Abstract
Paclitaxel and nab-paclitaxel differ in therapeutic efficacy and modulation of the tumor immune microenvironment, yet the molecular basis remains poorly defined. Here, based on a meta-analysis, we first show that treatment with nab-paclitaxel results in a higher overall response rate and pathological complete response compared to paclitaxel in female patients with breast cancer. Notably, TREM2 expression in macrophages is elevated in primary tumors of paclitaxel- but not nab-paclitaxel-treated female patients. In metastatic breast cancer, TREM2+ macrophage infiltration is increased in primary tumors. In breast cancer models in female mice, paclitaxel, but not nab-paclitaxel, promotes lung metastasis by recruiting TREM2+ macrophages to primary tumors. Mechanistically, paclitaxel enhances the ATF3-FGF2 axis in breast cancer cells; secreted FGF2 activates the EGR1-TREM2-EMT cytokine axis in macrophages. Genetic ablation of Trem2 or pharmacologic targeting with antisense oligonucleotides suppress paclitaxel-induced breast cancer lung metastasis in vivo. Collectively, our findings demonstrate that paclitaxel, but not nab-paclitaxel, stimulates TREM2 expression and expands TREM2+ macrophages, suggesting that TREM2 targeting could enhance paclitaxel efficacy while limiting metastasis.
Similar content being viewed by others
Introduction
Breast cancer is the most frequently diagnosed malignancy and the leading cause of cancer-related mortality among women worldwide1,2. Taxanes, including paclitaxel (PTX) and nab-paclitaxel (Nab-PTX), remain first-line chemotherapeutic agents, with Nab-PTX comprising PTX bound to albumin nanoparticles3,4. By exploiting the endogenous albumin transport pathway, Nab-PTX enhances tumor-specific drug delivery, increasing intratumoral drug concentration and potentiating antitumor effects5. Nab-PTX demonstrates efficacy even in patients with disease progression following prior PTX therapy6,7. However, the comparative clinical benefits of Nab-PTX versus PTX remain controversial.
Our recent study has revealed that Nab-PTX remodels the myeloid compartment, expanding mast cells and pro-inflammatory macrophages, which synergize with immune checkpoint blockade (ICB) to enhance therapeutic efficacy. In contrast, PTX promotes an immunosuppressive environment by increasing populations of exhausted CD8⁺ T cells, regulatory T cells (Tregs), and suppressive macrophages, which may underlie its limited clinical effectiveness8. In addition, murine studies report that PTX can promote breast cancer lung metastasis through extracellular vesicles9 or by enhancing host angiogenesis10,11,12,13,14. However, the precise mechanisms by which PTX actively reshapes the tumor immune microenvironment toward a suppressive state remain unclear.
Triggering receptor expressed on myeloid cells-2 (TREM2) is a single-pass transmembrane immune receptor expressed on myeloid cells and implicated in multiple cancers. Its role appears context-dependent, exhibiting both tumor-promoting and tumor-suppressing functions15,16,17,18. Notably, TREM2+ macrophages are enriched at invasive margins of breast tumors in patients and murine models19, suggesting a potential contribution to breast cancer lung metastasis. TREM2 depletion or blockade has been reported to enhance checkpoint immunotherapy efficacy by potentiating T cell responses15. Previously, we demonstrated that PTX impairs the expansion of CXCL13-expressing T cells, thereby limiting checkpoint immunotherapy outcomes in patients with breast cancer8,20. We therefore hypothesized that TREM2 may underlie the poor therapeutic efficacy associated with PTX treatment, and sought to determine whether PTX modulates TREM2+ macrophages and whether Nab-PTX operates via a distinct mechanism.
Here, we show that PTX, but not Nab-PTX, induces the expansion of TREM2+ macrophages, promoting lung metastasis in murine breast cancer models. Meta-analysis confirms that PTX exhibits inferior antitumor efficacy compared with Nab-PTX in patients with breast cancer. Mechanistically, PTX activates the ATF3-FGF2 axis in tumor cells, which triggers an EGR1–TREM2–EMT cytokine axis in macrophages. Targeting TREM2 attenuates PTX-driven metastasis and enhances therapeutic responses. These findings reveal a specific pro-metastatic effect of PTX and suggest that combining PTX with TREM2-targeted therapy could improve clinical outcomes.
Results
TREM2+ macrophages infiltrate breast cancer tissues of patients treated with PTX but not Nab-PTX
Accumulating clinical evidence highlights distinct efficacy profiles between PTX and Nab-PTX6,7,8,20. To quantitatively compare their antitumor efficacy, we systematically searched PubMed, Embase, and the Cochrane Library, identifying 2,833 candidate studies. The literature search and selection process are detailed in Supplementary Fig. 1. After rigorous abstract screening and full-text review to remove duplicates and ineligible studies (Supplementary Fig. 2), 17 studies comprising 6,486 patients were included (Supplementary Tables 1 and 2). A comprehensive overview of the study characteristics, including the year of the latest published results, is presented in Supplementary Table 3. These consisted of eight randomized controlled trials (RCTs), four retrospective studies, one single-arm clinical trial, and four cohort studies21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43.
Among 4,171 patients from 11 studies with available objective response rate (ORR) data, Nab-PTX treatment yielded a significantly higher ORR than PTX (OR 1.42, 95% CI 1.01–2.00; I2 = 70%) (Fig. 1a). Sensitivity analysis confirmed robustness of the results (Supplementary Fig. 3). Subgroup analyses by line of therapy, disease stage, and study design (Supplementary Figs. 4–6) indicated that heterogeneity in ORR primarily arose from studies including later-line treatments and advanced disease. Nab-PTX demonstrated a pronounced efficacy advantage in later-line patients. Progression-free survival (PFS) analysis in advanced-disease patients showed no overall difference (HR 0.91; 95% CI 0.78–1.07), but subgroup analysis revealed significantly prolonged PFS with Nab-PTX in later-line therapy (HR 0.77; 95% CI 0.65–0.91), consistent with ORR findings (Supplementary Figs. 7–9).
a Pooled efficacy comparisons of ORR in patients who received Nab-PTX versus PTX in breast cancer therapy. The blue boxes signify the effect size for each individual study, with their size reflective of the weight assigned to each study in the analysis. The whiskers extending from each blue box delineate the 95% confidence interval (CI) for the effect size of the respective study. The odds ratio (OR) was calculated using a random-effects model. Statistical significance was assessed by a two-sided test, with P < 0.05 considered statistically significant. b Pooled efficacy comparisons of the pCR in patients who received Nab-PTX versus PTX in breast cancer therapy. The blue boxes signify the effect size for each individual study, with their size reflective of the weight assigned to each study in the analysis. The whiskers extending from each blue box delineate the 95% confidence interval (CI) for the effect size of the respective study. The odds ratio (OR) was calculated using a random-effects model. Statistical significance was assessed by two-sided test, with P < 0.05 considered statistically significant. c Schematic of experimental design for patient samples. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pd UMAP plot and bar graph showing identified cell clusters of infiltrated macrophages of breast cancer patient tissues with indicated treatment and their proportion in the indicated groups. e Box plots showing the TREM2 expression levels in macrophage subsets across the indicated experimental groups. Two-sided Wilcoxon test. In the box plots, the center line corresponds to the median, box corresponds to the interquartile range (IQR), and whiskers 1.5 × IQR. f Violin-box plots showing the TREM2 expression levels in identified cell clusters of infiltrated macrophages using the scRNA-seq data. g Representative immunofluorescence staining of tissues from patients who received PTX treatments or not. h Representative immunofluorescence staining of tissues from patients who received Nab-PTX treatments or not. ns, not significant. Source data are provided as a Source Data file.
Seven studies (3829 patients) provided pathological complete response (pCR) data using the ypT0/is ypN0 definition44,45. Patients receiving Nab-PTX-containing neoadjuvant therapy exhibited a 9.1% higher pCR rate than those receiving PTX (OR 1.69, 95% CI 1.38–2.06; I2 = 28%) (Fig. 1b). Sensitivity analysis revealed that the findings were unaffected after each study was sequentially excluded (Supplementary Fig. 10). Subgroup analysis indicated that heterogeneity primarily arose from study design differences (Supplementary Fig. 11). Collectively, these data demonstrate that PTX exhibits inferior antitumor efficacy relative to Nab-PTX in patients with breast cancer.
Our previous scRNA-seq study indicated that PTX treatment increases infiltration of immunosuppressive TREM2+ macrophages in primary breast tumors20. To investigate differential effects of PTX and Nab-PTX on TREM2+ macrophages, we analyzed our previously published scRNA-seq dataset8, which included 7 PTX-treated and 12 Nab-PTX-treated patients (Fig. 1c). Tumors from PTX-treated patients showed a significant increase in TREM2+ macrophages compared with Nab-PTX-treated tumors (Fig. 1d, e and Supplementary Fig. 12a). Cluster-specific analysis revealed elevated TREM2 expression in CCL3L1, FOLR2, FTH1, IL1B, ISG15, MT1X, NUPR1, and RNASE1 macrophages in PTX-treated tumors (Fig. 1f), suggesting that selective expansion of these cells contributes to the reduced therapeutic efficacy of PTX relative to Nab-PTX.
To validate these findings in patient tissues, we performed immunofluorescence co-staining against the macrophage marker CD68 and TREM2 on primary tumor samples from 20 patients (5 per treatment group). Consistent with the sequencing data, PTX-treated tumors exhibited a significantly higher density of TREM2+ macrophages than Nab-PTX-treated tumors (Fig. 1g, h; Supplementary Figs. 12b, c and Supplementary Table 5). High TREM2 expression correlated with reduced overall survival in patients receiving chemotherapy or endocrine therapy (Supplementary Fig. 12d). In addition, another cohort of immunofluorescence co-staining of tumors from 12 PTX-treated and 16 untreated patients further confirmed PTX-induced TREM2 upregulation (Supplementary Figs. 12e, f and Supplementary Table 6). scRNA-seq of orthotopic tumors in mice recapitulated these observations, showing increased TREM2+ macrophages following PTX treatment (Supplementary Figs. 13a–i). Collectively, these data suggest that PTX-induced expansion of TREM2+ macrophages may underlie its limited antitumor efficacy.
