Abstract
Epstein–Barr virus (EBV) can infect B cells and epithelial cells, and cause lymphomas and various epithelial malignancies. During epithelial cell infection, EBV employs a complex combination of viral glycoproteins and host receptors. However, the exact mechanism and whether a dominant receptor exists remain unclear. Here we identify desmocollin 2 (DSC2) as a dominant EBV entry receptor for epithelial cell infection using CRISPR–Cas9 screening. Knockout of DSC2 reduced EBV infection in both nasopharyngeal and gastric epithelial cell lines, and infection was rescued when DSC2 expression was restored. Expression of human DSC2 in non-EBV-susceptible hamster cell lines enabled susceptibility to EBV. Furthermore, we found that DSC2 directly binds to EBV glycoprotein H/glycoprotein L through its extracellular domain, particularly the preEC–EC2 regions, which could be targeted by polyclonal antibodies, therefore blocking EBV infection in primary epithelial cells. DSC2 enabled virus entry independent of Ephrin receptor A2. These findings could aid development of currently unavailable animal models and support development of targeted therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Source data files are provided with this paper. Illumina sequencing reads from the CRISPR screens can be accessed via the NCBI Sequence Read Archive (SRA BioProject ID: PRJNA1273314). The key raw data of all the functional experiments in this work have been deposited in the Research Data Deposit public platform (www.researchdata.org.cn) under accession number RDDB2025796481. Source data are provided with this paper.
Code availability
The analysis was performed using standard protocols with previously described computational tools. No custom code was used in this study.
References
Damania, B., Kenney, S. C. & Raab-Traub, N. Epstein–Barr virus: biology and clinical disease. Cell 185, 3652–3670 (2022).
Young, L. S., Yap, L. F. & Murray, P. G. Epstein–Barr virus: more than 50 years old and still providing surprises. Nat. Rev. Cancer 16, 789–802 (2016).
Farrell, P. J. Epstein–Barr virus and cancer. Annu. Rev. Pathol. 14, 29–53 (2019).
Zhong, L. Y. et al. Research landmarks on the 60th anniversary of Epstein–Barr virus. Sci. China Life Sci. 68, 354–380 (2025).
Hutt-Fletcher, L. M. EBV glycoproteins: where are we now? Future Virol. 10, 1155–1162 (2015).
Shannon-Lowe, C. & Rowe, M. Epstein–Barr virus entry; kissing and conjugation. Curr. Opin. Virol. 4, 78–84 (2014).
Bu, G. L., Xie, C., Kang, Y. F., Zeng, M. S. & Sun, C. How EBV infects: the tropism and underlying molecular mechanism for viral infection. Viruses 14, 2372 (2022).
Chen, J. & Longnecker, R. Epithelial cell infection by Epstein–Barr virus. FEMS Microbiol. Rev. 43, 674–683 (2019).
Tanner, J., Weis, J., Fearon, D., Whang, Y. & Kieff, E. Epstein–Barr virus gp350/220 binding to the B lymphocyte C3d receptor mediates adsorption, capping, and endocytosis. Cell 50, 203–213 (1987).
Sun, C. et al. Structural basis of Epstein–Barr virus gp350 receptor recognition and neutralization. Cell Rep. 44, 115168 (2025).
Ogembo, J. G. et al. Human complement receptor type 1/CD35 is an Epstein–Barr virus receptor. Cell Rep. 3, 371–385 (2013).
Sathiyamoorthy, K. et al. Structural basis for Epstein–Barr virus host cell tropism mediated by gp42 and gHgL entry glycoproteins. Nat. Commun. 7, 13557 (2016).
Li, Q. et al. Epstein–Barr virus uses HLA class II as a cofactor for infection of B lymphocytes. J. Virol. 71, 4657–4662 (1997).
