Abstract
The NLRP3 inflammasome plays a pivotal role in the innate immune system, orchestrating the activation of caspase-1 and the release of proinflammatory cytokines IL-1β and IL-18 in reaction to microbial infections and cellular damage. Despite its crucial function in defending against pathogens, the dysregulated activation of the NLRP3 inflammasome has been associated with various inflammatory disorders. In the current investigation, promising plant-derived alkaloids compounds have been discovered as targeted inhibitors against multiprotein NLRP3 using an in-silico drug development approach. The repurposing of natural compounds as anti-inflammatory agents remains a relevant approach for identifying promising early interventions to prevent and manage inflammatory diseases. In this molecular docking study targeting Chain A of the NLRP3 inflammasome protein, eight plant-derived alkaloids renowned for their anti-inflammatory properties were chosen. Docking analysis of the selected alkaloids showed the lowest/best binding energies of less than − 10 Kcal/mol against NLRP3 Chain A, based on this docking result, which is regarded as an exceptional binding score. Notably, Oxyacanthine, Magnoflorine, Corynoline, and Berbamine demonstrated the most favourable binding energies, displaying unique interactions within the binding pocket of the NACHT/PYD domain of NLRP3 Chain A among all compounds investigated. These findings highlight the potential of these alkaloids as promising therapeutic candidates specifically targeting this trans-activating NACHT/PYD domain of NLRP3 Chain A in the context of anti-inflammatory interventions. Protein-protein interactions (PPIs) play an important role in elucidating protein function and drug interactions. To identify bioactive compounds with anti-inflammatory potential, a functional protein network was constructed from publicly available PPI data. As a result, the findings of this in-silico study may cause researchers to emphasize more on alkaloids when considering natural plant products for the treatment of various illnesses that target the inflammatory intermediates. This computational approach predicted ligands that may modulate inflammatory proteins and support host immunity. However, further in vitro and in vivo studies are still needed to validate these in-silico findings before clinical use. In summary, analysing PPI networks can aid discovery of therapeutic candidates, but experimental validation remains essential.
Similar content being viewed by others
Introduction
These disorders encompass cryopyrin-associated periodic syndromes, Alzheimer’s disease, diabetes, and atherosclerosis. The intricate balance of NLRP3 inflammasome regulation is essential for maintaining immune homeostasis, and its dysregulation may contribute to the pathogenesis of these inflammatory conditions. While the precise mechanism underlying NLRP3-mediated disorders remains not fully elucidated, numerous phytochemicals and medicinal plants have been identified for their ability to mitigate inflammatory disorders without inducing adverse effects in other areas.
The innate immune system is the body’s first line of defences against infections and other foreign invaders. It provides immediate, nonspecific protection, and it is present from birth, providing a rapid response to a wide range of pathogens. The innate immune system plays a crucial role in preventing the entry, multiplication and spread of harmful microorganisms. Pattern recognition receptors (PRRs) identify distinct microbial elements known as pathogen-associated molecular patterns (PAMPs) or damage-associated molecular patterns (DAMPs). These molecular patterns, produced by internal stress, are recognized by PRRs, initiating subsequent inflammatory pathways aimed at eradicating microbial infections and facilitating the recovery of impaired tissues. Among different active key components of immune systems Pattern-Recognition Receptors (PRRs) is one of the germline codded pattern receptors that is activated in response to external and internal invading patterns. This initiates the activation of downstream inflammatory response pathways, crucial for repairing damaged tissues and eliminating both the microbe and the associated infection. The activation of the NLRP3 inflammasome dependent caspase-1 is amongst one of the inflammatory pathways comprised of a multiprotein activating complex that plays a crucial role in the innate immune system. It is formed by the assembly of NOD-like receptor family pyrin domain-containing 3 (NLRP3), the adapter protein ASC (apoptosis-associated speck-like protein containing a caspase recruitment domain), and pro-caspase-11,2,3,4. Activation of the NLRP3 inflammasome is triggered by various danger signals, such as microbial products, ATP, and environmental irritants. Upon activation, pro-caspase-1 is cleaved into active caspase-1, which subsequently processes pro-inflammatory cytokines like interleukin-1β (IL-1β) and interleukin-18 (IL-18)5. The NLRP3 inflammasome is of particular interest as it plays a prominent role in various human disorders and is a recognized member of the NLR family, which comprises 22 members in humans and over 34 members in mice6. This inflammatory response is implicated in various diseases, including autoimmune disorders, metabolic diseases, and neurodegenerative conditions. Researchers study the NLRP3 inflammasome to better understand its role in health and disease, with the goal of developing therapeutic interventions.
The integral unit of NLRs typically comprises the Nucleotide-binding oligomerization domain (NACHT) positioned centrally, the leucine-rich repeat (LRR) situated at the C-terminus, and both the CARD and pyrin domain (PYD) located at the N-terminus. Subsequent to the detection of signals by the LRR domain from danger-associated molecular patterns (DAMPs) or pathogen-associated molecular patterns (PAMPs), the PYD domain engages in an anchoring interaction with the PYD domain in ASC (Apoptosis-associated speck-like protein containing a CARD)7.
NLRs typically consist of three segments: (1) carboxy-terminal LRRs, responsible for stimulus recognition (e.g. from DAMPS and PAMPS; (2) an oligomerization- and nucleotide binding-domain facilitating self-oligomerization; and (3) an amino-terminal region featuring CARD, pyrin, acidic transactivating, or baculovirus inhibitor of apoptosis protein (IAP) domains involved in protein-protein interactions8,9. For the NLRP3 sensor, a carboxy-terminal LRR domain, a central NACHT domain, and an amino-terminal PYD domain are present. When combined with an adaptor (ASC or PYCARD) and an effector (caspase 1), they constitute the inflammasome10. However, the NLRP3 inflammasome requires two signals for activation: priming and activation. Initially, cells must undergo priming via toll-like receptor (TLR) activation to induce the expression of NLRP3, caspase-1, and the pro-IL-1β inflammasome component through the type I interferon (IFN) or NF-κB pathway11.
