Abstract
Excessive dieting (ED), a common weight-control strategy, often causes neurological and emotional disturbances, yet its gut-brain interaction mechanisms remain unclear. Employing a short-term dietary (SDR) adult male rabbit model, we found that SDR can induced cerebral cortex up-regulation of the immune-related genes (e.g., C1QC, SAA3) enriched in NF-kappa B signaling pathways, contrasted with down-regulation of sex hormone-related genes (e.g., PRLR, SPA17) implicated in metabolic homeostasis. Furthermore, dysregulated expression of metabolic genes (e.g., PPM1J, GALNT18) in the cecum of the SDR group interacted to impair the immune protection pathways related to intestinal mucosa. Then, SDR significantly increased the cecal Firmicutes/Bacteroidetes ratio (from 3.38 to 5.57) and reduced microbial diversity. Specifically, beneficial bacteria involved in tryptophan metabolism and neurotransmitter synthesis (e.g., Bacteroidales_bacterium, Alistipes_indistinctus) decreased, whereas bile acid-metabolizing bacteria (e.g., Clostridium_sp._CAG:710, Ruminococcus_sp._Marseille-P6503) linked to increase energy metabolism. The top 20 genes from the brain-gut axis analysis (e.g., ITPR1, CAMK4, CDK5R1) were enriched in critical neural pathways like axon guidance, GABAergic synapse, and long-term potentiation. Notably, key neurodevelopmental genes (e.g., GPR37, GPX3) correlated with these microbial shifts, implicating oxidative stress, synaptic plasticity, and mitochondrial function in microbiota-host crosstalk. This study highlights a “microbial-metabolism-neural” axis in SDR, providing novel targets for future obesity intervention strategies.
Similar content being viewed by others
Introduction
Weight loss has always been a main topic of concern for middle-aged and elderly obese people. It can not only reduce body fat but also effectively lower the risk of obesity-related metabolic diseases, such as cardiovascular and cerebrovascular diseases, type 2 diabetes (T2D), Alzheimer’s disease (AD), hypertension and hyperlipidemia1. For obesity intervention methods, compared with drug intervention, dietary intervention has become a non-dependent way to lose weight. Studies have indicated that restrictive diets, involving controlled nutrient intake, fasting, and intermittent eating, can serve as a primary intervention strategy for obesity in its early stages2. These measures mainly aim at controlling the intake of nutrients but are prone to rebound and may involve multiple physiological metabolic mechanisms, such as diet-induced immune cells in adipose tissue, mitochondrial metabolic remodeling, calcium ion signal transformation and substrate circulation, as well as increased activity of the sympathetic nervous system. In these physiological processes, various dietary patterns impact gut flora and its metabolites, which in turn stimulate the vagus nerve to regulate brain metabolism directly or indirectly. This regulation influences internal energy balance and central appetite reward signals in the metabolic processes of obesity3. Prior research indicates that Leptin (LEP), ghrelin, and glucagon-like peptide 1 (GLP-1) in the brain-gut axis directly control appetite, affecting the metabolic interactions between the gut and the microbial community, thereby influencing feeding behavior4. These effects may be linked to intestinal host genes and cerebral cortex metabolic patterns influenced by intestinal microbial metabolism. Nevertheless, the precise interactive mechanisms between host gene regulation in the brain-gut axis and gut microbial population metabolism, which impact nutrient digestion and absorption, remain incompletely understood.
Limiting nutrient intake is a relatively effective way to lose weight. However, we need a high-quality weight loss that enhances metabolic function and promotes physical health. Dietary restriction, involving fasting or severe caloric reduction for 12–48 h alternated with regular food intake, is recognized as an effective approach for weight loss and lifespan extension by reducing nutrient intake5. In some diverse dietary pattern studies6, it has been indicated that very low-carb ketogenic diets (VLCKD), high-fiber diets, and intermittent fasting effectively alleviate obesity-related inflammation and reduce body fat. Weight loss through high-protein calorie restriction diets leads to distinct alterations in gut microbiome composition, including increases in Akkermansia sp. and Bifidobacterium sp. and reductions in Prevotella sp7. However, the precise connections between brain energy homeostasis and gut microbiota under various dietary patterns, particularly restricted diets, remain incompletely understood. Recent researches have demonstrated that dietary obesity can induce dysregulation of neuron transcription in the cerebral cortex of mice, involving excitatory neurons and glial cells, particularly microglia8,9. Conversely, studies have shown that intermittent fasting can shift brain energy metabolism patterns, enhance mitochondrial activity in hippocampal neurons, regulate intestinal microbial circulation to generate beneficial metabolites such as 3-indolepropionic acid, serotonin, short-chain fatty acids, or tauroursodeoxycholic acid, thereby conferring resistance to metabolic diseases10,11.
In randomized clinical trials12 on time-restricted eating, it was found that early time-restricted eating (eTRE) every day for 14 weeks was more effective in weight loss, improving diastolic blood pressure and mood than eating over a 12-h period. On the other hand, dietary restriction can also cause insufficient energy supply to the body, leading to irreversible emotional and cognitive problems in the metabolism of the brain-gut axis, such as depression, anxiety, and abnormal behaviors13. Recent clinical statistical analysis indicates a lack of research on the influence of intermittent fasting (IF) on brain-derived neurotrophic factor (BDNF) in healthy, overweight/obese individuals and patients with metabolic diseases, leaving uncertainty about its impact on brain cognition. Therefore, the relationship between dietary patterns and brain cognition remains unclear14. We hypothesize that the limitation of nutritional levels in various diet patterns may alter brain energy metabolism homeostasis and raise the risk of cognitive diseases.
The brain-gut axis plays a crucial role in regulating the body’s metabolic balance15,16. The gut microbiota has been found to influence immune response, intestinal function, host feeding behavior, and central nervous system (CNS) metabolism via the vagus nerve17,18. Disruption of the gut microbiota has been associated with various CNS diseases, including Parkinson’s disease (PD), Autism spectrum disorder (ASD), attention deficit hyperactivity disorder (ADHD), and Rett syndrome (RTT)19,20. Recent studies have identified specific metabolites such as bile acid, butyric acid, succinate, red acid, urinary asbestos A, and asparagine, derived from changes in intestinal microorganism abundance like Bacteroides, Ruminococcus, and Clostridium butyricum (CCFM1299), that are involved in regulating lipid metabolism signaling pathways across different body tissues, including PKA, PPARa, PGC-1a, and mTORC1 signaling pathways21,22. We should pay attention to the mechanism of brain-gut interaction under different dietary conditions, especially nutrient limitation. However, the underlying mechanisms by which gut microbiota may contribute to alterations in brain-related metabolic patterns under restrictive diet conditions are still not fully understood.
The study aims to explore transcriptional changes in lipid metabolism in the cerebral cortex caused by dietary restrictions and their potential influence on the body’s energy equilibrium. This will involve analyzing the correlation between microorganisms and the cerebral cortex in rabbits to lay the groundwork for future interventions and treatments of metabolic disorders associated with weight loss and obesity in humans.
Results
Neuropathological and transcriptional programming of the cerebral cortex in response to dietary restriction
Through the construction of a feed restriction model, we observed a significant decrease in body weight and a remarkable reduction in perirenal fat mass in the restricted group compared to the control group, as clearly illustrated in Supplementary Fig. 1. However, pathological examination with HE staining showed no obvious inflammatory response in the brain-intestinal tissues among each group, as shown in Supplementary Fig. 2. To better explain the interaction between restricted diet induced intestinal microbiome in the cecum and the cerebral cortex, the transcriptional differences between the two groups in the left cerebral cortex tissues of rabbits were measured. The transcriptomic sequencing of the cerebral cortex tissue samples between Co and Re showed that the clean bases data amount of each sample was distributed in 13.13–14.92G, with a Q30 base and an average GC content proportion of 93.45–94.97% and 51.60–53.04%, respectively. Then, the number of coding genes in each sample was mapped, analyzed and ranged from 17,139 to 17,240, as shown in Supplementary Table 1. By comparing the expression levels of genes between the two groups, a total of 370 significantly differentially expressed genes were identified from 17,191 genes using the screening criteria of ∣log₂FC∣ ≥ 1 and p-value < 0.05, among which 275 were up-regulated and 95 were down-regulated, such as RPS29, TMEM256, C1QC, LGALS3, etc, as shown in Fig. 1A, B. Enrichment analysis showed the top 30 GO enrichment terms (Fisher’s exact test, q < 0.05) of these DEGs in the following: biological processes, including immune response, regulation of immune response, adaptive immune response, chemotaxis, and innate immune response, etc; cellular component, including the external side of plasma membrane, cell surface, plasma membrane, immunological synapse, integral component of membrane, etc; molecular functions, including antigen binding, C-C chemokine receptor activity, folic acid binding, immunoglobulin receptor binding, signaling receptor activity, etc, as shown in Fig. 1C. Moreover, KEGG pathway enrichment analysis (Fig. 1D) showed that up-regulated DEGs were mainly enriched in pathways (Fisher’s exact test, q < 0.05), such as the Intestinal immune network for IgA production, Viral myocarditis, Hematopoietic cell lineage, Primary immunodeficiency, Cell adhesion molecules (CAMs), Epstein-Barr virus infection, Asthma, Allograft rejection, Cytokine-cytokine receptor interaction, and the NF-kappa B signaling pathway. On the other hand, the downregulated DEGs were mainly enriched in pathways (Fisher’s exact test, q < 0.05), such as Malaria, Bile secretion, Complement and coagulation cascades, ECM-receptor interaction, Glycine, serine and threonine metabolism, and Staphylococcus aureus infection. Furthermore, the expression levels and interaction relationships of these inflammatory factors indicated that CD40, CD40LG, SYK, TNFSF13B and TRAF4 played significant regulatory roles in the protein interaction network from STRING database analysis, as shown in Fig. 1E, F, and Supplementary Table 2, 3. Therefore, we speculated that the dietary restriction could induced a deceleration in amino acid metabolism and an elevation in the transcriptional levels of inflammatory factors in rabbit cerebral cortex tissue.
