Abstract
Systematic characterization of the molecular states of cells in livestock tissues is essential for understanding the cellular and genetic mechanisms underlying economically and ecologically important physiological traits. Here, as part of the Farm Animal Genotype-Tissue Expression (FarmGTEx) project, we describe a comprehensive reference map including 1,793,854 cells from 59 bovine tissues in calves and adult cattle, spanning both sexes, which reveals intra-tissue and inter-tissue cellular heterogeneity in gene expression, transcription factor regulation and intercellular communication. Integrative analysis with genetic variants that underpin bovine monogenic and complex traits uncovers cell types of relevance, such as spermatocytes, responsible for sperm motility and excitatory neurons for milk fat yield. Comparative analysis reveals similarities in gene expression between cattle and humans, allowing for the detection of relevant cell types to study human complex phenotypes. This Cattle Cell Atlas will serve as a key resource for cattle genetics and genomics, selective breeding and comparative biology.
Similar content being viewed by others
Main
Cattle, domesticated over 10,000 years ago1, have a crucial role in converting indigestible fiber feed into protein-rich food for humans, such as beef and milk, which are essential for human nutrition and health. To meet increasing global demand for safe animal food products while minimizing the production-associated negative impacts on animal welfare and the environment (for example, greenhouse gas emissions and zoonotic diseases)2, it is essential to understand the genetic and molecular mechanisms underlying various phenotypes of economic and ecological importance in cattle.
As of 23 December 2024, 192,336 genomic loci have been associated with around 500 different complex traits in cattle3. A large proportion of these variants reside in noncoding genomic regions, have small individual effects on phenotypic variation and influence complex traits via modulation of gene regulation. In this regard, many studies have explored the molecular mechanisms underlying complex traits at the tissue expression level by integrating genome-wide association study (GWAS) datasets and molecular quantitative trait loci (molQTLs). For instance, the Cattle Genotype-Tissue Expression (CattleGTEx) project4 linked gene expression in over 20 tissues with 43 significant economically important traits and provides valuable insights into their gene regulatory mechanisms. However, tissues are generally heterogeneous mixtures of distinct cell types and states. Because of the rapid development of single-cell sequencing technology, single-cell transcriptome atlases have been constructed for many organisms5,6. However, in cattle, previous studies on single-cell transcriptomic studies were limited to specific tissue types, including the rumen7,8,9, peripheral blood10,11, skeletal muscle12 and the digestive system13,14. Therefore, there is a need to comprehensively catalog different bovine cell types and states across several tissue types and biological contexts, which will substantially contribute to our understanding of the genetic and molecular architecture underlying many phenotypes in cattle.
In this study, we built a comprehensive Cattle Cell Atlas (CattleCA) for the livestock research community by generating and analyzing single-cell RNA sequencing (scRNA-seq) and single-nucleus RNA-seq (snRNA-seq) data from 1,793,854 cells across 59 tissue types in 15 animals, spanning both sexes (Fig. 1). We characterized 131 distinct cell types and assessed cellular heterogeneity in terms of gene expression, transcription factor (TF) regulation, and intra-tissue and inter-tissue cellular communication. Leveraging the CattleCA, we highlight specific cell types and states associated with monogenic conditions and complex traits. Furthermore, we explored the evolutionary conservation of the transcriptome between cattle and humans at the single-cell level, revealing shared cellular mechanisms underlying complex traits and diseases in humans. Thus, the CattleCA (https://ngdc.cncb.ac.cn/cattleca/) is a valuable resource for cattle genetics and genomics, immunology, precision breeding and comparative biology.
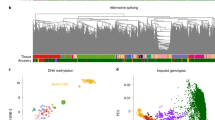
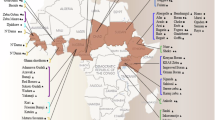
a, Overview of the CattleCA data, analysis and its application. b, Schematic diagram of 59 cattle tissues collected in this study. The numbers of samples and cells/nuclei per tissue are shown in parentheses. c, Uniform manifold approximation and projection (UMAP) visualization of CattleCA colored according to the different cell types. All cell types are categorized into seven cell lineages; cell type annotation and the corresponding cell number are provided in the legend on the right. The figure was created in BioRender.
Results
The scope of CattleCA
After removing low-quality cells and doublets (Methods, Supplementary Fig. 1 and Supplementary Tables 1 and 2), 1,506,438 single cells and 287,416 single nuclei (hereinafter referred to as cells) from one fetus, four calves and ten adults, including 234,802 from males and 1,559,052 from females, were used for the subsequent analyses (Supplementary Fig. 2 and Supplementary Table 3). The average numbers of expressed genes and read counts were 1,783 and 5,752, respectively, across 59 tissues (Supplementary Fig. 3). By integrating all high-quality cells in tissues using Harmony15 (Supplementary Figs. 4, 5 and 6c,d), we annotated 131 cell types based on canonical marker genes, representing seven distinct cell lineages: immune (n = 679,021), endothelial (n = 308,878), epithelial (n = 268,626), stromal (n = 240,771), nerve (n = 182,450), muscle (n = 81,438) and germline (n = 4,404) cells (Fig. 1c, Supplementary Fig. 6a and Supplementary Tables 4 and 5). These cells clustered well based on cell lineage types rather than processing methods, sequencing platforms and tissue types (Fig. 1c, and Supplementary Fig. 6b–d), with cell type abundance ranging from 33 for circulating epithelial cells to 233,497 for blood vascular endothelial cells (Fig. 1c). On average, 12 distinct cell types were identified per tissue, ranging from seven in the oviduct to 21 in the ileum (Supplementary Figs. 7 and 8, and Supplementary Table 6). Among the 131 annotated cell types, 67 were observed only in one tissue type, whereas immune, endothelial and epithelial cells were found across 58, 49 and 41 tissues, respectively (Supplementary Figs. 7 and 8, and Supplementary Table 6). For instance, blood vascular endothelial cells were ubiquitously present in 45 tissues, whereas alpha and beta cells were exclusively detected in the pancreas (Supplementary Fig. 9). Furthermore, cell cycle analysis indicated that cells from the germline lineage were predominantly enriched in the G2/M phase, reflecting their active growth and preparation for DNA replication. Conversely, epithelial, muscle, stromal and endothelial cells were primarily enriched in the G0/G1 phase, indicating a quiescent state with no active division (Supplementary Fig. 6e and Supplementary Table 7). We offer a comprehensive resource of TF and cellular communication regulation across several tissues, cell types and sexes in cattle (Extended Data Figs. 1 and 2, Supplementary Fig. 10 and Supplementary Note).
Cellular heterogeneity in mammary glands and testis
To study cellular heterogeneity in tissues, we took mammary glands as an example because of their importance in milk production. We categorized a total of 2,670 mammary epithelial cells into eight distinct clusters, including six luminal secretory cell subtypes (LumSec, ME0–5), characterized by high expression of ELF5 and LTF; a luminal hormone-responsive cell cluster (LumHR, ME6), marked by PGR and ESR1 expression; and a basal-myoepithelial cell cluster (ME7), defined by MYLK and KRT17 expression (Fig. 2a,b). These cell clusters exhibited distinct spatial distribution patterns and functional characteristics in mammary glands (Fig. 2b,c). Among luminal secretory cells, ME0–3 were strongly associated with milk biosynthesis, as indicated by high lactation scores16 (Fig. 2d and Supplementary Table 8). Genes upregulated in ME0, including HSPH1, HSPA1A and JUN, were significantly enriched in the regulation of ATP-dependent activity, heat response and DNA-templated transcription (Fig. 2b). Moreover, both ME0 and ME2 were enriched for immune-related genes (for example, PIGR, LPO and SPP1), suggesting their potential involvement in immune regulation under the stress associated with lactation17 (Fig. 2e). ME1 and ME6 were significantly associated with milk (MY), fat (FY) and protein (PY) yield, based on candidate genes identified through GWAS in 16,188 Holstein cows using scPagwas18 (Fig. 2f). LumHR cells exhibited high expression of genes encoding hormone receptors for estrogen, progesterone, prolactin and growth factors (ESR1, PGR, PRLR and GHR), suggesting their potential roles in hormonal and growth factor signaling. As a hormone-responsive epithelial cell type, ME6 might promote cell adhesion and signal transmission, ultimately supporting the overall function of the mammary gland during lactation19 (Fig. 2g–i). Furthermore, ME7 was identified as a potential responder to oxytocin, with high expression of genes involved in the oxytocin signaling pathway, including OXTR and PLCB1, coordinating contractile processes within the mammary gland for milk ejection20 (Fig. 2j). These findings provide insights into the cellular complexity of the mammary gland during lactation. Additionally, we highlight bovine spermatogenesis and the evolutionary conservation of key TFs of spermatogenesis in mammals (Supplementary Fig. 11, Extended Data Fig. 3 and Supplementary Note).
a, UMAP plot displaying the subtypes of mammary epithelial cells. b, Expression heatmap for the top 120 DEGs in eight mammary epithelial subtypes, with cell count (top), selected marker genes (left) and significantly enriched Gene Ontology (GO) terms for the clustered module (right). c, Heatmap showing the odds ratios (ORs) of mammary epithelial cell subtypes in each anatomical structure of the mammary gland. d, Box plot showing the lactation gene signature scores for the epithelial cell subtypes in the mammary gland (n = 4). The y axis represents the module score derived from genes associated with lactation-related processes; the x axis represents cell subtypes. The central band in the box plot represents the median, the box boundaries represent the 25% to 75% percentiles, and the whiskers extend 1.5 times the interquartile range (IQR). The red dashed line indicates the baseline lactation score (0). e, Expression levels of immune genes in different mammary epithelial cell subtypes. Dot size represents the expressed percentage; the color represents the average expression of genes in each subtype. f, Heatmap showing the mammary epithelial cell subtypes significantly associated with MY, FY and PY. *FDR < 0.05, **FDR < 0.01. g, Outgoing signaling pathways in mammary epithelial cell subtypes. Dot size represents the percentage contribution. h, Specific outgoing communication signaling patterns in ME6. Dot size represents the percentage contribution. i, Specific ligand–receptor pairs in ME6-specific outgoing communication pathways. Dot size represents the P significance level; dot color represents the maximum communication probability. j, Expression levels of genes in the oxytocin signaling pathway in different mammary epithelial cell subtypes. Dot size represents the expressed percentage; dot color represents the average expression of genes in each subtype.
