Abstract
Microglia survey and regulate central nervous system myelination during embryonic development and adult homeostasis. However, whether microglia–myelin interactions are spatiotemporally regulated remains unexplored. Here, by examining spinal cord white matter tracts in mice, we determined that myelin degeneration was particularly prominent in the dorsal column (DC) during normal aging. This was accompanied by molecular and functional changes in DC microglia as well as an upregulation of transforming growth factor beta (TGF)β signaling. Disrupting TGFβ signaling in microglia led to unrestrained microglial responses and myelin loss in the DC, accompanied by neurological deficits exacerbated with aging. Single-nucleus RNA-sequencing analyses revealed the emergence of a TGFβ signaling-sensitive microglial subset and a disease-associated oligodendrocyte subset, both of which were spatially restricted to the DC. We further discovered that microglia rely on a TGFβ autocrine mechanism to prevent damage of myelin in the DC. These findings demonstrate that TGFβ signaling is crucial for maintaining microglial resilience to myelin degeneration in the DC during aging. This highlights a previously unresolved checkpoint mechanism of TGFβ signaling with regional specificity and spatially restricted microglia–oligodendrocyte interactions.
Similar content being viewed by others
Main
Microglia, the resident immune cells of the central nervous system (CNS), are pivotal for maintaining CNS homeostasis and are actively involved in various neurobiological processes. Recent studies reveal phenotypical heterogeneity of microglia across CNS regions1. This diversity in microglia is understood to be the consequence of distinct microenvironmental niches, such as locally secreted molecules and cell–cell interactions. Despite recent findings suggesting distinct characteristics of microglia between the white and gray matter (GM) tracts, understanding of spatially resolved microglial heterogeneity is still in its infancy, with less known about the spinal cord compared to the brain2,3,4,5. Microglia can interact with and regulate the function of other cell types in the CNS, including oligodendrocytes and the myelin that they produce. Reciprocally, perturbations in oligodendrocytes and myelin also affect transcriptomic and functional changes in microglia6,7.
Oligodendrocytes in the spinal cord originate from different developmental waves of oligodendrocyte precursor cells (OPCs), with the spinal cord dorsal column (DC) hosting a pool of OPCs that are of a distinct developmental origin8,9,10. These dorsally derived OPCs myelinate axons in the DC and some lateral tracts, and exhibit different transcriptomic profiles and responses to injury compared to ventrally derived OPCs10,11,12,13. However, it remains unclear whether the myelin in the DC has distinct characteristics during adulthood and aging, and how the microenvironment in the DC impacts myelin health.
The TGFβ1–TGFβR1/2 axis regulates microglia–oligodendrocyte interactions during embryonic development and adulthood, and its abrogation results in disruption of myelin integrity, as demonstrated by our previous work and that of others14,15,16. TGFβ signaling is important for microglial development and acts as a checkpoint to anchor a homeostatic state of microglia in adulthood17,18. Whether there is regional heterogeneity of microglial dependence on TGFβ signaling, and if this contributes to vulnerability to demyelination in certain regions, has not been hitherto explored.
In this study, we investigated spatiotemporal alterations in myelination patterns and microenvironmental cues within the spinal cord, identifying a region-specific demand for TGFβ signaling that fine-tunes microglia–myelin interactions within the DC.
Results
Age-dependent accumulation of myelin degeneration in the DC
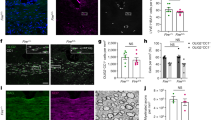
To investigate differences in myelination patterns throughout the spinal cord, we employed transmission electron microscopy (TEM) to compare myelin–axon ultrastructure in the DC and the ventral column (VC). These are two anatomically separated spinal cord white matter tracts seeded by distinct OPC pools during development8,9,10. As aging imposes error-prone myelination patterns and leads to myelin degeneration7,19,20, we conducted a comparative analysis of mice of different ages, including groups aged 2–3, 6–8, 12–13 and 16–18 months. In all age groups, we observed that axon density was much higher in the DC than in the VC, whereas the axon size in the DC was smaller than in the VC (Extended Data Fig. 1a–c), consistent with previous findings21,22. To investigate changes in myelin ultrastructure, we calculated the g ratio, defined as the ratio of the axon diameter to the outer diameter of the myelinated fiber. We determined that the g ratio in the VC remained unchanged during aging, but that it significantly decreased in the DC, starting between 8 and 12 months of age. (Fig. 1a–c). This reduction in the g ratio may be due to the decompaction of the myelin sheath, the out-folding of the myelin or myelin detachment from the axon leading to an increased periaxonal space. Consistent with this decreased g ratio, we noted marked enlargement of the periaxonal space in the DC but not in the VC of 12–13-month-old and 16–18-month-old mice (Fig. 1d–f). Subsequent TEM analyses revealed more abnormal myelin features within the DC with aging, particularly the appearance of redundant myelin (Fig. 1g–i). Immunostaining also confirmed an increased accumulation of degraded myelin basic protein in the DC of 12-month-old and older mice (Extended Data Fig. 1d–f). Taken together, our findings reveal differences in the properties of axon–myelin units between the DC and VC, confirming DC-specific myelin degeneration that begins during normal aging at approximately 8–12 months of age.
a, Representative TEM images of axon–myelin units in the DC of mice aged 2–3 months, 6–8 months, 12–13 months and 16–18 months, with myelin pseudocolored in magenta. Scale bar, 2 μm. b, Statistical analyses of the g ratio (axon diameter to outer myelin diameter) in the DC and VC across age groups. c, Correlation between axon diameter and myelin thickness of individual axon–myelin units in the DC. d, TEM images of DC axon–myelin units showing the periaxonal space (magenta) across age groups. Scale bar, 2 μm. e, Statistical analyses of periaxonal space size in the DC and VC across age groups. f, Correlation between axon diameter and periaxonal space in individual axon–myelin units of the DC. g, Representative features of myelin abnormalities in the DC, with the red arrow highlighting the specific abnormalities in each category. Scale bar, 2 μm. h, The percentage of total myelin abnormalities in myelinated axons of the DC and VC across age groups. i, The percentage of individual myelin abnormality features in myelinated axons of the DC and VC across age groups. For TEM analysis, n = 10, 5, 6 and 9 mice for the 2–3, 6–8, 12–13 and 16–18 month age groups, respectively; 100–250 axon–myelin units per region per mouse were analyzed for g ratio and periaxonal space size, and more than 300 axon–myelin units per region per mouse were analyzed for myelin abnormalities. Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01 and ***P < 0.001; two-way ANOVA with Tukey’s post hoc test was used for b, e and h. In i, analyses of redundant myelin, myelin fragment and infold were performed using one-way ANOVA with Dunnett’s post hoc test, while the other features were analyzed using Kruskal–Wallis with Dunn’s post hoc test.
Spinal cord TGFβ signaling and microglial state are spatiotemporally regulated
Since we observed myelin degeneration in the DC beginning between 8 and 12 months, we next sought to characterize age-dependent global molecular changes in the spinal cord by performing transcriptomic analysis of spinal cord homogenates from 2–3-month-old and ~12-month-old mice using the NanoString nCounter platform. This employs direct molecular barcodes to detect transcripts and thereby maintains high sensitivity and specificity without reverse transcription or amplification of the mRNA samples. The differentially expressed genes (DEGs) between the 2–3-month-old and ~12-month-old spinal cords predominantly clustered into six gene sets using an unbiased clustering algorithm (Fig. 2a). These included genes associated with microglial function, TGFβ signaling, myogenic regulatory factor (MRF) binding and proteosome activity, most of which had enhanced expression in the older spinal cord, while genes involved in myelination were suppressed. Gene Ontology (GO) analyses and pathway enrichment analyses also pinpointed an enhancement of biological processes and molecular functions linked to microglial activity (Extended Data Fig. 2a).
a, DEGs from NanoString nCounter analysis comparing the spinal cords of female mice aged 2–3 month and 12 months were analyzed using Cytoscape with the StringApp and ClusterMaker2 plugins. The gradient color of the genes represents the log2 fold change between 12-month-old and 2–3-month-old mice, with red indicating upregulation and blue indicating downregulation in the 12-month group (n = 3–5 mice per group). b,c, Western blot analysis of TGFβR2, TGFβ1, pSMAD2, SMAD2/3 and β-actin expression in the spinal cord homogenates (b) and the quantification of gene expression (c); n = 4 or 5 mice per group. d, The percentage of TGFβ1+ CNS cells in spinal cords analyzed by high-dimensional flow cytometry, including CD11b+CD45+CX3CR1+ microglia (MG), CD11b−CD45−GLAST+ astrocytes (ASTRO), CD11b−CD45−O4+ oligodendrocyte lineage cells (OPC/OL), CD11b−CD45−GLAST−O4− other CNS cells (n = 6 and 5 mice for the 2–3 month and 12–13 month age groups, respectively). e,f, RNAscope of Tgfb1 (green puncta) showing the spatial distribution of Tgfb1 in the spinal cords of mice of different ages, as well as in microglial Tgfb1-deficient mice (5 days post-tamoxifen administration in Cx3cr1CreER: Tgfb1fl/fl mice; MG-Tgfb1KO) as a negative control (scale bar, 10 μm) (e) and the quantification of Tgfb1 puncta in the DC and VC (f); n = 6 mice per group. g,h, Staining of IBA1 (red), LipidSpot (green) and nuclei (Hoechst, blue) in the DC and VC of the spinal cords of mice across age groups (g) and the quantification of LipidSpot+ lipid droplets (h). Scale bar, 10 μm; n = 6 mice per group. Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01 and ***P < 0.001. A two-tailed unpaired t-test was used for c. A two-tailed Mann–Whitney test was used for comparisons between ages within each cell type in d, and a Kruskal–Wallis test with Dunn’s post hoc test was used for cross-cell type comparisons within each age group. A two-way ANOVA with Tukey’s post hoc test was used for f. A two-tailed Wilcoxon matched-pairs signed-rank test was used for comparisons between VC and DC within each age group in h, and a Kruskal–Wallis test with Dunn’s post hoc test was used for cross-age comparisons within each region.
TGFβ signaling is critical for maintaining microglial homeostasis, and it acts as a checkpoint to suppress microglial activation18,23,24. We recorded that in both mouse and human spinal cords, the genes involved in the TGFβ signaling pathway were upregulated with aging (Extended Data Fig. 2b,c). Using western blotting, we further confirmed an age-dependent increase in the protein expression of TGFβ1 and TGFβR2 in the spinal cords (Fig. 2b,c). However, SMAD2 expression and its phosphorylation remained unchanged despite increased TGFβ1 expression in the older spinal cord, suggesting that higher levels of TGFβ1 are required to maintain a similar level of downstream SMAD signaling during aging. In addition, the ratio between the TGFβ1 latent complex and its active form was increased in the older spinal cord, suggesting that the efficiency of liberating bioactive TGFβ1 from the latent TGFβ complex was also compromised during aging. To pinpoint the cellular source of increased TGFβ1 during aging, we performed high-dimensional flow cytometry and confirmed that microglia are the main source of TGFβ1, with production significantly increasing with age (Fig. 2d and Extended Data Fig. 3a–c).
Since myelin degeneration increases in the DC with aging (Fig. 1) and TGFβ signaling regulates microglia–myelin interactions14,15,16, we next investigated whether the enrichment of TGFβ1 also exhibits a spatial preference during normal aging. Using RNAscope we examined the spatial distribution of Tgfb1 in the spinal cord. Consistent with the flow cytometry results, RNAscope analysis confirmed that Tgfb1 was specifically localized in IBA1+ microglia (Fig. 2e,f). Moreover, we noted that Tgfb1 was markedly elevated in DC microglia compared to in VC microglia, with this disparity becoming increasingly pronounced during aging. In parallel, we performed microdissection of the DC, VC and the GM and analyzed their respective mRNA expression of genes involved in the TGFβ signaling pathway. We consistently observed higher levels of TGFβ signaling-related genes in the DC compared to in either the VC or GM, respectively, and this was further amplified with aging (Extended Data Fig. 2d). These data suggest that the DC harbors a TGFβ1-enriched microenvironment that sustain microglial TGFβ signaling during aging.
Since microglia monitor the microenvironment and are the primary source of TGFβ1, we hypothesized that the accumulation of myelin degeneration in the DC during normal aging may induce specific changes in the states of DC microglia. Using flow cytometry we first confirmed that microglia in the spinal cord of 12–13-month-old mice exhibited elevated expression of surface markers compared to those in the 2–3-month-old mice (Extended Data Fig. 2e,f). These included markers identified as DEGs from the NanoString analysis (Fig. 2a), such as GPNMB, CD45 (Ptprc) and CD74, as well as Dectin-1, a marker indicative of increased microglial activity25. Lipid droplet accumulation is often associated with alterations in microglial phagocytosis, metabolism and responses to damage or to changes in the microenvironment26,27,28. To further investigate this, we performed LipidSpot staining and observed an age-dependent increase in lipid droplet accumulation in older microglia across both regions, with a higher accumulation in the DC compared to the VC starting at 12 months (Fig. 2g,h). Taken together, these data demonstrate that aging induces DC-specific changes in TGFβ signaling and microglial state.
Disruption of microglial TGFβ signaling leads to microglial activation and demyelination in the DC
Given that TGFβ signaling maintains microglial homeostasis, it is counterintuitive that DC microglia exhibit a more reactive state during normal aging, despite the enriched TGFβ signaling in this region. To mechanistically understand the biological function of increased TGFβ signaling in the DC with aging and its potential effect on microglia, we conducted loss-of-function studies to disrupt TGFβ signaling in microglia. We used 8–12-month-old Cx3cr1CreER:Tgfbr2fl/fl (knockout (KO)) mice, as myelin degeneration in the DC starts to appear at this age. Tamoxifen administration induces Tgfbr2 deletion in Cx3cr1-expressing microglia, thereby preventing binding of TGFβ ligands and subsequent recruitment of TGFβR1 and SMAD signaling transduction29 (Fig. 3a). Transcriptomic analyses revealed that Tgfbr2-deficient microglia downregulated microglia homeostatic gene expression (Extended Data Fig. 4a–c) and concurrently upregulated genes involved in inflammatory responses, phagocytosis, cell adhesion, antigen presentation, cholesterol metabolism and matrix metalloproteinases (Extended Data Fig. 4c–e). This microglial molecular signature resulting from the disruption of TGFβ signaling is consistent with previous reports14,18,30. NanoString analysis also confirmed kinetic molecular changes in the spinal cord following microglial Tgfbr2 deletion (Extended Data Fig. 5a–f). Specifically, we noted that the previously described microglia neurodegenerative phenotype or disease-associated microglia (DAM) signature was highly induced, including expression of genes such as Apoe, Trem2, Lpl, Itgax, Lirlb4a, Ccl6 and Clec7a (Extended Data Fig. 5c–e)31,32. Conversely, genes involved in myelination were downregulated, whereas genes involved in oligodendrocyte differentiation were upregulated (Extended Data Fig. 5f).
