Abstract
Cell cycle regulation is well understood in opisthokonts (fungi and metazoans) but not in plants or Apicomplexa, as some cell cycle regulators are not conserved. In opisthokonts, cell cycle progression requires the dephosphorylation of cyclin-dependent kinase (CDK) by the CDC25 phosphatase. Plants have no CDC25, and thus their mechanisms of cell cycle regulation remain elusive. Here we show that the BSL1 phosphatase dephosphorylates CDKB1 to promote mitotic entry in Chlamydomonas. Alterations of BSL1 or CDKB1 block mitotic entry after DNA replication. BSL1 shows dynamic localization through the cell cycle at the basal bodies, spindle poles and cleavage furrow. CDKB1 is hyperphosphorylated at the Thr14 and Tyr15 residues in the bsl1 mutant and in wild-type cells treated with DNA replication inhibitors. BSL1 binds to CDKB1 and dephosphorylates CDKB1 pThr14/pTyr15 in vitro. Phospho-mimicking alterations of Thr14/Tyr15 inactivate CDKB1 function, whereas phospho-blocking alterations cause sensitivity to DNA replication inhibitors, which delay cytokinesis in wild-type cells more than in cells expressing unphosphorylatable mutant CDKB1. These results indicate that CDKB1 Thr14 and Tyr15 are phosphorylated to block mitotic entry before DNA replication is complete, and BSL1 dephosphorylates CDKB1 to promote mitosis. Our study demonstrates that BSL1, a phosphatase conserved in plants and Apicomplexa but absent in fungi and animals, is a CDKB1-activating mitosis-promoting factor that has evolved additional signalling functions in receptor kinase pathways in higher plants.
Similar content being viewed by others
Main
Progression through the eukaryotic cell cycle is highly regulated to ensure faithful propagation of the genetic material to daughter cells1. The mechanisms of cell cycle regulation have been studied extensively in fungal and animal model systems, which represent the opisthokont branch of eukaryotes, but these mechanisms remain poorly understood in other branches of eukaryotes, including plants and medically relevant parasites belonging to the Apicomplexa phylum2,3. In both fungi and metazoans, orderly progression through sequential cell cycle events is orchestrated by cyclin-dependent kinase (CDK), whose activity is regulated by not only the rising and falling level of cyclin but also checkpoint-controlled phosphorylation4. CDK is inactivated due to the phosphorylation of threonine 14 and tyrosine 15 residues to prevent cell cycle progression when DNA is damaged or incompletely replicated5,6. Dephosphorylation of pThr14 and pTyr15 by the CDC25 phosphatase is required to activate CDK and recover from checkpoint arrest4,5,6,7,8. Dysfunctions in CDC25 cause defects in mitotic entry control, leading to genome instability and cancers9. This core CDK–CDC25 cell cycle regulation mechanism is conserved in fungi and animals. However, CDC25 is absent in plants and Apicomplexa3,10, which includes plastid-containing parasites causing major human diseases such as malaria. How cell cycle progression is regulated without CDC25 in plants and apicomplexan parasites remains a major mystery in biology.
Plants have evolved a plant-specific family of CDKs, named CDKB, in addition to CDKA, the orthologue of fungal/animal Cdk1/Cdc2 (refs. 11,12,13,14). Genetic studies indicate that CDKA plays a dominant role in G1/S transition and CDKB1 is essential for mitotic entry12,15,16, in contrast to a single CDK acting in both transitions in yeast17. How checkpoint signalling regulates CDKA and CDKB1 activities and thereby transitions into S and M phases in plants is not fully understood18. The Thr14 and Tyr15 residues are conserved in CDKA and CDKB1, but their functions remain elusive. Altering Thr14/Tyr15 of Arabidopsis CDKA suggested that their phosphorylation inactivates CDKA but is not required for normal response to DNA replication stress19. The function of CDKB1 Thr14/Tyr15 phosphorylation has not been reported. Without CDC25 and functional evidence for the importance of CDK phosphorylation during replication stress, recent research has focused on transcriptional and protein–protein interaction mechanisms that modulate CDK activities in plants18.
With multiple technical advantages over multicellular higher plants, the unicellular green alga Chlamydomonas reinhardtii is a powerful model system for genetic studies of cell division20. Screening for temperature-sensitive mutants identified conserved eukaryotic cell cycle regulators and plant-specific genes required for mitosis in Chlamydomonas14,21. These include a mutation in the BSL1 phosphatase gene that blocks mitosis after DNA replication, a phenotype similar to the cyclin b (cycb) and cdkb mutants22. The BSL1 family of proteins is characterized by an amino-terminal Kelch repeat domain and a carboxy-terminal phosphatase domain, and is thus also named protein phosphatase with Kelch-like domain (PPKL). BSL1 is universally conserved throughout the green plant kingdom and apicomplexan parasites but is absent from fungi and metazoans23,24,25. Arabidopsis contains four BSL family members, BSU1 (bri1 Suppressor1) and BSL1, 2 and 3 (BSU1-like 1, 2 and 3), which have been studied extensively as components of receptor kinase pathways that regulate brassinosteroid promotion of growth26,27, immune signalling28 and stomatal development29,30,31. However, the Arabidopsis triple mutants lacking BSL1, 2 and 3 are embryonic lethal23, whereas Chlamydomonas contains a single BSL1 paralogue that is essential for mitosis14, suggesting that the BSL members share an ancestral and essential role in Chlamydomonas and higher plants. Here we investigate the function of BSL1 in cell cycle regulation in Chlamydomonas. Our results indicate that mitotic entry is arrested by the phosphorylation of CDKB1 Thr14/Tyr15 and promoted by BSL1-mediated dephosphorylation.
The BSL1 phosphatase is essential for mitosis in Chlamydomonas
To understand the function of BSL1 in mitosis, we characterized the cell cycle phenotypes resulting from various loss-of-function alleles of bsl1 mutants. The Chlamydomonas bsl1-1 mutant carries a missense L46P substitution in the N-terminal Kelch domain of BSL1 and is inviable at 33 °C, a temperature that does not affect the growth of wild-type cells. The phenotype of bsl1-1 was rescued by a full-length genomic sequence of Chlamydomonas BSL1 (Cre01.g050850) fused with a C-terminal mVenus–3xFLAG tag (BSL1–VF; Fig. 1a), but not by a variant protein with a substitution in a conserved phosphatase active site residue (BSL1(D619N)–VF)32 (Fig. 1b and Extended Data Fig. 1), indicating that the phosphatase activity is essential for BSL1 function. We disrupted the native BSL1 locus (bsl1-ko) using CRISPR–Cas9-guided genome editing in a strain that expresses the BSL1–VF protein from an unlinked locus (Extended Data Fig. 2). After crossing to the wild type, meiotic progeny carrying only the bsl1-ko allele without the BSL1–VF rescue construct could not be recovered (>16 recombinant tetrads tested), consistent with an essential function for BSL1.
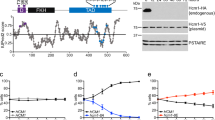
a, Diagram of the ProBSL1::BSL1-Venus::paroR construct showing the locations of the bsl1-1 mutation, the insertion in bsl1-ko and the D619N substitution. b, Anti-FLAG immunoblot (IB) showing BSL1–VF and BSL1(D619N)–VF in the bsl1-1 and bsl1-ko backgrounds. WT, wild type. c, Growth assay of the indicated strains. Cells grown at 21 °C were spotted in fivefold serial dilutions and then grown at 21 °C or 33 °C. Note the lethality of bsl1-1 and of bsl1-1::BSL1D619N-VF at 33 °C. d, Anti-FLAG and anti-BSL1 immunoblots showing the expression of BSL1, BSL1(L46P)–VF and BSL1–VF, and the absence of endogenous BSL1 in bsl1-ko. The experiment was performed once.
We tracked cell division in wild-type and bsl1-1 cells at restrictive temperatures via time-lapse microscopy. The cells were synchronized in G1 at 21 °C, incubated at 33 °C until the onset of mitosis and then imaged with a temperature-controlled microscope. We used a strain that expresses the microtubule end-binding protein (EB1) fused to the red fluorescent protein mScarlet (EB1–mSc), which marks cytoplasmic microtubules, the mitotic spindle and the basal bodies33,34. The cells also express a bleomycin resistance protein fused to GFP (ble–GFP), which was previously shown to localize in the nucleus, except for a brief period in mitosis coinciding with spindle formation and loss of nuclear integrity35.
The cell cycle in wild-type Chlamydomonas is characterized by a long G1 phase, during which the cells can grow to over eightfold their starting size without DNA synthesis or cell division20. An interphase cell maintains a pair of cilia anchored to the cell via two basal bodies. In wild-type cells grown at 33 °C, the onset of cell division was marked by the movement of the nucleus36 towards the basal bodies at the anterior end of the cell (Fig. 2a and Supplementary Video 1). In our videos, this nuclear movement was rapidly followed by the splitting of the two basal bodies, which moved to opposite sides of the nucleus near the future spindle poles. The ble–GFP signal disappeared from the nucleus during metaphase, indicating the breakdown of the nuclear envelope integrity. Subsequently, the cell performed nuclear division, cleavage furrow formation and cytokinesis.
a–c, Time-lapse images of a wild-type cell (a, representing 14 cells) and two bsl1-1 cells (b,c, representing 44 and 12 of 56 cells, respectively) grown at 33 °C. In each panel, the upper row shows an overlay of ble–GFP (yellow) and EB1–mSc (magenta), and the lower row shows the corresponding bright-field images. The time (minutes) of each frame is relative to the moment the nucleus contacts basal body (t = 0). The white arrow indicates the anterior notch formed after nuclear movement, the white arrowheads indicate basal body separation and the red arrowheads indicate cleavage furrows. Scale bars, 5 μm. d, Tetrads from a cross of bsl1-ko BSL1-VF × wild type were grown for 0, 24, 38 and 68 h at 22 °C. The genotypes are indicated. The bsl1-ko genotype is inferred. The bsl1-ko cell performs three to four divisions before lysing at the 8–16-cell stage (between 38 and 68 h, indicated by an asterisk; n = 8 tetrads). Scale bar, 40 µm. e,f, Time-lapse microscopy showing ble–GFP (yellow, nucleus) in meiotic progeny from a cross (ble-GFP EB1-mSc × bsl1-ko BSL1-VF) at the indicated hours after the start of imaging. Scale bars, 5 µm. In wild-type cells (e), each nuclear division results in two uniform-sized nuclei (n = 6 cells), whereas bsl1-ko cells (f) exhibit nuclear division failure (white arrow) and nuclear fragmentation (white arrowheads) (n = 5 cells).
The bsl1-1 mutant grown at 33 °C showed normal initial movement of the nucleus towards the basal body region (Fig. 2b,c and Supplementary Video 2). However, after nuclear migration, we observed no spindle formation or disappearance of ble–GFP from the nucleus. The bsl1-1 cells did form cleavage furrows associated with EB1–mSc after a variable delay (Fig. 2b,c), suggesting that cleavage furrow formation does not require BSL1 or that the residual BSL1 activity in bsl1-1 at 33 °C is sufficient for cleavage furrow formation. In most bsl1-1 cells, the undivided nucleus was found on one side of the ingressing furrow (Fig. 2b and Supplementary Video 2). In about 20% (12/56) of the cells, the furrow bisected the nucleus, generating nuclear fragments of varying size (Fig. 2c and Supplementary Video 3). In cells grown at 30 °C, spindles formed but were randomly oriented, the duration of metaphase was significantly prolonged and the centrin–mCherry (mCh) marker, which always localizes near the spindle poles in the wild type, was clearly mislocalized away from the spindle poles in bsl1-1 (Extended Data Fig. 3a). At the permissive temperature (21 °C), most bsl1-1 cells behaved like the wild type, but we occasionally observed misoriented spindles and slightly prolonged metaphase duration (Extended Data Fig. 3c,d). These results suggest that BSL1 is essential for spindle formation and that sufficient BSL1 activity is required for the precise organization of the spindle apparatus.
