Abstract
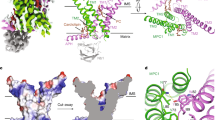
MPC1 and MPC2 are two well-known components of the mitochondrial pyruvate carrier (MPC) complex maintaining MPC activity to transport pyruvate into mitochondria for tricarboxylic acid (TCA) cycle entry in mammalian cells. It is currently unknown whether there is an additional MPC component crucially maintaining MPC complex activity for pyruvate mitochondrial import. Here we show that ALDH4A1, a proline-metabolizing enzyme localized in mitochondria, serves as a previously unrecognized MPC component maintaining pyruvate mitochondrial import and the TCA cycle independently of its enzymatic activity. Loss of ALDH4A1 in mammalian cells impairs pyruvate entry to mitochondria, resulting in defective TCA cycle entry. ALDH4A1 forms an active trimeric complex with MPC1–MPC2 to maintain the integrity and oligomerization of MPC1–MPC2 and facilitates pyruvate transport in an in vitro system. ALDH4A1 displays tumour suppression by maintaining MPC complex activity. Our study identifies ALDH4A1 as an essential component of MPC for pyruvate mitochondrial import, TCA cycle entry and tumour suppression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data supporting the findings of this study are available from the corresponding author on reasonable request. Metabolomics and mass spectrometry data are available via Figshare at https://figshare.com/s/ddce7916d4d7b38e8b3b (ref. 47) (metabolomics) and https://figshare.com/s/749b9763ce937570f931 (ref. 48) (mass spectrometry). Source data are provided with this paper.
References
Gray, L. R., Tompkins, S. C. & Taylor, E. B. Regulation of pyruvate metabolism and human disease. Cell Mol. Life Sci. 71, 2577–2604 (2014).
Olson, K. A., Schell, J. C. & Rutter, J. Pyruvate and metabolic flexibility: illuminating a path toward selective cancer therapies. Trends Biochem. Sci. 41, 219–230 (2016).
Bergers, G. & Fendt, S. M. The metabolism of cancer cells during metastasis. Nat. Rev. Cancer 21, 162–180 (2021).
Elia, I. et al. Breast cancer cells rely on environmental pyruvate to shape the metastatic niche. Nature 568, 117–121 (2019).
Bricker, D. K. et al. A mitochondrial pyruvate carrier required for pyruvate uptake in yeast, Drosophila, and humans. Science 337, 96–100 (2012).
Herzig, S. et al. Identification and functional expression of the mitochondrial pyruvate carrier. Science 337, 93–96 (2012).
Rauckhorst, A. J. & Taylor, E. B. Mitochondrial pyruvate carrier function and cancer metabolism. Curr. Opin. Genet. Dev. 38, 102–109 (2016).
Vanderperre, B., Bender, T., Kunji, E. R. & Martinou, J. C. Mitochondrial pyruvate import and its effects on homeostasis. Curr. Opin. Cell Biol. 33, 35–41 (2015).
Bader, D. A. et al. Mitochondrial pyruvate import is a metabolic vulnerability in androgen receptor-driven prostate cancer. Nat. Metab. 1, 70–85 (2019).
Vanderperre, B. et al. Embryonic lethality of mitochondrial pyruvate carrier 1 deficient mouse can be rescued by a ketogenic diet. PLoS Genet. 12, e1006056 (2016).
Vigueira, P. A. et al. Mitochondrial pyruvate carrier 2 hypomorphism in mice leads to defects in glucose-stimulated insulin secretion. Cell Rep. 7, 2042–2053 (2014).
Schell, J. C. et al. A role for the mitochondrial pyruvate carrier as a repressor of the Warburg effect and colon cancer cell growth. Mol. Cell 56, 400–413 (2014).
Zou, H. et al. MPC1 deficiency accelerates lung adenocarcinoma progression through the STAT3 pathway. Cell Death Dis. 10, 148 (2019).
Tian, G. A. et al. MPC1 deficiency promotes CRC liver metastasis via facilitating nuclear translocation of β-catenin. J. Immunol. Res. 2020, 8340329 (2020).
Bensard, C. L. et al. Regulation of tumor initiation by the mitochondrial pyruvate carrier. Cell Metab. 31, 284–300 e287 (2020).
Tang, X. P. et al. Mitochondrial pyruvate carrier 1 functions as a tumor suppressor and predicts the prognosis of human renal cell carcinoma. Lab. Invest. 99, 191–199 (2019).