Antigen-presenting immune cell heterogeneity
A total of 777,873 immune cells from 58 tissues (excluding the retina) were identified and annotated into 29 cell types (Fig. 3a). These immune cells were clustered according to five cell lineages (T and natural killer (NK) cells, myeloid cells, mast cells, plasma cells and neutrophils) rather than tissue types, suggesting that cells in the same lineage exhibit high transcriptional similarity (Fig. 3b). However, some immune cell types also displayed strong tissue specificity. For example, microglia in the brain formed a distinct transcriptional cluster separating from the broader myeloid lineage (Fig. 3b).
a, Bar chart displaying the number and proportion of 29 immune cell types in 58 tissues. The scatter plot represents the Shannon entropy values of immune cells in each tissue, with higher values indicating higher diversity. b, Heatmap showing the Pearson correlation of gene expression levels for each immune cell type, calculated based on the top 1,000 genes with the largest s.d. c, t-distributed stochastic neighbor embedding (t-SNE) visualization of annotated cell subtypes of myeloid cells, with cells color-coded according to subtype. d, Heatmap highlighting the average regulatory activity z-score of key TFs in 247 TFs that regulate the differentiation and maintenance of APC subtypes. e, Pseudotime trajectory analysis of monocyte and macrophage subtypes, with cell states colored on the trajectory tree. f, Statistical analysis of monocyte and macrophage subtype composition in each state. g, Heatmap showing the changes in gene expression of representative DEGs (q < 1 × 10−4) in pseudotime series, and GO enrichment analysis of DEGs reclustered into four clusters (bottom left). h, Representative pseudotime trajectory of marker genes in microglia across different states.
As antigen-presenting cells (APCs) are central to adaptive immunity by presenting antigens to lymphocytes, we focused on 169,610 myeloid cells across 44 tissues and annotated them into six macrophage subtypes (MA0–5), three monocyte subtypes (MO0–2) and five dendritic cell (DC) subtypes (DC0–4) (Fig. 3c and Extended Data Fig. 4a). A total of 135,186 macrophages were identified in 40 tissues, making them an appropriate model for exploring cellular heterogeneity across tissue microenvironments. Macrophage subtypes exhibit distinct patterns of tissue distribution, TF regulation and metabolic characteristics (Extended Data Figs. 4 and 5). IRF7 (ref. 21) and IRF8 (ref. 22) might regulate MA1, which was preferentially distributed in the intestine and might participate in sugar metabolism and fatty acid elongation, suggesting the strongest major histocompatibility complex class II antigen-presenting score (APS) among all the macrophage subtypes (Fig. 3d, Extended Data Figs. 4b–d and 5a–c and Supplementary Table 9). In contrast, MA4 in the liver showed the weakest APS (Extended Data Fig. 5c). These findings were consistent with the notion that the intestine has a high immune vigilance because of constant exposure to microbial antigens23, while the liver, a site of immune tolerance, exhibits a lower antigen-presenting activity to prevent excessive immune activation24. The pseudotime trajectory analysis revealed a separation of macrophage populations that aligns with known developmental stages in humans and mice25: one branch progresses toward an inflammatory phenotype, which is consistent with monocyte differentiation, while the other maintains markers associated with tissue-resident macrophages, which might be derived from embryonic progenitors (Fig. 3e–h). Additionally, TF analysis revealed that distinct TFs associated with monocyte recruitment and differentiation, such as SPI1 (ref. 26) and IRF7 (ref. 21), were strongly expressed in the monocyte-derived branch, whereas BHLHE41 (ref. 27) and IRF8 (ref. 22), which are responsible for tissue-resident macrophages, were more active in the other branch (Supplementary Fig. 12). We also provide insights into the heterogeneity and dynamics of B cell differentiation (Extended Data Fig. 6 and Supplementary Note).
Epithelial cells interact with immune cells in the intestine
We analyzed 278,584 epithelial cells across 41 tissues, identifying 50 subtypes (Supplementary Fig. 13a and Supplementary Table 10). Although most subtypes were tissue-specific, keratinocytes (36,114), spinous cells (22,672), basal cells (19,283) and goblet cells (GCs) (13,055) exhibited the highest cell counts and were identified in more than six tissues (Supplementary Fig. 13a). Cell types with similar biological functions were clustered together, such as chief cells, parietal cells, isthmus cells, mucous neck cells and pit cells, all of which have the function of secreting gastric protease and promoting digestion (Supplementary Fig. 13b). Additionally, three regions of the forestomach (rumen, reticulum and omasum) and six regions of the intestine (duodenum, ileum, jejunum, colon, cecum and rectum) were also clustered together (Supplementary Fig. 13c). Because of the importance of epithelial cells to the digestive system, we scrutinized 152,009 epithelial cells across 14 digestive tissues, uncovering eight cell-to-tissue modules with distinct biological functions and TF regulation (Fig. 4a). We further analyzed the cellular heterogeneity of the bovine forestomach and abomasum, provided evidence supporting the multi-origin hypothesis that the forestomach originates from the esophagus while the abomasum originates from the duodenum, and emphasized the critical role of rumen spinous cells in fatty acid absorption (Extended Data Figs. 7 and 8 and Supplementary Note).
a, Expression heatmap for the top 50 upregulated DEGs in 24 epithelial cells of 15 selected gastrointestinal tissues, showing significant Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways for the clustering module (left), specific TFs (middle) and corresponding tissue anatomical structures and cellular composition (right). b, Volcano plot illustrating the DEGs between paratuberculosis-seropositive and healthy cattle. DEGs were identified based on FDR < 0.05 and log2(fold change) > 1 using edgeR’s two-sided exactTest. c, Box plot showing the module scores of 17 disease-associated and 56 health-associated upregulated genes in epithelial cells of the intestine (n = 5), calculated using the AddModuleScore of Seurat. The central band in the box plot represents the median, the box boundaries represent the 25% to 75% percentiles, and the whiskers extend 1.5 times the IQR. A two-sided Wilcoxon rank-sum test was used. d, Bubble plot showing ligand–receptor pairs between GCs and immune cells, which were shared in at least four intestinal segments. e, t-SNE visualization of seven GC subtypes. Cells are color-coded according to cell subtype. f, Heatmap showing the top 20 upregulated DEGs and their markers for each GC subtype. The bar chart displays their GO terms and KEGG pathways using single-sample gene set enrichment analysis (GSEA). g, Heatmap representing the tissue distribution of GC subtypes; ORs were calculated and used to indicate preferences. **OR > 1.5 and FDR < 0.01. h, Heatmap displaying the signature scores of tuberculosis and paratuberculosis candidate genes from the Animal Quantitative Trait Loci (QTL) and KEGG databases across GC subtypes. i, Scatter plot highlighting tissues and GC subtypes associated with human IBD and CD using GWAS enrichment analysis with LDSC. j, Violin plot representing the expression of key ligand–receptor genes in GC subtypes and immune cells.
Bovine paratuberculosis, a chronic and severe enteritis, causes notable economic losses to the cattle industry annually28. To study the complex dynamics between intestinal epithelial and immune cells during disease progression, we analyzed transcriptomic data from five paratuberculosis-seropositive cattle and three healthy controls29 (Fig. 4b) and found that 17 upregulated DEGs in the diseased group exhibited significantly higher signature scores in GCs, suggesting a key role of GCs in the progression of intestinal inflammation (Fig. 4c). We observed significant ligand–receptor pairs shared between GCs and immune cells across six different intestinal segments (Fig. 4d and Supplementary Table 11). For instance, GCs communicate internally through the GUCA2A–GUCY2C ligand–receptor pair and interact with immune cells through pathways such as APP–CD74 and MIF–CD74_CXCR4. Furthermore, we classified GCs into seven subtypes (GC0–6), each with distinct biological functions and tissue distribution patterns (Fig. 4e–g). For instance, GC1, primarily found in the ileum and rectum, might participate in tight junctions, while GC5, located in the jejunum, might be involved in immune responses. Additionally, we found that the marker genes of GC5 were significantly overrepresented in 529 genes associated with tuberculosis and paratuberculosis (Fig. 4h and Supplementary Table 12). Using one-to-one orthologous genes between cattle and humans, we observed that the heritability of human inflammatory bowel disease (IBD) and Crohn’s disease (CD) was significantly enriched in genes with specific expression in the jejunum and reticulum, particularly in GC5 (Fig. 4i). Cellular communication analysis showed that GC5 exhibited a dysregulation in certain communication pathways, such as the CDH, JAM, GRN, MIF and SEMA4 signaling pathway (Supplementary Fig. 14 and Supplementary Table 13). Dysregulation of adhesion molecules like CDH1 (ref. 30) and F11R31 in GC5 might contribute to epithelial shedding and increased intestinal permeability, while upregulation of antigen-presenting genes like CD74 might promote antigen recognition and presentation32, which might collectively influence the progression of intestinal inflammation (Fig. 4j). These findings provide supportive evidence for the potential communication between GCs and immune cells.
