Abstract
Nidogen-1 (NID1) is a secreted glycoprotein widely distributed in basement membranes. NID1 interacts with extracellular matrix proteins such as collagen and laminin and has been implicated in the progression of various cancers. However, study on the role of NID1 in breast cancer is scarce and inconsistent. In this work, we found that the expression of NID1 is significantly lower in breast cancer tissue than in normal tissue. In addition, NID1 expression correlated negatively with a poor prognosis for breast cancer patients. Based on those findings, we speculated that NID1 might act as a cancer suppressor in breast cancer. To investigate the role of NID1 in breast cancer, we constructed NID1-overexpressing cell lines. NID1 overexpression decreased breast cancer cell proliferation, migration, and in vivo tumor growth. Moreover, glucose metabolism, which is known to enhance cancer cell proliferation and migration, was also decreased by NID1 overexpression. Mechanistically, NID1 overexpression downregulated hypoxia-inducible factor-1α (HIF-1α) expression at the transcription level. Furthermore, we found that NID1 reduced integrin β1 stability and downregulated the transcription of HIF-1α through the FAK/Src/NF-κB p65 signaling axis, which is downstream of integrin β1. Together, the results of this study demonstrate the tumor suppressive role of NID1 in triple-negative breast cancer.
Similar content being viewed by others
Introduction
The extracellular matrix (ECM) is composed of a complex of molecules that forms a three-dimensional structure. Many cellular processes, including cell proliferation, survival, motility, metabolism, and differentiation, are regulated by ECM1. Nidogen-1 (NID1), also known as entactin, is an ECM glycoprotein and a member of the nidogen family2. In the assembly of the basement membrane, NID1 acts as a bridge between ECM proteins such as laminin and collagen3. NID1 has also been reported to regulate the epithelial to mesenchymal transition (EMT) and metastasis in various cancers, including ovarian, liver, and colorectal cancer4,5,6. However, the role of NID1 in breast cancer is uncertain. Alečković et al.7 reported that NID1 overexpression induces lung metastasis in breast cancer and melanoma. But a study by Ferraro et al.8 showed that extracellular NID1 inhibits cell migration in the SK-BR-3 human breast cancer cell line.
Cancer cells exhibit abnormal metabolism compared with most normal cells. Unlike normal cells, which rely on mitochondrial oxidative phosphorylation to generate energy, some cancer cells rely on aerobic glycolysis, which is called the Warburg effect9. Aerobic glycolysis in cancer cells affects cell proliferation, migration, invasion, and resistance to treatment10,11. This unique metabolism requires large amounts of glucose and is accompanied by conversion of pyruvate to lactate for NAD+ regeneration12. Thus, cancer cells using aerobic glycolysis have increased expression of genes such as glucose transporter and lactate dehydrogenase-A (LDHA)13. Previous studies have reported that oncogenes such as Ras, Src, and c-Myc and transcription factors such as hypoxia-inducible factor-1 (HIF-1) regulate glycolysis-dependent cancer cell metabolism14. The metabolic state of cells is also known to be regulated by extracellular factors such as growth factor signaling, nutrients, and ECM components15. However, the correlation between basement membrane proteins such as nidogen and oncogenic metabolism has rarely been reported.
In this work, we found that NID1 expression correlates negatively with progression of breast cancer based on a patient database. In addition, triple-negative breast cancer (TNBC) cell lines overexpressing NID1 showed decreased cell proliferation, migration, invasion, and in vivo tumor growth compared with controls, and the glycolytic phenotype in TNBC cell lines was attenuated by NID1 overexpression. Furthermore, NID1 suppressed the expression of HIF-1α at the transcriptional level by inhibiting the integrin β1/FAK/Src/NF-κB p65 signaling axis. Taken together, our results suggest that NID1 can act as a tumor suppressor by causing metabolic shifts in TNBC.
Materials and methods
Cell culture
The MDA-MB-231 and Hs578T human breast cancer cell lines were obtained from the American Type Culture Collection (Manassas, VA, USA). MDA-MB-231 LM1 cell was provided by SJ Lee (Fibrosis and Cancer Targeting Biotechnology, Seoul, Korea). All cell lines were maintained in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. Cells were incubated at 37 ℃ in a humidified atmosphere with 5% CO2.
Generation of NID1-overexpressing cell lines
To generate NID1-overexpressing cell lines, HEK 293 T cells were transfected with pBABE-puro-vec or pBABE-puro-NID1, together with pCMV-VSVG and pCMV gag-pol (8:3:4 mass ratio). After 48 h, cell medium containing retroviral particles was collected and filtered with a 0.45 μm syringe filter. For transduction, cells were seeded at approximately 70% confluency and treated with culture medium containing retroviral particles and 10 μg/ml polybrene (Sigma-Aldrich). The NID1-overexpressing cell lines were selected by treatment with 10 μg/ml puromycin. The expression of NID1 was confirmed by western blotting and qRT-PCR.
Western blotting analysis
To prepare lysates, cells were washed with PBS and lysed with lysis buffer (0.1 mM EDTA, 150 mM NaCl, 1% NP-40, 0.1% Triton X-100, 0.1% SDS, 20 mM Tris-HCl, pH 7.4) supplemented with phosphatases and protease inhibitor (Roche, Basel, Switzerland). Protein concentrations were measured by standardization with a BCA protein assay kit (Thermo). Samples were mixed in SDS sample buffers, boiled for 10 min for denaturation, resolved by SDS-PAGE, and transferred to nitrocellulose membranes (Whatman, Dassel, Germany). The membranes were blocked with 5% skim milk in TBST and incubated with primary antibodies overnight. The next day, the membranes were incubated with HRP-conjugated secondary antibodies for 2 h, and then the proteins were visualized using Dyne ECL (Dyne, Seongnam, Korea). Protein bands were detected using X-ray film (AGFA, Mortsel, Belgium) or iBright CL1500 Imaging System (Invitrogen, Carlsbad, CA, USA). The list of antibodies used for the western blot analysis is provided in Table S1.