Cells carrying the bsl1-ko allele are inviable. To analyse the bsl1-ko mutant, we first crossed the complemented strain bsl1-ko BSL-VF to the wild type and monitored the colony growth of meiotic progenies. From complete tetrads, where all genotypes could be assigned, we found that bsl1-ko progeny died at the 8–16-cell stage (Fig. 2d). We then crossed bsl1-ko BSL1-VF to a wild-type strain expressing the markers ble–GFP and EB1–mSc and imaged meiotic progeny immediately after tetrad dissection via time-lapse microscopy (Supplementary Video 4). Under these conditions, the wild type remained viable for the duration of the video (48 h) and performed up to four rounds of cell division. Using ble–GFP as a nuclear marker, we observed tight coordination between nuclear division and cell division in the wild type, where each division resulted in two equal-sized daughter nuclei (Fig. 2e and Supplementary Video 5). In bsl1-ko cells, the first two nuclear divisions were indistinguishable from the wild type, probably because of parentally loaded BSL1 protein or mRNA. When bsl1-ko cells reached the eight-cell stage, we observed aberrant nuclear morphology and failure in nuclear division that resulted in fragmented ble–GFP signal (Fig. 2f and Supplementary Videos 4 and 6). Thus, the two loss-of-function mutants, bsl1-ko and bsl1-1, both display defective nuclear division.
BSL1 localizes to segregating basal bodies and mitotic spindle poles
To understand the dynamics of BSL1 during cell division, we imaged BSL1 (BSL1–VF) together with fluorescently tagged EB1 (EB1–mSc) (Fig. 3a and Supplementary Video 7). Interphase cells contained BSL1–VF near the basal body region, with weak signals around the nuclear periphery. When the cells entered prophase, the BSL1–VF signal increased dramatically near the segregating basal bodies and peaked near the spindle poles in metaphase when the spindle was fully formed. In cytokinesis, BSL1–VF relocalized to a band along the cleavage furrow (Fig. 3a). In daughter cells, BSL1–VF returned to its interphase configuration, with accumulation near the basal body region and the nucleus (Supplementary Video 8).
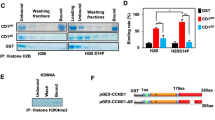
a, Confocal images of BSL1–VF and EB1–mSc at various cell cycle stages (minutes relative to the nucleus (n) contacting the basal bodies (BB)). EB1–mSc labels basal bodies (prophase), spindle microtubules (metaphase) and the cleavage furrow (CF) (cytokinesis). BSL1 colocalizes with EB1 in prophase and localizes strongly near the spindle poles (SP) in metaphase and weakly to the cleavage furrow. Similar observations were made in >100 cells. b,c, Confocal images of BLS1–mCh (yellow), BLD12–V (cyan, white arrowhead) and chloroplasts (magenta) in cells at interphase (9/14 cells) (b) and metaphase (13/13 cells) (c). Scale bars, 5 μm.
In Chlamydomonas, the basal bodies remain attached to the cell surface during mitosis, while the mitotic spindle assembles inside the nucleus37. To determine the spatial relationship of BSL1 to the interphase and mitotic basal bodies, we generated a fusion between the core basal body protein BLD12 and Venus (BLD12–V). In interphase, we observed a specific signal from BLD12–V near the flagellar base in non-dividing cells (Fig. 3b), consistent with the incorporation of the fusion protein into the centriole cartwheel structure38. In these cells, BSL1–mCh formed two filaments bridging the space between the basal bodies and the nucleus. In metaphase cells, the BLD12–V signal split into two dots, which remained close to the cell wall (Fig. 3c), and BSL1–mCh extended from the membrane-localized basal bodies towards the presumed spindle poles (Fig. 3c). The dynamic localization of BSL1 throughout the cell cycle, particularly the strong accumulation near the spindle poles, suggests that BSL1 plays a direct role in mitosis.
The bsl1-1 mutant is sensitive to a reduction in CDKB1 activity
Chlamydomonas contains a single CDKB gene (CDKB1), and one CYCB gene (CYCB1) that is probably the essential activator of CDKB1 activity13. The inactivation of CYCB1 or CDKB1 leads to a similar mitotic arrest phenotype as in the bsl1-1 mutant, with replicated DNA but no spindle formation and no nuclear division12,22. If BSL1 and CDKB1/CYCB1 act in a common pathway in mitosis, simultaneous partial loss of both genes may lead to a synthetic phenotype. We tested this possibility by crossing bsl1-1 to the temperature-sensitive cycb1-5 and cdkb1-1 mutants, which are not viable at a restrictive temperature (33 °C)12, and analysed the viability of the double mutant at semi-restrictive temperatures (Fig. 4). To analyse the bsl1-1 cycb1-5 and bsl1-1 cdkb1-1 double mutants, we dissected tetrads at a permissive temperature (21 °C), where all single mutants are viable without strong growth defects. After dissection, one set of tetrads was shifted to 26 °C, and the other set was maintained at 21 °C. One colony from each tetrad was unable to grow at 26 °C, suggesting that the double mutant lost viability while the single mutants were viable at the semi-restrictive temperature (Fig. 4a). The clones of a tetrad from a bsl1-1 × cdkb1-1 cross were genotyped and tested at a lower temperature of 24 °C, and the growth was scored daily (Fig. 4b,c). At 21 °C and 24 °C, both single mutants were fully viable, although bsl1-1 grew more slowly than the wild type at 21 °C. The bsl1-1 cdkb1-1 double mutant was indistinguishable from the bsl1-1 single mutant at 21 °C but showed a strong synthetic phenotype with reduced growth and viability at 24 °C, indicating a requirement for wild-type CDKB1 to maintain viability in the bsl1-1 mutant. These genetic interactions are indicative of synthetic lethality, which suggests that BSL1 and CYCB1/CDKB1 act in concert to promote mitosis.
a, Tetrads from a bsl1-1 × cycb1-5 cross were dissected at 21 °C and restreaked to duplicate plates at 21 °C and 26 °C. The bsl1-1 cycb1-5 double mutants (magenta boxes) show a synthetic lethal phenotype at 26 °C. b, Representative images of tetrads from a bsl1-1 × cdkb1-1 cross grown at 21 °C and 24 °C for four days. Scale bars, 100 µm. c, Colony sizes of the indicated strains dissected from bsl1-1 × cdkb1-1 crosses and grown at 21 °C (n = 4 tetrads) or 24 °C (n = 8 tetrads). The error bars indicate s.e.m.
BSL1 is phosphorylated in dividing cells
To test whether BSL1 is differentially phosphorylated in G1 and S/M phases, we performed anti-FLAG immunoprecipitation from synchronized BSL1-VF cdc20-1 cells in mid-G1 (0% M phase cells) and during cdc20-induced arrest (~30% M phase cells), with cdc20-1 as a control, and analysed the precipitants using liquid chromatography–tandem mass spectrometry (LC–MS/MS). The peptide coverage of BSL1 was similar in G1 (~85%) and S/M (~90%) cells. We identified 16 phosphorylated serine or threonine residues in BSL1 immunoprecipitated from the S/M-enriched samples (Extended Data Fig. 4a). The phosphosites were clustered either near the N-terminal Kelch domain or in the middle region between the Kelch and the phosphatase domains. Of these sites, five were also detected in G1 samples, indicating increased BSL1 phosphorylation as cells enter mitosis. Eight of the S/M-specific phosphosites are conserved in Arabidopsis BSLs; four residues are also present in apicomplexan homologues (Extended Data Fig. 4b). Overall, 8 of the 16 phosphosites contained a Ser/Thr followed by a proline, which matches the minimal consensus motif of proline-directed kinases, including CDKs.
BSL1 dephosphorylates CDKB1 pThr14/pTyr15
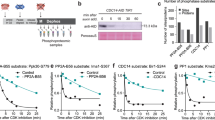
To identify BSL1-dependent dephosphorylation events, we performed a global quantitative phosphoproteomic analysis in cell-cycle-synchronized wild-type and bsl1-1 cells at four time points corresponding to newborn cells (t = 0, 21 °C), mid-G1 (t = 6 h, 33 °C), early-S/M (t = 8 h, 33 °C) and mid-S/M (t = 10 h, 33 °C) in the cell cycle. Microscopic analyses indicated that the bsl1-1 cells are delayed in cell division events (Extended Data Fig. 5a,b). Mass spectrometry analysis of flow-through peptide fractions showed an accumulation of S phase (for example, MCM3, RIR1 and CDKA) and mitotic (CYCB1 and CDKB1) proteins at the 8 h and 10 h time points (Extended Data Fig. 5c), consistent with the accumulation of cells with S/M phase morphology at these times. Interestingly, the accumulation of CYCB1 and CDKB1 levels was delayed in bsl1-1. Mass spectrometry analysis of the phosphopeptide fractions indicated that hundreds of proteins showed increased or decreased phosphorylation in bsl1-1. These include CDKB1, which was hyperphosphorylated at Thr14 and Tyr15 (IGEGpTpYGK) in bsl1-1 at the 8 h and 10 h time points (Extended Data Fig. 6a,b), in contrast to the decreased CDKB1 protein level at 8 h (Extended Data Fig. 5c). We further quantified the intensity ratio of IGEGpTpYGK between the bsl1-1 mutant and the wild type via parallel reaction monitoring mass spectrometry39 (Fig. 5a and Extended Data Fig. 6c–f). The intensity of the doubly phosphorylated peptide (IGEGpTpYGK) was 30 to 60 times higher in bsl1-1 than in the wild type (Fig. 5b and Extended Data Fig. 7a,b). The same peptide phosphorylated only on Tyr15 was moderately increased (about twofold) in bsl1-1 at the 10 h time point (Fig. 5b). In contrast, the abundance of phosphopeptides of CYCB1, PEPC2 and CDPKK was similar between bsl1-1 and the wild type (Fig. 5b and Extended Data Fig. 7c–e). These results indicate that BSL1 is required for efficient dephosphorylation of CDKB1 at Thr14 and Tyr15.
a, Quantification of CDKB1 phosphorylation in bsl1-1 and wild-type cells after 10 h at 33 °C. The bsl1-1 cells were labelled with heavy nitrogen (15N) and mixed with wild-type cells grown in 14N media before LC–MS/MS analysis. Each trace represents the intensity of a fragment ion of the IGEGpTpYGKVYK+2 peptide across the elution profile (dashed vertical lines) of the peptide. b, Quantification of the intensity ratio log2(bsl1-1/WT) of the doubly (pThr14 and pTyr15) and singly (pTyr15) phosphorylated CDKB1 peptides, and the control peptide from the PEPC2 protein, at the 8 h and 10 h time points. Each time point contains two repeats with isotope labels inversed. c, Parallel cultures of synchronized CDKB1–mCh cells. At 10 h, one culture was treated with 8 mM HU. Total protein extracts were analysed via anti-mCh immunoblots of phos-tag and SDS–PAGE gels. +P, phosphorylated CDKB1–mCh. Similar results were observed in two biological experiments. d, Wild-type CDKB1 and unphosphorylatable CDKB1(AF) were immunoprecipitated from HU-treated cells. CDKB1–mCh was treated with recombinant phosphatase domains from Chlamydomonas BSL1 (CrBSL1), Toxoplasma PPKL (TgPPKL) or Lambda phosphatase (λ). Negative controls were untreated (−) or treated with the non-phosphatase protein SPY. Dephosphorylation was measured as the intensity ratio of phosphorylated to unphosphorylated forms (+P/CDKB). The experiment was repeated once with similar results. e, Bio-layer interferometry showing a direct interaction between recombinant GST–CDKB1 and BSL1 proteins. GST–CDKB1 protein was loaded onto biosensors, dipped into the indicated concentrations of purified BSL1 protein for association (time 0) and transferred into buffer (red dashed line) for dissociation. The dissociation constant (Kd) was calculated via the ratio of the off and on rates (Koff/Kon). f, Dual-colour imaging of CDKB1–mCh and BSL1–Venus in a cell with two metaphase spindles. CDKB1–mCh localizes to the spindle pole (magenta arrow; similar results observed in three cells). Chlorophyll autofluorescence (magenta asterisk) is captured in the mCh channel. Scale bar, 5 µm. g, Normalized fluorescence intensity along a line bisecting both spindle poles (white dashed arrow in f).
