Abstract
Directed atomic fabrication using an aberration-corrected scanning transmission electron microscope (STEM) opens new pathways for atomic engineering of functional materials. In this approach, the electron beam is used to actively alter the atomic structure through electron beam induced irradiation processes. One of the impediments that has limited widespread use thus far has been the ability to understand the fundamental mechanisms of atomic transformation pathways at high spatiotemporal resolution. Here, we develop a workflow for obtaining and analyzing high-speed spiral scan STEM data, up to 100 fps, to track the atomic fabrication process during nanopore milling in monolayer MoS2. An automated feedback-controlled electron beam positioning system combined with deep convolution neural network (DCNN) was used to decipher fast but low signal-to-noise datasets and classify time-resolved atom positions and nature of their evolving atomic defect configurations. Through this automated decoding, the initial atomic disordering and reordering processes leading to nanopore formation was able to be studied across various timescales. Using these experimental workflows a greater degree of speed and information can be extracted from small datasets without compromising spatial resolution. This approach can be adapted to other 2D materials systems to gain further insights into the defect formation necessary to inform future automated fabrication techniques utilizing the STEM electron beam.
Similar content being viewed by others
Introduction
The rapidly evolving landscape of science and technology demands unprecedented precision in materials fabrication, pushing the boundaries of traditional manufacturing techniques. Atomically precise fabrication has emerged as a transformative approach that promises unparalleled control over the structure and properties of materials at the atomic scale. This precision is essential for unlocking breakthroughs in diverse fields, including quantum sensing and communications1,2,3,4. Recently, it has been shown that the electron beam can be used as an exceptionally precise tool for direct fabrication on the atomic scale, the approach dubbed the atomic forge5. As the pursuit of atom-by-atom construction of materials gains momentum2,6,7,8,9,10,11,12,13,14,15,16, it becomes imperative to deepen our understanding of the underlying processes involved in atomic disordering and reordering. This knowledge is crucial in informing the development of future atomic fabrication techniques for novel functional applications. By comprehending and identifying the mechanisms behind defect formation, it becomes possible to tailor defects within two-dimensional (2D) materials to exhibit specific electronic, magnetic, or optical properties17,18,19,20,21,22,23,24,25,26. Such tailored defects hold the potential to revolutionize the performance and functionality of 2D materials, opening exciting avenues for scientific exploration and technological innovation in diverse fields as the degree of control over materials fabrication can be understood down to the point defect scale. In this paper, we aim to explore the intricate relationship between defect formation processes and electron beam irradiation, offering insights into the future design principles for atomically precise fabrication and the creation of functional materials with tailored properties.
While traditional synthesis methods, such as top-down or bottom-up synthesis have been used extensively for 2D materials27,28, these methods do not reach the needed length scales required for atomic structure fabrication. With the advent of using the sub-Å e- beam of a scanning transmission electron microscope (STEM) as a direct fabrication tool in recent years, information has already been gathered on the effect that the e- beam irradiation has on various 2D materials11,12,13,14,15,16,29,30,31,32,33. This technique relies on the localized defect generation caused by the highly energetic electrons. The primary mechanism for electron beam damage is believed to be the knock-on effect, where the electron transfers enough kinetic energy to the nucleus to displace it from the atomic structure33,34,35,36. The knock-on energy is determined by the momentum transfer from the electron to the nucleus and the binding energy of the atom in the lattice. However, in most materials, atomic manipulation can be observed well below this knock-on threshold, stimulating the development of alternative theories for this process. In previous work, Susi proposed the additional role of the phonons to the displacement33. At the same time, Lingerfelt and Meunier proposed and developed theories based on the local electronic excitations to the antibonding states and subsequent isomerization of the excited bond, akin to the photochemical effects37,38. Additionally, recently TMD semiconductors/semimetals have been seen to suffer from beam induced charging. At slower scan speeds sample charging can modify the effective knock-on threshold energy. Regardless, by controlling these atomic interactions, different defects and structures can be written into the atomic structure of 2D materials11,12,13,14,15,16,29,30,31,32,33. However, the generation of these defects and formation of different defect structures need to be better understood and dose control cannot be relied upon as a control mechanism. Through the precise control with the e- beam, these defects can be fabricated below knock-on thresholds. Active monitoring of the acquired STEM images can help limit the overall damage to the surrounding atomic structure and aid in designing new atomic structures in future devices made with 2D materials.
Here we used the transition metal dichalcogenide (TMD) MoS2, as the model system to explore the structural changes associated with defect and nanopore formation. As a model system, defect engineering within TMDs have been of interest in recent years as different atomic defects39,40,41,42, nanopores17,24,43, and nanowire structures16,21,44,45,46 fabricated within the monolayer have been shown to have beneficial for electronic and optical properties. Using an external beam control system operating at high temporal resolution, the atomic reordering and disordering that initially takes place during e- beam irradiation was recorded. To further delve into the interatomic relation these images were then decoded using the ensemble learning-iterative training (ELIT) workflow47,48 to process the images and display atomic trajectories and the formation of different defect structures. This workflow allowed for more information to be extracted from small datasets without losing out on temporal or spatial resolution.
