Abstract
Foxp3-expressing regulatory T (Treg) cells protect against systemic autoimmunity. However, little is known about the significance of Treg cells in inflammation-experienced tissues. Here, we use an experimental autoimmune encephalomyelitis model and show that Treg cells accumulate and persist in the central nervous system (CNS) long after the resolution of the bulk of the inflammatory infiltrate. CNS-specific depletion of postinflammatory Treg cells, but not systemic depletion of Treg cells, results in autoimmune inflammatory flares in the CNS by residual local effector T cells. Expression of the NAD-consuming ectoenzyme CD38 is crucial for the functional adaptation of postinflammatory CNS Treg cells to a stressful microenvironment, in which access to interleukin-2 (IL-2) is limited. CD38 counteracts ADP-ribosylation of the IL-2 receptor and thus maintains its high sensitivity to IL-2. This fully functional high-affinity IL-2 receptor prevents the loss of tissue-resident antigen-specific Treg cells. These ‘stress-tolerant’ CNS Treg cells impede the collapse of immune homeostasis in the CNS once acute inflammation is controlled.
Similar content being viewed by others
Main
Foxp3+ regulatory T (Treg) cells have regained attention due to their ability to adapt to tissue niches and exert noncanonical functions in tissue homeostasis and regeneration1,2. Although the developmental trajectory of tissue-resident Treg cells is consistent with a thymic origin of these cells in most tissues, peripherally induced Treg cells might be prevalent in some barrier-associated tissues with a high load of commensal microbiota3. A widely accepted model of tissue Treg cell development suggests a multistep process, involving the imprinting of a tissue Treg cell precursor subset in secondary lymphoid tissue and the terminal instruction to tissue Treg cells by the specific milieu in the target tissue4,5. The dwell time of Treg cells in nonlymphoid tissues may vary depending on the tissue context6, and egress of Treg cells from these tissues, rejoining the circulation, may occur7. However, the description of Treg cell population dynamics and functional phenotype has mostly been performed in the steady state8,9. It is unclear how Treg cells behave in nonlymphoid tissues with low-grade chronic inflammation, as is observed in most autoimmune diseases, including multiple sclerosis.
It is well established that Treg cells are essential in re-establishing immune homeostasis in any adaptive immune response, where IL-2-driven feedforward loops lead to the expansion of effector T cells10. Once autoimmune inflammation in the central nervous system (CNS) has been initiated, thymus-derived Treg cells are recruited to the CNS11. In the target tissue, they adopt an effector phenotype, which is steered by a network of transcription factors, including BLIMP1 and BATF12,13,14. This transcriptional network also secures the preservation of Treg cell identity in the face of inflammation. Because highly activated effector T cells are resilient to Treg cell-mediated control11, it has been suggested that conventional T cell-intrinsic down-modulatory mechanisms must be initiated before efficient Treg cell-mediated suppression can take effect.
Treg cells that accumulate in the CNS in autoimmune inflammatory diseases (but also after stroke, in traumatic brain injury and perhaps even neurodegenerative diseases) were suggested to be involved in tissue repair and regulation of organ function by a series of different mechanisms15,16,17,18,19,20. The relevance of Foxp3+ Treg cells for these processes has been demonstrated in loss- and gain-of-function approaches. However, whether Treg cells that accumulate in the CNS after the peak of experimental autoimmune encephalomyelitis (EAE; a paradigmatic model for autoimmune CNS disease) actually contribute to the re-establishment and subsequent maintenance of immune homeostasis in the CNS is not known. The postinflammatory CNS is a hostile environment for Treg cells, as it is low in interleukin-2 (IL-2) due to the contraction of the conventional T (Tconv) cell compartment and simultaneously holds high levels of nicotinamide adenine dinucleotide (NAD+)21. High levels of extracellular NAD+ have been shown to induce apoptosis in Treg cells, in part by facilitating the ADP-ribosylation of the ATP-gated ion channel P2RX7 (ref. 22), which induces a gain-of-function of this molecule with a downstream increase in intracellular Ca2+ (ref. 23). Nevertheless, Foxp3+ Treg cells constitute a stable population in the postinflammatory CNS.
Here, we show that the NAD+ consuming enzyme CD38 controls stress resilience of CNS-resident Treg cells by preventing ADP-ribosylation and subsequent loss of function of the IL-2 receptor α-subunit (IL-2Rα; CD25). This function enables CNS Treg cells to survive despite a limited amount of IL-2 in the microenvironment and to exert compartmentalized immune control in the inflammation-experienced CNS.
Results
Treg cells persist in the CNS after recovery from an autoimmune response
To investigate the population dynamics and function of Treg cells in the CNS during recovery from autoimmune inflammation, we used the MOG35–55-induced EAE model in Foxp3 (green fluorescent protein (GFP)) reporter mice24. After an acute monophasic inflammatory episode, MOG35–55-induced EAE is characterized by a prolonged period of recovery, during which clinical symptoms improve and stabilize but usually never fully resolve (Fig. 1a). Although the neurological sequelae in this model are due to incomplete recovery from axonal damage caused during the acute phase of the disease, the inflammatory infiltrate in the CNS in the recovery phase is largely reduced. Here, we focused on this ‘re-established’ homeostatic condition in recovery and interrogated the CNS Treg cell compartment.
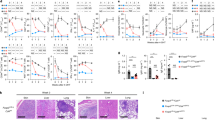
a, Mean EAE scores + s.e.m. (sample sizes as in c and d) in Foxp3 (GFP) mice following immunization. Naive (without immunization), preonset (day 8 after immunization), peak (day 15 after immunization) and recovery (day 29 after immunization). b, Cytograms with percentages of Tconv (CD3+CD4+Foxp3−) and Treg (CD3+CD4+Foxp3+) cells among total CD4+ T cells in the brain and spinal cord. c,d, Fractions and absolute (Abs.) counts of Tconv (c) and Treg cells (d) in the brain. Open symbols represent individual mice. Data are shown as mean + s.d. P values in c and d are from a two-way analysis of variance (ANOVA) with a Tukey’s honestly significant difference (HSD) test (naive n = 5, preonset n = 4, peak n = 3, recovery n = 3 biological replicates). e, EAE was induced in Cd4Cre × Rosa26LSL-mitoDendra2 mice, and inguinal lymph nodes (iLNs) were photoconverted at peak or during recovery. Two days later, photoconverted mitoDendra2RED (mDRED) CD4+ T cells were analyzed in the spleen (SPL) and CNS. f, Cytograms of the CD4+ T cell compartment with the fraction of photoconverted cells (%). In the bar graphs, each symbol represents an individual mouse. Data are shown as mean + s.d. P values are from an unpaired two-tailed Student’s t-test (peak n = 6, recovery n = 10 biological replicates). g, Photoconverted CNS Tconv (CD4+CD25−GITR−mDRED+) and Treg (CD4+CD25+GITR+mDRED+) cells were quantified. Each symbol represents an individual mouse. Data are shown as mean + s.d. P values are from an unpaired two-tailed Student’s t-test (n = 3 biological replicates). h, In vivo EdU incorporation was measured in C57BL/6 wild-type EAE mice at peak and during recovery. i, Gating was performed on extravascular (CD45.2−) near-IR dead−CD4+ T cells. The percentages of EdU+ Tconv (Foxp3−) and Treg (Foxp3+) cells were quantified. Symbols represent individual mice. Data are shown as mean + s.d. P values are from a two-way ANOVA with a Tukey’s HSD test (peak n = 4, recovery n = 3 biological replicates). j, In vivo Foxp3 fate mapping was conducted in Foxp3Cre-ERT2 × Rosa26LSL-tdTomato EAE mice. Mice were injected with tamoxifen (i.p.) for 5 days at the EAE recovery stage, and CNS cells were analyzed 14 days after the last injection. k, Representative cytogram with percentages of stable (Foxp3 (GFP)+tdTomato+) and unstable (Foxp3 (GFP)−tdTomato+) Treg cells within the CNS CD4+ T cell gate during EAE recovery. l, Stable and unstable CNS Treg cells were sorted and analyzed by bulk RNA-seq (n = 5 biological replicates). An enrichment plot for ‘IL-2–STAT5 signaling’ derived from GSEA comparing stable and unstable CNS Treg cells is shown; FDR, false discovery rate; NES, normalized enrichment score. m, CD25 expression of stable and unstable CNS Treg cells at the recovery phase. Each symbol represents an individual mouse. Data are shown as mean + s.d. P are values from a paired two-tailed Student’s t-test (n = 3 biological replicates); MFI, mean fluorescence intensity.
Although Tconv cell numbers declined significantly between the peak and early recovery phase (Fig. 1b,c), Treg cell numbers remained largely stable during this time frame in both the brain and spinal cord (Fig. 1b,d). Consequently, the local frequency of Treg cells within the CNS CD4+ T cell compartment increased significantly in the postinflammatory CNS (Fig. 1d). Notably, Treg cells persisting in the postinflammatory CNS were not evenly distributed but were located in certain niches around the third ventricle and in the meningeal space (Extended Data Fig. 1).
To gauge the contribution of the systemic immune compartment to the CNS Treg cell pool, we labeled mitoDendra2-expressing T cells in the draining lymph nodes of MOG-immunized mice at the peak of EAE and during recovery (day 30) and tracked photoconverted T cells to the spleen and CNS (Fig. 1e). Although CD4+ T cells still trafficked to the CNS during peak disease, the recruitment of T cells from the initial priming site to the CNS significantly dropped during recovery (Fig. 1f) and was negligible for Treg cells (Fig. 1g). Conversely, the fraction of proliferating CNS Treg cells was about 10% at the peak of EAE and dropped to 3% in the postinflammatory CNS (Fig. 1h,i). Together, these data suggest that the Treg cell pool in the postinflammatory CNS was essentially disconnected from its priming site in the systemic immune compartment and, to some extent, had self-renewing capacity.
To investigate the stability of postinflammatory Treg cells in their CNS environment, we used Foxp3Cre-ERT2 × Rosa26LSL-tdTomato mice to perform fate mapping of Foxp3+ Treg cells. Tamoxifen was administered at EAE recovery (Fig. 1j). This allowed us to categorize Treg cells into stable (Foxp3 (GFP)+tdTomato+) and unstable (Foxp3 (GFP)−tdTomato+) populations. We termed those Treg cells that persisted as stable Foxp3 expressors in inflammation-experienced tissues ‘stress-tolerant’ Treg cells. Notably, most CNS Treg cells maintained stable Foxp3 expression in the recovery phase, with only 15 to 20% of CNS Treg cells losing Foxp3 expression (Fig. 1k). To identify potential drivers of instability in recovery CNS Treg cells, RNA-sequencing (RNA-seq) analysis was performed in stable and unstable Treg cell populations isolated during the recovery phase (Extended Data Fig. 2a,b and Supplementary Tables 1 and 2). Stress-tolerant Treg cells expressed key immunosuppressive molecules, including Il2rb, Ctla4 and Areg. Conversely, unstable Treg cells exhibited increased expression of Bach2, a transcription factor known to reduce Foxp3 expression25, and Tcf7, which suppresses the transcription of genes co-bound by Foxp3 (ref. 26). These findings suggest that the downregulation of Foxp3 in unstable Treg cells is linked to the activity of Bach2 and Tcf7. By contrast, the dominant pathway associated with CNS Treg cell stability in the postinflammatory CNS was the ‘IL-2–STAT5 signaling’ pathway (Fig. 1l and Extended Data Fig. 2c). To confirm our RNA-seq findings in stress-tolerant Treg cells, we analyzed the expression of IL-2Rα (CD25) in stable and unstable Treg cells using flow cytometry. Consistent with the transcriptomic data, IL-2Rα expression was significantly lower in unstable than in stable CNS Treg cells (Fig. 1m), further highlighting the strong association between IL-2 signaling and Foxp3 stability in stress-tolerant Treg cells.