Chesnokova, L. S., Nishimura, S. L. & Hutt-Fletcher, L. M. Fusion of epithelial cells by Epstein–Barr virus proteins is triggered by binding of viral glycoproteins gHgL to integrins αvβ6 or αvβ8. Proc. Natl Acad. Sci. USA 106, 20464–20469 (2009).
Chesnokova, L. S. & Hutt-Fletcher, L. M. Fusion of Epstein–Barr virus with epithelial cells can be triggered by αvβ5 in addition to αvβ6 and αvβ8, and integrin binding triggers a conformational change in glycoproteins gHgL. J. Virol. 85, 13214–13223 (2011).
Xiong, D. et al. Nonmuscle myosin heavy chain IIA mediates Epstein–Barr virus infection of nasopharyngeal epithelial cells. Proc. Natl Acad. Sci. USA 112, 11036–11041 (2015).
Zhang, H. et al. Ephrin receptor A2 is an epithelial cell receptor for Epstein–Barr virus entry. Nat. Microbiol. 3, 1–8 (2018).
Chen, J. et al. Ephrin receptor A2 is a functional entry receptor for Epstein–Barr virus. Nat. Microbiol. 3, 172–180 (2018).
Wang, H. B. et al. Neuropilin 1 is an entry factor that promotes EBV infection of nasopharyngeal epithelial cells. Nat. Commun. 6, 6240 (2015).
Speck, P. & Longnecker, R. Epstein–Barr virus (EBV) infection visualized by EGFP expression demonstrates dependence on known mediators of EBV entry. Arch. Virol. 144, 1123–1137 (1999).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Montgomery, R. I., Warner, M. S., Lum, B. J. & Spear, P. G. Herpes simplex virus-1 entry into cells mediated by a novel member of the TNF/NGF receptor family. Cell 87, 427–436 (1996).
Shieh, M. T., WuDunn, D., Montgomery, R. I., Esko, J. D. & Spear, P. G. Cell surface receptors for herpes simplex virus are heparan sulfate proteoglycans. J. Cell Biol. 116, 1273–1281 (1992).
Geraghty, R. J., Krummenacher, C., Cohen, G. H., Eisenberg, R. J. & Spear, P. G. Entry of alphaherpesviruses mediated by poliovirus receptor-related protein 1 and poliovirus receptor. Science 280, 1618–1620 (1998).
Shukla, D. et al. A novel role for 3-O-sulfated heparan sulfate in herpes simplex virus 1 entry. Cell 99, 13–22 (1999).
Satoh, T. et al. PILRalpha is a herpes simplex virus-1 entry coreceptor that associates with glycoprotein B. Cell 132, 935–944 (2008).
Chesnokova, L. S., Valencia, S. M. & Hutt-Fletcher, L. M. The BDLF3 gene product of Epstein–Barr virus, gp150, mediates non-productive binding to heparan sulfate on epithelial cells and only the binding domain of CD21 is required for infection. Virology 494, 23–28 (2016).
Harrison, O. J. et al. Structural basis of adhesive binding by desmocollins and desmogleins. Proc. Natl Acad. Sci. USA 113, 7160–7165 (2016).
Callaway, E. Major AlphaFold upgrade offers boost for drug discovery. Nature 629, 509–510 (2024).
Miller, N. & Hutt-Fletcher, L. M. Epstein–Barr virus enters B cells and epithelial cells by different routes. J. Virol. 66, 3409–3414 (1992).
Connolly, S. A., Jardetzky, T. S. & Longnecker, R. The structural basis of herpesvirus entry. Nat. Rev. Microbiol. 19, 110–121 (2021).
Mohl, B. S., Chen, J., Park, S. J., Jardetzky, T. S. & Longnecker, R. Epstein–Barr virus fusion with epithelial cells triggered by gB is restricted by a gL glycosylation site. J. Virol. 91, e01255-17 (2017).
Chen, J., Schaller, S., Jardetzky, T. S. & Longnecker, R. Epstein–Barr virus gH/gL and Kaposi’s sarcoma-associated herpesvirus gH/gL bind to different sites on EphA2 to trigger fusion. J. Virol. 94, e01454-20 (2020).