To acquire the atomically resolved images required for these experiments, an aberration-corrected STEM was used in conjunction with an external beam control system that allows for simultaneous imaging and atomic fabrication. A custom-built beam control system was used for these studies due to the difficulties introduced from traditional STEM raster scanning. The standard usage of these methods results in high quality atomic resolution data, but at the expense of slower scanning rates that result in higher e- beam irradiation to the samples and less frequent updates on the beam-induced modifications. However, when using the STEM e- beam as the fabrication tool, greater control at higher temporal resolution is needed for automated feedback control. Therefore an external beam control system, that was developed at Oak Ridge National Laboratory, was used to control the scanning pathways the beam takes while simultaneously imaging6,8,30,49,50,51. Through this novel beam control system, a spiral scan path is used where single frequency drive signals to the scan coils is employed, enabling acquisition of atomically resolved images at speeds of up to 100 frames per second (fps)51. The utilization of a spiral path eliminates sharp changes in scan velocity from one line to the next and mitigates the flyback duration associated with traditional raster scan profiles, enabling much faster imaging speeds without distortion, improved temporal resolution from frame to frame, and therefore reduced electron dose to the material per frame while using standard beam steering magnetic coils. At the periphery the beam is moving faster likely causing less charge build-up localizing the beam irradiation effects to the center of the field of view (FOV) frames. Remaining image distortion resulting from high-speed imaging can be mitigated using subsequent image correction methods50,51. However, at such high imaging speeds, averaging times per pixel are lower and result in lower signal-to-noise ratios in each frame. Here we show that through the development and application of advanced post-processing techniques and sophisticated data analytics, we are able to extract critical information from the images (specifically, atomic positions and species) that can be used to better understand beam-induced material transformations as they occur. Furthermore, the beam control system is built with automation and feedback that is used to assess beam damage from frame to frame and automatically stop imaging after a specified damage threshold has been crossed (discussed further below).
In this study, a workflow incorporating deep learning (DL) networks was employed to decode individual frames captured during this rapid image acquisition. Distinguishing features in atomically resolved experiments, such as scanning tunneling microscopy (STM) and scanning transmission electron microscopy (STEM), differ from classifying objects commonly found in publicly available datasets like CIFAR and ImageNet52,53. Previous DL networks have been designed and implemented for precise and accurate image analysis of molecular54,55,56 and atomically resolved data57,58,59,60,61,62,63, but typically require high-quality data for precise model predictions. In this work, a network was utilized for image segmentation of nearly identical objects, namely atoms, while also quantifying prediction uncertainty to adapt to images originating from different experimental or simulation setups. Analysis of the acquired datasets presented challenges due to material lattice distortions, image drifts, and variations in imaging parameters, resulting in an out-of-distribution drift effect that reduces the accuracy of traditional deep convolutional neural network (DCNN) model predictions between microscope experiments. To address this issue, the ELIT workflow proposed and developed by Ghosh et al. was implemented here47,48. This approach demonstrated how a DL workflow could be applied to STEM data, allowing for adaptation to changes in imaging conditions. Within the ELIT workflow, multiple U-Net models were trained with random initialization parameters for the material system being studied61. The ensemble training phase aimed to select artifact-free models for feature selection, while the iterative phase directed the network’s attention towards recognized features to identify unknowns while considering pixel-wise uncertainty. By following this process, the models successfully identified the location and type of each atom within the frame with a high degree of certainty due to various parameters being altered between experiments. Through the utilization of the ELIT workflow, these small MoS2 datasets were effectively decoded, uncovering new insights into various materials systems while simultaneously increasing the speed and information content.
Results
Experimental decoding workflow during e- beam fabrication
The experimental workflow from data acquisition to processing and decoding can be seen in Fig. 1. The process starts with the creation of ideal simulated training to give a baseline for the models constructed later to iterate from (Fig. 1a). To begin crystal models of pristine molybdenum disulfide’s atomic structure, with a random assortment of sulfur sites deleted, were used as training data. These sulfur site point defects were introduced into the training data to simulate the defect generation that occurs during the e- beam fabrication therefore it was deemed necessary to train the model on the distinction of sites with both two overlapping sulfur atoms, as is seen in the ideal crystal structure, and sites with only a single sulfur atom. STEM images were then simulated using the variation of the multislice algorithm within the abTEM package developed by Madsen et al.64,65. To make these simulated images a .xyz crystal file of monolayer MoS2 was used for atomic coordinates that was 6 nm in thickness to create the potential with a give real-space sampling. This is then run through a multislice algorithm that makes use of an electron beam operating at 60 kV simulated using a plane wave function and a contrast transfer function that represents a given aperture and spherical aberration. Using these simulated images, corresponding circular masks with a fixed radius were constructed classifying each atomic site appropriately. Once created, these image and mask pairs were augmented through random cropping, rotations, zooming-in, contrast variations and through the application of blurring (Gaussian \(\sigma =[\mathrm{1,50}]\)), and addition of both Gaussian (\(\sigma =[3000,\,8000]\)) and Poisson (\(\sigma =[\mathrm{30,45}]\)) noise to replicate real experimental data more accurately. An example of a final simulated image mask pair can be seen in the block inset of Fig. 1a.
a Simulate the training data: The initial block consisted of constructing the simulated training data. First STEM images are simulated and then their associated labeled mask pairs created, followed by data transforms. b The Ensemble Learning and Iterative Training workflow is then demonstrated as the training and experimental data is input and several DCNN models are output comparing the fit between the model constructed off the simulated data vs. the experimental. These models are then compared against one another until an ideal model is selected. c A schematic of the experimental automated feedback-controlled beam control experiment that allows for in situ monitoring and study of the atomic disordering of MoS2 using high temporal resolution. d The output of the models from the ELIT workflow is then fed through post-processing filters before feature finding can be performed.
