Abstract
Environmental differences in genetic effect sizes, namely, gene–environment interactions, may uncover the genetic encoding of phenotypic plasticity1,2,3. We provide a cross-population atlas of gene–environment interactions comprising 440,210 individuals from European and Japanese populations, with replication in 539,794 individuals from diverse populations. By decomposing the contributions from age, sex and lifestyles, we delineate the aetiology of these gene–environment interactions, including a reverse-causality from a disease-related dietary change. Genome-wide analyses uncovered missing heritability and trait–trait relationships connected by the synergistic effects of genome and environments, which systematically affected polygenic prediction accuracy and cross-population portability. Single-cell projection revealed aging shift of pathways and cell types responsible for genetic regulation. Omics-level gene–environment analyses identified multiple sex-discordant genetic effects in lipid metabolism, informing clinical trial failures for genetically supported drug development. Our comprehensive gene–environment study decodes the dynamics of genetic associations, offering insights into complex trait biology, personalized medicine and drug development.
Similar content being viewed by others
Main
There has been great success in human genetics—particularly genome-wide association studies (GWAS)—at revealing disease pathophysiology and complex traits biology4. Genetic association mapping on multi-omics layers has covered proteomics5, metabolomics6 and single-cell RNA sequencing (scRNA-seq)7, providing granular insights into trait-associated genetic loci. However, such efforts focus on fixed genetic effects (more precisely, marginal effects), oversimplifying the intrinsic complexity of trait biology1. Essentially, human phenotypes show dramatic changes in response to multifactorial environmental exposures, including sex, senescence and lifestyle. Inter-individual heterogeneity in responses to environments has been shaped by genetic adaptation8,9 and affects present disease risks2 and drug efficacy10. Genetically, this phenotypic plasticity manifests as changes in genetic effect sizes across environmental factors (or, equivalently, changes in environmental effects across genotypes), namely, gene–environment (G×E) interactions3 (Fig. 1a). G×E interactions capture dynamic changes in genetic effects, unveiling the genetic regulation of phenotypic plasticity. In some traits, G×E interaction studies have begun to explain phenotypic variation not captured by marginal effects (that is, missing heritability) and disparities in polygenic risk prediction11. Therefore, identifying G×E interactions may contribute to mitigating health disparities and implementing personalized medicine precisely12.
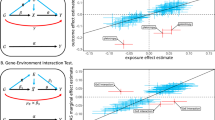
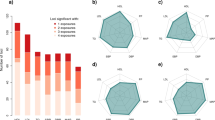
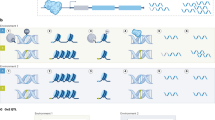
a, A schematic of gene–environment (G×E) interactions. b, Overview of the cohorts. EUR, European; AFR, African; AMR, American; EAS East Asian. c, Circular plot of genome-wide significant G×E interactions in UKB, estimated using two-sided linear regression for quantitative traits and Firth logistic regression for dichotomous traits. Dot sizes represent replication P values in UKB2 and BBJ2. Pleiotropic loci are indicated by lines, with the nearest genes to the lead variants. d, Numbers of genome-wide significant loci per trait in UKB. Bar colours represent trait categories; the colour intensity indicates the types of pleiotropy. e,f, As in d (e) and c (f), for BBJ (see Supplementary Table 1 for the sample sizes used in c–f). AST, aspartate aminotransferase; ALT, alanine aminotransferase; BC, red blood cell; BS, blood sugar; CAD, coronary artery disease; CRP, C-reactive protein; Eosino, eosinophils; Hb, haemoglobin; Ht, haematocrit; Lymph, lymphocytes; MCH, mean corpuscular haemoglobin; MCHC, mean corpuscular haemoglobin concentration; MCV, mean corpuscular volume; P, phosphate (UKB)/inorganic phosphorus (BBJ); Plt, platelets; PP, pulse pressure; sCr, serum creatinine; T2D, type 2 diabetes; T.Bil, total bilirubin; TC, total cholesterol; TG, triglycerides; TP, total protein; UA, uric acid.
Nevertheless, after decades of effort3, there is a limited number of established G×E interactions in humans13, and their biological interpretation remains underestablished14. Past studies have suffered from low replication rates15,16 due to low statistical power17, heavy multiple-testing burdens18, arbitrary filtering of genetic variants15 and, in some cases, imprecise statistical testing19. G×E interactions have been studied at scale only for limited traits and environments, primarily in European populations20. Therefore, a global overview of G×E interactions across phenotypes, environments and populations remains unknown.
Here, using the recent advent of population-scale biobanks21 and computationally efficient methods22, we conducted parallel genome-wide G×E interaction studies using UK Biobank (UKB) and Biobank Japan (BBJ) to provide a cross-population atlas of G×E interactions. We validated the identified G×E interactions in four independent cohorts with diverse populations, annotated the environmental contributors and assessed their impacts on heritability, polygenic prediction accuracy and responsible cell types. Multi-omics G×E analyses provided molecular insights into clinical G×E interactions. These multi-resolution analyses demonstrated that G×E interactions have pivotal roles in regulating dynamic phenotypic plasticity, informing personalized phenotype prediction and drug development.
G×E interactions in individual biobanks
To reliably detect G×E interactions, we divided UKB and BBJ into discovery and replication cohorts (UKB1, Nmax = 273,453; UKB2, Nmax = 38,149; BBJ1, Nmax = 166,757; BBJ2, Nmax = 65,373) (Fig. 1b and Supplementary Table 1). Targeting 38 biomarkers and 9 diseases, which spanned 10 categories (anthropometric, metabolic, proteins, kidney-related, electrolytes, liver-related, inflammatory, haematological, blood pressure and diseases), G×E interactions were tested for nine environmental factors individually and jointly, with P values aggregated per variant on the Cauchy distribution23 to assess genome-wide significance (Extended Data Fig. 1 and Supplementary Tables 2 and 3). The environmental factors included age, sex, ever-drinking, ever-smoking and current-smoking, and four clusters for diet and physical activity derived from questionnaire data (Extended Data Fig. 2 and Supplementary Table 4).
In UKB1, we identified 64 genome-wide significant G×E interactions at 45 loci spanning all trait categories (PG×E < 5.0 × 10−8), with 31 interactions at 23 loci remaining after Bonferroni correction (PG×E < 5.3 × 10−10), indicating that G×E interactions were widespread in human complex traits (Fig. 1c,d and Supplementary Tables 5 and 6). These included known interactions, G×Current-smoking at the HYKK locus for body mass index (BMI)24, G×Age at the UMOD locus for estimated glomerular filtration rate (eGFR)25, and G×E at the FTO locus for BMI driven by multiple environments such as physical activity, diet, age, drinking and smoking26, empirically validating our results. The remaining interactions—based on our curation of GWAS Catalog27—were not reported at PG×E < 5.0 × 10−8 (Supplementary Note 2 and Supplementary Table 7). In total, 16 loci overlapped with recent UKB G×E reports using different protocols13,28 (Supplementary Note 3 and Supplementary Table 8). We observed pleiotropy at 13 loci (10 intra- and 3 inter-categorical), with 2 inter-categorical loci showing distinct significant variants across trait categories (Extended Data Fig. 3), suggesting trait-category specificity of G×E pleiotropy in contrast to the broader pleiotropy of marginal effects29.
In BBJ1, 36 significant G×E interactions were detected across 15 loci (26 across 8 loci after Bonferroni correction) (Fig. 1e,f and Supplementary Table 6). These included the well-established locus in the European population, that is, the FTO locus for BMI, which we confirmed in the East Asian population (driven by G×Age and G×Ever-drinking). Other loci with PG×E < 5.0 × 10−8 have not been reported in the GWAS Catalog, emphasizing the importance of studying G×E interactions in non-European populations. Notably, 58% (21 out of 36) of G×E interactions were at the ALDH2 locus, which harbours an East-Asian-specific missense variant (rs671) with a strong dominant effect on alcohol metabolism30, consistent with its high pleiotropy for GWAS29.
Inflation was minimal in both cohorts (Supplementary Table 5), and the results were robust to phenotype normalization (Supplementary Table 9). A stepwise variable selection approach revealed that a mean of 2.4 environments contributed to G×E interactions (range = 1–7; Supplementary Table 6), supporting our approach of testing G×E interactions both individually and jointly across environments. In 91% (10 out of 11) of the intra-categorical pleiotropic loci, at least one shared environment contributed to all traits, suggesting that the combinations of trait categories and environments were major determinants of G×E pleiotropy.
Replication within populations
Of the 64 G×E interactions in UKB1, 23 were nominally replicated in UKB2 (PG×E < 0.05), and 6 remained significant after Bonferroni correction (Fig. 1c and Supplementary Table 6). In BBJ, 28 out of 36 G×E interactions were nominally replicated, and 19 were significant in BBJ2 (Fig. 1f and Supplementary Table 6). These included a G×Ever-drinking interaction at the ALDH2 locus for haemoglobin in BBJ1 (PG×E = 2.2 × 10−15; Pmarginal = 1.7 × 10−3), replicated in BBJ2 (PG×E = 2.9 × 10−9; Pmarginal = 8.9 × 10−3), highlighting context-specific effects that would be missed by marginal genetic tests. The same locus also showed a significantly replicated G×Ever-drinking interaction for type 2 diabetes (PG×E = 4.8 × 10−16 in BBJ1 and 1.2 × 10−6 in BBJ2). This interaction remained significant after adjusting for haemoglobin (PG×E = 8.1 × 10−15 in BBJ1), suggesting minimal mediation by haemoglobin. For replication, we stringently required consistent environments and effect directions; failure in either was deemed non-replicated, regardless of P values. Replication rates were comparable with those in GWAS, supporting the robustness of our findings (Supplementary Note 4).