Stress-tolerant Treg cells control compartmentalized CNS inflammation
To study the functional relevance of CNS-residing stress-tolerant Treg cells after resolution of acute inflammation (>day 35), we used Foxp3-DTR mice, which harbor a knock-in of the diphtheria toxin (DTx) receptor (DTR) gene expressed under the control of the endogenous Foxp3 locus, for transient ablation of Foxp3-expressing cells via administration of DTx27. Because a complete depletion of Treg cells from the CNS at late stages of EAE is challenging to achieve using the conventional systemic (intraperitoneal (i.p.) or intravenous (i.v.)) routes of DTx injection, we chose to administer low doses of DTx (15 ng) directly into the CNS using stereotactic intracerebroventricular (i.c.v.) injection (Fig. 2a). Local DTx delivery resulted in almost complete removal of CNS Treg cells, while the systemic compartment was only transiently (and partially) depleted of Foxp3+ Treg cells (Fig. 2b,c). Depletion of CNS Treg cells was accompanied by a rapid worsening of EAE symptoms (Fig. 2d and Extended Data Fig. 3), that is, the return of complete hind limb paralysis. Although some relapses were controlled again after 5 days, most DTx-treated mice succumbed to their relapses (Extended Data Fig. 3b,c). The disease relapses of i.c.v. DTx-injected Foxp3-DTR mice were not due to DTx-mediated CNS toxicity because i.c.v. DTx-injected wild-type control mice did not show any exacerbation of clinical symptoms. By contrast, efficient systemic depletion of Treg cells in recovered EAE mice by i.p. DTx injection, which largely spared the CNS Treg cell compartment, failed to trigger an EAE relapse (Fig. 2d).
a, EAE was induced in Foxp3 (GFP) and Foxp3-DTR mice. DTx was administered i.c.v. (15 ng) or i.p. (500 ng) during the recovery phase of EAE. Mice were killed 2 and 3 days after DTx injection, and spleen and CNS cells were isolated and analyzed by flow cytometry. b,c, The efficiency of Treg cell depletion 2 days after DTx injection is shown in the spleen and CNS by the Treg cell percentage (b) and the CNS Treg cell count (c). Each symbol in b and c represents an individual mouse. Data are shown as mean + s.d. P values were calculated using a one-way ANOVA with a Tukey’s HSD test (DTx i.c.v. control n = 6, DTx i.p. n = 7, DTx i.c.v. n = 8). d, EAE scores after DTx administration are shown as mean + s.e.m. (only including scores of mice followed up until the day 3 endpoint). P values were calculated using a two-way ANOVA with a Tukey’s post hoc test (DTx i.c.v. control n = 4, DTx i.p. n = 5, DTx i.c.v. n = 7 biological replicates). e, CD4+ T cell counts in the CNS at different time points after DTx administration. Each symbol represents an individual mouse. Data are shown as mean + s.d. P values were calculated using a two-way ANOVA with a Tukey’s HSD test (day 2: DTx i.c.v. control n = 6, DTx i.p. n = 7, DTx i.c.v. n = 8; day 3: DTx i.c.v. control n = 4, DTx i.p. n = 5, DTx i.c.v. n = 5 biological replicates). f, CD69 expression (MFI) on CNS CD4+ Tconv cells on day 3 after DTx treatment. Symbols represent individual mice. Data are shown as mean + s.d. P values were calculated using a one-way ANOVA with a Tukey’s HSD test (DTx i.c.v. control n = 4, DTx i.p. n = 5, DTx i.c.v. n = 5 biological replicates).
Only the near-complete (i.c.v. DTx), but not the partial (i.p. DTx), depletion of CNS Treg cells led to a marked increase in the amount of activated CD4+CD69+ T cells in the CNS of Foxp3-DTR mice by day 3 after DTx administration (Fig. 2e,f). In histologic analyses, T cells and myeloid cells primarily expanded at the sites of pre-existing inflammatory infiltrates after i.c.v. injection of DTx into post-EAE Foxp3-DTR mice (Fig. 3a,b). Although some effector cells were recruited from the systemic compartment (Fig. 3c), FTY720 did not prevent the massive activation of those Tconv cells already residing in the CNS (Fig. 3d). Their activation status was similar to the activation of CNS Tconv cells in non-FTY720-treated i.c.v. DTx-injected mice (Fig. 3d). Accordingly, FTY720 treatment failed to prevent EAE relapse following complete local depletion of recovery CNS Treg cells (Fig. 3e), suggesting that residual CNS Tconv cells are sufficient to drive a relapse following removal of local Treg cells.
a, EAE was induced in Foxp3 (GFP) and Foxp3-DTR mice. DTx was administered i.c.v. or i.p. during the recovery phase of EAE. Mice were killed 3 days after DTx injection. Brains were collected for immunohistochemistry staining. Arrows and squares indicate immune cell niches. Representative stainings for CD45, CD4 and IBA-1 are shown; scale bars, top row: 800 μm, bottom three rows: 50 μm. b, Quantification of CD45+ and CD4+ cells as percentages and absolute counts per mm2. Symbols represent individual mice. Data are shown as mean + s.d. P values were calculated using a one-way ANOVA with a Tukey’s HSD test (DTx i.c.v. control n = 3, DTx i.p. n = 5, DTx i.c.v. n = 9 biological replicates). c,d, EAE was induced in Foxp3-DTR mice, and DTx was administered i.c.v. during the recovery phase. FTY720 (FTY; 1 μg per gram per day) or PBS was administered i.p. from 24 h before to 72 h after DTx i.c.v. injection. CD4+ T cell counts in the CNS (c) and MFI of CD69 on CNS CD4+ Tconv cells (d) as determined by flow cytometry are shown. Symbols represent individual mice. Data are shown as mean + s.d. The P value in c was determined by a two-tailed unpaired Student’s t-test (DTx i.c.v. plus PBS n = 4, DTx i.c.v. plus FTY n = 3 biological replicates). e, EAE scores after i.c.v. DTx administration shown as mean + s.e.m. (DTx i.c.v. plus PBS n = 4, DTx i.c.v. plus FTY n = 4 biological replicates).
In summary, stress-tolerant CNS Treg cells, but not systemic Treg cells, are essential for maintaining immune homeostasis in postinflammatory EAE mice.
Stress-tolerant CNS Treg cells have a tissue-type transcriptome
To determine whether the changing inflammatory milieu within the CNS between the peak and the remission phase of EAE would shape the transcriptional profile of stress-tolerant CNS Treg cells, we performed RNA-seq of bulk CD4+Foxp3 (GFP)+ Treg cells and CD4+Foxp3 (GFP)− Tconv cells sorted by fluorescence-activated cell sorting (FACS) from the spleen and CNS on days 15 and 40 after EAE induction.
RNA-seq analysis revealed a total of 311 differentially expressed genes (DEGs; >1.5-fold, adjusted P < 0.05) between CNS Treg cells derived from EAE recovery and EAE peak, with 159 genes upregulated in recovery CNS Treg cells and 152 genes upregulated in peak CNS Treg cells (Extended Data Fig. 4a), consistent with a distinct positioning of CNS Treg cells along principal component 2 (PC2) according to the disease phase (Fig. 4a). The number of disease-phase conditional DEGs was smaller within splenic Treg cells, with only 15 upregulated and 6 downregulated genes in recovery splenic Treg cells versus peak splenic Treg cells. Compared to their disease-phase-specific splenic counterparts, both recovery and peak CNS Treg cells showed an upregulation of several genes that have previously been associated with a nonlymphoid tissue Treg cell phenotype, including Areg, Nfil3, Prdm1, Rora, Itgae, Icos, Il10, Il1rl1 and Il2ra (Extended Data Fig. 4b,c). Although most of the tissue Treg cell-associated genes did not undergo disease-phase-dependent changes in expression, the ‘TNF signaling via NF-κB’ and ‘IL-2–STAT5 signaling’ pathways were the two top upregulated pathways in recovery CNS Treg cells compared to peak CNS Treg cells (Extended Data Fig. 4a–h), suggesting that the optimal exploitation of limited amounts of IL-2 after the contraction of the IL-2-producing effector T cell population (see Fig. 1c) might become the most crucial determinant of their ‘resilience to stress’ in CNS recovery Treg cells.
EAE was induced in Foxp3 (GFP) reporter mice. Tconv and Treg cells were sorted from the spleen and CNS during peak EAE or the recovery phase (rec) and were subjected to bulk RNA-seq. a, PC analysis. Each symbol represents an individual mouse. b,c, Volcano plots showing genes with annotation to the Gene Ontology (GO) terms ‘GO:0009986 cell surface’ (b) and ‘GO:0003700 DNA binding transcription factor activity’ (c) with differential expression (≥1.5-fold, adjusted P < 0.05) between recovery phase and peak CNS Treg cells. DEGs are highlighted in orange. Adjusted P values were derived from DESeq2 using a two-tailed Wald’s test with correction for multiple testing using the Benjamini and Hochberg method (CNS Treg peak n = 4, CNS Treg recovery phase n = 5 biological replicates); TF, transcription factor. d, Heat map with hierarchical clustering of DEGs between CNS recovery phase and peak Treg cells. DEGs with the same phase-dependent regulation in CNS Tconv cells as in CNS Treg cells were subtracted to yield a CNS Treg private gene signature. e, Upregulated DEGs from the recovery phase CNS Treg private signature were used to perform GO enrichment analysis (gprofiler 2g:GOSt). P values are derived from Fisher’s one-tailed test, and adjusted P values were determined by the g:GOSt implemented g:SCS method for multiple testing correction. The top five enriched terms are marked and summarized in the table on the right; MF, molecular function; CC, cellular component; BP, biological process; KEGG, Kyoto Encyclopedia of Genes and Genomes; REAC, Reactome; WP, WikiPathways; Reg., regulation; multicell., multicellular; org., organismal; proc., process; Pos., positive; biol., biological. f,g, Volcano plots displaying only genes that are part of both the GO terms ‘response to stress’ (GO:0006950) and ‘cell surface’ (GO:0009986; f) or ‘DNA-binding transcription factor activity’ (GO:0003700; g) between CNS recovery phase and peak Treg cells. DEGs (≥1.5-fold, adjusted P < 0.05) from the recovery phase CNS Treg private ‘up’ signature are highlighted in purple. Adjusted P values are derived from DESeq2 using a two-tailed Wald’s test with correction for multiple testing using the Benjamini and Hochberg method.
