Abstract
How somatic cells acquire totipotency and subsequently develop into a whole plant (plantlet) remains a mystery in plant biology. Here we used three Kalanchoe species to address this fundamental question. By assembling high-quality chromosome-level reference genomes and conducting comparative genomic analyses, we reveal hidden signatures of gene expansion, contraction and loss during the evolution of Kalanchoe species and elucidate conserved temporal gene expression signatures and epigenetic states during plantlet formation. Remarkably, we uncover three innovations contributing to the plantlet formation in Kalanchoe. Specifically, our results suggest that the loss of the F-box gene LCR is a prerequisite for plantlet formation. Both gene duplication and increased chromatin accessibility of pluripotency-associated genes further create conditions that enhance the potential of plantlet formation. The previously uncharacterized gene KdLBD19 could be leveraged to improve crop transformation efficiency. Overall, this study reveals the genetic basis underlying the acquisition of totipotency and plantlet formation in Kalanchoe.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All high-throughput sequencing data used in this study, including short-read sequencing data, Nanopore long-read sequencing data, Hi-C sequencing data, RNA-seq sequencing data and ATAC-seq sequencing data, were deposited in the National Genomics Data Center (NGDC) under the accession number PRJCA051258. The assembled reference genomes of the three Kalanchoe species are available via figshare at https://doi.org/10.6084/m9.figshare.29109305 (ref. 106). Source data are provided with this paper.
References
Arnoux-Courseaux, M. & Coudert, Y. Re-examining meristems through the lens of evo-devo. Trends Plant Sci. 29, 413–427 (2024).
Eshed Williams, L. Genetics of shoot meristem and shoot regeneration. Annu. Rev. Genet. 55, 661–681 (2021).
Chen, Z., Debernardi, J. M., Dubcovsky, J. & Gallavotti, A. Recent advances in crop transformation technologies. Nat. Plants 8, 1343–1351 (2022).
Chen, C. et al. Plant regeneration in the new era: from molecular mechanisms to biotechnology applications. Sci. China Life Sci. 67, 1338–1367 (2024).
Vogel, G. How does a single somatic cell become a whole plant?. Science 309, 86 (2005).
Su, Y. H., Tang, L. P., Zhao, X. Y. & Zhang, X. S. Plant cell totipotency: insights into cellular reprogramming. J. Integr. Plant Biol. 63, 228–243 (2021).
Wang, F. X., Shang, G. D. & Wang, J. W. Towards a hierarchical gene regulatory network underlying somatic embryogenesis. Trends Plant Sci. 27, 1209–1217 (2022).
Garces, H. M. et al. Evolution of asexual reproduction in leaves of the genus Kalanchoe. Proc. Natl Acad. Sci. USA 104, 15578–15583 (2007).
Karami, O. et al. An Arabidopsis AT-hook motif nuclear protein mediates somatic embryogenesis and coinciding genome duplication. Nat. Commun. 12, 2508 (2021).
Gallois, J. L., Nora, F. R., Mizukami, Y. & Sablowski, R. WUSCHEL induces shoot stem cell activity and developmental plasticity in the root meristem. Genes Dev. 18, 375–380 (2004).
Zuo, J., Niu, Q. W., Frugis, G. & Chua, N. H. The WUSCHEL gene promotes vegetative-to-embryonic transition in Arabidopsis. Plant J. 30, 349–359 (2002).
Harding, E. W., Tang, W., Nichols, K. W., Fernandez, D. E. & Perry, S. E. Expression and maintenance of embryogenic potential is enhanced through constitutive expression of AGAMOUS-Like 15. Plant Physiol. 133, 653–663 (2003).
Boutilier, K. et al. Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell 14, 1737–1749 (2002).
Gazzarrini, S., Tsuchiya, Y., Lumba, S., Okamoto, M. & McCourt, P. The transcription factor FUSCA3 controls developmental timing in Arabidopsis through the hormones gibberellin and abscisic acid. Dev. Cell 7, 373–385 (2004).
Lotan, T. et al. Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93, 1195–1205 (1998).
Stone, S. L. et al. Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: implications for somatic embryogenesis. Proc. Natl Acad. Sci. USA 105, 3151–3156 (2008).