We further tested replication in independent cohorts: the European population in All of Us (Nmax = 208,700) and the East Asian population in two Japanese cohorts: (1) the Japan Multi-Institutional Collaborative Cohort–Hospital-based Epidemiologic Research Program at Aichi Cancer Center (J-MICC/HERPACC) (Nmax = 70,909), and (2) the Japan Public Health Center-Based Prospective Study (JPHC) (Nmax = 10,904) (Supplementary Table 10). Despite differences in lifestyle questionnaires, dietary clusters (for example, ‘Japanese cuisine (Washoku)’ in all Japanese cohorts, and ‘meat and cheese’ in UKB1 and JPHC) were consistently recovered across cohorts, supporting the robustness of our clustering approach (Extended Data Fig. 4). Bonferroni replication rates were 27% in All of Us (17 out of 64 trait–locus pairs) and 56% in J-MICC/HERPACC (20 out of 36; Extended Data Fig. 5a,b and Supplementary Table 11). Notably, the pleiotropic G×E interactions at the ALDH2 locus were replicated at 81% (17 out of 21) in J-MICC/HERPACC, with six also replicated in JPHC despite its smaller sample size.
Collectively, our approach using cross-population biobank resources thoroughly detected and validated G×E interactions across diverse trait categories and environments.
Cross-population consistency
Combining UKB and BBJ results yielded 94 trait–locus pairs across 54 loci. Six loci were shared between biobanks (40% of BBJ1 loci; Fig. 1c,f), often involving essential (‘core’) genes for the target phenotypes. For example, ALPL for alkaline phosphatase (ALP) was commonly driven by G×Sex, GGT1 for γ-glutamyl transpeptidase (GGT) by G×Sex, G×Age and G×Ever-smoking, and UMOD for eGFR by G×Age (Extended Data Fig. 5c–f).
To assess cross-population sharing more broadly, we examined replication in the other population’s discovery cohort. After excluding the ALDH2 locus (rs671) to avoid introducing bias due to its well-known East Asian specificity, 22 out of 73 interactions were nominally (and nine significantly) replicated (PG×E < 6.8 × 10−4). The conservative signal-sharing estimate (Storey’s π1; ref. 31) was 0.41, indicating moderate consistency of G×E interactions across populations. The cross-population-replicated loci included three that were originally from UKB1: G×Age at the APOE locus for total cholesterol; G×Sex at the ABCG2 locus for urate; and G×Sex and G×Physical-activity at the SURF6 locus for ALP (PG×E = 5.7 × 10−4, 3.1 × 10−5 and 1.5 × 10−4 in BBJ1, respectively). These shared G×E interactions suggested that cross-population meta-analyses would be beneficial. Indeed, we detected one additional G×E interaction through a meta-analysis across BBJ and UKB (Supplementary Note 5 and Supplementary Table 12).
Minor allele frequencies of the lead variants tended to be higher in the population where the G×E interactions were detected (Extended Data Fig. 5g). Population specificity (that is, dietary environments) and differing distributions of environments probably also contributed to the population-specific G×E detection (Extended Data Fig. 5h).
We further evaluated replication in the African and American populations in All of Us (Nmax = 70,558 and 66,556, respectively) and the Israeli population in the Human Phenotype Project (HPP; Nmax = 8,645). Three G×E interactions were significantly replicated in the African population—two of which overlapped with UKB1–BBJ1 shared loci (Extended Data Fig. 5a and Supplementary Table 13). In the American population, two G×E interactions for pulse pressure—primarily driven by G×Age—were significantly replicated. Although no G×E interaction reached significance in the Israeli population, possibly due to its small sample size, the top signal aligned with a UKB1–BBJ1 shared interaction. These findings demonstrated both shared and population-specific G×E interactions. Subsampling analyses revealed that detecting G×E interactions required biobank-scale sample sizes, and detected loci were not saturated (Extended Data Fig. 5i), encouraging future global collaboration to thoroughly capture worldwide G×E interactions.
Environments contributing to G×E
Gene–environment interactions can enhance locus interpretation by revealing context-specific genetic associations. In UKB1, diet-related environments contributed to five G×E interactions—all of which are at least nominally replicated. These included the ABCG2 locus, where association with eGFR was specific to non-consumers of ‘meat and cheese’ (PG×E = 1.5 × 10−14; Fig. 2a,b). Raw questionnaire data confirmed that low meat consumption unmasked the genetic effect (Extended Data Fig. 6). As ABCG2 encodes a primarily intestinal urate exporter32 and urate is the end-product of purine metabolism, high purine intake from meat may obscure the genetic effect.
a,b, Urate levels across rs4148155 genotypes (the ABCG2 locus) and ‘meat and cheese’ consumption in UKB1 (a) and UKB2 (b). Dots show 5,000 randomly sampled individual measurements per genotype; lines represent the linear regression lines using all participants. c,d, AST (c) and haemoglobin (d) levels by rs671 genotypes (the ALDH2 locus) and drinking status in BBJ1. Boxplots show the median, quartiles and 1.5× interquartile range (see Supplementary Table 14 for exact P values). e, Schematic of the mechanisms of action for warfarin, natto (fermented soybean) and DOACs. The heart icon is from TogoTV (2016 DBCLS TogoTV, CC-BY-4.0). f, Heatmap of the arrhythmia prevalence in BBJ1. g, As in f, but stratified by arrhythmia subgroups. h, Natto intake before and after warfarin initiation in BBJ atrial fibrillation or flutter patients with ≥1 prothrombin time-international normalized ratio (PT-INR) record per year, considering that regular monitoring of PT-INR is required during warfarin therapy. i, Odds ratios of rs72900155 (the PITX2 locus) for arrhythmia stratified the natto intake frequency. N = 109,642 and 49,116 for natto-consuming and non-consuming participants in BBJ1, and N = 44,954 and 18,155 for those in BBJ2, respectively. Data are presented as estimated values with 95% confidence intervals (Firth logistic regression). j, Heatmap of the atrial fibrillation or flutter prevalence in BBJ2, stratified by warfarin and DOAC medication.
In BBJ1, although ALDH2 primarily affects alcohol metabolism, multiple environments contributed to the pleiotropic G×E interactions at the ALDH2 locus after adjusting for G×Drinking interactions (Supplementary Note 6 and Supplementary Table 14). Stratified analysis in ever- and never-drinkers revealed that 12 of 19 biomarkers showed strong non-additive effects in ever-drinkers (Pnon-additive < 5.0 × 10−8; Fig. 2c and Supplementary Table 15), consistent with the dominant deleterious effect on alcohol metabolism of the lead variant (rs671). Four haematopoietic traits showed purely additive effects in never-drinkers (red blood cells, haemoglobin, haematocrit and white blood cells; Padditive < 5.0 × 10−8 and Pnon-additive > 0.05; Fig. 2d), opposite to the functional role and inheritance pattern of rs671. These effects were all replicated in BBJ2 (Padditive < 0.05/19 = 2.6 × 10−3 and Pnon-additive > 0.05). Given the long-range linkage disequilibrium (~2.44 Mb) with signs of recent selection33 at this locus, other causal variants or genes may underlie these haematopoietic associations. The region harbours haematopoiesis-related genes (for example, SH2B3 and PTPN11), whose roles in common-variant genetics warrant further investigation. To facilitate future research, we applied a deep learning model to prioritize variant–gene pairs with potential regulatory effects (Supplementary Note 7 and Supplementary Table 16). These loci together demonstrated the utility of G×E interactions for gaining biological insights into genetic loci.
A reverse-causal G×E interaction
In BBJ1, the PITX2 locus for arrhythmia showed a G×E interaction primarily driven by natto (fermented soybean) intake (PG×E = 2.8 × 10−12; PG×E = 2.1 × 10−10 when testing G×Natto alone; Extended Data Fig. 7). The lead variant, rs72900155, has been reported to be associated with atrial fibrillation29—a subgroup of arrhythmia. Clinically, warfarin, a long-standing anticoagulant, may link natto and atrial fibrillation. As vitamin K in natto reduces the anticoagulation effect of warfarin, patients on warfarin are advised to avoid it (Fig. 2e). In BBJ1, the arrhythmia prevalence was markedly high in the homozygous carriers of natto non-consumers (Fig. 2f), and this pattern was primarily driven by atrial fibrillation or atrial flutter, for which warfarin was the sole first-line anticoagulant before the launch of direct oral anticoagulants (DOACs) (Fig. 2g). In this subgroup, natto intake declined markedly after warfarin initiation in the same individuals (Fig. 2h), suggesting that this G×E interaction was driven by reverse causality from the disease to the environment.
The marginal effect size of rs72900155 was substantially larger for atrial fibrillation or flutter than for the other subgroups (0.49 (95% CI, 0.46–0.52) versus 0.13 (0.10–0.15)). Owing to this effect size heterogeneity, the overall effect size for arrhythmia would vary with the proportion of atrial fibrillation or flutter patients across natto intake strata, explaining the link between the reverse causality and the G×Natto interaction. Consistently, the G×Natto interaction was not significant in either subgroup when evaluated separately (PG×E = 0.84 for atrial fibrillation or flutter; 2.6 × 10−4 for the other subgroups).
This G×E interaction was not replicated in BBJ2 (PG×E = 0.40; Fig. 2i). Notably, this replication failure might be reasonable as BBJ1 participants were recruited from 2003 to 2008, whereas most BBJ2 participants (84.9%) were recruited from 2013 to 2017, and between these periods, DOACs replaced warfarin in more than half of atrial fibrillation patients in Japan34. As DOACs do not require natto restriction (Fig. 2e), the atrial fibrillation or flutter patients taking DOACs in BBJ2 did not show increased natto avoidance (Fig. 2j).
In summary, we identified the G×E interaction driven by reverse causality. Although machine-learning-based locus interpretation is increasingly investigated35, these results indicate that this technology is not readily applicable to G×E interactions, and careful interpretation by specialists is necessary to disentangle their causal mechanisms.