Protein precipitation from culture medium
Methanol/chloroform precipitation was performed to detect secreted proteins. Cells were seeded at a density of 5 × 105 cells per 60-mm dish. After 24 h, the culture media were harvested and centrifuged at 2000 rpm for 5 min to remove cell debris. After centrifugation, the culture media were subjected to methanol/chloroform precipitation, and the precipitated protein samples were analyzed by western blotting. Ponceau S staining was used for loading control for this analysis.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Whole cell RNA was extracted using Trizol reagent (MRC, Cincinnati, OH, USA), and RT-PCR was performed using an RT-PCR kit (Toyobo). qRT-PCR was performed using an SYBR qPCR kit (KAPA) in a Thermal Cycler Dice (Takara, Otsu, Shiga, Japan) according to the manual. Results were calculated relative to the control sample using the ΔΔCt method. The value of GAPDH was used as the internal control. The list of specific primers used for qRT-PCR is provided in Table S2.
Proliferation assay
Cells were seeded in 12-well plates at 2 × 104 cells per well. Every 24 h for 4 days, cells were harvested by trypsinization, resuspended in 1 ml of medium, and counted using a hemocytometer.
Two-dimensional colony formation assay
Cells were seeded in 6-well plates at 2 × 102 cells per well. After 10 days, the cells were washed with PBS and stained with 0.5% crystal violet for 30 min. The stained colonies were observed with an optical microscope.
Wound healing assay
Cells were seeded in 6-well plates and incubated to 90–95% confluency. A wound was generated by scraping the monolayer with a micropipette tip, and the cells were serum-starved overnight under normal incubation conditions. Wound closure was monitored microscopically at the indicated times.
Transwell migration and invasion assays
Cell migration assays were performed using transwell chambers (Corning, NY, USA) with 8-μm pores. The lower chamber was filled with DMEM containing 10% FBS. Next, 2 × 104 cells were suspended in DMEM without FBS and transferred to the upper chamber. After 18 h, cells that migrated through the chamber were fixed and stained with 0.5% crystal violet for 30 min. Those cells were washed twice with PBS and observed using an optical microscope. Cell invasion assays were performed using the same procedure except that the upper chamber was coated with Rat Collagen Type 1(Sigma-Aldrich, St. Louis, MO, USA) 2 h prior to cell seeding.
Immunofluorescence
At a density of 2 × 104 cells per well, cells were seeded in a 12-well plate with a coverslip. After 24 h, the cells were fixed and permeabilized using a 3.7% formaldehyde solution and 0.5% Triton X-100 for 10 min each. After that, blocking was performed for 30 min using 3% skim milk. To stain each target of interest, the primary antibody and fluorescence-conjugated secondary antibody were diluted in 1% skim milk and administered to the cells for 1 h each. To stain DNA, Hoechst 33342 dye (1 μg/ml) was administered to the cells for 10 min after target staining. Fluorescence was observed using a Nikon C2 Si plus confocal microscope (Nikon, Tokyo, Japan), and the data were analyzed using ImageJ software.
Cycloheximide chase assay
Cells were seeded in a 60-mm dish at a density of 5 × 105. After 24 h, 40 μg/ml cycloheximide (CHX) was administered to the cells, which were harvested at the indicated time points and subjected to western blotting.
Dual luciferase assay
Cells were seeded in a 24-well plate at a density of 5 × 104. After 24 h, 500 ng of reporter constructs and 5 ng of pRL-CMV were transfected into the cells using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA). After 24 h, luciferase activity was measured using a dual-luciferase reporter assay system (E1910, Promega, San Luis Obispo, CA, USA) according to the manufacturer’s protocol. Luminescence was measured using a GloMax 20/20 luminometer (Promega).
Cell viability assay
Cells were seeded in a 12-well plate at a density of 1 × 105. After 24 h, the cells were washed with PBS, and the medium was changed to glucose-free medium for glucose starvation (a control group was kept in glucose-containing medium). After incubation for 24 h, the cells were stained with trypan blue and counted manually. The viability of the cells was normalized to the viability of the control group.
Glucose uptake assay
Cells were seeded in a 96-well plate at a density of 1 × 104. After 24 h, the culture medium was changed to a 2-NBDG-containing medium, and the cells were incubated for 10 min. Then, the level of glucose uptake was measured using a glucose uptake cell-based assay kit (Cayman Chemical, Ann Arbor, MI, USA) according to the manufacturer’s protocol. The fluorescence intensity was measured using a microplate reader (Varioskan Flash, Thermo Scientific).
Lactate assay
Cells were seeded in a 60-mm dish at a density of 5 × 105. After 24 h, the culture medium was changed to phenol red-free medium, and the cells were incubated for another 24 h. The culture medium was harvested and centrifuged at 2000 rpm for 5 min to remove cell debris. Lactate assays were performed using a kit (Biovision, Milpitas, CA, USA) according to the manufacturer’s protocol. The colorimetric density was measured at 570 nm using a Multiscan EX device.
ATP assay
Cells were seeded in a 96-well plate at a density of 1 × 104. After 24 h, cellular ATP levels were examined using a CellTiter-Glo® luminescent cell viability assay kit (Promega) according to the manufacturer’s protocol. Luminescence was measured using a microplate reader (Varioskan Flash, Thermo Scientific).
Mitochondrial stress test
A mitochondrial stress test was performed using an XF Cell Mito stress test kit (Agilent Technologies). Cells were seeded in the XF cell culture microplate at a density of 1 × 104. After 24 h, the cell culture medium was replaced with basal medium containing 2 mM L-glutamine, 4 mM sodium pyruvate, and 25 mM glucose for 1 h. The oxygen consumption rate was measured in a Seahorse XF Pro analyzer (Agilent Technologies) with sequential addition of 1 μM oligomycin A, 0.5 μM FCCP, and 0.5 μM rotenone/antimycin A.