PTX-induced TREM2+ macrophages associate with lung metastasis
Metastasis is a leading cause of treatment failure and mortality in breast cancer46,47. TREM2+ macrophages have been reported at invasive margins of lung metastases19. To investigate the association between TREM2+ macrophage infiltration and lung metastasis in patients, we performed immunofluorescence co-staining of primary tumors from patients with (n = 5) or without (n = 5) lung metastases. The results confirmed significantly elevated TREM2+ macrophage infiltration in metastatic cases (Fig. 2a, b, Supplementary Fig. 14a and Supplementary Table 7). In addition, immunofluorescence co-staining in a larger cohort (n = 35 metastatic, n = 105 non-metastatic) corroborated these results (Supplementary Figs. 14b, c and Supplementary Table 8). Multivariate regression, stratified by tumor subtype, showed that metastasis was significantly associated with higher TREM2+ macrophages in Luminal B, HER2+ and triple-negative breast cancer (TNBC) (Supplementary Table 9). Tissue microarray analysis further confirmed elevated TREM2 levels in macrophages from metastatic patients (Supplementary Figs. 14d, e). Additional analyses controlling for lymph node positivity demonstrated that this factor independently predicted increased TREM2+ macrophages in Luminal A, HER2+ and TNBC tumors (Supplementary Tables 10, 11). Collectively, these data establish a robust clinical link between elevated TREM2+ macrophages and breast cancer metastasis.
a Representative immunofluorescence staining of tissues from patients with and without lung metastases. b Quantification of TREM2-expressing macrophages in tissues from patients with and without lung metastases (n = 5 patients per group). c Schematic of the experimental design in 4T1-Luc breast-tumor-bearing mice treated with DMSO or PTX. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pd Representative in vivo bioluminescence images of mice that received the indicated treatments (n = 5 mice per group). e Representative bright-field images (left) and hematoxylin and eosin (H&E) staining (right) of lungs from mice injected orthotopically with 4T1-Luc cells and treated as described in (c) (n = 5 mice per group). f Quantification (right) of the total numbers of lung-surface metastases (n = 6 mice per group). g Representative flow cytometry analysis showing a histogram (left) and quantification (right) of the proportion of TREM2-expressing macrophages in tumors (n = 3 mice per group). h Representative immunofluorescence staining of tumors in mice that received the indicated treatments. i Schematic of Nab-PTX treatment in Py8119 breast tumor-bearing mice. Mice were intraperitoneally injected with DMSO, PTX, PBS, or Nab-PTX at the indicated times. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pj Photographs (left) of tumors in mice that received the indicated treatments on day 14. Tumor weights (right) of mice that received the indicated treatments (n = 6 mice per group). k Tumor volumes (right) of mice that received the indicated treatments (n = 6 mice per group). l Representative bright-field images (left) of lungs from mice that received the indicated treatments. Quantification of the total numbers of lung-surface metastases is shown (right) (n = 6 mice per group). m Representative H&E-stained images of lungs from mice that received the indicated treatments. n, o Representative flow cytometric analysis of pseudo-color plots (n) and quantification (o) showing the proportion of TREM2-expressing macrophages in tumors (n = 4 mice per group). Data are shown as means ± S.D. and were analyzed by two-sided unpaired Student’s t test (b, f and g), two-sided one-way ANOVA followed by Tukey’s test (j, l and o) and two-sided two-way ANOVA followed by Tukey’s test (k). Source data are provided as a Source Data file.
To test whether TREM2+ macrophages contribute to reduced efficacy observed in PTX treatment, we treated orthotopic 4T1 tumor-bearing mice with PTX (9 mg/kg) or vehicle every two days for five doses (Fig. 2c). PTX suppressed primary tumor growth (Fig. 2d and Supplementary Figs. 15a, b) but paradoxically increased lung metastases (Fig. 2e, f). Flow cytometry and immunofluorescence confirmed increased TREM2+ macrophages in PTX-treated tumors (Fig. 2g, h; Supplementary Fig. 15c, d). MMTV-PyMT mice treated similarly exhibited suppressed tumor growth, enhanced lung metastasis, and increased tumor-associated TREM2+ macrophages (Supplementary Figs. 15e–k, 32a), along with elevated TREM2+ macrophages in peripheral blood, suggesting bone marrow recruitment (Supplementary Figs. 15l, 32b). These findings demonstrate that PTX promotes metastasis in concert with TREM2+ macrophage expansion.
We next compared PTX and Nab-PTX (both 9 mg/kg) in mouse models (Fig. 2i). Both treatments suppressed primary tumor growth similarly (Fig. 2j, k), but Nab-PTX did not promote lung metastasis (Fig. 2l, m) or induce TREM2+ macrophages (Fig. 2n and o and Supplementary Fig. 15m). Nab-PTX efficacy was comparable in Trem2+/+ and Trem2−/− mice without pro-metastatic effects (Supplementary Figs. 16a–c). These results indicate that the superior therapeutic profile of Nab-PTX over PTX stems not from increased tumoricidal potency, but from its inability to induce TREM2+ macrophage-dependent lung metastasis.
TREM2+ macrophages are required for PTX-promoted lung metastasis in breast cancer
To determine whether TREM2 is essential for PTX-promoted lung metastasis, Py8119 tumor-bearing Trem2+/+ and Trem2−/− mice were treated with PTX or vehicle (DMSO) intraperitoneally every two days for five doses (Fig. 3a). In Trem2+/+ mice, PTX significantly suppressed primary tumor growth yet paradoxically increased lung metastases relative to DMSO controls. In contrast, Trem2−/− mice exhibited similar tumor suppression by PTX but no increase in lung metastasis, with metastatic burden comparable between PTX and DMSO groups (Fig. 3b–e). Flow cytometry confirmed a higher proportion of TREM2⁺ macrophages in primary tumors and peripheral blood of PTX-treated Trem2+/+ mice compared to controls (Fig. 3f–h; Supplementary Figs. 32a, b). To investigate the relevance of TREM2 in inducing metastasis, we utilized a B16 melanoma model. TREM2 promoted metastasis in a TREM2-dependent manner (Supplementary Figs. 17a–c), suggesting a pro-metastatic function of TREM2 extending beyond breast cancer. Collectively, these results indicate that TREM2 is required for PTX-induced lung metastasis.
a Schematic of the experimental design. Trem2+/+ and Trem2−/− mice were orthotopically injected with Py8119 cells and treated with DMSO or PTX, respectively. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pb Photographs of representative tumors from mice that received the indicated treatments on day 14. c Tumor growth curves (left) and tumor weights (right) of Py8119 tumors in mice that received the indicated treatments (n = 6 mice per group). d Representative bright-field images (left) of lungs from Py8119 cell-bearing mice that received the indicated treatments. Quantification of the total number of lung-surface metastases is shown (right) (n = 6 mice per group). e Representative H&E-stained images of lungs from mice. f Representative flow cytometric analysis of histograms showing the proportion of TREM2-expressing macrophages in tumors. g Quantification of flow cytometric analysis showing the proportion of TREM2-expressing macrophages in tumors (n = 3 mice per group). h Representative flow cytometric analysis of pseudo-color plots showing the proportion of TREM2-expressing macrophages in the PB of mice (n = 3 mice per group). i Schematic of the experimental design. Trem2-/- mice were co-injected orthotopically with Py8119 cells and BMDMs. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pj Photographs of representative tumors from Trem2-/- mice that received the indicated treatments on day 17. k Tumor growth curves (left) and tumor weights (right) of Py8119 tumors in Trem2-/- mice (n = 6 mice per group). l Representative bright-field images (left) of lungs from Trem2-/- mice that received the indicated inoculations. Quantification of the total number of lung-surface metastases is shown (right) (n = 6 mice per group). Data are shown as means ± S.D. and were analyzed by two-sided unpaired Student’s t test (g), two-sided one-way ANOVA followed by Tukey’s test (c, d, k and l) and two-sided two-way ANOVA followed by Tukey’s test (c and k). Source data are provided as a Source Data file.
Given TREM2 expression on diverse myeloid populations, we next assessed whether TREM2⁺ macrophages specifically drive metastasis. A co-implantation model was established by injecting Py8119 cells with bone-marrow-derived macrophages (BMDMs) into mice, generating tumors enriched in TREM2⁺ macrophages (Fig. 3i). In Trem2−/− host mice, co-implantation with Trem2−/− BMDMs resulted in significantly reduced tumor growth compared to Trem2+/+ BMDMs (Fig. 3j, k). Consistently, lung metastasis was significantly attenuated in mice receiving Trem2−/− BMDMs (Fig. 3l and Supplementary Fig. 17d). This requirement for TREM2 in BMDMs to drive metastasis was further confirmed in Trem2+/+ host mice (Supplementary Figs. 17e–i). In vitro, conditioned medium (CM) from Trem2+/+ BMDMs significantly enhanced Py8119 cell migration and invasion compared to Trem2−/− BMDM medium (Supplementary Figs. 17j, k). These data collectively demonstrate that TREM2⁺ macrophages are essential for PTX-induced lung metastasis.
Targeted ASO-mediated Trem2 knockdown suppresses PTX-promoted lung metastasis
Based on the established role of TREM2 in PTX-induced metastasis, we tested whether targeted knockdown could mitigate this effect. Trem2-targeting antisense oligonucleotides (ASOs) or inactive ASOs were administered to Py8119 tumor-bearing mice in combination with PTX (Fig. 4a and Supplementary Figs. 18a–f and 31). Co-treatment with PTX and Trem2 ASOs significantly enhanced primary tumor suppression compared to PTX alone, ASOs alone, or PTX with inactive ASOs (Fig. 4b, c). Critically, Trem2 ASO co-treatment abrogated PTX-induced lung metastasis, markedly reducing metastatic burden relative to PTX plus inactive ASOs (Fig. 4d, e). This combination also extended survival without significant weight loss or overt organ toxicity (Fig. 4f and Supplementary Figs. 18g, h). These findings demonstrate that ASO-mediated TREM2 inhibition uncouples the antitumor from the pro-metastatic effects of PTX, thereby enhancing its therapeutic efficacy.
a Schematic of ASO treatment in Py8119 breast-tumor-bearing mice. Mice were intraperitoneally injected with DMSO or PTX and intratumorally injected with inactive ASOs or Trem2 ASOs at the indicated times. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pb Photographs (left) and tumor weights (right) of representative tumors in mice that received the indicated treatments on day 14. c Tumor growth curves of mice that received the indicated treatments (n = 6 mice per group). d Representative bright-field images of lungs from mice that received the indicated treatments. e Representative H&E-stained images (left) of lungs from mice that received the indicated treatments. Quantification of the total number of lung-surface metastases is shown (right) (n = 6 mice per group). f Survival curves of Py8119 breast tumor-bearing mice that received the indicated treatments (n = 6 mice per group). Data are shown as means ± S.D. and were analyzed by two-sided one-way ANOVA followed by Tukey’s test (b and e), two-way ANOVA followed by Tukey’s test (c) and log-rank test (f). Source data are provided as a Source Data file.