Su, C. et al. Molecular basis of EphA2 recognition by gHgL from gammaherpesviruses. Nat. Commun. 11, 5964 (2020).
Sun, C., Wang, L., Yang, X. X., Jiang, Y. H. & Guo, X. L. The aberrant expression or disruption of desmocollin2 in human diseases. Int. J. Biol. Macromol. 131, 378–386 (2019).
Tsai, M. H. et al. Spontaneous lytic replication and epitheliotropism define an Epstein–Barr virus strain found in carcinomas. Cell Rep. 5, 458–470 (2013).
Wang, B. et al. Integrative analysis of pooled CRISPR genetic screens using MAGeCKFlute. Nat. Protoc. 14, 756–780 (2019).
Zhu, Q. Y. et al. A potent and protective human neutralizing antibody targeting a novel vulnerable site of Epstein–arr virus. Nat. Commun. 12, 6624 (2021).
Sun, C. et al. A gB nanoparticle vaccine elicits a protective neutralizing antibody response against EBV. Cell Host Microbe 31, 1882–1897.e10 (2023).
Acknowledgements
We thank M. Masucci (Karolinska Institute, Sweden) for providing the EBV-positive Akata cell line; S.-W. Tsao (University of Hong Kong, Hong Kong SAR) for providing the TW03 cell lines; R. Longnecker and P. G. Spear (Northwestern University) for providing plasmids pCAGT7 and pT7EMCLuc; W. Hammerschmidt (Helmholtz Zentrum München) for providing plasmid p2670. This paper was edited by Life Science Editors. This work was supported by the National Key Research and Development Program of China (2022YFC2305400 to H.Z., 2022YFC3400900 to M.-S.Z.); the National Natural Science Foundation of China (82372246 to H.Z., U24A20743 to M.-S.Z., 32441094 to M.-S.Z., 82402614 to C.S., 82030046 to M.-S.Z.); the Noncommunicable Chronic Diseases-National Science and Technology Major Project (2023ZD0501000 to Q.Z.); Cancer Innovative Research Program of Sun Yat-sen University Cancer Center (CIRP-SYSUCC-0006 to M.-S.Z.); Guangdong Science and Technology Department (2021A1515010734 to H.Z.); Shenzhen Fundamental Research Program (JCYJ20230807111219040, ZDSYS20220606100803007 to H.Z); China Postdoctoral Science Foundation (GZB20230886 to C.S., 2023M743998 to C.S., 2024T171080 to C.S.); and Postdoctoral Fellowship Program of CPSF (GZB20230886 to C.S.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.-S.Z., H.Z. and C.S. conceived of and designed the experiments, provided supervision, and wrote the paper. H.Z. conducted and analysed the key experiments and wrote the paper. Y.-C.L., D.P. and C.X. performed and analysed the key experiments. T.Z., Ying Li, Yan Li, Z.-Y.J., G.-L.B., M.-M.L., Y.-R.C., H.-X.F., R.-B.L., P.-H.W., W.-T.D., G.-X.Z., Y.-L.L. and P.H. performed the experiments. Q.Z. provided constructive suggestions for this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CRISPR/Cas9 screening and gene silencing identify DSC2 as a host factor for EBV infection in epithelial cells.