In yeast and animals, the phosphorylation of CDK on Thr14/Tyr15 is induced by DNA replication stress40. Hydroxyurea (HU), a ribonucleotide reductase inhibitor, induces replication stress in plants and animals and was previously shown to block DNA replication in Chlamydomonas41. We found that HU increased the proportion of phosphorylated CDKB1 with little effect on the CDKB1 protein level as cells entered the S/M phase (Fig. 5c). Phosphorylation of CDKB1 was lost in cells expressing a version of CDKB1 that carries unphosphorylatable T14A and Y15F substitutions (CDKB1(AF)–mCh; Fig. 5d). We performed quantitative proximity labelling with BSL1 fused to a version of BioID (BSL1–BioIDG3) that was engineered to function in the Chlamydomonas cytoplasm (Methods). The result showed increased labelling of CYCB1 (Extended Data Fig. 8), which is known to form a complex with CDKB13. To test whether BSL1 can dephosphorylate CDKB1 in vitro, we immunoprecipitated CDKB1–mCh from HU-treated cells as a substrate for in vitro phosphatase assays. The results showed that CDKB1 was dephosphorylated effectively by BSL1 and Lambda phosphatase and less effectively by Toxoplasma PPKL, and was not dephosphorylated after incubation with buffer only or with SPY protein as negative controls (Fig. 5d). Bio-layer interferometry assays, using recombinant proteins purified from Escherichia coli, indicated that BSL1 interacts with CDKB1 in vitro with a Kd of 900 nM (Fig. 5e). Confocal microscopy showed that BSL1 and CDKB1 colocalize transiently near the spindle poles in metaphase (Fig. 5f,g), consistent with previous observations of CYCB1 localization to the spindle poles13. Together, the in vivo and in vitro evidence indicates that BSL1 dephosphorylates CDKB1 pThr14 and pTyr15.
CDKB1 phosphorylation is essential for the response to DNA replication stress
Phosphorylation at Thr14/Tyr15 is known to inhibit Cdk1/Cdc2 activity in yeast and animals42. To test the effect of CDKB1 phosphorylation in Chlamydomonas, we analysed phosphomimetic (Asp14/Tyr15, Thr14/Glu15 and Asp14/Glu15) and unphosphorylatable (Ala14/Phe15, CDKB1(AF)) variants fused to mCh. None of the phosphomimetic variants rescued the cdkb1-1 mutant at the restrictive temperature (Extended Data Fig. 9a), suggesting that the phosphorylation of CDKB1 blocks its function. The unphosphorylatable variant, CDKB1(AF)–mCh, complemented the cdkb1-1 mutant but supported slower growth in colony size than CDKB1–mCh at the restrictive temperature (Extended Data Fig. 9b), indicating that CDKB1 phosphorylation is dispensable for viability but required for optimal growth. At permissive temperatures, we observed similar growth rates between the cdkb1-1 CDKB1-mCh and cdkb1-1 CDKB1AF-mCh strains, indicating that the functional CDKB1-1 is dominant over CDKB1(AF) (Extended Data Fig. 9c). Expression of CDKB1(AF) did not suppress bsl1-1 (Extended Data Fig. 9d), implying that BSL1 plays additional roles besides dephosphorylating CDKB1 pThr14/pTyr15. However, microscopic analysis indicated that CDKB1(AF) advanced cleavage furrow formation in bsl1-1 (Extended Data Fig. 9e), suggesting that the phosphorylation of CDKB1 contributes to delayed furrowing in bsl1-1 mutant cells. Similar to wild-type CDKB1 (Extended Data Fig. 5c), the accumulation of CDKB1(AF) was reduced in bsl1-1 cells (Extended Data Fig. 9f), suggesting that BSL1 promotes the stability of CDKB1 independently of dephosphorylation at Thr14/Tyr15. We expressed a phosphatase-dead mutant BSL1 in the CDKB1AF bsl1-1 strain and tested its growth under a restrictive temperature. The results show that BSL1 phosphatase activity is required even in the presence of unphosphorylated CDKB (Extended Data Fig. 9g),
Checkpoint-mediated phosphorylation of Cdk1 Thr14/Tyr15 delays the onset of mitosis until DNA replication is finished, when dephosphorylation by Cdc25 triggers exit from the delay in fission yeast40 and animals43. Since CDKB1 is the main driver of mitosis in Chlamydomonas, we tested the role of CDKB1 phosphorylation in response to DNA damage and replication stress. We found that cdkb1-1 mutant cells complemented with CDKB1(AF)–mCh were highly sensitive both to HU (Fig. 6a) and to the polymerase inhibitor aphidicholin, but not to the DNA-damage-inducing drug zeocin44 (Extended Data Fig. 10a). The bsl1-1 mutant cells also showed increased sensitivity to HU (Extended Data Fig. 10b), probably due to the additive effects of bsl1-1 and HU on CDKB1 phosphorylation; this is consistent with the synthetic phenotype of the bsl1-1 cdkb1-1 double mutant (Fig. 4), supporting the role of BSL1 in exit from HU-induced mitotic arrest. We imaged cdkb1-1 mutant cells complemented with CDKB1 or CDKB1(AF) via time-lapse microscopy in the presence of HU (Fig. 6b,c). After 12 h of incubation on control media (Tris Acetate Phosphate (TAP)) at 33 °C, cdkb1-1 mutant cells expressing wild-type CDKB1 or CDKB1(AF) formed microcolonies of 8–16 daughter cells, whereas the cdkb1-1 mutant cells did not divide after 12 h and lysed by 30 h (Fig. 6b). A low concentration (1 mM) of HU caused post-S/M lethality in CDKB1(AF) cells without an obvious impact on the wild type by 30 h, whereas a high concentration of HU (10 mM) delayed the appearance of cleavage furrows and caused lethality in all strains by 30 h (Fig. 6b,c). These results show that HU delays cell cycle progression in the wild type and this delay was largely bypassed in CDKB1(AF) cells, indicating that the phosphorylation of CDKB1 Thr14/Tyr15 delays mitotic entry and cytokinesis under replication stress to prevent cell division with incompletely replicated DNA. Taken together, these results demonstrate that replication stress increases CDKB1 Thr14/Tyr15 phosphorylation, which inactivates CDKB1 and delays mitotic entry and cytokinesis; BSL1-mediated dephosphorylation activates CDKB1 and promotes mitotic entry in Chlamydomonas.
a, Growth and survival of the wild type, cdkb1-1 and cdkb1-1 complemented with CDKB1–mCh and CDKB1(AF)–mCh. Cells were spotted (fivefold dilution) on TAP and TAP + 1 mM HU, and grown at 21 °C and 33 °C. b,c, Microscopic analysis of cell cycle dynamics of individual cells. Panel b shows selected time-lapse images of representative cells taken at 12 h and 30 h on TAP and TAP + HU media at 33 °C. The black arrows point to the microcolonies containing 8–16 cells at 12 h. The cdkb1-1 mutant arrests with a ‘notch’ morphology (white arrowhead), indicating mitotic arrest. The black arrowhead points to representative cells that have undergone complete lysis. Scale bar, 5 μm. Panel c shows the quantification of cleavage furrow formation in TAP and 10 mM HU. n indicates the number of cells analysed. P indicates the result of Fisher’s exact test for each time point. NS, not significant. The experiment was repeated twice with similar results.
Discussion
Control of mitotic entry is crucial both for maintaining genome integrity and for endoreduplication in development. The mechanism of mitotic entry is well understood in opisthokonts, which include fungi and metazoans. The core mechanism involves keeping CDK inactive through the phosphorylation of the Thr14 and Tyr15 residues before DNA replication is complete and DNA damage is repaired, and rapidly activating CDK through dephosphorylation by CDC25. However, the mechanism controlling mitotic entry is not understood in other clades of eukaryotes2,45. In particular, the absence of CDC25 has created a mystery about how cell cycle progression is regulated in plants and Apicomplexa. Our study resolves this mystery by identifying BSL1 as a phosphatase that dephosphorylates and activates CDKB1, the main kinase driving mitosis in Chlamydomonas. We show that the inactivation of CDK by the phosphorylation of Thr14 and Tyr15 is conserved, but different phosphatases have evolved to activate CDK in opisthokonts and the green lineage of eukaryotes (Fig. 7). Furthermore, unlike CDC25 and Cdk1 in mammals, which act on both G1/S and G2/M transitions46,47,48, the plant-specific BSL1–CDKB1 pathway appears to be dedicated specifically to the control of mitosis.
In fungi and metazoans, mitosis is triggered by the activation of Cdk1, which is inactivated during replication stress by phosphorylation at Thr14/Tyr15 and subsequently activated by Cdc25-mediated dephosphorylation. Similarly, in Chlamydomonas, CDKB1 is essential for mitosis and is inactivated by phosphorylation at Thr14/Tyr15. However, the dephosphorylation of CDKB1 Thr14/Tyr15 is mediated by BSL1. In addition, BSL1 promotes CDKB1 accumulation independent of Thr14/Tyr15 dephosphorylation, possibly by inhibiting an unknown protein (X) that mediates CDKB degradation. +p/−p, phosphorylation/dephosphorylation. Triangular arrowheads indicate positive regulation, diamond-shaped arrowheads indicate inhibitory regulation and dashed lines indicate hypothetical mechanisms.
Our genetic, cell biological and biochemical evidence supports a model where CDKB1 and BSL1 act synergistically to drive mitosis. Cells carrying the temperature-sensitive alleles of bsl1-1 and cdkb1-1 mutants both arrest in mitosis after DNA replication and show synthetic lethal interactions at semi-restrictive conditions, suggesting that these alleles function in a common pathway. The loss of BSL1 increases the phosphorylation of CDKB1 Thr14 and Tyr15 and decreases the protein levels of CDKB1 and CYCB1, indicating that BSL1 is required for CDKB1 dephosphorylation and activation. Several lines of evidence show that BSL1 directly dephosphorylates CDKB1, including in vitro binding and dephosphorylation assays, colocalization of both proteins to the spindle poles during mitosis and in vivo association between BSL1 and CYCB1 detected via BSL1–TurboID proximity labelling.
The phosphorylation of Thr14 and Tyr15 in CDK-family kinases interferes with catalysis and is a key mechanism for controlling cell cycle progression in fungi and animals49. However, a functional role for CDK Thr14/Tyr15 phosphorylation has been elusive in the plant kingdom. Alterations of Thr14/Tyr15 of CDKA1 did not affect plant responses to replication stress in Arabidopsis19. This observation, along with the absence of CDC25, prompted the hypothesis that Thr14/Tyr15 phosphorylation is not critical for mitotic regulation in plants19. Here we demonstrate that the phosphorylation of Thr14/Tyr15 in CDKB1 is a key mechanism of mitotic regulation in Chlamydomonas. Inhibiting DNA replication via HU enhances the phosphorylation of CDKB1 Thr14/Tyr15, which apparently inhibits CDKB1 function, as CDKB1 containing substitutions of Thr14 and Tyr15 with phosphomimetic residues was unable to rescue the cdkb1-1 mutant. CDKB1 is essential for mitosis and cytokinesis12. It was reported previously that HU blocks mitosis but not cytokinesis in Chlamydomonas41. However, our microscopic analysis showed that HU delays cytokinesis. Consistent with HU-induced phosphorylation inhibiting CDKB1, cells expressing non-phosphorylatable CDKB1(AF) showed a reduced delay of cytokinesis and consequently reduced survival when treated with DNA replication inhibitors. In the absence of replication inhibitors, these cells are viable, but their colonies grow more slowly than the wild type, probably because some cells are lost due to premature mitosis/cytokinesis before completing DNA replication. DNA synthesis could be slowed by nutrient limitation near the end of each cell division cycle. These results indicate that replication stress causes CDKB1 Thr14/Tyr15 phosphorylation, which inhibits CDKB1 function and delays mitosis and cytokinesis till DNA replication is complete.
