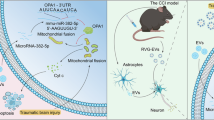
Abstract
Traumatic brain injury (TBI), a leading cause of death and disability globally, often results in functional impairments due to complex pathological mechanisms and limited therapeutic options. Recently, astrocytes, the most abundant glia in the central nervous system, have been recognized as therapeutic targets for TBI due to their roles in neuroinflammation and neuroprotection. While NeuroD1-based gene therapy demonstrates therapeutic potential by reversing glial scar, suppressing neuroinflammation, and repairing brain tissue, the underlying mechanisms remain incompletely understood. To elucidate the molecular and cellular mechanisms involved, we conducted single-cell RNA sequencing using cortical stab injury mice of TBI to compare the effects of vehicle treatment with those of NeuroD1-based gene therapy. Single-cell RNA sequencing reveals significant shifts in cellular composition and astrocyte subtypes. These changes involve upregulated synaptic assembly and myelination pathways alongside downregulated mitochondrial and metabolic functions in TBI mice. Conversely, NeuroD1-based gene therapy restores mitochondrial and metabolic functions and attenuates aberrant synaptic and myelination processes. This single-cell transcriptomic dataset provided a valuable resource for mechanistic and therapeutic research for TBI.
Similar content being viewed by others
Background & Summary
Traumatic brain injury (TBI) is defined as an alteration in brain structure or function resulting from traumatic external mechanical forces, such as impacts, penetrating injuries, blasts, or rapid acceleration/deceleration movements. TBI is manifested by a complex array of pathological changes such as cerebral edema, glial scar, neurodegeneration, neuroinflammation, and perturbations in cerebral perfusion1,2,3,4. Among all neurological disorders and injuries, TBI exhibits the highest incidence and poses a substantial global public health concern and associated healthcare burden5.
The clinical management of TBI involves various interventions, including surgery, pharmacotherapy, regenerative therapies, neuromodulation, and rehabilitation. Surgical approaches focus on controlling intracranial pressure and may involve decompressive craniectomy6. In the acute phase, pharmacological strategies utilize hyperosmolar agents like NaCl to alleviate elevated intracranial pressure7,8. Regenerative therapies are an emerging field, with studies showing the effectiveness of neural stem cell (NSC) administration in rats, grafting human embryonic stem cell-derived cerebral organoids in mice, and using autologous bone marrow mononuclear cells in pediatric patients9,10,11. Neuromodulation strategies, such as brain, spinal cord, and vagus nerve stimulation, are under investigation12,13,14. Rehabilitation follows a multidisciplinary approach aimed at enhancing recovery, reducing disabilities, and facilitating social reintegration15,16. Despite these varied strategies, current treatments face significant limitations, including surgical risks, side effects from medications, and challenges in achieving lasting efficacy with regenerative approaches, neuromodulation, and rehabilitation.
The management of TBI constitutes a complex process necessitating a multidisciplinary and integrated approach to achieve optimal outcomes. In recent years, therapeutic strategies for TBI targeting neurorestorative processes, neuroinflammation mitigation, and glial scar attenuation have gained significant attention and shown promising results17,18,19,20,21. Among these interventions, NeuroD1-based gene therapy has emerged as a potential TBI treatment strategy, simultaneously targeting neuroregeneration, reducing neuroinflammation, and promoting glial scar elimination22,23,24,25,26,27.
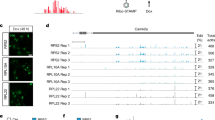
To advance the clinical translation of NeuroD1-based gene therapy for TBI, we aimed to comprehensively elucidate its molecular and cellular mechanisms of action. We employed a mouse model of cortical stab injury to investigate the therapeutic mechanisms underlying NeuroD1-based gene therapy in TBI (Fig. 1a). At 3 days post-injury, AAV vectors encoding either green fluorescent protein (GFP, negative control) or NeuroD1 were stereotaxically injected into the lesion core. Tissue samples were then harvested for histological and single-cell RNA sequencing (scRNA-seq) analyses at 7 and 14 days post-injection (dpi). Histological assessment revealed that NeuroD1 overexpression significantly mitigated TBI-induced tissue damage. Immunofluorescence analyses demonstrated a pronounced reduction in cortical cavity size in NeuroD1-transduced animals compared to GFP controls (Fig. 1b-d, Sup Fig. 1a-b, 2a–c). Concurrent analyses revealed a substantial attenuation of neuro-inflammation in the NeuroD1-treated group, as evidenced by a significant decrease in activated microglia (quantified by Iba1+ cell density, Fig. 1e–g, Sup Fig. 1c, d, note that the reduction at D7 was slightly less than that at D14).
NeuroD1 promotes brain tissue repair and attenuates microglial activation following TBI. (a) Experimental scheme. Mice received a TBI, (red), 3 days later, AAV9 GFAP::GFP (green) or AAV9 GFAP:: NeuroD1-GFP (blue) were injected in the lesion area. This figure was created using BioRender and is reproduced under license (https://biorender.com). (b) Representative coronal sections immediately after TBI model (dashed lines indicate injury boundaries). Nuclei, DAPI (blue). (c) Representative coronal sections (dashed lines delineate injury boundaries). Left: AAV9 GFAP::GFP; right: AAV9 GFAP:: NeuroD1-GFP. Nuclei, DAPI (blue); NeuroD1 (red, scale bar, 500 µm). Insets (scale bar, 50 µm) show NeuroD1 expression in the lesion area. (d) Quantification of cortical lesion volume from brain slices (n = 5 mice per group; one-way ANOVA with Tukey’s post-hoc test; mean ± s.d.; **P < 0.001). (e) Microglial activation in the lesion area. Iba-1 (purple, scale bar, 500 µm). (f) High-magnification images (scale bar, 50 µm) showing microglial morphology. (g) Quantification of the intensity of Iba-1+ cells (f; n = 5-6 mice per group; one-way ANOVA with Tukey’s post-hoc test; mean ± s.d.; ****P < 0.0001).