The potential role of hepatocytes in milk production
As the liver has a crucial role in energy metabolism during lactation in dairy cows33, we annotated a total of 18,205 hepatocytes from the liver and bile duct into ten subtypes (HPE0–9; Fig. 5a,b). These subtypes aligned well with known liver architecture in both cattle and other animals like humans and mice34. The observed spatial distribution of hepatocyte subtypes along the pericentral to the periportal axis was consistent with well-established liver zonation principles (Fig. 5c), where hepatocytes in different zones exhibit distinct metabolic functions (Fig. 5d). For example, enrichment of mammalian target of rapamycin signaling, ABC transporters and bile acid biosynthesis pathways in HPE0–3 corresponded to the known pericentral zone, which might be responsible for drug metabolism and bile acid synthesis. Similarly, HPE4–6 might be involved in fatty acid metabolism, glycolysis and gluconeogenesis, which was in line with the metabolic functions typically associated with the interzonal and periportal regions. Furthermore, enrichment of lactation-related genes in HPE4–7 suggested species-specific adaptations in dairy cattle and the liver’s important role in supporting high metabolic demands for milk production (Fig. 5e). In addition, HPE2 was significantly enriched in GO terms related to gastrulation, digestive system development and regionalization, suggesting its potential role in developmental stemness and as the main source of new hepatocytes in steady-state tissue renewal35 (Fig. 5f). We further observed the activation of key genes involved in lipid localization, lipid transport, protein hydrolysis regulation and immune response throughout hepatocyte development (Fig. 5g,h). Cattle hepatocytes showed pronounced enrichment in tryptophan metabolism, fatty acid metabolism and peroxisome proliferator-activated receptor (PPAR) signaling pathways, compared to those of humans, underscoring their specialized role in lactation (Fig. 5i,j).
a, UMAP plot of ten hepatocytes subtypes (HPE0–9), with cells color-coded according to subtype. b, Bubble plot displaying the standardized expression of the marker genes of each cell subtype. c, Pearson correlation of hepatocyte subtypes between cattle, humans and mice, calculated based on 92 spatial localization marker genes in human and mouse hepatocytes. A two-sided Wilcoxon rank-sum test was used. *Padj < 0.05, **Padj < 0.01, ***Padj < 0.001. d, Heatmap showing the GO terms and KEGG pathways of upregulated DEGs from each hepatocyte subtype using single-sample gene set variation analysis. e, UMAP plot displaying key cell types identified using GWAS enrichment analysis with scPagwas, integrating GWAS data with MY, FY and PY. The P value was calculated using a two-sided hypergeometric test, adjusted using multiple comparisons (FDR). f, GSEA of GO terms was performed using the fgsea package. Genes were ranked according to the log2(fold change) of DEGs in HPE2; a one-sided permutation test with 10,000 iterations was used to calculate the P values. g, UMAP plot depicting the developmental trajectory of hepatocyte subtypes over a pseudotime series. h, Heatmap showing the changes in gene expression of representative DEGs (q < 1 × 10−4) in the pseudotime series, and GO enrichment analysis of DEGs reclustered into two clusters (bottom left). i, Volcano plot illustrating the DEGs between cattle and human hepatocytes. j, GO terms of DEGs between cattle and human hepatocytes. The P value was calculated using a one-sided hypergeometric test and adjusted for multiple comparisons (FDR). ECM, extracellular matrix; NES, normalized enrichment score; TGFβ, transforming growth factor beta.
Cellular basis and mechanisms underlying monogenic disorders
To explore whether CattleCA could serve as a powerful resource for the dissection of the cellular basis and mechanisms underlying monogenic conditions in cattle, we compiled 183 causal genes associated with 145 bovine disorders from the Online Mendelian Inheritance in Animals database36 and divided them into ten trait domains based on their phenotypic manifestations (Supplementary Table 14). We detected 2,677 cell-type-specific genes (z-score > 0.75) across 129 distinct cell types (excluding PVALB+ GABAergic neurons and proliferative cells) (Fig. 6a). Our enrichment analysis revealed significant overlaps between cell-lineage-specific genes and the causal genes of disorders. For instance, muscle-cell-specific genes were significantly enriched with muscle disorder genes (false discovery rate (FDR) < 0.05), germline cells with reproduction disorders and epithelial cells with skin disorders (Fig. 6b).
a, Cell type correlation based on the expression levels of highly expressed genes (z-score > 0.75) specific to each of the 129 cell types using Euclidean distance. b, Correlations between seven cell lineages and ten disorder domains. Dot size represents log2(OR) values; the color represents the significant levels of a two-sided Fisher’s test. c, Expression level of nine epithelium-specific skin disorder genes in different cell lineages. The star above the violin represents a significant expression difference between this cell lineage and epithelial cells based on a two-sided Wilcoxon rank-sum test. The central band in the box plot represents the median, the box boundaries represent the 25% to 75% percentiles, and the whiskers extend 1.5 times the IQR. d, Expression levels of nine epithelium-specific skin disorder genes in different epithelial cell types. Dot size represents log10 counts per million (CPM) values and the color represents the z-score of genes in each cell type. e, Putative cellular communication between epithelial cells and other cell types implicating skin disorder genes. Cell types (inner color) in different tissues (outer color) are connected by putative interactions (dotted lines) between a ligand (left square) expressed in one epithelial cell and a receptor (right square) expressed in the other cell types. Bold italic indicates a skin-disorder-related gene. f, The expression distribution of LAMA3 in the placenta primarily occurs in UTCs; it is associated with the skin disorder EB. g, Reclustering and trajectory analysis of UTCs in the placenta. The distribution of LAMA3 expression (log10(CPM)) in different cell subtypes through the trajectory line is shown. h, Putative cellular communication between UTC subtypes and other cell types in the placenta. i, Expression levels of muscle disorder genes in different muscle cell types. j, Expression distribution of MYBPC1 in the esophagus. k, Reclustering and trajectory analysis of SMCs in the esophagus. The distribution of MYBPC1 expression (log10(CPM)) in different cell subtypes through the trajectory line is shown.
Nine of 27 genes related to skin disorders exhibited significantly higher expression in epithelial cells compared to other cell lineages (Fig. 6c,d and Extended Data Fig. 9a). To explore the cell–cell interaction of nine epithelium specifically expressed disease genes, we performed cellular communication analysis, identifying laminin, encoded by LAMA3 and LAMC2, as a key ligand in uninucleate trophoblast cells (UTCs) (found in the placenta) and progenitor cells (present in the jejunum and duodenum), respectively (Fig. 6e and Supplementary Fig. 15). LAMA3 showed high expression in placental UTCs, while LAMC2 exhibited elevated expression in intestinal progenitor cells (Fig. 6f and Extended Data Fig. 9b). LAMA3 and LAMC2 have crucial roles in regulating skin strength and resiliency37, with mutations disrupting laminin assembly and leading to epidermolysis bullosa (EB) in cattle38. A previous study also reported that laminin deficiency can affect trophoblast differentiation and embryonic development39. Trajectory analysis of UTCs revealed specific expression of LAMA3 toward the end of UTC differentiation (Fig. 6g and Extended Data Fig. 9c). Furthermore, we annotated UTCs into seven subtypes (UTC0–6; Extended Data Fig. 9d). Among them, LAMA3 was predominantly expressed in UTC2 and UTC3 (Fig. 6g and Extended Data Fig. 9c). Notably, UTC2 and UTC3, in which LAMA3 has an important role as a ligand-encoding gene, exhibited stronger cellular communication with other cell types in the placenta compared to other five subtypes, indicating a potential correlation between EB and UTC2/UTC3 (Fig. 6h, Supplementary Fig. 16 and Extended Data Fig. 9e). Furthermore, the marker genes of UTC2 and UTC3 were significantly enriched in several membrane pathways, including the basement membrane, the sarcolemma and the apical plasma membrane (Extended Data Fig. 9f). These pathways regulate cell–cell adhesion and affect the structure and stability of the basement membrane zone, which is the key pathogenic mechanism underlying EB40. In addition, LAMA3, positioned within these pathways, provides a compelling hypothesis that LAMA3, by participating in membrane function as a ligand-encoding gene in UTC2 and UTC3, may potentially contribute to EB in cattle (Supplementary Table 15). Furthermore, we observed the specific cellular gene expression patterns for muscle, reproduction and blood/immune disorders (Fig. 6i–k, Supplementary Fig. 17 and Supplementary Note).
Cellular basis and mechanisms underlying complex traits
To explore whether CattleCA could contribute to unraveling some of the cellular basis and mechanisms underlying complex traits in cattle, we collected GWAS summary statistics for 55 complex traits, representing milk production (n = 30), male fertility (n = 5), coat color (n = 6), IgG (n = 10), body conformation (n = 3) and health traits (n = 1). To prioritize the cell types involved in these complex traits, we conducted a trait–cell type enrichment analysis using scPagwas18 and revealed certain associations between cell lineages and complex traits. For instance, nerve cells were associated with milk production and germline cells with sperm traits (Fig. 7a and Supplementary Fig. 18).
a, Correlations between cell lineages and complex traits. The heatmap represents the significant level of each cell lineage with traits. Top, The gray squares represent trait groups. b,c, Correlations between milk production (b) and male fertility (c) traits and cell types. Each circle represents a cell type–trait association. The x axis represents cell lineages sorted alphabetically. d–f, Correlation between cell types and traits in tissues. d, Cerebral cortex. e, Esophagus. f, Testis. The violin plot presents the TRS for each cell type with the corresponding trait. The red line represents the significance level (−log10(P)) for each cell type with the corresponding trait under a two-sided hypergeometric test. g, Active KEGG pathways in the top significant trait-relevant cell types. Dot size represents the significance level (−log10(P)) of pathway activity; the dot color represents the significance level (FDR) after multiple comparisons of the enrichment between active pathway genes and top trait-relevant genes in the given cell type.
For milk production traits, the most significantly associated cell types were neurons. Excitatory neurons, specifically, exhibited significant associations with milk FY (P = 8.25 × 10−19) and capric acid (C10:0; P = 1.51 × 10−20), alongside amacrine cells displaying associations with lauric acid (C12:0; P = 4.3 × 10−16; Fig. 7b). This finding is consistent with our previous observation at the bulk tissue level in cattle, where a strong association between neurobiology and milk production traits was noted41, and aligns with previous research in humans suggesting reciprocal regulation between neuronal activity and lipid metabolism42. Additionally, we observed that excitatory neurons in the cerebral cortex and amacrine cells in the retina exhibited significant associations and high trait-relevant scores (TRS) for FY (P = 8.87 × 10−6), C10:0 (P = 8.75 × 10−6) and C12:0 (P = 1.59 × 10−2) compared to other cell types, underscoring the important roles of these two tissues in regulating milk fat content (Fig. 7d and Supplementary Fig. 19a–d). Furthermore, skeletal muscle cells (SMCs) showed a significant association with C12:0 (P = 9.14 × 10−20; Fig. 7b), which is consistent with their pivotal role in fatty acid use and intracellular fatty acid homeostasis43. Further analysis revealed that SMCs in the esophagus (P = 6.89 × 10−13) and tongue (P = 1.03 × 10−12) showed significant associations with C12:0, indicating a prominent role for these two tissues in fatty acid regulation (Fig. 7e and Supplementary Fig. 19c). According to the pathway activity analysis conducted with scPagwas, neuronal-excitement-related and lipolysis-related pathways, like thermogenesis (P = 1.15 × 10−198), regulation of lipolysis in adipocytes (P = 2.07 × 10−207) and glutamatergic synapse (P = 2×10−197) had significant activity in excitatory neurons, amacrine cells and SMCs, respectively; the genes in these pathways had significant enrichment with fatty acid trait-relevant genes (Fig. 7g and Supplementary Fig. 19f). In addition, we observed that luminal cells, closely related to lactation and milk production traits16, exhibited a particularly significant association with the production of pentadecanoic acid (C15:0) (Fig. 7b), along with active involvement in pathways regulating lipolysis in adipocytes (P = 2.11 × 10−218), PI3K/Akt (P = 1.73 × 10−292) and retrograde endocannabinoid signaling pathways (P = 3.2 × 10−286) (Fig. 7g).