To identify upstream and downstream determinants of the stress-tolerant CNS Treg cell functional phenotype, we first analyzed genes differentially expressed in recovery CNS versus peak CNS Treg cells, which encoded cell surface proteins or DNA-binding proteins with transcription factor activity (Fig. 4b,c). We then performed functional enrichment analyses, only selecting those DEGs that were not at the same time significantly regulated in recovery versus peak Tconv cells to define a CNS Treg cell-exclusive signature of up- and downregulated transcripts in the recovery phase (Fig. 4d and Supplementary Table 3). Functional enrichment analyses with the upregulated genes revealed a few significantly regulated cellular processes, including ‘response to stress’ (Fig. 4e). Notably, the genes that were exclusively upregulated in recovery CNS Treg cells and at the same time intersected the GO terms ‘response to stress’ and ‘cell surface’ comprised Nrp1, Cxcr6, Cxcr4, Sdc4, H2-Aa, H2-Ab, Lag3, Fasl and Cd38 (Fig. 4f). Those recovery CNS Treg cell DEGs that intersected the GO terms ‘response to stress’ and ‘transcription factor activity’ comprised Rbpj, Nr4a1, Nr4a2, Nfatc1, Klf10, Zeb2 and Irf4 (Fig. 4g). Together, these data suggest that T cell antigen receptor (TCR) signaling and IL-2 signaling are defining features of the functional phenotype of stress-tolerant Treg cells residing in the chronically inflamed CNS. Interestingly, CD38 was strongly linked to this Treg cell phenotype.
Stress-tolerant Treg cells and tissue Treg cells share features
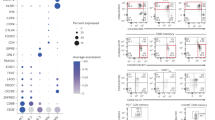
To test whether stress-tolerant Treg cells in the postinflammatory CNS are distinct from bona fide tissue Treg cells, we compared their transcriptome with the transcriptomes of Treg cells isolated from the CNS of unmanipulated naive mice and prototypic tissue Treg cells isolated from the visceral adipose tissue (VAT). Due to the scarcity of Foxp3+ Treg cells in the CNS of naive mice, the cell input had to be adjusted to a lower number in this RNA-seq analysis, resulting in a lower overall sequencing depth across all samples. Nevertheless, PC analysis of the 500 top most variable genes revealed a clear separation along PC1 of the transcriptome of CNS-derived Treg cell populations from the transcriptome of steady-state VAT Treg cells, in part driven by the considerably higher expression of tissue Treg-associated genes such as Areg, Klrg1, Gata3 and Il1rl1 in VAT Treg cells (Fig. 5a). However, stress-tolerant Treg cells also showed a clear overlap with VAT Treg cells for the expression of other tissue Treg-associated genes (Extended Data Fig. 5a), suggesting that stress-tolerant Treg cells adopt key features of bona fide tissue Treg cells. Notably, within the CNS Treg cell compartment, the transcriptome of stress-tolerant Treg cells resembled that of naive CNS Treg cells, and both separated along PC2 from the transcriptome of Treg cells isolated during the peak of disease (Fig. 5a), indicating that the stress-tolerant CNS Treg cell transcriptional profile reverts back to a more steady-state-like gene expression. However, despite the similarities to the transcriptome of homeostatic CNS Treg cells, gene set enrichment analysis (GSEA) again confirmed an enrichment in the IL-2 signaling pathway in stress-tolerant CNS Treg cells versus naive CNS Treg cells (Fig. 5b and Extended Data Fig. 5b–d), whereas naive CNS Treg cells were characterized by more prominent oxidative phosphorylation than stress-tolerant postinflammatory CNS Treg cells (Fig. 5c and Extended Data Fig. 5b–d). A selection of Treg cell-associated surface markers, including CD69, CXCR4 and CD38, was markedly expressed in stress-tolerant CNS Treg cells and to a lesser extent in naive CNS Treg cells, whereas KLRG1 and ST2 were highly upregulated in VAT Treg cells (Fig. 5d,e). Together, these data indicate that postinflammatory stress-tolerant CNS Treg cells adopt a CNS-specific tissue signature28 with a particular dependence on IL-2.
Foxp3 (GFP)+ Treg cells were isolated from VAT and the CNS of naive mice and EAE mice at the peak (n = 3) and recovery stages and were subjected to bulk RNA-seq (VAT naive n = 3, CNS naive n = 4, CNS peak n = 3, CNS recovery n = 4 biological replicates). a, PC analysis. b,c, Enrichment plots for ‘IL-2–STAT5 signaling’ (b) and ‘Oxidative phosphorylation’ (c) derived from GSEAs comparing CNS recovery and CNS naive Treg cells. d, CD4+Foxp3+ Treg cells from the naive CNS, VAT and the EAE CNS (peak and recovery) were analyzed by flow cytometry. Flow data were pooled (CNS naive n = 3, CNS peak n = 4, CNS recovery n = 3, VAT naive n = 3 biological replicates) and downsampled to 300 Treg cells per group. Treg cells were clustered by FlowSOM and visualized using uniform manifold approximation and projection (UMAP). e, Heat map displaying marker expression in Treg cell metaclusters.
Lack of CD38 impairs the regulatory capacity of CNS Foxp3+ Treg cells
CD38 was upregulated in Treg cells in the inflamed CNS, whereas we did not observe disease-phase-dependent upregulation of CD38 in Tconv cells (Fig. 6a). CD38 is the major NAD+ hydrolyzing ectoenzyme29. In fact, CD38 is indispensable for maintaining the concentration of NAD+ in the microenvironment at low levels. High levels of NAD+, in turn, drive the ADP-ribosylation of cell surface molecules by ADP-ribosyltransferase 2b (ARTC2.2), a toxin-related, GPI-anchored ADP-ribosyltransferase expressed in T cells. ARTC2.2 has multiple target molecules that can gain or lose activity through ADP-ribosylation30.
a, EAE was induced in Foxp3 (GFP) mice, and CNS cells were collected and analyzed by flow cytometry to assess CD38 expression on Tconv (CD4+Foxp3−) and Treg cells (CD4+Foxp3+) across EAE stages. Symbols represent individual mice. Data are shown as mean + s.d. P values were calculated using a two-way ANOVA with a Tukey’s HSD test (naive n = 10, peak n = 10, recovery n = 6 biological replicates). b, EAE was induced in recipient DEREG mice (CD45.1) and donor Cd38+/+ or Cd38−/− mice (CD45.2). Treg cells in recipient mice were depleted by i.p. DTx injection at days 9 and 10 after EAE induction. In total, 105 sorted splenic donor Treg cells (CD4+CD25+GITR+CD45.2+) were transferred into CD45.1+ DEREG recipients. Treg protector S+16a nanobody was i.p. administered to donor mice 30 min before euthanasia; d.p.i., days post immunization. c, Mean EAE scores + s.e.m. of recipient mice. P values were determined using a two-way ANOVA with a Tukey’s post hoc test (without Treg cell transfer (wo) n = 5, Cd38+/+ Treg cell transfer n = 4, Cd38−/− Treg cell transfer n = 5 biological replicates). d–f, Donor CD4+CD45.2+ cells were analyzed and quantified in recipient lymph nodes (d and e) and CNS tissue (d and f). Each symbol in e and f represents an individual mouse, and the data are shown as mean + s.d.; Transf., transfer. g, BM from Foxp3-DTR mice (CD45.1/CD45.2) mixed 1:1 with either wild-type Cd38+/+ (CD45.2) or Cd38−/− BM (CD45.2) was transferred into sublethally irradiated Rag1−/− recipients, and EAE was induced in BM-reconstituted mice. DTx was administered i.c.v. (15 ng) in stable EAE mice on day 28 after MOG35–55 immunization, and CNS cells were collected 3 days after DTx i.c.v. injection. h, Proportions of Treg cells (TCRβ⁺CD4⁺Foxp3⁺) were analyzed in the Foxp3-DTR compartment (CD45.1+) and in the CD38-sufficient or CD38-deficient compartment (CD45.2+) within the CNS of MBMCs. i, Fraction of CD38⁺ cells among total CNS Treg cells 3 days after DTx injection. Symbols represent individual mice. Data are shown as mean + s.d. P values were calculated using an unpaired two-tailed Student’s t-test (Cd38+/+ MBMCs n = 4, Cd38−/− MBMCs n = 5 biological replicates). j, Changes in EAE scores (ΔEAE scores) after DTx injection shown as mean + s.e.m. (Cd38+/+ MBMCs n = 4, Cd38−/− MBMCs n = 5 biological replicates).
To test whether CD38 expression by Treg cells is relevant in autoimmune CNS inflammation, we developed an experimental model in which we adoptively transferred Treg cell populations enriched for MOG-reactive Treg cells that were either sufficient or deficient in CD38 into MOG-immunized host mice whose Treg cell compartment was acutely depleted immediately before Treg cell transfer (Fig. 6b). The transfer of MOG-activated wild-type Treg cells attenuated the EAE severity of recipient mice compared to control EAE mice that were Treg cell depleted but did not receive a subsequent Treg cell transfer (Fig. 6c). By contrast, the adoptive transfer of MOG-activated CD38-deficient Treg cells failed to control EAE symptoms of recipient mice that became as sick as control animals without transfer of Treg cells (Fig. 6c), suggesting that Cd38−/− Treg cells had a Treg cell-intrinsic deficiency to regulate EAE. Because CD38-deficient Treg cells were equally recruited to the CNS as wild-type Treg cells (Fig. 6d–f), we speculated that CD38 was directly or indirectly involved in the formation and maintenance of a productive effector Treg cell population in the CNS.
To gauge the particular significance of CD38 expression for the functional phenotype of stress-tolerant postinflammatory Treg cells in nonlymphoid tissues (especially in the CNS), we generated a mixed bone marrow chimera (MBMC) model by transplanting a 1:1 mixture of wild-type Foxp3-DTR+ (CD45.1/CD45.2) and ‘nondepletable’ Foxp3-DTR− (CD45.2) donor cells of either wild-type or Cd38−/− origin into Rag1−/− recipient mice (Fig. 6g). CNS-specific depletion of DTR-expressing wild-type Treg cells in these MBMCs by local DTx injection during stable post-peak EAE allowed us to follow disease progression in a scenario in which the remaining CNS-residing Treg cells were either wild-type or CD38 deficient (Fig. 6g–i). Notably, the Tconv cell compartment was composed of either only wild-type cells (Cd38+/+ MBMC) or a 1:1 mixture of wild-type and Cd38−/− T cells (Cd38−/− MBMCs; Fig. 6h). Consistent with our observation that the severity of EAE was reduced in a gene dose-dependent manner in globally CD38-deficient mice (Extended Data Fig. 6a), Cd38−/− MBMCs experienced a milder disease course than Cd38+/+ MBMCs (Extended Data Fig. 6b). Nevertheless, following local depletion of Foxp3-DTR Treg cells, the remaining nondepletable wild-type Treg cells in Cd38+/+ MBMCs were sufficient to maintain a stable EAE score (Fig. 6j and Extended Data Fig. 6b), even when faced with a fully CD38-competent CNS-resident Tconv cell compartment. By contrast, Cd38−/− MBMCs, with only Cd38−/− CNS Treg cells remaining after local depletion of the wild-type Treg cell fraction, rapidly deteriorated and experienced an EAE relapse with similar kinetics and intensity as we had observed after the complete depletion of CNS Treg cells (Fig. 6j and Extended Data Fig. 6b; see also Fig. 2d). These data highlight that cell-autonomous expression of CD38 in CNS-resident Treg cells is indispensable for adequate control of local immune homeostasis in the chronically inflamed CNS in vivo.