This study presents a scRNA-seq dataset (10x Genomics) comprising the cortex of healthy male FVB mice, TBI mice injected with control AAV, and TBI mice injected with NeuroD1-expressing AAV. This resource elucidates the cell-type-specific transcriptional landscapes that drive brain damage following TBI, as well as the brain repair processes induced by NeuroD1-based gene therapy, with particular attention to the dynamics of astrocyte subclusters and the implications of differentially expressed genes (DEGs). Our findings not only offer pathological insights for mechanistic studies on TBI, but also provide a valuable resource for identifying potential therapeutic targets and interventions.
Methods
Animals
All mice used in this study are adult FVB/NJ male mice aged 3-4 months. All mice were purchased from Guangdong Zhiyuan Biopharmaceutical Technology Co., Ltd. All mice used in this study received a standard 12 h light-dark cycle and were provided with sufficient water and food. The number of animals used and the suffering of animals were minimized during the experiment. The experimental protocol was approved by the Experimental Animal Ethics Committee of Jinan University, China (approval number: IACAC-20230515-02).
Adeno-associated virus (AAV) injection
To achieve precise adeno-associated virus (AAV) injection into specific regions of the mouse brain, we used stereotactic brain injection. Mice were anesthetized by intraperitoneal injection of 1.25% Avertin (Sigma, T48402) at a dose of 20 ml/kg and placed in a stereotactic apparatus. AAV injection was performed using an injection pump (RWD Life Science, China) and glass microelectrodes. We injected the NeuroD1-expressing AAV into the cortical lesion of a randomly assigned cortical hemisphere (a random integer generator was used for allocation: odd numbers designated the left side, and even numbers designated the right side), and delivered the GFP-expressing AAV to the contralateral hemisphere to serve as a control. All AAVs (5E11 GC/ml) were produced by Guangzhou Pack Gene Biotech Co., Ltd., China. AAV injections were carried out at an infusion rate of 100 nl/min, with each injection site receiving 1.5 μl of AAVs. After injection, the electrode was kept in place for at least 5 minutes to ensure adequate virus diffusion, and then slowly withdrawn28.
TBI mouse model
Using the cortical stab injury method, construct a stable TBI mouse model. Mice were anesthetized by intraperitoneal injection of 1.25% Avertin (Sigma, T48402) at a dose of 20 mL/kg. Under anesthesia, the mice were placed in a stereotactic device, and the skull and anterior fontanelle were exposed through a midline incision. The mouse head was leveled in the anterior-posterior and left-right directions using a stereotactic instrument (RWD Life Science, China) to ensure the accuracy of positioning. Taking the anterior fontanelle as the origin, the coordinates of the cortical injury were determined according to the brain atlas: AP: −1.0 mm; ML: ± 1.5 mm; DV: −1.8 mm. A small hole with a diameter greater than 1 mm was drilled on the skull above the cortex using a cranial drill (RWD Life Science, China). Subsequently, a sterilized blunt needle (diameter 1 mm, custom-made) was slowly inserted into the cortex, held for 3 minutes, and then slowly withdrawn to simulate needle puncture injury23.
Immunohistochemistry
First, the brain tissue was vibratome-sectioned with a thickness of 35 μm. The brain sections were placed in a 24-well plate and washed three times with PBS buffer (0.01 M, pH 7.4), each time for 10 minutes. Then, the sections were incubated at room temperature for 1 hour in a PBS solution containing 5% donkey serum and 0.3% Triton X-100 to block non-specific binding. Next, the samples were incubated with the primary antibodies at 4 °C overnight: [chicken anti-GFP (Abcam, Ab13970, 1:1000); rat anti-GFAP (Invitrogen, 13-0300, 1:1000); rabbit anti-Iba1 (Wako, #019-19741, 1:1000); rabbit anti-NeuroD1 (Abcam, Ab205300, 1:1000)]. The samples were washed three times in PBS containing 0.1% Triton X-100, then added with the secondary antibody mixture containing DAPI nuclear staining (diluted 1:1000), and incubated in the dark at room temperature for 2 hours. The samples were washed three times in PBS containing 0.1% Triton X-100, and the brain slices were mounted on slides with mounting medium and covered with coverslips. They were stored at 4 °C in the dark for imaging.
Microscopic imaging
Obtain large-field mosaic images at low magnification using an ordinary fluorescence microscope for overall analysis. The objective lens used is 10x, and the fluorescence intensity is determined based on the actual staining condition. Use a laser confocal microscope to obtain high-quality detailed images and statistical graphs. The magnification of the detailed images is selected based on the size of the required image. The acquisition of statistical graphs: first, scan the entire area to be statistically analyzed through the 10x lens, and then use the 40x lens to randomly select the field of view through the “Position” function and take images with a resolution of 1024 × 1024. After obtaining the images, process and analyze them using the ZEN Blue software.
Statistical analysis
The immunohistochemical data were analyzed and processed using ZEN 2.3 blue edition, including image truncation and maximum fluorescence intensity superimposed on the z-axis. GraphPad Prism 8 software was used for data analysis. one-way ANOVA was used with Tukey’s post hoc test, and statistical significance was expressed as *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 showed significant differences.