For male fertility traits, the most significant association was observed between sperm motility (SMOT) and spermatocytes across 129 cell types (P = 1.11 × 10−7; Fig. 7c). Spermatocytes undergo a complex process of differentiation after meiosis, ultimately maturing into spermatids. This is a crucial step that directly affects the quality of sperm produced44. This finding was further reinforced by the significant association observed between spermatocytes in the testis and SMOT (P = 2.83 × 10−10) within the testis tissue level, with the function of active pathways in spermatocytes primarily focused on nutrient metabolism (Fig. 7f,g). These pathways also had significant enrichment with SMOT-relevant genes of spermatocytes, indicating their potential association with sperm production and maturation (Fig. 7g). Additionally, plasmacytoid DCs derived from peripheral blood mononuclear cells (PBMCs) were significantly associated with semen concentration per ejaculate trait (P = 2.06 × 10−5 in the global atlas; P = 7.31 × 10−4 in PBMCs; Fig. 7c and Supplementary Fig. 19e), which is consistent with our previous findings regarding the involvement of immune cells in male fertility traits41. This aligns with reports in humans indicating a significant association between DC abundance and sperm quality, suggesting a potential contribution of DC-mediated immune responses suboptimal to male fertility or infertility45. Pathway analysis revealed significant activity in distal convoluted tubule cells predominantly involving the regulation of the hypothalamic gonadotropin-releasing hormone (GnRH) signaling pathway (P = 8.83 × 10−250; Fig. 7g). Furthermore, signal transduction pathways, like the dopaminergic synapse, had significant activity in cone photoreceptor cells (P = 5.59 × 10−265; Fig. 7g). Additionally, the cell types most associated with coat color were type B intercalated cells, luminal cells and amacrine cells; for IgG levels, they were type B intercalated cells, CD8+ T cells and SMCs; and for body and health, they were chondrocytes, epithelial stem cells and amacrine cells (Extended Data Fig. 10). In summary, the association of specific cell types with complex traits provides a cellular perspective for pinpointing the genetic regulatory mechanisms underlying important cattle traits.
Comparison of cross-species cell transcriptome similarity
We analyzed the public single-cell transcriptome datasets from 30 human tissues (Supplementary Table 16) and found considerable conservation at cellular expression, TF regulation and communication between cattle and humans, offering insights into cross-species similarities (Fig. 8a–g, Supplementary Figs. 20–24 and Supplementary Note).
a, MetaNeighbor area under the receiver operating characteristic (AUROC) values calculated using shared genes of the same cell types in cattle and humans. b,c, Regulon specificity score (RSS) of TFs in immune (b) and epithelial (c) cells. The scatter plot represents TFs that were identified in both cattle and humans; the x axis shows the RSS of each TF in cattle and the y axis shows the RSS in humans. Representative TFs were considered significant (RSS > 0.25) and are color-labeled. The r represents the Pearson correlation coefficient (PCC) measuring the linear association of RSS values between the two species. The corresponding P value was calculated based on a two-sided t-test and adjusted for multiple comparisons (FDR). d–g, Cellular communication in the small intestine (d,e) and liver (f,g) between cattle and humans. The network diagram shows the analysis of cellular communication between cattle and humans; the column stacking diagram represents the proportion of the two species in the signaling pathway. Red represents pathways significantly upregulated in cattle, while blue represents pathways significantly upregulated in humans; black represents pathways that are not significant in either cattle or humans. h,i, Heritability enrichment of cell types for orthologous marker genes with MS (h) and IBD (i). The y axis represents the significance level (−log10(P)) of heritability enrichment and the x axis represents all cell types identified in the corresponding cattle tissue. P values were derived from the heritability enrichment analysis and adjusted for multiple comparisons (FDR). The red dashed line represents the significance threshold of FDR = 0.05. The figure was created with BioRender.com.
To explore whether the CattleCA resource can contribute to the explanation of genetic and cellular mechanisms underlying complex human traits and diseases, we analyzed the heritability enrichment of 43 human traits and diseases using linkage disequilibrium score regression (LDSC) on the orthologous marker genes across 49 cattle tissues. Significant enrichment of heritability for human complex traits and diseases was found in corresponding tissues in cattle (Supplementary Fig. 25a). For instance, the orthologous marker genes in cattle jejunum, colon and reticulum showed significant enrichment of heritability for human IBD and celiac disease, while markers in the blood, cecum, ileum and pituitary gland showed enrichment for multiple sclerosis (MS) (FDR < 0.05).
We analyzed four diseases—MS, IBD, rheumatoid arthritis (RA) and celiac disease—because of their significant heritability enrichment and the potential links in their mechanisms of onset and progression in humans and cattle46. We calculated the heritability enrichment for these conditions across seven tissues by extending the categories of orthologous marker genes to cell types. Remarkably, certain cell types showed significant heritability enrichments, enhancing our understanding of their genetic and molecular basis. For instance, CD4+ T cells exhibited substantial heritability enrichment for MS and RA, while CD8+ T cells showed enrichment for CD and IBD (Fig. 8h,i and Supplementary Fig. 25b,c). Moreover, several cell types displayed significant heritability enrichments for diseases that have not been extensively studied. For example, microglia in the pituitary gland showed significant enrichment for MS (Fig. 8h), while differentially expressed genes (DEGs) in pituitary microglia were enriched in glycerophospholipid and ether lipid metabolism (Extended Data Fig. 4d), suggesting that alterations in lipid metabolism pathways in microglia may contribute to the pathogenesis of MS47. Additionally, GCs in the colon were significantly enriched for IBD (Fig. 8i) and had strong communication with immune cells (Fig. 4d,h–j), suggesting their potential protective roles in the intestinal epithelial barrier.
Discussion
By generating and analyzing the gene expression of 1,793,854 cells across 59 tissue types in 15 cattle, we annotated 131 cell types to build the CattleCA and provided a web portal for the community to explore and query all the results. The following comprehensive inter-tissue and intra-tissue analyses of cellular heterogeneity advanced our understanding of bovine lactation and health. For instance, in mammary epithelial cells, we explored how specialized subpopulations act and contribute to prolonged milk synthesis under high metabolic and complex environmental demands, providing insights into the cellular complexity of the mammary gland. Because the liver has a central role in volatile fatty acid metabolism, gluconeogenesis and nitrogen cycling33, we investigated subtypes of hepatocytes associated with lactation, inferred their spatial distribution and found that those in dairy cattle might have a pivotal role in protein and lipid metabolism, as well as energy supply for lactation compared to human hepatocytes. Integrating this resource with large-scale population genetics data facilitated the detection of relevant cell types and stages for complex phenotypes. For instance, among all the annotated cell types, spermatocytes and excitatory neurons showed the strongest association with SMOT and milk FY, respectively, in dairy cattle. This knowledge can further serve as biological priors for prioritizing causal genes and variants and improving the prediction accuracy of complex phenotypes41. The cross-species comparative analysis revealed a high similarity in gene expression, regulation and cellular communication between cattle and humans at single-cell resolution. These findings will contribute to understanding the cellular and evolutionary mechanisms underlying zoonoses associated with cattle, such as tuberculosis, salmonellosis and ringworm48, facilitating the development of bovine models for certain human diseases49.
Although it provides a valuable resource for the cattle genetics and genomics community, the current CattleCA has some limitations: (1) our current samples were limited to Holstein cattle, a dairy breed of global economic value. It is thus imperative to encompass a broader range of cattle breeds, such as beef cattle; (2) more biological and environmental contexts should be considered in the future development of CattleCA, such as embryonic stages, healthy status and diet changes because some cell types (for example, macrophages) and states may only be observed in certain contexts; (3) additional single-cell omics data, such as epigenome, proteome and spatial transcriptome, are required for accurately and precisely annotating cell types and stages50; and (4) further functional experiments are needed to validate the proposed regulatory mechanisms, including ligand–receptor interactions in cellular communication and TFs influencing cell differentiation trajectories. A more comprehensive CattleCA will enable a deeper understanding of the dynamic landscape of cellular function across diverse biological and environmental contexts, advancing our understanding of the molecular mechanisms underlying complex phenotypes and environmental adaptations in cattle and even in humans.
Methods
Ethics statement
The experimental procedures were approved by the Institutional Animal Care and Use Committee at the China Agricultural University (approval no. DK996), Northwest A&F University (approval no. DK20230113) and the Animal Care Committee of Zhejiang University (approval no. ZJU202017326).
scRNA-seq, snRNA-seq and data analyses
This study analyzed the scRNA-seq and snRNA-seq transcriptome data of 152 samples representing 59 tissues from 15 Holstein cattle (Fig. 1b, Supplementary Fig. 2, Supplementary Table 3 and Supplementary Note).
Identification of cell clusters
Seurat v.4.0.6 (ref. 51) was used to perform unsupervised clustering. Libraries from the same tissue were merged and underwent normalizing and scaling. Harmony v.0.1.1 (ref. 15) was used to correct four batch effects (sources, methods, platforms and individuals) with the resetting parameters (lambda = 1, theta = 0.5). Variable genes were determined using the Seurat’s FindVariableGenes function with default parameters (selection.method = vst, nfeatures = 2,000). Clusters were identified using the FindClusters function (resolution = 0.5) implemented in Seurat using the top 30 principal components and subsequently visualized using the runUMAP function (reduction = harmony). Artificial annotation was performed on each cell cluster based on the marker genes reported in the relevant scientific literature.
Cell cycle index estimation
To obtain additional insights into the dynamic information about cell states, a cell cycle index was computed for each cell type using the CellCycleScoring function in Seurat51. Cells were categorized into G0/G1, S and G2/M phases, denoting distinct cell cycle stages.