In-gel lactate dehydrogenase (LDH) isoenzyme assay
Cells were seeded in a 60-mm dish at a density of 5 × 105. After 24 h, the cells were harvested, washed with PBS, and lysed in lysis buffer (10 mM Tris-HCl, pH 7.4, 10 mM NaCl, 3 mM MgCl2, 0.1% Triton X-100) for 30 min. The lysate was extracted by centrifugation, and the protein concentration was quantified using a BCA protein assay. Each sample was loaded onto an 8% polyacrylamide gel without SDS. Electrophoresis proceeded for 2 h at 4 °C and 120 V. After electrophoresis, the gels were stained with 10 ml of staining solution (0.1 M lithium lactate, 1.5 mM NAD + , 0.1 M Tris-HCl, pH 8.6, 10 mM NaCl, 5 mM MgCl2, 0.3 mg phenazine methosulphate, and 2.5 mg nitro blue tetrazolium) for 10 min at 37 °C. The value of each subunit was quantified using ImageJ software.
Immunoprecipitation
Cells were washed in cold PBS and lysed in IP buffer (150 mM NaCl, 1% Triton X-100, 0.5% sodium deoxycholate, 2 mM EDTA, 10% glycerol, 50 mM Tris, pH 7.4) supplemented with phosphatases and protease inhibitor (Roche, Basel, Switzerland). Cell lysates containing 1 mg of total protein in lysis buffer were incubated with the appropriate primary antibodies overnight at 4 ℃. Antibody-bound proteins were precipitated with Protein A/G PLUS-Agarose (Santa Cruz Biotechnology) overnight at 4 °C. The beads were washed three times with cold PBS and eluted in 1 × SDS sample buffer by boiling for 10 min. Immunoprecipitates were resolved by SDS-PAGE and analyzed by western blotting.
In vivo tumor growth assay
Eight-week-old female NSG mice (Orient Bio, Seoul, Korea) were used in this study. For the in vivo tumor growth assay, 2 × 106 MDA-MB-231 control or NID1-overexpressing cells were suspended in 100 μl of a 1:1 mixture of PBS and Matrigel and 1.5 × 106 MDA-MB-231 LM1 control/NID1 overexpressing cells were suspended in 150 μl PBS and injected through the 4th mammary fat pad of each mouse. After injection, the tumor volume was measured using calipers every 3 or 4 days until sacrifice. The tumor volume was calculated using the following formula: (length × width × width)/2. Mice were euthanized using CO2 at the end point of the experiment.
Bioinformatics
NID1 expression data from diverse normal and tumor tissues were obtained and visualized using TIMER 2.0 (http://timer.cistrome.org/)16. NID1 mRNA expression data from breast cancer patients in the TCGA databases were obtained from cBioPortal (http://cbioportal.org/)17,18. For survival analyses of breast cancer patients, GSE1456 and integrated cohorts in the Kaplan–Meier plotter (http://kmplot.com/analysis/)19 were used. Gene set enrichment analysis (GSEA) was performed using gene expression data from METABRIC and gene sets from the Molecular Signature Database20,21.
Statistical analysis
All experiments were conducted using at least three independent trials. Data are presented as mean; error bars represent standard deviation (SD). Statistical significance between two groups was calculated using Student’s t-test. For comparisons of 3 or more groups, a one-way ANOVA (post hoc Tukey’s test) was used to calculate statistical significance. The Kaplan–Meier survival analysis was comparatively analyzed using the log-rank test. For all experiments, a P value of 0.05 or less was considered statistically significant. GraphPad Prism software 8.0 was used for statistical analysis.
Ethical approval
Approval Statement: The experiments employed in this study have been approved by the institutional animal care and use committee (IACUC) in Hanyang University (HY-IACUC-24-0002). IACUC carefully reviewed the study protocol, making sure that the protocol meets ethical standards and safeguards the rights of the animals.
ARRIVE: The experiments in this manuscript were conducted following the ARRIVE guidelines (http://arriveguidelines.org).
Accordance Statement: The research method and data analysis have been designed to ensure compliance with scientific rigor and integrity. The authors affirm that the content of this manuscript is an accurate representation of the research conducted.
Results
NID1 expression is negatively correlates with breast cancer progression.
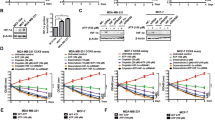
To identify the clinical significance of NID1 in breast cancers, we compared NID1 expression data for multiple cancer and normal tissue samples from public TCGA datasets. Those data showed that NID1 expression was significantly higher in tumors of GBM, HNSC, KIRC, STAD, THCA compared to their normal tissues. In tumors of BLCA, BRCA, CESC, KIRP, PRAD and UCEC, expression of NID1 was significantly lower than in normal tissues. (Fig. 1A). In the detailed analysis of breast cancer patient datasets, the expression of NID1 was significantly lower in TNBC than in the other subtypes (Fig. 1B). The survival analysis results also showed that NID1 expression correlated negatively with relapse-free survival (RFS) and overall survival (OS) of breast cancer patients (Fig. 1C). Based on those results, we hypothesized that NID1 plays a tumor-suppressive role in breast cancer by inhibiting oncogenic features. To investigate the role of NID1 in breast cancer, we generated stable NID1-overexpressing cell lines from the TNBC cell lines MDA-MB-231 and Hs578T. NID1 overexpression in total lysates and culture media was confirmed by Western blot analysis (Fig. 1D). NID1 overexpression significantly reduced the proliferation and colony-forming ability of the TNBC cell lines (Fig. 1E and F). In addition, in vivo tumor growth assays using immunodeficient mice showed that tumor formation by MDA-MB-231 NID1-overexpressing cells was significantly delayed compared with that by the control cell line (Fig. 1G–I). Given that Matrigel is known to contain NID122, we established the MDA-MB-231 LM1 NID1 overexpressing cell line to investigate the effect of NID1 on tumor growth without the use of Matrigel (Fig. S1A). MDA-MB-231 LM1 cells are derived from lung lesions that have undergone metastasis and can form tumors in NSG mice in the absence of Matrigel23. The data in Fig. S1 corroborates the data in Fig. 1I, suggesting that NID1 in Matrigel may not mask the difference in growth rate between NID1 overexpressing cells and control cells. Collectively, these results suggest that NID1 is a negative regulator of breast cancer progression.