Single-cell dissociation and library construction
Single-cell RNA sequencing was performed on cerebral motor cortex tissues collected from a cohort of 15 adult male FVB/NJ mice. The cohort included 3 naïve controls and 12 mice subjected to TBI. The mice subjected to TBI were further divided into two subgroups receiving either GFP or NeuroD1. A small cylinder-shaped region of the cerebral motor cortex tissues was collected from within 1 mm diameter of the lesion site from the TBI mice, or from a corresponding region in the control mice. To obtain sufficient number of cells, mouse cortical tissues were pooled into 3 samples: first sample combined tissues from 3 mice in the Normal group; second sample pooled tissues from 6 mice in the TBI + GFP group across 2 time points (D7 and D14, 3 mice per time point); and third sample pooled tissues from 6 mice in the TBI + NeuroD1 group across the same time points (D7 and D14, 3 mice per time point). These 3 pooled samples were then processed for subsequent cell dissociation and sequencing. Sequencing was conducted by Singleron Biotechnologies following tissue dissociation into a single-cell suspension. Animals were anaesthetized with 1.25% tribromoethanol and transcardially perfused with ice-cold artificial cerebrospinal fluid (ACSF). The mouse brains were rapidly dissected and placed in cold ACSF, and were micro-dissected under a stereomicroscope to remove meninges and white matter. Tissue fragments were transferred to chilled tubes and dissociated into a single-cell suspension using the Papain Dissociation System (Worthington) according to the manufacturer’s protocol. The suspension was adjusted to 2–2.5 × 105 cells mL−1 in 0.04% BSA/PBS. Approximately 20,000–30,000 cells were loaded onto a 150,000-well SCOPE-chip™ microfluidic array. Poisson distribution under gravity ensured ≤ 1 cell per microwell, targeting recovery of ~7,000 cells. Millions of uniquely barcoded magnetic beads (cell barcodes and UMIs) were then introduced, again yielding ≤ 1 bead per well. Following on-chip lysis, poly-adenylated mRNA was captured via oligo(dT) hybridization. Beads were pooled, and captured RNA was reverse-transcribed and PCR-amplified (12–14 cycles). Amplified cDNA was tagmented, end-repaired, A-tailed and ligated to Illumina-compatible adapters to generate sequencing libraries (average insert 350 bp).
Primary analysis of raw read data
Primary analysis of raw sequencing data (scRNA-seq) was performed to generate comprehensive gene expression profiles. The raw reads were processed using CeleScope v1.15.0 (Singleron Biotechnologies) with optimized parameters to ensure accurate data interpretation. Initially, barcoded sequences and unique molecular identifiers (UMIs) were demultiplexed from the R1 reads and subjected to correction algorithms to minimize errors. Subsequently, R2 reads underwent stringent quality control measures, including the removal of adapter sequences and poly A tails to eliminate sequence artifacts. The cleaned R2 reads were then aligned to the GRCm38 (mm10) transcriptome using STAR v2.6.1a, a splicing-aware aligner, to ensure precise mapping of reads to their respective genomic loci. Uniquely mapped reads were subsequently assigned to annotated genes using FeatureCounts v2.0.1, which leverages the transcriptional context for robust gene quantification. Reads sharing identical cell barcodes, UMIs, and gene assignments were collated to construct a high-resolution gene expression matrix, facilitating downstream analytical workflows. This systematic approach ensured the generation of reliable and reproducible transcriptomic data for subsequent biological interpretations.
Filtering cell doublets and RNA contamination
Cell doublets were identified based on the expression profiles of canonical cell type markers. Clusters demonstrating enrichment of markers specific to multiple cell types were excluded from subsequent analyses to ensure data accuracy and specificity.
To address RNA contamination and doublets in downstream analyses, we applied DecontX29 for contamination estimation and removal and DoubletFinder30 for doublet identification and exclusion.
Quality control, dimensionality reduction, and clustering (Scanpy)
To ensure data quality and facilitate downstream analysis, Scanpy v1.8.2 was employed under Python 3.7 for quality control, dimensionality reduction, and clustering. Each sample dataset underwent stringent filtering based on the following criteria: exclusion of cells with gene counts below 200 or in the top 2% of gene counts, removal of cells with UMI counts in the top 2%, exclusion of cells with mitochondrial content exceeding 15%, and elimination of genes expressed in fewer than 5 cells. The raw count matrix was normalized by total counts per cell and transformed logarithmically. The top 2,000 variable genes were selected using the “Seurat” flavor. Principal Component Analysis (PCA) was performed on the scaled variable gene matrix, retaining the top 20 principal components for clustering and dimensionality reduction. The Louvain algorithm with a resolution parameter of 1.2 was applied, resulting in 26 distinct cell clusters. These clusters were visualized using Uniform Manifold Approximation and Projection (UMAP) and t-Distributed Stochastic Neighbor Embedding (t-SNE)31.
Batch effect correction using harmony
To address the batch effect between samples, we employed Harmony v1.0, a robust and widely-used computational tool for batch effect correction. This method leverages harmonization techniques to align datasets across experimental conditions while preserving biological variability. For our analysis, the top 20 principal components (PCs) derived from the earlier PCA step were used as input for Harmony. These PCs were selected to capture the majority of the variance in the dataset, ensuring that both biological and technical variations were appropriately accounted for. By applying Harmony, we effectively corrected for batch effects, enabling a more accurate comparison of gene expression profiles across samples. This approach ensured that downstream analyses, such as clustering and differential expression analysis, were based on biologically meaningful differences rather than technical artifacts. The use of Harmony v1.0 provided a statistically rigorous framework for batch effect correction, ensuring the reliability and reproducibility of our findings.
Cell type annotation
The cell type annotation for each cluster was determined based on canonical markers from the reference database SynEcoSysTM (Singleron Biotechnology), which integrates markers from CellMakerDB, PanglaoDB, and recent studies focused on single-cell sequencing data.
Subtyping of major cell types
To achieve a high-resolution characterization of astrocytes, cells from the specific cluster were isolated and subjected to re-clustering following the same analytical pipeline described above, with the clustering resolution parameter set to 0.5. This approach enabled a more detailed and refined analysis of astrocytic heterogeneity32.
Differentially expressed genes analysis (scanpy)
DEGs were identified using the scanpy.tl.rank_genes_groups() function, employing the Wilcoxon rank sum test. Genes were considered DEGs if they were expressed in more than 10% of cells in either of compared group and exhibited an average log2 fold change (log2FC) greater than 0.25. Statistical significance was assessed using the Benjamini-Hochberg correction with a p-value threshold of 0.05. This approach ensured the identification of genes with biologically meaningful expression differences while maintaining statistical rigor.