Distribution analysis of cell type preference
To characterize the tissue distribution of mammary epithelial cells, ORs were calculated and used to indicate preferences. Specifically, for each cell subtype i and tissue j, a 2 × 2 matrix was constructed, which contained the number of cells of cell subtype i in tissue j, the number of cells of cell subtype i in other tissues, the number of cells of non-i-cell subtype in tissue j and the number of cells of non-i-cell subtype in other tissues. Then, a Fisher’s exact test was applied to this matrix; thus, the OR and corresponding P value could be obtained. P values were adjusted using the Benjamini–Hochberg method implemented in the R function p.adjust. An OR value > 1.5 indicated that cell type i was preferentially distributed in tissue j; an OR value < 0.5 indicated that cell type i was not preferentially distributed in tissue.
Pseudotime trajectory analysis
Monocle2 v.2.26 (ref. 52) was used to infer the state transition of the cell types and subtypes. The unique molecular identifier count matrix of the cells was used to create the CellDataSet object and then filter out the genes expressed in fewer than ten cells. Genes with q < 0.01 were identified as DEGs using the differentialGeneTest function and sorted according to q value using the setOrderingFilter function. The pseudotime trajectory was constructed using the DDRTree algorithm using default parameters. The dynamic expression changes of selected marker genes in pseudotime were visualized using the plot_genes_in_pseudotime and plot_pseudotime_heatmap functions. To explore the process of spermatogenesis, germline cells were extracted from the testis and then redescended for clustering with pcs = 15 and resolution set to 0.5. To explore the mechanism of monogenic disorder genes in the process of cell differentiation, UTCs and SMCs were extracted from the placenta and esophagus, respectively. Batch effects were corrected using Harmony v.0.1.1 and then redescended for clustering with pcs = 30 and resolution set to 0.1. To explore the developmental process of cell types across tissues, cells were extracted and merged from each tissue, batch effects were corrected using Harmony v.0.1.1 and then redescended for clustering with pcs = 15 and resolution = 0.4 for myeloid cells. Two thousand cells from each macrophage and monocyte cluster were randomly selected for trajectory analysis. For B cells and hepatocytes, their resolutions were 1 and 0.5, respectively. Monocle3 v.1.3.1 (ref. 53) was then used to reconstruct the cellular differentiation trajectory. Specifically, subsets were separated into large and well-divided partitions using the cluster_cells function and the principal graph was fitted in each partition using the learn_graph function. The resulting principal graph, revealed on the UMAP as ‘skeleton lines’, indicates the differentiation trajectories.
Cellular communication analyses
Cellular communication analyses were implemented using the CellChat v.1.6.1 R package54 for each tissue in label-based mode. Default parameters were used, except that min.cells was set to ten, which allows filtering out cell types with fewer than ten cells. All annotated cell types were classified based on their cell lineage. Interactions between different cell types were then aggregated and the average intensity was computed to assess the comprehensive dynamics of cellular communication networks. Furthermore, cellular communication patterns in diverse tissues were examined in detail. To compare cellular communication across tissues, the cellular communication analysis in specific tissues was first performed separately; then, the datasets were merged using mergeCellChat. Finally, the netVisual_diffInteraction function was used to compare and analyze differences in communication strength.
Gene regulatory network analysis
Gene regulatory network inference was performed using the Python package pySCENIC v.0.12.1 (ref. 55) with default parameters. The raw counts, derived from the Seurat object, were based on one-to-one homologous genes between humans and cattle using the HomoloGene database (release 1.4.68.19.3.27; https://github.com/oganm/homologene; Supplementary Table 17). The human TF list (https://resources.aertslab.org/cistarget/tf_lists/allTFs_hg38.txt) was used as a reference to identify co-expression modules using the GRNBoost2 algorithm. Then, regulons were obtained by detecting the genes directly targeted by the TF and removing other genes based on the enrichment of motifs within 10 kb from the target transcription start site using the cisTarget databases (Homo sapiens, hg38, refseq_r80, SCENIC+ databases, gene-based database (https://aertslab.org/)). Using AUCell, the regulon activity score was measured as the area under the recovery curve. The activities associated with each cell type were evaluated by calculating the RSS56. The Jensen–Shannon divergence was calculated according to the TF expression; the TF-specific score was defined as 1-√Jensen–Shannon divergence. The z-score was then calculated to normalize the TF-specific score to predict the basic TF in each cell type according to the following formula: \(z\;{\rm{score}}=\frac{{x}_{{ij}}-{\mu }_{i}}{{\sigma }_{i}}\); where xij was the RSS for TF j in cell type i, μi was the average RSS of all the TFs in cell type i and σi was the RSS s.d. of all TFs in cell type i. The correlation coefficient r and P value were calculated using the corr.test function (method = pearson, adjust = fdr). For each cell type in a given tissue, a TF was deemed ‘positive’ in that particular cell type if its regulon was positive in more than 25% of cells, and it also exhibited a z-score exceeding 0.75.
Regulon modules were identified based on the Connection Specificity Index (CSI) using scFunctions (https://github.com/FloWuenne/scFunctions/)57. First, the PCC of the activity scores was evaluated for each pair of regulons. Next, for a fixed pair of regulons, A and B, the corresponding CSI was defined as the fraction of regulons whose PCC with A and B was lower than the PCC between A and B. To conduct a global comparison, we extracted 10% of cells from each tissue cluster, retaining all cells from clusters with fewer than 500 cells, and identified regulons using pySCENIC based on co-expression and motif enrichment. This process was repeated five times, generating ~200,000 cell sets per iteration and identifying ~300 regulons. Hierarchical clustering with Euclidean distance was performed based on the CSI matrix to identify different regulon modules. We also used CSI > 0.7 as a cutoff to build the regulon association network to investigate the relationship between different regulons.
Function enrichment analysis
GO and KEGG analyses were performed using clusterProfiler v.4.0 (ref. 58). The GO terms and KEGG pathways of selected genes were enriched in the org.Bt.eg.db and bta databases, using the enrichGO and enrichKEGG functions, respectively, with a threshold parameter of pvalueCutoff = 0.05. GSVA v.1.49.6 (ref. 59) and fgsea v.1.24.0 (ref. 60) were used to investigate functional differences between cell subtypes. The bovine KEGG and GO databases were downloaded from the Molecular Signatures Database (MSigDB) using msigdbr v.7.5.1 (https://github.com/igordot/msigdbr), after which the gsva and gsea functions were used to calculate the pathway scores in the pathway for each subtype-specific gene set.
aPEAR v.1.0.0 (ref. 61). was used to objectively summarize the enrichment results by using the similarity between pathway gene sets and representing them as interconnected cluster networks. Each cluster was assigned a meaningful name that highlights the main biological theme in the experiment.
Analysis of cell type diversity
The Shannon entropy62 was calculated to evaluate cell type diversity in each tissue according to the formula \(-{\sum }_{{\rm{i}}}({{\rm{p}}}_{{\rm{i}}}\times {\log }_{2}({{\rm{p}}}_{{\rm{i}}}))\), where pi is the proportion of cell type in cell class i for each tissue. They were then plotted in R v.4.2.0 using ggplot2 v.3.4.1 and ComplexHeatmap v.2.15.4 (ref. 63).
Cell type conservation analysis
MetaNeighbor 1.18 (ref. 64) was used to provide a measure of the replicability of cell types. The highly variable genes were identified using the variableGenes function and the correlations between cell types were determined based on AUROC values using MetaNeighborUS analysis.
Cell function scoring
The AddModuleScore function in Seurat was used to score each signature in each cell subtype; gene sets to measure antigen presentation ability (Supplementary Table 9) were obtained from the MSigDB database (www.gsea-msigdb.org/gsea/msigdb). The classically activated M1 macrophages gene sets are SOCS1, NOS2, TNF, CXCL9, CXCL10, CXCL11, CD86, IL1A, IL1B, IL6, CCL5, IRF5, IRF1 and CCR7; the selectively activated M2 macrophages gene sets are IL4R, CCL4, CCL18, CCL22, MARCO, VEGFA, CTSA, CTSB, TGFB1, MMP9, CLEC7A, MSR1, IRF4, CD163, TGM2 and MRC1. Data were compared using a Wilcoxon rank-sum test (****P < 0.0001; NS = P > 0.05). Functional gene sets were also used to evaluate the lactation ability of epithelial cells in the mammary gland, the metabolic function of macrophages and the function of B cells (Supplementary Tables 8 and 9). The association between GC subtypes and diseases was evaluated using five gene sets, including a total of 529 genes, with two gene sets from the animal QTL database (www.animalgenome.org/cgi-bin/QTLdb/index), two from the KEGG database and one set of upregulated genes identified in the disease group29 (Supplementary Table 12). The metabolic functions of macrophage subtypes were evaluated using scMetabolism and the AUCell method, based on the KEGG metabolic database65.
High-dimensional weighted correlation network analysis
The hdWGCNA v.0.2.24 (ref. 66) was used to perform high-dimensional weighted gene co-expression network analysis based on single-cell data. First, we input the genes expressed in at least 5% of cells and used the MetacellsByGroups function to construct the metacell gene expression matrix. Then, the TestSoftPowers function was used to determine the soft power. The ConstructNetwork function was used to build the co-expression network. All analyses were conducted according to standard procedures (https://smorabit.github.io/hdWGCNA/articles/hdWGCNA.html).
Cattle monogenic disorder gene collection
A total of 634 monogenic disorders in cattle were assembled from the Online Mendelian Inheritance in Animals database (www.omia.org/home). Candidate genes associated with 145 disorders were identified. To facilitate systematic analysis, these disorders were classified into ten groups based on their phenotypic manifestation, including conditions related to blood and immune systems, connective tissue, embryonic development, embryonic lethality, metabolic processes, muscle function, neural disorders, reproductive issues, skin-related disorders and tissue-specific conditions (Supplementary Table 14). To ensure the robustness of the statistical analyses, a minimum threshold of more than five candidate genes was imposed for each disorder group.
Cell-type-specific gene identification
Cell-type-specific genes were identified using the z-score. Initially, the count matrix underwent a transformation into the CPM values; the average was computed in each cell type to create a pseudobulk expression matrix67. Subsequently, tspex v.0.6.2 (ref. 68) was used to calculate the z-score values under the log10-transformed CPM matrix. Genes exhibiting a z-score > 0.75 in each cell type were designated as cell-type-specific genes; cell-type-specific genes in a given cell lineage were defined as cell-lineage-specific genes.
