivative cells, which were induced into senescence with doxycycline at D10. Cells were injected into the left armpit of anesthetized mice in a 100 μL suspension of RPMI 1640 medium, forming a subcutaneous tumor. Each group consisted of six mice (n = 6). To enable tracking of senescent LU-RAS cells, they were pre-stained with CellBriteTM NIR790 Cytoplasmic Membrane Dye (Biotium #30079; 786/820 nm) prior to injection. Tumor growth and fluorescence signals from A549-GFP and senescent LU-RAS cells were monitored twice weekly using the PerkinElmer IVIS Lumina imaging system. Fluorescence signals were standardized for each image and normalized using ImageJ macros for image processing. Tumor signal intensity was further analyzed semi-manually using ImageJ macros, with data expressed as fluorescence integrated density. Tumor size was monitored with caliper every 4 days until 28 days. After the experimental endpoint, mice were euthanized, and tumors were excised, fixed, and subjected to sectioning and staining procedures, including H&E staining, SA-β-Gal staining, and immunostaining with antibodies for IL-6 (Proteintech, #21865-1-AP, 1:100), IGFBP2 (Proteintech, #11065-3-AP, 1:100), CCND1 (Proteintech, # 26939-1-AP, 1:1000), and STAG2 (Proteintech, #19837-1-AP, 1:100). Tumor volume was calculated using the formula: Tumor volume (mm3) = width2 × length × 0.52.
Statistics analysis and reproducibility
Statistical analyses were conducted using a two-sided unpaired Student’s t test for single comparisons with GraphPad Prism 9.0 and Microsoft Excel 2019. A p < 0.05 was considered statistically significant. Data are presented as means ± SD. To ensure reproducibility, blots were repeated in three biological replicates, as detailed in the Methods and figure legends. RNA-seq, ChIP-seq, WGBS, and COGC-seq experiments were performed with two biological replicates, whereas micrographs and other biochemical experiments were conducted independently with four biological replicates per condition. All experimental results were successfully reproduced. Sample sizes were chosen based on previous experiments, and no statistical method was used to predetermine sample size for all experiments. For box plots, the centerline indicates the median, the box represents the 25th to 75th percentiles, and whiskers denote the minimum and maximum values. Correlations were assessed using Pearson’s correlation coefficient (r). The specific n values for each dataset are provided in the corresponding figure legends. Raw sequencing data have been deposited in the Gene Expression Omnibus database (accession number GSE295083, GSE295084, GSE295087), while raw mass spectrometry data are accessible via the iProX repository under the identifier PXD063140.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data of COGC-seq and ChIP-seq is available in the Gene Expression Omnibus database under the accession number GSE295087. The raw data of DNA methylation whole-genome bisulfite sequencing is available in the Gene Expression Omnibus database under the accession number GSE295084. The raw data of RNA-seq is available in the Gene Expression Omnibus database under the accession number GSE295083 [https://www.ncbi.xyz/geo/query/acc.cgi]. The raw mass spectral data in our study is available via iProX with identifier PXD063140. The raw data and files of MD simulation are available via Github [https://github.com/ATOMS-xjtlu/NuRD] and Zenodo [https://doi.org/10.5281/zenodo.17062795]. All additional data used to generate figures are provided in Supplementary Data 1–7. Source data are provided with this paper.
Code availability
The input code of MD simulation are available via Github [https://github.com/ATOMS-xjtlu/NuRD] and Zenodo [https://doi.org/10.5281/zenodo.17062795]. All additional code used to generate data are provided in methods.
References
Collado, M., Blasco, M. A. & Serrano, M. Cellular senescence in cancer and aging. Cell 130, 223–233 (2007).
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
Di Micco, R., Krizhanovsky, V., Baker, D. & d’Adda di Fagagna, F. Cellular senescence in ageing: from mechanisms to therapeutic opportunities. Nat. Rev. Mol. Cell Biol. 22, 75–95 (2021).
Hernandez-Segura, A., Nehme, J. & Demaria, M. Hallmarks of cellular senescence. Trends Cell Biol. 28, 436–453 (2018).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. Hallmarks of aging: an expanding universe. Cell 186, 243–278 (2023).
Dong, Z. et al. Cellular senescence and SASP in tumor progression and therapeutic opportunities. Mol. Cancer 23, 181 (2024).
Takasugi, M., Yoshida, Y., Hara, E. & Ohtani, N. The role of cellular senescence and SASP in tumour microenvironment. FEBS J. 290, 1348–1361 (2023).
Faget, D. V., Ren, Q. & Stewart, S. A. Unmasking senescence: context-dependent effects of SASP in cancer. Nat. Rev. Cancer 19, 439–453 (2019).
