Abstract
The retinoic acid-inducible gene I (RIG-I) receptor senses cytoplasmic viral RNA and activates type I interferons (IFN-I) and downstream antiviral immune responses. How RIG-I binds to viral RNA and how its activation is regulated remains unclear. Here, using IFI16 knockout cells and p204-deficient mice, we demonstrate that the DNA sensor IFI16 enhances IFN-I production to inhibit influenza A virus (IAV) replication. IFI16 positively upregulates RIG-I transcription through direct binding to and recruitment of RNA polymerase II to the RIG-I promoter. IFI16 also binds to influenza viral RNA via its HINa domain and to RIG-I protein with its PYRIN domain, thus promoting IAV-induced K63-linked polyubiquitination and RIG-I activation. Our work demonstrates that IFI16 is a positive regulator of RIG-I signalling during influenza virus infection, highlighting its role in the RIG-I-like-receptor-mediated innate immune response to IAV and other RNA viruses, and suggesting its possible exploitation to modulate the antiviral response.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The MS proteomics data have been deposited with the ProteomeXchange Consortium via the PRIDE54 partner repository (https://www.ebi.ac.uk/pride/) with the dataset identifiers PXD020723 and PXD020723. The accession numbers for the RNA sequencing data are GSE157609 and GSE158122. Source data are provided with this paper.
References
Collins, P. J. et al. Crystal structures of oseltamivir-resistant influenza virus neuraminidase mutants. Nature 453, 1258–1261 (2008).
Wang, M. Z., Tai, C. Y. & Mendel, D. B. Mechanism by which mutations at His274 alter sensitivity of influenza A virus N1 neuraminidase to oseltamivir carboxylate and zanamivir. Antimicrob. Agents Chemother. 46, 3809–3816 (2002).
Deyde, V. M. et al. Surveillance of resistance to adamantanes among influenza A(H3N2) and A(H1N1) viruses isolated worldwide. J. Infect. Dis. 196, 249–257 (2007).
Furuse, Y., Suzuki, A. & Oshitani, H. Large-scale sequence analysis of M gene of influenza A viruses from different species: mechanisms for emergence and spread of amantadine resistance. Antimicrob. Agents Chemother. 53, 4457–4463 (2009).
Eisfeld, A. J., Neumann, G. & Kawaoka, Y. At the centre: influenza A virus ribonucleoproteins. Nat. Rev. Microbiol. 13, 28–41 (2015).
Blasius, A. L. & Beutler, B. Intracellular Toll-like receptors. Immunity 32, 305–315 (2010).
Takeuchi, O. & Akira, S. Innate immunity to virus infection. Immunol. Rev. 227, 75–86 (2009).
Pang, I. K. & Iwasaki, A. Inflammasomes as mediators of immunity against influenza virus. Trends Immunol. 32, 34–41 (2011).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Gack, M. U. et al. TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 446, 916–920 (2007).
Oshiumi, H. et al. The ubiquitin ligase Riplet is essential for RIG-I-dependent innate immune responses to RNA virus infection. Cell Host Microbe 8, 496–509 (2010).
Jiang, X. et al. Ubiquitin-induced oligomerization of the RNA sensors RIG-I and MDA5 activates antiviral innate immune response. Immunity 36, 959–973 (2012).
Loo, Y. M. & Gale, M. Jr. Immune signaling by RIG-I-like receptors. Immunity 34, 680–692 (2011).
Dawson, M. J. & Trapani, J. A. The interferon-inducible autoantigen, IFI 16: localization to the nucleolus and identification of a DNA-binding domain. Biochem. Biophys. Res. Commun. 214, 152–162 (1995).
Orzalli, M. H., Conwell, S. E., Berrios, C., DeCaprio, J. A. & Knipe, D. M. Nuclear interferon-inducible protein 16 promotes silencing of herpesviral and transfected DNA. Proc. Natl Acad. Sci. USA 110, E4492–E4501 (2013).
Johnson, K. E. et al. IFI16 restricts HSV-1 replication by accumulating on the hsv-1 genome, repressing HSV-1 gene expression, and directly or indirectly modulating histone modifications. PLoS Pathog. 10, e1004503 (2014).
Almine, J. F. et al. IFI16 and cGAS cooperate in the activation of STING during DNA sensing in human keratinocytes. Nat. Commun. 8, 14392 (2017).
Goubau, D., Deddouche, S. & Reis e Sousa, C. Cytosolic sensing of viruses. Immunity 38, 855–869 (2013).