CD38 is crucial for maintenance of antigen-specific CNS Treg cells
Foxp3+ Treg cells need to be activated in an antigen-specific manner to efficiently control effector T cells in EAE31. Therefore, the most direct link to the failure of CD38-deficient Treg cells to control EAE would be the lack of MOG-specific Treg cells within the population of Cd38−/− CNS Treg cells. Indeed, in the CNS, MOG35–55–I-Ab tetramer-binding Treg cells were virtually entirely comprised in the CD38+ compartment, remarkably during EAE recovery (Fig. 7a–c).
EAE was induced in wild-type C57BL/6 mice. a,b, Percentages of MOG–I-Ab tetramer-binding Tconv and Treg cells in the CNS at different stages of EAE. c, CD38 expression in MOG–I-Ab− versus MOG–I-Ab+ Tconv and Treg cells across EAE stages. Each symbol in b and c represents an individual mouse. Data are shown as mean + s.d. P values were calculated using a two-way ANOVA with a Tukey’s HSD test (n = 5 biological replicates). d, Mixed BM from Cd38+/+ (CD45.1) and Cd38−/− (CD45.2) mice was transferred into sublethally irradiated Rag1−/− recipients. EAE was induced in BM-reconstituted mice, and CNS cells were collected 20 days after MOG35–55 immunization. Percentages of MOG–I-Ab-binding Cd38+/+ CD45.1+ and Cd38−/− CD45.2+ Tconv (TCRβ+CD4+Foxp3−; e) and Treg cells (TCRβ+CD4+Foxp3+; f). Each symbol in e and f represents an individual mouse. Data are shown as mean + s.d. The P value was calculated using a paired two-tailed Student’s t-test (n = 6 biological replicates).
To test whether CD38 status determined the fraction of MOG-specific Tconv and Treg cells, we induced EAE in MBMCs, whose hematopoietic compartment consisted of wild-type (CD45.1) and Cd38−/− (CD45.2) cells, and read out the fraction of MOG35–55–I-Ab-binding cells in Tconv and Treg cells in each cellular compartment that were present side-by-side in the same chronically inflamed CNS (day 20 after immunization; Fig. 7d). Notably, although CD38 expression was irrelevant for the fraction of MOG-specific Tconv cells in the CNS, the fraction of MOG-specific CNS Treg cells was significantly lower in CD38-deficient cells than in CD38-sufficient T cells (Fig. 7e,f). Together, these data indicate that the presence of antigen-specific Foxp3+ Treg cells, but not antigen-specific Tconv cells, in the inflamed CNS is dependent on the cell-intrinsic expression of CD38. This may be particularly relevant for high-affinity antigen-specific Treg cells because MOG–I-Ab tetramer staining identifies high-affinity MOG-specific Treg cells32.
Stress-tolerant CNS Treg cells are dependent on CD38 for sensing IL-2
To dissect the mechanism by which CD38 contributes to the maintenance of antigen-specific Treg cells in the postinflammatory CNS, we referred to its known function as an NAD+-consuming enzyme and speculated that Treg cells might be more susceptible to NAD+-induced cell death than Tconv cells in the chronically inflamed CNS. To directly test this hypothesis, we induced EAE in Foxp3 (GFP) reporter mice and, in the recovery stage, challenged the mice by i.c.v. injection of NAD+ in the absence or presence of an inhibitor of ARTC2.2 (Fig. 8a). Although NAD+ injection directly resulted in the apoptosis of around 90% of all CNS Treg cells, CNS Tconv cells were spared from apoptosis (Fig. 8b–d). Interestingly, i.c.v. injection of NAD+ did not affect Treg cells or Tconv cells in the systemic immune compartment (Extended Data Fig. 7a–c). In the CNS of recovery EAE mice, NAD+-induced apoptosis was entirely rescued by the co-injection of a blocking nanobody to ARTC2.2 together with NAD+ (Fig. 8b–d), suggesting that ADP-ribosylation by ARTC2.2 was the key downstream mediator of NAD+-induced cell death in vivo. Also, these data were consistent with an in vivo effect of NAD+ and not an artificial release of NAD+ from cells during the preparation procedure33. Although the gain-of-function of P2RX7 through ADP-ribosylation has previously been shown to drive cell death23, the expression of P2RX7 was not different between Treg cells and Tconv cells in the postinflammatory CNS. Yet, Tconv cells were relatively resistant to NAD+-induced apoptosis, whereas Treg cells were not. Therefore, we explored alternative Treg cell-specific ADP-ribosylation targets. Because IL-2 signaling had turned out to be a major determinant of stress-tolerant CNS Treg cells, we first focused on the IL-2Rα subunit, which is highly expressed in antigen-specific Treg cells (Extended Data Fig. 8a–c). Here, we made use of the prior observation that the R178 ADP-ribosylated hypofunctional form of IL-2Rα is not recognized by the 7D4 clone of anti-CD25, whereas the PC61.5 clone of anti-CD25 detects both the native and the ADP-ribosylated form of IL-2Rα34. To investigate IL-2Rα ADP-ribosylation status, wild-type or CD38-deficient Treg cells were exposed to NAD+ in vitro in the absence or presence of ARTC2.2-inhibiting nanobodies. Although IL-2Rα was mostly not ADP-ribosylated in wild-type Treg cells, 80% of IL-2Rα was ADP-ribosylated in CD38-deficient Treg cells. Notably, wild-type Treg cells were essentially resistant to further ADP-ribosylation induced by NAD+ exposure. By contrast, IL-2Rα became fully ADP-ribosylated through NAD+ in CD38-deficient Treg cells, a process that was prevented by a blocking nanobody to ARTC2.2 (Fig. 8e,f). In line with the idea that ADP-ribosylation of IL-2Rα leads to deficient IL-2 signaling through the high-affinity IL-2R expressed in Treg cells (Fig. 8g), NAD+ exposure reduced STAT5 phosphorylation significantly more in CD38-deficient Treg cells than in wild-type Treg cells in an ARTC2.2-dependent manner (Fig. 8h,i). Finally, IL-2Rα ADP-ribosylation was significantly more pronounced in CD38-deficient Treg cells than in their wild-type counterparts, even when Treg cells of both genotypes were cocultured together in the same dish during NAD+ exposure (Fig. 8j), suggesting that cell-intrinsic expression of CD38 was crucial to achieve low levels of NAD+ in the proximal vicinity of the cell and, thus, prevent ADP-ribosylation of its surface molecules. Together, these data illustrate that Treg cell-autonomous expression of CD38 is essential for their perception of restricted amounts of IL-2, as would occur in ‘stressful’ nonlymphoid environments.
a,b, EAE was induced in Foxp3 (GFP) mice. NAD+ or PBS was administered i.c.v. (100 µg) with or without anti-ARTC2.2 i.p. injection (50 µg) during the recovery phase of EAE (a). Mice were killed 1 day after treatment, and Annexin V and LD Violet staining was performed on isolated CNS Tconv and T cells (b). c,d, Percentage of Annexin V+ (c) and total counts of viable (Annexin V−LD Violet−) Tconv and Treg cells (d). Each symbol in c and d represents an individual mouse. Data are shown as mean + s.d. P values were calculated using a one-way ANOVA followed by a Tukey’s HSD test (n = 4 biological replicates). e, Splenocytes from Cd38+/+ and Cd38−/− mice were preincubated with or without NAD+ (30 µM) and anti-ARTC2.2 (1 µg ml−1). CD4+Foxp3+ Treg cells were analyzed for CD25 ADP-ribosylation using clones 7D4 and PC61.5 of anti-CD25 by flow cytometry. f, Percentage of 7D4+ within PC61.5+ Treg cells. Each symbol represents an individual mouse. Data are shown as mean + s.d. P values were calculated using a two-way ANOVA with a Tukey’s HSD test (n = 3 biological replicates). g, Model for CD38 inhibiting the ADP-ribosylation (ADPR) of CD25: CD38 on Treg cells metabolizes extracellular NAD+, competitively inhibiting ADP-ribosyltransferase (ART) activity, thereby preserving the high-affinity IL-2R conformation and sustaining STAT5 phosphorylation; NAM, nicotinamide. h, As in e, following preincubation, cells were stimulated with mouse IL-2 (1.25 ng ml−1) and analyzed for STAT5 Y694 phosphorylation (pSTAT5 Y694) in Treg cells by flow cytometry. i, Percentage of pSTAT5 Y694+CD4+Foxp3+ Treg cells. Each symbol represents an individual mouse. Data are shown as mean + s.d. P values were calculated by a two-way ANOVA with a Tukey’s HSD test (n = 3 biological replicates). j, Cd38+/+ and Cd38−/− splenocytes were labeled with different anti-CD4 fluorochromes, mixed 1:1 (2.5 × 106 each) in the same well and incubated with a twofold serial titration of NAD+ with or without anti-ARTC2.2 (1 µg ml−1). The percentages of 7D4+ cells within the PC61.5+ Treg cell population are shown as mean + s.d. for the wild-type and CD38-deficient compartments (n = 4 biological replicates); WT, wild-type.
Discussion
In the present study, we show that Foxp3+ Treg cells persist in the CNS for extended periods of time after the contraction of pathogenic effector T cells. These CNS-resident Treg cells are indispensable for maintaining the re-established immune homeostasis in the CNS after an autoimmune inflammatory episode, as their site-specific depletion immediately triggers a relapse. The long-term persistence of Treg cells in the postinflammatory CNS is dependent on the efficient exploitation of limited amounts of IL-2, which selects for antigen-specific Treg cells with high expression of the high-affinity IL-2R CD25. These stress-tolerant Treg cells, at the same time, must express CD38 to cope with enhanced levels of extracellular NAD+, which otherwise dampen Treg cell-intrinsic IL-2 signaling due to ADP-ribosylation of IL-2Rα. Therefore, the expression of the NAD+ hydrolase CD38 critically determines the resilience of CNS Treg cells to the hostile environment of the postinflammatory CNS and, thus, is a hallmark feature of stress-tolerant Treg cells.