Enrichment analysis between cell types and monogenic disorders
The correlation between cell-type-specific and monogenic disorder genes was assessed using chi-squared and Fisher’s exact tests, using a 2 × 2 matrix consisting of intersected genes, disorder genes, cell-type-specific genes and all genes. A chi-squared test was used when the expected frequencies for all elements exceeded five; otherwise, a Fisher’s exact test was used. Subsequently, FDR values were computed using the Bonferroni method to address multiple comparisons. Additionally, ORs were also calculated to validate the accuracy of significant results based on the same 2 × 2 matrix.
Gene coding DNA sequence region alignment between humans and cattle
The DNA sequences of the gene coding DNA sequence region were downloaded from GenBank (www.ncbi.nlm.nih.gov/genbank/). Sequence alignment was performed based on the Basic Local Alignment Search Tool method (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
GWAS analysis
The single-nucleotide polymorphism (SNP) BeadChip and phenotypic data of 16,188 Chinese Holstein cows were assembled, including 9,045 bovine 150K BeadChip, 1,505 bovine 80K BeadChip and 5,638 bovine 50K BeadChip, from the Dairy Association of China. The phenotypic data included 55 traits, spanning milk production, sperm, coat color, IgG, body conformation and health. All SNP BeadChip data were mapped to the bovine reference genome (ARS-UCD1.2); 132,961 SNPs were obtained after imputing to the 150K level using Beagle v.5.1 (ref. 69). Then, they were imputed again to the genome-scale sequence level using a high-quality sequencing imputation panel of 28,166,177 SNPs based on 3,530 cattle. After filtering out low-quality SNPs with ‘dosage R2 < 0.9, minor allele frequency < 0.05 and Hardy–Weinberg Equilibrium test result P < 0.0001’, 8,535,460 SNPs were obtained. Finally, GWAS analysis was performed using Genome-wide Complex Trait Analysis v.1.94.0 (ref. 70) with the ‘--fastGWA-mlm--pca 5--grm’ option; the results were visualized using CMplot v.4.4.1 (ref. 71).
Enrichment analysis between cell types and complex traits
scPagwas v.1.3.0 (ref. 18) was used to perform the enrichment analysis between cell types and complex traits. It uses a polygenic regression model to prioritize a set of trait-relevant genes and uncover trait-relevant cell subpopulations by incorporating pathway activity-transformed scRNA-seq data with GWAS summary data. To enhance the comprehensiveness of our results, 319 human KEGG pathways downloaded from the KEGG database (www.genome.jp/kegg) were used after eliminating duplicates and converting homologous genes. The Boot_evaluate function was used to identify the significant trait-relevant cell types and calculate the TRS. The scGet_PCC function was used to prioritize the top trait-relevant genes by ranking the PCCs. Genes with the top 50 PCC values were defined as trait-relevant genes in each cell type. In addition, the scPagwas_perform_score function was applied to perform pathway activity analysis and define the significance of active pathways in each cell type based on the singular value decomposition method. Enrichment analysis between trait-relevant genes and active pathway genes was performed based on Fisher’s exact and chi-squared tests in each cell type.
Cross-species cellular transcriptome similarity analysis
All cross-species analyses at the single-cell level were conducted using 18,148 one-to-one homologous genes between humans and cattle, which were downloaded from the HomoloGene database (release 1.4.68.19.3.27; Supplementary Table 17). For each cell type, 500 cells were randomly selected. Cell type conservation across species was assessed using MetaNeighbor v.1.181 (ref. 64). For the cellular communication analysis, significant homologous receptor–ligand pairs in cattle were detected by referencing the human database using the CellChat v.1.6.1 R package54. The netVisual_diffInteraction function was used to analyze differences in communication strength across tissues and species. For the TF analysis, the TF datasets for cattle and humans were separately inferred using pySCENIC v.0.12.1 (ref. 55). The correlation coefficient r and P value were calculated using the corr.test function (method = pearson, adjust = fdr).
LDSC
LDSC72 was used to detect whether the heritability of a phenotype was enriched around highly specifically expressed genes in a given tissue or cell type. All SNPs associated with the trait were obtained from publicly available data. The comparison of tissue contributions involved the selection of the top 200 DEGs (log2(fold change) ≥ 1.5 and FDR ≤ 0.05), which were sorted using the FDR from least to most for each tissue as a category. All DEG categories for tissues were collectively input to run the LDSC for traits. Subsequently, P values were computed using the Benjamini–Hochberg method to account for multiple comparisons. Similarly, the top 200 DEGs of each cell type cluster meeting the same criteria were selected for analysis; the results were visualized using ggplot2 v.3.4.1.
Statistics and reproducibility
No statistical method was used to predetermine sample size but our sample sizes are similar to those reported in previous publications5,6. The details of data exclusions for each specific analysis are available in the Methods. The experiments were not randomized. The investigators were not blinded to allocation during the experiments and outcome assessment. Data normalization and transformation were carefully performed to ensure that all data being analyzed met the assumptions of the statistical tests used.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All scRNA-seq and snRNA-seq data newly generated in this study are available for download from the Sequence Read Archive under accession no. PRJNA1119173. The processed datasets and expression profiles of annotated cell types are available via the CattleCA web portal (https://ngdc.cncb.ac.cn/cattleca/). The GWAS summary statistics are freely available at Zenodo via https://doi.org/10.5281/zenodo.16572998 (ref. 73). Source data are provided with this paper.
Code availability
All computational code is freely available at GitHub (https://github.com/FarmGTEx/CattleCellAtlas_pipeline_V0) and at Zenodo (https://doi.org/10.5281/zenodo.15721498) (ref. 74).
References
Bruford, M. W., Bradley, D. G. & Luikart, G. DNA markers reveal the complexity of livestock domestication. Nat. Rev. Genet. 4, 900–910 (2003).
Hawkins, J. W. et al. High-yield dairy cattle breeds improve farmer incomes, curtail greenhouse gas emissions and reduce dairy import dependency in Tanzania. Nat. Food 3, 957–967 (2022).
Hu, Z.-L., Park, C. A. & Reecy, J. M. Bringing the Animal QTLdb and CorrDB into the future: meeting new challenges and providing updated services. Nucleic Acids Res. 50, D956–D961 (2021).
Liu, S. et al. A multi-tissue atlas of regulatory variants in cattle. Nat. Genet. 54, 1438–1447 (2022).
Han, X. et al. Construction of a human cell landscape at single-cell level. Nature 581, 303–309 (2020).
Wang, F. et al. Endothelial cell heterogeneity and microglia regulons revealed by a pig cell landscape at single-cell level. Nat. Commun. 13, 3620 (2022).
Gao, Y. et al. Single-cell transcriptomic analyses of dairy cattle ruminal epithelial cells during weaning. Genomics 113, 2045–2055 (2021).
Wu, J.-J. et al. Microbiota-host crosstalk in the newborn and adult rumen at single-cell resolution. BMC Biol. 20, 280 (2022).
Xue, M.-Y. et al. Investigation of fiber utilization in the rumen of dairy cows based on metagenome-assembled genomes and single-cell RNA sequencing. Microbiome 10, 11 (2022).
Gao, Y. et al. Single-cell transcriptomic and chromatin accessibility analyses of dairy cattle peripheral blood mononuclear cells and their responses to lipopolysaccharide. BMC Genomics 23, 338 (2022).
Gu, F. et al. Gut microbiome is linked to functions of peripheral immune cells in transition cows during excessive lipolysis. Microbiome 11, 40 (2023).
Wang, L. et al. A single-cell atlas of bovine skeletal muscle reveals mechanisms regulating intramuscular adipogenesis and fibrogenesis. J. Cachexia Sarcopenia Muscle 14, 2152–2167 (2023).
Wu, J.-J. et al. Cross-tissue single-cell transcriptomic landscape reveals the key cell subtypes and their potential roles in the nutrient absorption and metabolism in dairy cattle. J. Adv. Res. 37, 1–18 (2022).
Wu, J.-J. et al. Age- and microbiota-dependent cell stemness plasticity revealed by cattle cell landscape. Research 6, 0025 (2023).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Twigger, A.-J. et al. Transcriptional changes in the mammary gland during lactation revealed by single cell sequencing of cells from human milk. Nat. Commun. 13, 562 (2022).
Vlasova, A. N. & Saif, L. J. Bovine immunology: implications for dairy cattle. Front. Immunol. 12, 643206 (2021).
Ma, Y. et al. Polygenic regression uncovers trait-relevant cellular contexts through pathway activation transformation of single-cell RNA sequencing data. Cell Genom. 3, 100383 (2023).
Seachrist, D. D. & Keri, R. A. The activin social network: activin, inhibin, and follistatin in breast development and cancer. Endocrinology 160, 1097–1110 (2019).
Iovino, M. et al. Oxytocin signaling pathway: from cell biology to clinical implications. Endocr. Metab. Immune Disord. Drug Targets 21, 91–110 (2021).
Green, I. D. et al. Macrophage development and activation involve coordinated intron retention in key inflammatory regulators. Nucleic Acids Res. 48, 6513–6529 (2020).
Li, L., Jin, H., Xu, J., Shi, Y. & Wen, Z. Irf8 regulates macrophage versus neutrophil fate during zebrafish primitive myelopoiesis. Blood 117, 1359–1369 (2011).
Delfini, M., Stakenborg, N., Viola, M. F. & Boeckxstaens, G. Macrophages in the gut: masters in multitasking. Immunity 55, 1530–1548 (2022).
Thomson, A. W. & Knolle, P. A. Antigen-presenting cell function in the tolerogenic liver environment. Nat. Rev. Immunol. 10, 753–766 (2010).
Mass, E., Nimmerjahn, F., Kierdorf, K. & Schlitzer, A. Tissue-specific macrophages: how they develop and choreograph tissue biology. Nat. Rev. Immunol. 23, 563–579 (2023).
Jego, G. et al. Dual regulation of SPI1/PU.1 transcription factor by heat shock factor 1 (HSF1) during macrophage differentiation of monocytes. Leukemia 28, 1676–1686 (2014).
Rauschmeier, R. et al. Bhlhe40 and Bhlhe41 transcription factors regulate alveolar macrophage self-renewal and identity. EMBO J. 38, e101233 (2019).