It is noteworthy that the amino-terminal PYD domain serves as a transactivating transducer component in the NLRP3 chain. As mentioned earlier, this region initiates the activation of the ASC (apoptosis-associated speck-like protein containing a caspase recruitment domain) adapter protein by engaging with its PYD domain. Subsequently, the CARD domain of ASC interacts with the neighboring CARD domain of caspase 1, culminating in the autocatalytic activation of dormant procaspase-1 into effector caspase-1. This cascade of events enhances the inflammatory cytolytic activity. The active auto-phosphorylation property of the NOD domain of NLRP3, facilitated by the ATPase, plays a crucial role in the process of activated oligomerization and assembly12. MCC950, a well-established NLRP3 inhibitor, has been identified as specifically targeting this ATPase activity. Research has shown that inhibiting the NLRP3 inflammasome can significantly reduce inflammation, and MCC950 is one of the most well-studied inhibitors in this regard. MCC950 functions by specifically blocking NLRP3 activation, which in turn decreases the production of key pro-inflammatory cytokines such as IL-1β and IL-18. Numerous studies have demonstrated the effectiveness of MCC950 in reducing inflammation in a variety of inflammatory disease models. For example, research conducted by Coll et al. (2015)13 revealed that MCC950 markedly reduced IL-1β levels and eased inflammation in models of systemic inflammation and multiple sclerosis in mice. Additionally, MCC950 has been shown to provide protection in various conditions marked by acute and chronic inflammation, including diseases like gout, Alzheimer’s, and atherosclerosis. These studies highlight the importance of targeting NLRP3 to control inflammation, and the role of inhibitors like MCC950 should be clearly acknowledged to emphasize its therapeutic potential in inflammatory diseases.This discovery sheds light on a pivotal step in the regulatory mechanisms of NLRP3 activation and underscores the significance of ATPase inhibition in the modulation of NLRP3-associated processes14,15. Therefore, in this study the amino-terminal PYD/ NACHT domain of NLRP3 chain A serves as the key target site for the alkaloid compounds binding to the transducing component of this inflammasome protein. Because of its unique response to inflammatory triggers, NLRP3 is actively being pursued as a promising target for drug development aimed at treating autoimmune and inflammatory diseases16.
In the realm of disease associations, NLRP3 emerges as a central player, implicated in various conditions such as neurological disorders, including Parkinson’s disease, Alzheimer’s disease, and multiple sclerosis17. Additionally, its involvement extends to type 2 diabetes18, obesity19, atherosclerosis20,21, and gout22. Particularly noteworthy is the substantial role played by the NLRP3 inflammasome in generating IL-1β, a key factor in the pathogenesis of gout. Considering the limitations and side effects associated with current therapeutic interventions, targeting NLRP3 presents a promising avenue for future therapeutic strategies23. Another compelling example lies in the context of atherosclerosis, where NLRP3 overexpression has been identified in the aorta of individuals with coronary atherosclerosis24,25.
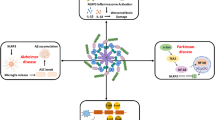
In drug discovery and development, phytochemicals are gaining prominence as alternative therapeutic and pharmacological agents. Their administration is associated with limited or no side effects, and they exhibit a unique mechanism of action26,27. Notably, their considerable chemical diversity enhances their therapeutic interactions with diverse biological targets, setting them apart from synthetic drugs. Representing an essential reservoir of numerous bioactive natural compounds, natural sources have played a crucial role in drug discovery. A significant observation from 1981 to 2014 underscores that only 35% of approved small-molecule drugs were synthetic, underscoring the pivotal contribution of natural molecules as potential drug candidates28. Within this context, natural agents influencing NLRP3 modulation emerge as promising leads against NLRP3-mediated diseases. Hence, in this insilico overview of phytochemicals that inhibit the NLRP3 inflammasome and associated diseases. The exploration of phytochemical inhibitors targeting NLRP3 inflammasome activation holds promise for unveiling novel therapeutic strategies in the treatment of NLRP3-associated disorders. Our current study delves into the exploration of potential anti-inflammatory alkaloids identified in phytochemical studies. All the selected alkaloids in this studies have been previously utilised as a potential inhibitor of inflammation as represented in Table 1. We view these compounds as promising candidates for drug development, specifically targeting the PYD domain of NLRP3 Chain A. This exploration is grounded in the traditional utilization of these alkaloids, presenting them as novel therapeutic strategies for addressing disorders associated with NLRP3. Figure 1. illustrates the representation of the molecular mechanisms by which the targeted alkaloids can modulate the NLRP3 inflammasome, offering insights into potential therapeutic strategies for conditions involving excessive inflammation.
The molecular mechanisms by which alkaloids can modulate the NLRP3 inflammasome, thus shedding light on novel possible therapeutic strategies against diseases related to excessive inflammation.
In our investigation, we utilize eight potential alkaloids that have undergone previous in vitro and in vivo testing for potential clinical efficacy as candidates for anti-inflammatory drugs. These alkaloids, alongside conventional anti-inflammatory drugs, undergo investigation using molecular docking, coupled with ADMET studies, pharmacokinetic evaluation, and analyses of drug likeliness, among other parameters. This comprehensive assessment is conducted to elucidate their actual binding molecular targets within the protein chain of NLRP3 inflammasomes29.
Materials and methods
Retrieval and optimisation of structure
The 3.840 Å resolution complex 3 D structures of Human NLRP3-deltaPYD hexamer (6NPY.pdb) was retrieved from RCSB Protein Data Bank (https://www.rcsb.org/). Using the SPDVB-4.10 version, initial tasks were first completed for protein preparation, including charge assignment, solvation parameters, and fragmental volumes (Morris et al., 2009). Using AutoDock4 Tool for molecular docking, the protein molecules were further improved30.
Secondary structure predictions
Using the Self-Optimized Prediction Method with Alignment (SOPMA) online web server at https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_sopma.html, protein secondary structures were proposed (Geourjon & Deléage, 1995)31. SOPMA predicts four conformational states: helix, beta sheets and bridges, turns, and coils.
Preparation of targeted single chain unit of NLRP3
This complex protein consists of a hexamer unit of delta PYD NLRP3 subunits. Using Biovia Discovery Studio 2021, we first removed the heteroatoms (including water molecules) and the repeated hexamer chains, retaining only the single chain A unit for analysis. The complete molecular unit of the protein comprises NACHT, LRR, and PYD domains, all organized together with chain A. This single protein sequence for chain A includes a total length of 1,034 amino acids. The complete 3D structure of NLRP3-deltaPYD hexamer and retrieved single chain molecule is represented in the Fig. 2. Using SwissProtein Data Base Viewer version 4.10, the crystallographic structures of the NACHT, LRR and PYD proteins were analyzed for errors and broken chains, and then repaired32. After target chain (chain A) preparation the polar hydrogen atoms were added for a more realistic depiction of hydrogen bonding patterns during molecular docking simulations and the chain was saved in AutoDock 4.2 readable PDB(.pdb) format for further modifications.