Pathway enrichment analysis
To elucidate the functional roles of astrocytes and their subclusters, we performed pathway enrichment analysis using the Gene Ontology (GO) database, facilitated by the “clusterProfiler” R package v4.0.033. Pathways with an adjusted p-value (p adj) < 0.05 were deemed significantly enriched. Selectively enriched pathways were visualized via bar plots to highlight key biological processes. Additionally, Gene Set Enrichment Analysis (GSEA) was conducted on 18,160 genes across the sham, GFP, and NeuroD1 groups34. For GSVA-based pathway enrichment, the average gene expression of each cell type served as input. Gene Ontology gene sets, encompassing molecular function (MF), biological process (BP), and cellular component (CC) categories, were utilized as reference sets to identify significantly enriched pathways, providing insights into the molecular mechanisms underlying astrocyte diversity and function35.
Data Records
Raw paired-end sequencing reads were deposited in the NCBI Sequence Read Archive under accession SRP64556336 and the Genome Sequence Archive of the National Genomics Data Center, China37,38, under accession CRA03136339. The exact FASTQ filenames for each experimental group are: Normal: (NCBI: control, SRR36068013, Normal_224943_R1.fastq.gz, Normal_224943_R2.fastq.gz, https://www.ncbi.nlm.nih.gov/sra/?term=SRR36068013; GSA: CRR2213892_r1.fastq.gz, CRR2213892_r2.fastq.gz, https://ngdc.cncb.ac.cn/gsa/browse/CRA031363/CRR2213892). TBI + GFP: (NCBI: TBI-GFP SRR36068012 GFP_145357_R1.fastq.gz, GFP 145357_R2.fastg.gz, https://www.ncbi.nlm.nih.gov/sra/?term=SRR36068012; https://www.ncbi.nlm.nih.gov/Traces/index.html?view=run_browser&acc=SRR36068012&display=data-access GSA: CRR2213893_r1.fastq.gz, CRR2213893_r2.fastq.gz, https://ngdc.cncb.ac.cn/gsa/browse/CRA031363/CRR2213893). TBI + NeuroD1: (NCBI: TBI-ND1 SRR36068011 ND1_145356_R1.fastq.gz, ND1_145356_R2.fastq.gz, https://www.ncbi.nlm.nih.gov/sra/?term=SRR36068011; https://www.ncbi.nlm.nih.gov/Traces/index.html?view=run_browser&acc=SRR36068011&display=data-access GSA: CRR2213894_r1.fastq.gz, CRR2213894_r2.fastq.gz, https://ngdc.cncb.ac.cn/gsa/browse/CRA031363/CRR2213894). All files are publicly accessible under the indicated accession numbers.
Additionally, the gene-expression matrices for all three samples have been deposited in the NCBI Gene Expression Omnibus (GEO) under accession number GSE31617240 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc = GSE316172). Each dataset comprises three compressed text files: barcodes, features (genes), and the sparse count matrix. The exact filenames are as follows. Normal (control): GSM9446291_Normal_barcodes.tsv.gz, GSM9446291_Normal_genes.tsv.gz, GSM9446291_Normal_matrix.mtx.gz. TBI + GFP (TBI-GFP): GSM9446289_GFP_barcodes.tsv.gz, GSM9446289_GFP_genes.tsv.gz, GSM9446289_GFP_matrix.mtx.gz. TBI + NeuroD1 (TBI-ND1): GSM9446290_ND1_barcodes.tsv.gz, GSM9446290_ND1_genes.tsv.gz, GSM9446290_ND1_matrix.mtx.gz. All matrix files are publicly accessible through the GEO accession page.
Data Overview
Cell type classification in normal, TBI and NeuroD1 treated mice
Following stringent cell and gene quality filtering based on the criteria outlined in the “quality control, dimensionality reduction, and clustering (Scanpy)” section, a total of 36,119 cells from the Normal group, 37,997 cells from the TBI + GFP group, and 23,169 cells from the TBI + NeuroD1 group were included in the subsequent analysis. Based on the expressing patterns of the top 10 highly expressed marker genes, 12 distinct cell clusters were identified (Fig. 2a). The tSNE dimensionality reduction plot highlights the expression patterns of NeuroD1/GFP in individual cells (Fig. 2b). The top 3 marker genes for each cluster, for instance, astrocytic AQP4, Aldoc, and Gfap, were minimally detected in other cell types, further validating the accuracy of the clustering (Fig. 2c). Dimensionality reduction was applied to visualize the distribution of these clusters (Fig. 2d–f). Comparative analysis among the Normal, TBI + GFP, and TBI + NeuroD1 groups revealed a significant reduction in the percentages of astrocytes and microglia in the TBI + NeuroD1 group, alongside an increase in oligodendrocytes, neurons, and choroid plexus epithelial cells in the TBI + NeuroD1 group (Fig. 2g).
Cell type classification of cells from mouse cortex. (a) Schematic overview of the experimental design. This figure was created using BioRender (https://biorender.com) and is reproduced under license. (b) tSNE visualization of NeuroD1/GFP expression across different cell types. The color bar indicates the relative levels of NeuroD1/GFP expression, with brighter colors representing higher expression. (c) The top 3 cell type-specific DEGs. (d–f) The cell type distribution in the Normal group (d), the TBI + GFP group (e) and the TBI + NeuroD1 group (f). (g) Bar plot showing the relative proportions of each cell types across three independent samples. Clusters are color-coded as shown in the color legend.