A Volcano plot analysis of DEGs between the two groups. The gray color represents the non-significant DEGs, and the red and blue colors represent the significant DEGs. The horizontal axis shows the log2FC, and the vertical axis shows the −log10 P-value. B Cluster heatmap analysis of significantly DEGs within different groups of samples. In the figure, each small square represents a gene, and the color of each square indicates the expression level of the corresponding gene. Red color represents highly expressed genes while blue color indicates lowly expressed genes. The dendrogram on the left side of the figure shows the clustering analysis results of different genes from various samples. C Enrichment plot of GO analysis for DEGs. In the bar chart, filter out the GO entries corresponding to the three categories whose number of differential genes is greater than or equal to 3. Sort the top 10 entries for each GO entry according to the −log10 p-value from small to large, and display them in the bar chart. D Enrichment analysis of KEGG pathways involved by the DEGs. The Y-axis in the graph corresponds to the names of pathways; The X-axis represents the ratio of DEGs in a specific pathway to the total number of genes in that pathway. A higher value indicates a greater proportion of DEGs in the pathway; The bubbles on the plot represent the number of DEGs-the larger the bubble, the more DEGs are present in that particular pathway. The significantly enriched GO terms and KEGG pathways (Fisher’s exact test, q < 0.05) in (C) and (D) are shown in “Result” section. E Line plot analysis of the differences in mRNA expression levels of immune-related genes in the cerebral cortex between the two groups. The horizontal axis represents the sample names, and the vertical axis represents log2 expression. Different colors and shapes in the graph represent the differences in gene expression levels. F Correlation network diagram of immune-related genes in the cerebral cortex between the two groups. The size of the dots in the figure represents the importance of the gene in the network. Red dots indicate that the gene is upregulated, while green dots indicate that the gene is downregulated. The color of the lines connecting the two points ranges from yellow to red, indicating an increasing correlation.
Differential host gene analysis of the cecal tissues between Co and Re
The transcriptomic sequencing of the cecal samples between Co and Re showed that the clean bases data amount of each sample was distributed in 6.89–7.35 G, which contained the Q30 base (distributed in 93.99–94.32%) and an average GC content of 53.13%. Each sample was aligned to the reference genome (GCF_009806435.1). It was found that the number of coding genes in each sample ranged from 15,377 to 15,953, as shown in Supplementary Table 4. By comparing the expression levels of genes between the two groups, 1603 significant DEGs (DEGs) (218 upregulated and 1385 downregulated) were selected a total of from 15,777 DEGs under the criteria of ∣log₂FC∣ ≥ 1 and p-value < 0.05, as shown in Fig. 2A. The top 20 genes were selected from the DEGs and a cluster heatmap was drawn for subsequent analysis, such as PLA2G4C, CD79B, BAC2, APOE, etc, as shown in Fig. 2B. Enrichment analysis showed the top 10 GO enrichment terms of these DEGs (Fisher’s exact test, q < 0.05) (Fig.2C) in the following: biological processes, including B cell activation, positive regulation of T cell proliferation, adaptive immune response, inflammatory response, and immune response; cellular component, including integral component of lumenal side of endoplasmic reticulum membrane, clathrin-coated endocytic vesicle membrane, MHC class II protein complex, integral component of plasma membrane, and the plasma membrane; molecular functions, including MHC class II protein complex binding, SH3/SH2 adapter activity, peptide antigen binding, tumor necrosis factor receptor binding, and MHC class II receptor activity. KEGG pathway enrichment analysis (Fig. 2D) showed that the upregulated DEGs were mainly involved in pathways (Fisher’s exact test, q < 0.05), such as Drug metabolism—other enzymes, Ascorbate and aldarate metabolism, Pentose and glucuronate interconversions, Ether lipid metabolism, Porphyrin and chlorophyll metabolism, Legionellosis, Tight junction, p53 signaling pathway, Drug metabolism-cytochrome P450, and Steroid hormone biosynthesis. On the other hand, the downregulated DEGs (Fisher’s exact test, q < 0.05) were mainly enriched in pathways, such as Hematopoietic cell lineage, Asthma, Cell adhesion molecules (CAMs), Allograft rejection, Intestinal immune network for IgA production, Rheumatoid arthritis, Autoimmune thyroid disease, Viral myocarditis, NF-kappa B signaling pathway, and Cytokine-cytokine receptor interaction. Therefore, it was speculated that the dietary restriction potentially enhanced regulators of non-carbohydrate conversion while suppressing cecal immune responses.
A Volcano plot analysis of DEGs between the two groups. The gray color represents the non-significant DEGs, and the red and blue colors represent the significant DEGs. The horizontal axis shows the log2FC (Fold change), and the vertical axis shows the −log10 P-value. B Cluster heatmap analysis of significantly DEGs within different groups of samples. In the figure, each small square represents a gene, and the color of each square indicates the expression level of the corresponding gene. Red color represents highly expressed genes, while blue color indicates lowly expressed genes. The dendrogram on the left side of the figure shows the clustering analysis results of different genes from various samples. C Enrichment plot of GO analysis for DEGs. In the bar chart, filter out the GO entries corresponding to the three categories whose number of differential genes is greater than or equal to 3. Sort the top 10 entries for each GO entry according to the −log10 p-value from small to large, and display them in the bar chart. D Enrichment analysis of KEGG pathways involved by the DEGs. The Y-axis in the graph corresponds to the names of pathways; The X-axis represents the ratio of DEGs in a specific pathway to the total number of genes in that pathway. A higher value indicates a greater proportion of DEGs in the pathway; The bubbles on the plot represent the number of DEGs—the larger the bubble, the more DEGs are present in that particular pathway. The significantly enriched GO terms and KEGG pathways (Fisher’s exact test, q < 0.05) in (C) and (D) are shown in “Result” section.
Cross-tissue gene co-expression network analysis between the cecal and cerebral cortical transcriptomes
To better understand the degree of correlation in transcriptional differences between the two organ tissues, Venn diagram analysis was performed on the transcriptional differences in the cerebral cortex and cecum tissues between the Re and Co, as shown in Fig. 3A. The results showed that 15,298 differential genes (DEGs) were co-expressed in the two organs. Then, weighted gene co-expression network analysis (WGCNA) results showed that these DEGs were divided into 26 modules, as shown in Fig. 3B. The module correlation coefficient analysis (Fig. 3C) showed that the module correlation coefficient of MEpurple gene was positively correlated with the cerebral cortex samples (R > 0.5, p < 0.05), but negatively correlated with cecal samples (R < −0.59, p < 0.05). Therefore, we selected the modular gene cluster with the highest co-expression correlation. Then, enrichment analysis showed the top 30 GO enrichment terms (Fisher’s exact test, q < 0.05) of these module gene clusters (6542 DEGs) as follows (Fig. 2D): biological processes, including regulation of synaptic vesicle exocytosis, nervous system development, chemical synaptic transmission, signal transduction, homophilic cell adhesion via plasma membrane adhesion molecules, etc; cellular component, including glutamatergic synapse, cell junction, neuronal cell body, axon, dendrite, postsynaptic density, neuron projection, etc; molecular functions, including ion channel binding, SNARE binding, ubiquitin-protein transferase activity, syntaxin-1 binding, voltage-gated potassium channel activity, etc. The top 20 KEGG pathways (Fisher’s exact test, q < 0.05) showed that Axon guidance, Synaptic vesicle cycle, GABAergic synapse, Long-term potentiation and other pathways were the main pathways, as shown in Fig. 3E. These pathways are related to neural activity and were chosen for our research key pathways. We conducted co-expression correlation analysis on the significantly DEGs (P < 0.05) involved in these pathways between the two organ tissue samples. We screened out the top 20 genes with strong differences in co-expression coefficients as the key genes for our subsequent research, such as ITPR1, CAMK4, ATP6V1G2, PRKCB, CDK5R1, PLA2G4C, ATF2 and GPX3, as shown in Fig. 3F and Table 1. Then, we hypothesized that the dietary restriction induced remodeling of energy metabolism in the gut-brain axis, particularly through co-expression of neuromodulatory factors that regulate metabolic balance.
Note: A Analysis of cecal and cortical cross differential genes by Venn diagram. The red circle represents the differential genes in cerebral cortex tissue and the blue circle represents the differential genes in cecum tissue. B Construct a hierarchical clustering tree based on correlation coefficients of DEGs. Different branches of the clustering tree represent distinct gene modules, with each color indicating a separate module. By utilizing weighted correlation coefficients of genes, genes are classified according to their expression patterns, grouping those with similar patterns into the same module. C Analysis of associations between different modules and tissue samples. In the figure, different colors represent different gene modules. The small squares within each module correspond to the module genes of the samples. The values in the upper part of the module represent the correlation coefficients between the expression levels of the module genes and the samples within the corresponding sample group squares. The P-value for the significance test of differential expression of each gene among different sample groups is calculated using the T-test (Pearson correlation coefficient). D Enrichment plot of GO analysis for the module genes. In the bar chart, filter out the GO entries corresponding to the three categories whose number of differential genes is greater than or equal to 3. Sort the top 10 entries for each GO entry according to the -log10 p-value from small to large, and display them in the bar chart. E Enrichment analysis of KEGG pathways involved by the module genes. The y-axis in the graph corresponds to the names of pathways; The X-axis represents the ratio of DEGs in a specific pathway to the total number of genes in that pathway. A higher value indicates a greater proportion of DEGs in the pathway; The bubbles on the plot represent the number of DEGs-the larger the bubble, the more DEGs are present in that particular pathway. The significantly enriched GO terms and KEGG pathways (Fisher’s exact test, q < 0.05) in (D) and (E) are shown in “Results” section. F The figure displays a co-expression correlation heatmap analysis of neuro-metabolic pathway genes within modular gene clusters. Genes on the y-axis are derived from cecal tissue samples, while those on the x-axis are from cerebral cortex tissue samples. The small squares within the plot depict the correlation patterns of gene expression levels between the tissue samples: red signifies a positive correlation, blue indicates a negative correlation, and darker shades reflect stronger correlations. Statistically significant Pearson correlations are denoted as follows: *p < 0.05; **p < 0.01.