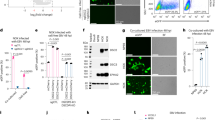
(a) Flow cytometry gating strategy for CRISPR/Cas9 screening. Briefly, HEK293-Cas9 cells were stably transfected with the human CRISPR knockout pooled library (GeCKO v2) and subsequently incubated with or without EBfaV-GFP (~50 encapsidated EBV genomes per cell). After 24 h, the EBV-uninfected group (Mock) was used as a negative control for flow cytometry gating, and in the EBV-infected group (EBV infection), GFP-negative cell populations (GFP-) were sorted, propagated, and re-infected with EBfaV-GFP to confirm the loss of susceptibility to EBV infection. (b) TW03 and AGS cells were transfected with ARHGEF39, ATXN2L, DSC2, ANGPTL1 siRNAs (siARHGEF39, siATXN2L, siDSC2 or siANGPTL1, respectively) or control siRNA (siCtrl), then analyzed by RT-qPCR to detect the knockdown efficiency of indicated siRNAs, Results were quantified relative to the housekeeping gene beta-actin (ACTB), and then normalized to siCtrl. (c) Representative flow cytometry plots of Fig. 1c. One-way (b) analysis of variance (ANOVA) with Dunnett’s post-test. Mean ± s.d. of three technical replicates (b) are shown. Data are representative of two (a, b, c) independent experiments.
Extended Data Fig. 2 DSC2 upregulation restores EBV infectivity in both DSC2 knockout and non-EBV-susceptible cells.
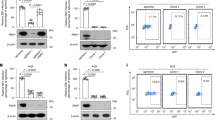
(a) Representative flow cytometry plots of Fig. 1f. (b) Representative fluorescence microscopy images of Fig. 1g, h. (c, d) CHO-K1 and BHK-21 cells were stably transfected with Myc-tagged human DSC2 (hDSC2) or empty vector (EV), then incubated with EBV (~100 encapsidated EBV genomes per cell) following flow cytometry analysis (c), and the flow cytometry quantification data are shown in (d). (e) Representative flow cytometry plots of Fig. 2b. One-way (d) analysis of variance (ANOVA) with Tukey’s post-test (d). Mean ± s.d. of three biological replicates (d) are shown. Data are representative of three (a, b, c, d, e) independent experiments.
Extended Data Fig. 3 FITC-labeled-gH/gL co-localizes with DSC2 expressed on TW03 cells.
(a) Validation experiments for the specificity of the rabbit anti-DSC2 antibody (10809-R004, Sino Biological) for immunofluorescence assay. WT and DSC2 knockout AGS single-cell clones (AGS-sgDSC2-c1) were stained with rabbit anti-DSC2 antibody and Alexa 488-conjugated secondary antibody, and imaged as green. Nuclei were stained with DAPI (blue). Scale bars,10 μm. (b) TW03 cells were incubated with 200 nM biotin-labeled gH/gL at 4 °C for 1 h. biotin-labeled gH/gL was stained with streptavidin-FITC and imaged as green. DSC2 was stained with rabbit anti-DSC2 antibody (10809-R004, Sino Biological) and Alexa 594-conjugated secondary antibody and imaged as red. Nuclei were stained with DAPI (blue). Scale bars,10 μm. Enlarged images are shown on the right. Data are representative of two (a, b) independent experiments.
Extended Data Fig. 5 Interaction pattern for DSC2-gH/gL complex in Alphafold3 model.
The detailed interactions for DSC2-gH/gL complex, composed of DSC2 preEC, EC1 and EC2 and gH. The interacting residues between DSC2 and gH/gL are shown as sticks. The zoomed box showed the detailed polar bonds between DSC2 preEC, EC1 or EC2 and gH. The residue labels were marked beside the sticks.
Extended Data Fig. 6 Ectopic expression of human CD21 slightly enhances EBV infection in non-EBV-susceptible cell line.
(a) Representative flow cytometry plots of Fig. 3g. (b, c) CHO-K1 cells were stably transfected with Myc-tagged human DSC2 (hDSC2), human CD21 (hCD21), or empty vector (EV), then incubated with EBV (~100 encapsidated EBV genomes per cell) followed by flow cytometry analysis (b), and the flow cytometry quantification data are shown in (c). One-way (c) analysis of variance (ANOVA) with Dunnett’s post-test (c). Mean ± s.d. of three biological replicates (c) are shown. Data are representative of three (b, c) independent experiments.
Extended Data Fig. 7 DSC2 restores EBV infectivity in EPHA2 knockout epithelial cells.