Li, X. et al. MPC1 and MPC2 expressions are associated with favorable clinical outcomes in prostate cancer. BMC Cancer 16, 894 (2016).
D’Aniello, C., Patriarca, E. J., Phang, J. M. & Minchiotti, G. Proline metabolism in tumor growth and metastatic progression. Front. Oncol. 10, 776 (2020).
Elia, I. et al. Proline metabolism supports metastasis formation and could be inhibited to selectively target metastasizing cancer cells. Nat. Commun. 8, 15267 (2017).
Phang, J. M. & Liu, W. Proline metabolism and cancer. Front. Biosci. 17, 1835–1845 (2012).
Phang, J. M., Liu, W., Hancock, C. N. & Fischer, J. W. Proline metabolism and cancer: emerging links to glutamine and collagen. Curr. Opin. Clin. Nutr. Metab. Care 18, 71–77 (2015).
Pemberton, T. A. et al. Structural studies of yeast delta(1)-pyrroline-5-carboxylate dehydrogenase (ALDH4A1): active site flexibility and oligomeric state. Biochemistry 53, 1350–1359 (2014).
Tanner, J. J. Structural biology of proline catabolic enzymes. Antioxid. Redox Signal. 30, 650–673 (2019).
Morris, S. M. Jr. Regulation of enzymes of the urea cycle and arginine metabolism. Annu. Rev. Nutr. 22, 87–105 (2002).
Geraghty, M. T. et al. Mutations in the delta1-pyrroline 5-carboxylate dehydrogenase gene cause type II hyperprolinemia. Hum. Mol. Genet. 7, 1411–1415 (1998).
Valle, D., Goodman, S. I., Applegarth, D. A., Shih, V. E. & Phang, J. M. Type II hyperprolinemia. Delta1-pyrroline-5-carboxylic acid dehydrogenase deficiency in cultured skin fibroblasts and circulating lymphocytes. J. Clin. Invest. 58, 598–603 (1976).
Srivastava, D. et al. The three-dimensional structural basis of type II hyperprolinemia. J. Mol. Biol. 420, 176–189 (2012).
Servet, C., Ghelis, T., Richard, L., Zilberstein, A. & Savoure, A. Proline dehydrogenase: a key enzyme in controlling cellular homeostasis. Front. Biosci. 17, 607–620 (2012).
Du, J., Zhu, S., Lim, R. R. & Chao, J. R. Proline metabolism and transport in retinal health and disease. Amino Acids 53, 1789–1806 (2021).
Tavoulari, S. et al. The yeast mitochondrial pyruvate carrier is a hetero-dimer in its functional state. EMBO J. 38, e100785 (2019).
Compan, V. et al. Monitoring mitochondrial pyruvate carrier activity in real time using a BRET-based biosensor: investigation of the Warburg effect. Mol Cell 59, 491–501 (2015).
Brinda, K. V. & Vishveshwara, S. Oligomeric protein structure networks: insights into protein–protein interactions. BMC Bioinform. 6, 296 (2005).
Nooren, I. M. & Thornton, J. M. Diversity of protein–protein interactions. EMBO J. 22, 3486–3492 (2003).
Bender, T. & Martinou, J. C. The mitochondrial pyruvate carrier in health and disease: to carry or not to carry? Biochim. Biophys. Acta 1863, 2436–2442 (2016).
Nagampalli, R. S. K. et al. Human mitochondrial pyruvate carrier 2 as an autonomous membrane transporter. Sci. Rep. 8, 3510 (2018).
Halestrap, A. P. The mechanism of the inhibition of the mitochondrial pyruvate transportater by alpha-cyanocinnamate derivatives. Biochem. J. 156, 181–183 (1976).
Zhang, W. et al. Lactate is a natural suppressor of RLR signaling by targeting MAVS. Cell 178, 176–189 (2019).
Huangyang, P. & Simon, M. C. Hidden features: exploring the non-canonical functions of metabolic enzymes. Dis. Model. Mech. 11, dmm033365 (2018).
Xu, L., Phelix, C. F. & Chen, L. Y. Structural insights into the human mitochondrial pyruvate carrier complexes. J. Chem. Inform. Model. 61, 5614–5625 (2021).
Chan, C. H. et al. Pharmacological inactivation of Skp2 SCF ubiquitin ligase restricts cancer stem cell traits and cancer progression. Cell 154, 556–568 (2013).