The experimental data was acquired using an aberration corrected Nion UltraSTEM100 operating at 60 kV with a spatial resolution of ~1 Å. 60 kV for STEM imaging was chosen to limit the dose applied to the MoS2 slowing any defect generation kinetics as much as possible while still maintaining atomic resolution capabilities. The custom-built scanning feedback and electron beam control system was controlled using LabView software to acquire atomically resolved image datasets with the high angle annular dark field (HAADF) detector. A schematic of this system’s operation can be seen in Fig. 1c. The spiral beam pathway discussed previously was used for these experiments to maximize the temporal resolution without sacrificing spatial resolution. Prior to each fabrication step, predetermined parameters were set for the maximum number of spirals (i.e., the maximum e- irradiation dosage) as well as the spiral size and speed. These spirals are made to maintain content frequency drive signal to the scan coils to avoid frequency dependance. The dose profile can be controlled to a certain degree, but by default there is less dose in the periphery as there is a lower density of the scan path points at the edge of the FOV. For these experiments the spiral field of view was maintained at around 1.5 to 2.5 nm in diameter for atomic resolution, with a temporal resolution ranging from 5 to 100 fps (200-10 milliseconds) with a varying dwell time at each position dependent on the overall frame time. During the fabrication process, the beam control system acquired an image after each scan and plotted the average intensity of each frame, which, as the formation of defects began within the MoS2 monolayer, lowered as the process continued, supporting the notion of material ejection. After reaching a pre-determined intensity threshold, the fabrication process was stopped, and the experimental datasets were exported to study the atomic disordering that took place prior to nanopore formation. Due to the need to increase this system’s speed, the exported data results in 64×64 pixels per frame, therefore pre-processing of this data was needed prior to decoding through the DCNN models. The data was first resized to 512 × 512 pixels, and images were cropped appropriately to match the square image training data pixel to angstrom ratio of 20 pix/Å (Fig. 2a) before applying similar blurring and noise application as was previously used for the training data. For these datasets a gaussian blurring function with a standard deviation of 3 and a Chambolle noise applicator (with a weight of 0.3) were applied to the dataset frames. These settings were used to simulate high resolution STEM images taken with a high pixel to angstrom ration than the originally acquired rapid spiral scans (Fig. 2b).
Initial (i) and Final (ii) frames taken of the atomic disordering shown at different stages of decoding. a The raw experimental image data before any processing. b Pre-processing of the experimental data was performed by first denoising the data and then using a gaussian filter. c Post DCNN decoded image error variance between of identified atomic sites. d The decoded results of the ELIT workflow are plotted over the gaussian-filtered experimental frame. As seen, there are several sites that are labeled incorrectly, resulting in clusters of atoms that are not realistic. e The results of the nearest neighbor filter with the redundant points and incorrect labeling removed from the frame. Every frame shown has the same field of view of 2.56 × 2.56 nm.
With both simulated and experimental data acquired, the ensemble learning and iterative training workflow (ELIT) workflow, as demonstrated by Ghosh et al.47, was used to identify features from the low resolution data using these datasets as the initial training set and inputs respectively (Fig. 1b). This approach requires the training of the models on the material system being studies, in this case, MoS2, however after the development of the training data the models are robust at decoding datasets with sparse information resulting from the rapid acquisition. The use of multiple models within the workflow also allows for the generation of a mean ensemble prediction and uncertainty as a measure of confidence in the system. Using this mean uncertainty the area most associated with unexpected behavior can be highlighted for further study. The variance between the mean can be seen in Fig. 2c for the initial and final frame. From these images the edges around each atom sites are found to be the most uncertain as different models in the ensemble define the boundary of the atom site differently. From this variance it is also shown that the models have a harder time identifying the sulfur sites from one another as the conditions of each model in the ensemble vary in what is classified as a S1 vs S2 site. This is primarily where the errors in the decoding occur from. It is also worth noting that the sites along the edge of the frame are found to be much more uncertain due to edge effects from the blurring. However as a result of the upscaling of the data, this approach resulted in the overdetermination of the atomic sites, so to avoid this problem several post-processing steps were taken before features could be extracted. As Fig. 2d shows, the output from these models unrealistically oversampled the certain atomic sites on several frames, labeling overlapping atoms on single sites. A nearest neighbor filter was created and used to correct this issue as the last step in Fig. 1d. This filter consisted of three major steps: a nearest neighbor distance filter, using atom finding to locate the correct sites, and finally a filter refinement. The first step consisted of a nearest neighbor distance filter that found the clusters of problem sites by setting a distance threshold of \(\sim\)0.85 Å, which is about half the distance between the nearest atoms to avoid single sites being identified as different classes multiple times. This was followed by implementing atom finding through the STEMtool package developed by Mukherjee et al.66 on the decoded images for each class, being Mo sites (red points), projected S sites with two atoms (S2, yellow points), and projected S sites with only one S atom (S1, green points), to find the correct center of mass for each site. After these coordinates were found, they were cross-correlated with the clusters of decoded coordinates found from the first step, with the mislabeled sites being deleted. This left us with a fully decoded image and associated coordinate list for each frame within the dataset as shown in Fig. 2e. This decoded data could then be used for feature finding.
Effect of temporal resolution on atomic decoding
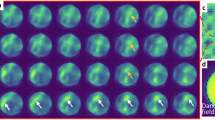
Due to the effectiveness of the ELIT workflow in decoding the experimental data, the temporal resolution was able to be pushed to much higher limits using this beam control system while still being capable of gathering structural information from the atomic disordering. Frames taken at different temporal resolutions and their associated decoded frames are shown in Fig. 3. Scan rates varying from 10 to 100 fps were performed using the external beam control system. The time between the initial and final frames were kept relatively consistent with total experiment times being 11.8, 9.5, 9.6, and 8.9 seconds respectively. The same beam conditions were used for all of these experimental spiral scans, therefore maintaining the same dose rate of \(1.6\times {10}^{5}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\cdot {\rm{s}}\), and only altering when/where the electron interact with the MoS2 lattice. Frames were then successfully decoded as seen in the ii and v frames, with red representing the Mo atoms, green the S2 sites, and blue the S1 sites. The processed data with the corrected coordinates can also be seen with the same color distinctions as used previously. The pre-processing and decoding begin to breakdown at 100 fps as the bright Mo are the only sites that can be reliably identified due to lack of current for each scan. This is due to the inherent nature of the abrupt and random shifts associated with HAADF imaging making the tracking of lighter atoms very susceptible to noise. At the slower scan speeds the atomic displacement of the lighter atomic species are confirmed to not be the result of noise artifacts until the 50 fps when the increase in noise leads to a greater degree in uncertainty in atom displacement tracking. However even at 50 fps, with each frame being acquired every 20 ms, atomic resolution is still fully achievable, and the S sites can be correctly identified as containing a VS (S1) or two S atoms (S2).