Selected protein chain validation
The selected NLRP3 chain A was validated using the Swiss-Model ExPASy web server, and these computational assessments confirmed the high quality of the homology model developed with the SWISS MODEL (https://swissmodel.expasy.org/) server. Additionally, the Protein Structure Analysis (ProSA) web tool (https://prosa.services.came.sbg.ac.at/) evaluated the protein model quality by comparing its Z-score to native structures. By analyzing dihedral angles and Z-scores, the in-silico assessments provided critical confirmation of the modelled protein structure prior to further computational analyses.
Phytochemicals ligands retrieval and selection
The PubChem database (https://pubchem.ncbi.nlm.nih.gov) provided the ligand compounds’ chemical composition, structural characteristics, and canonical SMILES. CORINA, an online program, was used to obtain the compound’s PDB structures using canonical SMILES. Tables 1 and Table 2 provides relative information on the phytochemicals employed in the study. Based on their prior investigation’s findings about their anti-inflammatory abilities, each phytoconstituents was approved and considered for docking studies.
Preparation of ligands
Using the Chem Draw and tool in Chem Office-16, AGP structures was used to optimized the structures. Then, using the Chem3D tool, they were transformed into 3D structures and saved as .sdf or. mol files for later use. All the 2D chemical structures are represented in Fig. 3. The structures of all the compounds were depicted using ACD/ChemSketch and ChemBio3D Ultra 12.0 (www.cambridgesoft.com), which was also used to optimise ligand geometry and run MM2 energy minimization on the 3D structures33. The OpenBabel program was utilized to transform the ligands’ three-dimensional (3D) structure from sdf to pdb file format34.
Molecular docking simulation and visualisation
Molecular docking simulations were performed using AutoDock v4.2.6 (for docking with the Lamarckian Genetic Algorithm, LGA) and AutoDock Vina v1.2.0 (for docking scoring and binding affinity predictions) with the standard scripting method to investigate the binding interactions of ligands with protein Chain-A of NLRP3. The ligand and receptor input files were prepared using AutoDockTools, with grid box settings tailored to ensure flexible docking of the ligands.
Protein and ligand preparation involved the following steps
-
Removal of water molecules and N3 heteroatom from the protein.
-
Addition of polar hydrogens and charges to both protein and ligands using Kollman and Gasteiger methods.
Docking parameters included
-
Grid box centered at 118.184, -170.692, -128.025 with spacing of 1.000 and dimensions of 62 × 96 × 66.
-
Lamarckian Genetic Algorithm (LGA) for optimization and stabilization of the protein structure.
Docking procedure
Each ligand was docked thrice. The 10 conformations generated for each docking run were evaluated based on binding energy. The conformation with the lowest binding energy was selected as the final representative pose. Final poses were visualized as 2D diagrams using Biovia Discovery Studio Visualizer v19.1.0.18287 to illustrate ligand-protein interactions. To quantify the binding strength and inhibitory potential of the docked ligands, their inhibition constants (Ki) and inhibitory efficiencies (Ki^-1) were calculated using the following equations:
Where:
-
ΔG represents the binding energy derived from docking simulations.
-
R is the gas constant (1.987 × 10^-3 kcal/mol).
-
T is the absolute temperature (298.15 K).
Visual analysis of the docking results was performed using PyMol and Biovia Discovery Studio 2021 to gain further insights into the ligand-protein interactions.
Grid generation and molecular docking analysis
Using AutoDock 4.2 and AutoDock Vina, all the chosen phytochemicals were docked into the binding site of the NLRP3 protein’s Chain A, adhering to the protocol outlined by Morris et al. (1998)43. Using the modeling tool and visualizer in BIOVIA Discovery Studio, protein optimization was carried out as part of chain preparation by first eliminating water and other atoms and then adding a polar hydrogen group. The grid-based approach and blind docking were used to enable quick assessment of the conformational binding energy. The target proteins were placed into the grid box, which was created using AutoDock 4.2. While enclosing the active residues of the protein, the XYZ coordinates were changed as per the size of the protein. It guaranteed that ligand molecules and proteins would bind precisely in that defined area. The XYZ dimension was 60 × 90 × 66 and the grid spacing was 1 A°. The active site served as the grid’s center, and the macromolecule’s XYZ coordinates were 118.184 170.692 128.025 A°. For docking, all other settings were left at their initial configurations. The values for exhaustiveness and energy range are 10 and 4, accordingly.
Pre-docking process
For parameterized optimization of the improved docking simulation in the AutoDock 4.2, the ligand and proteins were pre-processed. The partial kollman and gasteiger charges, which are based on electron density and improve computation accuracy, were added in order to optimize the protein (Gasteiger and Marsili 1980)44. After allocating AD4 type atoms, the parametrized pdb protein file was saved as a pdbqt file. While the ligand was optimized by choosing and detecting the root and futher saved to pdbqt file.
Enrichment and string network analysis for NLRP3 protein
In the pursuit of categorizing the cellular pathways associated with the NLRP3 protein, KEGG annotations were extracted from the KEGG database (https://www.genome.jp/kegg/pathway.html)45. The intention behind this retrieval was to systematically assess the amalgamation of interactive information emanating from NLRP3 and its affiliated proteins, thereby implying pathways entwined in diverse biological functions. For the exploration of potential interactions among genes linked to NLRP3, the Search Tool for Retrieval of Interacting Genes (STRING) database (https://string-db.org) was harnessed46. STRING, functioning as a predictive and integrative platform for protein–protein interactions, enabled the identification of functional relationships and intricate interactions between proteins. Employing the STRING tool facilitated the examination of potential gene interactions associated with NLRP3, revealing a network characterized by nodes, edges, average node degrees, protein–protein interaction (PPI) enrichment p-values, as well as average and local clustering coefficients. Notably, STRING not only served as a tool for elucidating gene interactions but also provided a comprehensive platform suggesting plausible biological processes, molecular functions, and cellular components associated with the candidate genes under scrutiny. This multifaceted analysis, derived from the STRING database, allowed for a nuanced understanding of the functional implications of NLRP3 and its associated proteins within cellular pathways. The integration of KEGG annotations and STRING database analyses contributes to a holistic exploration of the cellular pathways and interactive relationships involving NLRP3. This combined approach not only enriches our comprehension of the molecular landscape associated with NLRP3 but also provides valuable insights for further scientific inquiry and potential therapeutic interventions.
Atlas Map of protein contacts
The Protein Contacts Atlas (https://pca.mbgroup.bio) is an interactive web database for visualizing atomic-level interactions between ligands, molecules, proteins, and nucleic acids. It utilizes residue-residue interaction networks and computational modeling of crystal structures to enable multi-scale visualization of complexes. By precomputing structural data from experimental and predictive sources, this tool allows interactive exploration of contacts within and between biological molecules from proteins to small ligands.