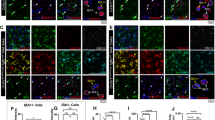
a, A schematic illustration of Tgfbr2 deletion blocking TGFβ ligand binding in microglia of Cx3cr1CreER:Tgfbr2fl/fl mice. b–d, Immunofluorescent staining of IBA1 (green), MHC-II (magenta) and nuclei (Hoechst, blue) in cervical spinal cord sections of Tgfbr2fl/fl (WT) and Cx3cr1CreER:Tgfbr2fl/fl (KO) mice (~12 months old) at 7, 20, 30 and 60 days post-tamoxifen injection (b), with the quantification of microglia number (c) and mean fluorescent intensity (MFI) of IBA1 and MHC-II (d). Scale bar, 200 μm; n = 4, 3, 3, 5 and 5 mice for each time point. e,f, Flow cytometry analyses comparing the expression of microglial surface markers MHC-II and CD45 in CD11b+CD45+Ly6C−Ly6G− cells microdissected from the DC, VC and GM regions of adult WT and Cx3cr1CreER:Tgfbr2fl/fl mice at 18 or 28 days post-tamoxifen administration (e) and quantification comparing the frequency of MHC-IIhigh and CD45high microglia across the different spinal cord regions (f); n = 3, 5, and 4 mice for each group. g,h, Staining of fluoromyelin (green), IBA1 (red) and nuclei (Hoechst, blue) in cervical spinal cord sections of ~12-month-old WT and KO mice at 20, 30 or 60 days post-tamoxifen administration (g) and quantification comparing the percentages of demyelinated area between DC and VC (D30) (h). Scale bar, 200 μm, n = 4 mice. i, Representative immunofluorescent staining of IBA1+ microglia (green), PLP1+ myelin (magenta) and TUJ1+ axons (cyan) in the DC of KO mice 25 days post-tamoxifen administration, with an orthogonal display illustrating the spatial arrangement of axons surrounded by microglia. A 3D reconstruction of microglia encompassing the axon–myelin units is shown below. Scale bar, 3 μm. j, Representative TEM images in the DC and VC of ~12-month-old WT and KO mice (25 days post-tamoxifen administration). Scale bar, 20 μm. The insets below indicate degenerated axons (orange arrows) and redundant myelin (red arrows). Scale bar, 2 μm. k,l, Quantification of myelinated axons (k) and myelin abnormalities (l) in the DC and VC of WT and KO mice; n = 4 and 3 mice for WT and KO_D25, respectively. m,n, Spontaneous behavioral deficits in KO mice were assessed using the disease score (m) and four-paw hanging wire behavioral test (n); n = 18 mice for the young female (YF) and older female (OF) groups, and n = 11 and 12 mice for the young male (YM) and older male (OM) groups, respectively. Young mice: 2–3 months; older mice: 8–12 months. Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01 and ***P < 0.001. A two-way ANOVA with Sidak’s post hoc test was used for c and d. Cross-region comparisons within each group in f were analyzed using repeated-measures one-way ANOVA with Geisser–Greenhouse correction, followed by Dunnett’s post hoc test. A two-tailed paired or unpaired t-test was used for h and i, respectively. Multiple unpaired t-tests with false discovery rate (FDR) correction were used for k. The cumulative disease score in m was analyzed using the Kruskal–Wallis with Dunnett’s post hoc test, and the mean time in the four-paw hanging wire test in n was analyzed using one-way ANOVA with Tukey’s post hoc test. p.t., post-tamoxifen administration. Panel a created with BioRender.com.
Histologically we first stained IBA1+ microglia and MBP+ myelin in a segment of spinal cord wholemounts followed by iDISCO tissue clearing and 3D imaging. We observed substantial microglia accumulation in the DC of Cx3cr1CreER:Tgfbr2fl/fl mice, accompanied with loss of MBP+ myelin (Extended Data Fig. 6a). The 3D reconstruction further demonstrated that Tgfbr2-deficient microglia extensively adhered to myelin in the DC (Extended Data Fig. 6b). Subsequent histological staining revealed a progressive increase in the number and immunoreactivity of IBA1+ microglia within the DC, characterized by elevated expression of the activation marker MHC-II, while the VC exhibited markedly less MHC-II positivity (Fig. 3b–d). We further compared the expression of surface markers relevant to microglial activation in microglia isolated from the DC, VC and GM, respectively, using flow cytometry at 18 (D18) and 28 days (D28) post-Tgfbr2 deletion (Fig. 3e,f). This revealed that the percentages of MHC-IIhigh or CD45high microglia were much higher in the DC compared to in other spinal cord regions. This DC-specific microglial activation was accompanied by myelin loss restricted to the DC, but not observed in the VC, as revealed by fluoromyelin staining, which became evident at approximately 30 days after the deletion of microglial Tgfbr2 (Fig. 3g,h). Furthermore, microglia in the DC displayed a gigantic foamy morphology with ingested myelin components (Fig. 3g). Costaining with the myelin protein PLP1 and the axon marker Tuj1 demonstrated that in the absence of TGFβ signaling, microglia in the DC were closely intertwined with the axons, compressing the axon–myelin units (Fig. 3i). To further confirm the spatially heterogeneous microglial response following Tgfbr2 deletion, we performed transcriptomic analyses of microglia isolated from microdissected DC, VC and GM of the spinal cord (Extended Data Fig. 7a). Compared to Tgfbr2-deficient microglia from the VC and GM, those in the DC were enriched in biological processes such as cell–cell adhesion and inflammatory response, and upregulated genes such as Il1b and Tnf, which are known to induce apoptosis of oligodendrocytes33,34 (Extended Data Fig. 7b–d).
At the ultrastructural level, TEM analyses revealed a marked reduction in the frequency of myelinated axons in the DC compared to in the VC following microglial Tgfbr2 deletion, along with increased abnormalities such as axonal degeneration and loose, redundant myelin in the DC (Fig. 3j–l). Collectively, these findings suggest that following Tgfbr2 deletion, microglia in the DC become excessively activated, resulting in severe myelin damage.
To compare the functional impact of myelin degeneration observed in the DC, we assessed mice for behavioral deficits and performed behavioral tests. We included both male and female mice at young (2–3 months) and older (8–12 months) ages to compare the impact of age and gender. Microglial Tgfbr2-deficient mice developed neurological impairments including deficits in tail, gait, urinary and limb functions, as well as weight loss. Female mice exhibited more severe symptoms than did males, with older males being more affected than younger ones (Fig. 3m). In the four-paw hanging wire test, microglial Tgfbr2-deficient mice were unable to complete the test, with older mice performing worse than younger ones, while young males were only partially affected after 6 weeks (Fig. 3n).
In summary, our findings reveal that, during aging, microglia in the DC require continuous TGFβ signaling as a regulatory checkpoint to prevent excessive responses and myelin damage.
snRNA-seq reveals a subset of microglia in the DC sensitive to the loss of TGFβ signaling
To gain an in-depth molecular understanding of how different cell types in the spinal cord respond to the disruption of microglial TGFβ signaling, we utilized single-nucleus RNA-sequencing (snRNA-seq) of freshly dissected spinal cord tissues from 12-month-old Cx3cr1CreERTgfbr2fl/fl mice at 10 (D10), 20 (D20) and 30 (D30) days post-tamoxifen administration, as well as Tgfbr2fl/fl littermate controls (wild type (WT)). This approach involved snap freezing and mechanical dissociation of the tissues to obtain nuclei for sequencing, providing a more accurate representation of the cellular landscape and minimizing gene expression artifacts induced by enzymatic digestion35,36. Following quality control, we included 182,019 nuclei from 11 samples across the four conditions. Using uniform manifold approximation and projection (UMAP) clustering followed by cell cluster annotation, we identified major cell types in the spinal cords (Extended Data Fig. 8a,d). We primarily noted dynamic changes in the proportion of glial cells at D20 and D30 following microglial Tgfbr2 deletion (Extended Data Fig. 8b,c). Specifically, the microglia and pre-mature oligodendrocyte (pre_Oligo) populations expanded over time, whereas the mature oligodendrocyte (Oligo) population decreased substantially.
Since we observed a temporal expansion of the microglia population we first pooled microglia from all conditions (11,313 nuclei) and subsequently performed separate subclustering, resulting in four major subclusters (Fig. 4a). We observed a notable expansion of microglia subcluster 1 after Tgfbr2 deletion, characterized by high expression of genes related to DAM signature genes (Trem2, Apoe and Clec7a), antigen presentation (H2-Aa, Ciita and Cd74), lipid processing (Lrp1, Cd36, Plin2 and Abca1) and genes associated with myelin interaction (Lgals3, Gpnmb and Mgll)37,38,39,40,41 (Fig. 4b–d). Lgals3 and Gpnmb are also signature genes of a previously identified microglial subset transiently present in postnatal white matter that phagocytoses newly formed oligodendrocytes and myelin3. We therefore designated microglia subcluster 1 as TGFβ signaling-sensitive microglia (TSM). By comparing marker gene expression we identified subcluster 3 as homeostatic microglia, while subcluster 2 was probably involved in cell adhesion and extracellular matrix (ECM) remodeling and was designated as tissue-remodeling microglia (TRM). Subcluster 0 was enriched with myelin-related genes but downregulated microglial homeostatic markers, resembling the previously identified myelin transcript-enriched microglia (MyTE)41,42. GO analyses revealed that TSM were involved in inflammatory responses, cytokine production, cell adhesion and phagocytic activity, with upregulated genes associated with endolysosomal function and lipid-droplet formation (Fig. 4e).
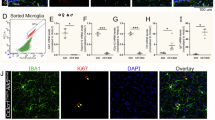
a, A UMAP plot of spinal cord microglia from 12-month-old female Tgfbr2fl/fl (WT) mice and Cx3cr1CreER:Tgfbr2fl/fl (KO) mice at D10, D20 or D30 post-tamoxifen administration (downsampled to 500 cells per group per time point for presentation). b, The kinetic fold change (compared to WT condition) of each microglial subcluster in a over time. c, The proportion of each microglial subcluster in the total microglial population in WT and KO_D30 conditions. Subclusters 0, 1, 2 and 3 in a and b are denoted as MyTE, TSM, TRM or homeostatic microglia (HM), respectively. d, A bubble plot depicting the expression of marker genes across different subclusters. Bubble size represents the percentage of cells expressing each gene within a subcluster and color indicates the average normalized expression level. Marker genes were identified using Seurat’s FindAllMarkers function (two-sided Wilcoxon rank-sum test with Benjamini–Hochberg correction for multiple comparisons). e, GO enrichment analysis based on the upregulated DEGs in TSM compared to other subclusters; log2 fold change >1, adjusted P value <0.001. f–h, Immunostaining validation of TSM signature markers GPNMB and Galectin-3 (Lgals3) in the spinal cords of 12-month-old WT and KO mice (scale bar, 100 or 20 μm), with quantification of GPNMB and Galectin-3 MFI (h) and insets in g showing costaining of F4/80–GPNMB and IBA1–Galectin-3. n = 6, 3, 3, 5 and 5 mice for each group. Data are presented as mean ± s.e.m. *P < 0.05 and **P < 0.01; multiple paired t-tests with FDR correction.
To assess the spatial distribution of TSM, we validated the kinetic expression of GPNMB and Galectin-3 (encoded by Lgals3) in the spinal cord following microglial Tgfbr2 deletion using immunostaining. We observed a time-dependent upregulation of GPNMB and Galectin-3 in the DC compared to in the VC (Fig. 4f–h). At an advanced stage (D60), the upregulation of Galectin-3 and GPNMB was also evident in the lateral columns, areas where microglia accumulation was also evident at a later stage following Tgfbr2 deletion (Fig. 3b). Costaining with F4/80 or IBA1 further confirmed the colocalization of GPNMB and Galectin-3 in microglia displaying a foamy morphology in the DC (Fig. 4g). This indicates that the induced TSM are mainly located in the DC region, highlighting their engagement in the DC demyelinating pathology. Collectively, these findings confirm that a microglia subset, primarily residing in the DC, is highly reliant on TGFβ signaling.