a–d Initial (i) and final (iv) frames of taken at different scan rates at 10, 20, 50 and 100 fps. These slides were then decoded into the Mo (red), S2 (green), and S1 (blue) classes in (ii) and (v) respectively. The corrected coordinates are overlaid onto the processed frames (iii) and (vi) respectively. Every frame has the same field of view of 1.6 × 1.6 nm.
With the image datasets fully decoded into the possible atomic sites, information can be obtained as to the stoichiometry, structure and atomic movement that takes place prior to the nanopore formation. By retaining the classifier information, the exact number of Mo and S atoms within the field of view being irradiated can be tracked and recorded along with the associated dose. Using these decoded frames the average dosage (an approximate beam current of 34 pA) required to cause initial atomic disordering that act as the nucleation site for the formation of sulfur single vacancy line (SVL) defects, point defects, and nanopores can be calculated and tracked for each experimental milling process. On average through the 50 experimental runs, we find that initial disordering takes a dose of \(3.3\pm 2.3\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\) to begin. After initial deformation, an additional \(1.3\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\) is needed to form SVLs, when they did form over 37 of the runs. From disordering to formation of a vacancy point defect, or hole, an additional dose of \(5.7\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\) is applied to the MoS2 field of view, which occurred during 38 runs. The dosage required to induce atomic transformations was found to generally be dependent on the speed of the scanning parameters with the slower scans with longer dwell times inducing atomic transformations more rapidly. This was also made evident from control experiments using a coherent beam with the same beam current, dose rate, and irradiated area which found that a much lower dose of \(4.3\times {10}^{5}\) and \(6.3\times {10}^{5}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\) were needed to initially disorder the lattice and then form a nanopore, respectively. This shows that the spiral scans used in this study help control the formation of defects within the lattice by slowing the disordering. However, the dosage required for each respective transformation was found to be irregular even within similar settings indicating other parameters outside of only dosage influencing the formation of these defects. Additionally, trends in the location of the initial disordering and defect can be studied in each milling dataset. As expected, the nucleation of the nanopore is entirely dependent on the location of the atomic disordering with the center of the frame requiring on average, lower dosage to begin forming in the center of the frame. When the initial defects begin forming at the edge of the frame, they take much longer to grow due to lower local dosage. It was also found that smaller fields of view result in the disordering further from the center of the field of the view.
Outside of general dosage trend, atomic tracking can be conducted between frames. Tracking is done by finding the closest identified atom between frames within a set distance threshold, if this distance is overcome, the atom has either been ejected or completely displaced. However, this threshold is large enough to observe the atomic shift both Mo and S species exhibit during disordering. Through observing these trajectories the trends in the movement of atoms relative to their neighbors can be studied and related to the dose applied to provide a more detail explanation of the respective atomic disordering and defect formation. The exact atomic movements seen before defect formation were dependent on the defect type. For example when forming SVL defects the atoms pulled together or apart and for forming holes the atoms that were not ejected shift away from the hole as the lattice shrinks around these edges. The dose required for the formation of these defect types were tracked and are discussed later. Several examples of the spatiotemporal trajectories of single atomic sites at different acquisition rates were gathered in a brush diagram in Fig. 4 allowing for the atomic disordering of both Mo (red) and S (blue for S2 site and green for S1 site or a single S vacancy) to be clearly observed. By tracking the classifier of the atom sites the formation of single and double VS is clearly shown as well.
a–d Analysis from 5, 10, 20 and 50 fps acquired milling processes. (i) The initial processed frame with the selected Mo and S site selected to be tracked. (ii) A three dimensional plot that tracks the location and movement and change in the class of the atomic site with red = Mo, blue = S2, and green = S1 site or single VS. (iii) The final processed frame showing the location of the atomic species or where the site was removed. Image FOV in (a) 3.0 × 3.0 nm and (b–d) 1.6 × 1.6 nm.
In Fig. 4a, the 5 fps experiment demonstrates the clear switch between S2 and S1 sites throughout the milling process seen in the tracking plot in Fig. 4b(ii). As a split in the lattice forms (the final result seen in Fig. 4a(iii)) prior to the opening of the nanopore the tracking become erratic and the distance between selected neighboring Mo and S sites increases before the S site disappears in some frames around 100 frames (20 s). Figure 4b shows a smaller field of view at a faster acquisition of 10 fps and clearly shows the formation of sulfur single vacancy line (SVL) defects. In the final frame in (iii) several of these SVL defects converge together before a nanopore begins to form. In the tracking plot in (ii) one can clearly see when the sulfur vacancy, that makes up one SVL forms, which is quickly followed by the Mo and S site to move closer together as SVLs cause a distortion as the lattice compresses around frame 50 (or 5 seconds). In the 20 fps experiment in Fig. 4c, the nanopore forms early in the top left corner of the field of view around frame 20 (or 1 s). As the nanopore grows, the Mo and S atom tracking can be seen to slowly shift as the lattice is compressed while the nanopore grows. Additionally, the S site remains S2 even after extended e- beam irradiation, this was likely due to the formation of the nanopore. Finally in the 50 fps experiment in Fig. 4d, the more random movement of the tracking column associated with higher temporal resolution better captures erratic nature of atomic motion within crystal structures while being exposed to e- beam irradiation. However at this point noise artifacts being introduced to the system by limited data leads to more uncertainty as the emergence of larger shifts in Fig. 4d(ii) become more common. All of these examples clearly emphasize that atomic structural information can be acquired even at 50 fps without relying upon highly specialized equipment or modifications to the electron optics. Using this workflow, atomic columns can be tracked and changes such as sites moving from S2 to S1 sites can be followed and correlated to the dosage and relationship to their nearest neighbors.