In fungi and animals, CDK and CDC25 form a positive feedback loop where they activate each other, and the active CDK then promotes mitotic entry. In fission yeast, the only essential function of CDC25 is to dephosphorylate CDK Thr14/Tyr15, as CDC25 is no longer essential when an unphosphorylatable Cdc2(Y15F) is present5. Our study shows that the role of CDC25 (that is, the dephosphorylation of pThr14/pTyr15 of mitotic CDK) is played by BSL1 in plants. However, the mitotic arrest of the bsl1-1 mutant is not suppressed by CDKB1(AF), suggesting that BSL1 has another essential function in addition to dephosphorylating CDKB1 pThr14/pTyr15. This additional function of BSL1 seems to be related to CDKB1 accumulation, as the bsl1-1 mutation not only increased CDKB1 Thr14Tyr15 phosphorylation but also reduced the accumulation of CDKB1, CYCB1 and CDKB(AF) proteins. BSL1 may stabilize CYCB1–CDKB1 through a phosphatase-independent mechanism, such as protein–protein interaction. However, expressing a phosphatase-dead mutant BSL1 in the bsl1-1 CDKB1AF background did not rescue viability, suggesting that BSL1’s phosphatase activity is essential even when CDKB1 is unphosphorylated at Thr14/Tyr15. Another possibility is that BSL1 dephosphorylates a protein involved in regulating CDKB1 and CYCB1 protein levels. Such regulation of both CDKB1 activity and accumulation would provide tighter control of CDKB1 function and mitotic entry.
BSL1 may have additional functions beyond regulating CDKB1. The dynamic subcellular localization of BSL1 suggests its multiple actions at different subcellular locations and cell cycle phases. The various defects in spindle organization and orientation in bsl1-1 cells grown at semi-restrictive temperatures also suggest that BSL1 is required for both entry and proper execution of mitotic progression. Phosphoproteomic experiments showed that bsl1-1 increased the phosphorylation of many proteins. Our genetic, microscopic and proteomic studies thus suggest that BSL1 has broad functions in addition to dephosphorylating CDKB1 Thr14/Tyr15. Future studies of additional BSL1 substrates will elucidate additional signalling pathways and mechanisms that regulate cell division in Chlamydomonas, including the mechanism of regulating CDKB1 and CYCB1 accumulation.
BSL1 and CDKB1 appear to play different roles in cleavage furrow formation (cytokinesis), which is abolished in the cdkb1-1 mutant but only delayed in the bsl1-1 mutant. As noted in previous studies using HU to inhibit DNA replication41 or with cell-cycle-blocked mutants20, cleavage furrow initiation does not depend on the completion of DNA replication or mitosis in Chlamydomonas. Our analyses show that HU delays cleavage furrow formation in wild-type cells, but the delay is much reduced in cells expressing the CDKB1(AF) protein, consistent with HU causing CDKB1 Thr14/Tyr15 phosphorylation and inactivating CDKB1. Similar to HU-treated cells, the bsl1-1 mutant cells show a delayed cleavage furrow, which is partially suppressed by CDKB1(AF). Together, our results support the model that DNA replication stress causes the phosphorylation of CDKB1 Thr14/Tyr15 to prevent premature mitotic entry and cytokinesis before DNA is fully replicated; the BSL1-mediated dephosphorylation and stabilization of CDKB1 trigger progression through mitosis and cytokinesis.
Interestingly, CDKB1(AF) cells showed no increased sensitivity to zeocin, which induces double-stranded DNA breaks, possibly due to distinct response pathways for different types of DNA stress. In Chlamydomonas, BSL1 and CDKB1 are required only for mitotic entry, whereas CDKA1 plays a dominant role in the G1/S transition12. Such distinct functions of CDKA and CDKB1 could enable different checkpoints to specifically control distinct cell cycle transitions. Consistent with this notion, the phosphorylation of CDKA1 Thr14/Tyr15 was shown to inactivate CDKA but play no obvious role in replication stress in Arabidopsis19. These observations raise the possibility of a CDKB1-independent pathway for the DNA damage response and a CDKB1-dependent replication stress pathway in plants.
In animals, replication checkpoint signalling leads to CDK phosphorylation by the Wee1 kinase to safeguard DNA replication and mitotic entry50. WEE1 is conserved in plants. The Chlamydomonas WEE1 is expressed in S/M cells, but its function has not been reported51. The Arabidopsis thaliana WEE1 (AtWEE1) is activated by replication stress, and the loss-of-function mutants are hypersensitive to the replication-inhibitory drug HU in growth but insensitive in mitotic arrest, indicating that AtWEE1 plays a role in replication stress52. Intriguingly, AtWEE1 was shown to mediate HU-induced phosphorylation of CDKA but not CDKB52. However, the alteration of CDKA Thr14 and Tyr15 to non-phosphorylatable residues did not affect the plant’s sensitivity to replication stress19, whereas genetic evidence supports a key role for CDKBs in mitotic entry in algae and Arabidopsis12,16. Our study suggests that the possible WEE1–CDKB1 connection should be examined in Chlamydomonas and Arabidopsis in future studies (Fig. 7). AtWEE1 was recently reported to phosphorylate the E3 ubiquitin ligase FBL17, leading to the accumulation of CDK inhibitors and the inhibition of cell cycle progression53; how this inhibitory pathway is reversed to mediate checkpoint exit and mitotic entry remains unknown. Future studies are required to illustrate the full replication checkpoint pathways in algae and higher plants.
The regulation of mitotic entry is not only crucial for genome integrity but also important for development in multicellular organisms16,54. BSL1 and its homologues have evolved additional roles, particularly in receptor kinase signalling in land plants such as Arabidopsis. Members of the BSL family were initially identified in Arabidopsis as components of the brassinosteroid signalling pathway26, acting downstream of the BRI1 receptor kinase to inactivate the GSK3-like kinase BIN2 by dephosphorylating its conserved tyrosine residue27. Recent studies have revealed additional roles of BSLs in receptor kinase pathways that regulate immunity and stomata development, acting upstream of mitogen-activated protein kinase modules, with signalling specificity enabled by phosphocoding and scaffolding28,29,30. In Arabidopsis, CDKB1 is known to play a role in mitotic entry and endoreduplication16. An intriguing possibility to be tested in the future is that receptor-kinase-mediated signalling may regulate endoreduplication through a BSL–CDKB1 connection in certain land plants. Our study in Chlamydomonas suggests that BSL1 is a cell cycle regulator that has evolved additional signalling functions during the evolution of land plants. It is conceivable that BSLs act as a signalling hub that integrates diverse developmental and environmental signals to regulate cell division, differentiation and endoreduplication in plants.
Our findings raise interesting questions about the evolution of BSL/PPKL phosphatases. The BSL/PPKL family is conserved in plants and Apicomplexa but absent in fungi and metazoans3, and thus their presence coincides with the absence of CDC25 in evolution. BSL/PPKL is essential for viability in Toxoplasma gondii and is thus considered an ideal target for anti-parasite drug development25. Our in vitro phosphatase assay shows that TgBSL/PPKL can dephosphorylate Chlamydomonas CDKB1. The function of BSL/PPKL in mitosis seems ancestral and conserved, and such conservation of a cell cycle regulator would add a key piece of supporting evidence for the algal origin of Apicomplexa. Future studies of BSL/PPKL hold great potential for improving agricultural productivity and managing human diseases caused by apicomplexan parasites.
Methods
Strains and culture conditions
The wild-type Chlamydomonas strains used are isogenic with CC-4402 (mt+) and CC-4403 (mt−) (Supplementary Table 1). The bsl1-1 mutant was isolated after UV mutagenesis14 and backcrossed eight times to CC-4402/4403 before analysis. The cycb1-5 and cdkb1-1 were described previously13. Chlamydomonas crosses were performed according to standard methods55, and meiotic products were dissected using a Zeiss Axioscope A1 equipped with a microdissection needle. Strains were propagated on TAP media at 21 °C with constant illumination (~20 µmol m−2 s−1). For selection, paromomycin (Sigma P9297) and hygromycin B (EMD Millipore 400052) were used at 10 µg ml−1 in TAP agar. For metabolic labelling, strains were passaged twice on TAP media containing heavy 15NH4Cl (Cambridge Isotopes, NLM4675) instead of regular 14NH4Cl before being used for experiments.
Asynchronous populations of cells in liquid culture were synchronized in G1 by washing the cells once in water and once in Tris Phosphate (TP) media, followed by incubation in TP media for 12 h at 21 °C in the dark. For time-course experiments, one of two protocols was used. In the plate protocol, ~2.5 × 107 G1 cells were spread on a 12 cm × 12 cm TAP agar plate and incubated at 33 °C under 50 µmol m−2 s−1. In the liquid culture protocol, TAP medium was inoculated to ~OD750 0.1 and grown at 33 °C under 120 µmol m−2 s−1. Cell division was scored visually by the appearance of cleavage furrows via bright-field microscopy using a ×40 objective. Wild-type cells typically initiated division at 8–10 h in the plate protocol and at 11–13 h in the liquid protocol.
Plasmid construction
The BSL1 gene (Cre01.g050850) was amplified from CC-4403 genomic DNA using bsl-1F/R and bsl-2F/R primers and introduced into pCR-Blunt II-TOPO (Thermo 450245) to generate pFT1. For the oligonucleotides, see Supplementary Table 2. A 2-kb promoter fragment was amplified from CC-4403 genomic DNA with bsl2kb-F and bsl2kb-R and introduced in pCR-Blunt II-TOPO to generate pFT2. The plasmid pFT4, for the expression of BSL1 in Chlamydomonas from the synthetic H/R promoter (ProH/R::gBSL1-Venus-APHVIII), was generated by amplifying CrBSL1 from pFT1 with bsl-cp4F and bsl-cp4R and introducing it into pMO449 (ref. 56) via Gibson assembly (New England Biolabs, E2611S). The native promoter version pFT40 (ProBSL1::BSL1-mVenus-3xFLAG) was generated by inserting the BSL1 promoter from pFT2 into PfoI+HpaI-digested pFT4 via Gibson assembly. The phosphatase-dead version BSL1(D619N) was generated via circular mismatch PCR from pFT40. The CDKB1–mCh (pKA1) and CDKB1(AF)–mCh (pKA17) plasmids were provided by F. Cross (Rockefeller University). The phosphomimic CDKB1 variants (CDKB1(DY/TE/DE)–mCh) were generated via mismatch PCR from pKA1. The BioIDG3 sequence corresponds to the G3 generation in the directed evolution of TurboID from BioID (see Supplementary Table 1 in Branon et al.57), including the substitutions Q65P, I87V, R118S, E140K, Q141R, A146Δ, S150G, L151P, V160A, T192A, M209V and I305V. The BioIDG3 sequence was codon-optimized for Chlamydomonas and combined with pFT40 to generate the construct pFT57 (ProBSL1::BSL1-BioIDG3-NeonGreen) and the control construct pFT58 (ProBSL1::BioIDG3-NeonGreen).
Transformation
Electroporation was performed essentially as described previously58 with 1.2 µg of linearized plasmid per approximately 2 × 108 cells. A 10-µl plasmid digestion reaction was performed in FastDigest buffer (Thermo Scientific). The cells were spun down and washed twice in electroporation buffer (10 mM N-cyclohexyl-2-aminoethanesulfonic acid (pH 9.25), 40 mM sucrose, 10 mM sorbitol). Then, 125 µl of cell suspension was added to the digested plasmid, mixed by pipetting and added to a 2-mm electroporation cuvette (Bulldog Bio). Electroporation was performed using a NEPA21 instrument (Bulldog Bio) with the following settings: voltage (poring pulse 250 V, transfer pulse 20 V), length (poring 8 ms, transfer 50 ms), interval (poring 50 ms, transfer 50 ms), no. of pulses (poring 2, transfer 5), decay (poring 40%, transfer 40%) and polarity (poring +, transfer +/− alternating). Impedance was generally 0.33–0.37 Ohm. Electroporated cells were transferred to 8 ml TAP + 40 ml sucrose and incubated for ~20 h at 22 °C in dim light (<5 µmol m−2 sec−1), then spun down and plated on selective media. Transformants were selected after seven days. Transformation with EcoRV-digested pFT40 typically resulted in ~100 colonies after selection at 33 °C and ~1,000 colonies when selected at 21 °C on 10 µg ml−1 paromomycin.