To evaluate causal directions at other loci, we leveraged repeat biomarker measurements to sort out temporal ordering from environments to phenotypes, and performed time-to-event Cox analyses for disease onset and overall survival (Supplementary Note 8 and Supplementary Tables 17 and 18). We identified a significant G×E interaction for overall survival at the ALDH2 locus driven by interactions with sex, ever-drinking and age (P = 1.7 × 10−11), suggesting potential G×E effects on human lifespan.
Pleiotropic G×E effects on diseases
We conducted phenome-wide G×E interaction analyses to assess pleiotropy on diseases (Supplementary Table 19). In UKB1, we detected one additional G×E interaction at the APOE locus for dyslipidaemia primarily driven by G×Sex (PG×E = 3.8 × 10−7; false discovery rate (FDR) < 0.05), consistent with the G×E interactions in the main analyses for cholesterol biomarkers (total cholesterol, triglycerides and low-density lipoprotein cholesterol (LDL-C); Extended Data Fig. 8 and Supplementary Table 20). In BBJ1, 11 G×E interactions were also significant. The ALDH2 locus exhibited widespread G×E pleiotropy across diseases, including the established G×Drinking interaction for oesophageal cancer36. We also observed a G×Age interaction at the HLA-DQB1 locus for rheumatoid arthritis (PG×E = 1.2 × 10−6), originally detected for asthma in BBJ1 and immune cells in UKB1 (lymphocytes, eosinocytes and white blood cells), suggesting shared G×E effects across immune phenotypes. These results showed that the G×E interactions for clinical biomarkers also affected disease statuses through pleiotropy.
Genome-wide heritability
We estimated G×E heritability to evaluate consistency across populations at the genome-wide level37. Evaluating individual environments, 14 and 12 trait–environment pairs were significant in UKB1 and BBJ1, respectively (FDR < 0.05; Supplementary Table 21), although statistical power was limited by multiple testing burden. When aggregating across environments, G×E heritability was significantly positive for seven traits in UKB1 and 11 traits in BBJ1, including four overlapping traits: height, BMI, high-density lipoprotein cholesterol (HDL-C) and diastolic blood pressure (DBP) (Fig. 3a–c). G×E-to-marginal heritability ratio was much larger for BMI than height (0.100 (95% CI, 0.057–0.142) versus 0.028 (0.007–0.048) in UKB1; 0.245 (0.130–0.360) versus 0.062 (0.009–0.115) in BBJ1), replicating a previous report in the European population38 and suggesting a shared G×E architecture across populations for the anthropometric traits. Other traits with significant G×E heritability showed ratios ranging from 0.03 to 0.52, indicating heterogeneity in G×E contributions across traits (Supplementary Table 21). Notably, G×E heritability across quantitative traits was significantly correlated between biobanks (Spearman’s ρ = 0.41, P = 0.011; Fig. 3d), suggesting moderately concordant G×E contributions across populations.
a, G×E heritability (h2) in UKB1, aggregated across all environmental factors (Methods). Asterisks denote FDR < 0.05. BUN, blood urea nitrogen; DL, dyslipidaemia; MAP, mean arterial pressure; WBC, white blood cells. b, As in a, for BBJ1. c, Heritability of G×E interactions and marginal effects for traits with significantly positive G×E heritability in UKB1 (left) and BBJ1 (right). d, Scatter plot of G×E heritability in UKB1 and BBJ1. Regression line estimated by Deming regression to account for estimation errors in both x and y axes. P values were estimated with a two-sided Spearman’s rank correlation test. For a–d, data are presented as estimated values with 95% confidende interval (see Supplementary Table 1 for the sample sizes). e, Significant genetic correlations (Rg, FDR < 0.05) of marginal effects (grey lines) and G×E interactions (coloured lines), estimated in UKB1. The widths of lines represent the strength of genetic correlations; dashed lines represent negative genetic correlations. f, As in e, for BBJ1.
We next estimated the cross-trait correlation of G×E interactions. Significant correlations were observed for 20 and 29 trait–environment pairs in UKB1 and BBJ1, respectively (FDR < 0.05; Supplementary Table 22). Although marginal genetic correlations formed a single cluster, G×E correlations were clustered by trait categories (Fig. 3e,f). Notably, the same environmental factors mediated G×E correlations across populations: current smoking in liver-related traits; sex and dietary consumption in blood pressure traits; and age and sex in renal-related traits. These trait–environment relationships were consistent with known epidemiology and recovered without previous clinical input, suggesting that the trait–environment relationships were embedded in the genome-wide G×E architecture.
Unfiltered approach for G×E detection
Past studies have often limited G×E analyses to prefiltered variants to reduce multiple testing burden15,18. A common approach is variance quantitative trait loci (vQTL) analysis, which tests associations between genotypes and phenotypic variance without requiring environmental measurements13. In Supplementary Note 9, we examined overlaps between G×E interactions and vQTL. G×E loci were 14.6-fold enriched for vQTL compared with GWAS loci, supporting vQTL as an effective prefiltering strategy13. However, vQTL missed most G×E interactions (54.8% in UKB1 and 80.6% in BBJ1; Supplementary Table 23) and their detection was sensitive to phenotype normalization. Moreover, vQTL heritability did not correlate with G×E heritability across traits. These results underscore the necessity of using environmental data explicitly for comprehensive G×E detection.
Influence on polygenic prediction
Polygenic score (PGS)-based disease risk prediction is actively explored, but environmental differences within and across populations can reduce its prediction accuracy for specific traits11,39, potentially exacerbating health disparities. This might affect other traits in general, considering the G×E heritability for broad trait categories. To systematically assess environmental effects on polygenic prediction, we stratified the discovery and replication cohorts by environments into two groups (for example, ever-smokers and never-smokers; younger and older halves of the group). We performed GWAS and constructed PGS within individual strata of the discovery cohorts, and evaluated their prediction accuracy in the strata of the replication cohorts (Fig. 4a).
a, Overview of PGS construction and evaluation of within-population prediction accuracy and cross-population portability. b,c, Examples of within-population prediction accuracy (R2): PP in UKB stratified by age (b) and uric acid in BBJ stratified by drinking status (c). Asterisks represent FDR < 0.05. d, BMI distribution stratified by G×E-PGS, and that from marginal PGS. Boxplots show median, quartiles and 1.5× interquartile range whiskers. N = 30,683 and 40,226 for male and female participants, respectively. e, UMAP of the Tabula Sapiens scRNA-seq data. We used the 10x Genomics subset and subsampled 100 cells per tissue–anatomy–cell type combination (Methods). Cell types defined in the original study. f,g, Single-cell associations with PP genetics in the younger (f) and older (g) groups. h, UMAP of the monkey artery scRNA-seq data. Cell types defined in the original study. i, Difference in single-cell associations between the older and younger groups.
Among the 26 trait–environment–biobank triplets with significantly positive G×E heritability, 20 exhibited significant intra-population differences in prediction accuracy in at least one stratum (FDR < 0.05; Fig. 4b,c, Extended Data Fig. 9a and Supplementary Table 24). Prediction accuracy was generally the highest when applied to the same environmental group at PGS construction. Excluding one related to a UKB1-specific environment cluster (BMI–fish-and-vegetable), 11 out of 25 triplets also showed significant differences in cross-population portability (Extended Data Fig. 9b–d), although the prediction accuracy was generally attenuated.
Polygenic scores constructed from G×E interactions (G×E-PGS)40 consistently significantly explained phenotypic variance in independent cohorts (11 out of 25 (9 out of 25) trait–environment pairs within (across) populations; Supplementary Table 25). Notably, a G×Sex-based PGS constructed in BBJ1 successfully stratified BMI in opposite directions between sexes in J-MICC/HERPACC, capturing the polygenic architecture of sex differences (Fig. 4d). This stratification was apparent even for each sex and among individuals with similar marginal PGS, suggesting that PGS extended to two dimensions could enhance phenotype prediction. Indeed, a model incorporating G×E-PGS improved BMI prediction accuracy by 16% over a model without G×E-PGS (R2 = 0.128 versus 0.110). By contrast, gains in prediction accuracy were modest for other trait–environment pairs (Supplementary Table 25). Larger sample sizes and methods tailored for G×E-PGS construction are warranted to fully realize the potential of G×E interactions in precision medicine.
Collectively, these observations demonstrated that environmental factors systematically impacted intra- and cross-population PGS prediction accuracy, and incorporating G×E interactions could enhance genetic risk prediction of human complex traits.
Aging shift of pulse pressure genetics
As genome-wide G×E architecture recapitulated epidemiologically plausible trait–environment relationships, we reasoned that G×E interactions could uncover biological mechanisms underlying the genetic dynamics of complex traits. We focused on G×Age interactions for pulse pressure, given their strong signals: all four G×E loci in UKB1 were driven by G×Age, G×Age heritability was significantly positive and PGS prediction accuracy varied across age groups (Supplementary Tables 6 and 21, and Fig. 4b).
We divided BBJ1 and UKB1 into two equal-sized age groups and conducted GWAS for pulse pressure within each group. Cross-population meta-analyses of MAGMA gene-set analyses41 revealed that vascular smooth muscle contraction was enriched in younger individuals and cellular senescence enriched in older individuals (Extended Data Fig. 9e). When projecting polygenic effects onto tissue-wide scRNA-seq data from Tabula Sapiens7,42, blood vessel cell types were significantly associated with pulse pressure in both age groups, whereas their relative strength of associations differed by age (Fig. 4e–g and Supplementary Table 26). To examine this closely, we repeated the analysis using a scRNA-seq dataset of monkey arteries43 (Fig. 4h). In younger individuals, genetic effects were associated with smooth muscle cells (P = 8.3 × 10−3 and 9.4 × 10−3 for the two subtypes), whereas in older individuals they were associated with a subgroup of coronary endothelial cells (P = 2.3 × 10−3) (Fig. 4i and Extended Data Fig. 9f,g). As endothelial cells are central to vascular senescence and atherosclerosis44, these results support the findings of the gene set enrichment analysis.