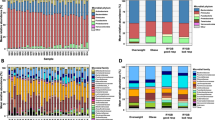
Linking dietary restriction-induced gut microbial shifts to host transcriptional responses in the cecum
Based on the clean read data and genome assembly, ORF prediction was performed between Co and Re, resulting in 20,685,283 ORF numbers (>200 bp). The results are shown in Supplementary Table 5. By comparing each sample’s clean reads with a set of non-redundant genes, a total of 9,481,157 unigenes were predicted and found in the intergroup samples. The number of genes in the Re samples was significantly higher than that in the Co samples, as shown in Fig. 4A. Similarly, anosim analysis was conducted based on the gene clusters in the inter-group samples. The distribution of gene cluster abundance in each sample between Re and Co was analyzed using the Anosim method (Fig. 4B). The gene clusters in the Re group were significantly lower than those in the Co group in this experiment (R = 0.37, P < 0.01), confirming that the differences between the experimental groups were significant and had biological significance. Moreover, species annotation was obtained through analysis of the NR library, and its abundance was calculated using the total gene abundance corresponding to the species. The results of microbial composition analysis showed that Firmicutes, Bacteroidetes, Proteobacteria, Verrucomicrobia, Tenericutes, Candidatus_Melainabacteria, Actinobacteria, Spirochaetes, Euryarchaeota and Fusobacteria were the most abundant bacterial phyla between the two groups, as shown in Fig.4C, and found that the Firmicutes/Bacteroidetes (F/B) ratio increased from 3.38 in the Co to 5.57 in the Re. At the genus level, Clostridia, Clostridiales, Lachnospiraceae, Ruminococcaceae, Bacilli, Bacteroidales, Firmicutes, Porphyromonadaceae, Akkermansia glycaniphila, and Clostridiaceae were the most abundant bacterial genera in the two groups, as shown in Fig. 4D. The microbial diversity index analysis found that the Goods coverage index of the two groups was higher than 98%, indicating a high microbial detection rate in all samples. The Chao1 and ACE indices in the Re were higher than those in the Co, indicating that the abundance of microbial species in the Re was higher than that in the Co. The Simpson index and Shannon index showed no significant difference between groups, but two index value of microflora in Re were lower than that in Co, suggesting that the diversity of microbial species in the Re had slightly decreased, as shown in Table 2. The results of species difference between the two groups were analyzed using the Wilcoxon rank-sum method and showed that the relative abundances of the top 10 microbes (P < 0.05) at the species level had significant differences between the two groups, as shown in Fig. 4E. The relative abundance of Bacteroidales_bacterium, Alistipes_indistinctus, Duncaniella_freteri, Bacteroides_ovatus, and Flavobacteriales_bacterium was reduced, while the relative abundance of Clostridiaceae_bacterium, Clostridium_sp._CAG:710, Clostridium_sp._CAG:452, Ruminococcaceae_bacterium_FB2012, Ruminococcus_sp._Marseille−P6503) were increased. The above significant differences information on the top 30 genus/species were shown in Table 4. Furthermore, the LDA (Linear Discriminant Analysis) method was employed to analyze the main metabolic pathways of the differential microorganisms in Fig. 4F. The results indicated that the top 10 metabolic pathways (Fisher’s exact test, q < 0.05) were the main pathways in the RE group, including Alanine aspartate and glutamate metabolism, Peptidoglycan_biosynthesis, Glycerolipid_metabolism, etc. In contrast, the top 10 metabolic pathways in the Co group were the main pathways, including Thiamine metabolism, Vitamin B6 metabolism, and N-Glycan biosynthesis, among others. Based on the weighted co-expression analysis of cerebral cortex and cecum genes, we conducted co-expression heatmap analysis of these 10 significant species and 38 key neurometabolic pathway genes in the cecum, as shown in Fig. 4G. The results showed that 20 key genes with high correlation were screened out to have a strong correlation with these 10 species (Pearson’s correlation coefficient >0.8, p < 0.05). By constructing correlation networks, it was found that 28 genes (PRKCB, GRIN3A, RYR1, CPLX3, PSMA3, PPP1R1B, PLA2G4C, etc.) were highly correlated with 10 microbial species in the co-expression networks, as shown in Fig. 4H and Table 3. Therefore, these correlation analysis results indicated that the dietary restriction augmented the functional capacity of cecal microbiota for amino acid biosynthesis and non-glycosidic metabolite transformation, engaging in cross-talk with host genetic regulators to preserve cecal metabolic equilibrium.
A The number of genes between the two groups was analyzed by Venn diagram. The blue circle represents Re group and the purple circle represents Co group. B Anosim method analysis of the gene clusters differences between the two groups. Horizontal is the group information, vertical is the distance information. “Between” refers to the difference between two groups, and the other refers to the difference within each group. The R values generally range from 0 to 1. The R value close to 1 indicates that the difference between groups is greater than the difference within groups, and an R value close to 0 indicates that there is no significant difference between and within groups. The reliability of statistical analysis is represented by p-value, and P < 0.05 indicates statistical significance. C, D Significant differences at the phylum and genus levels, respectively. Columns represent samples, and different colors represent different annotation information. Branch points represent different samples, different colors represent different groups, and the closer the branches, the more similar the samples. E Box plots showing the top 10 significantly different species between Co and Re. The horizontal axis is sample grouping. The relative abundance of the corresponding species is shown longitudinally. F The bar chart of LDA value distribution shows the significantly enriched pathways of differential microorganisms with LDA Score greater than the set value (default set to 2). The length of the bar chart represents the influence size of the differential functional pathway (i.e., LDA Score), and the color of the bar chart represents the respective groups. G Analysis of co-expression of top 20 key genes of neurometabolic pathways in the top 10 key microbial species and cecum tissues. In the heatmap, horizontal represents different genes, vertical represents significantly different microorganisms, and cluster analysis is performed on both horizontal and vertical data. Moreover, the color transformation from red to blue in the heat map represents different correlation coefficients, from positive correlation to negative correlation. The significant differences (“*”p < 0.05, “**” p < 0.01 and “***” p < 0.001) are indicated. H Construction of co-expression networks and analysis of interaction between flora and genes. The line between the two in the network diagram goes from red to gray, indicating that the correlation goes from strong to weak. The shape of each node from large to small represents the weight of the node in the correlation network from large to small.
Alterations in cecal neurometabolites of rabbits following dietary restriction
To further clarify the metabolic relationship between the cecal microbiota and the corresponding host, we conducted a comprehensive analysis of the metabolites in the cecal contents. The PLS-DA method was used to analyze the relationship model between metabolite expression and samples between Co and Re. Its results showed that the evaluation parameters in the model were all greater than 0.7 (R2 X = 0.743, R2 Y = 0.971 and Q2 = 0.77), indicating that the samples conformed to the model conditions, as shown in Fig. 5A. Based on the select parameter values |log2 (FoldChange)| > 0, P-value < 0.05 and VIP > 1 of the metabolites, the Pearson correlation analysis was conducted between 30 key differentially expressed metabolites (DEMs) related to neuro-metabolism and 30 different bacterial species in the cecum, as shown in Fig. 5B and Tables 4–6. The results indicated that Prevotella_copri, Bacteroidales_bacterium, Muribaculaceae_Flavobacteriales_bacterium, Roseburia_inulinivorans, and Parasutterella_excrementihominis, etc. have a strong positive correlation with the synthesis of tryptophan metabolites and neurotransmitter-like metabolites, while they have a negative correlation with bile acid metabolites and ion channel-related excitatory metabolites. Conversely, the species of the Clostridium genus (Clostridium_sp._CAG:452, Clostridium_sp._CAG:417 and Clostridium_sp._CAG:451, etc.), the Firmicutes_bacterium_HGW-Firmicutes-16, the Ruminococcus genus (Ruminococcus_sp._YE71, Ruminococcus_sp._CAG:579 and Ruminococcus_sp._Marseille-P6503, etc.) showed the opposite result. Functional enrichment analysis of all the DEMs showed that they were mainly enriched in the pathways (Fisher’s exact test, q < 0.05) of Arginine biosynthesis, Vitamin digestion and absorption, Vitamin B6 metabolism, ABC transporters, FoxO signaling pathway, and Huntington disease. (Fig. 5C). Furthermore, we screened out 20 intestinal metabolites related to brain neural metabolism from the metabolites and found that gamma-aminobutyric acid (GABA) and L-tryptophan (P < 0.05) were down-regulated in the Re, while butyric acid and bile acid were significantly upregulated (P < 0.05), as shown in Fig. 5D and Table 7. Then, we drew the interaction diagram between the microorganisms in the cecum and the metabolic activities of the cerebral cortex, as shown in Fig. 5E. In the figure, we focused on predicting the relationship between restricted feeding and the secretion of L-tryptophan, deoxycholic acid, GABA and other key neuro-metabolic derivatives by the key bacterial communities in the cecum, and the functional changes in the cerebral cortex. Consequently, we postulate that modifications in the secretion of crucial neuro-metabolites by cecal microorganisms as a result of restricted feeding could enter or indirectly influence the cerebral cortex tissue through the peripheral circulation. This process is anticipated to govern its immune and metabolic functions, subsequently influencing brain cognition, anxiety, and depression, and resulting in suboptimal weight loss effects. Fig. 6.