Hecht, V. et al. The Arabidopsis SOMATIC EMBRYOGENESIS RECEPTOR KINASE 1 gene is expressed in developing ovules and embryos and enhances embryogenic competence in culture. Plant Physiol. 127, 803–816 (2001).
Thakare, D., Tang, W., Hill, K. & Perry, S. E. The MADS-domain transcriptional regulator AGAMOUS-LIKE15 promotes somatic embryo development in Arabidopsis and soybean. Plant Physiol. 146, 1663–1672 (2008).
Tang, L. P. et al. Time-resolved reprogramming of single somatic cells into totipotent states during plant regeneration. Cell 188, 6923–6938 e6918 (2025).
Zhang, T. Q. et al. A two-step model for de novo activation of WUSCHEL during plant shoot regeneration. Plant Cell 29, 1073–1087 (2017).
Iwase, A. et al. WIND1 promotes shoot regeneration through transcriptional activation of ENHANCER OF SHOOT REGENERATION1 in Arabidopsis. Plant Cell 29, 54–69 (2017).
Iwase, A. et al. The AP2/ERF transcription factor WIND1 controls cell dedifferentiation in Arabidopsis. Curr. Biol. 21, 508–514 (2011).
Kareem, A. et al. PLETHORA genes control regeneration by a two-step mechanism. Curr. Biol. 25, 1017–1030 (2015).
Banno, H., Ikeda, Y., Niu, Q. W. & Chua, N. H. Overexpression of Arabidopsis ESR1 induces initiation of shoot regeneration. Plant Cell 13, 2609–2618 (2001).
Daimon, Y., Takabe, K. & Tasaka, M. The CUP-SHAPED COTYLEDON genes promote adventitious shoot formation on calli. Plant Cell Physiol. 44, 113–121 (2003).
Garces, H. M., Koenig, D., Townsley, B. T., Kim, M. & Sinha, N. R. Truncation of LEAFY COTYLEDON1 protein is required for asexual reproduction in Kalanchoe daigremontiana. Plant Physiol. 165, 196–206 (2014).
Blount, Z. D., Lenski, R. E. & Losos, J. B. Contingency and determinism in evolution: replaying life’s tape. Science https://doi.org/10.1126/science.aam5979 (2018).
Saul, F. et al. Subgenome dominance shapes novel gene evolution in the decaploid pitcher plant Nepenthes gracilis. Nat. Plants 9, 2000–2015 (2023).
Stankowski, S. et al. The genetic basis of a recent transition to live-bearing in marine snails. Science 383, 114–119 (2024).
Chomicki, G. et al. Convergence in carnivorous pitcher plants reveals a mechanism for composite trait evolution. Science 383, 108–113 (2024).
Jaillon, O. et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, 463–467 (2007).
Zhu, S. et al. Genomic insights on the contribution of balancing selection and local adaptation to the long-term survival of a widespread living fossil tree, Cercidiphyllum japonicum. New Phytol. 228, 1674–1689 (2020).
Badouin, H. et al. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 546, 148–152 (2017).
Rensing, S. A. Gene duplication as a driver of plant morphogenetic evolution. Curr. Opin. Plant Biol. 17, 43–48 (2014).
Alvarez, J. P., Furumizu, C., Efroni, I., Eshed, Y. & Bowman, J. L. Active suppression of a leaf meristem orchestrates determinate leaf growth. eLife https://doi.org/10.7554/elife.15023 (2016).
Ejaz, M., Bencivenga, S., Tavares, R., Bush, M. & Sablowski, R. ARABIDOPSIS THALIANA HOMEOBOX GENE 1 controls plant architecture by locally restricting environmental responses. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2018615118 (2021).
Pi, L. et al. Organizer-derived WOX5 signal maintains root columella stem cells through chromatin-mediated repression of CDF4 expression. Dev. Cell 33, 576–588 (2015).
Wang, Q., Kohlen, W., Rossmann, S., Vernoux, T. & Theres, K. Auxin depletion from the leaf axil conditions competence for axillary meristem formation in Arabidopsis and tomato. Plant Cell 26, 2068–2079 (2014).