NID1 negatively correlated with the progression of breast cancer. (A) Comparison of NID1 expression in various normal and tumor tissues. ACC: Adrenocortical Carcinoma, BLCA: Bladder Urothelial Carcinoma, BRCA: Breast Invasive Carcinoma, CESC: Cervical and Endocervical Cancer, CHOL: Cholangiocarcinoma, COAD: Colon Adenocarcinoma, ESCA: Esophageal Carcinoma, GBM: Glioblastoma Multiforme, HNSC: Head and Neck Cancer, KICH: Kidney Chromophobe, KIRC: Kidney Renal Clear Cell Carcinoma, KIRP: Kidney Renal Papillary Cell Carcinoma, LAML: Acute Myeloid Leukemia, LGG: Lower Grade Glioma, LIHC: Liver Hepatocellular Carcinoma, LUAD: Lung Adenocarcinoma, LUSC: Lung Squamous Cell Carcinoma, MESO: Mesothelioma, OV: Ovarian Serous Cystadenocarcinoma, PAAD: Pancreatic Adenocarcinoma, PCPG: Pheochromocytoma and Paraganglioma, PRAD: Prostate Adenocarcinoma, READ: Rectum Adenocarcinoma, SARC: Sarcoma, SKCM: Skin Cutaneous Melanoma, STAD: Stomach Adenocarcinoma, TGCT: Testicular Germ Cell Tumors, THCA: Thyroid Carcinoma, THYM: Thymoma, UCEC: Uterine Corpus Endometrial Carcinoma, UCS: Uterine Carcinosarcoma, UVM: Uveal Melanoma. (B) NID1 expression in data from breast cancer databases. Expression data were obtained from TCGA. (C) Kaplan–Meier survival analysis of breast cancer patients based on NID1 expression levels (GSE1456). (D) Establishment of NID1-overexpressing cell lines using MDA-MB-231 and Hs578T cells. Western blot analysis of total cell lysates with the indicated antibodies; GAPDH was used as the loading control. (E) Cell proliferation assays were performed using the NID1-overexpressing cell lines. (F) Colony formation assays were performed using the NID1-overexpressing cell lines. (G) Representative images showing xenograft tumors of MDA-MB-231 control and NID1-overexpressing cells at 7 weeks post-injection. (H) In vivo tumor growth curve from a xenograft mouse. (I) The tumor weights of each xenograft mouse were quantified 7 weeks post-injection. The data in (F, G, I), and J are presented as mean ± SD. P values in A were calculated using the Wilcoxon test; in (B and C), they were calculated using one-way ANOVA with a post-hoc Dunnett multiple comparison test; in (D), they were calculated using the log-rank test; in (F, G, I, and J), they were calculated using Student’s t test (*P < 0.05; **P < 0.005; ***P < 0.0005).
NID1 suppresses cell migration, invasion, and the EMT in TNBC cell lines
Because TNBC has greater metastatic potential than other breast cancer molecular subtypes24, we investigated whether NID1 affects cell migration ability. In the wound healing assay and transwell migration assay, we found that NID1 overexpression significantly reduced the migration ability of TNBC cells (Fig. 2A, B). Consistent with those results, invasion ability was significantly reduced in the invasion assay using Collagen type 1-coated transwells (Fig. 2C). Also, the protein and mRNA levels of EMT markers were downregulated in the NID1-overexpressing cell lines (Fig. 2D–F). Because NID1 acts after secretion from cells, we tested conditioned medium from the MDA-MB-231 NID1-overexpressing cell line (NID1 CM) and found that it also reduced the expression of the mesenchymal marker vimentin and cell migration ability (Fig. S2). These results indicate that NID1 reduces cell migration, invasion, and EMT marker expression in TNBC cell lines.
NID1 downregulated cell migration and EMT marker expression. (A) Wound-healing assay of the MDA-MB-231 and Hs578T NID1-overexpressing cell lines. The healed rate of each wound area was measured at 24 h intervals. (B–C) (B) Transwell migration and (C) invasion assays were performed with the MDA-MB-231 and Hs578T NID1-overexpressing cell lines. (D) Immunofluorescence analysis showed decreased expression of the mesenchymal marker vimentin (scale bar = 100 μm). (E) Western blot analysis of total cell lysate with the indicated antibodies showed decreased expression of EMT marker proteins. (F) RT-qPCR analysis showing the mRNA expression of EMT markers in the indicated cell lines. The values were normalized to the GAPDH level. The data in (A, B, C, and F) are presented as mean ± SD. P values in (A, B, C, and F) were calculated using Student’s t test (* P < 0.05; ** P < 0.005; *** P < 0.0005).
NID1 overexpression reduces glycolytic phenotypes
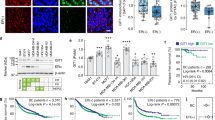
Metabolic reprogramming is frequently observed in TNBC cells because it helps to maintain malignant phenotypes such as high proliferation rates and metastatic ability25,26. Because our results showed that NID1 overexpression downregulated the proliferation and migration ability of TNBC cell lines, we hypothesized that metabolism reprogramming might play a role in the NID1-mediated suppression of oncogenic features. To test that hypothesis, we performed a GSEA using gene expression data from triple-negative breast cancer patients. Interestingly, NID1 expression correlated negatively with glycolysis-related gene signatures (Fig. 3A). Next, we examined cell viability assays using glucose-free medium to determine whether NID1 affects the glycolysis dependency of breast cancer cells. Those results showed that NID1-overexpressing cells showed higher viability following glucose deprivation than did the controls (Fig. 3B). Furthermore, glucose uptake, lactate production, and ATP production, all the major hallmarks of the glycolytic phenotype, decreased in NID1-overexpressing cells (Fig. 3C–E). We examined NID1-induced changes in mitochondrial respiratory capacity and found that the basal and maximal respiratory capacities were increased by NID1 overexpression compared with control cells. This result demonstrates that cells with NID1 overexpression are less dependent on glycolysis and have enhanced oxidative phosphorylation capacity compared with the controls (Fig. 3F). Western blot analysis showed that Glut3, LDHA, and PKM2, the proteins that act to shift the metabolism of the cells to glycolysis, were decreased, and LDHB, an isoenzyme that acts in opposition to LDHA, was increased in NID1-overexpressing cells (Fig. 3G). Also, native in-gel assays showed that NID1 overexpression shifted the LDH isoenzyme balance toward the LDHB-rich isoenzyme (LDH2), whereas the control cells shifted toward the LDHA-rich isoenzymes (LDH5 and LDH4) (Fig. 3H). These results suggest that metabolic reprogramming might be responsible for the suppression of oncogenicity by NID1.