Birch, J. & Gil, J. Senescence and the SASP: many therapeutic avenues. Genes Dev. 34, 1565–1576 (2020).
Kim, Y. M. et al. Implications of time-series gene expression profiles of replicative senescence. Aging cell 12, 622–634 (2013).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Huang, W., Hickson, L. J., Eirin, A., Kirkland, J. L. & Lerman, L. O. Cellular senescence: the good, the bad and the unknown. Nat. Rev. Nephrol. 18, 611–627 (2022).
Ma, J., Hou, C. & Wu, C. Demystifying the O-GlcNAc code: a systems view. Chem. Rev. 122, 15822–15864 (2022).
Slawson, C. & Hart, G. W. O-GlcNAc signalling: implications for cancer cell biology. Nat. Rev. Cancer 11, 678–684 (2011).
Zhao, Y. et al. O-GlcNAc signaling: implications for stress-induced adaptive response pathway in the tumor microenvironment. Cancer Lett. 598, 217101 (2024).
Zachara, N. E. & Hart, G. W. O-GlcNAc a sensor of cellular state: the role of nucleocytoplasmic glycosylation in modulating cellular function in response to nutrition and stress. Biochim. Biophys Acta Gen. Subj. 1673, 13–28 (2004).
Chen, P.-H., Chi, J.-T. & Boyce, M. Functional crosstalk among oxidative stress and O-GlcNAc signaling pathways. Glycobiology 28, 556–564 (2018).
Liu, Y. et al. Proteomic profiling and genome-wide mapping of O-GlcNAc chromatin-associated proteins reveal an O-GlcNAc-regulated genotoxic stress response. Nat. Commun. 11, 5898 (2020).
Fan, J. et al. O-GlcNAc transferase in astrocytes modulates depression-related stress susceptibility through glutamatergic synaptic transmission. J. Clin. Investig. 133, e160016 (2023).
Zhang, X., Shu, X. E. & Qian, S.-B. O-GlcNAc modification of eIF4GI acts as a translational switch in heat shock response. Nat. Chem. Biol. 14, 909–916 (2018).
Comer, F. I. & Hart, G. W. O-GlcNAc and the control of gene expression. Biochim. Biophys. Acta Gen. Subj. 1473, 161–171 (1999).
Sakabe, K., Wang, Z. & Hart, G. W. -N-acetylglucosamine (O-GlcNAc) is part of the histone code. Proc. Natl. Acad. Sci. 107, 19915–19920 (2010).
Lewis, B. A. & Hanover, J. A. O-GlcNAc and the epigenetic regulation of gene expression. J. Biol. Chem. 289, 34440–34448 (2014).
Tang, X. et al. The PTM profiling of CTCF reveals the regulation of 3D chromatin structure by O-GlcNAcylation. Nat. Commun. 15, 2813 (2024).
Liu, Y. et al. FOXA1 O-GlcNAcylation–mediated transcriptional switch governs metastasis capacity in breast cancer. Sci. Adv. 9, eadg7112 (2023).
Wang, L. et al. Chromatin-associated OGT promotes the malignant progression of hepatocellular carcinoma by activating ZNF263. Oncogene 42, 2329–2346 (2023).
Ling-Hang, M., Wei, Z. & Jia-Jia, W. Enrichment approaches of O-GlcNAcylated proteins. Chin. J. Biochem. Mol. Biol. 40, 463–473 (2024).
Loison, I. et al. O-GlcNAcylation inhibition redirects the response of colon cancer cells to chemotherapy from senescence to apoptosis. Cell Death Dis. 15, 762 (2024).
Lee, J.-S. & Zhang, Z. O-linked N-acetylglucosamine transferase (OGT) interacts with the histone chaperone HIRA complex and regulates nucleosome assembly and cellular senescence. Proc. Natl. Acad. Sci. 113, E3213–E3220 (2016).
Efimova, E. V. et al. O-GlcNAcylation enhances double-strand break repair, promotes cancer cell proliferation, and prevents therapy-induced senescence in irradiated tumors. Mol. Cancer Res. 17, 1338–1350 (2019).
White III, C. W. et al. Age-related loss of neural stem cell O-GlcNAc promotes a glial fate switch through STAT3 activation. Proc. Natl. Acad. Sci. 117, 22214–22224 (2020).
Boulard, M., Rucli, S., Edwards, J. R. & Bestor, T. H. Methylation-directed glycosylation of chromatin factors represses retrotransposon promoters. Proc. Natl. Acad. Sci. 117, 14292–14298 (2020).