Jonsson, K. L. et al. IFI16 is required for DNA sensing in human macrophages by promoting production and function of cGAMP. Nat. Commun. 8, 14391 (2017).
Kerur, N. et al. IFI16 acts as a nuclear pathogen sensor to induce the inflammasome in response to Kaposi sarcoma-associated herpesvirus infection. Cell Host Microbe 9, 363–375 (2011).
Monroe, K. M. et al. IFI16 DNA sensor is required for death of lymphoid CD4 T cells abortively infected with HIV. Science 343, 428–432 (2014).
Orzalli, M. H. et al. cGAS-mediated stabilization of IFI16 promotes innate signaling during herpes simplex virus infection. Proc. Natl Acad. Sci. USA 112, E1773–E1781 (2015).
Unterholzner, L. et al. IFI16 is an innate immune sensor for intracellular DNA. Nat. Immunol. 11, 997–1004 (2010).
Ansari, M. A. et al. Herpesvirus genome recognition induced acetylation of nuclear IFI16 is essential for its cytoplasmic translocation, inflammasome and IFN-beta responses. PLoS Pathog. 11, e1005019 (2015).
Dunphy, G. et al. Non-canonical activation of the DNA sensing adaptor STING by ATM and IFI16 mediates NF-ΚB signaling after nuclear DNA damage. Mol. Cell 71, 745–760.e5 (2018).
Thompson, M. R. et al. Interferon γ-inducible protein (IFI) 16 transcriptionally regulates type I interferons and other interferon-stimulated genes and controls the interferon response to both DNA and RNA viruses. J. Biol. Chem. 289, 23568–23581 (2014).
Cao, L. et al. P200 family protein IFI204 negatively regulates type I interferon responses by targeting IRF7 in nucleus. PLoS Pathog. 15, e1008079 (2019).
Chang, X. et al. IFI16 inhibits porcine reproductive and respiratory syndrome virus 2 replication in a MAVS-dependent manner in MARC-145 cells. Viruses https://doi.org/10.3390/v11121160 (2019).
Kim, B. et al. Discovery of widespread host protein interactions with the pre-replicated genome of CHIKV using VIR-CLASP. Mol. Cell 78, 624–640 (2020).
Monzon-Casanova, E. et al. The RNA-binding protein PTBP1 is necessary for B cell selection in germinal centers. Nat. Immunol. 19, 267–278 (2018).
Grellscheid, S. N. et al. Molecular design of a splicing switch responsive to the RNA binding protein Tra2β. Nucleic Acids Res. 39, 8092–8104 (2011).
Li, H. et al. RNA helicase DDX5 inhibits reprogramming to pluripotency by miRNA-based repression of RYBP and its PRC1-dependent and -independent functions. Cell Stem Cell 20, 462–477.e6 (2017).
Herdy, B. et al. Analysis of NRAS RNA G-quadruplex binding proteins reveals DDX3X as a novel interactor of cellular G-quadruplex containing transcripts. Nucleic Acids Res. 46, 11592–11604 (2018).
Di Giammartino, D. C. et al. RBBP6 isoforms regulate the human polyadenylation machinery and modulate expression of mRNAs with AU-rich 3′ UTRs. Genes Dev. 28, 2248–2260 (2014).
Li, T., Diner, B. A., Chen, J. & Cristea, I. M. Acetylation modulates cellular distribution and DNA sensing ability of interferon-inducible protein IFI16. Proc. Natl Acad. Sci. USA 109, 10558–10563 (2012).
Baum, A., Sachidanandam, R. & Garcia-Sastre, A. Preference of RIG-I for short viral RNA molecules in infected cells revealed by next-generation sequencing. Proc. Natl Acad. Sci. USA 107, 16303–16308 (2010).
Jakobsen, M. R. & Paludan, S. R. IFI16: at the interphase between innate DNA sensing and genome regulation. Cytokine Growth Factor Rev. 25, 649–655 (2014).
Li, D. et al. STING-mediated IFI16 degradation negatively controls type I interferon production. Cell Rep. 29, 1249–1260 (2019).
Merkl, P. E. & Knipe, D. M. Role for a filamentous nuclear assembly of IFI16, DNA, and host factors in restriction of herpesviral infection. mBio https://doi.org/10.1128/mBio.02621-18 (2019).
Antiochos, B., Matyszewski, M., Sohn, J., Casciola-Rosen, L. & Rosen, A. IFI16 filament formation in salivary epithelial cells shapes the anti-IFI16 immune response in Sjogren’s syndrome. JCI Insight https://doi.org/10.1172/jci.insight.120179 (2018).