Post dock computational ADME/T and Pharmacokinetics studies
Drug-likeness and in silico ADMET studies
The SwissADME, ProTox 247,48 online tool, along with the Molinspiration property engine, was used to provide computational predictions about the toxicity, pharmacokinetics, drug-likeness, and ADME of several drugs. Determined by49 Lipinski’s rule of five, molecular weights (MW), hydrogen bond acceptor (HBA), hydrogen bond donor (HBD), topological polar surface area (TPSA), octanol/water partition coefficient (LogP), and rotatable bond count (RB) were all determined through physicochemical analysis49. Results started to show up soon as the canonical smiles of all compounds were submitted to the Molinspiration, ProTox 2 and ADMET/SAR programmes. A CSV file containing the SwissADME database’s results was downloaded. Mutagenic, tumorigenic, reproductively active, and irritating were listed as the expected toxicities50. The Molinspiration database (http://www.molinspiration.com) was used to forecast the bioactivities of phytochemicals. The substances’ expected bioactivities included GPCR ligand, nuclear receptor ligand, kinase inhibitor, protease inhibitor, enzyme inhibitor, and ion channel modulator51. All ligands’ gastrointestinal absorption and access to the brain were predicted using the BOILED-Egg graph (Brain Or Intestinal permeation method)47,48. These characteristics are essential when looking for a new treatment, hence all ligands make ideal candidates for being possible medications (Figs. 12–14). The Swiss ADME service calculated the bioavailability radar to determine how closely the molecule resembled the medication. For ligands, lipophilicity (LIPO) ranged between 1.01 < XlogP3 < 2.03, whereas polarity (POLAR) fell between 54.56 and 111.46 TPSA, where TPSA stands for topological polar surface area.
Collection of experimental ADMET data
In addition to the computational predictions, experimental ADMET data for several of the selected compounds were obtained from published literature to allow for comparison and validation of the in silico results. Relevant studies that provided experimental data on solubility, absorption, and toxicity for similar alkaloids were identified from some peer-reviewed articles (e.g., Hodges et al., 2019; Wu et al., 2022; Zhang et al., 2023). This data was used to assess the accuracy of the computational predictions by comparing the predicted and experimentally validated values.
Data comparison and analysis
The predicted ADMET values were compared with available experimental data to evaluate the consistency and accuracy of the computational predictions. A detailed comparison was carried out for key ADMET properties, and any discrepancies between the predicted and experimental results were noted and discussed. This comparison aims to highlight the potential limitations of in silico approaches and the importance of experimental validation in drug development.
Bioactivity score and bio-availability radar
Ligand bioactivity scores were assessed using the Molinspiration software, available online at http://www.molinspiration.com/. Canonical SMILES of ligands sourced from PubChem were employed in this analysis. Properties scrutinized encompassed G-protein coupled receptors (GPCR), enzyme inhibitors (EI), kinase inhibitors (KI), nuclear receptor ligands (NRL), and ion channel modulators (ICM). Additionally, the bioavailability of the ligands was determined using Swiss ADME (http://www.swissadme.ch/index.php), which rapidly indicates whether a compound exhibits oral bioavailability. This integrated approach provides a comprehensive evaluation of ligand properties, essential for understanding their potential biological activities and therapeutic relevance.
The crystal structure and binding pocket of Chain A of NLRP3.
2D chemical structure of all the compounds.
The predicted secondary structure of the Chain A of NLRP3 using SOPMA software, whereas h: Alpha helix, e: Extended strand, t: Beta-turn and c: Random coil. The graphical representation of secondary elements in Chain A of NLRP3 protein (blue: alpha helix, red: extended, green: beta turn, yellow: random coil).
Human NLRP3 receptor-binding domain Z-score and plot of residue scores. (A) ProSA-web z-scores of all protein chains. (B) Energy plot. (C) Highest and lowest energies of protein.( ProSA-web server: https://prosa.services.came.sbg.ac.at/prosa.php).
Swiss-modelled prediction for chain A of NLRP3. (A) QMEAN Z-Scores (B) Modelled structure (C) Ramachandran Plots (D) Complete protein sequence of Chain A of NLRP3(931 amino acid) (https://swissmodel.expasy.org/).
Result and discussion
Molecular docking was first performed to predict protein-ligand interactions by minimizing ligand energy and calculating binding energies. Ligand binding inhibits the target enzyme, demonstrating the potential to alter its biochemical activity. Docking algorithms relate a drug’s structure to its cytotoxicity by evaluating inhibitory and activator properties of the ligand with the receptor protein. In summary, molecular docking studies predicted inhibitory interactions between phytochemical ligands and target proteins to assess their potential biochemical effects. To perform the target protein analysis, Cryo-EM structure of NLRP3 bound to NEK complex (PDB ID: 6NPY) was were retrieved from PDB databank from https://www.rcsb.org/structure/6NPY. The NEK7 were served as an HETATOM(heteroatom) and was removed along with the water molecule (Fig. 2).
Secondary structure prediction
The secondary structure prediction of chain A in protein NLRP3 revealed a composition of 29.08% α-helix (h), 27.12% extended strand (e), 11.44% beta-turn (t), and 32.35% random coil (c) elements. The results are visualized through two graphs, one illustrating the prediction and the other displaying score curves for all predicted states (Fig. 4). Additionally, pertinent parameters such as window width and the number of states utilized for the prediction are presented, offering a comprehensive insight into the methodology employed for this structural analysis. Figure 3 represents all the 2D Structures of phytochemicals used in this study.
Protein chain validation
The selected protein chain exhibited a Z-score within the range of native structures, validating its high quality as assessed by ProSA and Swiss- model analysis (Fig. 5 and Fig. 6). The QMEAN Z-Score of 0.72 for NLRP3 (Fig. 6A) indicates perfect selected structures score. ProSA analysis of human NLRP3 yielded a Z-score of -10.22, validating overall quality by comparing total energy deviation to random configurations. The ProSA results (Fig. 5) plot acquired protein structures from various sources differentiated by color (Fig. 5A), allowing assessment of the selected NLRP3 protein chain against experimental structures. These computational validations confirm the quality of the selected protein chain.