TSM phagocytose myelin and accumulate lipid droplets
We next investigated the functional role of TSM. As we had demonstrated remarkable myelin loss in the DC following Tgfbr2 deletion and that TSM were enriched with genes involved in phagosome and endolysosome activity as well as lipid droplet formation, we hypothesized that the TSM are highly phagocytic and would ingest more myelin. We first characterized the phagocytic activity of Tgfbr2-deficient microglia. Cultured primary microglia isolated from the spinal cords of Tgfbr2fl/fl and Cx3cr1CreERTgfbr2fl/fl mice (20 days post-tamoxifen administration) were incubated with pHrodo red-labeled myelin, which fluoresces red upon ingestion into the acidic lysosomes of microglia. Time-lapse live-cell imaging revealed a significantly increased uptake of myelin by Tgfbr2-deficient microglia (Fig. 5a,b). To further validate whether TSM were more phagocytic we incubated ex vivo spinal cord single-cell suspensions with pHrodo red-labeled myelin and used the TSM signature marker Galectin-3 as a proxy to identify the TSM subset using flow cytometry. We noted that the Galectin-3high microglia were specifically induced in Cx3cr1CreERTgfbr2fl/fl mice and confirmed significantly enhanced myelin uptake by Tgfbr2-deficient microglia (Fig. 5c,d,f). We further compared Galectin-3low and Galectin-3high microglia from the spinal cords of Cx3cr1CreERTgfbr2fl/fl mice and confirmed that Galectin-3high microglia exhibited elevated myelin phagocytosis activity compared to their Galectin-3low counterparts (Fig. 5e,g).
a,b, Time-lapse imaging (IncuCyte S3) of cultured spinal cord microglia from 8–12-month-old Tgfbr2fl/fl and Cx3cr1CreER:Tgfbr2fl/fl mice (D20 post-tamoxifen administration) incubated with pHrodo red-labeled myelin (scale bar, 50 μm; n = 3–4 wells) (a); ingested myelin fluoresces with a red signal, and the red signal intensity (normalized to cell confluence) in each group is presented over time (b). c–g, Spinal cord single-cell suspensions from 8–12-month-old Tgfbr2fl/fl and Cx3cr1CreER:Tgfbr2fl/fl mice (D20) were incubated with pHrodo red-labeled myelin for 1 h, followed by flow cytometric analysis (n = 5 or 8 mice). Shown are representative flow cytometry plots and quantification of the percentage of Galectin-3high microglia between the two groups (c,d), of the percentage of pHrodo-red+ microglia between the two groups (e,f) and the percentage of pHrodo Red+ microglia in Galectin-3low and Galectin-3high populations in the Cx3cr1CreER:Tgfbr2fl/fl mice (g). FMO, fluorescence-minus-one control for pHrodo Red. h, Representative immunofluorescent staining from two independent experiments with similar results showing IBA1 (green), PLIN2 (red) and nuclei (Hoechst, blue) in the VC and DC of 8–12-month-old Tgfbr2fl/fl and Cx3cr1CreER:Tgfbr2fl/fl (D20) mice. Scale bar, 10 μm. i–k, Staining of IBA1 (green), LipidSpot (magenta) and nuclei (Hoechst, blue) in the DC of 8–12-month-old Tgfbr2fl/fl mice and Cx3cr1CreER:Tgfbr2fl/fl mice at D20 or D60 post-tamoxifen administration (i) and quantification of LipidSpot puncta in the DC and VC (j). Scale bar, 20 μm; n = 3–4 mice per group. The insets in i show staining of IBA1 (green), Galectin-3 (yellow), LipidSpot (magenta) and nuclei (Hoechst, blue) in the DC of Cx3cr1CreER:Tgfbr2fl/fl mice (D60), with the quantification of LipidSpot puncta between Galectin-3low and Galectin-3high IBA1+ cells (k); n = 4, with ~50 cells analyzed per mouse. Data are presented as mean ± s.e.m. *P < 0.05 and ***P < 0.001; a two-way ANOVA with Sidak’s post hoc test was used in b. A two-tailed unpaired t-test was used in d and f, and a two-tailed paired t-test was used in g and k. Multiple paired t-tests with FDR correction were used in j.
Excessive myelin uptake poses a burden for microglia in handling the ingested lipids43. Accordingly, we characterized a temporal accumulation of lipid droplets in the DC of the spinal cord following Tgfbr2-deletion, as revealed by Plin2 and LipidSpot staining (Fig. 5h–j). In addition, Galectin-3high microglia accumulated more lipid droplets compared to Galectin-3low microglia (Fig. 5i,k). Taken together, these results demonstrate an elevated myelin-ingesting capacity of TSM.
Microglial Tgfbr2 deletion alters the landscape of oligodendrocyte lineage cells in the spinal cord
We next addressed how loss of TGFβ signaling in microglia would affect oligodendrocyte lineage cells in the spinal cord. Previous studies have revealed heterogeneity of mouse oligodendrocytes in the juvenile and adult CNS, and six mature oligodendrocyte (MOL) subclusters were reported44,45. Among the MOL subclusters, MOL5/6, characterized by high expression of Grm3, Ptgds and Il33, appeared to be more distinct from MOL1–4, while MOL2 exhibited specific marker genes such as Klk6, Hopx, Anxa5 and Pcsk6 (refs. 11,46,47). To explore potential changes in oligodendrocytes in Cx3cr1CreERTgfbr2fl/fl mice, we combined the oligo, pre_Oligo and OPC populations identified by snRNA-seq (Extended Data Fig. 8a) and performed subclustering of a total of 49,112 nuclei. We identified different subclusters of oligodendrocyte lineage cells based on their marker gene expression, including Pdgfra+ OPC, committed oligodendrocyte progenitors (COP) or newly formed oligodendrocytes (NFOL) positive for Bcas1 and Tcf7l2, Opalin+/Ctps+ myelin-forming oligodendrocytes (MFOL), Grm3+/Ptgds+ MOL5/6, MOL2 signature-enriched MOL1–4 and a recently identified disease-associated oligodendrocyte subcluster (DA-MOL) expressing Serpina3n46,47,48 (Fig. 6a–e and Extended Data Fig. 7a–c). We recorded a substantial loss of MOLs, with MOL1–4 appearing to be more affected than were MOL5/6 (Fig. 6a,c). This loss of MOL was accompanied by the expansion of OPC, COP/NFOL, MFOL and DA-MOL. Notably, DA-MOLs were specifically induced following microglial Tgfbr2 deletion (Fig. 6c).
a, A t-SNE plot of oligodendrocyte lineage cells from 12-month-old female WT mice and KO mice at D10, D20 and D30 post-tamoxifen administration (downsampled to 5,000 cells per group per time point for presentation). b, A bubble plot showing the expression of signature genes across different subclusters. Bubble size indicates the percentage of cells expressing the gene in the subcluster and the color scale bar indicates the normalized expression of the gene. c, The kinetic fold change (compared to WT condition) of each subcluster in a over time. d, A volcano plot depicting the DEGs between MOL1–4 and MOL5/6 (log2 fold change (FC) >1, adjusted P value <0.00; two-sided Wilcoxon rank-sum test with Benjamini–Hochberg correction). e, A volcano plot depicting the DEGs between DA-MOL and other MOLs (log2 fold change >1, adjusted P value <0.001; two-sided Wilcoxon rank-sum test with Benjamini–Hochberg correction). f,g, Spatiotemporal validation of DA-MOL in the spinal cord by staining of CC-1 (green), Serpina3n (magenta), fluoromyelin (white) and nuclei (Hoechst, blue) (f), with the inset showing the magnified image in the DC of KO_D20 mice, and quantification of Serpina3n MFI (g). Scale bar, 50 μm or 20 μm; n = 3, 5 and 4 mice for each group. Data are presented as mean ± s.e.m. *P < 0.05 and **P < 0.01; multiple paired t-tests with FDR correction.
We next validated the presence and spatial distribution of DA-MOLs in the spinal cord by immunostaining with the DA-MOL marker Serpina3n and the mature oligodendrocyte marker CC-1. We confirmed that the Serpina3n+CC-1+ DA-MOLs already appeared by D20 following microglial Tgfbr2 deletion, preceding substantial myelin loss, and that they were predominantly present in the DC but rarely in the VC (Fig. 6f,g). However, despite the generation and accumulation of DA-MOLs in the DC, they failed to synthesize myelin and ensheathe axons, as evidenced by the lack of fluoromyelin staining in this region at the experimental endpoint (D60) (Fig. 6f). Collectively, our results demonstrate that microglial Tgfbr2 deletion restructures the landscape of oligodendrocyte lineage cells in the spinal cord and induces a disease-associated oligodendrocyte subcluster in the DC.
DC microglia require autocrine TGFβ1 to maintain resilience to myelin degeneration
Our results confirm the requirement of TGFβ signaling in maintaining microglial resilience and preventing uncontrolled myelin damage in the DC. However, the cellular source of the TGFβ ligands required by DC microglia remained unclear. TGFβ1 is predominantly expressed by microglia compared to other CNS cell types (Fig. 2d) and is functionally irreplaceable by TGFβ2/3 for adult microglial homeostasis49. We noted that while expression of Tgfbr1 and Tgfbr2 were downregulated in the spinal cords of Cx3cr1CreERTgfbr2fl/fl mice following tamoxifen administration (Extended Data Fig. 10a,b), Tgfb1 expression was significantly upregulated along with the expression of Nrros, which encodes an essential anchoring protein LRRC33 for the integrin-mediated liberation and activation of latent TGFβ15 (Extended Data Fig. 10c,d). Furthermore, in microglia sorted from microdissected DC, VC and GM of Cx3cr1CreERTgfbr2fl/fl mice, we recorded higher Tgfb1 expression in DC microglia, which was independent of the expression of Tgfbr2 (Extended Data Fig. 10e). In addition, among the microglial subclusters identified through snRNA-seq analysis, Tgfb1 was predominantly expressed in TSM compared to in other subclusters (Extended Data Fig. 10f).
This suggested a potential compensatory mechanism to upregulate TGFβ1 production when microglia are unable to sense TGFβ ligands owing to the absence of receptors. We therefore hypothesized that autocrine TGFβ1 is essential for DC microglia to prevent myelin damage in the DC. To test this, we bred Cx3cr1CreERTgfb1fl/fl mice to delete Tgfb1 in microglia following tamoxifen administration (Fig. 7a). We noted a similar demyelinating pathology and aberrant microglial responses in the DC following Tgfb1 deletion as was observed in the Cx3cr1CreERTgfbr2fl/fl mice (Fig. 7b). Using high-dimensional flow cytometry analyses and unbiased clustering we uncovered a distinct separation between Tgfb1-deficient spinal cord microglia and those isolated from Tgfb1fl/fl littermate controls (Fig. 7c). In addition, this analysis identified a Galectin-3high subcluster following microglial Tgfb1 deletion, which was similar to the Lgals3+/Galectin-3+ TSM identified by snRNA-seq in Cx3cr1CreERTgfbr2fl/fl mice. Immunostaining with GPNMB and Galectin-3, signature markers of TSM, further confirmed a significant upregulation in the DC following microglial Tgfb1 deletion (Fig. 7d–f). Consistent with the phenotype reported in the Cx3cr1CreERTgfbr2fl/fl mice, microglial Tgfb1-deficient mice also developed severe neurological and behavioral deficits that were exacerbated with aging, although the disease progression was slower compared to through the deletion of Tgfbr2 (Fig. 7g,h). Taken together, these results demonstrate that microglia in the DC rely on autocrine TGFβ1 signaling to prevent damage to myelin.
a, Cx3cr1CreER:Tgfb1fl/fl mice were employed to delete TGFβ1 production in microglia. b, In 12-month-old Cx3cr1CreER:Tgfb1fl/fl mice (D30), a similar demyelinating pathology and microglial activation in the DC were evident, as observed in the Cx3cr1CreER:Tgfbr2fl/fl mice. Scale bar, 50 μm. c, t-SNE plots from high-dimensional flow cytometry reveal segregation of the microglia population between Tgfb1fl/fl and Cx3cr1CreER:Tgfb1fl/fl mice (D60). The mapped distribution of F4/80 (green), CD45 (yellow), MHC-II (purple), Galectin-3 (orange) and MSR1 (blue) indicates cells with high marker expression in the t-SNE plots. n = 4 mice per group. d–f, Staining of Galectin-3 (cyan), GPNMB (red) and nuclei (Hoechst, blue) in the VC and DC of Tgfb1fl/fl and Cx3cr1CreER:Tgfb1fl/fl mice (D30) (d), with quantification of Galectin-3 (e) and GPNMB MFI (f). Scale bar, 50 μm; n = 4 and 5 mice for the Tgfb1fl/fl and Cx3cr1CreER:Tgfb1fl/fl groups, respectively. g,h, Comparison of clinical scores (g) and four-paw hanging wire behavioral tests (h) between young (2–3 months; YF) and older female (8–12 months; OF) Cx3cr1CreER:Tgfb1fl/fl mice following tamoxifen administration. Tgfb1fl/fl mice and Cx3cr1CreER:Tgfbr2fl/fl mice (OF) were included as negative and positive control groups, respectively. n = 7, 10 and 7 mice for the Cx3cr1CreER:Tgfbr2fl/fl (OF), Cx3cr1CreER:Tgfb1fl/fl (OF) and Cx3cr1CreER:Tgfb1fl/fl (YF) groups, respectively. Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01 and ***P < 0.001; multiple paired t-tests with FDR correction were used for comparisons between DC and VC in e and f. The cumulative disease score in g was analyzed using the Kruskal–Wallis with Dunnett’s post hoc test, and the mean time in the four-paw hanging wire test in h was analyzed using the one-way ANOVA with Tukey’s post hoc test. Schematic in panel a created with BioRender.com.
Discussion
Our study reveals that an error-prone myelination pattern in the DC, which develops during normal aging, presents a physiological challenge to DC microglia that is otherwise manageable through upregulated TGFβ signaling. This functions as an efficient checkpoint mechanism, preventing further aging-induced damage to DC myelin by primed yet resilient microglia. However, in the absence of Tgfbr2 or Tgfb1, this checkpoint is absent and microglia exhibit an aberrant activation state that primarily results in myelin loss in the DC (Fig. 8). In addition, we identify a microglia subcluster in the DC that is hypersensitive to the loss of TGFβ signaling, becoming excessively phagocytic toward myelin. A disease-associated oligodendrocyte subcluster also expands in the DC in attempt to replenish the vacant oligodendrocyte niche but fails to myelinate axons. Our study thus reveals region-specific heterogeneity in microglial dependence on TGFβ signaling, underscores the vulnerability of DC myelin when TGFβ signaling is insufficient and indicates an age-dependent increased dependence on TGFβ signaling.
In a homeostatic state, microglia in the DC survey myelin health and regulate myelination. However, with aging, myelin in the DC undergoes ultrastructural changes indicative of myelin decompaction, which can be sensed by microglia. Such perturbations in myelin trigger low-grade microglial activation and immune responses, consequences of chronic exposure to degraded myelin fragments during the aging process. At this stage, microglia remain resilient owing to a microenvironment in the DC that reinforces microglial TGFβ signaling, which helps suppress excessive microglial immune responses. This state can be described as an inflammaging adaptive state, existing before reaching a critical threshold that transitions into an irreversible microglial inflammatory response, leading to further myelin damage. A critical tipping point occurs when TGFβ signaling is not sustained, which microglia in the DC particularly rely on for resilience. This leads to unchecked microglial activation and subsequent pathological myelin damage, culminating in a maladaptive state. The balls in the lower images symbolize the complex system of the DC, the green lane represents a healthy state spectrum, while the red lane represents a pathological state. The schematic representation of the tipping points in a complex dynamical system draws inspiration from Simons et al.63 and was created with BioRender.com.