As pulse pressure was defined as the difference between systolic and diastolic blood pressure (SBP and DBP, respectively), we estimated age-stratified genetic correlations among these traits. Although the genetic correlation (Rg) with pulse pressure remained stable for SBP, that for DBP declined in older individuals (from Rg of 0.46 (95% CI, 0.40–0.53) to 0.27 (0.19–0.35); Extended Data Fig. 9h), indicating relatively increased SBP influence with aging. Cross-age genetic correlation for pulse pressure was modest as expected (Rg = 0.42 (0.35–0.49); Extended Data Fig. 9i).
Collectively, these results suggested that the age-related changes in pulse pressure genetics are driven by increasing SBP influence over DBP, reflecting a shift from smooth muscle-mediated regulation in youth to an endothelial-driven SBP increase with vascular aging. Our results demonstrated that G×E interactions can reveal dynamic trait biology missed by typical GWAS.
Sex-discordant regulation in metabolites
In addition to single-cell analysis, molecular QTL mapping can offer granular biological insights into genetic loci. We analysed 2,924 proteins (Nmax = 28,561 in UKB1 and 2,153 in BBJ1; Supplementary Table 27) and 325 metabolites (Nmax = 153,410 in UKB1 and 89,040 in BBJ1; Supplementary Table 28) for 57 lead variants. We identified 13 significant protein–locus pairs in meta-analyses across biobanks (8 loci; FDRG×E < 0.05; Supplementary Table 29), with the ALDH2 and SURF6 loci reaching genome-wide significance. For metabolites, 2,326 significant metabolite–locus pairs (38 loci) were detected, and 650 (15 loci) passed genome-wide significance, aided by their large sample sizes (Supplementary Table 30). These omics-level G×E interactions covered 70% (40 out of 57) of loci, with the same environments driving both omics and clinical G×E interactions at all loci, indicating that most clinical G×E interactions were detectable at the molecular level.
The relationship between G×E and marginal P values exhibited five distinct patterns across 15 loci with the genome-wide significant metabolite G×E interactions (Fig. 5a and Extended Data Fig. 10): (1) high PG×E–Pmarginal correlation, (2) G×E-specific signals for 1–2 lipid metabolites, (3) bimodal distribution at the TNFAIP8 locus, (4) significant PG×E for 1–2 non-lipid metabolites; and (5) much smaller Pmarginal for most metabolites. The first three patterns were related to lipid metabolites. In the first pattern, metabolites highly correlated with clinical lipid biomarkers formed distinct clusters with a broad P -value spectrum, suggesting that clinical lipid biomarkers may adequately capture genetic and G×E structure at these loci.
a, P values of marginal effects and G×E interactions for metabolites in five representative loci (from left to right: UGT2B17, CETP, TNFAIP8, CPS1 and TRIB1), estimated by two-sided linear regression. Triangles denote P < 1.0 × 10−50 (see Extended Data Fig. 10 for all 15 loci with genome-wide significant G×E interactions). LDL_TG_pct, triglycerides to total lipids in LDL percentage. b, Effect sizes of rs12720908 (the CETP locus) on LDL_TG_pct in UKB1, stratified by sex and age (see Supplementary Table 31 for sample sizes). c, Hazard ratios of LDL_TG_pct on all-cause and CAD-caused mortality in UKB1, stratified by sex and statin medication at registration (see Supplementary Table 32 for sample sizes). In b and c, data are presented as estimated values with 95% CI.
In the second pattern, the nearest genes were involved in lipid metabolism, and strong G×E-specific signals were detected for lipid metabolites not highly correlated with clinical biomarkers, suggesting that direct metabolite measurement was necessary to detect G×E at these genes. For these loci, sex was the top contributor in 86% of G×E metabolite QTL (568 out of 664 metabolite–locus pairs). Among them, cholesteryl ester transfer protein (CETP) is an intriguing target of genetics-driven drug discovery. Although CETP inhibitors showed promise in GWAS for raising HDL-C and lowering LDL-C to reduce coronary artery disease risk45, several were discontinued in phase 3 trials. The CETP locus showed a G×E-specific signal for the percentage of triglycerides in LDLs (LDL_TG_pct; PG×E = 1.8 × 10−12, Pmarginal = 0.98). Given the role of CETP in exchanging triglycerides from LDLs and other lipoproteins with cholesteryl esters from HDL, LDL_TG_pct might represent a key metabolic process for this protein. Notably, effect directions differed by sex (Fig. 5b and Supplementary Table 31), and LDL_TG_pct predicted all-cause and coronary artery disease mortality in both sexes (hazard ratio per unit of s.d. in UKB1: 1.23 (95% confidence interval, 1.21–1.25) and 1.25 (1.18–1.32), respectively; Fig. 5c and Supplementary Table 32). As the known causal variant (rs1801706) showed increasing effects on CETP expression46,47, these results suggested that CETP inhibition might decrease LDL_TG_pct in men but increase it in women, potentially leading to an increased female mortality risk. We also confirmed a previously reported female-specific effect on clinical LDL-C using the same UKB data10 (Extended Data Fig. 11), although LDL-C hazard ratios were not significantly positive, probably due to statin use. These effects might together help explain the clinical trial failure. We also examined the other loci with the second pattern. All G×E-specific signals showed opposite-effect directions between sexes (Extended Data Fig. 12a), suggesting that sex-discordant genetic regulation may be common in lipid metabolism. Some effect sizes were also varied by age, possibly reflecting age-dependent declines in sex hormones, which warrants further investigation.
The TNFAIP8 locus showed a bimodal P value distribution (third pattern). Marginal effects acted mainly through the subfraction of triglycerides, whereas multiple HDL metabolites (especially very large HDL, XL_HDL) exhibited sex-discordant effects (Extended Data Fig. 12b–d). The nearest gene, TNFAIP8, is implicated in oncogenesis and inflammation, and has recently been shown to bind lipid messengers48. The adjacent gene, HSD17B4, encodes 17-β-hydroxysteroid dehydrogenase 4, a multi-functional enzyme involved in fatty acid and sex steroid metabolism49. This multi-functionality might underlie the bimodal pattern, though distinct genetic influences on HDL and triglyceride subfractions are also possible. As fine-mapping and co-localization analyses could not pinpoint causal variants (Supplementary Note 10), further experiments are warranted to characterize this locus.
Motivated by these G×E metabolite QTLs, we expanded the metabolome-wide G×E analysis to genome-wide variants, identifying 30 (11) genome-wide significant loci in the UKB1 (BBJ1), yielding 736 (228) metabolite–locus pairs. Among these, 16 (4) loci in the UKB1 (BBJ1) passed the Bonferroni threshold. Notably, G×Sex interactions at the ALDH1A2 and ZNF259 loci were observed in both cohorts (Supplementary Table 33), and several loci showed sex-discordant effects (Extended Data Fig. 13 and Supplementary Table 34), again underscoring the sex-specific genetic architecture of metabolome regulation. Although not Bonferroni-significant, we also detected current-smoker-specific, never-drinker-specific and G×E-only associations (Extended Data Fig. 13), which warrant further validation and functional studies.
Collectively, our analysis demonstrated that omics G×E studies could provide granular insights into the molecular basis of genetic effect plasticity, particularly for the prominent sexual dimorphism in lipid metabolism.
Discussion
We provided a G×E atlas across the genome, phenomes and environments in two populations and tested replication in diverse populations, substantially expanding the catalogue of human G×E interactions. Leveraging this atlas, we demonstrated that G×E interactions yielded both granular and holistic insights into biological dynamics at the locus, genome-wide, single-cell and molecular levels. At the locus level, G×E analyses revealed underlying biological mechanisms and highlighted the need for careful interpretation by specialists. At the genome-wide level, G×E interactions affected trait heritability, PGS prediction accuracy and cross-population portability, emphasizing the value of incorporating environmental context into genetic prediction. Single-cell- and omics-level analyses uncovered age-, sex- and other environment-specific effects, underscoring the dynamic and molecular nature of G×E interactions. Although moderate G×E sharing was observed across populations, population-specific signals and limited data from underrepresented groups highlight the need for more diverse cohorts with detailed environmental and omics data (Supplementary Note 11). In conclusion, we provided a rich resource for future genetic studies, establishing the importance of G×E interactions in decoding the dynamics of complex trait biology, refining personalized medicine and informing drug development.
Methods
Biobank Japan
The BBJ is a prospective hospital-based biobank with 267,289 participants, all of whom were diagnosed with at least one of the target diseases of BBJ by physicians at the cooperating hospitals50,51,52. All of the participants provided written informed consent approved by the ethics committees of the Institute of Medical Sciences, the University of Tokyo and RIKEN Center for Integrative Medical Sciences. The BBJ comprises two cohorts, which were genotyped separately: the first (BBJ1, N = 182,536) and second (BBJ2, N = 68,534) cohorts. The participants in BBJ1 were genotyped with the Illumina HumanOmniExpressExome BeadChip or a combination of the Illumina HumanOmniExpress and HumanExome BeadChip, whereas the participants in BBJ2 were genotyped with the Illumina Asian Screening Array. All BBJ1 participants and 17% of the BBJ2 participants (N = 11,716) were recruited from 2003 to 2008. The remaining BBJ2 participants (N = 56,818) were recruited from 2013 to 2017.
Definition of the discovery and replication cohorts
We used BBJ1 (BBJ2) as the discovery (replication) cohort.
Quality control of genotype data
We conducted a quality control of the participants and the genotypes, and excluded sample relatedness in BBJ1 via the same approach described previously53. The genotype data were imputed with 1000 Genomes Project Phase 3 (N = 2,504) and Japanese whole-genome sequencing data (N = 1,037) using Minimac3 software54. We excluded variants with an imputation quality of Rsq < 0.7 or a minor allele frequency (MAF) of less than 0.01, resulting in 7,444,735 autosomal variants analysed in total. We analysed 166,757 participants of the Japanese population as estimated by the visual inspection of principal component analysis (PCA).