Molecular docking analysis
All the compounds (alkaloids) were selected on the on the basis of their anti-inflammatory properties and were downloaded from Pubchem database. Prepared Chain A of NLRP3 protein were docked with all the selected alkaloids. The docking score of all the significant interactions (above ~ 10 kcal/mol) is shown in Table 3. All the compounds showed outstanding binding energies with the respective to Chain A of NLRP 3 protein. Tranilast (Synthetic Positive Control) and Oridonin (Natural Positive Control) were taken as controls for the target as it has been currently used and have shown good efficacy as selective drugs against NLRP3 induced inflammasoms. The results of binding affinities, inhibition constant (Ki) and ligand efficiency (L.E) obtained are shown in Table 3. The docking results presented in Table 1 indicate a strong interaction between the selected bioactive compounds and key amino acid residues in the binding pocket of Chain A of the NLRP3 protein. Notably, Oxyacanthine exhibited the highest binding affinity (-15.2 kcal/mol), with pi-cation and charge-charge interactions involving crucial residues such as GLU 375, HIS 724, and ARG 172. Similarly, Berbamine demonstrated a binding affinity of -13.9 kcal/mol, showing robust interactions with ARG 167, TRP 416, and ILE 521. The control compounds, Tranilast and Oridonin, also exhibited significant binding affinities. Oridonin showed a notable binding affinity of -11.6 kcal/mol, primarily through conventional hydrogen bonding and carbon-hydrogen interactions, further validating the binding potential of the selected phytochemicals when compared to known positive controls. These molecular interactions emphasize the importance of hydrogen bonding, pi-cation, and charge-charge interactions in stabilizing the ligand-protein complex. The inhibition constants (Ki values) further suggest the potential efficacy of these compounds, with Berbamine and Oxyacanthine displaying lower Ki values (0.122 µM and 0.1679 µM, respectively), indicating a stronger inhibitory potential against Chain A of the NLRP3 protein. While in vitro experiments are necessary to confirm these findings, our current study lays a solid foundation for future exploration of these compounds as potential NLRP3 protein inhibitors. Due to current resource constraints, additional in vitro validation is beyond the scope of this research. Nevertheless, the docking data, presented in Tables 3 and 4, offers valuable insight into the binding interactions and supports further experimental investigation.
Calculation for inhibition constant, binding energies and ligand efficiency
-
The inhibition constant (Ki) (µM/nM) for the ligands was calculated from the binding energy (ΔG) using the equation Ki (µM) = exp (ΔG/RT), where R is the gas constant (1.985 × 10 − 3 kcal/mol − 1 K − 1) and T is the temperature (298.15 K). This allowed prediction of Ki values for the ligand interactions from the computed binding energies.
-
The Gibbs free binding energy was calculated using udergiven formula:
ΔG = − RTlnKi …… 1 equation.
Ki = e[−ΔGRT] ……2 equation.
where R = Gas constant (1.987 × 10–3 kcal/mol); T = 298.15 K (absolute temperature); Ki = Inhibition constant; ΔG = Binding energy (the binding free energy).
-
The ligand efficiency was calculated using the under given formula:
Ligand Efficiency(LE) = − B.E ÷ Heavy atoms (H.A).
LE scale = 0.873e − 0.026× H.A − 0.064.
FQ = LE ÷ LE scale.
STRING protein network and KEGG enrichment analysis of NLRP3
The study employed KEGG analysis to explore the biological significance of the identified target and potential involvement of NLRP3 protein. Through protein-protein interaction (PPI) network analysis, a set of 19 key targets, including NEK 7, PYCARD, DHX33 along with GBPs (GTPase binding proteins) were identified through PPI network analysis. These targets were primarily associated with biological processes linked to NLRP3 and its associated protein. The KEGG enrichment analysis revealed signalling pathways closely linked to inflammation system, as illustrated in Fig. 7. The STRING protein interaction network of NLRP3 was analyzed using the STRING database. Specific evidence lines represented by colored links between nodes denote functional associations, with confidence indicated by internode distances per Bayesian scoring (Fig. 8). Co-expression analysis utilized gene correlation testing across large transcriptomic and proteomic datasets to infer functional relationships. Observed bovine gene co-expression is depicted in heatmaps (Fig. 9), where color intensity in triangle matrices indicates confidence of association between co-expressed proteins. Additionally, gene co-occurrence similarities provided computed protein similarity scores between organisms, with color intensity describing the degree of inter-protein similarity (Fig. 9A and B). This comprehensive examination enhances our understanding of the intricate interplay among these targets and their relevance to critical pathways implicated in both inflammation and its related systems.
Contacts Atlas of NLRP3 chain A
The Protein Contacts Atlas tool provided atomic-level visualization of the NLRP3 protein complex. Nodes in the network represent nucleic acids or proteins, with links between nodes denoting interaction surfaces between subunits. Chord plots displayed secondary structure contacts in NLRP3 (Fig. 10A). Asteroid plots showed the residue/ligand-centric atomic neighbourhood (Fig. 10B). Scatter plots provided quantitative insights per residue (Fig. 10C). Together, these models enabled multi-scale study of the NLRP3 complex from secondary structure contacts to atomic interactions using network graph, asteroid plot, and scatter plot visualizations.
Bioactive and bioavailability score
Table 5 showed the Molinspiration bioactivity predicted score of all the alkaloids. Predicted data suggests that this class of nitrogenous organic compounds possesses diverse potential biological activities. Notably, they may interact with G protein-coupled receptors (GPCRs), modulate ion channels, inhibit kinases, bind strongly to nuclear receptors, mildly inhibit proteases, and moderately inhibit enzymes. These predictions offer valuable insights into the compounds’ potential pharmacological effects and pave the way for further experimental investigations into their specific biological actions. Our results highlight five natural compounds – Magnoflorine, Corynoline, Columbamine, Palmatine, and Oridonin – as potential NLRP3 protein interactors. The evaluated scores indicate that values greater than 0.00 correspond to high activity, scores ranging from − 0.5 to − 0.00 suggest moderate activity, and scores lower than − 0.5 imply inactivity52. Out of all the ligands, Magnoflorine, Corynoline, Columbamine Palmatine, Tranilast, Oridonin (Natural compound) were found to have good bioactivity scores. For GPCR and ICM Magnoflorine showed the highest score of 0.37 and 0.78, respectively; palmatine and columbamine showed the highest score of 0.82;0.74 for ICM and 0.90;0.81 for EI; Corynoline had the highest score of 0.28 for GPCR. The remaining scores are represented in the table, which indicates the scoring profile for all the ligand compounds. Out of the two selected Standard drug Oridonin indicate the most significant bioactive score for ICM, NR and EI. Their predicted and comparable bioactivity scores to control compounds suggest that these phytochemicals could serve as alternative or adjuvant therapeutic agents for NLRP3-related inflammatory diseases.