Xu, C. et al. SIRPgamma-expressing cancer stem-like cells promote immune escape of lung cancer via Hippo signaling. J. Clin. Invest. 132, e141797 (2022).
Cai, Z. et al. Phosphorylation of PDHA by AMPK drives TCA cycle to promote cancer metastasis. Mol. Cell 80, 263–278 (2020).
Wei, M. C. et al. tBID, a membrane-targeted death ligand, oligomerizes BAK to release cytochrome C. Genes Dev. 14, 2060–2071 (2000).
Wittig, I., Braun, H. P. & Schagger, H. Blue native PAGE. Nat. Protoc. 1, 418–428 (2006).
Hsu, C. C. et al. Inositol serves as a natural inhibitor of mitochondrial fission by directly targeting AMPK. Mol. Cell 81, 3803–3819 (2021).
Hsu, C. C. et al. IMPA1-derived inositol maintains stemness in castration-resistant prostate cancer via IMPDH2 activation. J Exp. Med. 221, e20231832 (2024).
Hsu, C.-C. et al. Metabolomics_ALDH4A1.xlsx. Figshare https://figshare.com/s/ddce7916d4d7b38e8b3b (2025).
Hsu, C.-C. et al. Proteomics_IP_ALDH4A1.xlsx. Figshare https://figshare.com/s/749b9763ce937570f931 (2025).
Acknowledgements
We thank J.-C. Martinou for sharing several key plasmids in this study. We acknowledge the support of the Wake Forest Baptist Comprehensive Cancer Center Cell & Cellular Imaging & Flow Cytometry Shared Resources, supported by the National Cancer Institute’s Cancer Center Support Grant (P30CA012197). We also acknowledge the support of Proteomics Core from Duke University School of Medicine. This research was supported partly by start-ups from Wake Forest University School of Medicine and Duke University School of Medicine, Endowed Professor Funds from Anderson Discovery Professor for Cancer Research and Fred and Janet Sanfilippo Distinguished Professor, and NIH grants (R01CA256158, R01CA248037 and R01CA270617) to H.K.L. The funders had no role in the study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
C.-C.H., C.-Y.W., R.K.M. and H.-K.L. designed experiments and wrote and edited the manuscript. C.-C.H., C.-Y.W., R.K.M. and Z.C. performed experiments and analysed the data for all figures. C.-C.H., R.K.M., Z.C., V.P., R.K., L.B., B.-S.P., T.C., Y.-R.C, H.-E.W., Y.J., H.G. and C.-Y.L provided technical support, comments and suggestions.
Corresponding author
Ethics declarations
Competing interests
H.K.L. is a consultant for Stablix, Inc. and Chang Gung University of Science and Technology. All other authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Luca Scorrano and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 ALDH4A1 protein levels are downregulated in various human cancers and its loss predicts poor survival outcome.
(a, b) The gene expression of ALDH4A1 patterns across diverse human cancer and normal tissues from GENT (gene expression database of normal and tumor tissues). Significant test results P < 0.001 by two-sample t-test were indicated on web-accessible database of GENT (http://gent2.appex.kr/gent2/). The centre lines in the boxes represent median values. The box edges, upper and lower whiskers indicate the interquartile range (IQR; from the 1 to 3.7), the largest value smaller than 3.7 and the smallest value larger than 1 (a) and Oncomine publicly accessible database (b) P < 0.001 by Mann-Whitney U test. (c) The overall survival rate of patients with high or low ALDH4A1 expression from PrognoScan. P < 0.05 by long-rank test. (d) The database extracted from Kaplan-Meier Plotter was subjected to Kaplan-Meier analysis. (e) The percentage of cases from TCGA database (https://portal.gdc.cancer.gov) indicates loss of copy number variation (CNV) of ALDH4A1 in several cancer types including adenomas and adenocarcinomas from brain, liver, lung, stomach, pancreas, bladder, kidney, etc.
Extended Data Fig. 2 ALDH4A1 suppresses cancer cell growth, migration, sphere formation.