NID1 suppressed aerobic glycolysis in breast cancer cell lines. (A) A GSEA was performed using mRNA expression data from TNBC patients in METABRIC. NES; normalized enrichment score. (B) Cell viability assays were performed with the indicated cell lines after 24 h of glucose depletion. (C) Glucose uptake assays were performed in the NID1-overexpressing cell lines. (D) Lactate production assays were performed in the NID1-overexpressing cell lines. (E) ATP assays were performed in the NID1-overexpressing cell lines. (F) The oxygen consumption rate (OCR) of MDA-MB-231 NID1-overexpressing cell was measured using a Seahorse XF Analyzer. OM oligomycin; R/A rotenone/antimycin A. (G) Western blot analysis of total cell lysates with the indicated antibodies; GAPDH was used as the loading control. (H) Native in-gel LDH isoenzyme assays were performed with the indicated cell lines. The representative values are the normalized value of each subunit to the value of the control cell line. Data in (B, C, D, E and F) are presented as mean ± SD. P values in (B, C, D, and E) were calculated using Student’s t test (*P < 0.05; **P < 0.005; ***P < 0.0005).
Overexpression of NID1 decreases HIF-1α and its transcriptional activity
Next, we investigated the molecular mechanism underlying the NID1-induced metabolic shift. Previous studies demonstrated that HIF-1α, the alpha subunit of the transcription factor HIF-1, is a master regulator of aerobic glycolysis in diverse cancers27. Therefore, we measured the expression of HIF-1α in the NID1-overexpressing cell lines. Western blots showed that the HIF-1α protein level was lower in the NID1-overexpressing cell lines than in the control cells (Fig. 4A). We also found that the mRNA levels of HIF-1α were significantly reduced in the cells with NID1 overexpression (Fig. 4B). The dual luciferase assay using the hypoxia response element (HRE)-luc construct confirmed that the transcriptional activity of HIF-1 was reduced in the NID1-overexpressing cell lines (Fig. 4C). Consistent with those results, the mRNA levels of genes known to be targets of HIF-1α were significantly changed in NID1-overexpressing cells (Fig. 4D) The genes previously shown to be positively regulated by HIF-1α, LDHA, PKM2, HK2, VEGFA28,29 were downregulated by NID1-overexpression while LDHB, a gene downregulated by HIF-1α28 was upregulated by NID1. Finally, we found that exogenous treatment with recombinant human NID1 (rhNID1) also downregulated HIF-1 activity in the TNBC cell lines (Fig. 4E). These results show that the reduction in glycolytic phenotypes caused by NID1 was due to a reduction in HIF-1α, a known master regulator of glycolysis.
NID1 downregulated HIF-1α and its target gene expression. (A) Western blot analysis of total cell lysates with the indicated antibodies; GAPDH was used as the loading control. (B) RT-qPCR analysis showing the mRNA expression of HIF1A in the indicated cell lines. Values were normalized to GAPDH. (C) Dual luciferase assays were performed using HRE (hypoxia response element)-luc constructs. pRL-CMV was used as the internal control. (D) RT-qPCR analysis showing the mRNA expression of HIF-1α target genes in the indicated cell lines. Values were normalized to GAPDH. (E) Dual luciferase assays were performed using HRE-luc constructs. pRL-CMV was used as the internal control. Each assay was performed after 24 h of treatment with rhNID1 (100 ng/ml. Data in (B, C, D, and E) are presented as mean ± SD. P values in (B, C, D, and E) were calculated using Student’s t test (*P < 0.05; **P < 0.005; ***P < 0.0005).
NID1 decreases HIF-1α expression via the integrin β1/FAK/Src/NF-κB signaling axis
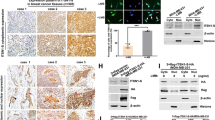
Next, we sought the molecular pathway by which NID1 induces HIF-1α downregulation. Because NID1 is an ECM protein, we focused on receptors that interact with it. A previous report demonstrated that NID1 physically interacts with integrin β130. Therefore, we assessed the expression of integrin β1 to see if it changed in the NID1-overexpressing cell lines. Interestingly, the total protein level of integrin β1 was significantly decreased in NID1 overexpressing cells, but the mRNA level was not changed (Fig. 5A, B). Based on those results, we speculated that downregulation of integrin β1 by NID1 did not occur at the transcriptional level. We performed a CHX chasing assay to determine whether the stability of integrin β1 was changed by NID1. Those results showed that integrin β1 degradation after CHX administration was significantly faster in the NID1-overexpressing cell lines than in the control cells (Fig. 5C, D). Moreover, a ubiquitination assay showed increased polyubiquitination of integrin β1 in cells overexpressing NID1, and the decrease in integrin β1 level could be restored by treatment with the 26S proteasome inhibitor MG132 (Fig. 5E). Next, we examined the activation of focal adhesion kinase (FAK)/Src/NF-κB p65 signaling, a well-known downstream signaling axis of integrin β1, in the NID1-overexpressing cell lines. We found that phosphorylation of FAK, Src, and NF-κB p65 was reduced in these cells (Fig. 5F). Furthermore, NF-κB reporter assays showed that NF-κB transcription activity was also reduced in NID1-overexpressing cells (Fig. 5G). Notably, NF-κB p65 is a transcription factor that regulates the expression of HIF-1α in cancer31. Therefore, the reduced activation of NF-κB p65 via integrin β1 might be involved in the reduction of HIF-1α caused by NID1. We also found that activation of the FAK/Src signaling axis was reduced after treatment with NID1 CM or recombinant human NID1 (rhNID1) in TNBC cells (Fig. S3A, B). In the survival analysis of breast cancer patients, NID1 significantly increased the RFS of patients with high integrin β1 expression, but it had no significant effect on the survival of patients with low integrin β1 expression (Fig. 5H), further suggesting the causative relationship between NID1 and integrin β1. However, it should be noted that the dataset in Fig. 5H is generated from multiple subtypes of breast cancer, not by TNBC alone. Taken together, our results suggest that NID1 reduces the progression of breast cancer by inhibiting the integrin β1-mediated FAK/Src/NF-κB/HIF-1α signaling axis (Fig. 5I).