Harris, N. B. & Barletta, R. G. Mycobacterium avium subsp. paratuberculosis in veterinary medicine. Clin. Microbiol. Rev. 14, 489–512 (2001).
Gao, Y. et al. Genome-wide association study of Mycobacterium avium subspecies paratuberculosis infection in Chinese Holstein. BMC Genomics 19, 972 (2018).
Muise, A. M. et al. Polymorphisms in E-cadherin (CDH1) result in a mis-localised cytoplasmic protein that is associated with Crohn’s disease. Gut 58, 1121–1127 (2009).
Khounlotham, M. et al. Compromised intestinal epithelial barrier induces adaptive immune compensation that protects from colitis. Immunity 37, 563–573 (2012).
Farr, L. et al. CD74 signaling links inflammation to intestinal epithelial cell regeneration and promotes mucosal healing. Cell Mol. Gastroenterol. Hepatol. 10, 101–112 (2020).
Trefts, E., Gannon, M. & Wasserman, D. H. The liver. Curr. Biol. 27, R1147–R1151 (2017).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Wei, Y. et al. Liver homeostasis is maintained by midlobular zone 2 hepatocytes. Science 371, eabb1625 (2021).
Nicholas, F. W. Online Mendelian Inheritance in Animals (OMIA): a record of advances in animal genetics, freely available on the Internet for 25 years. Anim. Genet. 52, 3–9 (2021).
Aumailley, M. et al. A simplified laminin nomenclature. Matrix Biol. 24, 326–332 (2005).
Kiritsi, D., Has, C. & Bruckner-Tuderman, L. Laminin 332 in junctional epidermolysis bullosa. Cell Adh. Migr. 7, 135–141 (2013).
Liu, M., Yin, Y., Yu, H. & Zhou, R. Laminins regulate placentation and pre-eclampsia: focus on trophoblasts and endothelial cells. Front. Cell Dev. Biol. 8, 754 (2020).
Uitto, J. & Richard, G. Progress in epidermolysis bullosa: genetic classification and clinical implications. Am. J. Med. Genet. C Semin. Med. Genet. 131C, 61–74 (2004).
Fang, L. et al. Comprehensive analyses of 723 transcriptomes enhance genetic and biological interpretations for complex traits in cattle. Genome Res. 30, 790–801 (2020).
Li, M. et al. Gut–brain circuits for fat preference. Nature 610, 722–730 (2022).
Watt, M. J. & Hoy, A. J. Lipid metabolism in skeletal muscle: generation of adaptive and maladaptive intracellular signals for cellular function. Am. J. Physiol. Endocrinol. Metab. 302, E1315–E1328 (2012).
Neto, F. T., Bach, P. V., Najari, B. B., Li, P. S. & Goldstein, M. Spermatogenesis in humans and its affecting factors. Semin. Cell Dev. Biol. 59, 10–26 (2016).
Duan, Y.-G. et al. Dendritic cells in semen of infertile men: association with sperm quality and inflammatory status of the epididymis. Fertil. Steril. 101, 70–77 (2014).
Naser, S. A., Sagramsingh, S. R., Naser, A. S. & Thanigachalam, S. Mycobacterium avium subspecies paratuberculosis causes Crohn’s disease in some inflammatory bowel disease patients. World J. Gastroenterol. 20, 7403–7415 (2014).
Ladakis, D. C. et al. Metabolomics of multiple sclerosis lesions demonstrates lipid changes linked to alterations in transcriptomics-based cellular profiles. Neurol. Neuroimmunol. Neuroinflamm. 11, e200219 (2024).
McDaniel, C. J., Cardwell, D. M., Moeller, R. B. Jr & Gray, G. C. Humans and cattle: a review of bovine zoonoses. Vector Borne Zoonotic Dis. 14, 1–19 (2014).
Saied, A. A., Metwally, A. A., Mohamed, H. M. A. & Haridy, M. A. M. The contribution of bovines to human health against viral infections. Environ. Sci. Pollut. Res. Int. 28, 46999–47023 (2021).
Zeng, H. K. What is a cell type and how to define it? Cell 185, 2739–2755 (2022).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Jin, S. Q. et al. Inference and analysis of cell–cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Suo, S. et al. Revealing the critical regulators of cell identity in the Mouse Cell Atlas. Cell Rep. 25, 1436–1445 (2018).
Fuxman Bass, J. I. et al. Using networks to measure similarity between genes: association index selection. Nat. Methods 10, 1169–1176 (2013).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Kerseviciute, I. & Gordevicius, J. aPEAR: an R package for autonomous visualization of pathway enrichment networks. Bioinformatics 39, btad672 (2023).
Eraslan, G. et al. Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376, eabl4290 (2022).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Crow, M., Paul, A., Ballouz, S., Huang, Z. J. & Gillis, J. Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor. Nat. Commun. 9, 884 (2018).
Wu, Y. et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 12, 134–153 (2022).
Morabito, S. et al. Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer’s disease. Nat. Genet. 53, 1143–1155 (2021).
Li, H. et al. Fly Cell Atlas: a single-nucleus transcriptomic atlas of the adult fruit fly. Science 375, eabk2432 (2022).
Camargo, A. P., Vasconcelos, A. A., Fiamenghi, M. B., Pereira, G. A. G. & Carazzolle, M. F. tspex: a tissue-specificity calculator for gene expression data. Preprint at Research Square https://doi.org/10.21203/rs.3.rs-51998/v1 (2020).
Browning, B. L., Zhou, Y. & Browning, S. R. A one-penny imputed genome from next-generation reference panels. Am. J. Hum. Genet. 103, 338–348 (2018).
Jiang, L., Zheng, Z., Fang, H. & Yang, J. A generalized linear mixed model association tool for biobank-scale data. Nat. Genet. 53, 1616–1621 (2021).
Yin, L. et al. rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study. Genomics Proteomics Bioinformatics 19, 619–628 (2021).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Han, B. FarmGTEx/CattleCellAtlas/GWAS_summary_V1. Zenodo https://doi.org/10.5281/zenodo.16572998 (2025).
Li, H. FarmGTEx/CattleCellAtlas_pipeline_V0. Zenodo https://doi.org/10.5281/zenodo.15721498 (2025).
Acknowledgements
D.S. was supported by the National Key R&D Program of China (no. 2021YFF1000700), the Program for Changjiang Scholar and Innovation Research Team in University (no. IRT_15R62) and the 2115 Talent Development Program of China Agricultural University. B.H. was supported by the National Natural Science Foundation of China (no. 32372836) and the Youth Elite Development Program of College of Animal Science and Technology, China Agricultural University. L.F. was supported by an Agriculture and Food Research Initiative Competitive grant no. 2022-67015-36215 (H.Z.) from the USDA National Institute of Food and Agriculture. G.E.L. was supported in part by Agriculture and Food Research Initiative Competitive grant nos. 2019-67015-29321 and 2021-67015-33409 from the USDA National Institute of Food and Agriculture. Y.H. was supported by the National Key R&D Program of China (no. 2024YFF1000100). J.C.H. was funded by the Biotechnology and Biological Sciences Research Council through Institute Strategic Programme Funding (grant no. BBS/E/RL/230002B). J.F.O. was supported by the Science Foundation Ireland Centre for Research Training in Genomics Data Science (grant no. 18/CRT/6214). B.L. was supported by the UK Biotechnology and Biological Sciences Research Council with grant no. BB/X009505/1. We thank the High-Performance Computing Platform of China Agricultural University (Beijing) for the support provided and the Xihe High-Performance Computing Platform of the National Research Facility for Phenotypic and Genotypic Analysis of Model Animals (Beijing).
Author information
Authors and Affiliations
Contributions
All authors made substantial contributions to the conception or design of the study; the acquisition, analysis or interpretation of the data; or the drafting or revision of the paper. D.S., L.F., B.H., H.S., Y.J. and G.E.L. conceived and designed the study. D.S., Y.J., H.S., Z.M., L. Liu. and J.H. provided samples and data. H.L., B.H., S.Z. and T.S. performed the bioinformatic analyses of the scRNA-seq and snRNA-seq data. Q.Z., W.Z., A.C., Y.S., W.Y., A.D., Y.F., M.J. and T.S. manually annotated the cell types. W.Z., Q.Z. and B.H. conducted the intra-tissue cellular heterogeneity analysis. W.Z. and B.H. conducted the inter-tissue cellular heterogeneity analysis. H.L. and W.Z. conducted the integrative analysis of the genetic variants. A.C., B.H. and Z.H.B. conducted the comparative analysis between cattle and humans. Y.H., Z.Z., D.Z. and Z.Y. built the CattleCA web portal. D.S., L.F. and B.H. contributed to the data and computational resources. D.S., L.F., G.E.L., Y.H., F.W., H.S., Y.J., W.L., W.T. and J.C.H. contributed to the critical interpretation of the analytical results before and during the manuscript preparation. B.H., H.L., W.Z., Q.Z. and A.C. drafted the manuscript. L.F., D.S., G.E.L., Y.H., F.W., H.S., Y.J., W.L., W.T., J.C.H., D.E.M., J.F.O., O.M., G.S., Y.L., L. Lin., C.L., Z.C., B.L. and Z.Z. revised the manuscript. All authors read, edited and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Transcriptional regulatory landscape across tissues and cell types in cattle.
(a) The connection specific index (CSI) matrix consisting of 156 regulons is divided into 6 modules (left). UMAPs illustrate the average AUCell score distribution in different regulon modules (in different colors). Wordcloud plots highlight the top 10 cell lineages and tissues exhibiting the highest average regulatory activity for each submodule, where color intensity and font size reflect the regulatory activity levels. Representative TFs and their corresponding binding motifs are also displayed (right). (b) Zoomed-in view of module 5 (including 23 regulons) identifies sub-module structures and their average regulatory activity in different cell types. (c) Bar plot shows the number of TFs regulating each cell lineage. The x-axis indicates TF counts, color-coded by category: cell-type specific TFs (active in a single cell type), lineage-specific TFs (active across multiple cell types within the same lineage), multi-lineage TFs (active across multiple lineages), and broadly active TFs (tau ≤ 0.85). (d) The TF regulatory network containing 123 highly specific TFs (tau > 0.85) exhibits tissue-specific, cell-type-specific, single-lineage-specific, and multi-lineage-regulated TFs. (e) Heatmap illustrates regulatory activity differences of 682 TFs between males and females. The color gradient indicates the difference in average regulon activity, highlighting a portion of the 45 regulons with an absolute difference greater than 0.15. To ensure unbiasedness, we retained 61 cell types with counts exceeding 50 in both males and females across 19 tissues, where at least one sex exhibited regulatory activity ( > 25% of cells showing regulatory activity).