A Partial Least Squares—Discriminant Analysis (PLS-DA) analysis of the differences in cecal metabolite samples between groups. The closer the interpretation rate R2Y (cum) and the prediction rate Q2 (cum) are to 1, the better the PLS-DA model can explain and predict the differences between the two groups of samples, representing a better predictive ability of the model. B Cluster heatmap for Pearson correlation analysis of the top 30 microbials and the top 30 metabolites profiles between two groups. In the figure, each column represents a different species, and each row represents the corresponding metabolite. Metabolites on the horizontal axis are color-coded according to functional categories: those in green denote neurotransmitter-related metabolites, red represents tryptophan derivatives, purple indicates neuropeptide analogs, and black corresponds to neuroexcitatory substances. Orange-red indicates a positive correlation, while blue indicates a negative correlation. The deeper the color, the stronger the correlation, and colors closer to white represent correlations closer to zero. The significance of the correlation (*p < 0.05, **p < 0.01 and ***p < 0.001) is indicated. C Bubble plot displaying enriched metabolic pathways for intergroup differential metabolites. The y-axis shows pathway names, and the x-axis represents the enrichment factor (Rich factor-differential metabolites/total metabolites in pathway), with higher values indicating greater enrichment. Point color corresponds to the enrichment p-value (green: higher p-value; red: lower p-value), and point size reflects the number of enriched metabolites. Pathway enrichment significance in figure was determined by Fisher’s exact test with FDR correction (Benjamini-Hochberg method, q < 0.05). D Differential analysis of neuroactive-related metabolites secreted by the cecal microbiota between groups. The plot displays the top 20 differential metabolites, consisting of the 10 most significantly upregulated (red) and the 10 most significantly downregulated (blue) metabolites based on p-value. The x-axis represents log₂(FoldChange), while the y-axis indicates metabolite names. Statistical significance is denoted by asterisks (“*” represents p < 0.05 and “**” represents p < 0.01). Dot size corresponds to VIP value. E Diagram showing the analysis of the interaction between the rabbit’s cecum and cerebral cortex under dietary restriction.
Discussion
Diet-induced weight loss through dietary control has become a common strategy in obesity intervention, and its safety is generally considered to be superior to drug treatment. However, the potential impacts of such dietary patterns on brain neuro-metabolism have not been fully explored. This study aims to systematically elucidate the mechanism of its effects on the neural network of the cerebral cortex, microbial metabolism, and related host gene regulation in adult New Zealand rabbits through the establishment of a short-term dietary (SDR) restriction model. The gut microbiota, as a key regulator of food digestion and nutrient absorption, can cross the blood-brain barrier by secreting various neuroactive compounds (such as short-chain fatty acids, amino acid derivatives, and bile acid metabolites), and affect the functional metabolism of the CNS through direct or indirect pathways23. Previous study10 has shown that short-term intermittent fasting can improve the behavioral disorders of diabetic mice through the microbial-metabolism-neurological axis, and increase the levels of microbial metabolites related to cognitive function. Clinical evidence24 also indicates that intermittent fasting (IF) can restore synaptic plasticity and integrity, reduce protein toxicity, and restore the homeostasis of the glial and immune systems in various neurodegenerative disease models (such as Alzheimer’s disease, Parkinson’s disease, Huntington’s disease, and amyotrophic lateral sclerosis). In this study, short-term high-intensity dietary restriction not only led to a significant decrease in body weight of New Zealand rabbits but also caused a series of neurobehavioral abnormalities, including drowsiness, slow response, disordered fur, and polyuria. This result is consistent with our previous report25, suggesting that such interventions may increase the potential risks of depression, anxiety, or mental disorders. From a systems physiology perspective, caloric restriction and intermittent fasting achieve the weight-loss effect through multiple synergistic pathways: (1) activating vagus nerve afferent signals; (2) regulating the systemic immune status; (3) reshaping the composition and metabolic characteristics of intestinal microbiota; and (4) regulating the central dopaminergic pathway. The dynamic balance of the “gut-brain-metabolism” axis constitutes a basic physiological adaptation mechanism for the body to energy restriction26. At the metabolic level, the body breaks down liver glycogen, mobilizes muscle protein, and utilizes adipose tissue to generate glucose and ketone bodies to maintain the energy supply of brain27. Elevated ketone levels are believed to optimize brain energy metabolism and may potentially become a key mechanism for improving cognitive function in neurodegenerative disorders (such as Parkinson’s and Alzheimer’s diseases)28. The acute, severe dietary restriction adopted in this study shows a significant difference from the reported neuroprotective effects of mild or intermittent fasting in previous studies. We speculate that this difference mainly stems from the intensity, duration, and adaptability of the fasting mode. Mild intermittent fasting can induce beneficial adaptive responses, including enhancing metabolic flexibility, activating cellular defense mechanisms, and optimizing communication between the gut and the brain. However, drastic and severe dietary restriction may exceed the compensatory capacity of the body, triggering energy supply crises, dysregulation of the stress system, intestinal flora disorders, and neurochemical imbalances, ultimately leading to temporary impairment of neurobehavioral functions. Therefore, we believe that even short-term dieting, if carried out in a rather intense manner, may still pose neurocognitive risks by causing metabolic reorganization in the brain. This is particularly likely to affect synaptic plasticity, neurotransmitter balance, and myelin maintenance functions. This potential risk should be given due attention, and a comprehensive assessment should be conducted in future studies.
In Alzheimer’s disease (AD) research, plasma exosome-derived complement C1q subcomponent subunit C (C1QC) has been found to collaborate with metalloproteinase domain-containing protein 10 (ADAM10), immunoglobulin-like domain-containing protein (A0A0G2JRQ6), complement component C9 (C9), platelet glycoprotein Ibβchain (GP1BB), and Ras suppressor protein 1 (RSU1) in disease pathogenesis29. Microglia, as the resident immune cells of the brain, contribute to neurodegenerative processes when abnormally activated and metabolically dysregulated30. Notably, the lipid metabolism-related factor SAA3 plays pivotal roles across multiple pathological contexts: it not only participates in macrophage-mediated proinflammatory responses (e.g., in acute kidney injury)31, but also suppresses cholesterol metabolism through the TLR4/c-Jun/Cyp7a1 pathway (e.g., in hypercholesterolemia)32, and synergizes with IL-1β to promote cancer cell metastasis33. Furthermore, galectin-3 (Gal-3), encoded by the LGALS3 gene, functions as an innate immune molecule that regulates microglial inflammatory responses via the TLR4/MyD88/NF-κB pathway34. In tauopathy models, Gal-3-associated microglial activation exacerbates neuroinflammation and degeneration35. Our study reveals that SDR restriction significantly upregulated the expression of genes (C1QC, SAA3, and LGALS3) in cerebral cortex tissues, suggesting that these molecules may modulate neurodegenerative processes by enhancing immune and inflammatory responses in the cerebral cortex tissue of the rabbits. Previous studies36 have found that intermittent fasting (IF) significantly reduces levels of corticosterone, interleukin-6, and tumor necrosis factor-α in the plasma of stressed mice, and improves cognitive function while protecting the brain from stress-induced damage by modulating inflammatory response pathways. Clinical intervention study37 has shown that an alternate-day fasting low-carbohydrate diet can modulate immune-related biomarkers in the body, influencing the progression of diabetes, cancer, and autoimmune diseases. Although IF can mitigate the progression of neurological and metabolic disorders by reducing inflammatory responses in various metabolic diseases, it remains unclear whether this dietary pattern poses potential unknown risks to neural and metabolic remodeling in the human brain. CD40 is a critical immune co-stimulatory molecule and a member of the tumor necrosis factor receptor superfamily (TNFRSF). Research on psychiatric disorders38 has revealed that CD40/CD40L signaling regulates T cell-B cell interactions, antibody production, and immune memory formation in adaptive immunity. It can trigger TRAF-mediated NF-κB and MAPK signaling pathways, promoting neuroinflammation, and is associated with brain inflammation in schizophrenia and depression. However, in the current study, the upregulation of the NF-κB signaling pathway involving CD40/TRAF in inflammatory factors in the cerebral cortex suggests that SDR restriction may induce inflammatory activation in the cerebral cortex. Unlike intermittent fasting, dietary restriction involves a comprehensive reduction in energy and nutrient intake to maintain the body’s basic needs. Therefore, it is speculated that long-term excessive dietary restriction may adversely affect functional metabolism in the brain.
The dietary restriction implemented for weight loss exert complex regulatory effects on the brain’s metabolism, immunity and neurological functions. Transthyretin (TTR) is a tetrameric protein mainly synthesized in the liver, and its has various mutant forms are closely associated with hereditary amyloidosis (hATTR)39. The misfolding of TTR can form ATTR amyloid fibrils, which deposit in tissues and cause neuropathy and cardiomyopathy40. Concurrently, caloric restriction downregulates the expression of the ACVRL1 gene, which is associated with congenital cerebral arteriovenous malformations (VOGM)41, but elevated plasma levels of ACVRL1 may increase the risk of dementia42, suggesting that these two genes could affect neurocognition through their functional expression to influence the risk of cerebrovascular abnormalities and cognitive decline. In this study, it was found that the mRNA expression of TTR and ACVRL1 genes in the cerebral cortex was highly downregulated after dietary restriction. This indicated that the dietary restriction method may make the potential cerebral vascular abnormalities and cognitive dysfunction through the regulatory mechanisms of these two genes. Moreover, in the process of regulating metabolism by sex hormones, the prolactin receptor (PRLR) is distributed in the limbic system of the brain (such as the hippocampus), and together with the pleiotropic neurohormone prolactin secreted by the pituitary gland, participates in the regulation of emotional behavior43. In a chronic mild stress-induced depression model, silencing of the PRLR gene inhibits apoptosis of hippocampal CA3 neurons by suppressing the JAK2-STAT5 signaling pathway and increasing the expression of brain-derived neurotrophic factor (BDNF), ultimately alleviating depression-like behaviors44. In the present study, we observed significant downregulation of genes associated with sex hormone genes regulation, including PRLR, sperm protein antigen 17 (SPA17), and DPY30 domain-containing 2 (DYDC2). Although SPA17 and DYDC2 have been primarily studied in the context of spermatogenesis and oncogenesis45,46, these findings suggest that the short-term dietary restriction may alleviate metabolic stress induced by cerebral nutrient deficiency through sex hormone-related pathways. Furtherly, the recent study47 has found that key inflammatory biomarkers: C-reactive protein (CRP), interleukin-6 (IL-6), and tumor necrosis factor-alpha (TNF-α) did not change or increase, but CRP levels frequently rise and often significantly increase, especially among overweight or obese individuals during long fasting periods. In this study, the short-terms dietary restriction activated inflammatory pathways in the cerebral cortex, including gut IgA immune networks, viral myocarditis pathways, and NF-κB signaling, potentially inducing low-grade inflammation. Simultaneously, it inhibits critical pathways such as complement coagulation cascades and glycine metabolism, which may reduce cerebral protein synthesis and remodel immune function. Therefore, while improving metabolism and providing neuroprotection, the dietary restriction may adversely affect immune homeostasis and cerebral anabolism. The overall effects require further research to weigh the benefits against potential risks.