Full frame views of atom tracking can be plotted and trends in the data can be clearly seen to a greater degree than single atomic columns. Highlighted here are examples from 10 and 50 fps datasets in Fig. 5. In the 10 fps the shift in the later frames as the lattice is compressed on the right side of the FOV can be seen as the SVL grows before the formation of a line defect neighboring the SVL. In the top view of the 50 fps scan, one can observe the atomic disordering near the bottom of the field of view as a nanopore nucleates and then expands. Throughout the reaction the nearest neighbor distance of each location can be tracked demonstrating the capability to track atomic disordering and lattice distortion of the structure and directly tie these changes to the dosage applied to the system. Additionally, when observing the global data, as expected, it was found that the percentage of Mo and S that left the field of view prior to the formation of a nanopore was independent of the speed of acquisition. To form the nanopore, the number of Mo decreased by a percentage of 7 ± 9% while a lot more S left the system at 51 ± 9%, which was consistent through the 50 experimental runs. The small decrease and large error associated with the Mo atom count can be attributed to the atomic disordering and lattice shift that occur as a nanopore forms. When considering the decrease in S, this much larger percentage of S removal is to be expected as S is preferentially removed from the MoS2 lattice during electron beam irradiation. Additionally, this percentage makes theoretical sense as, after nanopore formation, a common edge reconstruction in the MoS2 system is the formation of the Mo6S6 nanowire edge structure16,21. Decreasing the local quantity of S by roughly 50% satisfies the local stoichiometry needed to facilitate the formation of this structure.
a, b Atom tracking of full frames (i-iv) of different scans taken at 10 and 50 fps respectively going from pristine lattice (i) to the disordering and formation of defects (ii-iii) until the formation of the nanopore (iv). The decoded tracking of the full frame of the atom sites, viewed from both the top (v) and side (vi) view to visualize atomic motion in the xy plane as well as in time. Image FOV in both (a) and (b) (i–iv) is 1.6×1.6 nm.
Utilizing the high temporal resolution of the spiral scan pathways, the intermediate frames of the milling process can reveal insights into defect nucleation, evolution and eventual nanopore formation. By combining the rescaled HAADF STEM image, decoded image and dosage maps associated with the frames, the dose per pixel, and therefore per atom site, can be accurately tracked and associated with defect formation and hole nucleation through all intermediate steps. As a result of this more rapid acquisition and a greater dispersal of the e- beam irradiation across the frame, it was statistically found that higher dosage was required to induce disordering, defect and nanopore generation at the faster scan rates, enabling the defect formation process to be more closely examined. For example, the dosage needed at 10 fps (21 runs) vs 33 fps (3 runs) respectively for atomic disordering (\(1.9\pm 2.6\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA }}{^{2}}\) vs \(4.6\pm 3.9\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\)), defect formation (\(1.7\pm 1.5\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\) vs \(7.0\pm 5.0\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\)), and nanopore generation (\(5.5\pm 4.8\times {10}^{6}\,{{\rm{e}}}^{-}/{\text{\AA} }^{2}\) vs \(1.5\pm 0.018\times {10}^{7}\,{{\rm{e}}}^{-}/{\text{\AA }}^{2}\)) all increased. The required dosage required to form the different defect structures as a function of the frame rate is demonstrated in the plot seen in Figure S1. This type of analysis it is also worth mentioning the larger amount of error as defect and nanopore generation was very dependent on individual experiments as factors that are still being controlled such as drift correction, as the drift experienced can be seen by the leaning nature of the tracked atomic columns in Fig. 5, to guarantee the relation between dosage and defect generation in future studies.
Discussion
In summary, this study demonstrates the successful enhancement of temporal resolution in STEM imaging without direct modifications to the electron microscope optics. By employing custom scan control with image processing techniques and implementing the ELIT workflow to develop a DCNN, a significant increase in the speed of determination of atomic position and elemental species information at high speeds in STEM imaging was achieved. This approach facilitated a more comprehensive investigation of the kinetics involved in the defect formation process in 2D materials. The application of DCNN-enabled decoding of low-quality, high-temporal-resolution data provided valuable insights into the atomic disordering and reordering process. Specifically, the experiments focused on the MoS2 materials system, utilizing a beam control system capable of data acquisition at scanning rates of up to 100 fps, resulting in small datasets with resolutions as low as 64x64 pixels. Through image processing, atomic resolution was attained, and the ELIT workflow successfully decoded the data into Mo, S2, and S1 classes, enabling accurate tracking of these elements throughout the milling process. While demonstrated here in 2D TMD materials, such an experimental workflow can be employed in a variety of materials with only the construction of DCNNs dependent on an initial labeled image. This workflow enables the extraction of more useful information from small datasets without compromising temporal or spatial resolution. The integration of machine learning and beam control techniques in this manner holds the potential for the development of different electron beam control atomic manipulation methods along with a deeper understanding of early defect creation, nanopore formation and growth, and atomic edge state formation in 2D materials. While the scan rates were limited to 100 fps due to the engineering limits of the detectors and scan coils utilized, through improvements of these systems such as event-based detectors67 or the use of improved scan coils68 or pump-probe systems69,70 much greater temporal resolution can be achieved. However, experiments taken here relied on the standard annular detectors and scan coils, only altering beam pathways. These findings have implications for advancing atomic fabrication processes and fostering autonomous atomic-level control for the purpose of tailoring localized functional properties at the atomic scale.