CRISPR-guided knockout of CrBSL1
A culture of the complemented bsl1::BSL1-VF strain (50 ml, OD750 = 0.2) was spun down and resuspended in gametic autolysine enzyme (prepared from CC-4402/CC-4403 gametes) and incubated for 1 h at 22 °C. The loss of cell wall integrity was verified by treatment with 0.25% Triton X-100, which caused bursting of autolysine-treated cells. After autolysine treatment, the cells were heat-shocked for 30 min at 40 °C with gentle shaking. During heat-shock treatment, RNP complex was prepared. Single guide RNA (sgRNA) (100 pmol µl−1, Integrated DNA Technologies) and SpCas9 (61 pmol µl−1, Integrated DNA Technologies) were mixed in the following ratio: sgRNA 1.2 µl, SpCas9 1 µl, duplex buffer 2.8 µl (30 mM HEPES (pH 7.5), 100 mM KOAc). The mixture was incubated for 30 min at 37 °C and then kept on ice. 500 ng of APHVII PCR product (hygromycin resistance gene, amplified from plasmid pRAM103 (ref. 59) with MOP804 MOP626) was added to PCR tubes. Heat-shocked cells were spun down for 3 min at 2,000 rpm, washed twice in TAP + 40 mM sucrose and resuspended in ~500 µl TAP + 40 mM sucrose. Cell suspension (125 µl) was added to the APHVII PCR product, mixed by pipetting and then transferred to the sgRNA/Cas9 mixture. The mixture (125 µl) was used for electroporation with the above settings. Impedance in TAP + 40 mM sucrose was ~0.15 Ohm. Electroporated cells were transferred to 8 ml TAP + 40 ml sucrose and incubated for ~20 h at 22 °C in dim light (<5 µmol m−2 sec−1), then spun down and plated on TAP + 10 µg ml−1 hygromycin B. Transformants were selected after seven days, under 50 µmol m−2 sec−1 light. Eight of 12 hygromycin-selected transformants contained the APHVII gene inserted either in the native BSL1 locus or in the BSL1-VF transgene. No clone with simultaneous disruption of both loci was identified.
Time-lapse microscopy
For fluorescence time-lapse microscopy, cells were pre-grown in TAP media at 21 °C in constant illumination (20 µmol m−2 s−1) and synchronized in G1 using the TP-DARK method. Synchronized cells were grown in TAP liquid (~10–11 h) or TAP agar (8–9 h) before imaging. Temperature-sensitive strains were grown at 33–34 °C. Agar sandwiches were prepared by melting TAP-agar (1.5%), pipetting 275 µl onto an 18 mm × 18 mm coverslip (Avantor) and covering it with a second coverslip. When the agar had set, one coverslip was removed, and 2 µl of cell suspension was spotted on the agar surface. Mutant and wild-type control cells were spotted next to each other on the same agar slab. A 24 mm × 60 mm coverslip (Avantor, size 1.5) was placed on top of the cells, and the agar edges were sealed with grease to prevent drying of the agar during imaging. The 24 mm × 60 mm coverslip was placed on an inverted spinning disc confocal microscope (Olympus) equipped with a ×100 silicone-immersion objective (NA 1.35) and a heated stage insert (Tokai-Hit) set to hold 33–34 °C inside the chamber. mVenus and GFP fluorophores were excited with a 514-nm laser using a 440/514/561-nm beamsplitter and a 550/49-nm emission filter. mCh was excited with a 561-nm laser using a 440/514/561-nm beamsplitter and a 605/64-nm emission filter. For time-lapse imaging, images were collected every 3 min. This time interval ensured that every mitotic spindle would be captured, while minimizing phototoxicity to the cells.
For bright-field time-lapse microscopy, cells were synchronized as above and spotted on TAP-agar plates. For drug treatments, HU (Sigma-Aldrich, H8627), aphidicholin (EMD Millipore, 178273) or zeocin (InvivoGen, ant-zn-1) was added to TAP agar. The cells were incubated at 33 °C under ~80 µmol photons m−2 s−1 and imaged with a ×10 objective.
Statistics and reproducibility
Statistical tests were performed using R (v.4.5.0)60 and plotted with the ggplot2 package61. All sample mean comparisons were two-sided. Analysis of variance results (R function lm/aov) are presented as d.f. = df1,df2, F = F value and P = Pr(>F), where df1 and df2 are the within- and between-groups degrees of freedom. Tukey’s post-hoc tests (R function TukeyHSD) were performed if analysis of variance resulted in P < 0.05. The box plots show the median and 25th/75th percentiles, with whiskers at maximum 1.5 times the interquartile range. Significance groups from Tukey’s HSD post-hoc test are represented as letters above each box plot. t-tests were performed with the R function t.test. Pairwise two-sided tests for independence of rows (wild type versus CDKB(AF)) and columns (TAP versus HU) of cells with cleavage furrows in Fig. 6 were performed with the R function fisher.test. P < 0.05 indicates significant differences. The experiments were performed at least twice, except where specified.
Immunoprecipitation
Cells were collected via centrifugation, transferred to 50-ml tubes and washed once in immunoprecipitation (IP) buffer (40 mM HEPES (pH 7), 100 mM KCl, 100 sorbitol). The wet pellet was weighed, and an equal amount (w/v) of IP buffer was added. The resulting slurry was added dropwise to liquid nitrogen to make frozen cell pellets that were stored at −70 °C. The pellets were cryomilled in a Retsch cell grinder for 3 min at 30 Hz. Approximately 400 mg of milled powder per IP was thawed for 2 min at room temperature, then transferred to ice, and 1.5 ml of ice-cold IP buffer with protease inhibitors (Sigma no. A32695) and phosphatase inhibitor (Pierce no. 32957, 1 tablet per 5 ml) was added. The suspension was homogenized by pipetting and clarified via centrifugation for 10 min at 10,000 g and 4 °C. The supernatant was added to 100 µl of anti-FLAG magnetic beads (Sigma no. M8823), and the samples were incubated for 1 h at 4 °C with end-over rotation. The beads were washed four times with IP buffer. Bound proteins were eluted in SDS–PAGE loading buffer (Laemmli), without DTT (1% SDS, 10% glycerol, 62.5 mM Tris-HCl (pH 6.8)), for 10 min at 65 °C. A second elution was done in 100 µl of 2× Laemmli (2% SDS, 50 mM DTT) for 10 min at 65 °C. Generally, >90% of BSL1–FLAG eluted in the first elution. Sampled were separated on 7.5% Tris-glycine SDS–PAGE gels for in-gel digestion and mass spectrometry.
Immunoblot
Immunoblot analysis was performed according to standard methods, using monoclonal mouse anti-FLAG antibody (Sigma F1804, clone M2) and polyclonal rabbit anti-mCh (Abcam, ab167453) primary antibodies. The anti-BSL1 serum was raised against exon13 of CrBSL1 (amino acids 770–885 in Cre01.g050850.t2.1 Phytozome v.5.6) fused at its N terminus to maltose-binding protein. Total extracts for immunoblot analysis were prepared by mixing 30 µl of 2.5× Laemmli buffer (2.5% SDS (w/v), 125 mM Tris-HCl (pH 6.8), 12.5% glycerol, 0.0125% bromophenol blue), supplemented with 50 mM DTT, with 10 mg of wet cell pellet and heating for 10 min at 65 °C. The extracts were clarified via centrifugation, and approximately 7.5 µl was loaded per sample. Zn2+ phos-tag gel analysis was performed according to the method described by Kumar62.
Phosphatase assay
The phosphatase domains of Chlamydomonas BSL1 (amino acids 507–885 in Cre01.g050850.t2.1) and Toxoplasma PPKL (amino acids 532–934 in UniProt S7UP60) were fused at the N terminus to EcoRI/NotI-digested pSUMO DUet vector to create N-terminal 6xHis and SUMO tags. 6xHis–SUMO–SPY from Arabidopsis (AT3G11540) was included as a negative control. Proteins were produced in E. coli BL21 cells and partially purified via nickel column chromatography. For in vitro phosphatase assays, cells expressing CDKB1–mCh were synchronized as above and incubated for 13 h at 31 °C under 180 µmol m−2 s−1 white light. HU (8 mM) was added after 10 h. Cells were collected via centrifugation, and the pellet was resuspended in two volumes of ice-cold TBS (17.5 mM Tris-HCl (pH 7.6), 137 mM NaCl), added dropwise into liquid nitrogen and milled to a fine powder. To precipitate CDKB1–mCh, 150 mg of powder was resuspended in 400 µl of TBS with cOmplete protease inhibitors (Roche 04693132001, 1 tablet per 50 ml) and phosphatase inhibitors (Pierce no. 32957, 1 tablet per 5 ml), and 15 µl of RFP-Trap beads (Chromotek, RTMA-20) were added. After 1 h of incubation, rotating at 4 °C, the beads were washed five times with 1 ml of TBS and twice with phosphatase reaction buffer (50 mM Tris-HCl (pH 6), 5 mM MnSO4), then aliquoted into tubes. 1 µg of purified protein (BSL1-PP, PPKL-PP or SPY) or 200 U of Lambda phosphatase (Santa Cruz, sc-200312A) was added to CDKB1–mCh bound to RFP-Trap beads in a 20 µl volume and incubated for 3 h at room temperature. The beads were washed five times with 1 ml of TBS and heated for 10 min at 65 °C in 30 µl of 2.5× Laemmli buffer (2.5% SDS (w/v), 125 mM Tris-HCl (pH 6.8), 12.5% glycerol, 0.0125% bromophenol blue). Approximately 5 µl of the eluate was analysed via Zn2+ phos-tag gel according to the method described by Kumar62. The blots were probed with polyclonal anti-mCh antibody (Abcam, ab167453).
BioIDG3 proximity labelling
Strains expressing BSL1–BioIDG3–NeonGreen and the control BioIDG3–NeonGreen in the cdc20-1 background were labelled with 15N and synchronized in G1 as above. 500-ml TAP or TAP-15N cultures (OD750 ≈ 0.1) were transferred to 33 °C under 120 µmol light m−2 s−1 at t = 0. Biotin was dissolved in TAP or TAP-15N and added to the cultures at t = 11 h to a final concentration of 250 µM. The cultures were collected at t = 14 h via centrifugation for 5 min at 4,000 g. The pellet was weighted and resuspended in 200 µl of buffer (20 mM Tris-HCl (pH 7.4), 150 mM NaCl) per 100 mg of pellet and added dropwise to liquid nitrogen. The pellets were milled in a Retsch tissue homogenizer (M1000, 3 min, 30 Hz) to a fine powder. The sample and control powder were mixed in equal amounts by weight. Proteins were precipitated as above. Precipitated proteins were dissolved in 6 M guanidinium, with sonication, reduced (5 mM tris(2-carboxyethyl)phosphine, 45 min, 55 °C) and alkylated (10 mM iodoacetamide, 45 min, room temperature, dark). Protein concentration was determined (BioRad 5000001), and 1 µg of trypsin (Sigma, T6567) was added to 20 µg of protein in 1.5 M guanidine NH4HCO3 and incubated for 12 h at 37 °C. Trypsin was added again and incubated for 3 h at 37 °C. The reaction was acidified by adding 1% formic acid and desalted over C18 Sep-Pak resin (Waters no. 023590). Desalted peptides were resuspended in 1 ml of PBS (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH 7). 100 µl of M280 beads (Invitrogen 11205D) was equilibrated in PBS and added to the peptides, and the solution was incubated for 1 h at room temperature with constant agitation (800 rpm). The beads were washed once with PBS (5 min at room temperature and 800 rpm), once with PBS + 5% acetonitrile (5 min at room temperature and 800 rpm) and once with ddH2O + 5% acetonitrile (5 min at room temperature and 800 rpm). Biotinylated peptides were eluted three times with 200 µl of 0.2% trifluoroacetic acid, 0.1% formic acid and 80% acetonitrile (each elution 5 min, 75 °C, 800 rpm). The elutions were pooled, and peptides were dried via evacuation, resuspended in 200 µl of 5% acetonitrile and 0.1% trifluoroacetic acid and desalted using ZipTips (MilliporeSigma ZTC18S960). Desalted peptides were dried and analysed via LC–MS/MS.