PTX elevates TREM2 expression in macrophages via cancer cell-derived FGF2
To determine how PTX upregulates TREM2 in macrophages, we first assessed whether direct PTX exposure alters macrophage TREM2 levels. Neither flow cytometry nor western blot detected significant changes following direct PTX treatment compared to vehicle controls (Fig. 5a–c and Supplementary Figs. 19a and 32c), indicating that PTX does not act directly on macrophages. We therefore hypothesized that paracrine factors from the tumor microenvironment (TME) mediate the observed increase in TREM2, consistent with reports that tumor-resident macrophages are responsive to cancer cell-derived factors48,49,50.
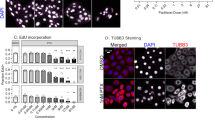
a Schematic of the experimental setup with THP1 or Raw264.7 cells with indicated treatment. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pb Quantification of proportion of TREM2 in THP1 and Raw264.7 cells with PTX. n = 3 biological independent samples. c Western blot of THP1 and Raw264.7 cells with indicated treatments. The experiment was independently repeated three times with similar results. d Schematic of the experimental strategy. CM was collected from tumor cells with the indicated treatment. TREM2 expression in macrophages incubated with the CM was assessed. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pe Quantification of proportion of TREM2 in THP1 incubated with CM of BT549 cells, SUM159 cells and MDA-MB-231 cells treated with PTX, in BMDM incubated with CM of Py8119 treated with PTX, in Raw264.7 incubated with CM from 4T1 and Py8119 cells treated with PTX, respectively. n = 3 biological independent samples. f Western blot analysis of TREM2 and related proteins in THP1, BMDMs, and Raw264.7 cells incubated with CM from BT549, SUM159, Py8119, and 4T1 cells, respectively. The experiment was independently repeated three times with similar results. g Representative immunofluorescence staining of THP1 cells incubated with BT549 CM. n = 3 biological independent samples. h Quantification of proportion of TREM2 in THP1 incubated with CM from BT549, SUM159, and MDA-MB-231 cells treated with Nab-PTX, respectively. n = 3 biological independent samples. i Quantification of the proportion of TREM2 in BMDM incubated with CM of Py8119 treated with Nab-PTX. n = 3 biological independent samples. j Cytokine array analysis of CM from BT549 cells treated indicated treatment. k Western blot analysis of the indicated proteins in Raw264.7 cells and BMDMs treated with recombinant FGF2. The experiment was independently repeated three times with similar results. l Quantification of the proportion of TREM2 in Raw264.7 cells and BMDMs treated with recombinant FGF2. n = 3 biological independent samples. m Schematic of the experimental strategy. CM was collected from tumor cells with the indicated treatment, and then pretreatment using an FGF2 neutralizing antibody. TREM2 expression in macrophages incubated with the CM was assessed. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pn Western blot analysis of the indicated proteins in Raw264.7 and BMDMs incubated with the indicated CM. The experiment was independently repeated three times with similar results. o Quantification of the proportion of TREM2 in Raw264.7 and BMDMs incubated with the indicated CM. n = 3 biological independent samples. p Representative multiplex immunofluorescence staining of CD68, TREM2 and FGF2 in tumors treated with PTX or Nab-PTX (n = 3 mice per group). The experiment was independently repeated three times with similar results. Data are shown as means ± S.D. and were analyzed by two-sided unpaired Student’s t test (b, e, h and i) and two-sided one-way ANOVA followed by Tukey’s test (l and o). Source data are provided as a Source Data file.
To test this, we treated macrophages with CM from breast cancer cells pre-exposed to PTX or DMSO (Fig. 5d). CM from PTX-treated cancer cells significantly increased TREM2 expression in macrophages, including BMDMs, confirming this effect in primary cells (Fig. 5e, f; Supplementary Figs. 19b–f and 32c). Membrane TREM2 expression was also elevated upon exposure to CM from PTX-treated cells (Fig. 5g and Supplementary Fig. 19g). Notably, Nab-PTX CM did not induce TREM2 upregulation in macrophages (Fig. 5h, i and Supplementary Figs. 19h, i, and 32c).
To identify the paracrine factor driving TREM2 induction, we performed a proteome cytokine array on CM from PTX- versus Nab-PTX-treated cancer cells. Cytokine arrays identify secreted chemokines and growth factors that modulate neighboring cells within tumors49,51,52. Among the candidates, fibroblast growth factor 2 (FGF2) was selectively and markedly upregulated in CM from PTX-treated cancer cells relative to DMSO controls. In contrast, Nab-PTX did not enhance FGF2 secretion (Fig. 5j), implicating PTX-specific FGF2 induction in macrophage TREM2 upregulation. Recombinant FGF2 alone was sufficient to robustly elevate TREM2 in macrophages (Fig. 5k, l and Supplementary Figs. 19j and 32c). Conversely, pre-incubation of PTX CM with a neutralizing FGF2 antibody effectively blocked TREM2 induction in both Raw264.7 cells and BMDMs (Fig. 5m–o and Supplementary Figs. 19k and 32c). To confirm these findings in vivo, multiplex immunofluorescence of primary tumors from PTX-treated mice demonstrated pronounced co-upregulation of TREM2 and FGF2, whereas Nab-PTX-treated tumors exhibited no appreciable changes (Fig. 5p and Supplementary Fig. 20).
To establish the functional importance of the FGF2/FGFR axis in PTX-driven metastasis, tumor-bearing mice were co-treated with an FGFR inhibitor (IGR) and PTX (Supplementary Fig. 21a). Co-administration of IGR significantly reduced both primary tumor growth and lung metastases compared with PTX alone (Supplementary Figs. 21b, c), accompanied by a marked decrease in tumor-infiltrating TREM2⁺ macrophages (Supplementary Figs. 21d and 32c). In vitro, FGFR inhibition abrogated TREM2 upregulation in Raw264.7 cells and BMDMs exposed to PTX CM (Supplementary Figs. 21e, f, and 32c). Together, these results demonstrate that PTX but not Nab-PTX elicits FGF2 secretion from cancer cells, which acts through FGFR signaling to elevate TREM2 expression on macrophages and drive metastasis.
ATF3 activates FGF2 transcription in PTX-treated cancer cells
To elucidate why FGF2 is upregulated in PTX- but not Nab-PTX-treated cancer cells, we sought transcription factors that could drive FGF2 expression. Using RNA-seq data combined with the GeneCard database, we identified two candidate regulators: ATF3 and SCRT1 (Fig. 6a). Correlation analysis revealed that ATF3 expression was more positively associated with FGF2 than SCRT1 in breast cancer (Fig. 6b and Supplementary Fig. 22a), and elevated ATF3 levels correlated with poor patient prognosis (Supplementary Fig. 22b). Consistently, PTX but not Nab-PTX significantly increased Atf3 expression in cancer cells (Fig. 6c). Overexpression of Atf3 in Py8119 cells was sufficient to markedly elevate Fgf2 levels compared to controls (Fig. 6d, e).
a Overlap of RNA-seq (n = 3) and PROMO public database analyses to predict transcription factors regulating TREM2 expression. b Correlations between FGF2 and ATF3 expression in breast cancer using TCGA databases. c qPCR analysis of Atf3 expression in the indicated cells. n = 4 (DMSO and PTX) or 3 (PBS and Nab-PTX) biological independent samples. d Western blot analysis of the indicated proteins in Py8119 cells transfected with ATF3-expressing or control vectors. The experiment was independently repeated three times with similar results. e qPCR analysis of Fgf2 expression in Py8119 cells transfected with Atf3-expressing or control vectors. n = 3 biological independent samples. f Luciferase activity in HEK293T cells transfected with the indicated reporters and Atf3-expressing or control vectors. n = 3 biological independent samples. g Abundance of Atf3 bound to the Trem2 promoter in Py8119 cells, as assessed by ChIP-qPCR. n = 3 biological independent samples. h ATAC-seq tracks showing the chromatin accessibility in the ATF3 loci for BT549 cells treated by PTX or Nab-PTX. n = 2 samples per group. Data are shown as means ± S.D. and were analyzed by two-sided unpaired Student’s t test (c, e and f) and two-sided two-way ANOVA followed by Šídák’s test (g). Source data are provided as a Source Data file.
To determine whether ATF3 directly regulates FGF2 transcription, we performed luciferase reporter assays using a vector containing the FGF2 promoter, which confirmed that ATF3 activates transcription at the FGF2 locus (Fig. 6f). Chromatin immunoprecipitation followed by quantitative PCR (ChIP-qPCR) further demonstrated ATF3 occupancy at the Fgf2 promoter in Py8119 cells (Fig. 6g). These results indicate that PTX upregulates FGF2 by inducing its transcriptional factor ATF3.
Given that PTX but not Nab-PTX increases ATF3 mRNA, we hypothesized that differential chromatin accessibility might underlie this effect. ATAC-seq analysis revealed that PTX broadly enhanced chromatin accessibility across multiple genes, including NFKB1, MAPK, ATF3, and IGF1R, compared to Nab-PTX (Supplementary Fig. 22c). Specifically, accessibility at the ATF3 locus was increased in PTX-treated Py8119 cells, whereas Nab-PTX-treated cells showed no change (Fig. 6h and Supplementary Fig. 22d). Together, these data suggest that PTX, but not Nab-PTX, promotes ATF3 expression by enhancing chromatin accessibility, which in turn drives FGF2 transcription and secretion.