DC-specific myelin loss is observed in both human and experimental conditions. Human diseases such as subacute combined degeneration, caused by deficiencies in copper and vitamin B12, or by long-term exposure to nitrous oxide, primarily present as myelopathy affecting the DC of the spinal cord, accompanied by accumulation of foamy microglia or macrophages50,51,52. If not treated promptly, this leads to irreversible nerve damage. In addition, patients with neurosyphilis also develop irreversible DC myelin degeneration with increased cerebrospinal fluid markers indicative of microglial activation53,54. Whether disrupted TGFβ signaling contributes to these human diseases remains to be explored. In mice, DC demyelination is observed in other experimental paradigms in which TGFβ signaling is disrupted. Mice lacking LRRC33, a microglial surface molecule that enables highly localized, integrin-dependent TGFβ activation, develop paraparesis, exhibit myelin loss in the DC and die by 5 months of age15. Our previous work also demonstrated that engraftment of Tgfbr2-deleted monocyte-derived macrophages into the microglia-depleted CNS damages myelin in the DC, leading to the development of a fatal neurological disease14. In our current study, we further demonstrate that deletion of microglial Tgfbr2 or Tgfb1 leads to similar DC pathology and neurological disease, which is exacerbated in older mice. Together, these findings underscore the vulnerability of DC myelin.
In the spinal cord, the functional differences between oligodendrocyte lineage cells from distinct developmental waves remain a topic of intense research. While the ventrally derived OPCs (emerging at embryonic day 12.5) dominate the oligodendrocyte lineage cells in the spinal cord (∼80%), dorsally derived OPCs (emerging at embryonic day 15.5) outcompete them for myelination of dorsal axons, as well as in the lateral corticospinal and rubrospinal tracts10. Transcriptomic and electrophysiological analyses reveal that in the developing CNS, OPCs generated from different developmental waves are quite similar10,55. However, recent studies employing in situ sequencing and hybridization reveal that a mature oligodendrocyte subcluster MOL2, enriched for marker genes such as Klk6, Anxa5 and Hopx, preferentially envelop axons of sensory neurons ascending in the DC11. Functional studies using adult mice also confirm that dorsal OPCs exhibit enhanced recruitment and differentiation following demyelinating injury12. Moreover, they are more vulnerable to age-related decline in their capacity to differentiate compared to their ventrally derived counterparts. In addition, the accumulation of DNA damage tends to induce apoptosis of dorsal OPCs, while ventral OPCs enter cell senescence13. This suggests that despite subtle differences during development, oligodendrocyte lineage cells originating from different waves may respond differently to aging conditions.
In agreement with this, our TEM results demonstrate that dorsal myelin undergoes more pronounced ultrastructural alterations during normal aging. We reveal that axon density in the DC is significantly higher, which may indicate an increased myelinating demand on oligodendrocytes in this region. In addition, the DC axons are, on average, smaller than those in the VC. It is proposed that axon caliber size serves as a physical cue that determines both the complexity of myelination and the myelinating capacity of oligodendrocytes: myelinating smaller axons is more challenging and energy demanding for oligodendrocytes owing to the difficulty of bending myelin beyond a certain curvature56,57. Axonal physical properties in the DC (higher density and smaller caliber) therefore probably introduce more stress on the oligodendrocytes to maintain myelination, which, over time with aging leads to error-prone myelination patterns. These ultrastructural changes in DC myelin during normal aging may, therefore, sensitize microglia and predispose them to a microglial attack in cases in which microglial TGFβ signaling fails to serve as a checkpoint.
Microglial phagocytosis is an important function for the clearance of CNS wastes. However, it must be well controlled and either too little or too excessive microglial phagocytic activity may lead to neurodegeneration. Microglia are the main effector cells in the CNS that phagocytose degraded myelin components, and efficient clearance of myelin debris by microglia is also a prerequisite for remyelination following demyelinating injuries58. Myelin is composed of a high proportion of lipids (approximately 70–85%) and cholesterol is one of its major components. Free cholesterol is hard to degrade and is cytotoxic. During the aging process, microglia are constantly exposed to myelin fragments owing to age-related myelin degradation, which burdens their lysosomal capacity and clearance function7. Excessive accumulation of cholesterol in microglia also forms cholesterol crystals and triggers inflammasome activation, as well as lipid droplet formation43. Recent studies have identified an ACSL1+ lipid droplet-associated microglial subset linked to the APOE4 genotype28,59,60. However, we did not observe ACSL1+ microglia in the DC of microglial Tgfbr2-deficient mice, suggesting a different mechanism underlying lipid droplet accumulation compared to that in APOE4 microglia. As a potential protective mechanism to restrain uncontrolled immune activation after being exposed to abnormal myelin, TGFβ signaling is reinforced to suppress microglial proinflammatory responses and to prevent their excessive phagocytic activities59,61,62. This mechanism renders sensitized or primed microglia more resilient to age-related CNS changes, helping them maintain an adaptive state that, if dysregulated, may lead to uncontrolled microglial immune activation and further irreversible CNS damage63 (Fig. 8).
Growing evidence suggests that excessive microglial phagocytosis of stressed but viable neurons and other neural cells contributes to multiple CNS pathologies and neurodegeneration, a process termed phagoptosis64,65. This is different from efferocytosis, in which cells undergo apoptosis and microglia remove them through phagocytosis. Instead, under certain perturbations such as in the absence of TGFβ signaling, microglia overreact to stressed but functional and viable neural cells, phagocytosing these cells and causing their death. However, recent studies suggest that in other CNS regions during pathological conditions, suppressing TGFβ signaling induces the DAM signature and enhances microglial phagocytosis of myelin debris or amyloid plaques, which is beneficial for promoting remyelination and preventing neurodegeneration in some pathological states59,61. These findings further underscore the importance of considering spatial information and CNS context (whether physiological or pathological) when manipulating TGFβ signaling. Our results thus strengthen the case for being cautious when using microglial TGFβ signaling inhibition as an attractive strategy for neurodegenerative conditions, such as Alzheimer’s disease. Furthermore, TGFβ pathway antagonists are rapidly emerging as effective and promising anticancer agents for preclinical investigations66,67. Our study therefore indicates a potential safety concern for patients receiving these therapies, especially for elderly patients.
Despite TGFβ being a potent ligand for microglial homeostasis, the cellular source of TGFβ that DC microglia rely on remained elusive. We demonstrate that deleting not only microglial Tgfbr2 but also Tgfb1 triggers an aggressive microglial response against myelin in the DC, confirming a hitherto overlooked TGFβ autocrine mechanism that microglia largely rely on. A recent landmark study underscores the crucial role of microglia in maintaining myelin integrity through the production of TGFβ1, acting on TGFβR1 receptors on oligodendrocytes16. Taken together with the findings from our study, we propose that microglia-derived TGFβ1 promotes myelin health in two distinct ways: (1) directly acting on oligodendrocytes to strengthen myelin integrity and (2) by acting on microglia as a checkpoint to maintain resilience to age-dependent myelin degeneration.
There are several unexplored findings from our current study that could be of interest for further investigation. We noted a sex difference in disease progression in the absence of microglial TGFβ signaling. Whether this is due to varying dependence on microglial TGFβ signaling or differences in damage resistance or spontaneous repair capacity between males and females warrants further investigation. Serpina3n+ DA-MOLs are gaining increasing interest as they emerge during the course of neurodegeneration and neuroinflammation, including during multiple sclerosis47,48,68. Whether they are passively responding to damage or are playing an active role in disease progression remains unclear. Despite being induced in demyelinating areas, the capacity for DA-MOLs to myelinate (at least in our context) appears abolished. It remains elusive whether the loss of myelination capacity is an inherent characteristic of DA-MOLs or if DA-MOLs upregulate prosurvival mechanisms, despite myelin synthesis being constantly disrupted by microglia lacking TGFβ signaling. A deeper characterization of this may shed light on our understanding of human diseases with insufficient remyelination. In agreement with previous findings, our snRNA-seq analyses also revealed that a subpopulation of microglia (MyTE) is enriched with myelin transcripts41,42. It is hypothesized that myelin transcripts can enter microglial nuclei following the phagocytosis of myelin through an as yet unidentified incorporation mechanism42. It will be intriguing to further explore how the incorporation of myelin transcripts in microglia affects their subsequent interaction with myelin and whether this process induces epigenetic reprogramming or innate immune training in microglia, rendering them more reactive to myelin. The interplay between microglia and CD8+ T cells in driving myelin degeneration in other CNS regions, such as the optic nerve, is gaining increasing attention69,70,71. Although we did not observe major infiltration of CD8+ T cells in the spinal cord of microglial Tgfbr2-deficient mice, we cannot completely rule out their potential involvement in our experimental paradigm. Previously, a white matter-associated microglial subset was reported to appear within white matter tracts in the brain in the context of aging or in response to demyelination6. While TSM can indeed be classified as a subset of white matter-associated microglial, the identification of this subset draws attention to spinal cord microglial diversity, which has long been overlooked in comparison with the focus on microglia in the brain, further deepening our understanding of microglial diversity within the white matter.
Overall, our study highlights the unique microenvironment of the DC during aging and underscores the critical role of TGFβ signaling in maintaining homeostatic microglia–myelin interactions in this region. We expect our findings to contribute to a deeper understanding of spatially distinct microglial heterogeneity and microglia–oligodendrocyte regulatory mechanisms. Furthermore, our results provide new insights into region-specific vulnerability to demyelination, shedding light on certain human diseases.
Methods
Ethics statement
All animal experiments complied with relevant ethical regulations and were approved by the North Stockholm Animal Ethics Committee under ethical permits N138/14, 8328-2019 and 23561-2022.
Experimental subjects
Experimental mice were bred and housed at the Comparative Medicine Department at Karolinska University Hospital, Sweden. Mice were maintained in climate-controlled and pathogen-free environment with regulated 12-h light/dark cycles. Cx3cr1CreER−YFP (#021160) and Tgfb1fl/fl (#065809) mice were obtained from the Jackson Laboratory. Tgfbr2fl/fl mice were kindly offered by M. Li at Sloan Kettering Institute. C57BL/6NTac mice (Taconic) were bred at the local animal facility. For tamoxifen administration, tamoxifen (T5648, Sigma) was dissolved in corn oil at 75 °C for 1 h. Mice were administered three injections of 4–5 mg tamoxifen in 200 µl corn oil (intraperitoneally) with a 1-day interval between each injection. The day mice received the last tamoxifen injection was counted as day 0 (D0). Most experiments used female mice of different ages to investigate age-related effects, while both males and females were included to compare disease progression and to analyze the spinal cord transcriptome using NanoString nCounter. For TEM analyses, female mice aged 2–3, 6–8, 12–13 and 16–18 months were used to examine age-related alterations in myelin ultrastructure. For NanoString nCounter analyses, we included both female and male mice aged 2–3 months or ~12 months to compare age-related molecular changes in the spinal cord, with the presented data obtained from female mice. For snRNA-seq, female mice were around 12 months old. To investigate age-related upregulation of TGFβ signaling in Fig. 2, young mice were 2–3 months old, while older mice were approximately 12 months or older. Microglia isolated from microdissected spinal cord regions for bulk RNA-sequencing analysis and flow cytometry were obtained from adult mice aged 4–5 months. Immunostaining and other flow cytometry results in Figs. 3–7 were obtained from Cx3cr1CreER:Tgfbr2fl/fl or Cx3cr1CreER:Tgfb1fl/fl mice aged 8–12 months. Detailed age information is provided in the figure legends. At the experimental endpoints (humane endpoint or latest 60 days post-tamoxifen administration), mice were euthanized by injection of pentobarbital sodium (intraperitoneally). No statistical methods were used to predetermine sample sizes but our sample sizes are similar to those reported in previous publications72,73,74. In all experiments, mice were randomly allocated to experimental groups.
TEM
Mice were perfused with 2.5% glutaraldehyde and 1% formaldehyde (>24 h) in 0.1 M phosphate buffer. Following fixation, the spinal cord segments were rinsed in 0.1 M phosphate buffer before postfixation in 2% osmium tetroxide in 0.1 M phosphate buffer, pH 7.4, at 4 °C for 2 h. The spinal cord segments were then dehydrated in ethanol followed by acetone and resin embedded in LX-112 (Ladd). Ultrathin sections (80–100 nm) were prepared using an EM UC7 (Leica) and subsequently stained with uranyl acetate and lead citrate for contrast. These sections were then analyzed with a Hitachi HT7700 transmission electron microscope (Hitachi High-Technologies) operating at 80 kV. Digital images of the sections were captured using a 2k × 2k Veleta CCD camera (Olympus Soft Imaging Solutions). The analysis for axon counts and axonal perimeter for each field of view of TEM images were automatically measured using the ‘Analyze Particles’ function in ImageJ/FIJI. The analyses for g-ratio calculation and periaxonal space were performed using MyelTracer75.