Bio-layer interferometry
The bio-layer interferometry assay was conducted using a Gator-Plus instrument (Gator Bio) following a protocol described previously63. In brief, anti-GST biosensors (Gator Bio no. 160042) were loaded with GST–CDKB1 or GST expressed in E. coli. These sensors were then immersed in wells containing varying concentrations of purified recombinant BSL1. After a 360-second association phase, the sensors were transferred to 1× PBS for dissociation. The dissociation constant (Kd) was calculated using Gator Bio’s data analysis software, on the basis of the Koff/Kon ratio.
Phosphopeptide enrichment
Wild-type and bsl1-1 cells were pre-grown in TAP-14N or TAP-15N agar plates, inoculated in liquid media and synchronized as above. G1 cells were plated on TAP-agar and incubated at 33 °C in continuous light. Cells were collected by scraping the plates with ice-cold water, added to crushed ice and pelleted via centrifugation in 50-ml tubes. An aliquot of collected cells was kept for microscopic scoring of cell division. The pellet was transferred to microfuge tubes, weighed and resuspended in buffer (17.5 mM Tris-HCl (pH 7.6), 137 mM NaCl with phosphatase inhibitors (Pierce no. 32957, 1 tablet per 5 ml); 200 µl of buffer used per 100 mg of pellet). Wild-type and bsl1-1 cells were mixed in equal amounts on the basis of pellet weight. The mixture was added dropwise to liquid nitrogen and stored at −70 °C. The frozen cells were cryomilled using a Retsch tissue homogenizer (M1000, 3 min, 30 Hz). Protein extraction buffer (100 mM Tris-HCl (pH 8), 2% w/v SDS, 1.2 v/v Triton X-100, 20 mM EGTA, 20 mM EDTA) was added (3 ml per gram of powder), vortexed for 1 min and heated for 10 min at 55 °C. Insoluble material was pelleted via centrifugation for 15 min at 16,000 g. The supernatant was extracted with phenol (pH 8) and the phases separated via centrifugation for 15 min at 16,000 g. The organic phase was re-extracted twice with 50 mM Tris-HCl (pH 8). Proteins were precipitated by adding ice-cold 100 mM ammonium acetate in MeOH with overnight incubation at −70 °C. The precipitated proteins were pelleted via centrifugation for 15 min at 16,000 g. The pellet was washed twice with 100 mM ammonium acetate in MeOH, washed twice with MeOH and air-dried for 10 min to remove traces of MeOH. Proteins were resuspended in 10% SDS for 15 min at 55 °C, followed by 15 min in a heated water bath (42 °C) sonicator (Fischer Scientific Ultrasonic Bath), and finally by sonication with a metal probe (Branson Digital Sonifier, 10% amplitude, 10 s). This resulted in near-complete solubilization of the precipitated proteins. The average yield was 2.3 mg of protein per 100 mg of cell pellet. The solution was made with 5% SDS and 50 mM triethylammonium bicarbonate (pH 8.5), and proteins were reduced (5 mM tris(2-carboxyethyl)phosphine, 45 min, 55 °C) and alkylated (10 mM iodoacetamide, 45 min, room temperature, dark). Approximately 5 mg of protein was digested using S-Trap midi columns (Protifi no. C02-midi-10) according to the manufacturer’s instructions, with 1:20 (w/w) trypsin (Sigma, T6567) per sample. Eluted peptides (~1 mg per sample) were desalted using C18 Sep-Pak resin (Waters no. 023590). Phosphopeptide enrichment was performed using TiO2 tips (Thermo no. 32993) according to the manufacturer’s instructions and analysed via LC–MS/MS. Phosphopeptide enrichment was >95%. Peptide concentrations in flow-through fractions were normalized and analysed via LC–MS/MS.
Mass spectrometry and data analysis
Peptides were analysed via LC–MS/MS on an Easy LC 1200 UPLC liquid chromatography system connected to an Orbitrap Eclipse quadrupole-orbitrap (Thermo Fischer) or a Q-Exactive HF hybrid quadrupole-orbitrap (Thermo Fischer) mass spectrometer in data-dependent acquisition mode. Precursor scans were between 375 and 1,600 m/z, and the 20 most intense multiply charged peptides were selected for fragmentation via higher-energy collision dissociation with normalized collision energy of 27. Tandem mass spectrometry peak lists were extracted using an in-house PAVA script and were searched using ProteinProspector v.6.4.9 (ref. 64) for metabolically labelled peptides essentially as reported previously65, against a database constructed from Chlamydomonas proteins (Phytozome v.5.6) with a decoy database of reversed peptides. Precursor and MS/MS tolerance was set to 10 ppm and 20 ppm, respectively. For phosphopeptide identification, serine/threonine/tyrosine phosphorylation was used as a variable modification. Phosphorylated CDKB1 peptides (IGEGpTpYGKVYK and IGEGpTpYGKVYK) and control peptides (pSEAFVGSQSVR, AApSGLGGDPTFTRR and DRGpSADFGPPVGTR) were detected in data-dependent acquisition mode. To quantify the relative intensity of phosphorylated CDKB1 peptides in bsl1-1 and wild-type cells, the peptides were re-injected with the same Easy LC 1200 UPLC liquid chromatography gradient and analysed using the Eclipse quadrupole-orbitrap in parallel reaction monitoring mode, as described previously39. The intensity of MS2 fragment ions across the elution peaks was plotted using Skyline v.23.1.0.268 (ref. 66), as described previously39.
Sequence alignments and accession numbers
Multiple sequence alignments were performed using the Muscle algorithm in Jalview67. The identifiers for A. thaliana are NP_192217.2 for AtBSL1, NP_001184935.1 for AtBSL2, NP_180289.3 for AtBSL3 and NP_171844.6 for AtBSU1. The identifier for Toxoplasma gondii PPKL is XP_002368432.1, and that for Plasmodium falciparum PPKL is XP_001348804.1. The Phytozome.org (v.5.6) identifiers for the Chlamydomonas genes mentioned in this study are Cre01.g050850 for BSL1, Cre08.g372550 for CDKB1, Cre08.g370401 for CYCB1, Cre06.g295700 for MCM3 and Cre12.g492950 for RIR1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are available in the main text or the Supplementary Information. The mass spectrometry raw data have been deposited at PRIDE with accession number PXD068429. Source data are provided with this paper.
Code availability
All code is available upon request.
References
Morgan, D. O. The Cell Cycle: Principles of Control (New Science Press, 2007).
Cross, F. R., Buchler, N. E. & Skotheim, J. M. Evolution of networks and sequences in eukaryotic cell cycle control. Phil. Trans. R. Soc. B 366, 3532–3544 (2011).
Freville, A. et al. Deciphering the role of protein phosphatases in Apicomplexa: the future of innovative therapeutics? Microorganisms https://doi.org/10.3390/microorganisms10030585 (2022).
Karlsson-Rosenthal, C. & Millar, J. B. Cdc25: mechanisms of checkpoint inhibition and recovery. Trends Cell Biol. 16, 285–292 (2006).
Gould, K. L. & Nurse, P. Tyrosine phosphorylation of the fission yeast cdc2+ protein kinase regulates entry into mitosis. Nature 342, 39–45 (1989).
Krek, W. & Nigg, E. A. Mutations of p34cdc2 phosphorylation sites induce premature mitotic events in HeLa cells: evidence for a double block to p34cdc2 kinase activation in vertebrates. EMBO J. 10, 3331–3341 (1991).
Millar, J. B. & Russell, P. The cdc25 M-phase inducer: an unconventional protein phosphatase. Cell 68, 407–410 (1992).
Kumagai, A. & Dunphy, W. G. The cdc25 protein controls tyrosine dephosphorylation of the cdc2 protein in a cell-free system. Cell 64, 903–914 (1991).
Sur, S. & Agrawal, D. K. Phosphatases and kinases regulating CDC25 activity in the cell cycle: clinical implications of CDC25 overexpression and potential treatment strategies. Mol. Cell. Biochem. 416, 33–46 (2016).
Boudolf, V., Inze, D. & De Veylder, L. What if higher plants lack a CDC25 phosphatase?. Trends Plant Sci. 11, 474–479 (2006).
Nowack, M. K. et al. Genetic framework of cyclin-dependent kinase function in Arabidopsis. Dev. Cell 22, 1030–1040 (2012).
Atkins, K. C. & Cross, F. R. Interregulation of CDKA/CDK1 and the plant-specific cyclin-dependent kinase CDKB in control of the Chlamydomonas cell cycle. Plant Cell 30, 429–446 (2018).
Pecani, K., Lieberman, K., Tajima-Shirasaki, N., Onishi, M. & Cross, F. R. Control of division in Chlamydomonas by cyclin B/CDKB1 and the anaphase-promoting complex. PLoS Genet. 18, e1009997 (2022).
Tulin, F. & Cross, F. R. A microbial avenue to cell cycle control in the plant superkingdom. Plant Cell 26, 4019–4038 (2014).
Boudolf, V. et al. B1-type cyclin-dependent kinases are essential for the formation of stomatal complexes in Arabidopsis thaliana. Plant Cell 16, 945–955 (2004).
Boudolf, V. et al. The plant-specific cyclin-dependent kinase CDKB1;1 and transcription factor E2Fa-DPa control the balance of mitotically dividing and endoreduplicating cells in Arabidopsis. Plant Cell 16, 2683–2692 (2004).
Fisher, D. L. & Nurse, P. A single fission yeast mitotic cyclin B p34cdc2 kinase promotes both S-phase and mitosis in the absence of G1 cyclins. EMBO J. 15, 850–860 (1996).
Herbst, J., Li, Q. Q. & De Veylder, L. Mechanistic insights into DNA damage recognition and checkpoint control in plants. Nat. Plants 10, 539–550 (2024).
Dissmeyer, N. et al. Control of cell proliferation, organ growth, and DNA damage response operate independently of dephosphorylation of the Arabidopsis Cdk1 homolog CDKA;1. Plant Cell 21, 3641–3654 (2009).
Cross, F. R. & Umen, J. G. The Chlamydomonas cell cycle. Plant J. 82, 370–392 (2015).
Breker, M., Lieberman, K. & Cross, F. R. Comprehensive discovery of cell-cycle-essential pathways in Chlamydomonas reinhardtii. Plant Cell 30, 1178–1198 (2018).
Tulin, F. & Cross, F. R. Cyclin-dependent kinase regulation of diurnal transcription in Chlamydomonas. Plant Cell 27, 2727–2742 (2015).
Maselli, G. A. et al. Revisiting the evolutionary history and roles of protein phosphatases with Kelch-like domains in plants. Plant Physiol. 164, 1527–1541 (2014).
Guttery, D. S. et al. A unique protein phosphatase with Kelch-like domains (PPKL) in Plasmodium modulates ookinete differentiation, motility and invasion. PLoS Pathog. 8, e1002948 (2012).
Yang, C., Doud, E. H., Sampson, E. & Arrizabalaga, G. The protein phosphatase PPKL is a key regulator of daughter parasite development in Toxoplasma gondii. mBio 14, e0225423 (2023).
Mora-Garcia, S. et al. Nuclear protein phosphatases with Kelch-repeat domains modulate the response to brassinosteroids in Arabidopsis. Genes Dev. 18, 448–460 (2004).