Hwang, I., Sheen, J. & Muller, B. Cytokinin signaling networks. Annu. Rev. Plant Biol. 63, 353–380 (2012).
Dewitte, W. et al. Arabidopsis CYCD3 D-type cyclins link cell proliferation and endocycles and are rate-limiting for cytokinin responses. Proc. Natl Acad. Sci. USA 104, 14537–14542 (2007).
Cheng, Y., Dai, X. & Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev. 20, 1790–1799 (2006).
Carroll, S. B. Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell 134, 25–36 (2008).
Fujii, S. et al. A stigmatic gene confers interspecies incompatibility in the Brassicaceae. Nat. Plants 5, 731–741 (2019).
Wang, Y. et al. The Arabidopsis RAD51 paralogs RAD51B, RAD51D and XRCC2 play partially redundant roles in somatic DNA repair and gene regulation. New Phytol. 201, 292–304 (2014).
Knauer, S. et al. A protodermal miR394 signal defines a region of stem cell competence in the Arabidopsis shoot meristem. Dev. Cell 24, 125–132 (2013).
Liu, L. et al. miR394 and LCR cooperate with TPL to regulate AM initiation. Nat. Commun. 15, 10156 (2024).
Lu, L. et al. miR394 enhances WUSCHEL-induced somatic embryogenesis in Arabidopsis thaliana. New Phytol. 238, 1059–1072 (2023).
Navarro-Cartagena, S. & Micol, J. L. Is auxin enough? Cytokinins and margin patterning in simple leaves. Trends Plant Sci. 28, 54–73 (2023).
Panchy, N., Lehti-Shiu, M. & Shiu, S. H. Evolution of gene duplication in plants. Plant Physiol. 171, 2294–2316 (2016).
Xu, C., Luo, F. & Hochholdinger, F. LOB domain proteins: beyond lateral organ boundaries. Trends Plant Sci. 21, 159–167 (2016).
Fan, M., Xu, C., Xu, K. & Hu, Y. LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res. 22, 1169–1180 (2012).
Omary, M. et al. A conserved superlocus regulates above- and belowground root initiation. Science 375, eabf4368 (2022).
Matsumura, Y., Iwakawa, H., Machida, Y. & Machida, C. Characterization of genes in the ASYMMETRIC LEAVES2/LATERAL ORGAN BOUNDARIES (AS2/LOB) family in Arabidopsis thaliana, and functional and molecular comparisons between AS2 and other family members. Plant J. 58, 525–537 (2009).
Nakayama, H., Leichty, A. R. & Sinha, N. R. Molecular mechanisms underlying leaf development, morphological diversification, and beyond. Plant Cell 34, 2534–2548 (2022).
Wu, L. Y. et al. Dynamic chromatin state profiling reveals regulatory roles of auxin and cytokinin in shoot regeneration. Dev. Cell 57, 526–542 e527 (2022).
Wang, F. X. et al. Chromatin accessibility dynamics and a hierarchical transcriptional regulatory network structure for plant somatic embryogenesis. Dev. Cell 54, 742–757 e748 (2020).
Zhang, T. Q., Chen, Y., Liu, Y., Lin, W. H. & Wang, J. W. Single-cell transcriptome atlas and chromatin accessibility landscape reveal differentiation trajectories in the rice root. Nat. Commun. 12, 2053 (2021).
Youngstrom, C., Wang, K. & Lee, K. Unlocking regeneration potential: harnessing morphogenic regulators and small peptides for enhanced plant engineering. Plant J. 121, e17193 (2025).
McCready, K. et al. TARGET OF RAPAMYCIN is essential for asexual vegetative reproduction in Kalanchoe. Plant Physiol. 189, 248–263 (2022).
Yang, X. et al. The Kalanchoe genome provides insights into convergent evolution and building blocks of crassulacean acid metabolism. Nat. Commun. 8, 1899 (2017).
Delaux, P. M. et al. Reconstructing trait evolution in plant evo-devo studies. Curr. Biol. 29, R1110–R1118 (2019).
Vlad, D. et al. Leaf shape evolution through duplication, regulatory diversification, and loss of a homeobox gene. Science 343, 780–783 (2014).