Fasting (or dietary restriction) can significantly influence the metabolic gene regulation of the intestinal immune system, and its effects are often bidirectional and highly dependent on the specific mode and duration of fasting. The study48 showed that long-term intermittent fasting can damage the health of the intestinal barrier, increase pro-inflammatory factors, disrupt the intestinal flora, increase pro-inflammatory metabolites, and exacerbate intestinal inflammation. Focusing on the balance between intestinal immunity and metabolism, the expression levels of metabolic enzymes can directly affect the absorption and transformation functions in the intestinal mucosa. The protein phosphatase Mg2 + /Mn2 + -dependent 1J (PPM1J) can influence muscle mass characteristics and glycerophospholipid composition through protein phosphorylation modifications49,50. As a member of the uridine diphosphate glucuronosyltransferase (UGTs) family, UGT2A1 is a key membrane-bound enzyme that catalyzes the conjugation of various endogenous (e.g., bilirubin) and exogenous compounds (e.g., drugs)51. Similarly, GALNT18 encodes a mucin glycosylase that promotes the formation of the intestinal mucosal barrier52. In this study, the gene expressions of PPM1J, UGT2A1, and GALNT18 were found to be significantly upregulated after dietary restriction, suggesting their potential roles in the repair of the intestinal mucosal barrier and the enhancement of epithelial cell metabolism. Meanwhile, phospholipase A2 group IVc (PLA2G4C) plays a key role in both MAPK signaling and glycerophospholipid metabolism. Studies53,54 have shown that gene polymorphisms in the PLA2G4 family are associated with the onset of psychiatric disorders, and the involved PLA2G2A/CYP2J2/PLA2G4C signaling pathway can regulate linoleic acid metabolism during placental ischemia. Asparaginase-like protein 1 (ASRGL1) cleaves isoaspartate residues to alter protein folding and proteolytic susceptibility. During the process of neural regulation, its overexpression can promote neuronal recovery, while its silencing leads to misfolded/phosphorylated proteins in the motor cortex of mice55. This study also observed upregulation of PLA2G4C and downregulation of ASRGL1, which are probably related to the metabolism of the intestinal nerve plexus. Then, we speculated that dietary restriction enhances intestinal energy conversion efficiency through the differential expression of metabolic genes in the cecal nerve plexus, thereby maintaining the body’s energy balance. Functional enrichment results further confirmed significant upregulation in metabolism-related signaling pathways (such as ascorbate and aldarate metabolism, pentose and glucuronate interconversions, and ether lipid metabolism, etc), while inflammation-related pathways (including the intestinal immune network for IgA production, rheumatoid arthritis, autoimmune thyroid disease, and NF-κB signaling pathway, etc) were significantly downregulated. Therefore, this interplay between metabolism and immunity may induce adaptive metabolic reprogramming in cecal tissues to compensate for intestinal metabolic deficiency caused by dietary restriction.
The brain-gut axis is a crucial regulatory circuit for substance metabolism, with its core mechanism centering on the interaction between gut microbiota changes and host genes affecting the body’s energy metabolism balance. From the perspective of molecular interactions, this study investigated the potential effects of dietary restriction on rabbit neurocognition by analyzing differential gene co-expression patterns between cecal and cerebral cortex tissues. The research56 found that intermittent fasting can increase the expression levels of intestinal Farnesoid X receptor (FXR), glucagon-like peptide-1 (GLP-1), and melanocortin-4 receptor (MC4R), while significantly reducing gastrointestinal peroxisome proliferator-activated receptor-gamma (PPAR-γ) expression, thereby alleviating high-fat diet-induced intestinal oxidative stress responses. Glutathione peroxidase 3 (GPX3) is an important endogenous antioxidant enzyme whose expression level changes are significantly associated with known pathological processes of obesity and insulin resistance57. The study58 has found that GPX3 knockout damages the mitochondrial inner membrane of adipocytes, leading to mitochondrial dysfunction, and its interaction with UCP1 participates in adipose browning, thereby increasing susceptibility to obesity. The zinc finger transcription factor EGR1 acts as a negative regulator of brown fat programming, and the inhibition of its expression increases the levels of thermogenic mitochondrial uncoupling protein 1 (UCP1), promotes spontaneous browning of subcutaneous white adipose tissue, and enhances fat burning and metabolic effects59. Meanwhile, research60 has also shown that GPR37, a G protein-coupled receptor, can activate the Hippo pathway in colon cancer cells, upregulate the expression of lactate dehydrogenase A (LDHA), and promote glycolysis. Additionally, it functions by modulating cell communication mechanisms that drive tissue damage and repair after ischemic stroke61. Notably, our analysis revealed that dietary restriction may induce distinct metabolic reprogramming phenomena in the gut-brain axis: the GPR37 and GPX3 expression showed significant upregulation in cecal tissue, the opposite trend was observed in the cerebral cortex. It is speculated that dietary restriction may induce distinct metabolic reprogramming phenomena along the gut-brain axis: while enhancing colonic metabolic compensation to maintain systemic energy supply, this adaptation may potentially lead to impaired mitochondrial energy metabolism in the cerebral cortex, thereby triggering inflammatory responses in the brain. Consequently, enhanced metabolic activity in cecal tissue likely plays a crucial regulatory role in systemic energy metabolism under dietary restriction conditions. Furthermore, the study62 found that prolonged fasting can trigger stress responses in the brain and exert relatively negative effects on immune cells, which may impair the ability to combat infections and increase the risk of heart disease. The cyclin-dependent kinase 5 regulatory subunit 1 (CDK5R1) promotes CNS development and neuronal migration, facilitating nerve repair through the CDK5/BDNF/TrkB pathway63. Reduced expression of the mitochondrial complex I-related gene (NDUFA2) is associated with a decreased risk of Alzheimer’s disease (OR = 0.96, P = 1.06 × 10-4)64. The calcium/calmodulin-dependent protein kinase IV (CaMK4), a multifunctional serine/threonine kinase, regulates both immune function and neural development, and its variants are associated with autism spectrum disorder65. Notably, expression profiling of neural function-related genes in this study revealed a more complex regulatory network: synchronous downregulation of CDK5R1 in both the cecum and cerebral cortex may affect neural repair processes; widespread upregulation of NDUFA2 suggests mitochondrial functional remodeling; and tissue-specific upregulation of CAMK4 in the cerebral cortex may be closely linked to neurodevelopmental abnormalities. Function enrichment analysis further confirmed that differentially expressed genes were significantly enriched in key neural metabolic pathways (Synaptic vesicle cycle, GABAergic synapse, long-term potentiation, glutamatergic synapse). These results provide molecular-level evidence explaining the neurocognitive changes induced by dietary restriction and, more importantly, highlight the necessity of implementing protective measures for neurological function during the process of humans dieting to lose weight.