Kim, T. W. et al. Brassinosteroid signal transduction from cell-surface receptor kinases to nuclear transcription factors. Nat. Cell Biol. 11, 1254–1260 (2009).
Park, C. H. et al. Deconvoluting signals downstream of growth and immune receptor kinases by phosphocodes of the BSU1 family phosphatases. Nat. Plants 8, 646–655 (2022).
Guo, X., Park, C. H., Wang, Z. Y., Nickels, B. E. & Dong, J. A spatiotemporal molecular switch governs plant asymmetric cell division. Nat. Plants 7, 667–680 (2021).
Kim, T. W., Michniewicz, M., Bergmann, D. C. & Wang, Z. Y. Brassinosteroid regulates stomatal development by GSK3-mediated inhibition of a MAPK pathway. Nature 482, 419–422 (2012).
Guo, X., Ding, X. & Dong, J. Dichotomy of the BSL phosphatase signaling spatially regulates MAPK components in stomatal fate determination. Nat. Commun. 13, 2438 (2022).
Myles, T., Schmidt, K., Evans, D. R., Cron, P. & Hemmings, B. A. Active-site mutations impairing the catalytic function of the catalytic subunit of human protein phosphatase 2A permit baculovirus-mediated overexpression in insect cells. Biochem. J. 357, 225–232 (2001).
Harris, J. A., Liu, Y., Yang, P., Kner, P. & Lechtreck, K. F. Single-particle imaging reveals intraflagellar transport-independent transport and accumulation of EB1 in Chlamydomonas flagella. Mol. Biol. Cell 27, 295–307 (2016).
Onishi, M., Umen, J. G., Cross, F. R. & Pringle, J. R. Cleavage-furrow formation without F-actin in Chlamydomonas. Proc. Natl Acad. Sci. USA 117, 18511–18520 (2020).
Ikui, A. E., Ueki, N., Pecani, K. & Cross, F. R. Control of pre-replicative complex during the division cycle in Chlamydomonas reinhardtii. PLoS Genet. 17, e1009471 (2021).
Salisbury, J. L., Baron, A. T. & Sanders, M. A. The centrin-based cytoskeleton of Chlamydomonas reinhardtii: distribution in interphase and mitotic cells. J. Cell Biol. 107, 635–641 (1988).
O’Toole, E. T. & Dutcher, S. K. Site-specific basal body duplication in Chlamydomonas. Cytoskeleton (Hoboken) 71, 108–118 (2014).
Nakazawa, Y., Hiraki, M., Kamiya, R. & Hirono, M. SAS-6 is a cartwheel protein that establishes the 9-fold symmetry of the centriole. Curr. Biol. 17, 2169–2174 (2007).
Reyes, A. V., Shrestha, R., Baker, P. R., Chalkley, R. J. & Xu, S. L. Application of parallel reaction monitoring in (15)N labeled samples for quantification. Front. Plant Sci. 13, 832585 (2022).
Rhind, N. & Russell, P. Tyrosine phosphorylation of cdc2 is required for the replication checkpoint in Schizosaccharomyces pombe. Mol. Cell. Biol. 18, 3782–3787 (1998).
Harper, J. D. I. & John, P. C. L. Coordination of division events in the Chlamydomonas cell cycle. Protoplasma 131, 118–130 (1986).
Morgan, D. O. Principles of CDK regulation. Nature 374, 131–134 (1995).
Donzelli, M. & Draetta, G. F. Regulating mammalian checkpoints through Cdc25 inactivation. EMBO Rep. 4, 671–677 (2003).
Cizkova, M., Slavkova, M., Vitova, M., Zachleder, V. & Bisova, K. Response of the green alga Chlamydomonas reinhardtii to the DNA damaging agent zeocin. Cells https://doi.org/10.3390/cells8070735 (2019).
White, M. W. & Suvorova, E. S. Apicomplexa cell cycles: something old, borrowed, lost, and new. Trends Parasitol. 34, 759–771 (2018).
Blomberg, I. & Hoffmann, I. Ectopic expression of Cdc25A accelerates the G(1)/S transition and leads to premature activation of cyclin E- and cyclin A-dependent kinases. Mol. Cell. Biol. 19, 6183–6194 (1999).
Santamaria, D. et al. Cdk1 is sufficient to drive the mammalian cell cycle. Nature 448, 811–815 (2007).
Gutiérrez-Escribano, P. & Nurse, P. A single cyclin–CDK complex is sufficient for both mitotic and meiotic progression in fission yeast. Nat. Commun. 6, 6871 (2015).
Welburn, J. P. I. et al. How tyrosine 15 phosphorylation inhibits the activity of cyclin-dependent kinase 2-cyclin A. J. Biol. Chem. 282, 3173–3181 (2007).
Elbaek, C. R., Petrosius, V. & Sorensen, C. S. WEE1 kinase limits CDK activities to safeguard DNA replication and mitotic entry. Mutat. Res. 819–820, 111694 (2020).
Bisova, K., Krylov, D. M. & Umen, J. G. Genome-wide annotation and expression profiling of cell cycle regulatory genes in Chlamydomonas reinhardtii. Plant Physiol. 137, 475–491 (2005).
De Schutter, K. et al. Arabidopsis WEE1 kinase controls cell cycle arrest in response to activation of the DNA integrity checkpoint. Plant Cell 19, 211–225 (2007).
Pan, T. et al. A novel WEE1 pathway for replication stress responses. Nat. Plants 7, 209–218 (2021).
Bouldin, C. M. & Kimelman, D. Cdc25 and the importance of G2 control: insights from developmental biology. Cell Cycle 13, 2165–2171 (2014).
Tulin, F. Mating and tetrad dissection in Chlamydomonas. Bio Protoc. 9, e3207 (2019).
Onishi, M. & Pringle, J. R. Robust transgene expression from bicistronic mRNA in the green alga Chlamydomonas reinhardtii. G3 (Bethesda) 6, 4115–4125 (2016).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
Yamano, T., Iguchi, H. & Fukuzawa, H. Rapid transformation of Chlamydomonas reinhardtii without cell-wall removal. J. Biosci. Bioeng. 115, 691–694 (2013).
Perlaza, K. et al. The Mars1 kinase confers photoprotection through signaling in the chloroplast unfolded protein response. Elife https://doi.org/10.7554/eLife.49577 (2019).
R Core Team R: A Language and Environment for Statistical Computing (R Core Team, 2014).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016).
Kumar, G. A simple method for detecting phosphorylation of proteins by using Zn(2+)-Phos-Tag SDS-PAGE at neutral pH. Methods Mol. Biol. 1853, 223–229 (2018).
Aizezi, Y. et al. Structure-based virtual screening identifies small-molecule inhibitors of O-fucosyltransferase SPINDLY in Arabidopsis. Plant Cell 36, 497–509 (2024).
Chalkley, R. J. et al. Comprehensive analysis of a multidimensional liquid chromatography mass spectrometry dataset acquired on a quadrupole selecting, quadrupole collision cell, time-of-flight mass spectrometer: II. New developments in Protein Prospector allow for reliable and comprehensive automatic analysis of large datasets. Mol. Cell Proteom. 4, 1194–1204 (2005).
Shrestha, R. et al. (15)N metabolic labeling quantification workflow in Arabidopsis using Protein Prospector. Front. Plant Sci. 13, 832562 (2022).
Schilling, B. et al. Platform-independent and label-free quantitation of proteomic data using MS1 extracted ion chromatograms in Skyline: application to protein acetylation and phosphorylation. Mol. Cell Proteom. 11, 202–214 (2012).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Acknowledgements
We thank A. Malkovskiy at the Carnegie Institution Imaging Facility. We thank A. Ikui (Brooklyn Collage) for providing the cycb1-5 mutant and F. Cross (Rockefeller University) for providing the ble–GFP strain. We thank Y. Schreckenberg for technical assistance. The CDKB–mCh (pKA1) and CDKB(AF)–mCh (pKA17) plasmids were constructed by K. C. Atkins and provided by F. Cross. The research reported in this publication was supported by funding from the European Union’s Horizon 2020 research and innovation programme (2014–2020) under Marie Skłodowska-Curie grant agreement no. 798198 (F.T.), the National Institute of General Medical Sciences of the National Institutes of Health (under award nos R01GM066258 to Z.-Y.W. and R01GM135706 and S10OD030441 to S.-l.X.), the National Science Foundation (MCB 1818383 to M.O. and J. R. Pringle) and the Carnegie Endowment Fund to the Carnegie Mass Spectrometry Facility.
Author information
Authors and Affiliations
Contributions
F.T., F.F.A. and Z.-Y.W. conceptualized the project. F.T., A.V.R., Y.A., Y.F., M.O., S.-l.X. and Z.-Y.W. developed the methodology. F.T. and Z.-Y.W. conducted the investigation. F.T. and Z.-Y.W. visualized the data. F.T., F.F.A. and Z.-Y.W. acquired the funding. F.T., F.F.A. and Z.-Y.W. were responsible for project administration. Z.-Y.W., A.G. and F.F.A. supervised the project. F.T., F.F.A. and Z.-Y.W. wrote the original draft of the manuscript. F.T., S.-l.X., M.O., F.F.A. and Z.-Y.W. reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Xingguang Deng, Chao Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 BSL1 phosphatase activity is essential for viability in Chlamydomonas.
a, Anti-Flag immunoblot analysis of five bsl1-1:BSL1-VF and four bsl1-1:BSL1D619N-VF transformants. Experiment performed once. b, Growth assays at 21oC with or without paromomycin (paro) and at 33oC without paromomycin. Paromomycin is the selection marker linked to BSL1-VF and BSL1D619N-VF.
Extended Data Fig. 2 CRISPR-Cas9-guided knock-out of BSL1.
a, Diagram of the bicistronic BSL1-Venus-3xFLAG (BSL-VF) construct showing Kelch and phsophatase (PP) domains. b, The APHVII hygromycin resistance cassette was delivered with or without Cas9 to bsl1-1:BSL1-VF, and hygromycin resistant strains were grown under the indicated conditions. Loss of paromomycin resistance and recovery of temperature sensitivity indicates APHVII integration into the BSL1-VF transgene (white boxes). c, Genotyping of transformants in (b). AluI cuts the wild-type BSL1 sequence. d, PCR amplification of APHVII insertion in transformants 1, 5, and 12, with primers in BSL1 and in the APHVII gene. e, Sanger sequencing of transformants 1, 5, 12. The APHVII gene (white block) is inserted in either the BSL1-VF transgene (#1, wild type CtA sequence, purple box) or in the bsl1-1 gene (#5 and #12). Clone #5 is referred to as bsl1-ko. f, Segregation analysis of the wild type crossed to transformants #1, #5 and #12 (6 tetrads of each cross shown). No hygR paroS recombinants (bsl1-ko, red box) could be isolated in #5 and #12, indicating that disruption of the bsl1-1 locus, without the BSL1-VF rescue construct, is lethal.
Extended Data Fig. 3 Spindle misorientation in the bsl1-1 mutant grown below 33oC.
a, Wild type and bsl1-1 mutant cells co-expressing EB1-mNeonGreen (green) and centrin-mCherry (magenta) imaged by time-lapse microscopy. The time (minutes) of each image is relative to the nuclear-basal body contact (NBBC, time 0). Nucleus (n) and basal body (BB) are marked. In wild type, the BBs segregate (time 3’), the mitotic spindle forms with centrin-mCh localized to the spindle poles (white asterisks). In bsl1-1, NBBC and BB separation appears similar to the wild type but one spindle pole does not recruit centrin-mCherry in metaphase (white arrow); the cleavage furrow MTs are more disorganized compared to the wild type; and the 2nd and 3rd divisions display highly aberrant spindles. Scale: 5 µm. b, The orientation of the mitotic spindle was measured as the angle (α) between one line through the centers of the spindle and pyrenoid (P) and one line through the two spindle poles. c, Quantification of spindle orientation in cells grown at 21 °C showed no difference between sample means (one-way ANOVA, df=3,37, F = 1.1, p = 0.35). At 30 °C, the average spindle angle of bsl1-1 and bsl1:BSL1D619N was different from the wild type (ANOVA, df=3,119, F = 9.2, p = 1e-5). Occasional spindle misorientation was observed in bsl1-1 at 21 °C (black arrow). d, Quantification of the duration of metaphase at 21 °C indicated longer metaphase duration in bsl1-1 (p = 0.006, two-sided Welch test on sample means, df=5.1). e, Quantification of the duration of metaphase at 30 °C indicated longer metaphase duration in bsl1-1 and bsl1-1:BSL1D619N (one-way ANOVA, df=3,93, F = 12.01, p = 1e-6).