By evaluating all these parameters, a bioavailability radar profile was predicted and the graphical output for all the alkaloid compounds are presented in Fig. 11 This radar plot helps in assessing the viability of new drug candidates. For a phyto-compound to be considered drug-like, its radar plot must fall within the optimal pink area across all parameters. Two essential determinants of bioavailability are flexibility and polarity. Compounds with over 10 rotatable bonds, indicating high flexibility, are predicted to have low oral bioavailability. Compounds with a topological polar surface area between 20 and 130 Å2 have optimal polarity for high oral bioavailability. By assessing flexibility and polarity along with other key parameters, bioavailability radars can effectively predict the oral absorbability of novel compounds in a drug discovery pipeline. Based on radar plot data all the compound seems to fulfil the minimum criteria to be orally bioavailable (Fig. 8).
Drug-likeness and in silico ADMET studies
In silico pharmacokinetic analysis revealed the phytochemicals have favorable drug-likeness and ADMET properties comparable to standard drugs, including high gastrointestinal bioavailability but low blood-brain barrier permeability as predicted by ADMET modelling (Fig. 11) (Data are tabulated for metabolism and toxicity in Table 6 and Table 7). All the alkaloids Corynoline, Columbamine, Berberine, Palmatine and Tranilast satisfied all of Lipinski’s criteria, while other alkaloids deviated in at most one parameter, still in the acceptable range (Table 8). Estimated solubility revealed the phytochemicals as moderately less soluble than the standards, except Oridonin which showed exceptionally high solubility high gastrointestinal absorption (GIA) and normal bioavailability scores were predicted for the ligands (Fig. 12). Importantly, Corynoline, Oxyacanthine, Columbamine, Berberine, Neferine and Tranilast were identified as non-substrates of p-glycoprotein, making them preferable for cancer treatment by avoiding efflux53,54. Regarding cytochrome P450 inhibition, the phytochemicals showed a balance of inhibitors/non-inhibitors of CYP3A4 and CYP1A2, minimizing risks of undesirable metabolic drug interactions55. In summary, in silico screening predicted that these alkaloid phytochemical compounds have favourable pharmacokinetics and ADME properties, including solubility, permeability, bioavailability, and limited interactions with clearance mechanisms, supporting their potential as therapeutic agents. In silico predictions for experimental ADMET properties for several of the alkaloids considered in this work have appeared. For example, Palmatine an alkaloid has significant pharmacological properties, including anticancer, antimicrobial, and anti-inflammatory effects. Palmatine shows dose-dependent growth inhibition on cancer cells, particularly breast cancer, and demonstrates synergistic effects when used with doxorubicin. The pharmacokinetics of palmatine highlight that it is metabolized primarily in the liver, with further research needed to better understand its bioavailability and excretion patterns. However, clearances that differ from both predicted and experimental values suggest the necessity for higher resolution and accuracy with such computational tools for truly reliable in vivo simulations. Similarly, experimental solubility and permeability problems of berberine were only partially confirmed by SwissADME and ADMET-SAR predictions. Besides, the comparison with the experimental data is essential for the validation of the performance of the computational models in drug discovery.
Water solubility and lipophilicity
The in-silico predictions indicate the phytochemical compounds have a wide range of lipophilicity (log Po/w 0.65–5.47) but are poorly water-soluble (log S -2.15 to -7.80) (Table 9). While the high lipophilicity may improve permeation into cells, it could also lead to solubility issues and potential toxicity. The compounds seem to violate Lipinski’s guidelines for lipophilicity, which may negatively impact oral absorption and bioavailability. However, the poor aqueous solubility suggests that specialized formulations like prodrugs or nanocarriers may be required to enable parenteral delivery. In summary, the high lipophilicity but low solubility presents both challenges and opportunities in developing the phytochemicals as therapeutic agents, necessitating innovative solutions to improve their delivery while retaining bioactivity.
Comparison of predicted and experimental ADMET properties
In this study, we utilized in-silico tools, including ADMET-SAR, SwissADME, and Protox II, to predict the ADMET properties of selected alkaloids. These predictions were then compared to experimental ADMET data collected from the literature for two of the compounds. The experimental data provided values for properties such as solubility, bioavailability, and toxicity, enabling us to evaluate the accuracy of the computational predictions. These studies provided validated ADMET profiles for some alkaloids that overlap with the compounds investigated in our research. A comparison of the predicted and experimental values is presented in Table 10. In general, there was a moderate to high agreement between the predicted and experimental values for several key ADMET properties, such as solubility and bioavailability. For example, the predicted solubility of Palmatine using SwissADME was consistent with the experimental data reported by Long et al. (2019)56, showing high correlation. Similarly, the predicted ADMET properties of Berbarine using SwissADME aligned with the experimental results from Ai et al. (2022)57. However, some discrepancies were noted, in predicted parameters from the experimental values. This deviation could be attributed to limitations in the in silico models or variations in experimental conditions, such as differences in compound formulation or testing environments. These comparisons underscore the value of using experimental data to validate computational predictions. While in silico methods provide a quick and efficient way to estimate ADMET properties, experimental validation remains critical to ensure the accuracy and reliability of the predictions. This integrated approach enhances the robustness of the ADMET analysis, providing a more comprehensive understanding of the pharmacokinetic profiles of the compounds under investigation.
KEGG analysis of NLRP3 involved inflammatory pathway.
STRING analysis NLRP3 chain A. Representing all the interacting sensor partners of the NLRP3 inflammasome (On the left Fig. A) and Guanylate-binding protein (GBP) an activator of NLRP3 inflammasome assembly (On the right Fig. B).
STRING analysis NLRP3 chain A. (A) Gene co-occurrence (B) Gene co-expression with legends.
Protein contact atlas of Human NLRP3 chain A: (A) Chord plot. (B) Asteroid plot. (C) Scatter plot.
The SwissADME radar depicts multiple bioactive drug-like molecules, with the red regions indicating characteristics related to lipophilicity, solubility, molecular weight, and versatility.