In BBJ2, we excluded participants with a low call rate (<0.98) and outliers from the Japanese Hondo (that is, the main islands) cluster estimated on the basis of PCA. We excluded the variants meeting the following criteria: (1) with a low call rate (<0.99); (2) with low minor allele counts (<5); and (3) with a Hardy–Weinberg equilibrium test P value of <1.0 × 10−10. We performed statistical phasing of the genotype data using Shapeit4 (ref. 55) and imputation using Minimac4 (ref. 56) with the same reference panel as used in the discovery cohort. After imputation, we excluded variants with an imputation quality of <0.7 or a MAF less than 0.01. We used King57 to exclude relatives within second degrees, resulting in 65,373 participants being analysed.
UK Biobank
The UKB is a population-based biobank with approximately 500,000 participants recruited between 2006 and 2010, aged 40–69 years58. Participants were genotyped using either the UK BiLEVE Axiom Array or UK Biobank Axiom Array. The genotypes were then imputed by IMPUTE4 software using a combination reference panel of the Haplotype Reference Consortium, UK10K and 1000 Genomes Project Phase 3. We accessed the UKB data under the project number 47821.
Definition of the discovery and replication cohorts
We included British European for the discovery cohort (UKB1), defined as the intersection of the self-reported British participants and the genetically ‘Caucasian’ participants (UK Biobank Data Fields 21000 and 22006), to strictly reduce the inflation of test statistics due to population stratification. We excluded one participant from every related pair within the third degree. For the replication cohort (UKB2), we included all other genetically ‘Caucasian’ participants and excluded one participant from every related pair within the second degree. We also excluded the participants related to any participants in UKB1 within the second degree.
Quality control of genotype data
We analysed 9,813,264 autosomal variants with an imputation quality of >0.7 and MAF of >0.01, which was equivalent to the threshold used in BBJ. We excluded participants with: (1) sex chromosome aneuploidy; (2) a mismatch between genetic and self-reported sex; or (3) outliers for heterozygosity or missing rate.
Descriptions of the independent replication cohorts, J-MICC/HERPACC (refs. 59,60,61), JPHC (ref. 62), All of Us (ref. 63) and HPP (ref. 64) are available in Supplementary Methods.
Quality control of phenotypes and environments
Clinical traits
The definition and quality control of clinical traits were summarized in Supplementary Tables 2 and 3. In brief, we obtained biomarker phenotypes from the initial assessment data for UKB and the medical records for BBJ. For BBJ, we used the biomarker phenotypes measured at the nearest dates to the baseline assessment for the main analyses, whereas we used those measured most recently for the temporal order analyses. For the UKB, we used the baseline assessment data for the main analyses, whereas we used the most recent data from the second to fourth revisit assessments for the temporal order analyses, namely, the first repeat assessment visit, the imaging visit, and the first repeat imaging visit. We applied the same quality controls for both biobanks, including (1) excluding participants with age <18 or age >85; (2) excluding participants with particular disease status that might have affected the phenotype values; (3) correction of phenotype values for participants taking anti-hypertensive medications or statins; (4) excluding outliers whose measured values were outside of three times the interquartile range (upper or lower quartile), or outside three standard deviations from the mean; and (5) applying natural log transformation for the phenotypes with right-skewed distributions. For the temporal order analyses, we restricted the data to those measured at least half a year after the baseline survey.
For disease statuses, we combined diagnoses data (ICD-10), operation data (OPCS-4) and self-reported illness and operation data for UKB. We determined the subtypes of diabetes mellitus based on ICD-10 codes and an established algorithm developed for UKB self-reported data65. We excluded participants with diabetes mellitus other than T2D, and patients with T2D who were also inferred to have type 1 diabetes from both cases and controls. Following a previous report66, we included ischaemic stroke patients as cases only if they had any evidence of stroke other than self-reports and excluded the participants with self-reported stroke from controls. For BBJ, we defined disease statuses based on the union of diagnoses by doctors at the cooperating hospitals and past medical history retrieved from electronic medical records. When testing T2D, we excluded the participants with diabetes mellitus other than T2D from both cases and controls. We defined clinical traits and performed quality control in the replication cohorts in the same manner, as detailed in Supplementary Methods.
Environmental factors
As the resolution of individual questionnaire items was limited due to their few discrete response options, we employed a clustering-based approach to summarize correlated items into high-resolution latent environments67 as summarized in Supplementary Table 4. The questionnaires for dietary consumption and physical activity were available for both UKB and BBJ, with low missing rates (0.01–0.27% in UKB1 and 1.4–13.3% in BBJ1), and were used in this manuscript. We converted the categorical responses into the continuous scale following ref. 68. For example, the responses to dietary consumption in BBJ were ‘Almost every day’, ‘3–4 days a week’, ‘1–2 days a week’ and ‘Rarely’, and we converted them into 7, 3.5, 1.5 and 0, respectively. We treated the responses ‘Do not know’ and ‘Prefer not to answer’ as missing values. For clustering analyses, we first regressed out age, sex, age2, age × sex, and age2 × sex from the environmental factors derived from questionnaires about dietary consumption and physical activities. We then performed consensus clustering analyses using the ConsensusClusterPlus R package69 for the environmental factors. We used (1 − Pearson’s correlation) as the distance between environmental factors and employed the hierarchical clustering algorithm with Ward’s method. We note that we multiplied the values of coffee consumption in UKB1 by −1 as we observed its strong negative correlation with tea consumption. We changed the number of consensus clusters from two to ten and defined the number of clusters based on the stability of the cumulative distribution function curve and the item tracking. We named these clusters based on the questionnaire included in the clusters and used the first principle component as their environmental values. We matched the direction of the first principle component to that of the raw questionnaire data included in individual clusters. The cluster scores were standardized to a mean of zero and a standard deviation of one. Finally, we excluded the outliers whose first principle component were outside of three standard deviations from the mean. The clustering of questionnaires for environmental factors in the replication cohorts was described in the Supplementary Methods and Supplementary Table 4.
Disease statuses used for the phenome-wide association study
In addition to the nine disease statuses used in the main analyses, we defined disease statuses based on ICD-10 in UKB1 and past medical history in BBJ1, as summarized in Supplementary Table 19. For the complications of diabetes mellitus (diabetic nephropathy and retinopathy), we restricted cases and controls to the T2D patients. We tested G×E interactions for the diseases with more than 1,000 cases in individual cohorts. Consequently, we tested 35 diseases for 49 lead variants in UKB1 and 52 diseases for 38 lead variants in BBJ1.
Metabolites measurement
We started from the internally quality controlled Nightingale NMR metabolome measurement data and removed its technical variation using the ukbnmr R package70. Briefly, this package removed the technical variation derived from: (1) the elapsed time from sample preparation to measurement, (2) the position (row and column) of the 96-well plate and (3) a series of measurement dates for each spectrometer. This package also calculated 76 useful biomarkers based on the ratio between directly measured ones, in addition to 249 biomarkers provided by Nightingale NMR (325 biomarkers in total; Supplementary Table 28). We then removed participants aged <18 or >85, or whose measured values were outside of three times the interquartile range (upper or lower quartile) or outside of three standard deviations from the mean. We also removed the following participants for particular biomarkers: participants with renal insufficiency (eGFR < 15 ml min–1 per 1.73 m2) for creatinine; participants with liver diseases, haematological malignancies, nephrotic syndrome or autoimmune diseases for albumin; or participants with diabetes mellitus for glucose and participants with autoimmune diseases for glycoprotein acetyls. We applied natural log transformation to the measurement values and standardized them to mean of 0 and s.d. of 1.
Protein expression measurement
We used the normalized Olink protein expression data (Olink Explore 3072). As implemented by Olink, these data were already bridge-normalized across measurement batches into the unit of normalized protein expression. As we did for other quantitative phenotypes, we removed participants aged <18 or >85, or whose measured values were outside of three times the interquartile range (upper or lower quartile), or outside of three standard deviations from the mean. We used 2,923 proteins with valid measurement data for both BBJ1 and UKB1 (Supplementary Table 27).
G×E interaction testing
All statistical tests were two-sided unless otherwise noted. As phenotypes, we targeted quantitative traits (clinical biomarkers, metabolites, and protein expression measurements), dichotomous traits (disease statuses), and time-to-event traits (disease onset for incidental cases and overall survival). For quantitative and dichotomous traits, we tested G×E interactions using GEM, a fast and scalable implementation of linear and logistic regressions with G×E interaction terms22 (Extended Data Fig. 1). We estimated the model-robust ‘sandwich’ standard errors to suppress the inflation of statistics22. As the case–control imbalance can cause the inflation of test statistics in the logistic regression, we re-estimated the effect sizes and P values using Firth logistic regression for the variants with P values <5.0 × 10−8 for dichotomous traits. For the time-to-event traits, we employed the Cox proportional hazard model implemented in the ‘survival’ R package. Individuals who had already developed the target disease at baseline or developed it within six months from baseline were excluded from the incidental case analyses.
We tested G×E interactions for different sets of environmental factors, as shown in Extended Data Fig. 1, and then aggregated P values across environmental sets on the Cauchy distribution23 to obtain per-variant P values. When P values were 0 (meaning that P values were <1.0 × 10−300), we estimated the precise P values using the multiple-precision floating-point arithmetic with the maximal precision of 10,000 bits, implemented in the Rmpfr R package. Regional G×E interactions were plotted using LocusZoom71. We defined a G×E interaction as significant at the variant level if the aggregated P value was below the genome-wide threshold of 5.0 × 10−8. We also reported the study-wide Bonferroni threshold for full transparency. We did not require that any individual environment’s interaction term be significant on its own, considering that G×E interactions can be driven by multiple environments. Nevertheless, we note that at least one raw P value was smaller than the Cauchy-combined P value in principle, as the Cauchy combination method is not a meta-analysis but rather returns a P value within the range of raw P values.