(a-d) Representative images (a and c) and quantification of cancer cell migration (b) and sphere formation (d) for Hep3B cells upon vec, 4A1WT, shLuc, sh4A1 (#1 and #2) and sh4A1 #1 with 4A1WT. (e-h) Representative images (e and g) and quantification (f and h) of cancer cell migration (e) and sphere formation (g) for Huh7 cells upon vec, 4A1WT, shLuc, sh4A1 (#1 and #2) and sh4A1 #1 with 4A1WT. Data are presented as mean ± s.d. by one-way ANOVA with Bonferroni post-hoc tests) from 3 biological replicates (n = 3); ***P < 0.001, compared to the shLuc. **, P < 0.01, ***, P < 0.001.
Extended Data Fig. 3 ALDH4A1 loss promotes cancer cell migration, cancer sphere formation, cell transformation and orthotopic liver cancer formation.
(a) Immunoblotting of HepG2 cells upon shLuc, sh4A1 (#1 and #2), shp53 (#1 and #2), and shRb (#1 and #2) using indicated antibodies. (b-e) Representative images and quantification of cancer cell migration (b and d) and sphere formation (c and e) for HepG2 cells upon vec, 4A1WT, shLuc, sh4A1 (#1 and #2) and sh4A1 #1 with 4A1WT, shp53 (#1 and #2) and shRb (#1 and #2). Data are presented as mean ± s.d. by one-way ANOVA with Bonferroni post-hoc tests from 3 biological replicates (n = 3); ***P < 0.001, compared to the shLuc. **, P < 0.01, ***, P < 0.001.
Extended Data Fig. 4 ALDH4A1 loss promotes cell transformation and orthotopic liver cancer formation.
(a, c) Representative images (a) and relative number of soft agar colony assay (c) for NIH3T3 cells upon shLuc, sh4A1#1, sh4A1#1 with restoration of 4A1WT. Data from (c) are presented as mean ± s.d by two-tailed unpaired t-test from 3 biological replicates (n = 3). **, P < 0.01. (b) Cell lysates from NIH3T3 cells upon shLuc, sh4A1#1 or sh4A1#1 with restoration of 4A1WT were subjected to immunoblotting assay with indicated antibodies. (d) Hep3B cells upon shGFP with luciferase (Luc) and sh4A1 (#1 and #2) with Luc were orthotopically injected into liver, followed by IVIS imaging after 4 weeks. Representative rainbow images of luminescence in livers of nude mice were shown. Rainbow scales are expressed in radiance (p/sec/cm2/sr). (e) Quantification of average radiance (p/sec/cm2/sr) from 5 mice in each group. Data are presented as mean ± s.d by two-tailed unpaired t-test from 5 biological replicates (n = 5). **, P < 0.01. *, P < 0.05.
Extended Data Fig. 5 Increased proline, ornithine or arginine levels do not account for cancer cell proliferation regulated by ALDH4A1.
(a, b, c) Colony formation assay in Hep3B cells treated with proline (a), ornithine (b) and arginine (c) with indicated concentration in Vec, 4A1WT and shLuc, sh4A1 (#1, #2) or sh4A1#1 restored with 4A1WT. (d) Cell proliferation of control and PRODH knockdown (shPRODH #1 and shPRODH #2) Hep3B were determined. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 5 biological replicates (n = 5). Data is no significance between PRODH knockdown and control cells.
Extended Data Fig. 6 ALDH4A1 is mainly localized in mitochondria and critically maintains mitochondrial pyruvate levels and mitochondrial oxidative phosphorylation independently of its enzymatic activity.