FGF2 promotes TREM2 expression by upregulating EGR1 in macrophages
To elucidate how FGF2 regulates TREM2 expression in macrophages, we first investigated candidate transcription factors using the GeneCards and EDCODE databases, identifying EGR1 as a prime candidate (Fig. 7a). Previous studies indicate that FGF2 induces EGR1 via ERK1/2 phosphorylation53. Consistently, western blot and qPCR analyses demonstrated that FGF2 stimulated ERK1/2 phosphorylation, upregulated downstream EGR1, and consequently increased TREM2 expression in macrophages (Fig. 7b). Nab-PTX CM, in contrast, did not activate ERK1/2 signaling (Fig. 7c). ATAC-seq of BMDMs exposed to CM from PTX- or Nab-PTX-treated cancer cells revealed no changes in chromatin accessibility at the Egr1 locus (Supplementary Fig. 23a and b), indicating that EGR1 induction occurs primarily through post-translational ERK activation rather than chromatin remodeling. Furthermore, treatment with the FGFR inhibitor IGR significantly reduced EGR1 levels in both Raw264.7 cells and BMDMs (Supplementary Fig. 23c), supporting the involvement of FGF2 signaling.
a Overlap of GeneCards and EDCODE public databases analyses to predict transcription factors that regulate TREM2. b Western blot analysis of the indicated proteins in BMDMs and Raw264.7 cells incubated with FGF2 (left) and with CM from Py8119 cells treated with PTX (right). The experiment was independently repeated three times with similar results. c Western blot analysis of the indicated proteins in Raw264.7 cells incubated with the indicated treatment. The experiment was independently repeated three times with similar results. d qPCR analysis of Trem2 expression in BMDMs transfected with Egr1-expressing vectors. n = 3 biological independent samples. e Western blot analysis of TREM2 expression in BMDMs transfected with Egr1-expressing vectors. The experiment was independently repeated three times with similar results. f Luciferase activity of HEK293T cells transfected with the indicated reporters and EGR1-expressing or control vectors. n = 3 biological independent samples. g Abundance of EGR1 bound to the TREM2 promoter in BMDMs assessed by ChIP-qPCR. n = 3 biological independent samples. h Schematic of the Transwell assay. The first CM was collected from tumor cells with indicated treatment, and the second CM was collected from macrophages incubated with the first CM. The migration and invasion capabilities of macrophages incubated with the first CM were assessed. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747pi Quantification of migration and invasion of Py8119 cells induced by BMDM CM incubated with CM from Py8119 cells treated with PTX, and BT549, SUM159, and MDA-MB-231 cells induced by THP1 CM incubated with CM from BT549, SUM159, and MDA-MB-231 cells treated with PTX (left) or Nab-PTX (right), respectively. n = 5 biological independent samples. j Cytokine array analysis of CM from BMDMs with or without TREM2. k Quantification of migration and invasion of Py8119 cells incubated with indicated proteins. n = 3 biological independent samples. l Quantification of migration and invasion of Py8119 cells induced by CM from BMDMs (Trem2+/+) with CDE-096. n = 3 biological independent samples. m Western blot analysis of EMT- stimulating proteins in BMDMs incubated with indicated proteins. The experiment was independently repeated three times with similar results. n Schematic illustration showing that FGF2 promotes ERK1/2 phosphorylation to upregulate EGR1, which increases TREM2 expression in macrophages. Upregulated TREM2 enhances the secretion of Serpin E1, HGF, CCL3, and CXCL2 from macrophages to tumor cells, facilitating tumor metastasis via EMT. Created in BioRender. Xing, Y. (2026) https://BioRender.com/nsp747p. Data are shown as means ± S.D. and were analyzed by two-sided unpaired Student’s t test (d, f, i and l), two-sided one-way ANOVA followed by Tukey’s test (k) and two-sided two-way ANOVA followed by Šídák’s test (g). Source data are provided as a Source Data file.
We next confirmed that EGR1 directly regulates TREM2 transcription. Overexpression of Egr1 in Raw264.7 cells robustly induced Trem2 expression (Fig. 7d, e), while luciferase reporter assays demonstrated that EGR1 activates transcription from the TREM2 promoter (Fig. 7f). ChIP-qPCR further confirmed EGR1 occupancy at the Trem2 promoter in Raw264.7 cells (Fig. 7g). Notably, high EGR1 expression correlates with poor prognosis in patients with breast cancer (Supplementary Fig. 23d). Together, these results indicate that FGF2 secreted by PTX-treated cancer cells activates the ERK pathway, elevates EGR1, and drives TREM2 transcription in macrophages.
TREM2-driven secretion of EMT-related cytokines promotes breast cancer metastasis
To investigate whether TREM2+ macrophages induced by PTX-treated cancer cells promote tumor cell migration and invasion, we performed Transwell assays. Macrophages were preconditioned with CM from PTX-treated cancer cells, and the resulting macrophage CM was applied to breast cancer cells in the upper chamber for 24 h (Fig. 7h). CM from macrophages preconditioned with PTX-treated cancer cell CM significantly enhanced cancer cell migration and invasion, whereas CM from Nab-PTX-treated cells had no effect (Fig. 7i and Supplementary Fig. 24a–g). Similarly, Trem2-/- BMDMs failed to enhance migration or invasion, regardless of conditioning with PTX- or Nab-PTX-treated CM (Supplementary Fig. 24h–j). Control experiments confirmed that PTX alone did not directly stimulate cancer cell migration or invasion (Supplementary Figs. 25a–e). Collectively, these findings indicate that the pro-migratory and pro-invasive effects are mediated by TREM2+ macrophages rather than direct effects of PTX on tumor cells.
To identify the paracrine factors secreted by TREM2+ macrophages that facilitate metastasis, we performed a cytokine array on CM from Trem2+/+ and Trem2−/− BMDMs. Serpin E1, HGF, CCL3, and CXCL2 were significantly elevated in Trem2+/+ BMDM CM relative to Trem2−/− CM (Fig. 7j), with corresponding increases in mRNA levels (Supplementary Fig. 26a). Direct PTX treatment of macrophages only modestly increased Serpin E1 expression and did not significantly affect the other genes (Supplementary Fig. 26b). In contrast, macrophages incubated with CM from PTX-treated cancer cells exhibited a robust ( ~ 7-fold) increase in Serpin E1 compared to control (Supplementary Fig. 26c), indicating that induction of these cytokines is primarily TREM2-dependent. Recombinant Serpin E1, HGF, CCL3, and CXCL2 enhanced Py8119 cell migration and invasion (Fig. 7k and Supplementary Fig. 26d), while neutralization of Serpin E1 in Trem2+/+ CM significantly impaired these effects (Fig. 7l and Supplementary Fig. 26e), confirming that Serpin E1 is essential for TREM2-driven pro-metastatic activity.
Immunofluorescence co-staining of primary tumor sections demonstrated that Serpin E1 expression was markedly upregulated in PTX-treated tumor-bearing mice, but not in Nab-PTX-treated mice (Supplementary Figs. 27a, b). Co-treatment with the FGFR inhibitor IGR significantly reduced Serpin E1 levels in tumor-associated macrophages relative to PTX alone (Supplementary Figs. 27c, d), confirming FGF2-dependent induction. Functional validation using an orthotopic Py8119 breast cancer model revealed that combining PTX with the Serpin E1 inhibitor PAI-039 markedly suppressed lung metastasis compared to PTX alone (Supplementary Figs. S28a–c). Furthermore, co-injection of Py8119 cells with Trem2−/− BMDMs overexpressing Serpin E1 (Trem2-/- Serpin E1OE) into Trem2−/− significantly enhanced tumor growth and lung metastasis relative to Trem2−/− BMDMs alone (Supplementary Figs. 28d–h), underscoring the requirement of Serpin E1 for PTX-induced metastasis.
Notably, Serpin E1, HGF, CCL3, and CXCL2 have previously been implicated in activating the epithelial-mesenchymal transition (EMT) program54,55,56,57. To determine their effect on EMT in breast cancer cells, we measured canonical EMT markers, observing downregulation of E-cadherin and upregulation of Vimentin by western blot (Fig. 7m). Multiplex immunofluorescence of clinical samples confirmed decreased E-cadherin and increased Vimentin expression in PTX-treated tumors, coinciding with expanded TREM2+ macrophages, whereas Nab-PTX-treated samples showed no such changes (Supplementary Figs. 29a and b). Collectively, these findings indicate that PTX-induced TREM2+ macrophages promote tumor cell migration and invasion by secreting EMT-stimulating cytokines (Fig. 7n).
Discussion
Our study demonstrates that the clinical efficacy of PTX in breast cancer is inferior to that of Nab-PTX, potentially due to their distinct effects on the tumor immune microenvironment. Previous work has attributed the superior efficacy of Nab-PTX to enhanced tumor penetration and retention via receptor-mediated transcytosis58,59,60, as well as to the avoidance of corticosteroid premedication, which is required for PTX and can dampen antitumor immune responses, thereby contributing to lower pCR rates61,62,63. We observed that PTX, but not Nab-PTX, increases TREM2+ macrophage infiltration in primary tumors. Given that TREM2+ macrophages promote breast cancer metastasis, this finding provides a potential explanation for the inferior antitumor efficacy of PTX (Fig. 8).
PTX, but not Nab-PTX, promotes lung metastasis by inducing TREM2+ macrophage recruitment. Mechanistically, PTX enhances the ATF3-FGF2 axis in breast cancer cells; secreted FGF2 activates the EGR1–TREM2–EMT cytokine axis in macrophages. Created in BioRender. Xing, Y. (2026) https://BioRender.com/6hxlbow.
Previous studies in murine mammary tumor models reported that PTX promotes metastasis through perivascular TIE2+ macrophages10,11,13, and by upregulating VEGFR1 on endothelial cells12 and increasing the vasculature properties14, stimulating angiogenesis. These studies revealed a pro-metastatic mechanism of PTX through angiogenesis, whereas our findings uncover a distinct mechanism, with PTX promoting lung metastasis by acting on host TREM2+ macrophages and modulating multiple cytokines within the TIME. Notably, the Macro_IL1B population was expanded in pre-PTX but was nearly absent in pre-Nab-PTX samples. Intriguingly, genetic or pharmacologic blockade of IL-1B signaling has been reported to accelerate metastasis64, suggesting that the Macro_IL1B subset may exert a net protective effect. However, the precise role of IL-1B remains ambiguous, and further preclinical studies are warranted.
Mechanistically, PTX does not directly enhance tumor cell metastasis. Instead, it induces tumor cells to secrete FGF2, which upregulates TREM2 in macrophages and stimulates the secretion of EMT-promoting cytokines, thereby facilitating tumor cell EMT and metastasis. However, Nab-PTX does not elicit these effects. Specifically, PTX increases chromatin accessibility at the ATF3 locus in tumor cells, leading to elevated ATF3 and its downstream target FGF2. Consequently, macrophages exposed to FGF2 upregulate EGR1 and TREM2. These results reveal how PTX drives infiltration of TREM2+ macrophages. In contrast, Nab-PTX reshapes myeloid cell infiltration toward pro-inflammatory macrophages and mast cells8, highlighting a mechanistic basis for its superior efficacy.
TREM2 is predominantly expressed on tumor-associated macrophages (TAMs) and myeloid-derived suppressor cells (MDSCs)65, and its inhibition has been shown to enhance checkpoint immunotherapy efficacy across multiple cancers, including sarcoma, colorectal, and breast cancers15. TREM2+ cells accumulate at the invasive margins of lung metastases in both humans and mice19, supporting their role in establishing a pro-metastatic niche. Here, we demonstrate that PTX-induced TREM2+ macrophages stimulate EMT in breast cancer, revealing a pro-metastatic function and providing a mechanistic explanation for the inferior efficacy of PTX relative to Nab-PTX. Furthermore, TREM2+ macrophages increased not only in primary tumors but also in peripheral blood, suggesting systemic circulation rather than confinement to the tumor site. Given that TREM2 deficiency promotes macrophage apoptosis66 and that depletion of immunosuppressive macrophages suppresses tumor growth and metastasis67, further studies are needed to clarify whether reduced lung metastasis in Trem2-/- mice results from macrophage depletion or specific TREM2 signaling disruption.