NID1 suppressed the FAK/Src/NF-κB p65 signaling axis via integrin β1 downregulation. (A) Western blot analysis of total cell lysates with the indicated antibodies. β-actin was used as the loading control. (B) RT-qPCR analysis showing the mRNA expression of ITGB1 in the indicated cell lines. Values were normalized to GAPDH. (C–D) Cycloheximide (CHX) chase assays were performed to measure the degradation of integrin β1 in the (C) MDA-MB-231 and (D) Hs578T NID1-overexpressing cell lines. Cells were treated with CHX (40 μg/ml) and harvested at the indicated times. (E) To detect ubiquitinated integrin β1, an immunoprecipitation analysis was performed with anti-integrin β1 antibody after a 24-h incubation with MG132 (10 μM). (F) Western blot analysis of total cell lysates with the indicated antibodies; GAPDH was used as the loading control. (G) Dual luciferase assays were performed using NF-κB-luc constructs. pRL-CMV was used as the internal control. (H) Kaplan–Meier survival analysis of breast cancer patients based on NID1 expression levels. (I) A schematic description of the NID1-mediated signaling pathway in breast cancer cells. Data in (B, C, D, and G) are presented as mean ± SD. P values in (B, C, D, and G) were calculated using Student’s t test; in H, they were calculated with the log-rank test (*P < 0.05; **P < 0.005; ***P < 0.0005).
Discussion
In the tumor microenvironment, ECM proteins play a crucial role in cancer progression by regulating cell growth, migration, metabolism, and immune functions32,33,34. Therefore, understanding ECM components is important for investigating interactions between the tumor microenvironment and cancer cells and ultimately for treating cancer.
In this study, we elucidated the tumor suppressive role of NID1, an ECM component, in breast cancer. We found that NID1 expression is significantly downregulated in aggressive breast cancers, and that NID1 overexpression inhibits cell proliferation, migration, and invasion in TNBC cell lines. In addition, NID1 suppresses the glycolytic phenotype that induces tumor growth and metastasis. A study has demonstrated that NID1 plays a regulatory role in the proliferation and migration/invasion of murine claudin-low mammary tumors35. In that study, the investigators observed that the knockdown of NID1 resulted in a decrease in the proliferation, migration, and invasion of murine claudin-low mammary tumor cells in vitro. This finding is contrary to our own observations. While the roles of many genes are known to be the same in mice and humans, there are reports indicating that experimental results in mice may not be reproduced in humans due to species differences36,37,38,39 Indeed, human NID1 and murine NID1 proteins share less than 85% amino acid sequence homology 40,41. Our research has demonstrated that the overexpression of NID1 reduces the proliferation, colony formation, migration, and invasion of TNBC cells in vitro. Additionally, we have shown that the overexpression of NID1 reduces tumor growth in vivo. We further found that these tumor suppressive effects are due to the downregulation of HIF-1α transcription that occurs through inhibition of the integrin β1/FAK/Src/NF-κB p65 signaling axis.
Breast cancer is the cancer most frequently diagnosed in women worldwide and the second leading cause of cancer deaths in women42,43. Among the molecular subtypes of breast cancer, TNBC has low expression of the estrogen receptor, progesterone receptor, and human epidermal growth factor receptor 2, which are the main therapeutic targets of breast cancer treatments44. Therefore, TNBC has poorer survival and higher recurrence rates than other breast cancer subtypes. TNBC also has aggressive characteristics, such as a high growth rate and high metastatic potential, that result in high energy demand45,46,47. The conversion of glucose to lactic acid in the presence of oxygen is called aerobic glycolysis, and it is a well-known metabolic property of cancer cells48. Recent studies have reported that 2-deoxyglucose and metformin, which inhibit glycolysis-dependent metabolism, increase the sensitivity of TNBC to chemotherapy49,50. In this study, we found that NID1 overexpression decreased glucose uptake, lactate production, and ATP production, which are all associated with the glycolytic phenotype. We also found that HIF-1α, the master regulator of glycolysis, was reduced at the transcriptional level in cells overexpressing NID1, and its downstream target genes, GLUT3, LDHA, PKM2, and HK2, were reduced accordingly. Therefore, the ability of NID1 to downregulate these enzymes could provide a new direction for cancer treatment.
HIF-1α is a subunit of HIF and acts as a transcription factor. The level of HIF-1α is increased not only in breast cancer, but also in a broad range of human cancer types, and it is associated with a poor prognosis in many cancers51. As a transcription factor, HIF-1α is responsible for the transcription of more than 100 target genes52. In addition to regulating the transcription of glycolytic enzymes, it is also known to affect tumor progression by regulating the transcription of genes related to cell proliferation and migration53. Therefore, the decrease we found in the transcript levels of EMT markers (ZEB1, VIM, SNAI1, SNAI2, and TWIST) in the NID1-overexpressing cells could be a direct result of the downregulation of HIF-1α. These results suggest that the direct reduction in HIF-1α signaling caused by NID1, as well as the reduction in the glycolytic phenotype, might affect the migratory ability of breast cancer cells.
Integrin β1 is a member of the integrin receptor family, which contains transmembrane glycoproteins that interact with the ECM. The integrin family has 24 αβ members, and they function as heterodimers54. Integrins also have tumorigenic functions, such as inducing metastasis, therapy resistance, stem cell plasticity, and energy metabolism reprogramming55,56. Integrin β1 has been previously reported to directly interact with NID130; in our results, the stability of integrin β1 was decreased in NID1-overexpressing cell lines via increased polyubiquitination. As the stability of integrin β1 decreased, we found that the phosphorylation of FAK/Src, well-known downstream signaling molecules of integrins57, also decreased. In a previous study, activation of the FAK/Src signaling pathway induced the phosphorylation of NF-κB p65 to increase its activity and regulate the transcription of genes, including HIF-1α31,58,59. Consistent with those reports, our results show that the phosphorylation of NF-κB p65 was reduced in NID1-overexpressing cells, as was the transcriptional activity of NF-κB. Based on those experimental results, we postulated that NID1 might regulate the glycolytic phenotypes of TNBC cells through the integrin β1/FAK/Src/NF-κB p65-mediated downregulation of HIF-1α. This reduced glycolytic phenotype would decrease the amount of energy in the cells and thus reduce their proliferation rate and metastatic potential. However, it remains unclear how the interaction between NID1 and integrin β1 leads to integrin β1 ubiquitination.