Morrone, S. R. et al. Cooperative assembly of IFI16 filaments on dsDNA provides insights into host defense strategy. Proc. Natl Acad. Sci. USA 111, E62–E71 (2014).
Hotter, D. et al. IFI16 targets the transcription factor Sp1 to suppress HIV-1 transcription and latency reactivation. Cell Host Microbe 25, 858–872 (2019).
Thapa, R. J. et al. DAI senses influenza a virus genomic RNA and activates RIPK3-dependent cell death. Cell Host Microbe 20, 674–681 (2016).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci Immunol. https://doi.org/10.1126/sciimmunol.aag2045 (2016).
Sui, H., Zhou, M., Chen, Q., Lane, H. C. & Imamichi, T. siRNA enhances DNA-mediated interferon lambda-1 response through crosstalk between RIG-I and IFI16 signalling pathway. Nucleic Acids Res. 42, 583–598 (2014).
Cao, L. et al. The nuclear matrix protein SAFA surveils viral RNA and facilitates immunity by activating antiviral enhancers and super-enhancers. Cell Host Microbe 26, 369–384 (2019).
Gao, Q. et al. A cell-based high-throughput approach to identify inhibitors of influenza A virus. Acta Pharm. Sin. B 4, 301–306 (2014).
Wei, F. et al. Induction of PGRN by influenza virus inhibits the antiviral immune responses through downregulation of type I interferons signaling. PLoS Pathog. 15, e1008062 (2019).
Manicassamy, B. et al. Analysis of in vivo dynamics of influenza virus infection in mice using a GFP reporter virus. Proc. Natl Acad. Sci. USA 107, 11531–11536 (2010).
Gavazzi, C. et al. An in vitro network of intermolecular interactions between viral RNA segments of an avian H5N2 influenza A virus: comparison with a human H3N2 virus. Nucleic Acids Res. 41, 1241–1254 (2013).
You, F. et al. ELF4 is critical for induction of type I interferon and the host antiviral response. Nat. Immunol. 14, 1237–1246 (2013).
Zhang, Y. et al. Identifying local and descending inputs for primary sensory neurons. J. Clin. Invest. 125, 3782–3794 (2015).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank F. You (Peking University), W. Liu (Institute of Microbiology, Chinese Academy of Sciences), Y. Zhu (Wuhan University) and Y. Chen (Wuhan University) for kindly providing cell lines, and W. Tang (Shandong University) for generously gifting the p204−/− mice. This work was supported by the National Key Research and Development Program of China (2016YFD0500204 to J.L.) and the National Natural Science Foundation of China (81960297 to F.W.).
Author information
Authors and Affiliations
Contributions
Z.J. and F.W. performed and analysed most of the experiments. Y.Z., T.W. and W.G. performed the AP–MS experiments. S.Y., H.S., J.P., Y.S., M.W. and Q.T. generated biochemical reagents. C.G. and K.-C.C. guided and analysed the data. F.W. and J.L. conceived and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Peter Staeheli, Aartjan te Velthuis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 IFI16 induction by IAV is dependent on viral replication.
a–c, IFI16 expression in a, THP-1, b, A549, and c, HEK293 was quantified by RT-qPCR. (d) p204 expression in lung tissues from PR8 virus-infected WT mice was determine. e, IFI16 expression in 1.0 MOI of PR8 virus-infected THP-1 cells was detected. f, IFI16 mRNA expression in 2 MOI of UV-inactivated and live PR8 virus-infected A549 cells was determined. g, IFI16 protein expression in 2 MOI of UV-inactivated PR8 virus-infected A549 cells was determined. h, IFI16 expression in the nuclear and cytoplasmic fractions of PR8 virus-infected A549 cells were determined. β-actin and H3 were used as purity markers for cytoplasmic and nuclear fractions, respectively. i, IFI16 expression in A549 cells transfected with poly(I:C) for 18 h was determined by Western blotting. j, IFI16 expression in A549 cells treated with IFN-γ for 18 h was determined. (k) Intracellular localization of IFI16 in A549 cells treated with poly(I:C) or IFN-γ for 12 h was determined. Scale bars, 45 μm and 5 μm (enlarged). l, A549 cells were infected with PR8 virus at 0, 6, 12 and 24 hpi. Cell lysates were then immunoprecipitated with anti-acetylated lysine. Bound proteins were analyzed by immunoblots with anti-IFI16 antibody. m, A549 cells were pre-incubated with C-646 for 2 h, then infected with PR8 virus for 1 h, washed and incubated in complete medium with or without C-646. IFI16 expression and viral NP protein in the nuclear and cytoplasmic fractions of PR8 virus-infected A549 cells at indicated time points were determined by Western blotting. β-Actin and H3 were used as purity markers for cytoplasmic and nuclear fractions, respectively. (a–d) and f, Data presented as means ± SD from three independent experiments. (e) and g–m, Data are representative of three independent experiments. Statistical significance in (a) to (d) and (f) was determined by unpaired two-tailed Student’s t-test.