Previous studies indicate that TREM2 depletion or blockade enhances T cell responses and improves checkpoint immunotherapy outcomes15. We previously reported that PTX impairs infiltration of CXCL13-expressing T cells, compromising combination checkpoint immunotherapy in breast cancer8,20. Here, we show that PTX stimulates TREM2+ macrophage expansion, contributing to its inferior monotherapy efficacy. scRNA-seq analysis of our previously published dataset8 further suggests that PTX-induced TREM2-overexpressing macrophages exert immunosuppressive effects on multiple immune populations, including T cells, B cells, mast cells, DCs, neutrophils, NK cells, and plasma cells (Supplementary Fig. 30). Thus, PTX-driven TREM2+ macrophage expansion likely impacts both chemotherapy and immunotherapy responses, supporting TREM2 as a potential therapeutic target to enhance clinical outcomes.
While our study focused on lung metastasis, breast cancer also disseminates to the liver, bone, and brain, and PTX is widely used in other malignancies such as lung cancer, melanoma, and esophageal cancer. Whether PTX promotes metastasis to these organs or in other tumor types warrants further investigation. Trem2−/− BMDMs fail to promote migration and invasion of Py8119 cells (Supplementary Figs. 17j, k), and PTX-induced lung metastasis is greatly reduced in Trem2−/− mice (Fig. 3a–h) or upon ASO-mediated TREM2 knockdown (Fig. 4), highlighting TREM2 as a critical mediator of PTX-driven metastasis. Collectively, these findings suggest that TREM2 blockade could enhance the efficacy of PTX and its combination with checkpoint immunotherapy15.
In addition, several limitations should be considered. First, although our data suggest that the inferior efficacy of PTX is linked to the expansion of TREM2+ macrophages, validation in larger patient cohorts is needed. Multivariable analyses stratified by tumor subtype, incorporating key covariates such as tumor size, grade, number of positive lymph nodes, patient age, and body mass index, are essential to fully assess the impact of PTX on TREM2+ macrophage infiltration. Second, while we identified PTX-treated cancer cells as a major source of FGF2, contributions from other tumor microenvironment components, such as cancer-associated fibroblasts, cannot be excluded. Third, in addition to FGF2, our cytokine array revealed that PTX-treated tumor cells secrete other factors, including GDF15, which may also modulate TREM2 expression in macrophages. Notably, tumor-derived GDF15 has been reported to promote tumor progression by regulating macrophage activity via the NF-κB pathway68. The functional role of these alternative cytokines warrants further investigation. Finally, although Trem2 ASO therapy demonstrated efficacy in preclinical models, its translational potential requires additional evaluation, including optimization of combination regimens and identification of patient populations most likely to benefit.
In conclusion, our study clarifies the distinct mechanisms underlying the differential efficacy of PTX and Nab-PTX in breast cancer. PTX, unlike Nab-PTX, promotes lung metastasis by upregulating TREM2 and expanding TREM2+ macrophages, which underlies its inferior therapeutic efficacy relative to Nab-PTX.
These results indicate that integrating PTX with TREM2-targeted therapy could enhance the clinical outcomes of PTX while limiting metastasis.
Methods
Ethical approval
This study complied with relevant ethical regulations, and all clinical samples were coded to maintain patient confidentiality and studies were approved by Tianjin Medical University Cancer Institute & Hospital Ethics Committee (Ethics approval number: bc20241748) and Peking University Biomedical Ethics Committee (Ethics approval number: PUIRB-YS2025237), in accordance with the declaration of Helsinki. All animal experiments were approved by the Peking University Biomedical Ethics Committee (Ethics approval number: DLASBD0408).
Patient samples
A total of formalin fixed and paraffin embedded tumor tissues from patients after treatment with indicated chemotherapy (Supplementary Tables 5 and 6), and tumor tissues from patients with/without lung metastasis (Supplementary Tables 7 and 8) were selected retrospectively by database extraction from the archives of Tianjin Medical University Cancer Institute & Hospital and Peking University Third Hospital (sex self-reported, as female is a high-risk group for breast cancer). Clinical and pathological characteristics of the patient tumors are presented in Supplementary Tables 5–8. All tumor tissue samples were obtained in compliance with the informed consent policy. We have obtained consent from the patients to publish clinical information. No participant remuneration was applicable. All clinical samples were coded to maintain patient confidentiality and studies were approved by Tianjin Medical University Cancer Institute & Hospital Ethics Committee (approval bc20241748) and Peking University Biomedical Ethics Committee (approval PUIRB-YS2025237), in accordance with the declaration of Helsinki.
Mice
Wild type C57BL/6 and BALB/C mice (Female, 6–8 weeks old) were purchased from the Laboratory Animal Center of Peking University Health Science Center. The MMTV-PyMT (FVB/N) strain was originally obtained from Z. Yu’s laboratory and was backcrossed for seven generations to the C57BL/6 background. Trem2-KO mice (Cat. NO. NM-KO-190402) were purchased from Shanghai Model Organisms Center Inc (SMOC).
The female knockout C57BL/6 J mice for Trem2 were generated using CRISPR-Cas9–based targeting and nonhomologous end joining. In founder lines, genotyping on genomic DNA from mouse tails was verified by multiplex polymerase chain reaction (PCR) to obtain Trem2−/− mice, which were further intercrossed to yield healthy offspring.
Mice were housed in a pathogen-free animal facility at Laboratory Animal Center of Peking University Health Science Center with a 12 h light/dark cycle, constant temperature and humidity, and fed standard rodent chow and water ad libitum. All animal experiments were approved by the Peking University Biomedical Ethics Committee, and the approval number is DLASBD0408. None of the animals used in these experiments exceeded ethical requirements. The maximum permitted tumor volume was 2,000 mm³. The body weight of the mice was maintained at 80% or above of their initial weight. Genomic DNA extracted from mouse tail biopsies was subjected to standard genotyping PCR using the primers specified in Supplementary Table 12. The reaction conditions were: 5 min at 94 °C; 35 cycles of 30 s at 94 °C, 30 s at 60 °C and 1 min at 72 °C, followed by 5 min at 72 °C and hold at 4 °C.
Cell lines
Human embryonic kidney cell line HEK293T (ATCC, CRL-3216), human breast cancer cell line SUM159 (CLS Cell Lines service, 305116), mouse melanoma cell lines B16-F10 (ATCC, CRL-6475) and mouse macrophage cell line Raw264.7 (ATCC, TIB-71) were cultured in Dulbecco’s Modified Eagle Medium (DMEM). Human breast cancer cell line BT549 (ATCC, HTB-122), Mouse breast cancer cell line 4T1-Luc (ATCC, CRL-2539-LUC2) and human macrophage cell lines THP1 (ATCC, TIB-202) were cultured in RPMI-1640 medium. Human breast cancer cell line MDA-MB-231(ATCC, CRM-HTB-26) was cultured in L15 medium. Mouse breast cancer cell line Py8119 (ATCC, CRL-3278) was cultured in Ham’s F12K medium. All medium were supplemented with 10% FBS (Invitrogen), 1% penicillin-streptomycin. All cell lines were maintained at 37 °C with 5% CO2 in a humidified atmosphere and passaged using 0.25% trypsin/0.02% EDTA for dissociation at 80% confluence. For transient transfection, 50%–60% confluent cells were transfected with plasmids or ASO using polyethylenimine (PEI, Polyscience, #24765), Lipo2000 (Invitrogen, #11668030) or transfection reagent (Zeta Life, #AD600150) according to the manufacture’s protocol. The FGF2 protein (HY-P73052AF) was purchased from MedChemExpress and used with indicated concentration. The Serpin E1 protein (HY-P71133), HGF protein (HY-P71133), CCL3 protein (HY-P7768AF) and CXCL2 protein (HY-P7258) were purchased from MedChemExpress and used at a concentration of 10 ng/mL. The CDE-096 (HY-120516) was purchased from MedChemExpress and used at a concentration of 10 μg/mL. The neutralizing FGF2 antibody (05117) was purchased from Millipore and used at a concentration of 10 μg/mL. The FGF2 inhibitor (IGR, HY-13311) was purchased from MedChemExpress and used at a concentration of 5 μg/mL.
Meta-analysis
Search Strategy
The meta-analysis adhered to the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) guidelines69. Two researchers (RZ and MX) independently conducted a systematic search across three major databases: PubMed, Embase, and the Cochrane Library. The keywords utilized and a detailed description of the search process are provided in Supplementary Table 1. The search was completed on December 17th, 2024, and was restricted to English-language publications. The inclusion criteria for the studies were as follows: (1) patients with a histologically and cytologically confirmed diagnosis of breast cancer, (2) an experimental intervention group receiving a treatment regimen that included Nab-PTX, and (3) a control group receiving a treatment regimen that included PTX. In addition, to ensure comparability, the experimental and control groups within each study were required to have identical treatment protocols, differing only in the type of taxanes administered.
Data extraction
In each trial included in this study, we systematically gathered essential information encompassing the first author, year of publication, region, the type of study, registration ID, and the total number of enrolled patients. In addition, we extracted detailed therapeutic regimens and pertinent data of specific outcomes, notably the objective response rate (ORR) and the pathologic complete response (pCR) for the intention-to-treat (ITT) population. ORR refers to the proportion of the complete and partial responses. The pCR rate refers to the proportion of cases without invasive carcinoma in both breast tissue and the axillary lymph nodes after taxane-containing neoadjuvant therapy. To ensure the utmost precision and reliability of the data, two independent researchers (RZ and MX) carried out the extraction process and subsequently cross-verified the information using a predefined table. Any discrepancies that arose during this process were diligently resolved through consensus, facilitated by the involvement of a third researcher (HM).
Bias assessment
Two researchers (RZ and MX) independently assessed the bias risk of RCTs using the risk of bias (RoB2) tool, and a third researcher (HM) was consulted when a disagreement arose between the two researchers. We also used the Newcastle-Ottawa scale (NOS) to assess the bias risk of cohort and retrospective studies70.