Gut microbiota is closely associated with host metabolism, immune function, and nervous system regulation. In disease associations, metagenome-wide association studies have identified significant correlations between specific microbial signatures and metabolic disorders. For instance, the abnormal proliferation of Bacteroidia and Eubacterium is strongly associated with an increased risk of type I diabetes (OR = 1.28)66. Bacteroidales species harbor reversibly regulated antibiotic resistance genes (ARGs), potentially prolonging ARG persistence in bacterial populations67. In obesity-related metabolic syndrome, Desulfovibrio expansion and Clostridia depletion are key features in aged metabolic syndrome models and patients68. Alistipes indistinctus and Porphyromonas correlate with lipid markers (total cholesterol, triglycerides, LDL-C) in acute myocardial infarction (AMI) patients, while Mucispirillum, Bilophila, and Stenotrophomonas associate with myocardial injury indicators (cTnI, CK, CK-MB, LDH, BNP)69. Moreover, in traditional Chinese medicine (e.g., Puren Dan, PRD) for type 2 diabetes (T2DM), treatment modulates gut microbiota—increasing Prevotella sp. 10(H), Parabacteroides sp. SN4, Flavobacteriales, Bacteroides massiliensis, Alistipes indistinctus, and Ruminococcus flavefaciens, which improve intestinal metabolism. Antiviral resistance studies have identified Oribacterium sp. GMB0313 and Ruminococcus sp. GMB0270 in hamster gut microbiota as being linked to viral resistance70. Collectively, the above results indicate that these bacterial strains are closely linked to intestinal lipid metabolism and immune responses, which is consistent with our findings. In this study, dietary restriction led to alterations in the relative abundance of the cecal microbiota. Among the top 10 bacterial species, the abundance of certain microbes, such as Bacteroides and Alistipes indistinctus, decreased, while that of others, including Ruminococcus, Verrucomicrobia, and Clostridium, increased. It slightly reduced the diversity of the microbiota and caused disorder in the intestinal microbial community metabolism (with an increase in the ratio of Firmicutes/Bacteroidetes). The functional prediction of the microbial community also confirms that these bacterial species with relatively higher abundance in the Re group were associated with the top 10 metabolic pathways (including Alanine aspartate and glutamate metabolism, Peptidoglycan biosynthesis, Glycerolipid metabolism, etc). This further indicated that the dietary restriction has led to the compensatory energy regulation between the microbiota and the intestinal tract. This might be one of the reasons for the rebound after dieting and weight loss. Furtherly, it is known that there are three types of metabolites that mediate the interaction between the gut microbiota and the host: tryptophan and its metabolites, short-chain fatty acids (SCFAs), and bile acids. Specific Bacteroides species (B. fragilis, B. uniformis, B. caccae) modulate tryptophan pathway metabolites and neurotransmitters, inducing anxiety, despair-like behaviors, and impaired hippocampal neurogenesis71. The mechanism involves dysregulation of the tryptophan-kynurenine metabolic pathway and activation of neuroinflammation. Tryptophan, an essential amino acid, is mainly catalyzed by the indoleamine 2, 3-dioxygenase (IDO) and tryptophan 2, 3-dioxygenase enzymes into the kynurenine pathway, generating kynurenic acid with neuroprotective effects or quinolinic acid (QUIN) with neurotoxicity. During the normal dynamic balance metabolism of kynurenine metabolism, it was found that tryptophan, kynurenine and 3-hydroxykynurenine (3-HK) can pass through the blood-brain barrier, while kynurenic acid and QUIN cannot. A decrease in tryptophan levels can reduce serotonin synthesis and exacerbate depressive symptoms; while an increase in kynurenine is associated with immune activation and inflammation72. In severe depression and schizophrenia, the imbalance in tryptophan metabolism leads to excessive kynurenic acid, causing dysfunction of the endogenous N-methyl-D-aspartate receptor and α7 nicotinic acetylcholine receptor (α7 nAChR), and subsequently triggering negative symptoms and cognitive deficits of schizophrenia73. Additionally, the immune imbalance caused by proinflammatory mechanisms contributes to the metabolic disorder of kynurenine metabolism in the pathogenesis of schizophrenia. Increased levels of IL-1β, IL-6, TNF-α, and IFN-γ promote the kynurenine metabolic pathway to produce excessive kynurenic acid, which leads to cognitive dysfunction and other symptoms74. Meanwhile, long-term intermittent fasting can promote the uptake of tryptophan, the precursor of serotonin, by muscles, reduce the levels of tryptophan in the blood and brain, and thereby lower the level of serotonin75. Our study found that SDR restrictions led to a decrease in tryptophan and its derivatives in the cecum, indicating that insufficient nutrition caused metabolic disorders in the microbiota. At the same time, the brain tissue may preferentially absorb tryptophan and enter the kynurenine metabolic pathway to maintain immune and neural functions, even though there may be relatively low levels of tryptophan and its metabolites in the peripheral blood. However, this metabolic process leads to an increase in kynurenic acid accumulation, disrupting the dynamic metabolic balance and promoting the occurrence of potential neuroinflammation and cognitive impairment. The elevation of Clostridium and Ruminococcus promotes an increase in SCFAs (such as butyrate, acetic acid and propionic acid), which strengthened the intestinal barrier, reduced inflammatory exudation, but reduces the synthesis of 5-hydroxytryptamine, gamma-aminobutyric acid and indole-3-propionic acid, suggesting a potential trend of neuroexcitability attenuation in the body. Abnormalities in bile acid metabolism, such as changes in glycodeoxycholic acid and taurodeoxycholic acid levels, have also been associated with depression76. In this study, cecal butyric acid and bile acid levels significantly increased after restricted feeding, suggesting that it may alleviate cognitive impairment caused by nutritional deficiency through a compensatory metabolic mechanism. These findings provide new insights into the underlying molecular mechanisms of the “microbial-gut-brain axis” in neurometabolic regulation under restricted diet-induced weight loss conditions. In addition, this study has some limitations. In the initial phase of this study, we considered that dietary restriction might affect overall brain metabolism; therefore, we selected cortical tissue as the key subject for transcriptomic sequencing analysis. However, given the hippocampus’s close association with cognitive function, although we evaluated its pathological changes, we did not perform transcriptomic analysis on hippocampal tissue. Future studies should involve more precise functional brain region investigations. It is well-established that severe deficiencies in energy and nutrient intake can lead to emotion-related disorders in humans, such as anxiety and depression. However, there is currently a lack of widely accepted standard paradigms for assessing depressive and anxiety-like behaviors in rabbits. The phenotypic data obtained in this experiment were based on behavioral observations of the rabbits. Future research should explore and establish a standardized behavioral assessment system for depression and anxiety in rabbits, similar to tail suspension and forced swim tests used in mice. Furthermore, although this experiment identified key bacterial species such as Bacteroides, Alistipes indistinctus, and Ruminococcus that were positively correlated with brain metabolic regulation, we have not yet conducted in vitro culture or fecal microbiota transplantation experiments to validate their functional roles in achieving a balance between weight loss and physiological function. This study preliminary elucidates the effects of dietary restriction on gene-gut microbiota interactions within the gut-brain axis of New Zealand rabbits, with particular focus on the potential regulatory mechanisms by which cecal gene expression and microbial alterations influence cerebral cortical neurometabolism under restricted feeding conditions. Together our data could provide new strategies for human dietary weight management.
Methods
Construction of the restricted feeding model
A total of 24 male New Zealand rabbits aged 270 days were randomly selected and divided into the control group (Co, n = 12) and the restricted diet group (Re, n = 12). All rabbits in this study were housed and fed at the Animal Experiment Center of Guizhou Medical University under standardized breeding conditions. Throughout the study, the rabbits received regular vaccinations and consistent care. The experimental protocol consisted of two phases: a 14-day pre-feeding period followed by a 28-day experimental period. The rabbits were individually housed in single cages and provided with 200 g of feed per rabbit twice daily. During the 14-day pre-feeding period, all rabbits received a standard commercial diet (200 g twice daily; detailed composition in Supplementary Table 6). In the subsequent 28-day experimental period, the feeding regimen for Re group rabbits was adjusted: they were fed 100 g twice daily for the first 20 days, followed by 80 g twice daily for the remaining 8 days. In contrast, Co group rabbits maintained the original feeding protocol (200 g twice daily). The design of this experiment was shown in Fig. 6. The commercial diets for the adult rabbits used during the trial were produced in the same lot and stored under the same conditions. The nutritional composition of the commercial feed was consistent with that reported in our previous study77. At the end of the experiment, the rabbits were euthanized using the air embolism method. The specific procedure was to rapidly inject 15 mL of air into the rabbit’s ear vein to cause its death, and the death was confirmed by observing the cessation of heart and breathing, as well as the dilation of the pupils.
Design of the experimental protocol of restricted feeding diet in New Zealand rabbits.
HE staining
The samples of hippocampal tissue and middle segment of the cecum tissue samples fixed with 4% formaldehyde were rinsed under running water and trimmed, then placed in plastic embedding cassettes for dehydration (75% ethanol for 6 h, 85% ethanol for 10 h, 95% ethanol for 4 h, absolute ethanol I for 2 h, absolute ethanol II for 2 h), clearing (xylene I for 20 min, xylene II for 15 min), and wax immersion for 3 h, followed by embedding the tissue blocks in paraffin. Subsequently, the Leica RM2235 microtome was used to cut the tissues into 5 µm-thick sections, which were flattened in warm water and mounted onto glass slides. The sections were then baked at 60 °C for at least 2 h. Next, the sections were dewaxed in xylene, rinsed under running water for 20 min, stained with hematoxylin for 30 min, washed again under running water for 20 min, differentiated with hydrochloric acid alcohol, and stained with eosin for 5 min. Finally, the sections were dehydrated through a graded alcohol series, cleared in xylene, and sealed with resinous mounting medium. Therefore, the pathological changes in the cerebral cortex and cecal samples within the group were observed and photographed under a microscopy imaging system.
Total RNA extraction and RNA-sequencing
Total RNA was isolated from six cecal samples and six right cerebral cortex tissues from the Co and Re groups using TRIzol reagent (Invitrogen, CA, USA) according to the manufacturer’s protocol. RNA concentration and purity were determined using a NanoDrop 2000 spectrophotometer (Thermo Scientific, USA). Sequencing libraries were prepared with the VAHTS Universal V6 RNA-seq Library Prep Kit, following the manufacturer’s instructions. Paired-end sequencing (150 bp) was performed on an Illumina NovaSeq 6000 platform by OE Biotech Co., Ltd. (Shanghai, China). Raw sequencing reads (FastQ format) were quality-filtered using fastp to obtain clean reads for each sample. Clean reads were then aligned to the reference genome (Oryctolagus cuniculus UM_NZW_1.0) using HISAT2. Gene expression levels were quantified as FPKM, and read counts per gene were generated using HTSeq-count.
DNA extraction and metagenomic sequencing
Total DNA was extracted from 12 cecal content samples (six each from Co and Re) using the QIAamp® Fast DNA Stool Mini Kit (Qiagen), with quality verified by NanoDrop2000 spectrophotometry and 1% agarose gel electrophoresis. Qualified DNA samples were fragmented to ~350 bp using a Covaris S220 ultrasonicator, size-selected with AMPure XP beads, and processed into sequencing libraries using the TruSeq Nano DNA LT Kit (Illumina). After Qubit quantification, libraries were subjected to 150 bp paired-end sequencing on an Illumina NovaSeq 6000 platform (OE Biotech Co., Ltd.), generating metagenomic data for downstream analysis.
RNA-Seq Data Analysis
Differentially expressed genes (DEGs) were identified using NOISeq and DESeq2 with stringent thresholds (∣log₂Foldchange (FC)∣ ≥ 1 and P-value < 0.05), followed by functional enrichment analysis through DAVID for Gene Ontology annotation and the KEGG database for pathway mapping, with all statistical analyses and visualizations performed using R software (v4.2.2) with ggplot2 and clusterProfiler packages. The statistical significance of the enrichment of gene sets in GO terms and KEGG pathways was analyzed using Fisher’s exact test. P-values were adjusted for multiple testing using the Benjamini-Hochberg false discovery rate (FDR) method. Terms with a q-value < 0.05 (Benjamini-Hochberg FDR) were considered significantly enriched.