CD38 is a multifunctional molecule with diverse roles in various cellular processes. It acts as an NAD+ consuming enzyme, thereby regulating both intracellular and extracellular NAD+ levels35,36. Additionally, CD38 functions as a receptor of PECAM-1 (CD31), influencing leukocyte migration37. Although PECAM-1/CD38 interactions might affect Treg cell migration into nonlymphoid tissues, our data indicate that Treg cells can migrate to the inflamed CNS without CD38. CD38+ Treg cells possess a survival advantage in nonlymphoid tissues. Conversely, CD38-deficient Treg cells fail to reduce extracellular NAD+ concentrations in their micromilieu and, as a consequence, are more susceptible to ARTC2.2/P2X7-mediated apoptosis, which is dependent on extensive Ca2+ influx through the ATP-gated P2X7 receptor ion channel due to ARTC2.2-mediated ADP-ribosylation of P2RX7 (refs. 38,39,40). However, although P2RX7 is similarly expressed in Tconv and Treg cells, the exaggerated susceptibility of Treg (but not Tconv) cells to NAD+-induced cell death in the CNS is likely due to further targets of ADP-ribosylation30, such as IL-2Rα, which is orders of magnitude more highly expressed in Treg cells than in Tconv cells. Here, ARTC2.2-mediated ADP-ribosylation leads to a loss-of-function of high-affinity IL-2Rα with reduced STAT5 activation. CD38 quenches ADP-ribosylation of IL-2Rα through removing membrane-proximal extracellular NAD+ (in a cell-autonomous manner) as a substrate of ARTC2.2. Additionally, CD38 also competes for intracellular NAD+, thereby influencing intracellular NAD+-dependent pathways. Intracellular NAD+ is an essential cofactor for sirtuins, which act as deacetylases of histones and, thus, play a role in gene silencing. Intracellular NAD+ is also a cofactor for poly(ADP-ribosyl) polymerases, which are involved in DNA damage repair. These functions of NAD+-dependent enzymes have been linked to cellular aging41. In the context of T cells, SIRT1 has been demonstrated to negatively regulate Treg cell function by decreasing the expression of Foxp3 and CTLA-4, while simultaneously increasing DNMT3a expression42. The absence of CD38 could lead to excess NAD+ availability, which may enhance SIRT1 activity and dismantle Treg cell stability. However, here, we did not observe a difference in Foxp3 expression in CD38-deficient Treg cells compared to wild-type Treg cells in the same environment. Furthermore, CD38 regulates intracellular Ca²⁺ levels via the cyclic adenosine 5′-diphosphate ribose (cADPR) signaling pathway, which may influence T cell differentiation43. Proper Ca²⁺ influx is critical for Treg cell functionality, and impaired Ca²⁺ signaling has been associated with the defective immunosuppressive function of Treg cells in individuals with multiple sclerosis44. Finally, Treg cells require CD38 for adenosine synthesis via the CD38/CD203a/CD73 ectoenzymatic pathway45. Yet, we observed a physical loss of antigen-specific Treg cells in the CNS when they lacked CD38 expression. Therefore, we suggest that CD38 is primarily necessary for the very maintenance of CNS Treg cells in the chronically inflamed CNS because it enables them to exploit limited sources of IL-2. Prior studies are consistent with the idea that the availability of IL-2 defines the physical and functional niche of Treg cells in nonlymphoid tissues16.
The chronically inflamed CNS is characterized by a high load of extracellular NAD+21. Therefore, we propose that CD38 expression by Treg cells is a key feature that determines their resilience to the ‘environmental’ stress of this niche. In addition, CNS Treg cells share much of what has been described as the tissue-resident Treg cell signature2,46. It is unresolved whether a universal tissue Treg cell signature can be defined or whether each tissue has its own unique Treg cell population, both in terms of TCR repertoire and functional phenotype. Although other investigators have found a comparable Treg cell TCR repertoire diversity in various tissue sites6, these data need to be interpreted with caution as it is virtually impossible to reach the saturation limit in single-cell sequencing of a fully polyclonal repertoire with currently available technologies. Our data would argue in favor of the requirement for a cognate engagement of postinflammatory CNS-resident Treg cells because the ablation of CD38 induces a more profound loss of MOG-specific Treg cells than of Treg cells that do not bind MOG35–55–I-Ab tetramers. Therefore, the availability of tissue-specific private antigens may require antigen-specific Treg cells, whose resilience to the local environment then depends on a universal tissue-type transcriptional program, including CD38.
Finally, local depletion of postinflammatory CNS Treg cells was sufficient to provoke an EAE relapse, whereas systemic Treg cell depletion was not. Postinflammatory Treg cells are, thus, not only adapted to their tissue niche, where they may fulfill nonimmune functions1,15,17,47, but also still absolutely essential to prevent the recurrence of inflammation in a metastable environment. In fact, CNS inflammation is rekindled from niches of persistent immune cells in the inflammation-experienced CNS when Treg cells are ablated. These findings establish a paradigm that could explain the increasing disconnect of immunopathology from the systemic immune compartment in chronic CNS autoimmunity. Defining their local interaction partners and growth, as well as maintenance, factors will open new avenues for manipulating CNS Treg cells as auditors of compartmentalized inflammatory processes that may drive immunopathology in progressive CNS autoimmunity.
Methods
Mice
Foxp3eGFP-Cre-ERT2 (Foxp3Cre-ERT2, IMSR_JAX:016961)48, Foxp3DTR (Foxp3-DTR, IMSR_JAX:016958)49, CD45.1 congenic (IMSR_JAX:002014)50 and B6.129S7-Rag1tm1Mom/J (Rag1−/−, IMSR_JAX:002216)51 mice were obtained from The Jackson Laboratory. C57BL/6J (wild-type, IMSR_JAX:000664) mice were from Charles River Laboratories. Foxp3 (GFP) reporter mice24 were provided by V. Kuchroo (Harvard Medical School and Brigham and Women’s Hospital). Cd38tm1Lnd (Cd38−/−, IMSR_JAX:003727) mice52 were provided by H.-W. Mittrücker (Medical Center Hamburg Eppendorf). Foxp3Cre-ERT2 mice and Rosa-CAG-LSL-tdTomato-WPRE::ΔNeo mice (Rosa26LSL-tdTomato, IMSR_JAX:007914)53 were mated to generate Foxp3Cre-ERT2 × Rosa26LSL-tdTomato mice. PhamT reporter mice (Cd4Cre × Rosa26LSL-mitoDendra2, that is, IMSR_JAX:022071 mated with IMSR_JAX:018385) were generated in-house54. DEREG mice (MMRRC_032049-JAX)27 were provided by T. Sparwasser (University Medical Center Mainz). We used 8- to 20-week-old disease score-matched male and female C57BL/6 mice in all experiments (see Hiltensperger et al.54), except for VAT, where we only used 20-week-old male mice55. Mice were housed in a specific pathogen-free facility with normal rodent chow at the Technical University of Munich. Protocols were approved by the Bavarian state authorities and conducted according to the relevant guidelines (ROB-55.2-2532.Vet_02-17-69, ROB-55.2-2532.Vet_02-17-234, ROB-55.2-2532.Vet_03-18-53, ROB-55.2-2532.Vet_02-20-1, ROB-55.2-2532.Vet_02-20-23, ROB-55.2-2532.Vet_02-20-187, ROB-55.2-2532.Vet_02-23-71, ROB-55.2-2532.Vet_02-23-118 and ROB-55.2-2532.Vet_02-25-68).
EAE model
Mice were immunized with MOG35–55 (200 μg, Auspep) emulsified in complete Freund’s adjuvant (500 mg Mycobacterium tuberculosis H37Ra, BD Difco). Mice received 200 ng or a suboptimal dose of 50 ng pertussis toxin (Sigma-Aldrich) i.v. on days 0 and 2 after immunization. Clinical scores were recorded as 0 (no disease), 1 (tail tone loss), 2 (hind limb paresis), 3 (hind limb paralysis), 4 (tetraplegia) and 5 (moribund). Peak disease occurred on days 12–15, with recovery starting around days 30–35.
BM chimeras
Six- to 8-week-old Rag1−/− mice were sublethally irradiated (7 Gy; two doses of 3.5 Gy, 3 h apart) and i.v. injected with 1 × 10⁷ CD90.2-depleted donor BM cells. Mice received enrofloxacin (0.1 mg ml−1) in their drinking water for 2 weeks after BM transplantation. Hematopoietic reconstitution was assessed using peripheral blood at 6 weeks, and the chimeras were immunized for EAE induction 8 weeks after BM grafting.
Tissue and cell isolation
Mice were killed and perfused with 20–30 ml of cold PBS via the left heart ventricle. Spleen tissue and lymph nodes were homogenized and filtered (70-µm strainers, Falcon), and red blood cells were lysed with 2 ml of FACS Lysing Solution (BD Biosciences) for 5 min. Cells were washed and resuspended in FACS buffer (2% fetal calf serum (FCS) in PBS). For CNS cell isolation, brains were dissected, and spinal cords were flushed out with PBS. CNS tissues were digested in 4 ml of digestion buffer (2.5 mg ml−1 collagenase D (Roche Diagnostics), 1 mg ml−1 DNase I (Roche Diagnostics) and 1 µg ml−1 S+16a anti-ARTC2.2 nanobody (BioLegend) in 10% FCS DMEM) at 37 °C for 45 min with shaking at 120 rpm, filtered and separated via Percoll gradient (70–37%, GE Healthcare). Cells were collected, washed and resuspended in cold FACS buffer. VAT from male mice was collected, digested in 3 ml of digestion buffer at 37 °C for 45 min, washed with 30 ml of FACS buffer and treated with 1 ml of FACS lysing solution for 5 min. Cells were washed, pelleted and resuspended in cold FACS buffer. BM was isolated from the femur and tibia. The bones were cut out from each end and flushed out with cold 10% FCS DMEM with a 21-gauge needle. The cells were collected, and red blood cells were lysed with 1 ml of FACS lysing solution for 5 min. The cells were then washed and resuspended in cold FACS buffer.
Cell sorting
Splenocytes and lymph nodes were pooled to isolate Treg cells. Briefly, CD4+ T cells were enriched from single-cell suspensions using a negative selection kit (Miltenyi Biotec). Cells were stained with antibodies for 15 min at 4 °C in the dark, washed and resuspended in FACS buffer. Sorting was conducted using a FACSAria III Cell Sorter (BD Biosciences), achieving >95% purity.
Flow cytometry
Cell suspensions were stained with Live/Dead Aqua, Live/Dead violet or Near-IR dyes (Thermo Fisher) for 15 min at 4 °C in PBS and washed with FACS buffer. Cells were incubated with an antibody cocktail targeting surface antigen and CD16/CD32 Fc block in FACS buffer for 15 min at 4 °C in the dark, washed twice and resuspended in FACS buffer. For Annexin V staining, cells were stained with an antibody cocktail in Annexin V binding buffer (BioLegend). For intracellular staining, cells were stained with antibodies to surface antigens, fixed with 4% paraformaldehyde (PFA) for 15 min at room temperature (protected from light) and washed with FACS buffer. Fixation and permeabilization were performed using a eBioscience Intracellular Fixation and Permeabilization Buffer Set (Invitrogen). Cells were incubated overnight at 4 °C with an intracellular antibody cocktail and Fc block in permeabilization buffer, washed and resuspended in FACS buffer.
For pSTAT5 Y694 staining, fixation and permeabilization were performed using a Transcription Factor Phospho Buffer Set (TFP, BD Pharmingen), following the manufacturer’s protocol. In short, after fixation and permeabilization, cells were incubated with anti-pSTAT5 Y694, anti-Foxp3 and anti-surface antigen with Fc block in TFP Perm/Wash Buffer for 50 min at 4 °C. Cells were washed twice with TFP Perm/Wash buffer and resuspended in FACS buffer for flow cytometry analysis. A list of the applied antibodies and resources is provided in Supplementary Table 4. Antibody dilutions are detailed in the Reporting Summary. Data were collected using CytoFLEX (Beckman Coulter), FACSAria III (BD Biosciences) or Aurora (Cytek Biosciences) and analyzed using FlowJo software (version 10.8.1, BD Biosciences).
Treg cell cluster visualizations were based on the data acquired on the Aurora system. Treg cells were identified as Aqua−Foxp3+CD4+ through primary gating in each sample in FlowJo. To standardize cell numbers across tissues, downsampling was conducted by pooling and selecting 300 Treg cells, which were then subjected to secondary FlowSOM (v3.0.18) clustering. Further subclustering of Treg cells was achieved using ClusterExplorer (v1.6.5). The clusters were visualized with uniform manifold approximation and projection through R and the FlowJo plugin packages.