Extended Data Fig. 2 Dynamics of cell-cell communication networks in cattle.
(a) Chordal diagram of the integrated cell lineage–cell lineage interaction network. (b) Scatter plot shows the interaction intensity and number of cellular interactomes among major cell lineages, clustered by the number of interactions. (c) Scatter plot shows 19 highly reliable secreted ligand-receptor pairs (probability > 0.2, p-value < 0.05, permutation test) between cell types in different tissues. Colors represent the probability of cellular communication. Red boxes represent tissue-specific ligand-receptor pairs. (d) Comparative analysis of cellular communication strength between cell types in the cerebral cortex, medulla oblongata, and cerebellum of males and females. To ensure unbiased comparison, we retained cell types with more than 50 cells in both sexes for each tissue and downsampled to ensure equal numbers of males and females for each cell type. The red line indicates increased communication in females, while the blue line indicates increased communication in males. (e) Scatter plot shows 6 highly reliable ligand-receptor pairs (probability > 0.2 and p-value < 0.05 after permutation test in males or females) involving excitatory neurons as receptors or ligands in the nervous system.
Extended Data Fig. 3 Single-cell atlas of spermatogenic cells in Holstein cattle.
(a) UMAP visualization of 14 annotated spermatogenic subtypes, with cells color-coded according to subtypes. (b) Bubble plot displays the standardized expression of marker genes for each spermatogenic subtype. (c-d) Box plots show the number of expressed genes (c) and normalized UMI counta (d) in spermatogenic subtype at different differentiation stages in testis (n = 6). The central band in the boxplot represents the median, the box boundaries represent the 25% to 75% percentiles, and the whiskers extend 1.5 × the interquartile range. (e) GWAS enrichment analysis shows spermatogenic subtypes significantly associated with the semen volume per ejaculate (SVPE), the initial sperm motility (SMOT), the sperm concentration per ejaculate (SCPE), the number of sperm per ejaculate (NSPE) and the number of motile sperm per ejaculate (NMSPE). *, **, and *** indicates FDR < 0.05, 0.01, and 0.001, respectively. (f) Pseudotime trajectory analysis of spermatogenic subtypes and cell states colored on the trajectory tree. Pie chart shows the proportion of subtypes on SPT_1 and SPT_2. (g) Heatmap shows the variation of regulatory activity of the 43 regulons over pseudotime, which were identified through rank-sum tests based on pseudotime and regulator activity (r > 0.2, FDR < 0.05). (h) Heatmap shows the changes in gene expression of representative DEGs (FDR < 1e-4) in pseudotime series, and GO enrichment analysis of DEGs re-clustered into four clusters (right). P-value was calculated using a one-sided hypergeometric test and adjusted by multiple comparisons (FDR). (i) Volcano plot illustrates the DEGs between SPT_1 and SPT_2. (j) Bar plot displays GO terms with up-regulated DEGs in SPT_1 and SPT_2. The p-value calculation is the same as in Figure h. (k) Bubble plot displays the standardized expression of key gene sets involved in spermatogenesis for SPT_1 and SPT_2.
Extended Data Fig. 4 Functional differences between myeloid cell subtypes.
(a) Bubble plot displays the standardized expression of marker genes for each cluster of myeloid cells. (b) Heatmap represents the tissue distribution, with odds ratios (OR) calculated and used to indicate preferences. * indicates OR > 1.5 and FDR < 0.05; ** indicates OR > 1.5 and FDR < 0.01. Bar chart shows the composition proportion of myeloid cell subtypes. (c) Heatmap displays the expression levels of up-regulated DEGs in myeloid cells in 40 different tissues, divided into 10 modules. (d) Network diagram represents the GO terms enriched by genes from each module.
Extended Data Fig. 5 Heterogeneity of macrophages across tissues.
(a) Heatmap shows the average regulatory activity z-score of 245 TFs of macrophages in 29 tissues. (b) Heatmap represents the key metabolic pathways enriched by macrophage subtypes. (c) Box plot shows the gene signature scores of MHC I (21 genes) and MHC II (26 genes) in macrophage subtypes (sample size = 11), calculated by AddModuleScore in Seurat 4.0.6. The central band in the boxplot represents the median, the box boundaries represent the 25% to 75% percentiles, and the whiskers extend 1.5 × the interquartile range. Statistical test was calculated based on Wilcoxon test with two-sided. The same sample size and statistical test were applied to the following Figures d-f. (d) Box plot shows the scores of M1 (14 genes) and M2 (16 genes) in macrophage subtypes, calculated by AddModuleScore of Seurat. (e) Box plot shows the scores of M1 and M2 in macrophage in 40 tissues, calculated by AddModuleScore of Seurat. (f) Box plot shows the pseudotime of each macrophage subtype.
Extended Data Fig. 6 Heterogeneity of B/plasma cells across tissues.
(a) UMAP visualization of all annotated major cell subtypes of B/plasma cells, with cells color-coded according to cell subtypes. (b) Bubble plot displays the standardized expression of marker genes for each B/plasma cell subtype. (c) Heatmap shows Pearson correlation of gene expression levels for each B/plasma cell type, calculated based on the top 1,000 genes with the largest standard deviation. (d) Heatmap represents the tissue distribution of B/plasma cell subtype, odds ratios (OR) were calculated and used to indicate preferences. * indicates OR > 1.5 and FDR < 0.05; ** OR > 1.5 and FDR < 0.01. (e) Bar chart shows the composition proportion of B/plasma cell subtypes of each tissue. (f) Heatmap shows the scores of gene signatures in B/plasma cell subtypes, calculated by AddModuleScore of Seurat. (g) Heatmap shows the average regulatory activity z-score of key TFs in 238 TFs that regulate differentiation and maintenance of B/plasma cell subtypes. (h) Trajectory analysis of B/plasma cell subtypes. (i) Curve plot shows the dynamic expression scores for high-affinity, low-affinity, activated, exhaustion, BCSR and CSR signatures in cells of four different differentiation pathways, respectively, along the inferred pseudotime. The center line indicates linear fit, and shaded lines indicate a 95% confidence interval. (j) Curve plot shows the dynamic expression scores for TFs in cells of four different differentiation pathways, respectively, along the inferred pseudotime. The center line indicates linear fit, and shaded lines indicate a 95% confidence interval.
Extended Data Fig. 7 Comparison of Species between bovine and human stomach.
(a) UMAP visualization of annotated cell subtypes of the human stomach and four cattle stomachs (abomasum, rumen, reticulum, and omasum). (b) The tree chart shows the hierarchical clustering of tissues based on gene expression levels of the top 1000 genes with the largest standard deviation (left), the scatter plot shows the number of cell types (middle), and the stacked bar chart shows the proportion of epithelial cells (right). (c) Network diagram shows the KEGG pathway enriched by up-regulated DEGs in each tissue. P-value was calculated based on hypergeometric test with one-sided. (d) Heatmap shows the TFs with the highest differences in regulatory specificity scores between each tissue in epithelial cells. (e) Trajectory analysis of epithelial cells in cattle stomach with cell state colored on the tree (upper left), the pie chart displays the cell type composition of each state (upper right), and tissue colored on the tree (bottom). (f) Pseudo-heatmap illustrates the dynamics of DEGs during cell fate at branch point 1 (left), and KEGG enrichment analysis of DEGs clustered into three clusters (right). P-value was calculated based on hypergeometric test with one-sided. (g) Pseudo-heatmap illustrates the dynamics of DEGs during cell fate at branch point 2 (left), and KEGG enrichment analysis of DEGs clustered into four clusters (right). P-value was calculated based on hypergeometric test with one-sided.
Extended Data Fig. 8 Heterogeneity of spinous cells across three forestomaches.
(a) Scatter plot shows the optimal soft threshold or power to make the constructed network more consistent with the scale-free topology. (b) Hierarchical clustering diagram shows the construction of a co-expression network based on the optimal soft threshold, dividing the genes into different modules to draw a gene clustering tree. The upper part is the hierarchical clustering tree of genes, and the lower part is the gene module. (c) Correlation bubble diagram depicts the associations between modules and spinous cell subtypes, as well as the GO terms of selected modules. P-value was calculated based on hypergeometric test with one-sided. (d) Module network diagram of the top 25 hub genes in the salmon, magenta, and greenyellow modules. (e) TF regulatory network diagram of the salomon, magenta, and greenyellow modules in the rumen.
Extended Data Fig. 9 Expression level, cell communication, and functional enrichment for skin disorder-related genes.
(a) Expression levels of 27 skin disorder-related genes in different epithelial cell types. The dot size represents log10CPM values and the color represents the z-score of genes in each cell type. The skin disorders corresponding to the genes are denoted in parentheses. (b) The expression distribution of LAMC2 in the duodenum primarily occurs in intestinal progenitor cells, which is associated with the skin disorder epidermolysis bullosa (EB). (c) Trajectory analysis of uninucleate trophoblast cells (UTCs) in the placenta. The dark to light blue represents the potential differentiation direction of UTCs. (d) UMAP of UTC subtypes in the placenta. (e) Cellular communication between UTC subtypes and other cell types in the placenta. The line thickness represents the communication strength between two cell types. (f) Expression levels and enriched GO terms of marker genes in UTC cell subtypes. The bubble plot shows the marker genes with the top three log2FoldChange values. Bubble size corresponds to the percentage of cells expressing each gene, and color reflects the average gene count in each subtype. GO terms in each subtype are listed (right). P-value was calculated based on hypergeometric test with one-sided.
Extended Data Fig. 10 Correlations between cell types and traits.
Correlations between cell types and coat color (a), IgG (b), and body and health (c) traits. Each circle represents a cell-type-trait association. The x-axis represents cell lineages sorted in alphabetical order.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–25, Tables 1–17, Note, Methods and References.
Source data
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, B., Li, H., Zheng, W. et al. A multi-tissue single-cell expression atlas in cattle. Nat Genet 57, 2546–2561 (2025). https://doi.org/10.1038/s41588-025-02329-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02329-5