Metagenomic data processing and analysis
Raw sequencing reads were quality-trimmed using Trimmomatic (v0.36)78, and the resulting paired-end clean reads were aligned to the Oryctolagus cuniculus UM_NZW_1.0 reference genome with HoCoRT: for host DNA removal79. De novo assembly was performed using MEGAHIT (v1.1.2)80, retaining scaftigs >500 bp for downstream analysis. Gene prediction was conducted via Prodigal (v2.6.3)81, followed by non-redundant gene catalog construction using CD-HIT (v4.5.7). Taxonomic profiling was achieved by aligning predicted genes against the NR database, with abundance calculations at multiple taxonomic levels (phylum to species). Statistical analyses included: (1) principal component analysis (PCA) of taxonomic/functional profiles using R (v3.2.0), (2) inter-group significance testing via ANOVA in SPSS 22.0, and (3) biomarker identification through LEfSe analysis of differentially abundant features.
Analysis of microbial metabolites
We utilized a standardized metabolomic preparation protocol: 60 mg of sample was homogenized with pre-chilled steel beads in 600 μL of methanol—water (4:1, v/v, containing 4 μg/mL L-2-chlorophenylalanine) using a grinder and ultrasonication. After incubation at –40 °C for 30 min and centrifugation (12,000 rpm, 4 °C, 10 min), 200 μL of supernatant was dried under nitrogen. The residue was reconstituted in 300 μL methanol–water (1:4, v/v), vortex-mixed, sonicated, incubated at–40 °C for 2 h, then recentrifuged. A 150 μL aliquot was filtered (0.22 μm) and stored at –80 °C until analysis. QC samples were prepared by mixing equal volumes of all extracts. LC—MS analysis was conducted on an ACQUITY UPLC I-Class Plus/QE Plus system. Separation used an ACQUITY UPLC HSS T3 column (100 × 2.1 mm, 1.8μm) at 45 °C, with mobile phase A (0.1% formic acid in water) and B (acetonitrile) at 0.35 mL/min. The gradient was: 5% B (0–2 min), 5–30% B (2–4 min), 30–50% B (4–8 min), 50–80% B (8–10 min), 80–100% B (10–14 min), hold 100% B (14–15 min), return to 5% B in 0.1 min and re-equilibrate for 1 min (total 16 min). Injection volume was 5 μL. MS data were acquired in both ESI+ and ESI—modes with spray voltages of 3800 V and—3000 V, respectively. Capillary and heater temperatures were 320 °C and 350 °C; sheath and auxiliary gas flows were 35 and 8 Arb. S-lens RF was 50. Full MS and MS/MS scans (m/z 100–1200) were performed at resolutions of 70,000 and 17,500, respectively, using stepped NCE (10, 20, 40).
Metabolomics data analysis
The raw metabolomics data were preprocessed using Progenesis QI v3.0 software. Metabolic features with over 50% within-group missing values (zero values) were removed, and remaining missing values were replaced with half of the corresponding minimum value. Compounds were filtered based on qualitative identification scores (Score); those scoring below 36 (out of 80) were deemed unreliable and excluded. Data from positive and negative ion modes were merged into a unified matrix for subsequent analysis. During statistical analysis, unsupervised PCA and supervised PLS-DA were employed to assess overall metabolic profile differences between groups. Differential metabolites were identified based on the criteria: VIP > 1, |Log₂FoldChange | >1, and p-value < 0.05. Enrichment analysis of metabolic pathways was then conducted using the KEGG database.
Protein–protein interaction network analysis
The protein-protein interaction network was constructed using STRING database and visualized in Cytoscape, with key hub genes identified through CytoHubba’s 12 topological algorithms including connectivity-based (Degree), clique-based (Maximum Clique Centrality), neighborhood-based (Maximum Neighborhood Component, Density of MNC, Edge Percolated Component), and shortest-path-based centralities (Bottleneck, Betweenness, Stress, Closeness, Radiality, Eccentricity), ensuring robust multi-algorithm identification of functionally significant network components.
Ethics statement
This study was approved by and conducted in strict accordance with the ethical standards of the Animal experimental ethical inspection form of Guizhou Medical University, Guizhou, China (No: 2201379).
Data availability
The datasets generated during the current study are available in the NCBI database [SRA] repository, accession number PRJNA1119243 (Metagenomic data), PRJNA1119171 (Cecum tissues transcriptome data) and PRJNA1274676 (Cerebral cortex tissues transcriptome data).
References
Ma, X. et al. PuRenDan alleviates type 2 diabetes mellitus symptoms by modulating the gut microbiota and its metabolites. J. Ethnopharmacol. 25, 322, 117627 (2024).
Forouhi, N. G. Embracing complexity: making sense of diet, nutrition, obesity and type 2 diabetes. Diabetologia 66, 786–799 (2023).
Asadi, A. et al. Obesity and gut–microbiota–brain axis: a narrative review. J. Clin. Lab Anal. 36, e24420 (2022).
Pena-Leon, V. et al. Novel mechanisms involved in leptin sensitization in obesity. Biochem. Pharm. 223, 116129 (2024).
Zanco, B. et al. A dietary sterol trade-off determines lifespan responses to dietary restriction in Drosophila melanogaster females. Elife 10, e62335 (2021).
Li, M. et al. Effect of 5:2 regimens: energy-restricted diet or low-volume high-intensity interval training combined with resistance exercise on glycemic control and cardiometabolic health in adults with overweight/obesity and type 2 diabetes: a three-arm randomized controlled trial. Diabetes Care 47, 1074–1083 (2024).
Dong, T. S. et al. A high protein calorie restriction diet alters the gut microbiome in obesity. Nutrients 12, 3221 (2020).
Patel, Y. et al. Obesity and the cerebral cortex: Underlying neurobiology in mice and humans. Brain Behav. Immun. 119, 637–647 (2024).
Gómez-Apo, E. et al. Structural brain changes associated with overweight and obesity. J. Obes. 16, 6613385 (2021).
Liu, Z. et al. Gut microbiota mediates intermittent—fasting alleviation of diabetes-induced cognitive impairment. Nat. Commun. 11, 855 (2020).
Mattson, M. P. et al. Intermittent metabolic switching, neuroplasticity and brain health. Nat. Rev. Neurosci. 19, 81–94 (2018).
Jamshed, H. et al. Effectiveness of early time-restricted eating for weight loss, fat loss, and cardiometabolic health in adults with obesity: a randomized clinical trial. JAMA Intern. Med. 182, 953–962 (2022).
Le, T. N. et al. Extreme weight loss and psychosis as presenting signs of thyrotoxicosis. Cureus 13, e14045 (2021).
Alkurd, R. et al. Effect of calorie restriction and intermittent fasting regimens on brain-derived neurotrophic factor levels and cognitive function in humans: a systematic review. Medicina 60, 191 (2024).
Wang, Q. et al. The microbiota–gut–brain axis and neurodevelopmental disorders. Protein Cell 14, 762–775 (2023).
Quigley, E. M. Microbiota-brain-gut axis and neurodegenerative diseases. Curr. Neurol. Neurosci. Rep. 17, 1–9 (2017).
Longo, S. et al. Microbiota-gut-brain axis: Relationships among the vagus nerve, gut microbiota, obesity, and diabetes. Acta Diabetol. 60, 1007–1017 (2023).
Richards, P. et al. The gut–brain axis: identifying new therapeutic approaches for type 2 diabetes, obesity, and related disorders. Mol. Metab. 46, 101175 (2021).
Hirayama, M. et al. Parkinson’s disease and gut microbiota. Ann. Nutr. Metab. 77, 28–35 (2021).
Góralczyk-Bińkowska, A. et al. The microbiota–gut–brain axis in psychiatric disorders. Int. J. Mol. Sci. 23, 11245 (2022).
Thibaut, M. M. et al. Crosstalk between bile acid-activated receptors and microbiome in entero-hepatic inflammation. Trends Mol. Med. 28, 223–236 (2022).
Tang, Y. et al. Skeletal muscles and gut microbiota-derived metabolites: novel modulators of adipocyte thermogenesis. Front. Endocrinol. 14, 1265175 (2023).
Li, B. et al. Melatonin ameliorates neuropsychiatric behaviors, gut microbiome, and microbiota-derived metabolites in rats with chronic sleep deprivation. Int. J. Mol. Sci. 24 (2023).
Hein, Z. M. et al. Intermittent fasting as a neuroprotective strategy: gut-brain axis modulation and metabolic reprogramming in neurodegenerative disorders. Nutrients 17, 2266 (2025).
Li, Y. H. et al. Intramuscular adipocyte and fatty acid differences between high-fat and control rabbit groups subject to a restricted diet. Vet. Med Sci. 7, 2051–2060 (2021).
Chelluboina, B. et al. Intermittent fasting induced cerebral ischemic tolerance altered gut microbiome and increased levels of short-chain fatty acids to a beneficial phenotype. Neurochem. Int. 178, 7 (2024).
Kersten, S. The impact of fasting on adipose tissue metabolism. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1868, 159262 (2023).
Gudden, J. et al. The effects of intermittent fasting on brain and cognitive function. Nutrients 13, 3166 (2021).
Cai, H. et al. Proteomic profiling of circulating plasma exosomes reveals novel biomarkers of Alzheimer’s disease. Alzheimers Res. Ther. 14, 181 (2022).
Cheng, J.et al. Early glycolytic reprogramming controls microglial inflammatory activation. J. Neuroinflammation 18 (2021).
Peng, Y. et al. Saa3 promotes pro-inflammatory macrophage differentiation and contributes to sepsis-induced AKI. Int Immunopharmacol. 127, 13 (2024).
Huang, S. et al. Osteocytes/osteoblasts produce SAA3 to regulate hepatic metabolism of cholesterol. Adv. Sci. 11, e2307818 (2024).
Zhang, C. et al. Pulmonary interleukin 1 beta/serum amyloid A3 axis promotes lung metastasis of hepatocellular carcinoma by facilitating the pre-metastatic niche formation. J. Exp. Clin. Cancer Res. 42, 166 (2023).