Immunofluorescence
Mouse spinal cord tissues were fixed with 4% paraformaldehyde for 24 h followed by dehydration in 20% and 30% sucrose at 4 °C. The dehydrated tissues were embedded with OCT cryo-mountant (45830, Histolab) and kept at −80 °C. OCT tissue chunks were sectioned (14 µm) in a Leica CM1850 cryostat and mounted on SuperFrost Plus Adhesion slides (J1800AMNZ, Epredia) kept at −20 °C. A mild antigen retrieval step was introduced by immersing the slides into boiled antigen retrieval solution (00-4955-58, Invitrogen) for 10 min, followed by washing with PBS and incubation with blocking buffer (10% normal goat or donkey serum and 0.3% Triton X-100 in PBS) for 1 h at room temperature. The following primary antibody were used: rabbit-anti-IBA1 (1:500, 019-19741, WAKO), goat-anti-IBA1 (1:400, ab5076, Abcam), rat-anti-MHC-II (1:200, 14-5321-82, Thermo), rat-anti-F4/80 (1:200, MCA497GA, Bio-Rad), rabbit-anti-GPNMB (1:500, ab188222, Abcam), rat-anti-Galectin-3 (1:200, 14-5301-82, Thermo), chicken-anti-PLP1 (1:200, NB100-1608, Novus Biologicals), mouse-anti-Tuj1 (1:100, 801213, BioLegend), rat-anti-MBP (1:500, MCA409S, Bio-Rad), R667-conjugated-anti-MBP (1:200, 130-131-152, Miltenyi Biotec), rabbit-anti-degraded myelin basic protein (1:1,000, AB5864, Millipore Sigma), rabbit-anti-PLIN2 (1:200, 15294-1-AP, Proteintech), mouse-anti-CC-1 (1:200, OP80, Millipore Sigma) and goat-anti-Serpina3n (1:200, AF4709, R&D Systems). Primary antibodies were diluted in staining buffer (1% BSA and 0.3% Triton X-100 in PBS) and incubated for overnight at 4 °C, followed by washing and staining with species-matched secondary antibodies for 1 h at room temperature. For Fluoromyelin (1:300, F34651, Invitrogen) and LipidSpot (1:1,000, 70069, Biotium) staining, sections were incubated for 30 min at room temperature. Nuclei staining was performed by incubating with Hoechst (62249, Thermo). Images were taken using a Zeiss LSM880 confocal microscope or Zeiss Axio Imager Z1 microscope, and images were analyzed using FIJI/ImageJ.
Microdissection of spinal cord tissue
After perfusion and collection of the spinal cords, they were either immersed in cold HBSS and microdissected right after for flow cytometry and cell sorting or immersed in RNAprotect Tissue Reagent (76104, Qiagen) for mRNA analyses. Microdissection was performed using a Leica MZ95 surgical microscope. The DC was gently pulled out using ophthalmic forceps curved with hook, and the VC was carefully dissociated using iris scissors and scalpels. GM was carefully separated with connecting white matter tissues and clipped out using iris scissors and tweezers.
Preparation of CNS single-cell suspensions
After perfusion, spinal cord tissue was flushed out from the spine and physically dissociated followed by enzymatic digestion with papain (1:100 diluted in L15 medium; LS003126, Worthington) and 0.2 mg ml−1 DNase I (10104159001, Roche). Myelin was removed using 38% isotonic Percoll (P1644, Sigma) by centrifugating at 800g (4× acceleration, no brake) for 10 min. Cell pellets were suspended for flow cytometry or cell sorting.
Flow cytometry
Spinal cord single-cell suspensions were stained with LIVE/DEAD Fixable Stain kits (1:1,000, L10119 or L34959, Invitrogen) to exclude dead cells. Data were acquired using a Cytek Aurora spectral flow cytometer or Beckman Coulter Gallios flow cytometer. The following panels of antibodies were used: (1) panel 1: CD11b-BV786 (clone M1/70, 417-0112-82, Thermo Fisher Scientific), CD45-AF488 (clone 30-F11, 103122, Biolegend), CX3CR1-AF700 (clone SA011F11, 149036, Biolegend), GPNMB-eFluor660 (clone CTSREVL, 50-5708-82, Thermo Fisher Scientific), Galectin-3-PE/Cy7 (clone M3/38, 125418, Biolegend), Dectin-1-PerCP-eFluor710 (clone bg1fpj, 46-5859-82, Thermo), CD74-BUV737 (clone In-1, 741798, BD), LAP/TGFβ1-BV421 (clone TW7-16B4, 141407, Biolegend), CD140a-BV480 (clone APA5, 746581, BD), O4-APC (clone REA576, 130-119-897, Miltenyi) and GLAST-PE (clone ACSA-1, 130-118-344, Miltenyi). Followed by surface marker staining, fixation and permeabilization of the cells for LAP/TGFβ1 staining were performed using the commercial kit (50-112-9060, Fisher Scientific) following the provided protocol. (2) Panel 2: CD11b-PerCP/Cy5.5 (clone M1/70, 101228, Biolegend), CD45-PE/Cy7 (clone 30-F11, 103114, Biolegend), Ly6C-PE (clone HK1.4, 128008, BioLegend), F4/80-APC (clone BM8, 123116, BioLegend), MHC-II-AF700 (clone M5/114.15.2, 107622, BioLegend). (3) Panel 3: CD11b-PerCP/Cy5.5 (clone M1/70, 101228, Biolegend), CD45-AF488 (clone 30-F11, 103122, Biolegend), Ly6C-AF700 (clone HK1.4, 128024, Biolegend), Ly6G-V450 (clone 1A8, 560603, BD) and Galectin-3-PE/Cy7 (clone M3/38, 125418, Biolegend). (4) Panel 4: CD11b-PerCP/Cy5.5 (clone M1/70, 101228, Biolegend), CD45-AF488 (clone 30-F11, 103122, Biolegend), Ly6C-AF700 (clone HK1.4, 128024, Biolegend), Ly6G-V450 (clone 1A8, 560603, BD), MHC-II-PE-Dazzle-594 (clone M5/114.15.2, 107648, Biolegend), MSR1-AF647 (clone 1F8C33, 154719, Biolegend), Galectin-3-PE (clone M3/38, 125405, Biolegend) and F4/80-BV421 (clone BM8, 123137, Biolegend). Flow cytometry data were analyzed using Kaluza or Flowjo 10 software. Dimensionality reduction and cluster identification were performed using the t-distributed stochastic neighbor embedding (t-SNE) plugin in Flowjo 10. A representative gating strategy for separating microglia and other neural cells is included in Extended Data Fig. 3.
Cell sorting
Single-cell suspensions from the microdissected spinal cord subregions (DC/VC/GM) were sorted using a BD Influx cell sorter. After myelin removal, the single-cell suspensions were stained with LIVE/DEAD Fixable Yellow Dead Cell Stain kit (1:1,000, L34959, Invitrogen) and surface antibodies: CD11b-PerCP/Cy5.5 (clone M1/70, 101228, Biolegend), CD45-PE/Cy7 (clone 30-F11, 103114, Biolegend), Ly6C-PE (clone HK1.4, 128008, BioLegend) and F4/80-APC (clone BM8, 123116, BioLegend). Microglia cells were sorted as CD11b+CD45+(int)Ly6C−F4/80+(int). Cell purity was determined to be >95% by checking the YFP expression of the gated Cx3cr1CreER-YFP cells. A total of 100 microglia from each region were collected in 5 µl single-cell lysis buffer (635013, Takara) and flash-frozen on dry ice.
Next-generation sequencing (bulk RNA-sequencing)
The sorted 100 cells from each spinal cord subregion were proceeded directly with cDNA synthesis using the SMART-Seq v4 Ultra Low Input RNA kit, and the library was prepared using the Nextera XT kit by staff at the Bioinformatics and Expression Analysis core facility at Karolinska Institutet. For each group, n = 3–5 biological samples were included. Next-generation sequencing was performed at the National Genomics Infrastructure at the Science for Life Laboratory using a NovaSeq 6000 S4 platform. Data normalization was performed using the DESeq2 method76. differential gene expression and KEGG_Pathway/GO enrichment analysis were performed using the online bioinformatic analysis platform developed by BGI (https://biosys.bgi.com), an R framework with installed R packages.
Nuclei isolation and snRNA-seq
Nuclei isolation was performed following a 10x Genomics protocol (CG000124) with modifications. Nuclei lysis buffer was freshly prepared: 10 mM Tris–HCl (T2194, Sigma), 10 mM NaCl (59222 C, Sigma), 3 mM MgCl2 (M1028, Sigma) and 0.1% Nonidet P40 substitute (74385, Sigma) in PBS. Spinal cord tissue (1-cm-long segment) was immersed in 2 ml lysis buffer and homogenized using a dounce homogenizer on ice until most nuclei were released when checking under a microscope. Homogenates were added with 5 ml Hibernate medium and passed through a 30 µm cell strainer, followed by centrifuging at 500g for 5 min at 4 °C. The pellets were washed twice by being resuspended in 10 ml washing buffer (2% BSA and 1 mM dithiothreitol (DTT) in PBS supplemented with RNase inhibitor) followed by centrifugation with the previous settings. After washing, the pellet was resuspended in 38% isotonic Percoll (P1644, Sigma) and centrifuged at 2,000g (4× acceleration, no brake) for 20 min at 4 °C. The floating myelin layer was carefully removed and the pellets were immediately resuspended in wash buffer in low-binding Eppendorf tubes with n = 3 samples for WT/D20/D30 conditions and n = 2 for D10 condition. Each sample contains pooled spinal cord cervical segments from two mice. Nuclei were counted and a final concentration of around 1,000 nuclei µl−1 was used. Library was constructed using the Chromium Next GEM Single Cell 3ʹ Reagent Kits v3.1 (Dual Index; CG000315 Rev), quality control was performed using a Bioanalyzer (Agilent), and the concentration was determined using a Qubit (Thermo Fisher) and KAPA Library Quantification kits (Roche) following the manufacturer’s protocols. Samples were sequenced on a NovaSeq 6000 (NovaSeq Control Software 1.7.5/RTA v3.4.4) with a 28nt(Read1)-10nt(Index1)-10nt(Index2)-90nt(Read2) setup using ‘NovaSeqStandard’ workflow in ‘SP’ mode flowcell. The Bcl to FastQ conversion was performed using bcl2fastq_v2.20.0.422 from the CASAVA software suite. The quality scale used is Sanger/phred33/Illumina 1.8+.
snRNA-seq preprocessing and unsupervised clustering
snRNA-seq data for each sample were preprocessed using the CellRanger count command in the CellRanger toolkit (version 7.2.0) provided by 10X Genomics for the alignment against the mouse reference genome (GRCm39), followed by filtering, barcode counting and unique molecular identifier counting to generate gene–barcode matrices. The Seurat package (version 5.1.0) was subsequently used to generate the gene expression matrix. For quality control, we kept high-quality cells with thresholds of 800–3,000 unique gene counts, and less than 5% mitochondrial counts. Additional quality control steps included ambient RNA removal using FastCAR (version 0.1.0) and doublet filtering with DoubletFinder (version 2.0.4)77,78. After quality control, the Seurat object of each sample was normalized using the SCTransform command and integrated using anchor-based CCA integration based on the 5,000 most variable genes for dimensional reduction and clustering. Principal component analysis was performed on the gene expression matrix and the shared nearest neighbor graph was constructed using the FindNeighbors command based on the top 50 principal components. The Louvian algorithm was applied on the nearest neighbor graph to find cell clusters using the FindClusters command with the resolution parameter of 0.2. The same principal components were used for dimensional reduction to generate the UMAPs for visualization.
Identification of cell types
Clusters of different cell types of a total of 182,019 cells were manually identified based on the expression of signature genes and annotated according to a harmonized atlas of the mouse spinal cord (Seqseek)79. Briefly, the FindAllMarkers command in the Seurat package was used to identify DEGs for each cluster. The cell type annotation for each cluster was confirmed by the elevated expression of known cell type signature genes (Extended Data Fig. 7): Astro for astrocytes (Aqp4, Gfap and Col23a1), MG for microglia (Ptprc, Itgam, Hexb, Mertk, Tgfbr1 and Csf1r), OPC for oligodendrocyte progenitor cells (Pdgfra, Cspg4, Olig2, Ptprz1 and Tnr), Pre_Oligo for pre-mature oligodendrocytes (Tcf7l2, Bcas1 and Fyn), Oligo for mature oligodendrocytes (Mbp, Plp1, St18, Prr5l, Mobp and Scd3), Endo for endothelial cells (Bsg, Pecam1 and Flt1), PC for pericytes (Vtn, Abcc9 and Rgs5), EPC for ependymal cells (Dnah12, Nnat and Cfap43), MC for meningeal cells (Lama1, Cemip and Cped1), CSF-cN for cerebrospinal fluid-contacting neurons (Pkd1l2, Pkd2l1 and Myo3b) and MN for motor neurons (Chat and Col6a6). Rbfox3, Syt1 and Snap25 were used as pan-neuron markers, in combination with Gad1/Gad2/Slc32a1 for inhibitory neurons and Slc17a6 for excitatory neurons.
Analysis of microglia cells and oligodendrocytes
Identified microglia (11,313 cells) and oligodendrocytes (49,112 cells) were extracted for the subsequent analyses. Principal component analysis was first performed on the gene expression matrix of extracted cells and the shared nearest neighbor graph was constructed with the top 20 principal components. The Louvian algorithm was applied on the nearest neighbor graphs to find subclusters, using a resolution parameter of 0.15 for microglia and 0.5 for oligodendrocytes. The same principal components were used for dimensional reduction to generate the UMAP for visualization. Genes specifically expressed in each subcluster were identified using the FindAllMarkers command in the Seurat package for annotation. Signature genes were defined as those with a log2 fold change >1 and an adjusted P value <0.001. Gene set enrichment analysis based on GO was conducted using the clusterProfiler package (version 4.12.6) with a significance cutoff of adjusted P value <0.05. For RNA velocity analysis80, loom files were generated for each sample. RNA velocity was performed using the scvelo (version 0.3.2) and scanpy (version 1.10.2) Python packages. Velocity plots were generated using the dynamical model.
NanoString nCounter analyses
The total RNA of the spinal cords was extracted using TRIzol Reagent (A33250, Invitrogen) following the manufacturer’s protocol. For each group, n = 3–5 biological samples were included. The concentration of the total RNA was measured using a Qubit RNA HS Assay kit (Q32852, Invitrogen) and samples were adjusted to an RNA concentration of 20 ng µl−1 before sending to KIGene Core Facility at Karolinska Institutet for analysis. The samples were first hybridized to the nCounter Mouse Glial Profiling Panel CodeSet (XT-CSO-M-GLIAL-12, NanoString). The CodeSet includes 757 probes for genes of interest, 13 probes for internal reference genes and 10 additional customized probes for Smad4, Smad7, Nrros, Tgfbr2, Tgfbr3, Smurf1, Tgfb2, Cxcr4, Adgre4 and Clec4b1. All the probes were single target-specific color-coded probes with around 100 bases in length and no reverse transcription or amplification process were needed. After hybridization, the target–probe mix was purified using an automated fluidic handing nCounter Prep Station and excessive probes were eliminated. The target-paired probes were subsequently immobilized in a sample cartridge, and the fluorescent reporter probes were analyzed using the nCounter Digital Analyzer. The mRNA reads were normalized with the 13 internal reference genes included in the CodeSet. Data normalization and further advanced analyses were performed using the NanoString nSolver Analysis Software following the provided guidelines. For differential gene expression analysis between two groups, a P value less than 0.05 was used as recommended.