Astrocyte clustering in normal, TBI and NeuroD1 treated mice
Astrocytes were classified into 7 distinct subclusters based on their top 3 DEGs (Fig. 3a). The tSNE dimensionality reduction plot highlights the expression patterns of NeuroD1/GFP in individual cells across various subclusters of astrocytes (Fig. 3b). Dimensionality reduction was applied to visualize the distribution of these subclusters (Fig. 3c–e). A comparative analysis across the 3 experimental groups revealed significant shifts in astrocyte composition: astrocyte subclusters 1 and 5 were the most abundant in the Normal group, astrocyte subcluster 7 was selectively expanded in the TBI + NeuroD1 group. Specifically, subcluster 7 constituted only 0.02% (1 cell) in the Normal group, the proportion increased to 1.59% (79 cells) in the TBI + GFP group, and further expanded to 24.84% (271 cells) in the TBI + NeuroD1 group. Notably, astrocyte subclusters 3 and 6 were enriched specifically in the TBI + GFP group, a pattern not shared in either the Normal or TBI + NeuroD1 groups. In contrast, astrocyte subcluster 4 was less prevalent in the Normal group compared to both the TBI + GFP and TBI + NeuroD1 groups. Furthermore, astrocyte subcluster 2 was uniquely detected in injury contexts (TBI + GFP and TBI + NeuroD1 groups, Fig. 3f).
Astrocyte subcluster landscape. (a) Violin plot showing the expression levels of the top 3 marker genes used to distinguish astrocytic subtypes. (b) t-SNE visualization of NeuroD1/GFP expression across different subclusters of astrocytes. The color bar indicates the relative levels of NeuroD1/GFP expression, with brighter colors representing higher expression. (c–e) t-SNE dimensionality reduction plots of cortical astrocytes from the Normal group (c), TBI + GFP group (d) and TBI + NeuroD1 group (e) colored by subclusters. (f) Bar chart displaying the relative proportions of astrocytic subclusters across three groups of samples. Clusters are color-coded as shown in the color legend.
Astrocytic differentially expressed genes and their associated signaling pathways following TBI and NeuroD1-based gene therapy
Differential gene expression analysis was performed between the Normal and TBI + GFP groups. Compared to the Normal group, the TBI + GFP group exhibited upregulation of many genes, including mt-Rnr1, mt-Rnr2, GFP, Gm28438, Fam213a, Hist1h2bc, Cmss1, Atp5f1, Atp5o and Gfap, and downregulation of many genes, including Atp5pb, Prxl2a, H2bc4, Ftl1, Myorg, Rabgef1, Pgm1, Atp5mpl, Micos10, and Hsd17b4 (Fig. 4a). GO enrichment analysis revealed that the top 5 up-regulated pathways in the TBI + GFP group relative to the Normal group were primarily associated with synapse assembly and organization (Fig. 4b). Conversely, the top 5 downregulated pathways were enriched for processes related to mitochondrial function, metabolite and energy generation, and cellular respiration (Fig. 4c).
Differentially expressed genes in astrocytes and GO pathway enrichment. (a) Heatmap of top 20 DEGs in astrocytes between the Normal and TBI + GFP groups. (b) GO enrichment of up-regulated astrocytic DEGs in TBI + GFP vs Normal group. (c) GO enrichment of down-regulated astrocytic DEGs in TBI + GFP vs Normal group. (d) Heatmap of top 20 DEGs in astrocytes between the TBI + GFP and TBI + NeuroD1 groups. (e) GO enrichment of up-regulated astrocytic DEGs in TBI + NeuroD1 vs TBI + GFP group. (f) GO enrichment of down-regulated astrocytic DEGs in TBI + NeuroD1 vs TBI + GFP group. (top 5; Color: -log₁₀(P-value); Size: Gene proportion).
Comparison between the TBI + NeuroD1 and TBI + GFP groups revealed significant downregulation of genes such as GFP, Cmss1, Trpm3, Lsamp, Pcdh9, mt-Rnr1, Gpc5, Mdga2, Magi2, Frmd4a, and significant upregulation of genes such as Cacng5, Fos, Fabp7, Vim, Junb, Sparc, Btg2, Jund, Olfm1, and Egr1 in the NeuroD1-treated group (Fig. 4d). GO term enrichment analysis indicated that these downregulated DEGs in the TBI + NeuroD1 group were enriched for pathways involved in synapse assembly and organization (Fig. 4e). GO term enrichment analysis also suggested that these upregulated DEGs in the TBI + NeuroD1 group were enriched for pathways involved in mitochondrial function, metabolite and energy generation, and cellular respiration (Fig. 4f).
In our study, both TBI modeling and NeuroD1 overexpression markedly influenced the composition of astrocyte subtypes. To investigate the transcriptomic alterations underlying these changes, we identified DEGs among astrocyte subclusters and conducted pathway enrichment analyses.
In astrocyte subcluster 3, many genes, including Myorg, Prxl2a, Atp5pb, Ftl1, Mtss2, H2bc4, Atp5mpl, Pgm1, Rmst, Hsd17b4 were substantial downregulated in the TBI + GFP group compared to the Normal group (Fig. 5a). Notably, these downregulated genes were associated with biological processes such as mitochondrial function, metabolite and energy generation, and cellular respiration (Fig. 5b). Moreover, when comparing the TBI + NeuroD1 group to the TBI + GFP group within astrocyte subcluster 3, many genes, such as Ptgds, Thrsp, Ly6e, lgfbp2, Sumf2, Tubb5, Atp5o, Bad, Robo2, and Tmsb4x, were upregulated following NeuroD1 treatment (Fig. 5c). Importantly, these DEGs clustered within pathways related to mitochondrial function, metabolite and energy generation, and cellular respiration (Fig. 5d).