After evaluating the significance of G×E interactions at the variant level, we determined the order of importance of G×E interaction terms by backward elimination from the regression model with all G×E interaction terms. Specifically, we repeatedly removed the G×E interaction term with the least improvement in the likelihood of the linear regression model or the largest Wald P value of the Firth logistic regression model. After removing all G×E interaction terms, we brought back the G×E interaction terms one by one in the order of their importance if the model fit was improved with a P value less than 0.05 (likelihood ratio test). We considered that the environmental factors contributed to G×E interactions if the corresponding interaction terms were included in the final model. In principle, this approach selects more parsimonious sets of environments than the Akaike information criterion and can determine the top environment contributing to individual G×E interactions.
We used the first 20 genetic principal components, genotype array, age2, age × sex and age2 × sex as covariates for the UKB. For the other cohorts, we used the first ten principle components, age2, age × sex and age2 × sex as covariates. Age and sex, as well as other environmental factors, were included as the interaction terms with genotypes, and the variables used for interaction terms were automatically included as covariates. We also included the status of the target diseases in BBJ as covariates for quantitative traits and overall survival. We included the status of statin medication as an additional covariate for metabolome measurements. For sex-specific diseases in the phenome-wide association study, we excluded sex from the environmental factors and age × sex and age2 × sex from covariates.
Following that previous cross-population GWAS used the distance-based locus definition21, we defined that the genome-wide significant variants were in the same locus if their distance was less than 500 kbp. For the genome-wide G×E interaction tests on the proteome and metabolome, we first targeted 56 lead variants of clinical G×E interactions, including those identified by meta-analyses. We then extended the targets on the metabolome to (1) genotyped variants in each cohort, (2) HapMap3 variants and (3) the variants targeted by the meta-analysis across BBJ and UKB, to reduce the computational cost. After defining the metabolome G×E loci based on their distance as above, we merged the metabolome G×E loci if they overlapped with the same locus defined for the clinical traits. See Supplementary Methods for further methodological details.
Ethics statement
This study was approved by the ethics committee of the University of Osaka (approval no. 734-18) and the ethics committee of the Graduate School of Medicine, the University of Tokyo (2023405G-(4)).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The genome-wide summary statistics of G×E interactions for both clinical phenotypes and the metabolome are publicly available at the NBDC Human Database (https://humandbs.dbcls.jp/en) with the accession ID hum0197.v26.374-traits.v1 and at the NHGRI-EBI GWAS Catalogue (https://www.ebi.ac.uk/gwas) with the accession IDs GCST90681837–GCST90690020. The UKB analysis was conducted under application no. 47821 (https://www.ukbiobank.ac.uk/). The BBJ data are available at the NBDC Human Database (https://humandbs.biosciencedbc.jp/en/) via accession IDs JGAS000114 and JGAS000412 (genotype), JGAS000561 (metabolome) and JGAS000785 (proteome). Human reference genome GRCh38, http://ftp.1000genomes.ebi.ac.uk/vol1/ftp/technical/reference/GRCh38_reference_genome/; Japanese human reference genome JG2.1.0, https://jmorp.megabank.tohoku.ac.jp/downloads/tommo-jg2.1.0-20211208; the enformer model v.1, https://www.kaggle.com/models/deepmind/enformer/tensorFlow2/enformer/1; East Asian LD block data, https://github.com/jmacdon/LDblocks_GRCh38/blob/master/data/pyrho_EAS_LD_blocks.bed.
Code availability
We conducted the analyses using the following publicly available tools: GEM v.1.1 (https://github.com/large-scale-G×E-methods/GEM), OSCA v.0.46 (https://cnsgenomics.com/software/osca), bgenie v.1.3 (https://jmarchini.org/bgenie/), LocusZoom v.1.4 (locuszoom.org), LDSC v.1.0.0 (https://github.com/bulik/ldsc), PRScs version 4 June 2021 (https://github.com/getian107/PRScs), MAGMA v1.09a (https://ctg.cncr.nl/software/magma), scDRS v.1.0.2 (https://martinjzhang.github.io/scDRS/), enformer (https://github.com/google-deepmind/deepmind-research/tree/master/enformer), shapeit4 v.4.1.2 (https://odelaneau.github.io/shapeit4/), minimac4 v.1.0.1 (https://github.com/statgen/Minimac4), plink2 v.2.00a6LM (https://www.cog-genomics.org/plink/2.0/), king v.2.2.5 (https://www.kingrelatedness.com/), FINEMAP v.1.4.2 (http://www.christianbenner.com/), GenomeStudio v.2.0.5 (https://support.illumina.com/downloads/genomestudio-2-0.html), Applied Biosystems Analysis Power Tools v.2.12.0 (https://www.thermofisher.com/jp/en/home/life-science/microarray-analysis/microarray-analysis-partners-programs/affymetrix-developers-network/affymetrix-power-tools.html), BCFtools/liftover (https://github.com/freeseek/score), Transanno v.0.4.5 (https://github.com/informationsea/transanno). We also conducted the analyses using the following R packages: logistf v.1.24, mixmeta v.1.2.0, GxEprs v.1.2, survival v.3.2-3, ukbnmr v.2.2, glmnet v.3.1-1, Rmpfr v.0.9-4 (https://cran.r-project.org/web/packages), ConsensusClusterPlus v.1.50.0 (https://bioconductor.org/packages/release/bioc/html/ConsensusClusterPlus.html), ACAT v.0.91 (https://github.com/yaowuliu/ACAT), Seurat v.4.3.0.1 (https://satijalab.org/seurat/) and susieR v.0.12.35 (https://stephenslab.github.io/susieR/).
References
Li, J., Li, X., Zhang, S. & Snyder, M. Gene-environment interaction in the era of precision medicine. Cell 177, 38–44 (2019).
Virolainen, S. J., VonHandorf, A., Viel, K. C. M. F., Weirauch, M. T. & Kottyan, L. C. Gene–environment interactions and their impact on human health. Genes Immun. 24, 1–11 (2022).
Hunter, D. J. Gene–environment interactions in human diseases. Nat. Rev. Genet. 6, 287–298 (2005).
Uffelmann, E. et al. Genome-wide association studies. Nat. Rev. Methods Prim. 1, 59 (2021).
Sun, B. B. et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 622, 329–338 (2023).
Karjalainen, M. K. et al. Genome-wide characterization of circulating metabolic biomarkers. Nature 628, 130–138 (2024).
Zhang, M. J. et al. Polygenic enrichment distinguishes disease associations of individual cells in single-cell RNA-seq data. Nat. Genet. 54, 1572–1580 (2022).
Fan, S., Hansen, M. E. B., Lo, Y. & Tishkoff, S. A. Going global by adapting local: a review of recent human adaptation. Science 354, 54–59 (2016).
Rees, J. S., Castellano, S. & Andrés, A. M. The genomics of human local adaptation. Trends Genet. 36, 415–428 (2020).
Legault, M. et al. Study of effect modifiers of genetically predicted CETP reduction. Genet. Epidemiol. 47, 198–212 (2023).
Kamiza, A. B. et al. Transferability of genetic risk scores in African populations. Nat. Med. 28, 1163–1166 (2022).
Sørensen, T. I. A., Metz, S. & Kilpeläinen, T. O. Do gene–environment interactions have implications for the precision prevention of type 2 diabetes? Diabetologia 65, 1804–1813 (2022).
Westerman, K. E. et al. Variance-quantitative trait loci enable systematic discovery of gene-environment interactions for cardiometabolic serum biomarkers. Nat. Commun. 13, 3993 (2022).
Westerman, K. E. & Sofer, T. Many roads to a gene-environment interaction. Am. J. Hum. Genet. 111, 626–635 (2024).
Dick, D. M. et al. Candidate gene–environment interaction research. Perspect. Psychol. Sci. 10, 37–59 (2015).
Joseph, P. G., Pare, G. & Anand, S. S. Exploring gene–environment relationships in cardiovascular disease. Can. J. Cardiol. 29, 37–45 (2013).
Aschard, H. A perspective on interaction effects in genetic association studies. Genet. Epidemiol. 40, 678–688 (2016).
Aschard, H. et al. Challenges and opportunities in genome-wide environmental interaction (GWEI) studies. Hum. Genet. 131, 1591–1613 (2012).
Boye, C., Nirmalan, S., Ranjbaran, A. & Luca, F. Genotype × environment interactions in gene regulation and complex traits. Nat. Genet. 56, 1057–1068 (2024).
Herrera-Luis, E., Benke, K., Volk, H., Ladd-Acosta, C. & Wojcik, G. L. Gene–environment interactions in human health. Nat. Rev. Genet. 25, 768–784 (2024).
Zhou, W. et al. Global biobank meta-analysis initiative: powering genetic discovery across human disease. Cell Genomics 2, 100192 (2022).
Westerman, K. E. et al. GEM: scalable and flexible gene–environment interaction analysis in millions of samples. Bioinformatics 37, 3514–3520 (2021).
Liu, Y. et al. ACAT: A fast and powerful P value combination method for rare-variant analysis in sequencing studies. Am. J. Hum. Genet. 104, 410–421 (2019).
Justice, A. E. et al. Genome-wide meta-analysis of 241,258 adults accounting for smoking behaviour identifies novel loci for obesity traits. Nat. Commun. 8, 14977 (2017).
Pattaro, C. et al. Genome-wide association and functional follow-up reveals new loci for kidney function. PLoS Genet. 8, e1002584 (2012).
Moore, R. et al. A linear mixed-model approach to study multivariate gene–environment interactions. Nat. Genet. 51, 180–186 (2019).
Cerezo, M. et al. The NHGRI-EBI GWAS Catalog: standards for reusability, sustainability and diversity. Nucleic Acids Res. 53, D998–D1005 (2025).