Conclusion
The findings are poised to capture the interest of researchers in the field of drug discovery targeting the NLRP3 inflammasome, particularly through the utilization of alkaloids. These alkaloids have the potential to function as phyto-drugs, addressing various related inflammatory disorders with natural phytocompounds. The approach involves leveraging natural bioactive components, and similar investigations can be conducted using various medicinal herbs. Moreover, our study opens avenues for further exploration into the effects of these phytocompounds in other inflammatory diseases of relevance. This computational study unveils the potential of bioactive compounds to proficiently bind to the PYD (NACHT) region of NLRP3, thereby inducing inhibition of its interaction with nearby effector domains. This inhibition, in turn, impedes the activation of the inflammasome cascade and its subsequent assembly. The findings suggest a promising avenue for the development of therapeutic strategies targeting NLRP3-mediated inflammation. To bolster these promising results, further investigation is imperative to elucidate the pharmacodynamics and kinetic properties of these phytochemicals. Understanding how these compounds interact with NLRP3 at a molecular level and the specific dynamics of these interactions will be crucial for optimizing treatment approaches. Moreover, a more in-depth exploration is warranted to unravel the detailed mechanism of action employed by these phytochemicals in effectively treating inflammation and associated diseases. This could involve dissecting the signaling pathways affected by the interaction, exploring downstream effects on cytokine release, and evaluating the broader impact on immune responses. In summary, while this computational study provides valuable insights into the inhibitory potential of bioactive compounds targeting the NLRP3 inflammasome, subsequent research endeavors should focus on establishing the pharmacokinetic properties of these compounds and delving into the intricacies of their mechanism of action. Such investigations will pave the way for the development of more targeted and effective therapeutic interventions for inflammatory conditions and related diseases. While molecular docking can aid hit/lead identification and optimization in drug design, experimental validation remains essential. The four hit compounds identified computationally require further in vitro and in vivo testing to confirm their therapeutic potential against inflammatory diseases. However, lack of funding currently limits experimental follow-up. Still, these in silico results provide initial targets that warrant future development pending financial support, ultimately aiming to find reliable and effective drugs or vaccines to treat detrimental inflammation.
BOILED-Egg diagram for all the selected compounds. The compounds that are grouped closely in the blood-brain region, exhibiting a TPSA between 20 to 70 and a WLOGP value between 1 to 4.
Molecule 1: Oxyacanthine.
Molecule 2: Neferine.
Molecule 3: Magnoflorine.
Molecule 4: Corynoline.
Molecule 5: Columbamine.
Molecule 6: Berberine.
Molecule 7: Berbamine.
Molecule 8: Palmatine.
Molecule 9: Tranilast (Positive Control).
Molecule 10: Oridonin (Positive control).
Data availability
All relevant data is included within the manuscript and its supplementary information.
References
Harry, G. J., Childers, G. M. & McPherson, C. A. An introduction to innate immunity in the central nervous system. In Advances in Neurotoxicology (Vol. 3, 1–34). Academic. (2019).
Schnappauf, O., Chae, J. J., Kastner, D. L. & Aksentijevich, I. The pyrin inflammasome in health and disease. Front. Immunol. 10, 1745 (2019).
Ren, K. & Torres, R. Role of interleukin-1β during pain and inflammation. Brain Res. Rev. 60 (1), 57–64 (2009).
Xu, S. et al. Inflammasome inhibitors: promising therapeutic approaches against cancer. J. Hematol. Oncol. 12 (1), 1–13 (2019).
Man, S. M. & Kanneganti, T. D. Regulation of inflammasome activation. Immunol. Rev. 265 (1), 6–21 (2015).
Ting, J. P. Y. et al. The NLR gene family: a standard nomenclature. Immunity. 28 (3), 285–287 (2008).
Martinon, F., Burns, K. & Tschopp, J. The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-β. Mol. Cell. 10 (2), 417–426 (2002).
He, Y., Hara, H. & Núñez, G. Mechanism and regulation of NLRP3 inflammasome activation. Trends Biochem. Sci. 41 (12), 1012–1021 (2016).
Franchi, L., Warner, N., Viani, K. & Nuñez, G. Function of nod-like receptors in microbial recognition and host defense. Immunol. Rev. 227 (1), 106–128 (2009).
Swanson, K. V., Deng, M. & Ting, J. P. Y. The NLRP3 inflammasome: molecular activation and regulation to therapeutics. Nat. Rev. Immunol. 19 (8), 477–489 (2019).
Bauernfeind, F. G. et al. Cutting edge: NF-κB activating pattern recognition and cytokine receptors license NLRP3 inflammasome activation by regulating NLRP3 expression. J. Immunol. 183 (2), 787–791 (2009).
Duncan, J. A. et al. Cryopyrin/NALP3 binds ATP/dATP, is an ATPase, and requires ATP binding to mediate inflammatory signaling. Proc. Natl. Acad. Sci. 104 (19), 8041–8046 (2007).
Coll, R. C. et al. A small-molecule inhibitor of the NLRP3 inflammasome for the treatment of inflammatory diseases. Nat. Med. 21 (3), 248–255 (2015).
Coll, R. C. et al. MCC950 directly targets the NLRP3 ATP-hydrolysis motif for inflammasome inhibition. Nat. Chem. Biol. 15 (6), 556–559 (2019).
Tapia-Abellán, A. et al. MCC950 closes the active conformation of NLRP3 to an inactive state. Nat. Chem. Biol. 15 (6), 560–564 (2019).
Mullard, A. NLRP3 inhibitors stoke anti-inflammatory ambitions. Nat. Rev. Drug Discovery. 18 (6), 405–408 (2019).
Eren, E. & Özören, N. The NLRP3 inflammasome: a new player in neurological diseases. Turkish J. Biology. 43 (6), 349–359 (2019).
Dixit, V. D. Nlrp3 inflammasome activation in type 2 diabetes: is it clinically relevant? Diabetes. 62 (1), 22 (2013).
Rheinheimer, J., de Souza, B. M., Cardoso, N. S., Bauer, A. C. & Crispim, D. Current role of the NLRP3 inflammasome on obesity and insulin resistance: a systematic review. Metabolism. 74, 1–9 (2017).
Karasawa, T. & Takahashi, M. Role of NLRP3 inflammasomes in atherosclerosis. J. Atheroscler. Thromb. 24 (5), 443–451 (2017).
Baldrighi, M., Mallat, Z. & Li, X. NLRP3 inflammasome pathways in atherosclerosis. Atherosclerosis. 267, 127–138 (2017).
Szekanecz, Z., Szamosi, S., Kovács, G. E., Kocsis, E. & Benkő, S. The NLRP3 inflammasome-interleukin 1 pathway as a therapeutic target in gout. Arch. Biochem. Biophys. 670, 82–93 (2019).
Zheng, F., Xing, S., Gong, Z. & Xing, Q. NLRP3 inflammasomes show high expression in aorta of patients with atherosclerosis. Heart Lung Circulation. 22 (9), 746–750 (2013).
Menu, P. & Vince, J. E. The NLRP3 inflammasome in health and disease: the good, the bad and the ugly. Clin. Experimental Immunol. 166 (1), 1–15 (2011).
Guo, H., Callaway, J. B. & Ting, J. P. Inflammasomes: mechanism of action, role in disease, and therapeutics. Nat. Med. 21 (7), 677–687 (2015).
Li, L., Sun, X., Zhao, D. & Dai, H. Pharmacological applications and action mechanisms of phytochemicals as alternatives to antibiotics in pig production. Front. Immunol. 12, 798553 (2021).