Statistical analysis
Statistical analysis was implemented using R, version 4.1.3 (http://www.r-project.org/). The pooled analysis results for ORR and pCR were presented as odds ratios (OR) with 95% confidence intervals. were reported in terms of odds ratios (OR) with their corresponding 95% confidence intervals. Heterogeneity across studies was assessed using the I2 test, where P < 0.1 or I2 > 50% indicated a significant heterogeneity, while P > 0.1 or I2 < 50% suggested no considerable heterogeneity. To ensure the robustness of our findings, a random-effects model was applied to all studies. Further, we employed meta-regression (Supplementary Table 4) and subgroup analyses to explore the sources of heterogeneity in our meta-analysis.
Single-cell RNA sequencing
Human published data from our previous study can be accessed from the Gene Expression Omnibus (GEO) under the accession number GSE266919. Mice scRNA-seq libraries were performed with the Chromium Single Cell 3’ Reagent Kits v3 (10× Genomics) according to the manufacturer’s instructions. Briefly, approximately 1 × 105 cells per FACS-sorted cell were washed with 0.04% bovine serum albumin Dulbecco’s PBS three times and were resuscitated to a concentration of 700 to 1200 cells/μL (viability, ≥ 85%). Cells were captured in droplets at a targeted cell recovery of cells. After the reverse transcription, emulsions were broken, and barcoded cDNA was purified with Gel Beads, followed by PCR amplification. The sequencing libraries were then constructed using amplified cDNA. The raw scRNA-seq data are available under accession code GSE285329. Before bioinformatic analysis, cells were filtered with unique molecular identifier counts below 30,000 and gene counts between 200 and 5000 to remove low-quality cells, and then filtered out the cell content of more than 10% mitochondrial genes of the total expressed genes. Then, the dimensionality reduction and clustering analysis was performed using Seurat v2.3. Normalize Data and ScaleData functions are used to normalize and scale all gene expression, and with the FindVariableFeatures function, the 3000 highly variable genes genes were identified for principal components analysis. The genes were divided into multiple groups with FindClusters. The identified clusters were then visualized in two-dimensional space produced with Uniform Manifold Approximation and Projection visualization.
Xenografts in mice
The 4T1-Luc tumor-bearing mouse model was established by orthotopically implanting 6 × 105 4T1-Luc cancer cells in 100 μL of PBS into the right second mammary fat pads of the BALB/C mice. For bioluminescence imaging, a solution of luciferase substrate D-luciferin (150 mg/kg) was intraperitoneally injected into the mice, followed by imaging using the In vivo imaging system (IVIS) (PerkinElmer).
About 8 × 105 Py8119 cells were orthotopically injected in the right mammary fat pads of C57BL/6 mice to build a Py8119 tumor-bearing mouse model. Mice were implanted subcutaneously with 1 × 106 B16-F10 cancer cells, and injected intravenously with 5 × 105 B16-F10 cancer cells to establish the tumor-bearing mouse models. For the co-implantation mouse model, Py8119 tumor cells and primary BMDMs were mixed at a ratio of 1: 2 and 1 × 106 total cells were orthotopically injected into the right second mammary fat pads of the C57BL/6 J mice. The tumor volumes were calculated according to the formula as Volume = (Length × Width2)/2. Mice were euthanized after indicated days. During the survival observation period, mice were euthanized once the tumor volume exceeded ethical requirements or when the mice showed paralysis, convulsions, or such phenomena, with the time of death recorded accordingly.
The PTX (HY-B0015) was purchased from MedChemExpress and used at a concentration of 9 mg/kg. The Nab-PTX (HY-P99974) was purchased from MedChemExpress and used at a concentration of 9 mg/kg. The FGF2 inhibitor (IGR, HY-13311) was purchased from MedChemExpress and used at a concentration of 20 mg/kg. The Serpin E1 inhibitor (PAI-039, HY-15253) was purchased from MedChemExpress and used at a concentration of 20 mg/kg.
Immunofluorescence staining
After being fixed with 4% paraformaldehyde, cells were blocked with 3% BSA and then incubated with primary antibodies against TREM2 (ab318262, Abcam, 1: 100) overnight at 4 °C, followed by Alexa Fluor 488 (GB25303, Servicebio, 1: 200) incubation. After being deparaffinized and rehydrated, paraffin sections were repaired in Tris-EDTA buffer and then followed similar steps. The paraffin sections were incubated with primary antibodies against TREM2 (ab318262, Abcam, 1: 100) and CD68 (66231-2-Ig, Proteintech, 1: 100) and with secondary antibodies against Alexa Fluor 488 (GB25303, GB25301, Servicebio, 1: 200) and Alexa Fluor 594 (GB28301, GB28303, Servicebio, 1: 200). Fluorescent images were acquired using ZEISS AXIOSCAN 7. Image analysis was performed using the HALO image analysis software (Indica labs). The specific analytical method using HALO software is as follows. We first selected representative areas and counted viable DAPI-positive cells while excluding necrotic areas. We then analyzed TREM2 expression on CD68-positive macrophages. Antibodies are listed in the Supplementary Table 13.
Flow cytometry analysis (FACS)
Flow cytometry was used for the analysis of TREM2 expression in macrophages. Briefly, the indicated macrophages were washed twice with cold wash-cytometry buffer and were stained with the indicated antibodies for 30 min and kept shielded from light, followed by the cold wash-cytometry buffer, and washed twice before analysis.
Flow cytometry was performed on tumor tissue samples and blood samples as previously described. Tumor tissues were collected and dissociated into homogenates using a Gentle MACS Dissociator with collagenase type I (1 mg/mL, Gibco) and collagenase type III (1 mg/mL, Gibco) for 30 min. Single-cell suspensions were washed with Red Cell Lysis Solution and filtered by nylon mesh filters and then washed with the cold wash-cytometry buffer. Blood samples were collected into collection tubes (EDTA-coated), followed by red blood cells lysis with Red Cell Lysis Solution. The cells were then resuspended with the cold wash-cytometry buffer. After being incubated with Fc receptor blocker for 10 min at 4 °C, the cells were then incubated with the antibodies (7-AAD (1 μL/test), BV605-anti-CD45 (1 μL/test), PerCP-Cy5.5-anti-CD45 (1 μL/test), FITC-anti-F4/80 (1 μL/test), PE-anti-CD11b (1 μL/test), APC-Cy7-anti-CD11b (1 μL/test) and APC-anti-TREM2(5 μL/test)) for 30 min at 4 °C. The cells were then washed and resuspended with the cold wash-cytometry buffer before analysis using a Beckman CytoFLEX (Beckman Colter), and data were analyzed using FlowJo software (BD Biosciences). Antibodies are listed in the Supplementary Table 13.
Lung metastasis analysis
Lungs were collected and washed by PBS, and then fixed in the Bouin’s fixative. The number of lung metastasis lesions were counted manually. After being embedded with paraffin, the fixed lung tissues were also embedded in paraffin blocks, sectioned at 5 μm thickness, and then stained with H&E to assess the structure.
Generation and culture of mouse BMDMs
Mouse macrophages were isolated from the bone marrow of 6–8 weeks old female C57BL/6 mice as previously described. Briefly, the lower limb was exposed by removing the skin. After collecting the femur and tibia, bone marrow cells were flushed with sterile PBS using a 26 G needles. Cell pellets were isolated by centrifuging at 500 × g at 4 °C for 5 min and resuspended in PBS. After being filtered through a 70 μm cell strainer and centrifuging, Cells were seeded in 10 mL complete DMEM with macrophage colony-stimulating factor (M-CSF) for 7-day continuous differentiation. On day 7, BMDM was assessed and then prepared for both in vitro co-culture and in vivo injection experiments.
Western blotting
Cells were lysed in RIPA lysis buffer supplemented with the cocktail inhibitor (Roche) and PhosSTOP (Roche). Samples supplemented with loading buffer were heated at 100 °C for 9 min and separated in SDS-PAGE, and then transferred to polyvinylidene fluoride (PVDF) membranes using standard techniques. After being blocked with 5% skim milk for 1 h, membranes were incubated with primary antibodies (TREM2 (ab305103, ab318262, Abcam, 1: 1000), FGF2 (11234-1-AP, Proteintech, 1: 800), p-ERK1/2 (4370 T, Cell Signaling Technology, 1: 1000), ERK1/2 (4695 T, Cell Signaling Technology, 1: 1000), EGR1 (22008-1-AP, Proteintech, 1:1000), E-Cadherin (3195 T, Cell Signaling Technology, 1: 500), Vimentin (5741, Cell Signaling Technology, 1: 500), Actin (sc-58673, Santa Cruz Biotechnology, 1: 2000), Tubulin (ab7291, Abcam, 1: 2000)) overnight at 4 °C, followed by being incubated with HRP-conjugated secondary antibodies for 1 h at room temperature. The membranes were visualized with the EZ-ECL Chemiluminescence Detection Kit for HRP (Biological Industries, # 20-500-1000) by ChampChemiTM (SageCreation). Antibodies are listed in the Supplementary Table 13.
RNA extraction and quantitative real-time PCR
Total RNA was isolated from cells using Trizol reagent (Invitrogen) according to the manufacture’s instruction. RNA was reverse-transcribed to cDNA using the HiScript II Q RT SuperMix for qPCR ( + gDNA wiper) (Vazyme). qPCR was performed using FastStart Essential DNA Green Master in a reaction system of 10 μL. All PCR reactions were run at least in triplicate, and the target mRNA expression was normalized to actin, respectively. All PCR reactions were run at least in triplicate, and the target mRNA expression was normalized to actin, respectively. Relative expression of each target gene was calculated by the 2-ddCt method. All primers are shown in Supplementary Table 12.
Cytokine array
Volume of indicated medium was analyzed using the Proteome Profiler Mouse XL Cytokine Array Kit (R&D System, ARY028) according to the manufacturer’s instructions.
Bulk RNA sequencing
The total RNA was isolated from cells using Trizol reagent (Invitrogen) according to the manufacture’s instruction, followed by being fragmented into shorter fragments with buffer. Double-stranded cDNA was then synthesized using those short fragments.
The resulting cDNA was performed with complex method according to the manufacture’s protocol. The sequence library was prepared on an Illumina NovaSeq 6000 platform. Align Reads to the Reference Genome, we took the GRch38 genome as the reference genome for this project. Bulk RNA-seq data of BT549 cells, SUM159 cells and MDA-MB-231 cells are available at the GEO (accession number GSE285485; https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE285485).