Human spinal cord tissue analyses
Human spinal cords were collected at Montpellier Hospital from organ-donor patients in strict agreement with French bioethics laws (articles L1232-1 and -6) and after approval by the French institution for organ transplantation. Approval was granted by the French Agence de la biomédecine (no. SPGED19-3-4692; 2 April 2019) for human spinal cord samples. Informed consent was obtained from the families by the organ procurement organization for this study. Surgery was performed as previously described81, and the thoracolumbar segments were immediately placed in liquid nitrogen before processing for cryosectioning and spatial transcriptomics. To enable gene expression analyses, the spinal cords were cryosectioned at 16-μm thickness and alternately placed on Superfrost microscope glass slides (Thermo Fisher Scientific) and Visium spatial gene expression slides (10X Genomics), after which they were stored at −80 °C for no more than 14 days before being used. Spatial gene expression libraries were generated using the Visium Spatial Gene Expression Kit from 10X Genomics as previously described82. Briefly, sections were fixed for 30 min in methanol, stained with hematoxylin and eosin, and imaged using the Metafer Slide Scanning system (MetaSystems). In total, Visium Spatial Gene Expression libraries from 12 spinal cord sections were prepared following the manufacturer’s protocol. Libraries were sequenced using the Illumina platform (NovaSeq 6000 and NextSeq 2000). Sequenced ST libraries were processed using the Space Ranger pipeline (10X Genomics). Reads were aligned to the human reference genome (ENSEMBL genome assembly, release 93) and annotated using GRCh38-3.0.0 to obtain expression matrices. In total, 12 tissue sections were analyzed. The count matrix was enriched for protein-coding and long intergenic noncoding RNA genes. The count matrix was filtered for all hemoglobin-related genes, MALAT1 and mitochondrial and ribosomal protein-coding genes. Spots with fewer than 500 genes and genes expressed in fewer than five spots were excluded from the analysis of the three postconception time points. Analyses, including data normalization, dimensionality reduction and clustering, of ST data were performed jointly using the Seurat and STUtility packages. Normalization was conducted using variance stabilizing transformation (SCTransform). Genes of interest were plotted across all the tissue sections.
Clinical scoring
Clinical symptoms were examined after tamoxifen administration. We assessed weight loss, tail tonus, tail lifting, gait pattern, dragging of legs, incontinence, paralysis, fur abnormality and breathing abnormality. Mice were assigned one point for the presence of each of the indicated symptom. Mice that reached humane endpoint or died were scored 15 from the day they were euthanized or died.
Hanging wire test
The four-paw hanging wire test was performed once per week. Mice were placed on a wire cage lid and inverted over a cage with soft beddings. The time until each mouse fell from the lid was recorded. The cutoff time was set to 180 s and normal mice could all reach this time. Two trials were performed for each mouse if the mouse did not stay until 180 s during the first trial, and the longest time was recorded.
RT–qPCR
Microdissected DC, VC and GM were processed using a RNeasy Lipid Tissue Mini kit (74804, Qiagen) to extract total RNA with on-column DNase digestion using a RNase-Free DNase set (79254, Qiagen). cDNA was synthesized using an iScript kit (1708891, Bio-Rad) and quantitative PCR with reverse transcription (RT–qPCR) was performed with SYBR green reactions (1708886, Bio-Rad) in a 384-well plate run in a Bio-Rad CFX384 Touch Real-Time PCR Detection System. The sequences (5′ to 3′) for the predesigned SYBR green primers (Sigma) used were as follows: Smad7 (FW): CTCTGTGAACTAGAGTCTCC; Smad7 (RE): GAAGTTGGGAATCTGAAAGC; Tgfb1 (FW): GGATACCAACTATTGCTTCAG; Tgf1 (RE): TGTCCAGGCTCCAAATATAG; Tgfbr1 (FW): CCTGAAGTTCTAGATGATTCC; Tgfbr1 (RE): CTTCATGGATTCCACCAATAG; Tgfbr2 (FW): CCAGGATGAATCTGGAAAAC; Tgfbr2 (RE): TAATCCTTCACTTCTCCCAC; Tgfbr3 (FW): GGTAGCTGTAGACAAAGATTC; Tgfbr3 (RE): CTGCACCACAATAGAGTTATAG; Nrros (FW): CATTGAATCCTCTGGAACTG; Nrros (RE): CTCCATTCCACAATCAAGAAG; Smad4 (FW): CCAGCTACTTACCATCATAAC; Smad4 (RE): GAGCAGGATGATTGGAAATG; Smurf2 (FW): CTAGCAGAGAGAAGAGTCAG; Smurf2 (RE): GTCCTTTGTTCATAGCCTTC; Smurf1 (FW): CCAAATAGTGGTCAGTTTACAG; Smurf1 (RE): CAGTACCATCTGTATATGGGG.
Western blotting
Spinal cords were homogenized and sonicated in RIPA buffer (R0278, Sigma) supplemented with protease/phosphatase inhibitors (1861261, Thermo) and phenylmethylsulfonyl fluoride (36978, Thermo). Homogenates were centrifuged (11,000g for 20 min at 4 °C) and the supernatants were determined using a BCA protein assay kit (23225, Thermo). Samples with equal amounts of proteins were added to 4× Laemmli sample buffer (1610747, Bio-Rad) supplemented with DTT (10197777001, Sigma) and electrophoresed in 4–20% Mini Protean TGX precast protein gels (4561093, Bio-Rad), followed by transferring to a 45 mm nitrocellulose membrane (1620115, Bio-Rad). After blocking with 5% non-fat dried milk in TBS-T (20 mM Tris–HCl and 137 mM NaCl with 0.1% Tween-20 at pH 7.6) for 30 min at room temperature, the membrane was incubated overnight at +4 °C with the following primary antibodies: anti-TGFβR2 (1:1,000, SAB5700739, Sigma), anti-TGFB1 (1:1,000, MA121595, Thermo), anti-SMAD2/3 (1:1,000, 8685 T, CST) and anti-phospho SMAD2 (1:1,000, AB3849-I, MERCK). After rinsing, the blots were incubated with HRP-linked secondary antibodies for 2 h at room temperature The blots were incubated with Clarity Western ECL (170-5061, Bio-Rad) and chemiluminescence was detected using a Bio-Rad Chemidoc Imaging System. Band intensity was quantified using the Bio-Rad Image Lab software, and data were normalized to β-actin as the reference protein.
Spinal cord wholemount staining and imaging
Spinal cord staining was performed following the iDISCO+ protocol as previously described83. Perfused spinal cord sections were dehydrated in an increasing series of 60%, 80%, 100% and 100% methanol (Sigma-Aldrich) diluted in water (each wash lasting 1 h). The samples were then chilled at 4 °C, followed by an overnight incubation in 66% dichloromethane/methanol (DCM; Sigma-Aldrich). The samples were washed twice in 100% methanol for 30 min and then bleached overnight at 4 °C in a 5% hydrogen peroxide (Sigma-Aldrich) methanol solution. Rehydration was achieved by incubating the samples in a series of 80%, 60%, 40% and 20% methanol (1 h each) followed by 1 h in PBS at room temperature with shaking. Samples were then washed for 1 h in PBS containing 0.2% Triton X-100 (PTx.2, Sigma-Aldrich) and permeabilized by a 24 h incubation at 37 °C in permeabilization solution, composed of 20% dimethyl sulfoxide (DMSO; Sigma-Aldrich) and 2.3% glycine (Sigma-Aldrich) in PTx.2. Samples were first blocked in blocking solution, containing 6% donkey serum (Jackson ImmunoResearch) and 10% DMSO in PTx.2 for 24 h at 37 °C. The primary antibodies were added to PTwH (PBS containing 0.2% Tween-20 (Sigma-Aldrich) and 0.1% of a 10 mg ml−1 heparin stock solution (Sigma-Aldrich)). The samples were incubated in the primary antibody anti-Iba-1 (1:200; Wako, 019-19741) for 3 days at 37 °C, then washed in PTwH (four to five times until the next day) and finally incubated for 2 days with the secondary antibody (donkey anti-rabbit IgG, Alexa Fluor 555; Invitrogen, A-31572) and the primary antibody anti-MBP (1:50; Miltenyi, 130-131-152). After immunostaining, the samples were washed in PTwH (four to five times until the next day), embedded in 3% gelatin in PBS and dehydrated in an increasing methanol/water concentration series (20%, 40%, 60%, 80%, 100% and 100% for 1 h each and 100% overnight at room temperature), followed by a wash in 66% DCM in methanol for 3 hs at room temperature. Methanol was washed out with two 15-min washes in 100% DCM and samples were cleared in dibenzyl ether (Sigma-Aldrich) overnight and finally transferred into ethyl cinnamate (Sigma-Aldrich) for at least 1 day. The prepared spinal cords were affixed onto a holder and carefully positioned within a sample chamber saturated with ethyl cinnamate, undergoing imaging procedures with the UltraMicroscope Blaze system (Miltenyi Biotec). The instrument operated in Ultramicroscope Expert mode, incorporating a Multi Color 3D Mosaic measurement mode and Fast Tiling scan mode. Tailoring to varying sample sizes, a singular left light sheet illumination was chosen, with the objective set at 4× and zoom at 2.5×. The numerical aperture was 0.19. For the imaging of Iba1 protein, a laser of excitation wavelength 561 nm at 60% power was employed, while MBP protein imaging utilized a 640 nm laser at 80% power. To mitigate phototoxicity, the camera exposure time was minimized to 5 ms. Employing a z-stack imaging approach with a 2 µm step size, scanning spanned from the layer preceding sample visibility to the layer postsample disappearance, ensuring comprehensive image capture. The resulting pixel size, based on the specified parameters, measured 0.65 µm × 0.65 µm × 2 µm. To ensure uniformity, all parameters and settings were standardized across all samples. Following imaging, MACS iQ View (3D large volume software, version 1.0.0) was employed to stitch and reconstruct wholemount 3D images for further analysis.
RNAscope
RNAscope was performed using the RNAscope Multiplex Fluorescent Detection Reagents kit v2 (323136, ACD). Briefly, slides were washed in 1× PBS for 5 min and baked in the HybEZ Oven for 30 min at 60 °C, followed by dehydration with 50%, 70% and twice with 100% ethanol for 5 min at room temperature. Hydrogen peroxide (322335, ACD) was applied to the sections to quench endogenous peroxidase activity for 10 min at room temperature, followed by wash with MilliQ water. Target retrieval was performed by submerging the slides into 98 °C 1× codetection target retrieval solution (322000, ACD) for 5 min, followed by wash with MilliQ water and dehydration with 100% ethanol for 5 min at room temperature. A barrier was created around the tissue sections using an ImmEdge Pen (H-4000; Vector laboratories). Protease digestion was performed using Protease III (322337, ACD) for 30 min at 40 °C in the HybEZ Oven, followed by wash with MilliQ water. Then, 1× mouse Tgfb1-C2 probe (Mm-Tgfb1-no-XHs-C2, 443571-C2, ACD) was applied to the sections for hybridization at 40 °C in the HybEZ Oven for 2 h, followed by a wash with 1× wash buffer (310091, ACD). Probe amplification was performed sequentially with reagents AMP1, AMP2 and AMP3 for 30, 30 and 15 min, respectively, at 40 °C in the HybEZ Oven, with a MilliQ water rinse between each amplification cycle. Following the final wash with 1× wash buffer after the AMP3 cycle, HRP-C2 was applied for 15 min at 40 °C, after which the slides were washed in wash buffer and incubated with TSA Vivid dye 520 (7523, R&D) diluted 1: 2,000. After rinsing with 1× wash buffer, the HRP blocker solution was applied for 15 min at 40 °C, followed by a rinse with 1× wash buffer. The slides were then incubated with blocking buffer (10% goat serum and 0.3% Triton X-100 in PBS) for 1 h at room temperature, followed by incubation with rabbit-anti-IBA1 antibody (1:100, 019-19741, WAKO) for overnight at 4 °C. After washing in PBS and incubation with secondary antibody, the slides were then mounted with Prolong Glass antifade moutant (P36984, Thermo Fisher Scientific).
Myelin purification and labeling
Pure myelin was obtained using a previous protocol from our laboratory20. Briefly, myelin was isolated by mechanically homogenizing perfused brains in a homogenization buffer with PBS containing 0.32 M sucrose. After two washes with the homogenization buffer, a 0.85 M sucrose solution in PBS was added as an underlay to the CNS homogenate. The gradient was then centrifuged at 4,500g for 50 min. The myelin-containing interphase was collected and washed twice with water. The purified myelin was subsequently incubated with pHrodo-red dye (P36600, Fischer Scientific) as per the manufacturer’s instructions, followed by a final wash.
Time-lapse live-cell imaging
Primary microglia were seeded at a density of 15,000 cells per well in an Incucyte ImageLock 96-well microplate (BA-04856, Sartorius) and incubated with pHrodo-red-labeled myelin. Plates were immediately transferred to an incubator connected to an Incucyte S3 live-cell analysis instrument (Sartorius) in which wells were analyzed using 20× ocular magnification every hour for 24 h, generating time-lapse images/movies (excitation wavelength 585 nm and emission wavelength 635 nm). Cell confluence mask and red signal intensity were quantified and acquired from the Incucyte S3 software.