(a) Immunoblotting of the fractionation of whole cell lysates (WCL), cytoplasm and mitochondria from stably expressing shLuc or shALDH4A1 Hep3B cells using indicated antibodies. (b) Immunoblotting of mitochondrial fractions from stably expressing control or ALDH4A1 knockdown Hep3B cells using indicated antibodies. The representative data are shown from two independent experiments. (c, e, f) Pyruvate level in the cytosolic and mitochondrial extracts of shLuc or shALDH4A1 (c), treatment of DMSO (vehicle control) or MPC1 inhibitor UK5099 (0.1 mM) for 24 hr (e), and shLuc or shMPC1 (f) Hep3B cells were determined. Data are presented as mean ± s.d by two-tailed unpaired t-test from 3 biological replicates (n = 3). *, P < 0.05; **, P < 0.01; ***, P < 0.001. (d) The pyruvate levels in mitochondria and cytosol in control, ALDH4A1 knockdown, and ALDH4A1 knockdown with ALDH4A1WT or ALDH4A1S352L mutant restoration Hep3B cells. Data are presented as mean ± s.d. by one-way ANOVA with Bonferroni post-hoc tests from 3 biological replicates (n = 3); Red is cytosol part, and blue i smitochondria part. ***P < 0.001 for cytosol and mitochondria part; NS, not significant. (g) maximal respiratory capacity from (a) was calculated by OCR measurement at basal levels and upon FCCP treatment subtracted by non-mitochondrial respiratory. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 3 biological replicates (n = 3). ***, P < 0.001. (h) The OCR assay was performed in 5 μg of isolated mitochondria from Hep3B cells upon shLuc, sh4A1, or sh4A1 with restoration of either 4A1WT or 4A1 enzymatic dead mutant (4A1S352L) in a coupling state with the substrates, 10 mM succinate and 2 μM rotenone, followed by 4 mM ADP, 2.5 μg/ml oligomycin (oligo), 4 μM FCCP, or 4 μM Antimycin (AA) treatment. Data are presented as mean ± s.d. by two-way ANOVA from 5 biological replicates (n = 5). NS, non-significant. (i, j) RCR were calculated by state 3/state 4o (i) and state 3 u/state 4o (j) from (h). Data are presented as mean ± s.d. by two-tailed unpaired t-test from 5 biological replicates (n = 5). NS, non-significant. (k) The OCR assay was performed in 5 μg of isolated mitochondria from Hep3B cells upon shLuc, sh4A1, or sh4A1 with restoration of either 4A1WT or 4A1 enzymatic dead mutant (4A1S352L) in a coupling state with the substrates, 5 mM pyruvate and 5 mM malate, followed by 4 mM ADP, 2.5 μg/ml oligomycin (oligo), 4 μM FCCP, or 4 μM Antimycin (AA) treatment. Data are presented as mean ± s.d. by two-way ANOVA from 5 biological replicates (n = 5). NS, non-significant. shLuc vs. sh4A1, ***P < 0.001; sh4A1 vs. sh4A1 + 4A1WT, ***P < 0.001; sh4A1 vs. sh4A1 + 4A1S352L, ***P < 0.001. (l, m) RCR were calculated by state 3/state 4o (l) and state 3 u/state 4o (m) from (k). Data are presented as mean ± s.d. by two-tailed unpaired t-test from 5 biological replicates (n = 5). **, P < 0.01; ***, P < 0.001.
Extended Data Fig. 7 ALDH4A1 loss does not affect pyruvate dehydrogenase activity, and glutamine and fatty acid oxidation, but impairs pyruvate transport.
(a) Relative PDH activity upon shluc with vec, sh4A1 with vec, 4A1WT, 4A1S352L was shown. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 3 biological replicates (n = 3). *P < 0.05. Unpaired, two-tailed t-test; *P < 0.05, **P < 0.01, ***P < 0.001. (b) Maximal respiratory capacity from Fig. 3i was calculated by OCR measurement at basal levels and upon FCCP treatment subtracted by non-mitochondrial respiratory. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 3 biological replicates (n = 3). ***, P < 0.001. (c) Maximal respiratory capacity from Fig. 3j was calculated by OCR measurement at basal levels and upon FCCP treatment subtracted by non-mitochondrial respiratory. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 3 biological replicates (n = 3). ***, P < 0.001. (d) Mycoplasma PCR Detection Kit (Abcam, ab289834) were used to detect mycoplasma in cell culture medium. Mycoplasma positive control is provided by Kit to present PCR product of ~500 bp in length that indicates cell culture contaminated with mycoplasma. (e, f) Pyruvate was incubated with isolated mitochondria from stably expressing control (shLuc), shALDH4A1 (sh4A1), shMPC1, shLuc accompanied by vector expression, sh4A1 restored by 4A1 Hep3B cells in vitro for 1 hour. Pyruvate levels in mitochondria were determined by Pyruvate Assay Kit (BioVision, K609) according to the manufacturer’s instructions. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 3 biological replicates (n = 3). **, P < 0.01; ***, P < 0.001. (g) Kinetic of pyruvate uptake was determined by incubating 14C-pyruvate with isolated mitochondria from 2×107 Hep3B cells stably expressing shLuc or sh4A1 at indicated time points. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 3 biological replicates (n = 3). **, P < 0.01. (h) Pyruvate uptake was determined by using 14C-Pyruvate incubated with isolated mitochondria upon shLuc, sh4A1 (#1), shMPC1MPC/2 and shMPC1/MPC2 with sh4A1 (#1) in Hep3B cells. *, P < 0.05; **, P < 0.01. Data are presented as mean ± s.d. by two-tailed unpaired t-test from 3 biological replicates (n = 3). ***, P < 0.001. (i) Immunoblotting of Hep3B upon shLuc, sh4A1 (#1), shMPC1/MPC2 and shMPC1/MPC2 with sh4A1 (#1) using indicated antibodies.