Extended Data Fig. 2 IFI16 inhibits IAV viral replication.
a, A549 cells were transfected with IFI16-Flag plasmids or empty control for 24 h and then infected with PR8 virus at 2.0 MOI. mRNA and vRNA expression of NP and M genes at 6, 12 and 18 hpi were determined by RT-qPCR. Data are presented as means ± SEMs from three independent experiments. b, HEK293 cells were transfected with IFI16-Flag plasmids or empty control for 24 h and then infected with PR8 virus at 2.0 MOI. NP protein expression at 0, 6, 12 and 18 hpi was determined by Western blotting. β-Actin detection was used as loading control. (c) and (d), A549 cells were transfected with IFI16-targeting siRNA and negative control (NC) siRNA for 24 h, followed by infection with PR8 virus (MOI = 1). c, NP and M1 protein expression were determined by Western blotting at 0, 12, 18 and 24 hpi. β-Actin detection was used as loading control. d, Viral titers were determined by TCID50 assay at the 24 hpi. Data presented as means ± SD from three independent experiments. (b) and (c), Data are representative of three independent experiments. Statistical significance in (a) and (d) was determined by unpaired two-tailed Student’s t-test. ns = non-significant.
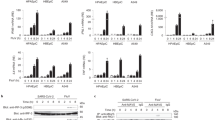
Extended Data Fig. 3 IFI16 enhances the activation of IFN-I pathway during IAV infection.
a to c, Serum-starved A549cells were transfected with IFI16-encoding plasmids or negative control, and then infected with 1.0 MOI of PR8 virus. Expression of IFN-β (a), ISG15 (b), IL-6 (c) at the indicated time points was determined by RT-qPCR. d to f, mRNA expression of IFN-α4 (d), IRF7 (e) and CXCL5 (f) in PR8 virus-infected IFI16+/+ and IFI16−/− A549 cells at 0, 6, and 12 hpi was determined by RT-qPCR. (g) Immunoblot analysis of RIG-I-triggered downstream signaling pathway in IFI16+/+ and IFI16−/− A549 cells after stimulation with 5′ppp-RNA for the indicated duration. h, IFI16+/+ and IFI16−/− A549 cells transfected with IFI16-Flag expression vectors or empty control for 24 h and then infected with PR8 virus at 1.0 MOI. RIG-I-triggered downstream signaling pathway at 0, 4, 8 and 12 hpi was assessed with indicated antibodies. a to f, Data presented as means ± SD from three independent experiments. g to h, Data are representative of three independent experiments. Statistical significance in (a) to (f) was determined by unpaired two-tailed Student’s t-test. ns = non-significant.
Extended Data Fig. 4 IFI16 binds RIG-I promoter and enhances RIG-I transcription.
a, RIG-I expression in A549 cells transfected with IFI16-Flag vectors was examined. b, HEK293 cells co-transfected with indicated plasmids for 24 h were treated with cycloheximide (CHX) for 12 and 24 h. Expression of RIG-I and IFI16 in cell lysates was detected. c, RIG-I expression in ifnar1−/−A549 cells transfected with IFI16-Flag vectors was examined. d, RIG-I mRNA expression in ifnar1−/− A549 cells transfected with IFI16-Flag plasmids for 24 h was determined. e, RIG-I expression in A549 cells transfected with IFI16-Flag plasmids for 24 h was determined. f, RIG-I expression in lung tissues from virus-infected WT and p204−/− mice (n = 3) was determined. g, Schematic diagram of the biotinylated probe sequences from the promoter of RIG-I gene and corresponding mutants. h, IB analysis of the binding ability between wild type (p1 and p2) and mutated (p2-mut1 to p2-mut5) probes, and IFI16 proteins. i, IFI16+/+ and IFI16−/− A549 cells were infected with PR8 virus for 12 h, followed by ChIP assay. j, Schematic diagram of full-length IFI16 vectors and truncated mutants. k, A549 cells were transfected with Flag-tagged full-length IFI16 vectors and truncated mutant plasmids. After 24 h transfection, nuclear extracts were incubated with non-biotinylated or biotinylated promoter sequence of RIG-I for 4 h. Nuclear extracts were examined for IFI16 and truncated mutant expression by Western blotting. l, A549 cells after Flag-tagged IFI16 vectors and truncated mutants transfection and PR8 infection was determined. d to f and i, Data are representative of three independent experiments (mean ± SD). (a), (b), (c), (h), (k), and (l), Data are representative of three independent experiments. Statistical significance in (d) to (f) and (i) was determined by unpaired two-tailed Student’s t-test. ns = non-significant.