Shuai, B., Reynaga-Pena, C. G. & Springer, P. S. The lateral organ boundaries gene defines a novel, plant-specific gene family. Plant Physiol. 129, 747–761 (2002).
Garces, H. & Sinha, N. Transformation of the plant Kalanchoe daigremontiana using Agrobacterium tumefaciens. Cold Spring Harb. Protoc. 2009, pdb prot5303 (2009).
Franco-Zorrilla, J. M. et al. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 39, 1033–1037 (2007).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Sun, H. J., Uchii, S., Watanabe, S. & Ezura, H. A highly efficient transformation protocol for Micro-Tom, a model cultivar for tomato functional genomics. Plant Cell Physiol. 47, 426–431 (2006).
Zhang, T. Q. et al. An intrinsic microRNA timer regulates progressive decline in shoot regenerative capacity in plants. Plant Cell 27, 349–360 (2015).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat. Commun. 11, 1432 (2020).
Hu, J. et al. NextDenovo: an efficient error correction and accurate assembly tool for noisy long reads. Genome Biol. 25, 107 (2024).
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Zhang, X., Zhang, S., Zhao, Q., Ming, R. & Tang, H. Assembly of allele-aware, chromosomal-scale autopolyploid genomes based on Hi-C data. Nat. Plants 5, 833–845 (2019).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Alonge, M. et al. RaGOO: fast and accurate reference-guided scaffolding of draft genomes. Genome Biol. 20, 224 (2019).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Bruna, T., Hoff, K. J., Lomsadze, A., Stanke, M. & Borodovsky, M. BRAKER2: automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genom. Bioinform. 3, lqaa108 (2021).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Bruna, T., Lomsadze, A. & Borodovsky, M. GeneMark-ETP significantly improves the accuracy of automatic annotation of large eukaryotic genomes. Genome Res. 34, 757–768 (2024).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, W609–W612 (2006).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Zhang, C., Rabiee, M., Sayyari, E. & Mirarab, S. ASTRAL-III: polynomial time species tree reconstruction from partially resolved gene trees. BMC Bioinformatics 19, 153 (2018).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: a resource for timelines, timetrees, and divergence times. Mol. Biol. Evol. 34, 1812–1819 (2017).
Sun, P. et al. WGDI: a user-friendly toolkit for evolutionary analyses of whole-genome duplications and ancestral karyotypes. Mol. Plant 15, 1841–1851 (2022).
Chen, Y. et al. A collinearity-incorporating homology inference strategy for connecting emerging assemblies in the Triticeae tribe as a pilot practice in the plant pangenomic era. Mol. Plant 13, 1694–1708 (2020).
Tang, H. et al. JCVI: a versatile toolkit for comparative genomics analysis. Imeta 3, e211 (2024).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience https://doi.org/10.1093/gigascience/giab008 (2021).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Meng, X.-R et al. Genome files: unravelling the predominant genetic paths for asexual reproduction in Kalanchoe. Figshare https://doi.org/10.6084/m9.figshare.29109305 (2025).
Acknowledgements
We thank J.-W. Wang and L. Xu (CAS Center for Excellence in Molecular Plant Sciences, Institute of Plant Physiology and Ecology, Chinese Academy of Sciences) for discussion; F. Cheng (Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences) for Zunla-1 seeds; PhD candidate G. Yang from our laboratory for assisting in tomato genetic transformation experiments; members from J.-W.W. lab for technical assistance; Y. Ma (Central laboratory of College of Horticulture, Nanjing Agricultural University) for assistance in using Sony MA900 cell sorter and ZEISS Axio Zoom V16; and H. Ma (Modern Agricultural Analysis and Testing Center of Nanjing Agricultural University) for help in using Olympus SpinSR. This work was supported by grants from the Natural Science Foundation of Jiangsu Province (BK20230995 to T.-Q.Z.), Fundamental Research Funds for the Central Universities (KJYQ2025021 and KJYQ2024025 to T.-Q.Z.), the Entrepreneurship and Innovation Doctoral Talent of Jiangsu Province (JSSCBS20220379 to T.-Q.Z.; JSSCBS20220360 to Y.-J.C.) and National Key Research and Development Program of China (2023YFF1000501 to T.-Q.Z.).