Methods
Sample preparation
Chemical vapor deposition was used to synthesize the MoS2 monolayers at a growth temperature of 750 °C using a hinged tube furnace (Lindberg/Blue) equipped with a 2 in. diameter quartz tube. SiO2 (300 nm)/Si was used as a growth substrate. These substrates were first cleaned with isopropanol and then spin-coated with perylene-3,4,9,10- tetracarboxylic acid tetrapotassium salt before being dried and placed face-down above an alumina crucible containing MoO3 powder (∼5 mg) (99.9%, Sigma-Aldrich). This crucible was then inserted into the center of the quartz tube. Another crucible was filled with S powder (∼0.7 g) (99.998%, Sigma-Aldrich) and inserted at a region located 20 cm upstream from the other crucible in the quartz tube. When the center of the tube reaches 750 °C center of the tube, the temperature at the S powder crucible was around 180 °C. The tube was then evacuated to ∼5 mTorr. At this point a 70 sccm (standard cubic centimeters per minute) flow of Ar at atmospheric pressure was established. The furnace was heated to 750 °C with a ramping rate of 30 °C/min and held at this temperature for 4 − 6 min. The furnace is allowed to naturally cool down to room temperature.
MoS2 monolayers were then transferred from the SiO2/Si substrate onto gold mesh holey carbon Quantifoil TEM grids using a wet transfer process for STEM imaging. PMMA (Poly(methyl methacrylate)) was spun coated onto the SiO2/Si substrate with monolayer MoS2 crystals on the surface at 500 rpm for 15 s followed by 3000 rpm for 50 s. The PMMA-coated sample was then placed into a 1 M KOH solution to remove the SiO2 substrate. This leaves the PMMA film with the MoS2 monolayer on the solution surface. The KOH residue was then rinsed from the sample film with DI water. A TEM grid was then placed into a funnel filled with DI water. Using a glass slide the washed PMMA/MoS2 film was then transferred to float on the DI water. As the water drains the film is then placed onto the TEM grid. The samples are then baked to dry at 80 °C before being placed into acetone to soak for 12 hours to remove remaining PMMA before a final rinse in IPA, leaving a clean monolayer surface. Prior to STEM experiments, the specimens were baked at 160 °C in vacuum overnight to reduce surface contamination.
STEM imaging
An aberration corrected Nion UltraSTEM100 electron microscope was used for all the STEM imaging experiments and operated at 60 kV, with a probe current of ~34 pA, and convergence semi-angle of 30 mrad. The custom-built scanning feedback control system described in this study was based on a National Instruments DAQ (PXIe-6124) and field-programmable gate array (FPGA) system (PXIe-7856R) in a PXIe-1073 chassis that was interfaced with the beam control system of the Nion UltraSTEM100. Input coordinates for spiral STEM scans were generated using customizable MatLab code then input into LabView to control STEM scan coils for positioning the focused STEM probe along well-defined spiral scan pathways. The custom scan control system has a maximum IO rate of 1 M Samples/s.
Data availability
Sample example experimental and simulated datasets along with the model used are available through the Python notebook located at https://github.com/mgboebinger/FaSTEM/.
Code availability
The functions used to simulate structures from DL predictions can be found at https://github.com/pycroscopy/atomai. The details of training DL networks used in this work are available through the python notebooks located at https://github.com/aghosh92/ELIT. The refinement procedure and temporal tracking can be found in https://github.com/mgboebinger/FaSTEM/. Additional refinement done through the use of the stemtool package found at https://github.com/stemtool/stemtool and simulated STEM images formed using the abTEM Python API found at https://github.com/abTEM/abTEM.
References
Iannaccone, G. et al. Quantum engineering of transistors based on 2D materials heterostructures. Nat. Nanotechnol. 13, 183–191 (2018).
Wyrick, J. et al. Atom‐by‐Atom Fabrication of Single and Few Dopant Quantum Devices. Adv. Funct. Mater. 29, 1903475 (2019).
Fölsch, S. et al. Quantum dots with single-atom precision. Nat. Nanotechnol. 9, 505–508 (2014).
Narang, P. et al. Quantum materials with atomic precision: artificial atoms in solids: ab initio design, control, and integration of single photon emitters in artificial quantum materials. Adv. Funct. Mater. 29, 1904557 (2019).
Kalinin, S. V., Borisevich, A. Y. & Jesse, S. Fire up the atom forge. Nature 539, 485–487 (2016).
Dyck, O. et al. Building structures atom by atom via electron beam manipulation. Small 14, 1801771–1801779 (2018).
Tai, K. L. et al. Atomic‐Scale Fabrication of In‐Plane Heterojunctions of Few‐Layer MoS2 via In Situ Scanning Transmission Electron Microscopy. Small 16, 1905516 (2020).
Dyck, O. et al. Variable voltage electron microscopy: Toward atom-by-atom fabrication in 2D materials. Ultramicroscopy 211, 112949 (2020).
Silver, R., et al. Atomic precision fabrication of quantum devices down to the single atom regime. SPIE Advanced Lithography. Vol. 11610. 2021: SPIE.
Khajetoorians, A. A. et al. Creating designer quantum states of matter atom-by-atom. Nat. Rev. Phys. 1, 703–715 (2019).
Su, C. et al. Engineering single-atom dynamics with electron irradiation. Sci. Adv. 5, eaav2252 (2019).