I-Ab tetramer staining
CNS and spleen cells were treated with neuraminidase (0.35 U ml−1, Sigma-Aldrich) and dasatinib (100 nM, Selleckchem) in serum-free DMEM at 37 °C with 5% CO2 for 25 min. Cells were then washed twice with FACS buffer, blocked with anti-mouse CD16/CD32 Fc block on ice for 15 min and stained with T-Select I-Ab MOG35–55 Tetramer-PE (Medical and Biological Laboratories, TS-M704-1) or the control T-Select I-Ab OVA323–339 Tetramer-PE (TS-M710-1) at room temperature for 2 h, with gentle resuspension every 30 min. After tetramer staining, cells were centrifuged (430g, 4 °C, 5 min), washed twice with FACS buffer and incubated with anti-phycoerythrin MicroBeads (Miltenyi Biotec) on a 4 °C shaker for 30 min to stabilize tetramer binding. The cells were washed with FACS buffer and subjected to surface and intracellular staining for flow cytometry analysis. The tetramer+ population was identified by gating on NIR−, Lin (CD19, CD8a, F4/80, Ly6G)− and MOG35–55 tetramer–PE+ cells, with the OVA323–339 tetramer as a negative control.
Bulk RNA-seq
Approximately 2,000 or 250 Treg or Tconv cells (viability > 90%) were sorted directly into TCL buffer (Qiagen) plus 1% 2-mercaptoethanol (Merk) and stored at −80 °C. RNA was extracted and purified using an RNeasy Plus Micro kit (Qiagen). RNA quality was assessed using an Agilent 2100 Bioanalyzer (Agilent Technologies). Library preparation followed a previously described protocol56. Barcode-specific cDNA was synthesized via reverse transcription with Maxima RT polymerase (Thermo Fisher) using an oligo(dT) primer containing barcodes, unique molecular identifiers (UMIs) and adaptor sequences. The 5′ end of the cDNA was then extended using primers complementary to the template switch oligonucleotide site, followed by amplification of full-length cDNA. The cDNA was fragmented using a Nextera XT kit (Illumina), and the 3′-end fragments were amplified with primers incorporating Illumina P5 and P7 overhangs. The P5 and P7 sites were modified to optimize cluster recognition during sequencing, allowing for sequencing of the cDNA in read 1 and the barcodes/UMIs in read 2.
Sequencing was performed on an Illumina NextSeq 500 platform with 75 cycles for read 1 and 16 for read 2. The sequencing data were processed using the Drop-seq pipeline57 to generate UMI tables organized by sample and gene. The GRCm38 reference genome was aligned with transcript and gene annotations derived from ENSEMBL release 75.
Differential gene expression analysis was conducted in R (v4.5.0) using the DESeq2 package (v1.48.1)58. Significant DEGs were defined as genes with a Benjamini–Hochberg-adjusted P value of <0.05 (in some cases 0.1 as indicated) and an absolute fold change in expression of >1.5. A log2 (fold shrinkage) estimation was performed using the R package glmGamPoi (v1.20.0)59. GO term annotations were derived from the R package org.Mm.eg.db (v3.21.0). For visualization, count data were transformed using the variance stabilizing transformation (‘vst’) function in DESeq2. GSEA was performed on DESeq2 normalized counts using the Broad Institute GSEA Desktop Application (v4.3.2) in conjunction with the Molecular Signatures Database (v2024.1.Mm), with permutation type ‘gene_set’ and an FDR threshold of 0.25 applied. GO enrichment analysis of adjusted P value ranked DEGs was performed using the gprofiler2 R package60.
In vivo cell proliferation assay
EAE was induced in C57BL/6 wild-type mice. At various time points during EAE progression, EdU (0.05 μg mg (body weight), Click-iT EdU, Sigma-Aldrich) was administered i.p. 18 h before euthanasia. Circulating cells were labeled with anti-CD45.2 BV650 (3 μg, BioLegend) via i.v. injection 3–5 min before euthanasia. EdU staining was performed according to the manufacturer’s protocol. Cells from the CNS and spleen were stained with surface antigens and fixed with 4% PFA (Thermo Fisher) at room temperature for 15 min in the dark. After two washes with FACS buffer, cells were incubated in a saponin-based fixation-permeabilization buffer at room temperature for 20 min, followed by centrifugation. A fresh EdU detection cocktail was prepared, and cells were incubated at room temperature for 30 min. After two washes with the saponin-based fixation-permeabilization buffer, cells were stained with anti-Foxp3 in the same buffer overnight at 4 °C. Cells were washed twice again, resuspended in FACS buffer and analyzed by flow cytometry.
In vivo T cell trafficking assay
Mice were anesthetized with a cocktail of medetomidine, midazolam and fentanyl, and the fur was removed using depilatory cream (Veet). For photolabeling, the inguinal lymph node on the immunization side was irradiated for 20 min using a 405-nm diode laser (LuxX 405-60, Soliton) as previously described54. Two days after photoconversion, the mice were killed, and cells from the CNS and spleen were collected to assess the migration of photolabelled lymphocytes.
In vivo Foxp3 fate mapping
For tamoxifen preparation, tamoxifen (Sigma-Aldrich) was dissolved in 100% ethanol (Merck) at a concentration of 1 g ml−1, vortexed at 37 °C for 15 min and mixed with Miglyol (Caelo) to achieve a final concentration of 10 mg ml−1. The solution was incubated and vortexed at 37 °C for 30 min to ensure complete dissolution. We followed the protocol by Jahn et al.61 for tamoxifen treatment. Foxp3Cre-ERT2 × Rosa26LSL-tdTomato mice were given daily i.p. injections of 1 mg 100 μl−1 tamoxifen for 5 days. Fourteen days after the last injection, the mice were killed, and their spleen and CNS cells were collected for either flow cytometry or RNA-seq analysis.
In vivo Treg cell depletion
EAE was induced in Foxp3-DTR and control Foxp3 (GFP) mice. During the EAE recovery phase (between 32 and 35 days after immunization, with a clinical score of less than 2), DTx (from Corynebacterium diphtheriae, Sigma-Aldrich) or PBS was administered either i.c.v. (15 ng) or i.p. (500 ng). The i.c.v. injection was performed according to the protocol described by Taylor et al.62, modified to administer 3 μl of DTx at a rate of 1 μl min−1. Buprenorphine (0.1 μg per mg (body weight)) was given before, after and 1 day after surgery for pain management. Treg cell depletion efficiency was confirmed by flow cytometry, and clinical scores were monitored daily from day 0 to day 3 after DTx injection. For FTY720 treatment, FTY720 (1 μg per mg (body weight), Sigma-Aldrich) or vehicle control was administered i.p. starting 1 day before the DTx i.c.v. injection and continuing daily until 1 day after surgery. Clinical scores were recorded from day 0 to day 3.
In vivo Treg cell suppression
EAE was induced in both donor (either Cd38+/+ or Cd38−/−, CD45.2) and recipient DEREG (CD45.1) mice. DTx (1 μg, 50 μg per kg (body weight)) was administered i.p. on days 9 and 10 after induction to deplete Treg cells in recipients. On day 11, donors were pretreated i.p. with 50 µg of S+16a (anti-ARTC2.2 nanobody, BioLegend) 30 min before euthanasia to prevent cell apoptosis. Donor splenic Treg cells (LIVE/DEAD Aqua⁻CD4⁺CD25 (clone 7D4)⁺GITR⁺CD45.2⁺) were sorted, and 105 Treg cells were transferred i.v. into recipients.
In vivo NAD+ i.c.v. injection
EAE was induced in Foxp3 (GFP) mice. During the recovery phase, mice received an i.c.v. injection of either freshly prepared NAD⁺ (Thermo Fisher; 100 μg in 3 μl of cold PBS) or vehicle control (3 μl of PBS). Buprenorphine (0.1 μg per mg (body weight)) was administered before and after surgery. Mice were killed 24 h after injection.
In vitro NAD+ incubation and IL-2 stimulation
Splenocytes (5 × 10⁶) from Cd38+/+ or Cd38−/− mice were incubated with or without NAD⁺ (concentrations are specified in the figure legends) and anti-ARTC2.2 (1 µg ml−1) in serum-free DMEM at 4 °C for 15 min. Cells were washed with cold FACS buffer, stained with antibodies and analyzed by flow cytometry. For the pSTAT5 Y694 assay, following the initial incubation, cells were stimulated with mouse IL-2 (Miltenyi Biotec, 1.25 ng ml−1) in DMEM containing 2% BSA at 37 °C for 10 min. Cells were immediately fixed and stained for anti-pSTAT5 Y694.
Histological analysis
Mice were killed and perfused with 20–30 ml of cold PBS via the left ventricle. Brains were dissected and fixed in 4% PFA at 4 °C for 48 h, washed with PBS and paraffin embedded. Representative 2-μm sections were cut with a rotary microtome, and immunohistochemistry was conducted using a Leica Bond Rxm autostainer with a Bond Polymer Refine Detection kit (DS9800, Leica Biosystems). Heat-induced epitope retrieval was performed by using citrate-based pH 6 Bond Epitope Retrieval Solution 1 (AR9961, Leica Biosystems). Antibody details are listed in Supplementary Table 4. Slides were then scanned with a Leica Aperio AT2 system and analyzed in QuPath (v0.6.0) using automated positive cell detection.
To generate multiplex immunostainings, 2-µm sections were deparaffinized and subjected to antigen retrieval (citrate buffer pH 6.0). To prevent myelin autofluorescence, slides were treated with 4.5% H2O2 and 20 mM NaOH in PBS under a light-emitting diode for 45 min twice at room temperature. Nonspecific binding was blocked with 2.5% goat serum in PBS, and samples were incubated overnight with rabbit anti-CD4 (Cell Signaling). Signals were detected with anti-rabbit horseradish peroxidase (Dako, K4003) with amplification (TSA vivid 520, Tocris, 7534). Following denaturation (Discovery Roche, CC2), slides were blocked with Fab fragment anti-mouse IgG (Jackson Immunoresearch) and 2.5% goat serum in PBS and incubated with rat anti-Foxp3 (eBioscience) overnight. Signals were detected with anti-rat horseradish peroxidase (Vectorlab) with amplification (TSA vivid 570, Tocris, 7535). Nuclei were stained with DAPI (Invitrogen), and slides were mounted in Fluoromount aqueous mounting medium (Sigma-Aldrich, F4680). Immunostained sections were scanned using an Akoya PhenoImager HT 2.0 (Akoya Biosciences).
For cryosections, brains were fixed in 4% PFA at 4 °C for 48 h and subjected to a 30% sucrose gradient for 48 h. Ten-micron cryosections (CM3050 S, Leica) were subjected to immunofluorescence stainings. The antibody details are listed in Supplementary Table 4. Images were acquired on an SP8 confocal microscope (Leica) and analyzed using LAS X software (v.3.5.6.21594, Leica).