Author information
Authors and Affiliations
Contributions
T.-Q.Z. designed the research. X.-R.M. performed genome assembly and most bioinformatic analysis. Q.-Q.W. generated transgenic plants and conducted shoot regeneration experiments. S.-L.Z. performed the plasmid construction, sampling collection, ATAC-seq library construction and confocal imaging. J.-L.W. optimized the DNA and RNA extraction methods for plantlets. C.-Z.Q., J.Y., Y.Z., and Y.-J.C. contributed to the collection and cultivation of plants. Z.-G.X. and J.-Y.X. provided suggestions on genome assembly. Y.-X.M. and Y.-J.C. contributed to technical help of the flow cytometer. Z.-Y.C., Y.-J.C., J.-Y.X. and Y.L. contributed to MS discussion. T.-Q.Z., X.-R.M., Q.-Q.W. and S.-L.Z. analysed the data. T.-Q.Z. wrote the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Xian Sheng Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Collection of Kalanchoe germplasm resources and genome size and ploidy analysis.
Systematically collected 43 Kalanchoe species with different plantlet forms and capacities; 9 species with differences in genome size, ploidy, and plantlet formation were displayed. Genome size and ploidy of Kalanchoe plants were estimated by flow cytometry using the A. thaliana genome size as a reference. Scale bar, 5.0 cm.
Extended Data Fig. 2 Karyotyping of K. daigremontiana and K-mer estimation of genome size for the three Kalanchoe species.
a, Root tissue cells of K. daigremontiana used for karyotyping to examine ploidy. K. daigremontiana is inferred to be diploid karyotype (2 N = 2X = 36). A representative image is shown from eight similar observations (n = 8). Scale bar, 5.0 μm. b-d, Genome survey estimation of genome size and heterozygosity for the three species. K-mer = 21 was used.
Extended Data Fig. 3 Assembly and annotation of chromosome-level reference genomes for three Kalanchoe species.
a, Heatmap showing Hi-C interaction signals for K. daigremontiana. b, Synteny between the assembled K. daigremontiana and K. fedtschenkoi genomes. c, Synteny between the assembled K. daigremontiana and K. marnieriana genomes. d, BUSCO assessment of genome annotation quality for the three species. e, Evaluation of K. daigremontiana genome assembly accuracy based on read mapping distribution across chromosomes.
Extended Data Fig. 4 Analysis of chromosomal collinearity and chromatin structure among three Kalanchoe species.
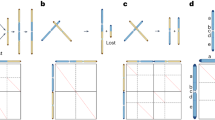
a, Genome-wide synteny features within each of the three species, and between them and V. vinifera or C. japonicum. Lines between chromosomes indicate synteny, with colored lines emphasizing the 1:2 syntenic relationships between representative V. vinifera or C. japonicum chromosomes and Kalanchoe chromosomes. b, d, f, Chromosomal collinearity analysis to identify chromosomal fragments derived from WGD events in K. daigremontiana (b), K. fedtschenkoi (d), and K. marnieriana (f). c, e, g, Overview of assembly completeness and structural characteristics in each chromosome. Chr03, Chr05, Chr06, Chr08, Chr09, Chr10 and Chr16 are gap-free assemblies containing centromeres in K. daigremontiana (c); Chr05, Chr07, Chr08, Chr09, Chr13, Chr14, Chr15, Chr16, Chr17, and Chr18 are gap-free assemblies containing centromeres in K. fedtschenkoi (e); Chr04, Chr05, Chr07, Chr08, Chr09, Chr10, Chr11, Chr13, Chr14, and Chr16 are gap-free assemblies containing centromeres in K. marnieriana (g).
Extended Data Fig. 5 Phylogeny and synonymous substitution rate analysis for the three Kalanchoe species.
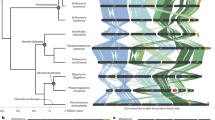
a, b, Phylogenetic relationships of the three Kalanchoe species within angiosperms inferred using two phylogenetic methods. (a) Concatenation-based method, numbers at divergence nodes indicate SH-aLRT support and bootstrap support percentages (1000 replicates). (b) Coalescent-based method, numbers at divergence nodes indicate quartet support for the main topology, the first alternative topology and the second alternative topology. c-e, Synteny analysis within chromosomes of each species, with different colors indicating Ks values. Circles highlight representative syntenic blocks between duplicated chromosomes, for example, between Chr04 and Chr07, Chr09 and Chr15.