Differentially expressed genes and GO pathway enrichment in astrocyte subclusters 3 and 6. (a) Heatmap of top 20 DEGs in astrocytes subcluster 3 between the Normal and TBI + GFP groups. (b) GO enrichment of down-regulated subcluster 3 DEGs in TBI + GFP vs Normal group. (c) Heatmap of top 20 DEGs in astrocytes subcluster 3 between the TBI + GFP and TBI + NeuroD1 groups. (d) GO enrichment of up-regulated subcluster 3 DEGs in TBI + NeuroD1 vs TBI + GFP group. (e) Heatmap of top 20 DEGs in astrocytes subcluster 6 between the Normal and TBI + GFP groups. (f) GO enrichment of down-regulated subcluster 6 DEGs in TBI + GFP vs Normal group. (g) Heatmap of top 20 DEGs in astrocytes subcluster 6 between the TBI + GFP and TBI + NeuroD1 groups. (h) GO enrichment of up-regulated subcluster 6 DEGs in TBI + NeuroD1 vs TBI + GFP group. Color: -log₁₀ adjusted P-value; Size: Gene proportion.
In astrocyte subcluster 6, many genes including Prx12a, Atp5pb, H2bc4, Myorg, Ftl1, Pgm1, Rabgef1, Atp5mpl, Hsd17b4, and Micos10 were downregulated in the TBI + GFP group compared to the Normal group (Fig. 5e). Pathway enrichment analysis revealed that the downregulated genes were associated with mitochondrial function, metabolite and energy generation, and cellular respiration (Fig. 5f). Furthermore, when comparing the TBI + NeuroD1 group to the TBI + GFP group within astrocyte subcluster 6, many genes such as Cadm3, Ptgds, Aebp1, Ostc, Prr5, Hes5, Fndc5, Ly6e, Qdpr, and Gfap were upregulated after NeuroD1 treatment (Fig. 5g). GO term enrichment analysis indicated that the upregulated genes were significantly associated with the mitochondrial function, metabolite and energy generation, and endoplasmic reticulum stress (Fig. 5h).
In astrocyte subcluster 4, compared to the Normal group, the TBI + GFP group showed downregulation of genes including Prxl2a, H2bc4, Atp5pb, Ftl1, Prss56, Crym, Rabgef1, Atp5mpl, Micos10, and Hsd17b4 (Fig. 6a). These DEGs were enriched in pathways related to mitochondrial function, metabolite and energy generation, and cellular respiration (Fig. 6b). In contrast, the TBI + NeuroD1 group exhibited upregulation of genes such as Egr1, Junb, Jun, Ptgds, Fos, Ler2, Tmsb4x, Btg2, Jund, and Slc6a6 in astrocyte subcluster 4 compared to the TBI + GFP group (Fig. 6c). These genes were associated with cytoplasmic translation, ribosomal functions, and protein synthesis and folding (Fig. 6d).
Differentially expressed genes and GO pathway enrichment in other astrocyte subclusters. (a) Heatmap of top 20 DEGs in astrocytes subcluster 4 between the Normal and TBI + GFP groups. (b) GO enrichment of down-regulated subcluster 4 DEGs in TBI + GFP vs Normal group. (c) Heatmap of top 20 DEGs in astrocytes subcluster 4 between the TBI + GFP and TBI + NeuroD1 groups. (d) GO enrichment of up-regulated subcluster 4 DEGs in TBI + NeuroD1 vs TBI + GFP group. (e) Heatmap of top 20 DEGs in astrocytes between the Normal (subclusters 1 and 5) and TBI + GFP (subclusters 2 and 7) groups. (f) GO enrichment of up-regulated DEGs in TBI + GFP (subclusters 2 and 7) vs Normal (subclusters 1 and 5) group. (g) Heatmap of top 20 DEGs in astrocytes subclusters 2 and 7 between the TBI + GFP and TBI + NeuroD1 group. (h) GO enrichment of up-regulated subcluster 2 and 7 DEGs in TBI + NeuroD1 vs TBI + GFP group. Color: -log₁₀ adjusted P-value; Size: Gene proportion.
Consistent with the astrocyte subcluster distribution (Fig. 3f), Ast-3, Ast-4, and Ast-6 were present across all experimental groups, differing only in their relative abundance. In contrast, Ast-1 and Ast-5 were detected exclusively in the normal group and were absent following injury. Conversely, Ast-2 and Ast-7 were either newly induced or significantly upregulated after injury. We compared with astrocyte subclusters 1 and 5 in the Normal group, subclusters 2 and 7 from the TBI + GFP group showed upregulated expression of genes including mt-Rnr2, mt-Rnr1, GFP, Hist1h2bc, Fam213a, Gm28438, Atp5f1, Atp5o, Cmss1, and Gm5514 (Fig. 6e). These genes were enriched in biological processes associated with protein degradation, neuronal myelination and cell migration and motility (Fig. 6f). Further analysis revealed downregulation of genes such as Gal, Ptgds, Ly6e, Sct, Tmsb4x, Olig1, Cpne9, Cdkn1a, Cadm3, and Vim in astrocyte subclusters 2 and 7 derived from the TBI + NeuroD1 group relative to those from the TBI + GFP group (Fig. 6g). These DEGs were enriched in pathways related to acylglycerol and neutral lipid metabolism (Fig. 6h).
Technical Validation
In this dataset, the Normal group captured 38,246 cortical cells, the TBI + GFP group captured 40,310 cortical cells, and the TBI + NeuroD1 group captured 24,842 cortical cells (Table 1). The high proportions of valid reads, Q30 bases in both barcodes and UMIs, and uniquely genome-mapped reads collectively indicate that the scRNA-seq library exhibits good quality and efficient mapping performance (Table 2). Cells exhibiting mitochondrial gene percentages exceeding 10% accounted for 0.38% in the Normal group, 8.34% in the TBI + GFP group, and 4.05% in the TBI + NeuroD1 group (Table 3), suggesting adequate cellular viability following cell harvesting. The three sample groups (Normal, TBI + GFP, and TBI + NeuroD1) captured ~25,000 to 40,000 cells, with an average sequencing depth ranging from 19,000 to 34,000 reads per cell. All three groups exhibited highly consistent median genes per cell (Normal: 1519; TBI + GFP: 1363; TBI + NeuroD1: 1671) with less than 15% variation. These values fell within the recommended range of 1,000–2,000 genes per cell for the 10x Genomics platform (Table 1). The high median gene counts, coupled with the overall data quality, provides a reliable foundation for subsequent cell subtype identification and differential gene expression analysis.