Statistical analyses
Analyses and graph generation were performed using GraphPad Prism 10 software. Normality and equal variances were formally tested before statistical analysis. Comparisons between two groups were performed using Student’s two-tailed unpaired or paired t-test, or the Mann–Whitney U test or Wilcoxon matched-pairs signed-rank test, as appropriate. For multiple group comparisons, a one-way ANOVA or the Kruskal–Wallis test was employed. Two-way ANOVA with Tukey’s post hoc test was used for comparisons involving two independent variables. P < 0.05 was considered statistically significant. Analysis of RNA-sequencing data was done with the DESeq2 package using Wald testing with Benjamini–Hochberg adjustment for multiple testing. Detailed statistical information is provided in the figure legends. Data collection and analysis were performed in a blinded manner, and no data points were manually excluded after collection.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Bulk RNA-sequencing data of microdissected microglia and NanoString nCounter raw data were uploaded to NCBI GEO database under the accession numbers GSE254915 and GSE283721, respectively. snRNA-seq data are available at BioProject with the accession number PRJNA1185825. The raw sequencing data related to genes in the TGFβ signaling pathway of the human spinal cord have been deposited to the European Genome-phenome Archive (EGA) with the accession number EGAD50000002049 (restricted until the end of 2027); in accordance with the European Data Protection Regulation (GDPR) and Karolinska Institutet’s data governance policy, these individual-level human sequencing data will be accessible upon request from the corresponding author, subject to institutional and ethical approval procedures. Source data are provided with this paper.
Code availability
The codes used for snRNA-seq analyses are available via Github at https://github.com/Kenny19920122/TGFb_snRNAseq.
References
Masuda, T., Sankowski, R., Staszewski, O. & Prinz, M. Microglia heterogeneity in the single-cell era. Cell Rep. 30, 1271–1281 (2020).
Easley-Neal, C., Foreman, O., Sharma, N., Zarrin, A. A. & Weimer, R. M. CSF1R ligands IL-34 and CSF1 are differentially required for microglia development and maintenance in white and gray matter brain regions. Front. Immunol. 10, 2199 (2019).
Li, Q. et al. Developmental heterogeneity of microglia and brain myeloid cells revealed by deep single-cell RNA sequencing. Neuron 101, 207–223.e10 (2019).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e6 (2019).
Grabert, K. et al. Microglial brain region-dependent diversity and selective regional sensitivities to aging. Nat. Neurosci. 19, 504–516 (2016).
Safaiyan, S. et al. White matter aging drives microglial diversity. Neuron 109, 1100–1117.e10 (2021).
Safaiyan, S. et al. Age-related myelin degradation burdens the clearance function of microglia during aging. Nat. Neurosci. 19, 995–998 (2016).
Vallstedt, A., Klos, J. M. & Ericson, J. Multiple dorsoventral origins of oligodendrocyte generation in the spinal cord and hindbrain. Neuron 45, 55–67 (2005).
Cai, J. et al. Generation of oligodendrocyte precursor cells from mouse dorsal spinal cord independent of Nkx6 regulation and Shh signaling. Neuron 45, 41–53 (2005).
Tripathi, R. B. et al. Dorsally and ventrally derived oligodendrocytes have similar electrical properties but myelinate preferred tracts. J. Neurosci. 31, 6809–6819 (2011).
Floriddia, E. M. et al. Distinct oligodendrocyte populations have spatial preference and different responses to spinal cord injury. Nat. Commun. 11, 5860 (2020).
Crawford, A. H., Tripathi, R. B., Richardson, W. D. & Franklin, R. J. M. Developmental origin of oligodendrocyte lineage cells determines response to demyelination and susceptibility to age-associated functional decline. Cell Rep. 15, 761–773 (2016).
Boda, E. et al. Molecular and functional heterogeneity in dorsal and ventral oligodendrocyte progenitor cells of the mouse forebrain in response to DNA damage. Nat. Commun. 13, 2331 (2022).
Lund, H. et al. Fatal demyelinating disease is induced by monocyte-derived macrophages in the absence of TGF-β signaling. Nat. Immunol. 19, 1–7 (2018.
Qin, Y. et al. A milieu molecule for TGF-β required for microglia function in the nervous system. Cell 174, 156–171.e16 (2018).
McNamara, N. B. et al. Microglia regulate central nervous system myelin growth and integrity. Nature 613, 120–129 (2023).
Utz, S. G. et al. Early fate defines microglia and non-parenchymal brain macrophage development. Cell 181, 557–573.e18 (2020).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Wang, F. et al. Myelin degeneration and diminished myelin renewal contribute to age-related deficits in memory. Nat. Neurosci. 23, 481–486 (2020).
Berglund, R. et al. Microglial autophagy-associated phagocytosis is essential for recovery from neuroinflammation. Sci.Immunol. 2020, eabb5077 (2020).
Saliani, A. et al. Axon and myelin morphology in animal and human spinal cord. Front. Neuroanat. 11, 129 (2017).
Duval, T. et al. Axons morphometry in the human spinal cord. Neuroimage 185, 119–128 (2019).
Deczkowska, A., Amit, I. & Schwartz, M. Microglial immune checkpoint mechanisms. Nat. Neurosci. 21, 779–786 (2018).
Buttgereit, A. et al. Sall1 is a transcriptional regulator defining microglia identity and function. Nat. Immunol. 17, 1397–1406 (2016).
Deerhake, M. E. & Shinohara, M. L. Emerging roles of Dectin-1 in noninfectious settings and in the CNS. Trends Immunol. 42, 891–903 (2021).
Marschallinger, J. et al. Lipid-droplet-accumulating microglia represent a dysfunctional and proinflammatory state in the aging brain. Nat. Neurosci. 23, 194–208 (2020).
Gouna, G. et al. TREM2-dependent lipid droplet biogenesis in phagocytes is required for remyelination. J. Exp. Med. 218, e20210227 (2021).
Haney, M. S. et al. APOE4/4 is linked to damaging lipid droplets in Alzheimer’s disease microglia. Nature 628, 154–161 (2024).
Spittau, B., Dokalis, N. & Prinz, M. The role of TGFβ signaling in microglia maturation and activation. Trends Immunol. 41, 836–848 (2020).
Zöller, T. et al. Silencing of TGFβ signalling in microglia results in impaired homeostasis. Nat. Commun. 9, 4011 (2018).
Krasemann, S. et al. The TREM2–APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581.e9 (2017).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Takahashi, J. L., Giuliani, F., Power, C., Imai, Y. & Yong, V. W. Interleukin-1β promotes oligodendrocyte death through glutamate excitotoxicity. Ann. Neurol. 53, 588–595 (2003).
Nakazawa, T. et al. Tumor necrosis factor-α mediates oligodendrocyte death and delayed retinal ganglion cell loss in a mouse model of glaucoma. J. Neurosci. 26, 12633–12641 (2006).
Guilliams, M. et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell 185, 379–396.e38 (2022).
Marsh, S. E. et al. Dissection of artifactual and confounding glial signatures by single-cell sequencing of mouse and human brain. Nat. Neurosci. 25, 306–316 (2022).
Hendrickx, D. A. E. et al. Gene expression profiling of multiple sclerosis pathology identifies early patterns of demyelination surrounding chronic active lesions. Front. Immunol. 8, 1810 (2017).
Karlsson, A. et al. Galectin-3 functions as an opsonin and enhances the macrophage clearance of apoptotic neutrophils. Glycobiology 19, 16–20 (2008).
Bernal-Chico, A. et al. Blockade of monoacylglycerol lipase inhibits oligodendrocyte excitotoxicity and prevents demyelination in vivo. Glia 63, 163–176 (2015).
Hernández-Torres, G. et al. A reversible and selective inhibitor of monoacylglycerol lipase ameliorates multiple sclerosis. Angew. Chem. Weinh. Bergstr. Ger. 126, 13985–13990 (2014).
Depp, C. et al. Myelin dysfunction drives amyloid-β deposition in models of Alzheimer’s disease. Nature 618, 349–357 (2023).
Schirmer, L. et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573, 75–82 (2019).
Cantuti-Castelvetri, L. et al. Defective cholesterol clearance limits remyelination in the aged central nervous system. Science 359, 684–688 (2018).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Foerster, S. et al. Developmental origin of oligodendrocytes determines their function in the adult brain. Nat. Neurosci. 27, 1545–1554 (2024).
Kukanja, P. et al. Cellular architecture of evolving neuroinflammatory lesions and multiple sclerosis pathology. Cell 187, 1990–2009.e19 (2024).
Pandey, S. et al. Disease-associated oligodendrocyte responses across neurodegenerative diseases. Cell Rep. 40, 111189 (2022).
Castelo-Branco, G., Kukanja, P., Guerreiro-Cacais, A. O. & Rubio Rodríguez-Kirby, L. A. Disease-associated oligodendroglia: a putative nexus in neurodegeneration. Trends Immunol. 45, 750–759 (2024).
Bedolla, A. et al. Adult microglial TGFβ1 is required for microglia homeostasis via an autocrine mechanism to maintain cognitive function in mice. Nat. Commun. 15, 5306 (2024).
Sun, H. Y., Lee, J. W., Park, K. S., Wi, J. Y. & Kang, H. S. Spine MR imaging features of subacute combined degeneration patients. Eur. Spine J. 23, 1052–1058 (2014).
Yoon, J. Y. & Klein, J. P. Subacute combined degeneration from nitrous oxide use. N. Engl. J. Med. 387, 832 (2022).
Nakamura, T. et al. A case of subacute combined degeneration of the spinal cord due to folic acid and copper deficiency. Brain Dev. 41, 111–115 (2019).
Zhou, J., Zhang, H., Tang, K., Liu, R. & Li, J. An updated review of recent advances in neurosyphilis. Front. Med. 9, 800383 (2022).
Li, W. et al. Increased CSF soluble TREM2 concentration in patients with neurosyphilis.Front. Neurol. 11, 576 (2020).
Marques, S. et al. Transcriptional convergence of oligodendrocyte lineage progenitors during development. Dev. Cell 46, 504–517.e7 (2018).
Bechler, M. E., Swire, M. & Ffrench-Constant, C. Intrinsic and adaptive myelination-A sequential mechanism for smart wiring in the brain. Dev. Neurobiol. 78, 68–79 (2018).
Lee, S. et al. A culture system to study oligodendrocyte myelination processes using engineered nanofibers. Nat. Methods 9, 917–922 (2012).
Lampron, A. et al. Inefficient clearance of myelin debris by microglia impairs remyelinating processes. J. Exp. Med. 212, 481–495 (2015).
Yin, Z. et al. APOE4 impairs the microglial response in Alzheimer’s disease by inducing TGFβ-mediated checkpoints. Nat. Immunol. 24, 1839–1853 (2023).
Victor, M. B. et al. Lipid accumulation induced by APOE4 impairs microglial surveillance of neuronal-network activity. Cell Stem Cell 29, 1197–1212.e8 (2022).
Bosch-Queralt, M. et al. Diet-dependent regulation of TGFβ impairs reparative innate immune responses after demyelination. Nat. Metab. 3, 211–227 (2021).
Islam, A. et al. Sustained anti-inflammatory effects of TGF-β1 on microglia/macrophages. Biochim. Biophys. Acta Mol. Basis Dis. 1864, 721–734 (2018).
Simons, M., Levin, J. & Dichgans, M. Tipping points in neurodegeneration. Neuron 111, 2954–2968 (2023).
Brown, G. C. Cell death by phagocytosis. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-023-00921-6 (2023).
Butler, C. A. et al. Microglial phagocytosis of neurons in neurodegeneration, and its regulation. J. Neurochem. 158, 621–639 (2021).
Kim, B.-G., Malek, E., Choi, S. H., Ignatz-Hoover, J. J. & Driscoll, J. J. Novel therapies emerging in oncology to target the TGF-β pathway. J. Hematol. Oncol. 14, 55 (2021).
Liu, S., Ren, J. & Ten Dijke, P. Targeting TGFβ signal transduction for cancer therapy. Signal Transduct. Target. Ther. 6, 8 (2021).
Kenigsbuch, M. et al. A shared disease-associated oligodendrocyte signature among multiple CNS pathologies. Nat. Neurosci. 25, 876–886 (2022).
Groh, J. et al. Accumulation of cytotoxic T cells in the aged CNS leads to axon degeneration and contributes to cognitive and motor decline. Nat. Aging 1, 357–367 (2021).
Kaya, T. et al. CD8+ T cells induce interferon-responsive oligodendrocytes and microglia in white matter aging. Nat. Neurosci. 25, 1446–1457 (2022).
Groh, J. et al. Microglia activation orchestrates CXCL10-mediated CD8+ T cell recruitment to promote aging-related white matter degeneration. Nat. Neurosci. 28, 1160–1173 (2025).
Zhu, K. et al. Myeloid cell-specific topoisomerase 1 inhibition using DNA origami mitigates neuroinflammation. EMBO Rep. 23, e54499 (2022).
Lund, H. et al. CD163+ macrophages monitor enhanced permeability at the blood-dorsal root ganglion barrier. J. Exp. Med. 221, e20230675 (2024).
Sun, J.-X. et al. Activation of TRPV1 receptor facilitates myelin repair following demyelination via the regulation of microglial function. Acta Pharmacol. Sin. 44, 766–779 (2023).
Kaiser, T. et al. MyelTracer: a semi-automated software for myelin g-ratio quantification. eNeuro 8, ENEURO.0558-20.2021 (2021).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Berg, M. et al. FastCAR: fast correction for ambient RNA to facilitate differential gene expression analysis in single-cell RNA-sequencing datasets. BMC Genomics 24, 722 (2023).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e4 (2019).
Russ, D. E. et al. A harmonized atlas of mouse spinal cord cell types and their spatial organization. Nat. Commun. 12, 5722 (2021).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bauchet, L. et al. Isolation and culture of precursor cells from the adult human spinal cord. Methods Mol. Biol. 1059, 87–93 (2013).