Multiplex immunofluorescence staining and analysis
Multiple immunofluorescence staining was performed with TissueGnostics Multiple IHC Assay Kit (Opal 7 Solid Tumor Immunology kit, PerkinElmer). After being deparaffinized and rehydrated, the antigen of paraffin sections repaired in Tris-EDTA buffer with microwave, the nonspecific antigen was blocked by block buffer. For multiple rounds of cyclic dyeing, slides were incubated with primary antibody (TREM2 (ab305103, ab318262, Abcam, 1: 100), CD68 (ab303565, Abcam, 1: 100; 66231-2-Ig, Proteintech, 1: 100), FGF2 (11234-1-AP, Proteintech, 1: 100), E-Cadherin (3195 T, Cell Signaling Technology, 1: 100), Vimentin (5741, Cell Signaling Technology, 1: 100)) at 4 °C overnight or 37 °C for 2 h, followed by incubating with secondary antibody at 37 °C for 30 min. After being treated with antigen retrieval buffer, slides were repeated the above steps for multiple staining. The nuclei were stained with DAPI for 10 min, and then photographed using ZEISS AXIOSCAN 7. Image analysis was performed using the HALO image analysis software (Indica labs). Antibodies are listed in the Supplementary Table 13.
Luciferase reporter assay
The indicated pGL4.10-promoter plasmid, a pRL-tk Renilla luciferase reporter plasmid, and a vector encoding the indicated transcription factor or a control vector were co-transfected into the indicated cells. The firefly and Renilla luciferase activities were measured with the Dual-Luciferase Reporter Assay System (Promega). Relative firefly luciferase activity was normalized to the Renilla luciferase activity.
ChIP-qPCR
A total of 5 × 107 indicated cells were cross-linked with 1% formaldehyde for 10 min at 4 °C and quenched with 125 Mm glycine for 5 min. After washing twice with PBS, pelleted cells were suspended in lysis buffer and incubated for 15 min on ice. Cells were centrifugated at 3000 × g for 5 min, followed by being resuspended in nuclear lysis buffer. The lysates were then sonicated for 6 min by ultrasonication. After centrifugation, the supernatants containing chromatin were diluted 5 times with dilution buffer and then incubated with antibodies (5 μg) overnight at 4°C. The bead-antibody complexes were incubated with chromatin at 4 °C for 4 h. The beads were then washed twice with ChIP buffer, high salt buffer and low salt buffer. The bound chromatin was eluted and reverse-crosslinked at 65 °C overnight. DNA was purified by the MinElute PCR purification kit. qPCR was then performed using SYBR Green master mix. ChIP-qPCR primers can be found in Supplementary Table 12.
ATAC-seq and data analysis
A total of 500,00 cells were harvested and performed according to the manufacturer instructions (Novoprotein). The sequence library was prepared on an Illumina HiSeq2500 machine (Novogene). Raw reads were aligned to the GRch38 and mm10 genome as the reference genome for this project. The distance between the transcription start site (TSS) of the peak-related gene and the peak was calculated. Peaks were performed using the Integrative Genomics Viewer software. ATAC-seq data of BT549 cells and BMDMs are available at the GEO (accession number GSE295918; https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE295918).
Transwell migration and invasion assay
Cells subjected to the indicated treatments were resuspended in serum-free medium and seeded into upper chambers of the Transwell with 8 mm micropores (Millipore). The bottom chamber was filled with 0.5 mL culture medium supplemented with 20% FBS. For the invasion assay, the upper chambers were prepared with 1 × MaxGel ECM (Sigma-Aldrich). After 24 h or 48 h of incubation at 37 °C, the cells remaining on the top membranes were removed, whereas the cells adhering to the lower surface were fixed, stained with 0.2% crystal violet solution and counted under a microscope, and then photographed using microscopy. Image analysis was performed using the Image J software (NIH).
Statistics and reproducibility
Statistical results are presented as means ± SD from at least three independent biological replicates. Student’s t test was used to compare the two difference experimental groups. Kaplan-Meier curves were used for survival curves. ANOVA was used for two or more groups. Correlations between variables were assessed using Spearman’s rank correlation. To minimize observer bias, investigators were blinded to group allocation during experiments and outcome assessment. All statistical analyses were conducted with GraphPad Prism 9.0 (GraphPad software) and SPSS (V26, IBM Crop., Armonk, NY, USA), with statistical significance set at P < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Published human data from our previous study8 can be accessed from GEO under the accession number GSE266919. Raw data for this dataset is not available but could be provided for scientific research upon request to Prof. Fei Ma at drmafei@126.com. Code used for all processing and analysis is also available upon request. The raw single-cell RNA sequencing data generated in this study have been deposited in the GEO database under accession code GSE285329. The raw bulk RNA sequencing data generated in this study have been deposited in the GEO database under accession code GSE285485]. The raw ATAC-seq data generated in this study have been deposited in the GEO database under accession code GSE295918. All remaining data supporting the findings described in this manuscript are available in the article and its Supplementary Information files and/or from the corresponding author upon request. Source data are provided inthis paper.
References
Weigelt, B., Peterse, J. L. & van ‘t Veer, L. J. Breast cancer metastasis: markers and models. Nat. Rev. Cancer 5, 591–602 (2005).
Mukama, T. et al. Risk-adapted starting age of screening for relatives of patients with breast cancer. JAMA Oncol. 6, 68–74 (2020).
DeMichele, A., Yee, D. & Esserman, L. Mechanisms of resistance to neoadjuvant chemotherapy in breast cancer. N Engl. J. Med. 377, 2287–2289 (2017).
Gampenrieder, S. P., Rinnerthaler, G. & Greil, R. Neoadjuvant chemotherapy and targeted therapy in breast cancer: past, present, and future. J. Oncol. 2013, 732047 (2013).
Guarneri, V., Dieci, M. V. & Conte, P. Enhancing intracellular taxane delivery: current role and perspectives of nanoparticle albumin-bound paclitaxel in the treatment of advanced breast cancer. Expert Opin. Pharmacother. 13, 395–406 (2012).
Montero, A. J., Adams, B., Diaz-Montero, C. M. & Gluck, S. Nab-paclitaxel in the treatment of metastatic breast cancer: a comprehensive review. Expert Rev. Clin. Pharmacol. 4, 329–334 (2011).
Brufsky, A. nab-Paclitaxel for the treatment of breast cancer: an update across treatment settings. Exp. Hematol. Oncol. 6, 7 (2017).
Zhang, Y. et al. Distinct cellular mechanisms underlie chemotherapies and PD-L1 blockade combinations in triple-negative breast cancer. Cancer Cell 43, 446–463 (2025).
Keklikoglou, I. et al. Chemotherapy elicits pro-metastatic extracellular vesicles in breast cancer models. Nat. Cell Biol. 21, 190–202 (2019).
De Palma, M., Biziato, D. & Petrova, T. V. Microenvironmental regulation of tumour angiogenesis. Nat. Rev. Cancer 17, 457–474 (2017).
De Palma, M. et al. Tie2 identifies a hematopoietic lineage of proangiogenic monocytes required for tumor vessel formation and a mesenchymal population of pericyte progenitors. Cancer Cell 8, 211–226 (2005).
Daenen, L. G. et al. Chemotherapy enhances metastasis formation via VEGFR-1-expressing endothelial cells. Cancer Res. 71, 6976–6985 (2011).
Karagiannis, G. S. et al. Neoadjuvant chemotherapy induces breast cancer metastasis through a TMEM-mediated mechanism. Sci. Transl. Med. 9, eaan0026 (2017).
Chang, Y. S., Jalgaonkar, S. P., Middleton, J. D. & Hai, T. Stress-inducible gene Atf3 in the noncancer host cells contributes to chemotherapy-exacerbated breast cancer metastasis. Proc. Natl. Acad. Sci. USA 114, E7159–E7168 (2017).
Molgora, M. et al. TREM2 Modulation Remodels the Tumor Myeloid Landscape Enhancing Anti-PD-1 Immunotherapy. Cell 182, 886–900 (2020).
Zhong, J. et al. Distinct roles of TREM2 in central nervous system cancers and peripheral cancers. Cancer Cell 42, 968–984 e9 (2024).
Esparza-Baquer, A. et al. TREM-2 defends the liver against hepatocellular carcinoma through multifactorial protective mechanisms. Gut 70, 1345–1361 (2021).
Park, M. D. et al. TREM2 macrophages drive NK cell paucity and dysfunction in lung cancer. Nat. Immunol. 24, 792–801 (2023).
Yofe, I. et al. Spatial and Temporal Mapping of Breast Cancer Lung Metastases Identify TREM2 Macrophages as Regulators of the Metastatic Boundary. Cancer Discov. 13, 2610–2631 (2023).
Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39, 1578–1593 (2021).
Lv, H. et al. Efficacy and safety of nanoparticle albumin‑bound paclitaxel compared with solvent‑based paclitaxel in adjuvant therapy for breast cancer: A retrospective study. Oncol. Lett. 28, 509 (2024).
Rugo, H. S. et al. Randomized phase III trial of paclitaxel once per week compared with nanoparticle albumin-bound Nab-Paclitaxel once per week or Ixabepilone with bevacizumab as first-line chemotherapy for locally recurrent or metastatic breast cancer: CALGB 40502/NCCTG N063H (Alliance). J. Clin. Oncol. 33, 2361–2369 (2015).
Rugo, H. et al. Abstract GS3-06: Long-term follow-up of CALGB 40502/NCCTG N063H (Alliance): A randomized phase III trial of weekly paclitaxel (P) compared to weekly nanoparticle albumin bound nab-Paclitaxel (NP) or ixabepilone (Ix) +/- bevacizumab as first-line therapy for locally recurrent or metastatic breast cancer (MBC). Cancer Res. 78, GS3-06 (2018).
Brufsky, A. et al. A phase II randomized trial of cobimetinib plus chemotherapy, with or without atezolizumab, as first-line treatment for patients with locally advanced or metastatic triple-negative breast cancer (COLET): primary analysis. Ann. Oncol. 32, 652–660 (2021).
Luhn, P. et al. Comparative effectiveness of first-line nab-paclitaxel versus paclitaxel monotherapy in triple-negative breast cancer. J. Comp. Eff. Res. 8, 1173–1185 (2019).
Han, X. & Wang, Z. Efficacy of albumin-bound paclitaxel in the treatment of advanced refractory breast cancer and its effect on serum resistin. J. Buon 25, 681–687 (2020).
Untch, M. et al. NAB-Paclitaxel improves disease-free survival in early breast cancer: GBG 69-geparSepto. J. Clin. Oncol. 37, 2226–2234 (2019).
Miles, D. et al. Final results from the PERUSE study of first-line pertuzumab plus trastuzumab plus a taxane for HER2-positive locally recurrent or metastatic breast cancer, with a multivariable approach to guide prognostication. Ann. Oncol. 32, 1245–1255 (2021).
Bachelot, T. et al. Preliminary safety and efficacy of first-line pertuzumab combined with trastuzumab and taxane therapy for HER2-positive locally recurrent or metastatic breast cancer (PERUSE). Ann. Oncol. 30, 766–773 (2019).