Usage Notes
This dataset offers a transcriptomic resource characterizing cortical cells in the healthy FVB mouse brain and delineating their dynamic responses following TBI and subsequent NeuroD1-based brain repair. By integrating transcriptomic profiling with cellular composition analysis, this resource serves three primary purposes:
-
1.
Healthy control benchmark: Publication of a detailed transcriptome dataset derived from the cerebral cortex of adult male FVB mice, serving as a critical reference for studies utilizing this strain in neurological disease research.
-
2.
TBI pathological insights: Quantification of post-TBI alterations in cortical cell populations (glia, neurons, immune cells, endothelial cells) and identification of differentially expressed genes (DEGs), offering mechanistic insights into TBI pathology and informing treatment strategies.
-
3.
Brain repair mechanisms: The dataset captures the molecular underpinnings of the brain repair process, revealing alterations in cell subtype proportions, gene expression, and metabolic pathways following gene therapy for TBI via astrocytic NeuroD1 overexpression. These findings not only advance our understanding of the therapeutic mechanisms of NeuroD1-based gene therapy, but also provide a foundational resource for exploring broader interventions for TBI-induced damage.
Importantly, a couple of key points should be considered when using this dataset. Both the TBI + GFP group and TBI + NeuroD1 group represent pooled tissue samples collected on day 7 and day 14 post-viral expression. Note there is a slight difference in the number of Iba1+ cells at these two times. Special attention is required during pseudotime analysis due to the nature of these pooled samples. The TBI model employed was a stab injury model, which is particularly suited for studying penetrating brain injuries.
Data availability
Raw sequencing reads reported in this paper have been deposited in the two public repositories: (1) the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) under accession SRP645563 (https://identifiers.org/ncbi/insdc.sra:SRP645563); (2) the Genome Sequence Archive (GSA) of the National Genomics Data Center, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession CRA031363 (https://ngdc.cncb.ac.cn/gsa/browse/CRA031363). Processed single-cell gene-expression matrices have been deposited in the NCBI Gene Expression Omnibus (GEO) under accession GSE316172 (https://identifiers.org/geo/GSE316172). All uploaded data are publicly accessible through the aforementioned portals.
Code availability
No custom code was used in this study.
References
Stocchetti, N. et al. Severe traumatic brain injury: targeted management in the intensive care unit. Lancet Neurol 16, 452–464, https://doi.org/10.1016/s1474-4422(17)30118-7 (2017).
Simon, D. W. et al. The far-reaching scope of neuroinflammation after traumatic brain injury. Nat Rev Neurol 13, 171–191, https://doi.org/10.1038/nrneurol.2017.13 (2017).
Muñoz-Ballester, C. & Robel, S. Astrocyte-mediated mechanisms contribute to traumatic brain injury pathology. WIREs Mech Dis 15, e1622, https://doi.org/10.1002/wsbm.1622 (2023).
Jha, R. M. et al. A single-cell atlas deconstructs heterogeneity across multiple models in murine traumatic brain injury and identifies novel cell-specific targets. Neuron 112, 3069–3088.e3064, https://doi.org/10.1016/j.neuron.2024.06.021 (2024).
Maas A. I. R. et al. Traumatic brain injury: progress and challenges in prevention, clinical care, and research. The Lancet Neurology 21, 1004–1060, https://doi.org/10.1016/s1474-4422(22)00309-x (2022).
Schizodimos, T., Soulountsi, V., Iasonidou, C. & Kapravelos, N. An overview of management of intracranial hypertension in the intensive care unit. Journal of Anesthesia 34, 741–757, https://doi.org/10.1007/s00540-020-02795-7 (2020).
Gharizadeh, N. et al. Hypertonic saline for traumatic brain injury: a systematic review and meta-analysis. Eur J Med Res 27, 254, https://doi.org/10.1186/s40001-022-00897-4 (2022).
Kochanek, P. M. et al. Comparison of Intracranial Pressure Measurements Before and After Hypertonic Saline or Mannitol Treatment in Children With Severe Traumatic Brain Injury. JAMA Netw Open 5, e220891, https://doi.org/10.1001/jamanetworkopen.2022.0891 (2022).
Wang, W. et al. Ultrasound-activated piezoelectric nanostickers for neural stem cell therapy of traumatic brain injury. Nat Mater 24, 1137–1150, https://doi.org/10.1038/s41563-025-02214-w (2025).
Bao, Z. et al. Human Cerebral Organoid Implantation Alleviated the Neurological Deficits of Traumatic Brain Injury in Mice. Oxid Med Cell Longev 2021, 6338722, https://doi.org/10.1155/2021/6338722 (2021).
Cox, C. S. Jr. et al. Autologous bone marrow mononuclear cells to treat severe traumatic brain injury in children. Brain 147, 1914–1925, https://doi.org/10.1093/brain/awae005 (2024).
Zhuang, Y. et al. Effects of short-term spinal cord stimulation on patients with prolonged disorder of consciousness: A pilot study. Front Neurol 13, 1026221, https://doi.org/10.3389/fneur.2022.1026221 (2022).
Rezaei Haddad, A., Lythe, V. & Green, A. L. Deep Brain Stimulation for Recovery of Consciousness in Minimally Conscious Patients After Traumatic Brain Injury: A Systematic Review. Neuromodulation 22, 373–379, https://doi.org/10.1111/ner.12944 (2019).
Beekwilder, J. P. & Beems, T. Overview of the clinical applications of vagus nerve stimulation. J Clin Neurophysiol 27, 130–138, https://doi.org/10.1097/WNP.0b013e3181d64d8a (2010).
Hassett, L. Physiotherapy management of moderate-to-severe traumatic brain injury. J Physiother 69, 141–147, https://doi.org/10.1016/j.jphys.2023.05.015 (2023).