Bernabeu, E. et al. Sex differences in genetic architecture in the UK Biobank. Nat. Genet. 53, 1283–1289 (2021).
Sakaue, S. et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat. Genet. 53, 1415–1424 (2021).
Koyanagi, Y. N. et al. Genetic architecture of alcohol consumption identified by a genotype-stratified GWAS and impact on esophageal cancer risk in Japanese people. Sci. Adv. 10, ade2780 (2024).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003).
Eckenstaler, R. & Benndorf, R. A. The role of ABCG2 in the pathogenesis of primary hyperuricemia and gout—an update. Int. J. Mol. Sci. 22, 6678 (2021).
Okada, Y. et al. Deep whole-genome sequencing reveals recent selection signatures linked to evolution and disease risk of Japanese. Nat. Commun. 9, 1631 (2018).
Okumura, Y. et al. Current use of direct oral anticoagulants for atrial fibrillation in Japan: findings from the SAKURA AF Registry. J. Arrhythm. 33, 289–296 (2017).
Nicholls, H. L. et al. Reaching the end-game for GWAS: machine learning approaches for the prioritization of complex disease loci. Front. Genet. 11, 350 (2020).
Matsuo, K. et al. Gene-environment interaction between an aldehyde dehydrogenase-2 (ALDH2) polymorphism and alcohol consumption for the risk of esophageal cancer. Carcinogenesis 22, 913–916 (2001).
Shin, J. & Lee, S. H. GxEsum: a novel approach to estimate the phenotypic variance explained by genome-wide GxE interaction based on GWAS summary statistics for biobank-scale data. Genome Biol. 22, 183 (2021).
Robinson, M. R. et al. Genotype–covariate interaction effects and the heritability of adult body mass index. Nat. Genet. 49, 1174–1181 (2017).
Ojima, T. et al. Body mass index stratification optimizes polygenic prediction of type 2 diabetes in cross-biobank analyses. Nat. Genet. 56, 1100–1109 (2024).
Jayasinghe, D. et al. Mitigating type 1 error inflation and power loss in GxE PRS: Genotype–environment interaction in polygenic risk score models. Genet. Epidemiol. 48, 85–100 (2024).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Tabula Sapiens Consortium* et al The Tabula Sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 376, eabl4896 (2022).
Zhang, W. et al. A single-cell transcriptomic landscape of primate arterial aging. Nat. Commun. 11, 2202 (2020).
Jia, G., Aroor, A. R., Jia, C. & Sowers, J. R. Endothelial cell senescence in aging-related vascular dysfunction. Biochim. Biophys. Acta 1865, 1802–1809 (2019).
Schmidt, A. F. et al. Cholesteryl ester transfer protein (CETP) as a drug target for cardiovascular disease. Nat. Commun. 12, 5640 (2021).
Kanai, M. et al. Insights from complex trait fine-mapping across diverse populations. Preprint at medRxiv https://doi.org/10.1101/2021.09.03.21262975 (2021).
Ganesan, M. et al. c.*84 G > A mutation in CETP is associated with coronary artery disease in South Indians. PLoS ONE 11, e0164151 (2016).
Niture, S., Moore, J. & Kumar, D. TNFAIP8: inflammation, immunity and human diseases. J. Cell. Immunol. 1, 29–34 (2019).
Huyghe, S., Mannaerts, G. P., Baes, M. & Van Veldhoven, P. P. Peroxisomal multifunctional protein-2: the enzyme, the patients and the knockout mouse model. Biochim. Biophys. Acta 1761, 973–994 (2006).
Nagai, A. et al. Overview of the BioBank Japan Project: study design and profile. J. Epidemiol. 27, S2–S8 (2017).
Hirata, M. et al. Overview of BioBank Japan follow-up data in 32 diseases. J. Epidemiol. 27, S22–S28 (2017).
Hirata, M. et al. Cross-sectional analysis of BioBank Japan clinical data: a large cohort of 200,000 patients with 47 common diseases. J. Epidemiol. 27, S9–S21 (2017).
Kanai, M. et al. Genetic analysis of quantitative traits in the Japanese population links cell types to complex human diseases. Nat. Genet. 50, 390–400 (2018).
Akiyama, M. et al. Characterizing rare and low-frequency height-associated variants in the Japanese population. Nat. Commun. 10, 4393 (2019).
Delaneau, O., Zagury, J.-F., Robinson, M. R., Marchini, J. L. & Dermitzakis, E. T. Accurate, scalable and integrative haplotype estimation. Nat. Commun. 10, 5436 (2019).
Fuchsberger, C., Abecasis, G. R. & Hinds, D. A. minimac2: Faster genotype imputation. Bioinformatics 31, 782–784 (2015).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Takeuchi, K. et al. Study profile of the Japan Multi-institutional Collaborative Cohort (J-MICC) study. J. Epidemiol. 31, JE20200147 (2021).
Hamajima, N. et al. Gene–environment interactions and polymorphism studies of cancer risk in the Hospital-based Epidemiologic Research Program at Aichi Cancer Center II (HERPACC-II). Asian Pac. J. Cancer Prev. 2, 99–107 (2001).
Koyanagi, Y. N. et al. Development of a prediction model and estimation of cumulative risk for upper aerodigestive tract cancer on the basis of the aldehyde dehydrogenase 2 genotype and alcohol consumption in a Japanese population. Eur. J. Cancer Prev. 26, 38–47 (2017).
Tsugane, S. & Sawada, N. The JPHC study: design and some findings on the typical Japanese diet. Jpn. J. Clin. Oncol. 44, 777–82 (2014).
The “All of Us” Research Program. N. Engl. J. Med. 381, 668–676 (2019).
Shilo, S. et al. 10 K:Aa large-scale prospective longitudinal study in Israel. Eur. J. Epidemiol. 36, 1187–1194 (2021).
Eastwood, S. V. et al. Algorithms for the capture and adjudication of prevalent and incident diabetes in UK Biobank. PLoS ONE 11, e0162388 (2016).
Woodfield, R., UK Biobank Stroke Outcomes Group, UK Biobank Follow-up and Outcomes Working Group & Sudlow, C. L. M. Accuracy of patient self-report of stroke: a systematic review from the UK Biobank Stroke Outcomes Group. PLoS ONE 10, e0137538 (2015).
Westerman, K. E. et al. Genome-wide gene–diet interaction analysis in the UK Biobank identifies novel effects on hemoglobin A1c. Hum. Mol. Genet. 30, 1773–1783 (2021).
Yamamoto, K. et al. Genetic footprints of assortative mating in the Japanese population. Nat. Hum. Behav. 7, 65–73 (2022).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573 (2010).
Ritchie, S. C. et al. Quality control and removal of technical variation of NMR metabolic biomarker data in ~120,000 UK Biobank participants. Sci. Data 10, 64 (2023).
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336 (2010).
Acknowledgements
We gratefully acknowledge the participants and investigators of the BBJ, UKB, J-MICC, HERPACC, JPHC, the National Institutes of Health’s All of Us Research Program and HPP. Part of the super-computing resource was provided by Human Genome Center, the Institute of Medical Science, The University of Tokyo. The J-MICC Study was supported by Grants-in-Aid for Scientific Research for Priority Areas of Cancer (grant 17015018) and Innovative Areas (grant 221S0001), and by the Japan Society for the Promotion of Science (JSPS) KAKENHI grant (grants 16H06277 and 22H04923 (CoBiA)) from the Japanese Ministry of Education, Culture, Sports, Science and Technology. This work was also supported in part by funding for the BBJ from the Japan Agency for Medical Research and Development (from April 2015 until now), as well as the Ministry of Education, Culture, Sports, Science and Technology (from April 2003 to March 2015). The HERPACC Study was supported by a Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan Priority Areas of Cancer (grant 17015018), Innovative Areas (grant 221S0001) and the JSPS KAKENHI Grants (grants JP16H06277 and 22H04923 (CoBiA), JP26253041,JP20K10463, JP23K16316 and 24K02697) and a Grant-in-Aid for the Third Term Comprehensive ten-year Strategy for Cancer Control from the Ministry of Health, Labour and Welfare of Japan. The JPHC Study was supported by the National Cancer Center Research and Development Fund (grants 23-A-31 (toku), 26-A-2, 29-A-4, 2020-J-4 and 2023-J-4; from 2011 until now), and a Grant-in-Aid for Cancer Research from the Ministry of Health, Labour and Welfare of Japan (from 1989 to 2010). S.Namba was supported by AMED (grants JP24tm0424228, JP24tm0524009, JP25kk0305032 and JP256f0137004), the Takeda Science Foundation and the Japan Foundation for Applied Enzymology. Y.Okada was supported by JSPS KAKENHI (grant 25H01057); AMED (grants JP24km0405217, JP24ek0109594, JP24ek0410113, JP24kk0305022, JP223fa627001, JP223fa627002, JP223fa627010, JP24zf0127008, JP24tm0524002, JP24wm0625504 and JP24gm1810011); JST Moonshot R&D (grants JPMJMS2021 and JPMJMS2024); Takeda Science Foundation; Ono Pharmaceutical Foundation for Oncology, Immunology and Neurology; Bioinformatics Initiative of Graduate School of Medicine, Institute for Open and Transdisciplinary Research Initiatives, Center for Infectious Disease Education and Research (CiDER) and Center for Advanced Modality and DDS (CAMaD) at The University of Osaka; and the RIKEN TRIP initiative (AGIS).