In conclusion, the results of this study demonstrate that NID1 expression reduces the HIF-1α expression level and modulates HIF-1α-mediated cellular events, culminating in the suppression of aerobic glycolysis, a hallmark of cancer. Furthermore, the suppression of aerobic glycolysis decreases the proliferation rate and metastasis potential, both of which are associated with a poor prognosis in breast cancer. Taken together, our results suggest a novel function of NID1 as a tumor suppressor and suggest its potential for use in the treatment of TNBC by targeting tumor metabolism.
Data availability
All data generated or analysed during this study are included in this published article
References
Pickup, M. W., Mouw, J. K. & Weaver, V. M. The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. 15(12), 1243–1253 (2014).
Ho, M. S. et al. Nidogens-extracellular matrix linker molecules. Microsc. Res. Tech. 71(5), 387–395 (2008).
Pujuguet, P. et al. Nidogen-1 regulates laminin-1-dependent mammary-specific gene expression. J. Cell Sci. 113(Pt 5), 849–858 (2000).
Zhou, Y. et al. NID1, a new regulator of EMT required for metastasis and chemoresistance of ovarian cancer cells. Oncotarget 8(20), 33110–33121 (2017).
Mao, X. et al. Nidogen 1-enriched extracellular vesicles facilitate extrahepatic metastasis of liver cancer by activating pulmonary fibroblasts to secrete tumor necrosis factor receptor 1. Adv. Sci. (Weinh) 7(21), 2002157 (2020).
Rokavec, M., Bouznad, N. & Hermeking, H. Paracrine induction of epithelial-mesenchymal transition between colorectal cancer cells and its suppression by a p53/miR-192/215/NID1 axis. Cell. Mol. Gastroenterol. Hepatol. 7(4), 783–802 (2019).
Aleckovic, M. et al. Identification of Nidogen 1 as a lung metastasis protein through secretome analysis. Genes Dev. 31(14), 1439–1455 (2017).
Ferraro, D. A. et al. Endothelial cell-derived nidogen-1 inhibits migration of SK-BR-3 breast cancer cells. BMC Cancer 19(1), 312 (2019).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science 324(5930), 1029–1033 (2009).
Feng, J. et al. Emerging roles and the regulation of aerobic glycolysis in hepatocellular carcinoma. J. Exp. Clin. Cancer Res. 39(1), 126 (2020).
Liu, C., Jin, Y. & Fan, Z. The mechanism of warburg effect-induced chemoresistance in cancer. Front. Oncol. 11, 698023 (2021).
Feron, O. Pyruvate into lactate and back: From the Warburg effect to symbiotic energy fuel exchange in cancer cells. Radiother. Oncol. 92(3), 329–333 (2009).
Shin, E. & Koo, J. S. Glucose metabolism and glucose transporters in breast cancer. Front. Cell Dev. Biol. 9, 728759 (2021).
Abdel-Wahab, A. F., Mahmoud, W. & Al-Harizy, R. M. Targeting glucose metabolism to suppress cancer progression: Prospective of anti-glycolytic cancer therapy. Pharmacol. Res. 150, 104511 (2019).
Sullivan, W. J. et al. Extracellular matrix remodeling regulates glucose metabolism through TXNIP destabilization. Cell 175(1), 117-132 e21 (2018).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48(W1), W509–W514 (2020).
Cerami, E. et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data (vol 2, pg 401, 2012). Cancer Discov. 2(10), 960–960 (2012).
Gao, J.J., et al., Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the cBioPortal. Science Signaling, 2013. 6(269).
Gyorffy, B., Discovery and ranking of the most robust prognostic biomarkers in serous ovarian cancer. Geroscience, 2023.
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34(3), 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U.S.A. 102(43), 15545–15550 (2005).
Kaur, S. et al. Non-matrigel scaffolds for organoid cultures. Cancer Lett. 504, 58–66 (2021).
Cui, Y. H. et al. FBXL14 abolishes breast cancer progression by targeting CDCP1 for proteasomal degradation. Oncogene 37(43), 5794–5809 (2018).
Garrido-Castro, A. C., Lin, N. U. & Polyak, K. Insights into molecular classifications of triple-negative breast cancer: Improving patient selection for treatment. Cancer Discov. 9(2), 176–198 (2019).
Sun, X. et al. Metabolic reprogramming in triple-negative breast cancer. Front. Oncol. 10, 428 (2020).
Munkacsy, G., Santarpia, L. & Gyorffy, B. Therapeutic potential of tumor metabolic reprogramming in triple-negative breast cancer. Int. J. Mol. Sci. 24(8), 6945 (2023).
Courtnay, R. et al. Cancer metabolism and the Warburg effect: The role of HIF-1 and PI3K. Mol. Biol. Rep. 42(4), 841–851 (2015).
Nam, K., Oh, S. & Shin, I. Ablation of CD44 induces glycolysis-to-oxidative phosphorylation transition via modulation of the c-Src-Akt-LKB1-AMPKalpha pathway. Biochem. J. 473(19), 3013–3030 (2016).
Taylor, C. T. & Scholz, C. C. The effect of HIF on metabolism and immunity. Nat. Rev. Nephrol. 18(9), 573–587 (2022).
Dedhar, S. et al. The receptor for the basement-membrane glycoprotein entactin is the integrin alpha-3 beta-1. J. Biol. Chem. 267(26), 18908–18914 (1992).
van Uden, P., Kenneth, N. S. & Rocha, S. Regulation of hypoxia-inducible factor-1alpha by NF-kappaB. Biochem. J. 412(3), 477–484 (2008).
Davis, G. E. & Senger, D. R. Endothelial extracellular matrix: Biosynthesis, remodeling, and functions during vascular morphogenesis and neovessel stabilization. Circ. Res. 97(11), 1093–1107 (2005).