Yen, H. C. et al. Assessing the impact of early progressive mobilization on moderate-to-severe traumatic brain injury: a randomized controlled trial. Crit Care 28, 172, https://doi.org/10.1186/s13054-024-04940-0 (2024).
Schiweck, J. et al. Drebrin controls scar formation and astrocyte reactivity upon traumatic brain injury by regulating membrane trafficking. Nat Commun 12, 1490, https://doi.org/10.1038/s41467-021-21662-x (2021).
Shi, G. et al. Inhibition of neutrophil extracellular trap formation ameliorates neuroinflammation and neuronal apoptosis via STING-dependent IRE1α/ASK1/JNK signaling pathway in mice with traumatic brain injury. J Neuroinflammation 20, 222, https://doi.org/10.1186/s12974-023-02903-w (2023).
Izzy, S. et al. Nasal anti-CD3 monoclonal antibody ameliorates traumatic brain injury, enhances microglial phagocytosis and reduces neuroinflammation via IL-10-dependent T(reg)-microglia crosstalk. Nat Neurosci 28, 499–516, https://doi.org/10.1038/s41593-025-01877-7 (2025).
Zhang, H. et al. Astrocyte-mediated inflammatory responses in traumatic brain injury: mechanisms and potential interventions. Front Immunol 16, 1584577, https://doi.org/10.3389/fimmu.2025.1584577 (2025).
Kerr, N. A. et al. Single cell RNA sequencing after moderate traumatic brain injury: effects of therapeutic hypothermia. J Neuroinflammation 22, 110, https://doi.org/10.1186/s12974-025-03430-6 (2025).
Chen, Y.-C. et al. A NeuroD1 AAV-Based Gene Therapy for Functional Brain Repair after Ischemic Injury through In Vivo Astrocyte-to-Neuron Conversion. Molecular Therapy 28, 217–234, https://doi.org/10.1016/j.ymthe.2019.09.003 (2020).
Zhang, L. et al. Development of Neuroregenerative Gene Therapy to Reverse Glial Scar Tissue Back to Neuron-Enriched Tissue. Frontiers in Cellular Neuroscience 14, https://doi.org/10.3389/fncel.2020.594170 (2020).
Kim, M. et al. In vivo neural regeneration via AAV-NeuroD1 gene delivery to astrocytes in neonatal hypoxic-ischemic brain injury. Inflammation and Regeneration 44, 33, https://doi.org/10.1186/s41232-024-00349-y (2024).
Wu, Z. et al. Brain‐Wide Neuroregenerative Gene Therapy Improves Cognition in a Mouse Model of Alzheimer’s Disease. Advanced Science 12, https://doi.org/10.1002/advs.202410080 (2025).
Chen, F. et al. Neurons derived from NeuroD1-expressing astrocytes transition through transit-amplifying intermediates but lack functional maturity. Science advances 11, eadw9296, https://doi.org/10.1126/sciadv.adw9296.
Kim, M. et al. In vivo neural regeneration via AAV-NeuroD1 gene delivery to astrocytes in neonatal hypoxic-ischemic brain injury. Inflamm Regen 44, 33, https://doi.org/10.1186/s41232-024-00349-y (2024).
Guo, Z. et al. In Vivo Direct Reprogramming of Reactive Glial Cells into Functional Neurons after Brain Injury and in an Alzheimer’s Disease Model. Cell Stem Cell 14, 188–202, https://doi.org/10.1016/j.stem.2013.12.001 (2014).
Yang, S. et al. Decontamination of ambient RNA in single-cell RNA-seq with DecontX. Genome Biol 21, 57, https://doi.org/10.1186/s13059-020-1950-6 (2020).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: Doublet Detection in Single-Cell RNA Sequencing Data Using Artificial Nearest Neighbors. Cell Syst 8, 329–337 e324, https://doi.org/10.1016/j.cels.2019.03.003 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biology 19, https://doi.org/10.1186/s13059-017-1382-0 (2018).
Zhang, Y. et al. SynEcoSys: a multifunctional platform of large-scale single-cell omics data analysis. bioRxiv: the preprint server for biology, 2023.2002.2014.528566, https://doi.org/10.1101/2023.02.14.528566 (2023).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287, https://doi.org/10.1089/omi.2011.0118 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550, https://doi.org/10.1073/pnas.0506580102 (2005).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7, https://doi.org/10.1186/1471-2105-14-7 (2013).
NCBI SRA NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP645563 (2025).
Zhang 张思思, S. et al. The GSA Family in 2025: A Broadened Sharing Platform for Multi-omics and Multimodal Data. Genomics Proteomics Bioinformatics 23, https://doi.org/10.1093/gpbjnl/qzaf072 (2025).
Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2025. Nucleic Acids Res 53, D30-d44, https://doi.org/10.1093/nar/gkae978 (2025).
NGDC GSA Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA031363 (2025).
NCBI GEO NCBI Gene Expression Omnibus https://identifiers.org/geo/GSE316172 (2026).
Acknowledgements
This work is supported by the National Natural Science Foundation of China (Grant No. 32571169), and the Program for Guangdong Introducing Innovative and Entrepreneurial Teams (Grant No. 2021ZT09Y552).
Author information
Authors and Affiliations
Contributions
W.L. supervised the entire project. S.Z. and S.L. performed the surgical procedures and stereotactic injections, conducted the immunostaining assays, and completed the wide-field and confocal imaging; SZ and SL prepared the samples for single-cell RNA sequencing; SZ, SL and RC conducted the statistical analysis; WL, RC and SZ wrote the manuscript. All authors approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
Gong Chen is a cofounder of NeuExcell Therapeutics Inc. The other authors have no conflicts of interest to declare.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, R., Zhang, S., Liu, S. et al. A single-cell transcriptomic dataset profiling traumatic brain injury and NeuroD1-based gene therapy in mice. Sci Data 13, 406 (2026). https://doi.org/10.1038/s41597-026-06788-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06788-1