Extended Data Fig. 8 ALDH4A1 maintains the integrity of MPC1/MPC2 complex and forms the oligomeric complex with MPC1/MPC2.
(a, b) Cell lysates from HepG2 (a) and Huh7 (b) upon shLuc expression were subjected to immunoprecipitation with IgG or 4A1 antibodies, followed by immunoblotting with indicated antibodies. The representative data are shown from 3 independent experiments. (c) Cell lysates from Hep3B upon shLuc with vector, sh4A1 with vector or 4A1WT expression were subjected to immunoprecipitation with IgG or Flag antibodies, followed by immunoblotting with indicated antibodies. The number indicated the quantification of proteins for blots determined by ImageJ. The representative data are shown from 3 independent experiments. (d), Hep3B cell lysates upon vehicle (Veh) or 0.1 mM UK5099 treatment for 24 hours were immunoprecipitated with IgG or 4A1, followed by crosslinking with BMH. The complexes were determined by immunoblotting via SDS-PAGE with indicated antibodies. The representative data are shown from 3 independent experiments. (e, f) Hep3B cell lysates upon vehicle (Veh) or 0.1 mM UK5099 treatment for 24 hours were immunoprecipitated with IgG, MPC1 (e) or MPC2 (f), followed by crosslinking with BMH. The complexes were determined by immunoblotting via SDS-PAGE with indicated antibodies. The representative data are shown from 3 independent experiments. (g) Recombinant MPC1 and MPC2 proteins incubated with or without 4A1 protein were subjected to 4%-16% BN-PAGE for MPC1 and MPC2 oligomerization or were subjected to SDS-PAGE for the levels of the total proteins used for normalization. The number indicated intensity of oligomerization normalized by total proteins determined by ImageJ. The representative data are shown from 2 independent experiments. (h, i) Recombinant MPC1 (h) and MPC2 (i) proteins incubated with or without 4A1 proteins were subjected to 4%-8% BN-PAGE for MPC1 and MPC2 oligomerization or were subjected to SDS-PAGE. * indicated oligomeric MPC1/MPC2 complexes in the presence of ALDH4A1 proteins. The representative data are shown from 2 independent experiments. (j) Recombinant MPC1 and MPC2 proteins incubated with or without 4A1 proteins were subjected to 4%-10% BN-PAGE, followed by mass spectrometry analysis from the bands boxed by red lines. The bands contain the peptides from MPC1, MPC2 and ALDH4A1 proteins identified by mass spectrometry. The representative data are shown from 3 independent experiments. (k) Recombinant MPC1 and MPC2 proteins incubated with 4A1 proteins were subjected to 4%-16% BN-PAGE. 1D native gel strip was proceeded for BN-PAGE again, followed by immunoblotting with indicated antibodies. Red arrows indicated the complex of ALDH4A1, MPC1 and MPC2 in 2D-BN/BN-PAGE. The representative data are shown from 2 independent experiments.
Extended Data Fig. 9 ALDH4A1 displays glycolytic suppression in steady-state and hypoxia conditions.