Kang, D. et al. Regulation of senescence-associated secretory phenotypes in osteoarthritis by cytosolic UDP-GlcNAc retention and O-GlcNAcylation. Nat. Commun. 16, 1094 (2025).
Palma, F. R. et al. Histone H3. 1 is a chromatin-embedded redox sensor triggered by tumor cells developing adaptive phenotypic plasticity and multidrug resistance. Cell Rep. 43, 113897 (2024).
Maya-Mendoza, A. et al. Immortalised breast epithelia survive prolonged DNA replication stress and return to cycle from a senescent-like state. Cell Death Dis. 5, e1351–e1351 (2014).
Zhang, Z. et al. O-GlcNAc homeostasis contributes to cell fate decisions during hematopoiesis. J. Biol. Chem. 294, 1363–1379 (2019).
Hao, Y. et al. Chemoproteomic and transcriptomic analysis reveals that O-GlcNAc regulates mouse embryonic stem cell fate through the pluripotency network. Angew. Chem. 135, e202300500 (2023).
Cheng, B. et al. Facile synthesis of clickable unnatural sugars in the unprotected and 1, 6-Di-O-Acylated forms for metabolic glycan labeling. Chem.–A Eur. J. 29, e202203054 (2023).
Martínez-Zamudio, R. I. et al. AP-1 imprints a reversible transcriptional programme of senescent cells. Nat. cell Biol. 22, 842–855 (2020).
Han, R. et al. Functional CRISPR screen identifies AP1-associated enhancer regulating FOXF1 to modulate oncogene-induced senescence. Genome Biol. 19, 1–13 (2018).
Ma, J. & Hart, G. W. O-GlcNAc profiling: from proteins to proteomes. Clin. Proteom. 11, 8 (2014).
Malard, F., Massman, L., Campagne, S. & Olivier-Van Stichelen, S. The O-GlcNAc database: introducing new features and tools developed from community feedback. Anal. Bioanal. Chem. 417, 879–884 (2025).
Brackertz, M., Gong, Z., Leers, J. & Renkawitz, R. p66α and p66β of the Mi-2/NuRD complex mediate MBD2 and histone interaction. Nucleic acids Res. 34, 397–406 (2006).
Gnanapragasam, M. N. et al. p66α–MBD2 coiled-coil interaction and recruitment of Mi-2 are critical for globin gene silencing by the MBD2–NuRD complex. Proc. Natl. Acad. Sci. 108, 7487–7492 (2011).
Low, J. K. et al. The nucleosome remodeling and deacetylase complex has an asymmetric, dynamic, and modular architecture. Cell Rep. 33, 108450 (2020).
Redmer, T. et al. JUN mediates the senescence associated secretory phenotype and immune cell recruitment to prevent prostate cancer progression. Mol. Cancer 23, 114 (2024).
Brackertz, M., Boeke, J., Zhang, R. & Renkawitz, R. Two highly related p66 proteins comprise a new family of potent transcriptional repressors interacting with MBD2 and MBD3. J. Biol. Chem. 277, 40958–40966 (2002).
Stirzaker, C. et al. Methyl-CpG-binding protein MBD2 plays a key role in maintenance and spread of DNA methylation at CpG islands and shores in cancer. Oncogene 36, 1328–1338 (2017).
Xia, L. et al. CHD4 has oncogenic functions in initiating and maintaining epigenetic suppression of multiple tumor suppressor genes. Cancer Cell 31, 653–668. e657 (2017).
Woo, C. M. et al. Mapping and quantification of over 2000 O-linked glycopeptides in activated human T cells with isotope-targeted glycoproteomics (Isotag). Mol. Cell. Proteom. 17, 764–775 (2018).
Xu, S., Zheng, J., Xiao, H. & Wu, R. Simultaneously identifying and distinguishing glycoproteins with O-GlcNAc and O-GalNAc (the Tn Antigen) in human cancer cells. Anal. Chem. 94, 3343–3351 (2022).
Schmitt, C. A., Wang, B. & Demaria, M. Senescence and cancer—role and therapeutic opportunities. Nat. Rev. Clin. Oncol. 19, 619–636 (2022).
Wang, L., Lankhorst, L. & Bernards, R. Exploiting senescence for the treatment of cancer. Nat. Rev. Cancer 22, 340–355 (2022).
Haugstetter, A. et al. Cellular senescence predicts treatment outcome in metastasised colorectal cancer. Br. J. Cancer 103, 505–509 (2010).
Thaçi, K. & Anthony, R. M. The importance of IgG N-glycosylation in Health, Disease, and Neonatal hemochromatosis. Glycosci. Ther. 1, 100002 (2025).