Susi, T. et al. Towards atomically precise manipulation of 2D nanostructures in the electron microscope. 2D Mater. 4, 1–042009 (2017).
Susi, T., Meyer, J. C. & Kotakoski, J. Manipulating low-dimensional materials down to the level of single atoms with electron irradiation. Ultramicroscopy 180, 163–172 (2017).
Tripathi, M. et al. Electron-beam manipulation of silicon dopants in graphene. Nano Lett. 18, 5319–5323 (2018).
Zagler, G. et al. Beam-driven dynamics of aluminium dopants in graphene. 2D Mater. 9, 035009 (2022).
Boebinger, M. G. et al. The atomic drill bit: Precision controlled atomic fabrication of 2D materials. Adv. Mater. 2210116 (2023).
Ritter, K. A. & Lyding, J. W. The influence of edge structure on the electronic properties of graphene quantum dots and nanoribbons. Nat. Mater. 8, 235–242 (2009).
Cheng, F. et al. Controlled growth of 1D MoSe2 nanoribbons with spatially modulated edge states. Nano Lett. 17, 1116–1120 (2017).
Xu, H. et al. Oscillating edge states in one-dimensional MoS2 nanowires. Nat. Commun. 7, 12904 (2016).
Hu, G. et al. Work Function Engineering of 2D Materials: The Role of Polar Edge Reconstructions. J. Phys. Chem. Lett. 12, 2320–2326 (2021).
Sang, X. et al. In situ edge engineering in two-dimensional transition metal dichalcogenides. Nat. Commun. 9, 2051 (2018).
Magda, G. Z. et al. Room-temperature magnetic order on zigzag edges of narrow graphene nanoribbons. Nature 514, 608–611 (2014).
Saab, M. & Raybaud, P. Tuning the magnetic properties of MoS2 single nanolayers by 3d metals edge doping. J. Phys. Chem. C. 120, 10691–10697 (2016).
Yin, X. et al. Edge nonlinear optics on a MoS2 atomic monolayer. Science 344, 488–490 (2014).
Fung, V. et al. Inverse design of two-dimensional materials with invertible neural networks. npj Comput. Mater. 7, 200 (2021).
Gutiérrez, H. R. et al. Extraordinary room-temperature photoluminescence in triangular WS2 monolayers. Nano Lett. 13, 3447–3454 (2013).
Mannix, A. J. et al. Synthesis and chemistry of elemental 2D materials. Nat. Rev. Chem. 1, 0014 (2017).
Dong, R., Zhang, T. & Feng, X. Interface-assisted synthesis of 2D materials: trend and challenges. Chem. Rev. 118, 6189–6235 (2018).
Qi, Z. J. et al. Correlating atomic structure and transport in suspended graphene nanoribbons. Nano Lett. 14, 4238–4244 (2014).
Dyck, O. et al. Atom-by-atom fabrication with electron beams. Nat. Rev. Mater. 4, 497–507 (2019).
Dyck, O. et al. Direct matter disassembly via electron beam control: electron-beam-mediated catalytic etching of graphene by nanoparticles. Nanotechnology 31, 245303 (2020).
Hudak, B. M. et al. Directed atom-by-atom assembly of dopants in silicon. ACS Nano 12, 5873–5879 (2018).
Susi, T., Meyer, J. C. & Kotakoski, J. Quantifying transmission electron microscopy irradiation effects using two-dimensional materials. Nat. Rev. Phys. 1, 397–405 (2019).
Kretschmer, S. et al. Formation of defects in two-dimensional MoS2 in the transmission electron microscope at electron energies below the knock-on threshold: the role of electronic excitations. Nano Lett. 20, 2865–2870 (2020).
Komsa, H.-P. et al. Two-dimensional transition metal dichalcogenides under electron irradiation: Defect production and doping. Phys. Rev. Lett. 109, 1–5 (2012).
Yoshimura, A. et al. First-principles simulation of local response in transition metal dichalcogenides under electron irradiation. Nanoscale 10, 2388–2397 (2018).
Yoshimura, A. et al. Quantum theory of electronic excitation and sputtering by transmission electron microscopy. Nanoscale 15, 1053–1067 (2023).
Lingerfelt, D. B. et al. Understanding beam-induced electronic excitations in materials. J. Chem. Theory Comput. 16, 1200–1214 (2020).
Wang, S. et al. Detailed atomic reconstruction of extended line defects in monolayer MoS2. ACS nano 10, 5419–5430 (2016).
Chen, Q. et al. Ultralong 1D vacancy channels for rapid atomic migration during 2D void formation in monolayer MoS2. ACS nano 12, 7721–7730 (2018).
Chen, J. et al. In situ high temperature atomic level dynamics of large inversion domain formations in monolayer MoS 2. Nanoscale 11, 1901–1913 (2019).
Lin, Y.-C. et al. Atomic mechanism of the semiconducting-to-metallic phase transition in single-layered MoS2. Nat. Nanotechnol. 9, 391–396 (2014).
Hu, G. et al. Superior electrocatalytic hydrogen evolution at engineered non-stoichiometric two-dimensional transition metal dichalcogenide edges. J. Mater. Chem. A 7, 18357–18364 (2019).
Lin, J. et al. Flexible metallic nanowires with self-adaptive contacts to semiconducting transition-metal dichalcogenide monolayers. Nat. Nanotechnol. 9, 436–442 (2014).
Murugan, P. et al. Assembling nanowires from Mo−S clusters and effects of iodine doping on electronic structure. Nano Lett. 7, 2214–2219 (2007).
Kibsgaard, J. et al. Atomic-scale structure of Mo6S6 nanowires. Nano Lett. 8, 3928–3931 (2008).