Nisar, A. et al. Phytochemicals in the treatment of inflammation-associated diseases: the journey from preclinical trials to clinical practice. Front. Pharmacol. 14, 1177050 (2023).
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 79 (3), 629–661 (2016).
Chen, Q. L., Yin, H. R., He, Q. Y. & Wang, Y. Targeting the NLRP3 inflammasome as new therapeutic avenue for inflammatory bowel disease. Biomed. Pharmacother. 138, 111442 (2021).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30 (16), 2785–2791. https://doi.org/10.1002/jcc.21256 (2009).
Geourjon, C. & Deléage, G. SOPMA: significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Comput. Appl. Biosciences: Cabios. 11 (6), 681–684. https://doi.org/10.1093/bioinformatics/11.6.681 (1995).
Dallakyan, S. & Olson, A. J. Small-molecule library screening by docking with PyRx. Methods Mol. Biol. 1263, 243–250 (2015).
Allinger, N. L. Conformational analysis. 130. MM2. A hydrocarbon force field utilizing V1 and V2 torsional terms. J. Am. Chem. Soc. 99 (25), 8127–8134 (1977).
O’Boyle, N. M. et al. Open Babel: an open chemical toolbox. J. Cheminform. 3, 33. https://doi.org/10.1186/1758-2946-3-33 (2011).
Murugananthan, G. & Mohan, S. Anti-arthritic and anti-inflammatory constituents from medicinal plants. J. Appl. Pharm. Sci. 3 (4), 161–164 (2013).
Asokan, S. M., Mariappan, R., Muthusamy, S. & Velmurugan, B. K. Pharmacological benefits of neferine-A comprehensive review. Life Sci. 199, 60–70 (2018).
Xu, T. et al. Magnoflorine: a review of its pharmacology, pharmacokinetics and toxicity. Pharmacol. Res. 152, 104632 (2020).
Liu, B. et al. Corynoline exhibits anti-inflammatory effects in lipopolysaccharide (LPS)-stimulated human umbilical vein endothelial cells through activating Nrf2. Inflammation. 41, 1640–1647 (2018).
Liu, X. et al. Anti-inflammatory and anti-nociceptive activities of compounds from Tinospora sagittata (Oliv.) Gagnep. Arch. Pharm. Res. 33, 981–987 (2010).
Jia, X. J. et al. Berbamine exerts anti-inflammatory effects via inhibition of NF-κB and MAPK signaling pathways. Cell. Physiol. Biochem. 41(6), 2307–2318 (2017).
Saeedi-Boroujeni, A. et al. Tranilast: a potential anti-inflammatory and NLRP3 inflammasome inhibitor drug for COVID-19. Immunopharmacol. Immunotoxicol. 43(3), 247–258 (2021).
Matsui, T. et al. Lansiumamide B and SB-204900 isolated from Clausena lansium inhibit histamine and TNF-α release from RBL-2H3 cells. Inflamm. Res. 62, 333–341 (2013).
Morris, G. M. et al. Automated docking using a lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 19 (14), 1639–1662 (1998).
Gasteiger, J. & Marsili, M. Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron. 36 (22), 3219–3228 (1980).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51 (D1), D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Szklarczyk, D. et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43 (D1), D447–D452 (2015).
Daina, A. & Zoete, V. A BOILED-Egg to predict gastrointestinal absorption and brain penetration of small molecules. ChemMedChem. 11 (11), 1117–1121 (2016).
Daina, A., Michielin, O. & Zoete, V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 7, 42717 (2017).
Lipinski, C. A. Lead-and drug-like compounds: the rule-of-five revolution. Drug Discovery Today: Technol. 1 (4), 337–341 (2004).
Daina, A., Michielin, O. & Zoete, V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 7 (1), 42717 (2017).
Sander, T., Freyss, J., von Korff, M. & Rufener, C. DataWarrior: an open-source program for chemistry aware data visualization and analysis. J. Chem. Inf. Model. 55 (2), 460–473 (2015).
Khan, T. et al. Molecular docking, PASS analysis, bioactivity score prediction, synthesis, characterization and biological activity evaluation of a functionalized 2-butanone thiosemicarbazone ligand and its complexes. J. Chem. Biol. 10 (3), 91–104. https://doi.org/10.1007/s12154-017-0167-y (2017).
Amin, M. L. P-glycoprotein inhibition for optimal drug delivery. Drug Target. Insights. 7, 27–34. https://doi.org/10.4137/DTI (2013).
Bansal, T., Jaggi, M., Khar, R. K. & Talegaonkar, S. Emerging significance of flavonoids as P-glycoprotein inhibitors in cancer chemotherapy. J. Pharm. Pharm. Sciences: Publication Can. Soc. Pharm. Sci. Societecanadienne Des. Sci. Pharmaceutiques. 12 (1), 46–78. https://doi.org/10.18433/j3rc77 (2009).
Ji, D., Xu, M., Udenigwe, C. C. & Agyei, D. Physicochemical characterisation, molecular docking, and drug-likeness evaluation of hypotensive peptides encrypted in flaxseed proteome. Current Research in Food Science, 3, 41–50. (2020). https://doi.org/10. 1016/j. crfs. 2020. 03. 001.
Long, J. et al. Palmatine: a review of its pharmacology, toxicity and pharmacokinetics. Biochimie. 162, 176–184 (2019).
Ai, X., Yu, P., Peng, L., Luo, L., Liu, J., Li, S., … Meng, X. (2021). Berberine:a review of its pharmacokinetics properties and therapeutic potentials in diverse vascular diseases. Frontiers in pharmacology, 12, 762654.
Author information
Authors and Affiliations
Contributions
Nilay Singh: Methodology, Data curation, Investigation, Writing - original draft. Manoj K. Pal: Methodology, Formal analysis. Promila Sharma: Supervision, Validation. Kumud Pant: Resources, Data curation, Formal analysis. Ragini Kahera and Himani Badoni: Reviewing & editing the original draft, Literature survey. Neetu Sharma and Bhawana Bhist: Conceptualization, Supervision, Methodology, Preparing the revision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Singh, N., Sharma, P., Pal, M.K. et al. Computational drug discovery of phytochemical alkaloids targeting the NACHT/PYD domain in the NLRP3 inflammasome. Sci Rep 15, 16677 (2025). https://doi.org/10.1038/s41598-024-79054-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-79054-2
Keywords
This article is cited by
-
Corynoline promotes apoptosis and inhibits proliferation of glioblastoma via regulating the STAT3/Bcl-2 signalling pathway
Naunyn-Schmiedeberg's Archives of Pharmacology (2025)