Liu, Y. et al. Proteomic profiling and genome-wide mapping of O-GlcNAc chromatin-associated proteins reveal an O-GlcNAc-regulated genotoxic stress response. Nat. Commun. 11, 1–17 (2020).
Itkonen, H. M. et al. High OGT activity is essential for MYC-driven proliferation of prostate cancer cells. Theranostics 9, 2183 (2019).
Guo, H. et al. O-linked N-acetylglucosamine (O-GlcNAc) expression levels epigenetically regulate colon cancer tumorigenesis by affecting the cancer stem cell compartment via modulating expression of transcriptional factor MYBL1. J. Biol. Chem. 292, 4123–4137 (2017).
Liu, Y. et al. O-GlcNAc elevation through activation of the hexosamine biosynthetic pathway enhances cancer cell chemoresistance. Cell Death Dis. 9, 1–12 (2018).
Fan, Q. et al. O-GlcNAc site-mapping of liver X receptor-α and O-GlcNAc transferase. Biochem. Biophys. Res. Commun. 499, 354–360 (2018).
Lubas, W. A. & Hanover, J. A. Functional expression of O-linked GlcNAc transferase: domain structure and substrate specificity. J. Biol. Chem. 275, 10983–10988 (2000).
Wulff-Fuentes, E. et al. The human O-GlcNAcome database and meta-analysis. Sci. Data 8, 25 (2021).
Nelson, Z. M., Leonard, G. D. & Fehl, C. Tools for investigating O-GlcNAc in signaling and other fundamental biological pathways. J. Biol. Chem. 300, 105615 (2024).
Hao, Y. et al. Next-generation unnatural monosaccharides reveal that ESRRB O-GlcNAcylation regulates pluripotency of mouse embryonic stem cells. Nat. Commun. 10, 4065 (2019).
Gao, J. et al. Proteomic analysis of the OGT interactome: novel links to epithelial–mesenchymal transition and metastasis of cervical cancer. Carcinogenesis 39, 1222–1234 (2018).
Liu, T.-W., Myschyshyn, M., Sinclair, D. A. & Vocadlo, D. J. A Chemical genetic method for monitoring genome-wide dynamics of O-GlcNAc turnover on chromatin-associated proteins. ACS Cent. Sci. 5, 663–670 (2019).
Yi-Wen, H., Cheng, Y. & Jun-Ping, Z. O-GlcNAcylation in regulating inflammatory signaling pathways. Chin. J. Biochem. Mol. Biol. 38, 990–998 (2022).
Fowler, C. B., Cunningham, R. E., O’Leary, T. J. & Mason, J. T. ‘Tissue surrogates’ as a model for archival formalin-fixed paraffin-embedded tissues. Lab. Investig. 87, 836–846 (2007).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Szklarczyk, D. et al. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
We thank Prof. Bo Cheng at Peking University for kindly providing 1,6-Pr2GalNAz. This study is supported by the National Natural Science Foundation of China (32171282, 32471331), Yubo Liu; Liaoning Province’s “Xingliao Talent Plan” Youth Top Talents (XLYC2203069), Yubo Liu; the Fundamental Research Funds for the Central Universities (DUT23YG114, DUT25YG244) and XJTLU Research Development Fund (RDF-23-01-100), S.W.
Author information
Authors and Affiliations
Contributions
Yubo Liu and J.Z. conceived and designed the study. N.Z., Yajie Liu, K.Y., L.H., and F.M. performed experiments, and R.Z. and X.Z. supervised experiments. N.Z. and R.Z. performed bioinformatics analyses, and Yubo Liu and W.W. supervised bioinformatics analyses. Qingbin Chen performed the preparation of all sequencing samples. N.Z. and Y.R. performed LC-MS/MS and processed the data. Qiushi Chen, X.L., and K.Z. analyzed the mass spectrometry data. N.Z. and Q.D. acquired images using confocal microscopy; J.W. and S.W. performed molecular dynamics simulation; N.Z. obtained the next-generation sequencing data. Yubo Liu, N.Z., H.H., and J.Z. analyzed data. N.Z. and Yubo Liu contributed to and supervised the image analysis and prepared figures. All authors contributed to the experiments and supported data analyses. Yubo Liu, N.Z., S.W., W.W., and R.Y. wrote and revised the manuscript. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Stéphan Hardivillé, Anne-Sophie Vercoutter-Edouart, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, N., Zhao, R., Zhong, X. et al. Time-resolved multiomics profiling reveals chromatin O-GlcNAc modification promotes senescence-associated transcriptional program. Nat Commun 17, 1406 (2026). https://doi.org/10.1038/s41467-025-68143-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68143-z