Author information
Authors and Affiliations
Consortia
Contributions
S. Namba and Y. Okada conceptualized the work, administered the project and acquired funding. S. Namba designed the methodology, performed data validation and visualizations. K. Sonehara, Y.N.K. and T. Yamaji curated the data. T. Kikuchi and S. Namba performed the formal analysis. T.O., R.E., G.S., Y.T, H.U., K.Y., Y. Ogawa, S. Namba, K. Sonehara and K. Suzuki conducted the investigations. A.K., S.H., S.K., H.Y., Y.N., Y. Okazaki, N.M., K. Motomura, H.K., A.H., H.I., M.H., M.N., I.O., S. Nakano, the BBJ, Y. Oda, Y.S., Y. Okada, M.I., N.S., K. Matsuo, T. Yamauchi, T. Kadowaki, K. Matsuda, Y.N.K., T. Yamaji and T.M. provided resources. M.I., N.S., K. Matsuo, T. Yamauchi, T. Kadowaki, Y. Okada and K. Matsuda supervised the project. S. Namba wrote the original draft; both S. Namba and Y. Okada reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Schematic overview of the G×E interaction analysis workflow.
a, Environment sets used to test G×E interactions. P-values were calculated for individual environmental sets and integrated on the Cauchy distribution to obtain final P-values. b, Workflow of G×E interaction tests. c, Workflow of defining the environments contributing to G×E interactions and their orders.
Extended Data Fig. 2 Consensus clustering of environmental questionnaires.
a, Consensus clustering of questionnaires about dietary consumption and physical activity in UKB1. b, Delta area plot for area under the cumulative distribution function (CDF) curve when changing the number of clusters from two to ten. c and d, Same as a and b, respectively, for BBJ1.
Extended Data Fig. 3 LocusZoom plots of inter-categorical pleiotropic loci in UKB1.
a–i, LocusZoom plots of the significant G×E interactions at the ALPL (a and b), HK1 (c–f), and APOE (g–j) loci. k–m, Hg19 coordinates of protein-coding genes in individual loci. Recombination rates were calculated using the European in-sample linkage disequilibrium information from UKB1. P-values were estimated by two-sided linear regression.
Extended Data Fig. 4 Consensus clustering of dietary questionnaires in independent replication cohorts.
Consensus clustering of food frequency questionnaires in J-MICC/HERPACC (a), JPHC (b), and HPP (c). Delta area plots show the area under the cumulative distribution function (CDF) curve when changing the number of clusters from two to ten.
Extended Data Fig. 5 Replication within and across populations.
a–b, Replication statuses of G×E loci other than the ALDH2 locus (a) and the ALDH2 locus (b). Only interactions with at least one nominally significant replication (PG×E < 0.05) are shown. EUR, the European population; EAS, the East Asian population; AFR, the African population; AMR, the American population. c–f, The eGFR distributions across rs77924615 genotypes (the UMOD locus) and age in UKB1 (c), UKB2 (d), BBJ1 (e), and BBJ2 (f), as a representative example of cross-population shared G×E interaction. Dots, 5,000 randomly sampled individuals per genotype; lines, regression lines using all participants. g and h, Minor allele frequencies (MAF) of the lead variants (g) and environmental distributions (h) in UKB1 and BBJ1. In g, cross-population shared loci are marked with crosses and labelled by nearest genes. Solid gray lines represent x = 0, y = 0, and y = x. i, Number of G×E loci detected with PG×E less than 5.0×10−8 in randomly subsampled participants in UKB1 and BBJ1. Subsample sizes: 10, 50, 100, 150, and 200 thousand in UKB1; 10, 50, and 80 thousand in BBJ1. Full-cohort results are also shown (Nmean = 253,773 in UKB1 and 133,117 in BBJ1).
Extended Data Fig. 6 Urate measurements stratified by rs4148155 and questionnaires about “meat and cheese” consumption.
Distributions of urate measurements across the rs4148155 genotypes (the ABCG2 locus) and the questionnaires related to the “meat and cheese” consumption in UKB1. Dots represent 5,000 randomly sampled individuals per genotype; lines represent regression lines using all participants.
Extended Data Fig. 7 Environmental contributions at the PITX2 locus.
Log-likelihood improvement for G×E models of rs72900155 (the PITX2 locus) for arrhythmia in BBJ1. Environmental factors were added stepwise to the null model (Methods).
Extended Data Fig. 8 Phenome-wide G×E interactions for disease statuses.
P-values of G×E interactions for disease statuses in UKB1 (a) and BBJ1 (b), estimated by two-sided Firth logistic regression. Top G×E-contributing environments are shown for interactions with FDR less than 0.05. Horizontal line represents FDR = 0.05 in each cohort. COPD, chronic obstructive pulmonary disease; AKD, acute kidney disease; CKD, chronic kidney disease.
Extended Data Fig. 9 Additional genome-wide properties of G×E interactions.
a, PGS prediction accuracy in the same population. X-axis: log2 ratio of prediction accuracy (R2) between “same-stratum” and “opposite-stratum” PGS (based on matching vs. non-matching environmental strata). Y-axis: P-values of R2 differences, estimated by two-sided Hotelling’s t test. Dashed line: FDR threshold of 0.05. b, Same as a, for cross-population portability. c and d, Examples of PGS prediction accuracy within (c) and across (d) populations, for hematocrit (Ht) stratified by sex. e, MAGMA one-sided gene-set enrichment analyses of age-stratified genetic effects on pulse pressure (PP), using the gene ontologies for biological processes. Dashed lines: FDR = 0.05 (vertical/horizontal); y = x (diagonal). f and g, Single-cell associations in the monkey artery scRNA-seq data for the younger (f) and older (g) groups. Significant cell types marked with ellipses (false discovery rate <0.05; one-sided Monte Carlo sampling implemented in scDRS). h, Genetic correlations across PP, systolic blood pressure (SBP), and diastolic blood pressure (DBP), estimated separately for the younger and older groups. i, Genetic correlations for the same trait between age groups. For h and i, shown are the inverse-variance weighted meta-analyses of UKB1 and BBJ1 (N = 127,933, 128,369, and 128,530 in UKB1 and N = 73,480, 73,815, and 73,750 in BBJ1 for PP, SBP, and DBP, respectively, for each age stratum); data are presented as estimated values with 95% CI.
Extended Data Fig. 10 Metabolome-wide G×E interactions at lead variants of clinical G×E loci.
P-values of marginal effects and G×E interactions for metabolites in all 15 loci with genome-wide significant G×E interactions, estimated by two-sided linear regression and categorized into five patterns. Triangles denote P-values less than 1.0 × 10−50.
Extended Data Fig. 11 Sexual dimorphic effects at the CETP locus on clinical lipid biomarkers.
a, Effect sizes of rs12720908 (the CETP locus) on clinical lipid biomarkers in UKB1, stratified by sex and age. b and c, Hazard ratios of clinical lipid biomarkers for all-cause (b) and CAD-caused mortality (c) in UKB1, stratified by sex and statin medication at baseline. Data are presented as estimated values with 95% CI; see Supplementary Tables 31, 32 for the sample sizes.
Extended Data Fig. 12 Sexual dimorphic effects for lipid metabolites at multiple loci.
a, Effect sizes of rs1065853 (the APOE locus), rs6065904 (the PLTP locus), and rs58542926 (the TM6SF2 locus) on lipid metabolites in UKB1, stratified by sex and age. L_LDL_PL_pct, phospholipids to total lipids in large LDL percentage; L_VLDL_FC_pct, free cholesterol to total lipids in large VLDL percentage; LDL_PL_pct, phospholipids to total lipids in LDL percentage; S_HDL_FC_pct, free cholesterol to total lipids in small HDL percentage; S_HDL_PL_pct, phospholipids to total lipids in small HDL percentage. b, Heatmap of marginal effects and G×E interactions of rs67328001 (the TNFAIP8 locus) on lipid metabolites, using P-values estimated by two-sided linear regression. The metabolite with the smallest Pmarginal (“triglycerides in small VLDL” [S_VLDL_TG]) and the one with the smallest PG×E (“concentration of very large HDL particles” [XL_HDL_P]) are shown in red. c and d, Effect sizes of rs67328001 (the TNFAIP8 locus) on the five metabolites with the smallest Pmarginal (c) and PG×E (d), stratified by sex and age. M_LDL_TG, triglycerides in medium LDL; HDL_TG_pct, triglycerides to total lipids in HDL percentage; S_LDL_TG, triglycerides in small LDL; XS_VLDL_TG, triglycerides in very small VLDL; XL_HDL_FC, free cholesterol in very large HDL; XL_HDL_L, total lipids in very large HDL; XL_HDL_C, cholesterol in very large HDL; HDL_size, average diameter for HDL particles. For a,c, and d, data are presented as estimated values with 95% CI; see Supplementary Table 31 for the sample sizes.
Extended Data Fig. 13 Genome-wide G×E interactions for metabolites.
a, Sexual dimorphic effects at four loci in UKB1. XXL_VLDL_TG_pct, the triglycerides to total lipids ratio in chylomicrons and extremely large VLDL; S_VLDL_CE_pct, the cholesteryl esters to total lipids ratio in small VLDL. b and c, Environment-specific effects in UKB1 for current-smoking at the LRRC4C locus (b) and never-drinking at the KLHL32 locus (c). L_VLDL_PL_pct, the phospholipids to total lipids ratio in large VLDL; S_VLDL_PL, phospholipids in small VLDL. d, Distribution of Pmarginal and PG×E at the SLC7A2 locus in UKB1, estimated by two-sided linear regression. e, Environment-stratified effect sizes at the SLC7A2 locus in UKB1. For the “tea and coffee” and “meat and cheese” environment clusters, participants were stratified into two equal-sized groups. Note that coffee consumption was multiplied by −1 to account for its negative correlation with tea consumption in UKB1; therefore, higher cluster values indicate more frequent tea consumption, whereas lower values indicate more frequent coffee consumption. For a–c and e, data are presented as estimated values with 95% CI; see Supplementary Table 34 for the sample sizes.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–11, Supplementary Figs. 1–3, Supplementary Methods and Supplementary References.
Supplementary Tables (download XLSX )
Supplementary Tables 1–34
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Namba, S., Sonehara, K., Koyanagi, Y.N. et al. A cross-population compendium of gene–environment interactions. Nature 651, 688–697 (2026). https://doi.org/10.1038/s41586-025-10054-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-10054-6