Wang, M. et al. Role of tumor microenvironment in tumorigenesis. J. Cancer 8(5), 761–773 (2017).
Lu, P., Weaver, V. M. & Werb, Z. The extracellular matrix: A dynamic niche in cancer progression. J. Cell Biol. 196(4), 395–406 (2012).
Jagroop, R., Martin, C. J. & Moorehead, R. A. Nidogen 1 regulates proliferation and migration/invasion in murine claudin-low mammary tumor cells. Oncol. Lett. 21(1), 52 (2021).
Gould, S. E., Junttila, M. R. & de Sauvage, F. J. Translational value of mouse models in oncology drug development. Nat. Med. 21(5), 431–439 (2015).
Stergachis, A. B. et al. Conservation of trans-acting circuitry during mammalian regulatory evolution. Nature 515(7527), 365–370 (2014).
Cheng, Y. et al. Principles of regulatory information conservation between mouse and human. Nature 515(7527), 371–375 (2014).
Young, R. S. et al. The frequent evolutionary birth and death of functional promoters in mouse and human. Genome Res. 25(10), 1546–1557 (2015).
“NM_002508.3 Sequence Report—Fasta Format”. Available from: https://www.ncbi.nlm.nih.gov/nuccore/NM_002508.3?from=12&to=3755&report=fasta (2024).
“NM_010917.3 Sequence Report—Fasta Format”. [15 Nov 2024]. Available from: https://www.ncbi.nlm.nih.gov/nuccore/NM_010917.3?from=36&to=3773&report=fasta (2024).
Trayes, K. P. & Cokenakes, S. E. H. Breast cancer treatment. Am. Fam. Phys. 104(2), 171–178 (2021).
Fahad Ullah, M. Breast cancer: Current perspectives on the disease status. Adv. Exp. Med. Biol. 1152, 51–64 (2019).
Yin, L. et al. Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res. 22(1), 61 (2020).
Anders, C. K. & Carey, L. A. Biology, metastatic patterns, and treatment of patients with triple-negative breast cancer. Clin. Breast Cancer 9(Suppl 2), S73-81 (2009).
Petrocca, F. et al. A genome-wide siRNA screen identifies proteasome addiction as a vulnerability of basal-like triple-negative breast cancer cells. Cancer Cell 24(2), 182–196 (2013).
Shen, L. L. et al. Metabolic reprogramming in triple-negative breast cancer through Myc suppression of TXNIP. Proc. Natl. Acad. Sci. U.S.A. 112(17), 5425–5430 (2015).
Warburg, O. On respiratory impairment in cancer cells. Science 124(3215), 269–270 (1956).
Varghese, E. et al. Targeting glucose metabolism to overcome resistance to anticancer chemotherapy in breast cancer. Cancers (Basel) 12(8), 2252 (2020).
Grundker, C. et al. Inhibition of aerobic glycolysis enhances the anti-tumor efficacy of zoptarelin doxorubicin in triple-negative breast cancer cells. J. Obstet. Gynaecol. Res. 45(7), 1334–1342 (2019).
Keith, B., Johnson, R. S. & Simon, M. C. HIF1 alpha and HIF2 alpha: Sibling rivalry in hypoxic tumour growth and progression. Nat. Rev. Cancer 12(1), 9–22 (2012).
Masoud, G. N. & Li, W. HIF-1alpha pathway: Role, regulation and intervention for cancer therapy. Acta Pharm. Sin. B 5(5), 378–389 (2015).
Tiburcio, P. D., Choi, H. & Huang, L. E. Complex role of HIF in cancer: the known, the unknown, and the unexpected. Hypoxia (Auckl) 2, 59–70 (2014).
Yousefi, H. et al. Understanding the role of integrins in breast cancer invasion, metastasis, angiogenesis, and drug resistance. Oncogene 40(6), 1043–1063 (2021).
Cooper, J. & Giancotti, F. G. Integrin signaling in cancer: Mechanotransduction, stemness, epithelial plasticity, and therapeutic resistance. Cancer Cell 35(3), 347–367 (2019).
Seguin, L. et al. Integrins and cancer: Regulators of cancer stemness, metastasis, and drug resistance. Trends Cell Biol. 25(4), 234–240 (2015).
Guo, W. & Giancotti, F. G. Integrin signalling during tumour progression. Nat. Rev. Mol. Cell Biol. 5(10), 816–826 (2004).
Lee, H. S. et al. Src tyrosine kinases mediate activations of NF-kappaB and integrin signal during lipopolysaccharide-induced acute lung injury. J. Immunol. 179(10), 7001–7011 (2007).
Lluis, J. M. et al. Dual role of mitochondrial reactive oxygen species in hypoxia signaling: Activation of nuclear factor-kappaB via c-SRC and oxidant-dependent cell death. Cancer Res. 67(15), 7368–7377 (2007).
Acknowledgements
This work was supported by National Research Foundation of Korea (NRF) grants funded by the Korean government (MIST): 2022R1A2C1003218, RS-2023-00243643 and Korea Basic Science Institute (National research Facilities and Equipment Center) grant funded by the Ministry of Education.(2023R1A6C101A009).
Author information
Authors and Affiliations
Contributions
Conceptualization: J.H.L., S.S., Y.K., I.S.; Methodology: J.H.L., S.S., Y.K., H.L.; Validation: J.H.L., S.S., Y.K., H.L., M.L., M.G.K, H.K.; Formal analysis: J.H.L., S.S., Y.K., H.L.; Investigation: J.H.L., S.S., Y.K., H.L.; Resources: K.M.L., I.S.; Writing—original draft: J.H.L., S.S.; Writing—review & editing: J.H.L., S.S., H.L., K.M.L, I.S.; Visualization: J.H.L., S.S., Y.K., H.L.; Supervision: I.S.; Project administration: I.S.; Funding acquisition: I.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, Jh., Son, S., Ko, Y. et al. Nidogen-1 suppresses cell proliferation, migration, and glycolysis via integrin β1-mediated HIF-1α downregulation in triple-negative breast cancer. Sci Rep 15, 10633 (2025). https://doi.org/10.1038/s41598-024-84880-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-84880-5