(a) DSC2 or EPHA2 knockout TW03 single-cell clones (sgDSC2-c1 or sgEPHA2-c1, respectively) were infected with EBV, and the mRNA expression of EBV encoded genes (EBNA1, BZLF1, BXLF2, and BALF4) was detected using RT-qPCR. Results were quantified relative to the housekeeping gene beta-actin (ACTB), and then normalized to WT. (b) Representative flow cytometry plots of Fig. 5c, d. One-way analysis of variance (ANOVA) with Dunnett’s post-test (a). Mean ± s.d. of four technical replicates (a) are shown. Data are representative of two (a) or three (b) independent experiments.
Extended Data Fig. 8 Cellular surface expression of DSC2 and EphA2.
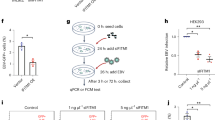
(a,b) The mRNA expression of EPHA2 (a) and DSC2 (b) in TW03, AGS, TECs, NPECs, CHO-K1 and BHK-21 cells were detected using RT-qPCR or analyzed by WB to detect protein levels. Results were quantified relative to the housekeeping gene beta-actin (ACTB). (c) The protein expression of EphA2 and DSC2 in TW03, AGS, CHO-K1 and BHK-21 cells was detected by WB. (d) TW03 and NPECs cells were stained with rabbit anti-DSC2 (10809-R004, Sino Biological) and Alexa 488-conjugated secondary antibody, and imaged as green, and with mouse anti-EphA2 and Alexa 594-conjugated secondary antibody and imaged as red. Nuclei were stained with DAPI (blue). Scale bars, 10 μm. Mean ± s.d. of three technical replicates (a, b) were used. Data are representative of three (a, b, c, d) independent experiments.
Extended Data Fig. 9 DSC2 restores EBV infectivity in EPHA2 knockout and EphA2 blocking antibody-treated epithelial cells.
(a) Representative flow cytometry plots of Fig. 5e, f. (b and c) TW03 cells stably transfected with Myc-tagged DSC2 (DSC2) or empty vector (EV) were preincubated with 50 μg ml−1 EphA2 blocking antibody, and then infected with EBV followed by flow cytometry analysis (b), and the flow cytometry quantification data are shown in (c). Results were normalized to EV-IgG group. Two-way (c) analysis of variance (ANOVA) with Tukey’s post-test (c). Mean ± s.d. of three biological replicates (c) were used. Data are representative of three (a,b, c) independent experiments.
Extended Data Fig. 10 Schematic showing the potential role of DSC2 in mediating EBV infection.
DSC2 is the dominant entry receptor for EBV infection in epithelial cells. The gene deletion of DSC2 resulted in the inability of EBV-susceptible epithelial cells to be infected by EBV even in the presence of EphA2 (left panel). Upregulation of DSC2 could still effectively mediate EBV infection in EPHA2 knockout EBV-susceptible epithelial cells (middle panel). Moreover, ectopic expression of human DSC2 enabled the cells to acquire susceptibility to EBV in non-EBV-susceptible cells (right panel). This figure was created with BioRender.com.
Supplementary information
Supplementary Tables 1–3 (download XLSX )
Supplementary Table 1. Results of MAGeCK analysis for CRISPR–Cas9 screen. Supplementary Table 2. siRNA sequences. Supplementary Table 3. Primer sequences for qPCR or RT–qPCR.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download TIF )
Unprocessed western blots.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download TIF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download TIF )
Unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download TIF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download TIF )
Unprocessed western blots.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, H., Li, YC., Pang, D. et al. Desmocollin 2 is a dominant entry receptor for Epstein–Barr virus infection of epithelial cells. Nat Microbiol 10, 2768–2780 (2025). https://doi.org/10.1038/s41564-025-02067-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02067-8
This article is cited by
-
Identified receptors shed light on Epstein–Barr virus infection
Nature Microbiology (2025)