Extended Data Fig. 5 IFI16 directly binds viral RNA during infection.
a, HEK293 cells were transfected with indicated vectors for 24 h. Cell lysates were incubated with biotin-labeled viral NP RNA and immunoprecipitated with streptavidin beads. Bound proteins were analyzed by immunoblots with anti-Flag antibody. b, HEK293 cells were transfected with HINa-GFP, HINb-GFP, and PYRIN-GFP expression vectors for 24 h. Cell lysates were incubated with influenza NS vRNA. Binding affinity between indicated proteins and NS vRNA was determined by MST assays. c, Purified GST-IFI16 proteins were incubated with fluorescein labeled influenza HA, NP, PA and PB2 vRNAs. Binding affinity was determined by MST assays. d, PCR detection of NP vRNA in eluted RNA from RIG-I-Flag, IFI16-Flag and indicated Flag-tagged IFI16 truncated constructs. The data represent means ± SD. (n = 3 independent experiments). e, PCR detection of NA vRNAs in eluted RNA from RIG-I-Flag, IFI16-Flag and indicated Flag-tagged IFI16 truncations. Data were normalized to vRNA from RIG-I-Flag immunoprecipitates. The data represent means ± SD. (n = 3 independent experiments). f, Confocal microscopy detecting the co-localization of endogenous IFI16 and RIG-I in PR8 virus-infected A549 cells at 0, 6 and 18 hpi. Nuclei were stained with DAPI (left). Scale bars, 10 μm. Quantification of the co-localization of IFI16 and RIG-I in cells (right). Means ±SD from 3 biological samples. g, Confocal microscopy of HEK293 cells transfected with RIG-I-mCherry plasmids, together with expression vectors for IFI16-GFP or GFP-tagged mutants. DAPI serves as a marker for nuclei (left). Scale bars, 20 μm. Quantification of the co-localization of IFI16 or mutants and RIG-I in cells (right). Means ±SD from 3 biological samples. h, Co-IP analysis of the interaction between Myc-tagged RIG-I and Flag-tagged full-length or mutant IFI16 in HEK293 cells. (i) Three-dimensional confocal microscopy of HEK293 cells co-transfected with plasmids encoding PYRIN-GFP and RIG-I-mCherry. DAPI serves as a marker for nuclei. All data are representative of three independent experiments. Scale bars, 5 μm.
Extended Data Fig. 6 RIG-I is involved in IFI16-mediated antiviral response in IAV infection.
a, IFI16+/+ or IFI16−/− A549 cells were infected with PR8 virus (MOI = 1) and then analyzed by PLA. The right panels are enlarged. Red point represents TRIM25 plus RIG-I complexes. Scale bars, 23.8 μm (left) and 7.45 μm (enlarged). b, IFI16+/+ or IFI16−/− A549 cells were infected with PR8 virus (MOI = 1) and then analyzed by PLA. The right panels are enlarged. Red point indicates K63 Ub plus RIG-I complexes. Scale bars, 40.3 μm (left) and 14.3 μm (enlarged). c, RIP-EMSA analysis of the binding of biotinylated PA vRNA to purified Flag-RIG-I protein by adding purified GST-IFI16 at dose of 50 and 200 ng. d, RNA pull-down analysis of the binding of biotinylated PA vRNA to purified Flag-RIG-I protein by adding purified GST-IFI16 at dose of 30, 60, and 120 ng. e, A549 cells were transfected with Flag-tagged IFI16, its truncated forms and control. RNA eluted from RIG-I immunoprecipitates were detected by RT-qPCR with specific primers of PA gene. f, A549 cells were transfected with Flag-tagged IFI16 and its truncated forms for 24 h and then infected with PR8 virus. Production of IFN-β in supernatants at 18 hpi was determined. g, Knockdown effect of MAVS-targeting siRNAs in A549 cells. h, IFI16+/+ or IFI16−/− A549 cells were transfected with control or MAVS-targeting siRNA #571 and after 24 h infected with PR8 virus. Production of IFN-β in supernatants at 18 hpi was determined (i) IFI16+/+ A549 cells were transfected with negative control or MAVS-targeting siRNA #571, and 24 h later, cells were transfected with Flag-IFI16 plasmids or control; 24 h later cells were infected with PR8 virus. Production of IFN-β in supernatants on 18 hpi were quantified. (e), (f), (h), and (i), Data are representative of three independent experiments (mean ± SD). (a) to (d) and (g), All data are representative of three independent experiments. Statistical significance in (e) to (f) and (i) was determined by unpaired two-tailed Student’s t-test. ns = non-significant.