Extended Data Fig. 6 Analysis of gene copy number and functional variation for key embryogenesis-related genes in Kalanchoe.
a, BBM maintains a single copy in all three species; no obvious variations in protein length or key domains were found. b, WUS maintains a single copy in K. daigremontiana and K. marnieriana, with 2 copies in K. fedtschenkoi; no obvious variations in protein length or key domains were found. c, AGL15 and its paralog AGL18 maintain a single copy in all three species; no obvious variations in protein length or key domains were found. d, ABI3 has 2 copies in all three species; no obvious variations in protein length or key domains were found. e, WOX2 has 2 copies in K. daigremontiana, and a single copy in K. fedtschenkoi and K. marnieriana; and WOX4 maintains a single copy in all three species; no obvious variations in protein length or key domains were found. f, LEC1 maintains a single copy in K. daigremontiana and K. marnieriana, with 2 copies in K. fedtschenkoi; however, LEC1 protein is truncated, and key domains are disrupted in K. daigremontiana and K. fedtschenkoi. L1L maintains a single copy in all three species, with no obvious variations in protein length or key domains. g, SERK1 and SERK2 maintain a single copy in all three species; no obvious variations in protein length or key domains were found. h, FUS3 has 3 copies in all three species; one copy encodes a novel protein domain, suggesting potential neofunctionalization. i, STM has 2 copies in all three species; no obvious variations in protein length or key domains were found. j, AHL15 has 2 copies in all three species; no obvious variations in protein length or key domains were found.
Extended Data Fig. 7 Temporal transcriptome analysis of plantlet formation.
a, Plantlet formation was divided into four representative stages based on the developmental gradient from leaf tip to leaf base. b, Tissue sections and toluidine blue staining analyzing the anatomical structures of plantlet formation at different stages. Yellow arrows and dashed lines identify nascent or forming plantlets. Each representative image is shown based on ten similar observations (n = 10). Scale bar, 100 μm. c, PCA plot showing transcriptomic differences among K. daigremontiana plantlet samples at four different stages. Please note that S4 and S3 show minor differences in PCA; to be consistent with K. fedtschenkoi and K. marnieriana, transcriptome data from S1 to S3 of K. daigremontiana were used for downstream analysis. d, PCA plot showing transcriptomic differences among K. fedtschenkoi plantlet samples at three different stages. e, PCA plot showing transcriptomic differences among K. marnieriana plantlet samples at three different stages. f, Numbers of up-regulated and down-regulated differentially expressed genes (DEGs) between different plantlet developmental stages.
Extended Data Fig. 8 Observing the dynamic plantlet formation in Kalanchoe.
a, Relative spatial positions of plantlet formation in leaves of K. daigremontiana, K. fedtschenkoi, and K. marnieriana. Plantlets of K. daigremontiana and K. fedtschenkoi are formed at leaf serrations, with K. daigremontiana generating a leaf-pedestal structure for attaching the plantlets. Plantlets of K. marnieriana form in the upper epidermis near the leaf margin. b-d, Tissue section and confocal imaging showing the dynamic plantlet formation (relative stages, S0 to S4) in K. daigremontiana, K. fedtschenkoi, and K. marnieriana, respectively. T represents the inferred totipotent cells of plantlets, V indicates the vascular tissue of the mother leaf, and SAM denotes the shoot apical meristem. The green dashed line indicates the outline of the leaf serration (b, c) or leaf margin (d). Please note that due to the narrow width between serrations on the leaves at early stage, the maximum projection imaging includes some leaf outlines from non-serration regions (b). M, C, U, and L respectively indicate the direction towards the leaf margin, leaf center, upper epidermis, and lower epidermis, and the yellow-colored arrow line represents the axial direction of plantlet formation. Each representative image is shown based on ten similar observations (n = 10). Scale bar, 100 µm.