Statistical analysis
Statistical analyses were performed using Prism 10 (GraphPad) or R (v4.4.0). No statistical methods were used to predetermine sample sizes, but our sample sizes were similar to those reported in previous publications54. Data distribution was assumed to be normal, but this was not always formally tested. In any case, individual data points are provided in the figures. Significance of the differences between two groups was evaluated using two-tailed Student’s t-tests. Differences among three or more groups were analyzed using a one-way or two-way ANOVA with a Tukey’s HSD post hoc test, and results are presented as mean values + s.d. Adjusted P values from DEGs were derived from DESeq2 using a two-tailed Wald’s test with correction for multiple testing using the Benjamini and Hochberg method. Unless stated otherwise in the figure legend, animals were assigned to experimental groups in an age- and sex-matched manner. Scoring of EAE was performed in a manner blinded to the experimental conditions. Unless otherwise stated in the figure legends, all animals were included in the analysis. The significance of differences in clinical scores was determined using two-way ANOVA with a Tukey’s post hoc test, and the results are presented as mean values + s.e.m. The significance of the survival rate was analyzed with a Mantel–Cox test. The individual sample sizes and statistical methods are provided in the figure legends. The individual P or adjusted P values are indicated in the figures.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequencing data have been deposited in the Gene Expression Omnibus database under accession code GSE311963. All other data are present in the article and Supplementary Information. Source data are provided with this paper.
References
Feuerer, M. et al. Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nat. Med. 15, 930–939 (2009).
Panduro, M., Benoist, C. & Mathis, D. Tissue Tregs. Annu. Rev. Immunol. 34, 609–633 (2016).
Russler-Germain, E. V., Rengarajan, S. & Hsieh, C. S. Antigen-specific regulatory T-cell responses to intestinal microbiota. Mucosal Immunol. 10, 1375–1386 (2017).
Li, C. et al. TCR transgenic mice reveal stepwise, multi-site acquisition of the distinctive fat-Treg phenotype. Cell 174, 285–299 (2018).
Korn, T. & Muschaweckh, A. Stability and maintenance of Foxp3+ Treg cells in non-lymphoid microenvironments. Front. Immunol. 10, 2634 (2019).
Burton, O. T. et al. The tissue-resident regulatory T cell pool is shaped by transient multi-tissue migration and a conserved residency program. Immunity 57, 1586–1602 (2024).
Delacher, M. et al. Single-cell chromatin accessibility landscape identifies tissue repair program in human regulatory T cells. Immunity 54, 702–720 (2021).
DiSpirito, J.R. et al. Molecular diversification of regulatory T cells in nonlymphoid tissues. Sci. Immunol. 3, eaat5861 (2018).
Fan, X. et al. CD49b defines functionally mature Treg cells that survey skin and vascular tissues. J. Exp. Med. 215, 2796–2814 (2018).
Abbas, A. K., Trotta, E., Dimitre, R. S., Marson, A. & Bluestone, J. A. Revisiting IL-2: biology and therapeutic prospects. 3, eaat1482 (2018)
Korn, T. et al. Myelin-specific regulatory T cells accumulate in the CNS but fail to control autoimmune inflammation. Nat. Med. 13, 423–431 (2007).
Cretney, E. et al. The transcription factors Blimp-1 and IRF4 jointly control the differentiation and function of effector regulatory T cells. Nat. Immunol. 12, 304–311 (2011).
Garg, G. et al. Blimp1 prevents methylation of Foxp3 and loss of regulatory T cell identity at sites of inflammation. Cell Rep. 26, 1854–1868 (2019).
Delacher, M. et al. Precursors for nonlymphoid-tissue Treg cells reside in secondary lymphoid organs and are programmed by the transcription factor BATF. Immunity 52, 295–312 (2020).
Ito, M. et al. Brain regulatory T cells suppress astrogliosis and potentiate neurological recovery. Nature 565, 246–250 (2019).
Yshii, L. et al. Astrocyte-targeted gene delivery of interleukin 2 specifically increases brain-resident regulatory T cell numbers and protects against pathological neuroinflammation. Nat. Immunol. 23, 878–891 (2022).
Dombrowski, Y. et al. Regulatory T cells promote myelin regeneration in the central nervous system. Nat. Neurosci. 20, 674–680 (2017).
Shi, L. et al. Treg cell-derived osteopontin promotes microglia-mediated white matter repair after ischemic stroke. Immunity 54, 1527–1542 (2021).
Marin-Rodero, M. et al. The meninges host a distinct compartment of regulatory T cells that preserves brain homeostasis. Sci. Immunol. 10, eadu2910 (2025).
Midavaine, E. et al. Meningeal regulatory T cells inhibit nociception in female mice. Science 388, 96–104 (2025).
Chiarugi, A. Inhibitors of poly(ADP-ribose) polymerase-1 suppress transcriptional activation in lymphocytes and ameliorate autoimmune encephalomyelitis in rats. Br. J. Pharmacol. 137, 761–770 (2002).
Aswad, F., Kawamura, H. & Dennert, G. High sensitivity of CD4+CD25+ regulatory T cells to extracellular metabolites nicotinamide adenine dinucleotide and ATP: a role for P2X7 receptors. J. Immunol. 175, 3075–3083 (2005).
Seman, M. et al. NAD-induced T cell death. Immunity 19, 571–582 (2003).
Bettelli, E. et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 441, 235–238 (2006).
Kim, E. H. et al. Bach2 regulates homeostasis of Foxp3+ regulatory T cells and protects against fatal lung disease in mice. J. Immunol. 192, 985–995 (2014).
Osman, A. et al. TCF-1 controls Treg cell functions that regulate inflammation, CD8+ T cell cytotoxicity and severity of colon cancer. Nat. Immunol. 22, 1152–1162 (2021).
Lahl, K. et al. Selective depletion of Foxp3+ regulatory T cells induces a scurfy-like disease. J. Exp. Med. 204, 57–63 (2007).
Wakim, L. M., Woodward-Davis, A. & Bevan, M. J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. Proc. Natl. Acad. Sci. USA 107, 17872–17879 (2010).
Zeidler, J. D. et al. Endogenous metabolism in endothelial and immune cells generates most of the tissue vitamin B3 (nicotinamide). iScience 25, 105431 (2022).
Leutert, M. et al. Identification of the mouse T cell ADP-ribosylome uncovers ARTC2.2 mediated regulation of CD73 by ADP-ribosylation. Front. Immunol. 12, 703719 (2021).
Stephens, L. A., Malpass, K. H. & Anderton, S. M. Curing CNS autoimmune disease with myelin-reactive Foxp3+ Treg. Eur. J. Immunol. 39, 1108–1117 (2009).
Sabatino, J. J. Jr., Huang, J., Zhu, C. & Evavold, B. D. High prevalence of low affinity peptide–MHC II tetramer-negative effectors during polyclonal CD4+ T cell responses. J. Exp. Med. 208, 81–90 (2011).
Georgiev, H. et al. Blocking the ART2.2/P2X7-system is essential to avoid a detrimental bias in functional CD4 T cell studies. Eur. J. Immunol. 48, 1078–1081 (2018).
Teege, S. et al. Tuning IL-2 signaling by ADP-ribosylation of CD25. Sci. Rep. 5, 8959 (2015).
Shubinsky, G. & Schlesinger, M. The CD38 lymphocyte differentiation marker: new insight into its ectoenzymatic activity and its role as a signal transducer. Immunity 7, 315–324 (1997).
Chini, C. C. S., Zeidler, J. D., Kashyap, S., Warner, G. & Chini, E. N. Evolving concepts in NAD+ metabolism. Cell Metab. 33, 1076–1087 (2021).
Deaglio, S. et al. Human CD38 (ADP-ribosyl cyclase) is a counter-receptor of CD31, an Ig superfamily member. J. Immunol. 160, 395–402 (1998).
Krebs, C. et al. CD38 controls ADP-ribosyltransferase-2-catalyzed ADP-ribosylation of T cell surface proteins. J. Immunol. 174, 3298–3305 (2005).
Hubert, S. et al. Extracellular NAD+ shapes the Foxp3+ regulatory T cell compartment through the ART2–P2X7 pathway. J. Exp. Med. 207, 2561–2568 (2010).
Andrejew, R. et al. The P2X7 receptor: central hub of brain diseases. Front. Mol. Neurosci. 13, 124 (2020).
Covarrubias, A. J., Perrone, R., Grozio, A. & Verdin, E. NAD+ metabolism and its roles in cellular processes during ageing. Nat. Rev. Mol. Cell Biol. 22, 119–141 (2021).
Beier, U. H., Akimova, T., Liu, Y., Wang, L. & Hancock, W. W. Histone/protein deacetylases control Foxp3 expression and the heat shock response of T-regulatory cells. Curr. Opin. Immunol. 23, 670–678 (2011).
Huang, X. & Rudensky, A. Y. Regulatory T cells in the context: deciphering the dynamic interplay with the tissue environment. Curr. Opin. Immunol. 89, 102453 (2024).
Schwarz, A. et al. Fine-tuning of regulatory T cell function: the role of calcium signals and naive regulatory T cells for regulatory T cell deficiency in multiple sclerosis. J. Immunol. 190, 4965–4970 (2013).
Horenstein, A. L. et al. A CD38/CD203a/CD73 ectoenzymatic pathway independent of CD39 drives a novel adenosinergic loop in human T lymphocytes. Oncoimmunology 2, e26246 (2013).
Burzyn, D., Benoist, C. & Mathis, D. Regulatory T cells in nonlymphoid tissues. Nat. Immunol. 14, 1007–1013 (2013).
Vasanthakumar, A. et al. The transcriptional regulators IRF4, BATF and IL-33 orchestrate development and maintenance of adipose tissue-resident regulatory T cells. Nat. Immunol. 16, 276–285 (2015).
Rubtsov, Y. P. et al. Stability of the regulatory T cell lineage in vivo. Science 329, 1667–1671 (2010).
Kim, J. M., Rasmussen, J. P. & Rudensky, A. Y. Regulatory T cells prevent catastrophic autoimmunity throughout the lifespan of mice. Nat. Immunol. 8, 191–197 (2007).
Shen, F. W. et al. Cloning of Ly-5 cDNA. Proc. Natl Acad. Sci. USA 82, 7360–7363 (1985).
Mombaerts, P. et al. RAG-1-deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Cockayne, D. A. et al. Mice deficient for the ecto-nicotinamide adenine dinucleotide glycohydrolase CD38 exhibit altered humoral immune responses. Blood 92, 1324–1333 (1998).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Hiltensperger, M. et al. Skin and gut imprinted helper T cell subsets exhibit distinct functional phenotypes in central nervous system autoimmunity. Nat. Immunol. 22, 880–892 (2021).
Vasanthakumar, A. et al. Sex-specific adipose tissue imprinting of regulatory T cells. Nature 579, 581–585 (2020).
Parekh, S., Ziegenhain, C., Vieth, B., Enard, W. & Hellmann, I. The impact of amplification on differential expression analyses by RNA-seq. Sci. Rep. 6, 25533 (2016).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ahlmann-Eltze, C. & Huber, W. glmGamPoi: fitting γ-Poisson generalized linear models on single cell count data. Bioinformatics 36, 5701–5702 (2021).
Kolberg, L., Raudvere, U., Kuzmin, I., Vilo, J. & Peterson, H. gprofiler2—an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Res 9, ELIXIR-709 (2020).
Jahn, H. M. et al. Refined protocols of tamoxifen injection for inducible DNA recombination in mouse astroglia. Sci. Rep. 8, 5913 (2018).