Ghosh, A. et al. Ensemble learning-iterative training machine learning for uncertainty quantification and automated experiment in atom-resolved microscopy. npj Comput. Mater. 7, 100 (2021).
Ziatdinov, M. et al. AtomAI framework for deep learning analysis of image and spectroscopy data in electron and scanning probe microscopy. Nat. Mach. Intell. 4, 1101–1112 (2022).
Jesse, S. et al. Direct atomic fabrication and dopant positioning in Si using electron beams with active real-time image-based feedback. Nanotechnology 29, 255303 (2018).
Sang, X. et al. Precision controlled atomic resolution scanning transmission electron microscopy using spiral scan pathways. Sci. Rep. 7, 43585 (2017).
Sang, X. et al. Dynamic scan control in STEM: spiral scans. Adv. Struct. Chem. Imaging 2, 1–8 (2016).
Krizhevsky, A. & Hinton G. Learning multiple layers of features from tiny images. (2009).
Deng, J. et al. Imagenet: A large-scale hierarchical image database. in 2009 IEEE conference on computer vision and pattern recognition. 2009. Ieee.
Ziatdinov, M., Maksov, A. & Kalinin, S. V. Learning surface molecular structures via machine vision. npj Comput. Mater. 3, 31 (2017).
Choe, J. et al. Direct imaging of structural disordering and heterogeneous dynamics of fullerene molecular liquid. Nat. Commun. 10, 4395 (2019).
Hoelzel, H. et al. Time-resolved imaging and analysis of the electron beam-induced formation of an open-cage metallo-azafullerene. Nat. Chem. 15, 1444–1451 (2023).
Rashidi, M. & Wolkow, R. A. Autonomous scanning probe microscopy in situ tip conditioning through machine learning. ACS nano 12, 5185–5189 (2018).
Yang, S.-H. et al. Deep Learning-Assisted Quantification of Atomic Dopants and Defects in 2D Materials. Adv. Sci. 8, 2101099 (2021).
Lee, K. et al. STEM Image Analysis Based on Deep Learning: Identification of Vacancy Defects and Polymorphs of MoS2. Nano Lett. 22, 4677–4685 (2022).
Ziatdinov, M. et al. Deep learning of atomically resolved scanning transmission electron microscopy images: chemical identification and tracking local transformations. ACS nano 11, 12742–12752 (2017).
Gordon, O. M. et al. Automated searching and identification of self-organized nanostructures. Nano Lett. 20, 7688–7693 (2020).
Horwath, J. P. et al. Understanding important features of deep learning models for segmentation of high-resolution transmission electron microscopy images. npj Comput. Mater. 6, 108 (2020).
Lee, C.-H. et al. Deep learning enabled strain mapping of single-atom defects in two-dimensional transition metal dichalcogenides with sub-picometer precision. Nano Lett. 20, 3369–3377 (2020).
Madsen, J. & Susi, T. The abTEM code: transmission electron microscopy from first principles. Open Res. Eur. 1, 24 (2021).
Madsen, J. & Susi, T. abTEM: ab Initio Transmission Electron Microscopy Image Simulation. Microsc. Microanalysis 26, 448–450 (2020).
Mukherjee, D. & Unocic, R. STEMTooL: An Open Source Python Toolkit for Analyzing Electron Microscopy Datasets. Microsc. Microanalysis 26, 2960–2962 (2020).
Peters, J. J. P. et al. Electron counting detectors in scanning transmission electron microscopy via hardware signal processing. Nat. Commun. 14, 5184 (2023).
Ishikawa, R. et al. High spatiotemporal-resolution imaging in the scanning transmission electron microscope. Microsc. (Oxf.) 69, 240–247 (2020).
Fu, X. et al. Direct visualization of electromagnetic wave dynamics by laser-free ultrafast electron microscopy. Sci. Adv. 6, eabc3456 (2020).
Fu, X. et al. Nanoscale-femtosecond dielectric response of Mott insulators captured by two-color near-field ultrafast electron microscopy. Nat. Commun. 11, 5770 (2020).
Acknowledgements
STEM experiments were performed at the Center for Nanophase Materials Sciences (CNMS), which is a US Department of Energy, Office of Science User Facility at Oak Ridge National Laboratory. The synthesis of MoS2 and development of the scan control was supported by the U.S. Department of Energy, Office of Basic Energy Sciences, Division of Materials Sciences and Engineering. This manuscript has been authored by UT-Battelle, LLC, under Contract No. DE-AC0500OR22725 with the U.S. Department of Energy. The United States Government retains and the publisher, by accepting the article for publication, acknowledges that the United States Government retains a non-exclusive, paid-up, irrevocable, world-wide license to publish or reproduce the published form of this manuscript, or allow others to do so, for the United States Government purposes. The Department of Energy will provide public access to these results of federally sponsored research in accordance with the DOE Public Access Plan (http://energy.gov/downloads/doe-public-access-plan).
Author information
Authors and Affiliations
Contributions
M.G.B. conducted STEM experiments, constructed DCNN, performed data analysis, and was the primary contributor in writing the manuscript. S.M., K.M.R. and A.G. provided assistance and guidance with the coding and data analysis with additional support in the construction of the DCNN from A.G., M.Z., and S.V.K.; K.X. synthesized MoS2 samples used in this study. S.J. provided experimental assistance with the beam control system during STEM experiments. RRU provided supervision and guidance for STEM experiments. All authors contributed to discussions and the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.V.K. is affiliated with npj Computation Materials as an Associate Editor. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Boebinger, M.G., Ghosh, A., Roccapriore, K.M. et al. Exploring electron-beam induced modifications of materials with machine-learning assisted high temporal resolution electron microscopy. npj Comput Mater 10, 260 (2024). https://doi.org/10.1038/s41524-024-01448-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41524-024-01448-7