(a) Seahorse ECAR assay normalized by the actual cell count during the experiment was performed in Hep3B cells upon shLuc with vector (vec), sh4A1 with vec, sh4A1 with 4A1WT and shLuc with 4A1WT, followed by the treatment of 0.5 μM Rotenone/Antimycin (Rot/AA) and 50 mM 2-DG. Data are presented as mean ± s.d by two-way ANOVA from 9 biological replicates (n = 9). shLuc+vec vs. sh4A1+vec, ***P < 0.001; shLuc+vec vs. shLuc+4A1, ***P < 0.001; sh4A1+vec vs. sh4A1 + 4A1WT, ***P < 0.001. (b) Immunoblotting of cancer cell lines under hypoxia condition with indicated antibodies. (c, e) Seahorse glycolysis stress assay normalized by the actual cell count during the experiment was performed in Hep3B cells upon shLuc, sh4A1 (#1 and #2), sh4A1 #2 with 4A1WT (c), or upon vec, 4A1WT (e), followed by the treatment of 10 mM glucose, 3 μM oligomycin and 50 mM 2-DG under hypoxia condition. Data are presented as mean ± s.d by two-way ANOVA from 3 biological replicates (n = 3). shLuc vs. sh4A1#1, ***P < 0.001; shLuc vs. sh4A1 #2, ***P < 0.001; sh4A1 #2 vs. 4A1WT, ***P < 0.001. (d, f) Seahorse ECAR assay normalized by the actual cell count during the experiment was performed in Hep3B cells upon shLuc, sh4A1 (#1 and #2), sh4A1 #2 with 4A1WT (d), or upon vec, 4A1WT (f), followed by the treatment of 0.5 μM Rotenone/Antimycin (Rot/AA) and 50 mM 2-DG under hypoxia condition. Data are presented as mean ± s.d by two-way ANOVA from 4 biological replicates (n = 4). shLuc vs. sh4A1 #1, ***P < 0.001; shLuc vs. sh4A1 #2, ***P < 0.001; sh4A1 #2 vs. 4A1WT, ***P < 0.001; vec vs. 4A1WT, ***P < 0.001. (g) Seahorse ECAR assay normalized by the actual cell count during the experiment was performed in Hep3B cells upon shLuc or sh4A1 treated with low or high glutamine (Gln), followed by the treatment of 0.5 μM Rotenone/Antimycin (Rot/AA) and 50 mM 2-DG. Data are presented as mean ± s.d by two-way ANOVA from 6 biological replicates (n = 6). shLuc+low Gln vs. sh4A1+low Gln, ***P < 0.001; shLuc+ high Gln vs. sh4A1+high Gln, ***P < 0.001; shLuc+low Gln vs. shLuc+high Gln, NS; sh4A1+low Gln vs. sh4A1+high Gln, NS. NS, non-significant. (h) Seahorse ECAR assay normalized by the actual cell count during the experiment was performed in Hep3B cells upon shLuc or sh4A1 treated with BSA or Palmitate, followed by the treatment of 0.5 μM Rotenone/Antimycin (Rot/AA) and 50 mM 2-DG. Data are presented as mean ± s.d by two-way ANOVA from 3 biological replicates (n = 3). shLuc+BSA vs. sh4A1+BSA, ***P < 0.001; shLuc+Palmitate vs. sh4A1+Palmitate, ***P < 0.001; shLuc+BSA Gln vs. shLuc+Palmitate, NS; sh4A1+BSA vs. sh4A1+Palmitate, NS. NS, non-significant. (i) Working model revealing the role of ALDH4A1 in maintaining an active MPC complex for mitochondrial pyruvate import and TCA cycle entry. In normal cells and cancer cells with intact ALDH4A1 expression, ALDH4A1 forms a trimeric complex with MPC1 and MPC2 to maintains an active MPC complex for facilitating pyruvate import into mitochondria, thereby subsequently driving TCA cycle entry. Deficiency of ALDH4A1 commonly found in diverse human cancers disrupts active MPC complex and impairs mitochondrial pyruvate import for TCA cycle entry, thus likely promoting Warburg effect leading to cancer cell growth and tumorigenesis.
Extended Data Fig. 10 Fold change of the metabolites involving in proline and urea cycle pathways according to metabolomic analysis.
The targeted LC/MS metabolic profiling of proline, pyruvate, glutamate, arginine, ornithine and histidine from control, ALDH4A1 knockdown, ALDH4A1 knockdown Hep3B cells with restoration of ALDH4A1WT or ALDH4A1S352L. P-values were calculated by two-tailed unpaired t-test from 3 biological replicates (n = 3).
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2.
Source data
Source Data Figs. 1–8 and Extended Dat Figs. 2–9 (download XLSX )
Statistical source data for Figs. 1–8 and Extended Data Figs. 2–9.
Source Data Figs. 1, 2 and 4–8, Source Data Extended Data Figs. 3, 4 and 6–9 (download PDF )
Unprocessed western blots for Figs. 1, 2 and 4–8 and source data for Extended Data Figs. 3, 4 and 6–9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hsu, CC., Wang, CY., Manne, R.K. et al. ALDH4A1 functions as an active component of the MPC complex maintaining mitochondrial pyruvate import for TCA cycle entry and tumour suppression. Nat Cell Biol 27, 847–862 (2025). https://doi.org/10.1038/s41556-025-01651-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01651-8