Extended Data Fig. 9 Expression patterns of representative pluripotency-associated genes during plantlet formation.
a, Expression patterns of WOX gene family in the three species during plantlet formation. b, Expression patterns of KNOX gene family in the three species during plantlet formation. c, Expression patterns of key somatic embryogenesis-related genes in the three species during plantlet formation. d, Expression patterns of key shoot organogenesis-related genes in the three species during plantlet formation. e, Expression patterns of multiple ATH1 copies in the three species during plantlet formation. f, Expression patterns of class Ia, class Id, class Ie, and class II LBD genes in the three species during plantlet formation. Black blocks indicate gene absence; gray blocks indicate no detectable expression.
Extended Data Fig. 10 Genome-wide analysis of chromatin accessibility features in leaves of plantlet and non-plantlet plants.
a, K-means clustering analysis and heatmap showing differentially accessible chromatin regions during plantlet formation. Representative genes within each peak cluster are highlighted. b, GO enrichment analysis of biological processes associated with genes corresponding to K3-K8 peak clusters. K5 and K6 are significantly enriched for biological pathways related to embryonic development and meristematic activity regulation. Adjusted P values resulting from a one-sided hypergeometric test with subsequent Benjamini-Hochberg correction are shown. c, Volcano plot showing differential peaks between immature embryos and differentiated leaves (1st leaf) in A. thaliana. Peaks upregulated ≥ 2-fold, corresponding to 3,538 genes, were defined as highly accessible genes of embryogenesis (HAG-embryogenesis). d, GO enrichment analysis of biological pathways for the 3,538 HAG-embryogenesis genes. Adjusted P values resulting from a one-sided hypergeometric test with subsequent Benjamini-Hochberg correction are shown. e, Comparative analysis of chromatin openness intensity for HAG-embryogenesis genes versus background genes in immature embryo and shoot apex tissues in A. thaliana. f, Volcano plot showing differential peaks between shoot apices and differentiated leaves (1st leaf) in A. thaliana. Peaks upregulated ≥ 2-fold, corresponding to 1,421 genes, were defined as highly accessible genes of organogenesis (HAG-organogenesis). g, GO enrichment analysis of biological pathways for the 1,421 HAG-organogenesis genes. Adjusted P values resulting from a one-sided hypergeometric test with subsequent Benjamini-Hochberg correction are shown. h, Comparative analysis of chromatin openness intensity for HAG-organogenesis genes versus background genes in immature embryo and shoot apex tissues in A. thaliana. Regions encompassing the coding sequence, upstream -3.0 Kb, and downstream 3.0 Kb of each gene were calculated. TSS: Transcription Start Site, TES: Transcription End Site. The coding region is shaded yellow.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–12, Text, Methods, source data for Figs. 10 and 12 and references.
Supplementary Table 1 (download XLSX )
Genome assembly and annotation.
Supplementary Table 2 (download XLSX )
Genomic data and orthologues used for evolutionary analysis in this study.
Supplementary Table 3 (download XLSX )
Calculations of HGVI in Kalanchoe.
Supplementary Table 4 (download XLSX )
RNA-seq analysis for plantlet formation.
Supplementary Table 5 (download XLSX )
Intersection of highly expanded genes and C8 cluster genes.
Supplementary Table 6 (download XLSX )
ATAC-seq analysis for plantlet formation.
Supplementary Table 7 (download XLSX )
Highly accessible genes of embryogenesis.
Supplementary Table 8 (download XLSX )
Highly accessible genes of organogenesis.
Supplementary Table 9 (download XLSX )
CAFE analysed gene expansion and contraction in Kalanchoe.
Supplementary Table 10 (download XLSX )
Structural variation in Kalanchoe.
Supplementary Table 11 (download XLSX )
The absence and presence of LCR in 545 Viridiplantae plant genomes.
Supplementary Table 12 (download XLSX )
Primers used in this study.
Source data
Source Data Figs. 1, 3, 4 and 6 (download XLS )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Meng, XR., Wang, QQ., Zhu, SL. et al. Unravelling the predominant genetic paths for asexual reproduction in Kalanchoe. Nat. Plants 12, 369–385 (2026). https://doi.org/10.1038/s41477-025-02214-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02214-3