Li, X. et al. Profiling spatiotemporal gene expression of the developing human spinal cord and implications for ependymoma origin. Nat. Neurosci. 26, 891–901 (2023).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Acknowledgements
We thank A. O. Guerreiro-Cacais for assistance with the application of ethical permits and the staff at our animal facility (Avdelningen för Komparativ Medicin, AKM) for their care of the animals. We express our gratitude to J. Han, X. M. Zhang, I. Benito-Cuesta, M. Khademi and S. Lewandowski at Karolinska Institutet for their invaluable support with experimental materials, laboratory management and participation in scientific discussions. We appreciate T. Sun’s input on the myelin-DAM signature. We also thank A. van Vollenhoven (Karolinska Institutet) for assistance with cell sorting, and L. Haag at EMil for help with electron microscopy. In addition, we appreciate A. Singh’s help with data mining and R. Fan for scientific inputs. Our thanks extend to the KIGene staff (K. Gell, A. Eriksson and M. Alvehus) and J. Zhang for their contributions in setting up the NanoString analysis. We are grateful to F. Fagerström, S. Fält and D. Brodin at the Bioinformatics and Expression Analysis facility (Karolinska Institutet) for their support with RNA-sequening setup. We acknowledge the support from the National Genomics Infrastructure in Stockholm, funded by Science for Life Laboratory, the Knut and Alice Wallenberg Foundation and the Swedish Research Council. We also thank SNIC/Uppsala Multidisciplinary Center for Advanced Computational Science for their assistance with massively parallel sequencing and access to the UPPMAX computational infrastructure. Special thanks to M. Li (Sloan Kettering Institute) for providing the Tgfbr2fl/fl mice. Many thanks to S. B. Ma for the support during the difficult times in San Francisco. We acknowledge funding support for our research from the Swedish Research Council/Vetenskapsrådet 2021-01926 (to R.A.H.), 2024-02509 (to H.L.) and 2023-06516 (to K.Z.), O.E. och Edla Johanssons vetenskapliga stiftelse (to H.L.), Foundation for Geriatric Diseases at Karolinska Institutet (to H.L.), Loo and Hans Osterman Foundation for Medical Research (to H.L.), Svenska Sällskapet för Medicinsk Forskning PD21-0017 (to H.L.), StratNeuro (to H.L.), Alltid Litt Sterkere (to R.A.H.), Neurofonden (to H.L. and K.Z.), China Scholarship Council 201700260280 (to K.Z.) and NIH-NINDS R01NS128021 and R21NS133891 (to S.P.J.F.).
Funding
Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Contributions
H.L. and K.Z. conceived the study, with inputs from R.A.H.; and H.S. H.L., K.Z. and R.A.H. designed the experiments. K.Z. performed most of the experiments and analyzed the data under the guidance of H.L. and R.A.H., with assistance from H.L. (mouse breeding, cell sorting and sample preparation for RNA-sequencing). All authors contributed substantially to the final manuscript and discussed the results: Y. Liu (single-cell data processing and analysis); V.J. (single-cell library preparation); J.-H.M. (mouse breeding and scoring); Y. Li, J.C.K., S.K. and P.U. (wholemount imaging and image analysis); E.M. and X.L. (human spinal cord data analysis); Z.A., J.-P.H. and J. Lundeberg (human spinal cord sample preparation); M.P. (mouse breeding and scoring); V.S. (cryosectioning and RT–PCR validation); J. Lin and Y.X. (immunostaining, mouse breeding and genotyping); H.S. (mouse scoring, flow cytometry, pHrodo-red labeling of myelin and western blotting); Y.G. (nuclei extraction); W.X. (RNAscope); and S.P.J.F. (inputs in oligodendrocyte biology).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Differences in axon-myelin properties between DC and VC.
a-c TEM analysis showing the density of myelinated axons (b) and the average axon size (c) in the DC and VC of mice aged 2-3 months, 12 months, and 16-18 months; scale bar=20μm; n = 10, 6, and 9 mice for the 2-3 mo, 12 mo, 16-18 mo age groups, respectively. d-f Immunostaining of MBP (red) and dMBP (green) in the DC and VC across age groups, with quantification of dMBP+ area in (e) and an inset of the DC of 18-23 month mice in (f); scale bar=50μm; n = 6 mice per group. Data are presented as mean ± SEM; *p < 0.05, **p < 0.01, ***p < 0.001; two-way ANOVA with Tukey’s post-hoc test.
Extended Data Fig. 2 Spatiotemporal alterations in TGF-β pathway genes in the spinal cord.
a Gene Ontology (GO) enrichment analyses of the DEGs from NanoString nCounter analysis between YM (2-3 months) and OM (12 months) reveal the Top 10 enriched Biological Processes (BP), Cellular Components (CC), and Molecular Functions (MF), while WikiPathway analysis highlighted all the enriched pathways. The bar charts represent the False Discovery Rate (FDR) in -log10 scale. b Heatmap showing the mRNA expression of genes in the TGF-β signaling pathway in mouse spinal cord from NanoString nCounter analysis. The gradient color represents the z-score of each gene across different samples, with red indicating higher expression and blue indicating lower expression of the gene across samples. c Bubble plot showing the mRNA expression of genes in the TGF-β signaling pathway in human spinal cord of subjects with varied ages. Bubble size represents the percentage of cells expressing the gene, and color scale bar represents the normalized expression value. d qRT-PCR results showing the mRNA expression of genes in the TGF-β signaling pathway across microdissected DC, VC and grey matter (GM) regions in young (2-3 months) and older mice (6-8 months); n = 9 mice for GM and DC, and n = 8 mice for VC. e, f Flow cytometry validation of microglial surface markers was conducted to compare 2-3-month-old (YM) and 12-13-month-old (OM) mice, including quantification (f) of the frequency of marker-positive cells within the total microglia population; n = 6 mice. Data are presented as mean ± SEM; *p < 0.05, **p < 0.01, ***p < 0.001. Multiple Mann-Whitney tests with FDR correction were used for comparisons between the age groups within each region in (d), and Kruskal-Wallis tests with Dunn’s post-hoc test were used for cross-region comparisons within each age group; two-tailed Mann-Whitney test or unpaired t-test was used in (f).
Extended Data Fig. 3 Flow cytometric analysis of the major neural cell types in the spinal cord.
a Flow cytometry gating strategy for separating major neural cell types in the spinal cord. b A flow cytometric contour plot of a BV421-TGF-β1 fluorescence-minus-one (FMO) control was used to set the TGF-β1⁺ gate. c Representative flow cytometric contour plots showing gated TGF-β1⁺ populations across cell types in 2–3-month-old and 12-13-month-old mice.
Extended Data Fig. 4 Transcriptomic analysis of microglia in the spinal cord following microglial Tgfbr2 deletion.
a A Venn diagram displays the number of differentially expressed genes (DEGs) in sorted microglia between Cx3cr1CreER:Tgfbr2fl/fl (KO) mice at 18 (D18) or 28 (D28) days post-tamoxifen administration, compared to Tgfbr2fl/fl (WT) mice. DEGs were identified using DESeq2 normalization (Log2 fold change≥2, q-value < 0.01). There were 668 DEGs at D18 compared to WT and 585 DEGs at D28 compared to WT. The 416 overlapping genes were defined as the core DEGs. b Top 50 hub genes from the 416 core DEGs and their interacting network were revealed using Cytohubba with the MCC algorithm. Genes in red bubbles represent upregulated genes following microglial Tgfbr2 depletion (compared to the WT group), while genes in blue bubbles represent downregulated genes. c Representative gene groups that were suppressed and induced following microglial Tgfbr2 deletion, compared to WT microglia. d, e The Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology Biological Process (GO_BP) enrichment analyses of the 416 core DEGs were performed using the Dr. Tom analysis platform (BGI), which employs the phyper function in R for two-sided hypergeometric testing. P values were adjusted for multiple comparisons using the Benjamini–Hochberg method to obtain q values, and pathways with q < 0.05 were considered significantly enriched. In the bubble plots, bubble size indicates the number of DEGs, while color represents the q value (red = low, blue = high). The ‘rich ratio’ denotes the proportion of DEGs annotated to each term relative to the total number of genes annotated to that term.
Extended Data Fig. 5 Profiling of molecular signatures in the spinal cord following microglial Tgfbr2 deletion.
a Schematic illustration of the experiment: spinal cords from 12-13-month-old Tgfbr2fl/fl (WT) and Cx3cr1CreER:Tgfbr2fl/fl mice (KO-D10, KO-D20, KO-D30 denote 10, 20, or 30 days post-tamoxifen injection, respectively) were homogenized, and total RNA was extracted for NanoString Glial Profiling Panel analysis, which covers 780 genes across different biological pathways involved in glial cell biology and neuronal regulation. b Unbiased Principal component (PC) analysis based on the normalized mRNA counts of all samples; n = 3-5 samples per group. c-e Volcano plots of mRNA count data show each gene’s -log₁₀(p) value versus its log2 fold change in the spinal cord of Cx3cr1CreER:Tgfbr2fl/fl mice at different time points following microglial Tgfbr2 deletion, compared with WT controls. Horizontal lines indicate thresholds for adjusted p values. Genes are highlighted when their adjusted p value falls below the specified significance cutoff (Benjamini–Hochberg correction for multiple comparisons). Differential expression analysis was performed using the NanoString nSolver Advanced Analysis module (two-sided test). f NanoString pathway scores were presented using a heatmap to illustrate how pathways involved in glial cell homeostasis and activation, inflammation and peripheral immune activation, glial cell regulation, cell stress and damage response, and neurotransmission vary across groups. Each square indicates the score of a specific pathway in one sample; high scores are indicated in orange, and low scores in blue.
Extended Data Fig. 6 iDISCO tissue clearing and wholemount imaging of spinal cord segments.
a A 3D view of spinal cord segments of 12-month-old Tgfbr2fl/fl (WT) and Cx3cr1CreER:Tgfbr2fl/fl (KO) mice (D25). Upper panel: whole 3D images and randomly selected slices showing Iba1 (in red, with expression strength indicated using a heatmap pseudo-colouring; left panel), MBP (green; middle panel) and merged (right panel) staining in the spinal cord. Scale bar=400μm. Bottom panel: corresponding magnified images in the DC, GM, and VC. Scale bar=150μm. b 3D reconstructed and segmented comparison in the DC of WT and KO mice. White scale bar=400μm; yellow scale bar=150μm. Representative images are from two independent experiments with similar results.
Extended Data Fig. 7 Transcriptomic analysis of microdissected microglia from different spinal cord regions following Tgfbr2 deletion.
a Different spinal cord regions of Cx3cr1CreER:Tgfbr2fl/fl mice at 28 days post-tamoxifen administration were microdissected and microglia from the DC, VC and GM were sorted for RNA-seq using FACS. b A total of 91 differentially expressed genes (DEGs) were revealed when comparing microglia from the DC with microglia from the VC and GM (Log2 fold change≥1, q-value < 0.05; DESeq2). The Gene Ontology Biological Process (GO_BP) enrichment analyses show the enriched terms of the 30 upregulated genes in the DC. c The 30 upregulated genes in the DC were analyzed using Cytohubba with the MCC algorithm, and the hub genes along with their interactions were presented. The colored bubbles represent the top 5 hub genes, in descending order from red, through orange, to yellow. d Heatmap displaying the relative expression of genes revealed in (c) across the GM, VC and DC regions. Schematic in a created with BioRender.com.
Extended Data Fig. 8 Annotation of cell clusters as revealed in the snRNA-seq analysis.
a Uniform manifold approximation and projection (UMAP) plot of cells from 12-month-old female Tgfbr2fl/fl (WT) mice and Cx3cr1CreER:Tgfbr2fl/fl (KO) mice at 10 (D10), 20 (D20) or 30 (D30) days post-tamoxifen administration (downsampled to 7, 500 cells per group/time point for presentation). Oligo: mature oligodendrocytes, OPC: oligodendrocyte progenitor cells, pre_Oligo: premature oligodendrocytes, MG: microglia, Astro: astrocytes, EPC: ependymal cells, Endo: endothelial cells, PC: pericytes, MC: meningeal cells, CSF-cN: cerebrospinal fluid-contacting neurons, MN: motor neurons, Neuro-Excit: excitatory neurons, Neuro-Inhib: inhibitory neurons. b Kinetic fold change (compared to WT condition) of cell clusters in (a) over time. c Abundance of cell clusters (in percentage) annotated in (a) between WT and D30 conditions. d A detailed bubble plot illustrating the signature genes that are enriched in each distinct cell cluster.
Extended Data Fig. 9 Subclustering of oligodendrocyte lineage cells from snRNA-seq.
a Oligodendrocyte lineage cells (Oligo, Pre_Oligo, and OPC populations from Extended Data Fig. 8) were subclustered into 15 Seurat subclusters as presented in the tSNE plot, with the number on top of each subcluster representing the Seurat ID. The Seurat subclusters with similar signature gene expression profile in (c) were combined into annotated cell populations. OPC, oligodendrocyte progenitor cells, COP: committed oligodendrocyte progenitor, NFOL: newly formed oligodendrocytes, MFOL: myelin-forming oligodendrocytes, MOL: mature oligodendrocytes, DA-MOL: disease-associated MOL. b t-SNE plot of cells from (a), with colors denoting different conditions/time points. c A heatmap showing the expression of population-specific signature genes across different Seurat subclusters. The color scale bar represents the z-score normalized expression of the gene in each Seurat subcluster (average value from 11 pseudobulk samples).
Extended Data Fig. 10 Expression of Tgfb1 following microglial Tgfbr2 deletion.
a-d Kinetic change of Tgfbr1, Tgfbr2, Tgfb1 and Nrros mRNA expression in spinal cord homogenates following microglial Tgfbr2 deletion (Cx3cr1CreER:Tgfbr2fl/fl mice) was analyzed using NanoString nCounter. Data are presented as mean ± SEM; n = 5, 3, 4, and 3 mice for the WT, KO_D10, KO_D20, KO_D30 groups, respectively. *p < 0.05, ***p < 0.001; one-way ANOVA with Dunnett’s post-hoc test. e A heatmap showing the mRNA expression of Tgfb1 in microglia sorted from DC, VC, and GM regions of WT or Cx3cr1creER:Tgfbr2fl/fl mice at 18 or 28 days post-tamoxifen administration. f A UMAP plot showing the expression of Tgfb1 across different microglial subclusters revealed by snRNA-seq.
Supplementary information
Source data
Source data Figs. 1–7 and Extended Data Figs1, 2 and 10 (download XLSX )
Statistical source data (a single file containing all source data).
Source Data (download PDF )
Unprocessed western blots and/or gels.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, K., Liu, Y., Min, JH. et al. TGFβ signaling mediates microglial resilience to spatiotemporally restricted myelin degeneration. Nat Neurosci 29, 617–631 (2026). https://doi.org/10.1038/s41593-025-02161-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-025-02161-4