Liu, Y. et al. Galectin-3 regulates microglial activation and promotes inflammation through TLR4/MyD88/NF-kB in experimental autoimmune uveitis. Clin. Immunol. 236, 108939 (2022).
Siew, J. J. et al. Galectin-3 aggravates microglial activation and tau transmission in tauopathy. J. Clin. Investig. 134, 17 (2024).
Cantoni, C. et al. Effects of dietary restriction on gut microbiota and CNS autoimmunity. Clin. Immunol. 235, 108575 (2022).
Akasheh, R. T. et al. Reduced serum PD-L1 and markers of inflammation in response to alternate day fasting with a low-carbohydrate intervention: a secondary analysis of a single-arm trial. Curr. Dev. Nutr. 9, 104566 (2025).
Maes, M. et al. Neuroimmune, metabolic and oxidative stress pathways in major depressive disorder. Nat. Rev. Neurol. 21, 473–489 (2025).
Dugo, K. et al. Hereditary transthyretin-related amyloidosis: genetic heterogeneity and early personalized gene therapy. Biomedicines 10, 2394 (2022).
Ueda, M. Transthyretin: its function and amyloid formation. Neurochem. Int. 155, 105313 (2022).
Zhao, S. et al. Mutation of key signaling regulators of cerebrovascular development in vein of Galen malformations. Nat. Commun. 14, 7452 (2023).
Beydoun, M. A. et al. Plasma proteomic biomarkers and the association between poor cardiovascular health and incident dementia: the UK Biobank study. Brain Behav. Immun. 119, 995–1007 (2024).
Jiang, J. et al. Prolactin deficiency drives diabetes-associated cognitive dysfunction by inducing microglia-mediated synaptic loss. J. Neuroinflammation 21, 295 (2024).
Tian, R. H. et al. Reducing PRLR expression and JAK2 activity results in an increase in BDNF expression and inhibits the apoptosis of CA3 hippocampal neurons in a chronic mild stress model of depression. Brain Res. 1725, 146472 (2019).
Grizzi, F. et al. Sperm protein antigen 17 and Sperm flagellar 1 cancer testis antigens are expressed in a rare case of ciliated foregut cyst of the common hepatic duct. Pathol. Res. Pract. 247, 154546 (2023).
Ma, R.-H. et al. Detection of aberrant DNA methylation patterns in sperm of male recurrent spontaneous abortion patients. Zygote 31, 163–172 (2023).
de Ciutiis, I. et al. Long-term fasting and its influence on inflammatory biomarkers: a comprehensive scoping review. Ageing Res. Rev. 110, 102797 (2025).
Fu, R. et al. Intermittent fasting exacerbates colon inflammation by promoting Th17 cell differentiation through inhibition of gut microbiota-derived indoleacrylic acid. World J. Gastroenterol. 31, 108815 (2025).
Huang et al. WIP1 phosphatase is a critical regulator of adipogenesis through dephosphorylating PPAR gamma serine 112. Cell Mol. Life Sci. 74, 2067–2079 (2017).
Ma, M. et al. PPM1J regulates meat quality feature and glycerophospholipids composition in broiler by modulating protein dephosphorylation. NPJ Sci. Food 8, 89 (2024).
Gaither, K. A. et al. Effects of alcohol consumption and tobacco smoking on the composition of the ensemble of drug-metabolizing enzymes and transporters in human liver. BioRxiv 17, 2024.05.14.594255 (2024).
Cheng, H. et al. Transcriptome analysis reveals modulations in glycosylation profiles of the mucosal barrier and their potential interaction with gut microbiota in weaned piglets. Anim. Nutr. 20, 226–238 (2025).
Yang, Z. et al. Altered distribution of fatty acid exerting lipid metabolism and transport at the maternal-fetal interface in fetal growth restriction. Placenta 139, 159–171 (2023).
Nadalin, S. et al. Association between PLA2 gene polymorphisms and treatment response to antipsychotic medications: a study of antipsychotic-naïve first-episode psychosis patients and nonadherent chronic psychosis patients. Prostaglandins Leukot. Ess. Fat. Acids 194, 102578 (2023).
Garcia-Montojo, M. et al. TDP-43 proteinopathy in ALS is triggered by loss of ASRGL1 and associated with HML-2 expression. Nat. Commun. 15, 4163 (2024).
Eraky, S. M. et al. The ameliorating effect of intermittent fasting on intestinal glucagon-like peptide 1 in rats fed a high-fat diet via the Farnesoid X receptor and the Melanocortin-4 receptor. Life Sci. 361, 123327 (2025).
Wodarski, A. et al. Association of glutathione peroxidase 3 (GPx3) and miR-196a with carbohydrate metabolism disorders in the elderly. Int. J. Mol. Sci. 25, 5409 (2024).
Song, Y. et al. Glutathione peroxidase 3 is essential for countering senescence in adipose remodelling by maintaining mitochondrial homeostasis. Redox Biol. 77, 103365 (2024).
Bléher, M. et al. Egr1 loss-of-function promotes beige adipocyte differentiation and activation specifically in inguinal subcutaneous white adipose tissue. Sci. Rep. 10, 15842 (2020).
Zhou, J. et al. GPR37 promotes colorectal cancer liver metastases by enhancing the glycolysis and histone lactylation via Hippo pathway. Oncogene 42, 3319–3330 (2023).
Bian, X. et al. GPR37 and its neuroprotective mechanisms: bridging osteocalcin signaling and brain function. Front. Cell Dev. Biol. 12, 1510666 (2024).
Janssen, H. et al. Monocytes re-enter the bone marrow during fasting and alter the host response to infection. Immunity 56, 783–796.e7 (2023).
Xia, L. et al. CDK5R1 promotes Schwann cell proliferation, migration, and production of neurotrophic factors via CDK5/BDNF/TrkB after sciatic nerve injury. Neurosci. Lett. 817, 137514 (2023).
Zheng, J. et al. Evaluating the efficacy and mechanism of metformin targets on reducing Alzheimer’s disease risk in the general population: a Mendelian randomisation study. Diabetologia 65, 1664–1675 (2022).
Kaiser, J. et al. Convergence on CaMK4: a key modulator of autism-associated signaling pathways in neurons. Biol. Psychiatry 97, 439–449 (2025).
Luo, M. et al. Gut microbiota and type 1 diabetes: a two-sample bidirectional Mendelian randomization study. Front. Cell Infect. Microbiol. 13, 1163898 (2023).
Yan, W. et al. Bacteroidales species in the human gut are a reservoir of antibiotic resistance genes regulated by invertible promoters. npj Biofilms Microbiomes 8, 1 (2022).
Petersen, C. et al. T cell–mediated regulation of the microbiota protects against obesity. Science 365, eaat9351 (2019).
Zuo, J. et al. Lipid alterations in acute myocardial infarction are associated with gut microbiota. Microbiol. Spectr. 13, e02370–02324 (2025).
Wang, M. et al. The gut microbe pair of Oribacterium sp. GMB0313 and Ruminococcus sp. GMB0270 confers complete protection against SARS-CoV-2 infection by activating CD8+ T cell-mediated immunity. Gut Microbes 16, 2342497 (2024).
Zhang, Y. et al. Bacteroides species differentially modulate depression-like behavior via gut-brain metabolic signaling. Brain Behav. Immun. 102, 11–22 (2022).
Pu, J. et al. Characterizing metabolomic and proteomic changes in depression: a systematic analysis. Mol. Psychiatry 30, 3120–3128 (2025).
Wang, Z. et al. Multiomics analyses reveal microbiome-gut-brain crosstalk centered on aberrant gamma-aminobutyric acid and tryptophan metabolism in drug-naïve patients with first-episode schizophrenia. Schizophr. Bull. 50, 187–198 (2024).
Suneson, K. et al. Inflammatory depression-mechanisms and non-pharmacological interventions. Int. J. Mol. Sci. 22, 1640 (2021).
Xie, K. et al. Intermittent fasting boosts sexual behavior by limiting the central availability of tryptophan and serotonin. Cell Metab. 37, 1189–1205.e7 (2025).
Jia, M. et al. Gut microbiota dysbiosis promotes cognitive impairment via bile acid metabolism in major depressive disorder. Transl. Psychiatry 14, 503 (2024).
Li, Y. et al. Comprehensive analysis of key host gene-microbe networks in the cecum tissues of the obese rabbits induced by a high-fat diet. Front. Cell Infect. Microbiol. 14, 1407051 (2024).
Chen, S. et al. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Rumbavicius, I. et al. HoCoRT: host contamination removal tool. BMC Bioinforma. 24, 1–371 (2023).
Li, D. et al. MEGAHIT v1.0: a fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Land, M. L. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 11, 119–119 (2010).
Acknowledgements
We thank J.N.C. and L.M.W. for their valuable comments and discussions. We also thank G.Z.W. for their valuable feedback on this manuscript. We thank X.L.Q. and W.H. for their support in providing the research platform. Finally, we thank the Molecular Biology Key Laboratory and Animal Experiment Center of Guizhou Medical University offer experimental platforms. This research was funded by Excellent Young Talents Plan of Guizhou Medical University [No. (2022)101], the Science, Technology Fund of Guizhou Provincial Health Commission [No. gzwkj2024-006], the National Natural Science Foundation of China [No. 32260215], the 2022 Health Sciences and Technology program of Hangzhou: General Project Category A [No. A20220084] and the Hangzhou Special Project for Biomedical and Health Industry [No. 2022WJC064].
Author information
Authors and Affiliations
Contributions
L.M.W. and G.Z.W. analyzed and interpreted the experimental data regarding the metagenomic metabolism and the transcriptome metabolism. J.N.C. was a major contributor to writing the manuscript. Y.H. conceived of and designed the study. X.L.Q. and W.H. provided valuable intellectual support throughout the course of the experiment. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Chen, J., Qi, X. et al. Microbiota-host crosstalk: the role of short-term dietary restriction in neurological and metabolic dysregulation. npj Biofilms Microbiomes 11, 226 (2025). https://doi.org/10.1038/s41522-025-00857-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-025-00857-w