Extended Data Fig. 7 IFI16 promotes RIG-I signaling.
a, The interaction between TRIM25, RIG-I, and K63-linked ubiquitination of RIG-I in PR8 virus-infected IFI16+/+ and IFI16−/− A549 cells. b, Co-IP analysis of RIG-I ubiquitination in HEK293 cells transfected with indicated plasmids. c, Cell lysates were prepared after indicated plasmids transfection and PR8 infection and then immunoprecipitated with control IgG or anti-Flag. Bound-RNA was extracted for qPCR analysis. d, IFI16+/+ and IFI16−/− A549 cells were infected with PR8 virus for 12 h. Cell lysates were then immunoprecipitated with IgG or anti-RIG-I. Bound-RNA was extracted for analysis. e, Schematic experimental procedure used in (f). f, RNA co-purified with RIG-I from IAV-infected IFI16+/+ and IFI16−/− A549 cells were transfected into A549 cells, and IFN-β was determined. g, Luciferase activity of HEK293 cells after transfection with IFI16 vectors or control and then infection with PR8 virus. h, Luciferase activity of HEK293 cells transfected with an IFN-β reporter plasmids, RIG-I vectors, and control or IFI16 vectors. i, Knockdown effect of RIG-I-targeting siRNAs in A549 cells. j, IFN-β in IFI16+/+ or IFI16−/− A549 cells after transfection with siRNA #2835 or control and infection with PR8 virus. k, IFN-β in IFI16+/+ cells after transfection with siRNA #2835 or control, IFI16-Flag or control plasmids, and infection with PR8 virus. l, NP protein levels in RIG-I−/− HEK293 cells after transfection with IFI16-Flag vectors or control and infection with PR8 virus were determined. m, RIG-I−/− HEK293 cells were transfected with indicated plasmids for 24 h and then infected with PR8 virus at 1.0 MOI. Viral titers were determined. (a) to (d), (i), and (l), Data are representative of three independent experiments. (c) to (d), (f) to (h), (j), (k) and (m), Data presented as means ± SD from three independent experiments, and the significance of the results was assessed using a parametric paired t-test (Student’s two-tailed t-test). ns = non-significant.
Extended Data Fig. 8 Schematic model to show IFI16 enhances RIG-I signaling in influenza virus infection.
Briefly, influenza virus infection upregulates IFI16 expression; IFI16 protein, in turn, enhances transcription of RIG-I. In addition, IFI16 protein, via its HINa domain, directly senses viral RNA and, via its PYRIN domain, interacts with RIG-I receptor to promote antiviral RIG-I signaling.
Supplementary information
Supplementary Information (download PDF )
The qPCR or PCR primers sequence used in this study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Jiang, Z., Wei, F., Zhang, Y. et al. IFI16 directly senses viral RNA and enhances RIG-I transcription and activation to restrict influenza virus infection. Nat Microbiol 6, 932–945 (2021). https://doi.org/10.1038/s41564-021-00907-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-021-00907-x
This article is cited by
-
Subcellular localization as a driver of protein function
Nature Reviews Molecular Cell Biology (2026)
-
Deciphering lactate/lactylation networks in AML: integrated scRNA-seq and transcriptomics reveal functions and prognostic model
BMC Cancer (2025)
-
Structural insights into the atypical filament assembly of pyrin domain-containing IFI16
The EMBO Journal (2025)
-
Cytosolic nucleic acid sensing as driver of critical illness: mechanisms and advances in therapy
Signal Transduction and Targeted Therapy (2025)
-
O-GlcNAc transferase plays dual antiviral roles by integrating innate immunity and lipid metabolism
Nature Communications (2025)