Loibl, S. et al. Dual HER2-blockade with pertuzumab and trastuzumab in HER2-positive early breast cancer: a subanalysis of data from the randomized phase III GeparSepto trial. Ann. Oncol. 28, 497–504 (2017).
Untch, M. et al. Nab-paclitaxel versus solvent-based paclitaxel in neoadjuvant chemotherapy for early breast cancer (GeparSepto-GBG 69): a randomised, phase 3 trial. Lancet Oncol. 17, 345–356 (2016).
Jain, M. M. et al. Paclitaxel injection concentrate for nanodispersion versus nab-paclitaxel in women with metastatic breast cancer: a multicenter, randomized, comparative phase II/III study. Breast Cancer Res. Treat. 156, 125–134 (2016).
Schmid, P. et al. Phase Ib study evaluating a triplet combination of ipatasertib (IPAT) atezolizumab (atezo), and paclitaxel (PAC) or nab-PAC as first-line (1L) therapy for locally advanced/metastatic triple-negative breast cancer (TNBC). Cancer Res. 79, CT049 (2019).
Schmid, P. et al. Abstract PS12-28: Phase 1b study evaluating a triplet combination of ipatasertib (IPAT), atezolizumab, and a taxane as first-line therapy for locally advanced/metastatic triple-negative breast cancer (TNBC). Cancer Res. 81, PS12–PS28 (2021).
Schmid, P. et al. First-Line Ipatasertib, Atezolizumab, and Taxane Triplet for Metastatic Triple-Negative Breast Cancer: Clinical and Biomarker Results. Clin. Cancer Res. 30, 767–778 (2024).
Guan, Z. Z. et al. Superior efficacy of a cremophor-free albumin-bound paclitaxel compared with solvent-based paclitaxel in chinese patients with metastatic breast cancer. Asia Pac. J. Clin. Oncol. 5, 165–174 (2009).
Gianni, L. et al. Comparing neoadjuvant nab-paclitaxel vs paclitaxel both followed by anthracycline regimens in women with ERBB2/HER2-negative breast cancer-the evaluating treatment with neoadjuvant abraxane (ETNA) trial: a randomized phase 3 clinical trial. JAMA Oncol. 4, 302–308 (2018).
Zhang, W. et al. Efficacy comparisons of solvent-based paclitaxel, liposomal paclitaxel, nanoparticle albumin-bound paclitaxel, and docetaxel after neoadjuvant systemic treatment in breast cancer. Nanomedicine 54, 102707 (2023).
Xie, F. et al. Efficacy of two-weekly nanoparticle albumin-bound paclitaxel as neoadjuvant chemotherapy for breast cancer. Nanomedicine 14, 1595–1603 (2019).
Gluz, O. et al. Nab-paclitaxel weekly versus dose-dense solvent-based paclitaxel followed by dose-dense epirubicin plus cyclophosphamide in high-risk HR+/HER2− early breast cancer: results from the neoadjuvant part of the WSG-ADAPT-HR+/HER2− trial. Ann. Oncol. 34, 531–542 (2023).
Huang, L. et al. Phase II trial of weekly nab-paclitaxel and carboplatin treatment with or without trastuzumab as nonanthracycline neoadjuvant chemotherapy for locally advanced breast cancer. Int. J. Nanomed. 10, 1969–1975 (2015).
Shi, W. et al. Nanoparticle albumin-bound paclitaxel-based neoadjuvant regimen: A promising treatment option for HER2-low-positive breast cancer. Nanomedicine 49, 102666 (2023).
Gradishar, W. J. et al. Phase III trial of nanoparticle albumin-bound paclitaxel compared with polyethylated castor oil-based paclitaxel in women with breast cancer. J. Clin. Oncol. 23, 7794–7803 (2005).
Hahnen, E. et al. Germline mutation status, pathological complete response, and disease-free survival in triple-negative breast cancer: secondary analysis of the geparSixto randomized clinical trial. JAMA Oncol. 3, 1378–1385 (2017).
Pohl-Rescigno, E. et al. Association of germline variant status with therapy response in high-risk early-stage breast cancer: a secondary analysis of the geparOcto randomized clinical trial. JAMA Oncol. 6, 744–748 (2020).
Huppert, L. A., Gumusay, O., Idossa, D. & Rugo, H. S. Systemic therapy for hormone receptor-positive/human epidermal growth factor receptor 2-negative early stage and metastatic breast cancer. CA Cancer J. Clin. 73, 480–515 (2023).
Akrida, I. et al. Epithelial to mesenchymal transition (EMT) in metaplastic breast cancer and phyllodes breast tumors. Med. Oncol. 41, 20 (2023).
Donzelli, S. et al. Paracrine signaling from breast cancer cells causes activation of ID4 expression in tumor-associated macrophages. Cells 9, 418 (2020).
Liu, M. et al. The crosstalk between macrophages and cancer cells potentiates pancreatic cancer cachexia. Cancer Cell 42, 885–903 (2024).
Li, D. et al. IFI35 regulates non-canonical NF-kappaB signaling to maintain glioblastoma stem cells and recruit tumor-associated macrophages. Cell Death Differ. 31, 738–752 (2024).
Mousset, A. et al. Neutrophil extracellular traps formed during chemotherapy confer treatment resistance via TGF-beta activation. Cancer Cell 41, 757–775 (2023).
Xiao, Y. et al. Cathepsin C promotes breast cancer lung metastasis by modulating neutrophil infiltration and neutrophil extracellular trap formation. Cancer Cell 39, 423–437 (2021).
Qi, M. et al. Targeting FAPalpha-expressing hepatic stellate cells overcomes resistance to antiangiogenics in colorectal cancer liver metastasis models. J. Clin. Invest. 132, e157399 (2022).
Tian, S. et al. SERPINH1 regulates EMT and gastric cancer metastasis via the Wnt/beta-catenin signaling pathway. Aging 12, 3574–3593 (2020).
Fu, J. et al. HGF/c-MET pathway in cancer: from molecular characterization to clinical evidence. Oncogene 40, 4625–4651 (2021).
Propper, D. J. & Balkwill, F. R. Harnessing cytokines and chemokines for cancer therapy. Nat. Rev. Clin. Oncol. 19, 237–253 (2022).
Hu, J. et al. Regulation of tumor immune suppression and cancer cell survival by CXCL1/2 elevation in glioblastoma multiforme. Sci. Adv. 7, eabc2511 (2021).
Yardley, D. A. nab-Paclitaxel mechanisms of action and delivery. J. Control Release 170, 365–372 (2013).
Desai, N., Trieu, V., Damascelli, B. & Soon-Shiong, P. SPARC Expression correlates with tumor response to albumin-bound paclitaxel in head and neck cancer patients. Transl. Oncol. 2, 59–64 (2009).
Desai, N. et al. Increased antitumor activity, intratumor paclitaxel concentrations, and endothelial cell transport of cremophor-free, albumin-bound paclitaxel, ABI-007, compared with cremophor-based paclitaxel. Clin. Cancer Res. 12, 1317–1324 (2006).
Arbour, K. C. et al. Impact of baseline steroids on efficacy of programmed cell death-1 and programmed death-ligand 1 blockade in patients with non-small-cell lung cancer. J. Clin. Oncol. 36, 2872–2878 (2018).
Jove, M., Vilarino, N. & Nadal, E. Impact of baseline steroids on efficacy of programmed cell death-1 (PD-1) and programmed death-ligand 1 (PD-L1) blockade in patients with advanced non-small cell lung cancer. Transl. Lung Cancer Res. 8, S364–S368 (2019).
Olnes, M. J. et al. Effects of Systemically Administered Hydrocortisone on the Human Immunome. Sci. Rep. 6, 23002 (2016).
Voloshin, T. et al. Blocking IL1beta pathway following paclitaxel chemotherapy slightly inhibits primary tumor growth but promotes spontaneous metastasis. Mol. Cancer Ther. 14, 1385–1394 (2015).
Wolf, E. M., Fingleton, B. & Hasty, A. H. The therapeutic potential of TREM2 in cancer. Front. Oncol. 12, 984193 (2022).
Cui, Y. et al. TREM2 deficiency aggravates renal injury by promoting macrophage apoptosis and polarization via the JAK-STAT pathway in mice. Cell Death Dis. 15, 401 (2024).
Linde, N. et al. Macrophages orchestrate breast cancer early dissemination and metastasis. Nat. Commun. 9, 21 (2018).
Ratnam, N. M. et al. NF-kappaB regulates GDF-15 to suppress macrophage surveillance during early tumor development. J. Clin. Invest. 127, 3796–3809 (2017).
Moher, D. et al. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst. Rev. 4, https://doi.org/10.1186/2046-4053-4-1 (2015).
Lo, C. K., Mertz, D. & Loeb, M. Newcastle-Ottawa Scale: comparing reviewers’ to authors’ assessments. BMC Med. Res. Methodol. 14, 45 (2014).
Acknowledgements
This study was supported by fundings from the National Natural Science Foundation of China (82230094 to H.Z., 82573243 to J.Z., 82372632 to J.Z., 82172972 to J.Z., 81972616 to H.Z.), the National Key Research and Development Program of China (2022YFA1104003 to H.Z., 2021YFC2501003 to H.Z.), Peking University Health Science Center-Michigan Medicine Joint institute for Translational and Clinical Research (BMU2025JI001 to H.Z.), and Peking University Medicine plus X Pilot Program-Key Technologies R&D Project (2024YXXLHGG006 to J.Z.). All schemes created in BioRender. Xing, Y. (https://BioRender.com/nsp747p) is licensed under CC BY 4.0.
Author information
Authors and Affiliations
Contributions
H.Z., J.Z., and Y.X. designed the research. Y.X. performed experiments. H.Z., J.Z., and Y.X. analyzed the data and wrote the manuscript. R.Z. and M.X. performed Meta-analysis. Q.L. and X.D. provided bioinformatics analysis. M.Y., J.H., and Y.S. offered experimental support for molecular and animal studies. C.L. and X.G. contributed to the collection of patient samples and performed the Immunohistochemical staining. Y.W. and Y.X. provided technique support. H.M. and F.M. provided support for clinic studies. H.Z. and J.Z. supervised this study. All authors revised and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Christos Sotiriou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xing, Y., Zhong, R., Li, Q. et al. Paclitaxel drives TREM2+ macrophage expansion underlying its inferior therapeutic efficacy compared to Nab-paclitaxel. Nat Commun 17, 2272 (2026). https://doi.org/10.1038/s41467-026-69060-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69060-5