Taylor, Z. V., Khand, B., Porgador, A., Monsonego, A. & Eremenko, E. An optimized intracerebroventricular injection of CD4+ T cells into mice. STAR Protoc. 2, 100725 (2021).
Acknowledgements
We are grateful to H.-W. Mittrücker (Universitätsklinikum Hamburg Eppendorf) for generously providing CD38-deficient mice. We would like to thank K. Schumann (University of Konstanz) and M. Schmidt-Supprian (Technical University of Munich School of Medicine) for fruitful discussions. We appreciate B. Lunk, M. Schmidt and V. Husterer (all Technical University of Munich School of Medicine) for their assistance with mouse colony management. We thank the Technical University of Munich Comparative Experimental Pathology Unit for expert support with histological analyses. This study was supported by the Deutsche Forschungsgemeinschaft (TRR274 (ID 408885537) to T.K., TRR355 (ID 490846870) to T.K. and A.M., GRK2668 (ID 435874434) to T.K. and EXC 2145 (SyNergy, ID 390857198) to T.K.) and by the Hertie Network of Clinical Neuroscience (to T.K.).
Funding
Open access funding provided by Technische Universität München.
Author information
Authors and Affiliations
Contributions
Conceptualization: H.-H.C., A.M. and T.K. Methodology: H.-H.C., S.T., D.R.N., R.K., T.G., I.W., H.D.M., I.B.-P., R.Ö., A.M.A., S.H., L.C.R., C.S., G.L., L.S., K.S., D.M., R.R., G.S. and B.S. Investigation: H.-H.C., S.T., D.R.N., R.K., T.G., I.W., H.D.M., A.M.A., S.H., L.C.R. and L.S. Visualization: H.-H.C., A.M. and T.K. Funding acquisition: A.M. and T.K. Supervision: A.M. and T.K. Writing, original draft: H.-H.C., A.M. and T.K. Writing, review and editing: A.M. and T.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Adrian Liston, Scott Zamvil and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Nick Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Stress-tolerant Treg cells populate distinct niches in the chronically inflamed CNS.
EAE was induced in Foxp3 (GFP) mice, and on day 45 after MOG35-55 immunization, CNS tissue was collected, and serial sections of the brain were analyzed for the presence of CD4+Foxp3+ T cells (a). Representative coronal section, scale bar, overview 500 μm; scale bars, high power magnifications 50 μm. (b, c) Absolute number of Treg cells per coronal section in relation to the distance from the bregma, around the third ventricle (b) or the meningeal space (c). (d) Visualization of the Treg cell hotspot in the third ventricle area in coronal mouse brain atlas schemes (according to Paxinos and Franklin, The mouse brain in stereotaxic coordinates). (e) Quantification of Foxp3+ Treg cells as a fraction of all CD4+ T cells in the third ventricle area and the meningeal space. (b, c, e) Sample size: n = 3 biological replicates.
Extended Data Fig. 2 Transcriptomic analysis of stable and unstable CNS Treg cells in chronic EAE.
In vivo Foxp3 fate mapping was conducted in Foxp3Cre-ERT2 x Rosa26LSL-tdTomato mice with MOG-induced EAE. Mice were injected with tamoxifen (i.p.) for 5 days at the EAE recovery stage. 14 days after the last tamoxifen injection, CNS cells were harvested and analyzed by flow cytometry. (a) Stable Treg cells (Foxp3 (GFP)+tdTomato+) and unstable Treg cells (Foxp3 (GFP)−tdTomato+) were sorted and analyzed by bulk RNA sequencing (n = 5 biological replicates). Volcano plots showing only genes with annotations to the GO terms ‘GO:0009986 cell surface’ and ‘GO:0003700 DNA binding transcription factor activity’ that exhibit differential expression (≥1.5-fold, adjusted P value < 0.05) between stable and unstable Treg cells. DEGs are highlighted in orange. (b) Heatmap showing the top 50 DEGs (≥1.5-fold, adjusted P value < 0.05) between stable and unstable Treg cells. Adjusted P values are derived from DESeq2 with a two-tailed Wald’s test and correction for multiple testing using the Benjamini and Hochberg method. (c) Gene set enrichment analysis of MSigDB Hallmark gene sets comparing stable and unstable CNS Treg cells.
Extended Data Fig. 3 Treg cells maintain immune homeostasis in the chronically inflamed CNS.
(a) EAE was induced in Foxp3 (GFP) and Foxp3-DTR mice. Diphtheria toxin (DTx) was administered intracerebroventricularly (i.c., 15 ng) or intraperitoneally (i.p., 500 ng) during the recovery phase of EAE. Mean EAE scores + SEM (DTx i.c. ctrl n = 3, DTx i.p. n = 5, DTx i.c. n = 6 biological replicates). (b) Survival of mice over 10 days following DTx injection. Survival curves were analyzed using the Mantel–Cox test (DTx i.c. ctrl n = 6, DTx i.p. n = 3, DTx i.c. n = 10 biological replicates), and corresponding individual EAE scores of mice are plotted in (c).
Extended Data Fig. 4 Transcriptomic analysis of spleen and CNS Treg cells in EAE peak and recovery stages.
Volcano plots displaying differentially expressed genes (≥1.5-fold, adjusted P value < 0.05) between recovery (Rec) and peak CNS Treg cells (a), peak CNS and peak spleen (SPL) Treg cells (b), and recovery CNS and SPL Treg cells (c). Adjusted P values in (a-c) are derived from DESeq2 using a two-tailed Wald’s test with correction for multiple testing using the Benjamini and Hochberg method (peak n = 4, recovery n = 5 biological replicates). Labeled genes comprise a selection of genes commonly associated with non-nonlymphoid tissue Treg cells (black: not significantly regulated, orange: DEG, light grey: not significant and not part of the selection of tissue Treg cell genes). (d) Summary of gene set enrichment analyses comparing recovery and peak CNS Treg cells using the MSigDb Hallmark gene set collection. (e, f) Enrichment plot of genes part of the ‘TNFα-signaling via NFκB’ (e) and the ‘IL-2 STAT-5 signaling’ signature (FDR, false discovery rate; NES, normalized enrichment score) (f). (g-h) Heatmap and hierarchical clustering of significantly upregulated genes (≥1.5-fold, adjusted P value < 0.05) that contribute to the leading-edge genes for the gene sets ‘TNFα-signaling via NFκB’ (g) and ‘IL-2 STAT-5 signaling’ (h) in recovery (Rec) vs. peak CNS Treg cells.
Extended Data Fig. 5 Transcriptomic analysis of VAT and naive CNS Treg cells.
(a) Volcano plot of differentially expressed genes (≥1.5-fold, adjusted P value < 0.1) of recovery (Rec) CNS Treg cells vs. VAT Treg cells. Labeled genes comprise a selection of genes commonly associated with a non-nonlymphoid tissue Treg cell phenotype (black: not significantly regulated, orange: selected DEG. Adjusted P values are derived from DESeq2 using a two-tailed Wald’s test with correction for multiple testing using the Benjamini and Hochberg method (VAT Treg cells n = 3, recovery CNS Treg cells n = 4 biological replicates) (b) Summary of gene set enrichment analyses comparing recovery (Rec) CNS Treg cells from EAE mice and naive CNS Treg cells using the MSigDb Hallmark gene set collection (n = 4 biological replicates). (c, d) Heatmap and hierarchical clustering of genes that contribute to the leading-edge genes for the gene sets ‘IL-2 STAT5 signaling’ (c) and ‘Oxidative phosphorylation’ (d) in Rec vs. naive CNS Treg cells.
Extended Data Fig. 6 EAE in CD38-deficient mice and MBMC mice.
(a) EAE was induced in wild-type (Cd38+/+), Cd38+/−, and Cd38−/− mice. Mean EAE scores + SEM. P values were calculated using a two-way ANOVA with Tukey’s post-test (Cd38+/+ n = 6, Cd38+/− n = 4, Cd38−/− n = 5 biological replicates). (b) Bone marrow from Foxp3-DTR mice (CD45.1/2) mixed 1:1 with either Cd38+/+ (CD45.2) or Cd38−/− bone marrow (CD45.2) was transferred into sublethally irradiated Rag1−/− recipients. EAE was induced in bone marrow-reconstituted mice. Diphtheria toxin (DTx) was administered intracerebroventricularly (i.c., 15 ng) in stable EAE mice on 28 days post MOG35-55-immunization. Mean EAE scores + SEM (Cd38+/+ MBMC n = 4, Cd38−/− MBMC n = 5 biological replicates).
Extended Data Fig. 7 Systemic Tconv and Treg cells are not affected by the injection of NAD+ into the lateral ventricle.
EAE was induced in Foxp3 (GFP) mice. NAD+ or PBS was administered intracerebroventricularly (i.c., 100 µg) with or without anti-ARTC2.2 injection (i.p., 50 µg) during the recovery stage of EAE. Mice were euthanized 1 day after treatment. Spleen (SPL) cells were isolated, and Annexin V and LD violet staining were performed on Tconv (CD4+Foxp3−) and Treg cells (CD4+Foxp3+), followed by flow cytometric analysis (a). (b) Percentage of Annexin V+ and (c) total counts of viable (Annexin V−LD violet−) splenic Tconv and Treg cells. Symbols in (b, c) represent individual mice (n = 4 biological replicates).
Extended Data Fig. 8 Antigen-specific Tconv and Treg cells express higher levels of CD25.
EAE was induced in Foxp3 (GFP) mice, and CNS cells were collected during the EAE recovery stage. (a) The percentage of CD25 expression was determined in MOG-I-Ab tetramer-binding Tconv and Treg cells. An OVA323-339-I-Ab tetramer was used as a control tetramer. CD25 expression (MFI) in MOG-I-Ab− vs. MOG-I-Ab+ Tconv (b) and Treg cells (c). Symbols in (b, c) represent individual mice. Data are means + s.d. P values were calculated using a paired two-tailed Student’s t-test (n = 8 biological replicates).
Supplementary information
Source data
Source Data Fig. 1 (download XLSX )
Numerical data (cell numbers, cell fractions and EAE scores).
Source Data Fig. 2 (download XLSX )
Numerical data (cell numbers, cell fractions and EAE scores).
Source Data Fig. 3 (download XLSX )
Numerical data (cell numbers, cell fractions and EAE scores).
Source Data Fig. 6 (download XLSX )
Numerical data (cell numbers, cell fractions and EAE scores).
Source Data Fig. 7 (download XLSX )
Numerical data (cell numbers and cell fractions).
Source Data Fig. 8 (download XLSX )
Numerical data (cell numbers and cell fractions).
Source Data Extended Data Fig. 1 (download XLSX )
Numerical data (cell numbers and cell fractions).
Source Data Extended Data Fig. 3 (download XLSX )
Numerical data (EAE scores).
Source Data Extended Data Fig. 6 (download XLSX )
Numerical data (EAE scores).
Source Data Extended Data Fig. 7 (download XLSX )
Numerical data (cell numbers and cell fractions).
Source Data Extended Data Fig. 8 (download XLSX )
Numerical data (cell numbers and cell fractions).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, HH., Tyystjärvi, S., Ruiz Navarro, D. et al. CD38 endows local antigen-specific Treg cells with stress resilience for control of compartmentalized CNS inflammation. Nat Immunol 27, 516–529 (2026). https://doi.org/10.1038/s41590-025-02416-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02416-z