Extended Data Fig. 4 BSL1 phosphorylation sites.
a, Diagram of BSL1 protein structure shows the relative locations of phosphosites identified only in S/M phase cells (red arrow) and those identified in both S/M and G1 cells (blue arrows). b, Multiple sequence alignment of Chlamydomonas BSL1, Arabidopsis BSL1,2,3 and BSU1, PPKL from Toxoplasma gondii (Tg) and Plasmodium falciparum (Pf). Asterisks: phosphosites, color codes as in (a) with a peptide expect value < 0.001 (see Source Data File) marked.
Extended Data Fig. 5 Proteomic analyses of the bsl1-1 mutant.
a, Quantification of cell cycle progression at 0, 6, 8 and 10 hrs at the restrictive temperature. Over 100 cells were counted per timepoint per sample. For quantitative mass spectrometry analysis, the Light and heavy N samples were combined as: 1 + 4 (FORWARD), and 2 + 3 (REVERSE). b, Example of cells from the respective category. This experiment was performed once. c, Spectra counts of selected proteins in wild type and bsl1-1. Spectra counts of BSL1L46P are reduced compared to wild type. Ribonucleotide reductase (RIR1) and the helicase MCM3 and CDKA accumulate with similar kinetics in wild type and bsl1-1. Accumulation of CYCB1 and CDKB1 is delayed in bsl1-1.
Extended Data Fig. 6 Targeted quantification of T14,Y15-phosphorylated CDKB1.
a, MS1 spectrum of the doubly phosphorylated CDKB IGEGpTpYGK peptide shows a stronger signal in bsl1-1 (14 N) than WT (15 N). b, MS2 spectrum shows the fragmentation pattern of the m/z = 687.79 peak in (a). (c-f), Targeted quantification of the IGEGpTpYGK peptide (phosphorylated on T14 and Y15, with +2 or +3 charge state) in PRM (parallel reaction monitoring) mode. Each colored curve represents the intensity of a specific MS2 ion at a given retention time. Peptides from t = 8 h and t = 10 h timepoints were analyzed. The bsl1-1 mutant cells were metabolically labeled with heavy nitrogen (15 N) in rep. 1. In rep. 2, the wild type was labeled with 15 N. c, t = 8 h, repeat 1; d, t = 8 h, repeat 2; e, t = 10 h, repeat 1; f, t = 10 h, repeat 2.
Extended Data Fig. 7 Summary of PRM MS2 quantification of CDKB1 and control phosphopeptides in bsl1-1 and wild type.
a-e, Each bar shows the total integrated peak intensity (MS2 quantification, parallel reaction monitoring mode, PRM) of the indicated phosphopeptides. Colors represent the integrated area of different MS2 ions across the elution peak. In all panels, rep.1 = 14N-WT vs 15N-bsl1-1, rep.2 = 14N-bsl1-1 vs 15N-WT. a, The doubly phosphorylated peptide IGEGpTpYGK is increased in bsl1-1 cells compared to wild type (15 N/bsl1-1 peak stronger in rep.1 and 14 N/bsl1-1 peak stronger in rep.2). Ion intensity as a function of retention time for this peptide is shown in Extended Data Fig. 6. b, The singly phosphorylated peptide IGEGTpYGK shows similar total integrated intensity in bsl1-1 and wild type. c, Quantification of phosphorylated CYCB1. d-e, Quantification of two phosphorylated control peptides, which show similar integrated intensity in bsl1-1 and wild type at t = 8 h and t = 10 h.
Extended Data Fig. 8 Quantitative proximity labeling indicates proximity between CYCB1 and BSL1.
BSL1 was fused to an engineered version of the biotin ligase BioID (BioIDG3-NeonGreen, see Methods). BioIDG3-NeonGreen was used as a control. Biotinylated peptides were purified and analyzed by mass spectrometry. a-b, Control cells were metabolically labeled with 15 N. a, MS1 spectrum (black arrow) of the CYCB1 peptide AAAGVK(Bio)AAAGLQAQNKR. The corresponding peak in the control sample (grey arrow) has lower intensity. b, MS2 fragmentation pattern for the peptide in (a). c-d, The experiment was repeated with the labeling inverted (heavy BSL1-BioIDG3-NG). c, MS1 spectrum of the peptide EK(Bio)AAVPVPADGVK (black arrow), d, MS2 fragmentation pattern of the peptide in (c). e, Table detailing intensity ratios of all CYCB1 peptides detected in four independent experiments. The median intensity ratio (BSL1/control) is 8.
Extended Data Fig. 9 Phosphorylation of CDKB1 T14 and Y15 is required for optimal growth.
a, The cdkb1-1 was transformed with wild-type CDKB1-mCherry, and variants containing AF, DY, TE, and DE substitutions at position T14 and Y15. Only wild type CDKB1 and the CDKB1AF variant could rescue cdkb1-1 at 33 °C. Transformation efficiency was assessed by plating half of the transformed cells on plates containing paromomycin at 21 °C. b-c, Quantification of colony size of individual clones of cdkb1-1 rescued by CDKB1-mCh and CDKB1AF-mCh. b, Colony size after 5 days growth at 33 °C. Inset: combined data for CDKB1-mCh and CDKB1AF-mCh indicates a smaller colony size for CDKBAF-mCh (one-way ANOVA, df=2,1375, F = 373.5, p = 2e-16), n=colonies scored. c, Colony size after 7 days growth at 21 °C (one-way ANOVA df = 3,82, F = 4.5, p = 0.005). n=colonies scored. d, Complementation experiments. The bsl1-1 and cdkb1-1 mutants were transformed with plasmids containing the BSL1 and CDKB1 genes. The bsl1-1 mutant was rescued by BSL1 but not by with wild-type CDKB1 or CDKB1AF. Transformed cells were split evenly. Half of the cells were plated on 10 μg/ml paromomycin at a permissive temperature (21 °C) as a transformation efficiency control. The other half was plated at 33 °C to select for complementation of the bsl1-1 or cdkb1-1 mutants. e, Timing of cleavage furrow formation in bsl1-1 and bsl1-1 CDKBAF. Each data point represents data from two independent experiments. 20-30 cells were counted in each experiment. f, α-mCh western blot of CDKB1AF-mCh in synchronized cells (BSL1 or bsl1-1). CDKB1-mCh runs as two bands. Reduced CDKB1 level also shown in Extended Data Fig. 5c. g, bsl1-1 CDKB1 and bsl1-1 CDKBAF cells were transformed with phosphatase-dead BSL1D619N-Venus or BSL1WT-Venus, and selected on paromomycin at 21 °C. Expression of BSL1 was verified by localization to spindle poles. Rescue of bsl1-1 was tested at 33 °C. Only BSL1WT rescued bsl1-1.
Extended Data Fig. 10 Mutations of CDKB1-T14/Y15 and BSL1 alter the sensitivity to DNA replication stress.
a, Cells expressing CDKB1AF are hypersensitive to the polymerase inhibitor aphidicholin, but not to the DNA-damage-inducing drug zeocin. cdkb1-1 cells complemented with either wild type CDKB1-mCherry (CDKB1) or unphosphorylatable CDKB1AF-mCherry (CDKB1AF, three independent clones). The density of each culture was normalized to OD750 ~ 0.4 and cells were spotted in 5-fold serial dilutions on TAP agar plates containing 50 μM aphidicholin or 2 μg/ml zeocin and incubated at 21 °C and 33 °C. b, Sensitivity of the bsl1-1 mutant to HU. Cells were spotted (5-fold serial dilutions) on TAP agar plates, and plates supplemented with 1 mM HU, and incubated at at 21 °C.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2.
Supplementary Video 1 (download AVI )
Dynamics of ble–GFP and EB1–mSc in mitosis. A wild-type cell expressing ble–GFP (a nuclear marker) and EB1–mSc (a microtubule marker) grown at 33 °C is shown. Acquisition every 3 min. Scale bar, 10 µm.
Supplementary Video 2 (download AVI )
Mitosis dynamics of ble–GFP and EB1–mSc in bsl1-1. A bsl1-1 cell expressing ble–GFP and EB1–mSc grown at 33 °C is shown. Acquisition every 3 min. Chloroplast division at t = 36 min, cleavage furrow at t = 72 min. The nucleus does not divide. Scale bar, 10 µm.
Supplementary Video 3 (download AVI )
Mitosis dynamics of ble–GFP and EB1–mSc in bsl1-1. A bsl1-1 cell expressing ble–GFP and EB1–mSc grown at 33 °C is shown. Acquisition every 3 min. Cleavage furrow at t = 51 min. No spindle forms. The ingressing furrow bisects the nucleus. Scale bar, 10 µm.
Supplementary Video 4 (download AVI )
Mitosis dynamics of ble–GFP in bsl1-ko. One tetrad from the cross bsl1-ko:BSL1-VF x EB1-mScarlet ble-GFP is shown. The bsl1-ko segregant was assigned on the basis of the lysis phenotype at the 8–16-cell stage (Fig. 3 and Extended Data Fig. 3a). The presence of BSL1–VF was assigned according to the localization of BSL1–VF to spindle poles in mitotic cells. bsl1-ko is recessive in the presence of BSL1–VF. The genotypes of the meiotic products are: (A) BSL1-VF EB1-mSc, (B) BSL1-VF, (C) ble-GFP EB1-mSc, (D) bsl-ko ble-GFP. The second allele of bsl1-ko may be in either A or B. Scale bar, 10 µm.
Supplementary Video 5 (download AVI )
Division of wild-type cells. A wild-type cell expressing ble–GFP (a nuclear marker) was imaged every hour. Scale bar, 5 µm.
Supplementary Video 6 (download AVI )
Division of bls1-ko cells. A bsl1-ko cell expressing ble-GFP was imaged every hour. Scale bar, 5 µm.
Supplementary Video 7 (download AVI )
Localization of BSL1–Venus throughout mitosis. A bsl1-1 cell complemented with BSL1–Venus (yellow) and co-expressing EB1–mSc (magenta) is shown undergoing mitosis. Acquisition every 3 min. Scale bar, 5 µm.
Supplementary Video 8 (download AVI )
Localization of BSL1–Venus through a multiple-fission cycle. A bsl1-1 cell complemented with BSL1–Venus (yellow) is shown. Acquisition every 1 h. The cell undergoes two divisions, producing four daughters. At exit from S/M, BSL1–Venus returns to the anterior end of the cell.
Source data
Source Data Fig. 1 (download PDF )
Unmodified blots for Fig. 1b.
Source Data Fig. 5 (download PDF )
Unmodified blots for Fig. 5c,d.
Source Data Fig. 6 (download PDF )
Two additional repeats of the experiment shown in Fig. 6c.
Source Data Figs. 3 and 6 and Extended Data Figs. 3–6 and 9 (download XLSX )
Multi-tab Excel file in which each tab contains source data for one main or extended data figure. Each data and legend tab is labelled with the respective figure number.
Source Data Extended Data Fig. 4 (download PDF )
Screenshots of ProteinProspector MS2 spectra for Extended Data Fig. 4.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tulin, F., Aizezi, Y., Reyes, A.V. et al. Mitotic entry is controlled by the plant-specific phosphatase BSL1 and cyclin-dependent kinase B. Nat. Plants 11, 2395–2408 (2025). https://doi.org/10.1038/s41477-025-02145-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02145-z