Abstract
Gastrointestinal nematode (GIN) infections, particularly caused by Haemonchus contortus, are a major concern in sheep farming, resulting in significant economic losses Genetic resistance, used alongside chemical control, nutrition, and pasture management, provides a sustainable strategy to mitigate these infections in industrial livestock systems The South African Dohne Merino sheep, known for their resilience and suitability for both wool and meat production, offer a potential genetic resource for breeding GIN resistance, providing an alternative to chemical control. This study utilised RNA-Seq and differential gene expression (DEG) analysis to investigate the molecular mechanisms underlying H. contortus infection in Dohne Merino sheep. Six adult ewes from Wauldby farm (Eastern Cape Province), naturally exposed to H. contortus, were selected. The animals were categorised into resistant (n = 3) and susceptible (n = 3) groups based on Estimated Breeding Values (EBVs) from faecal egg count (FEC) phenotypes. DEG analysis revealed 34 significantly differentially expressed genes (DEGs) related to immune responses and external stimuli, with involvement in key pathways such as Rap1 and PI3K-Akt signaling, linked to H. contortus resistance. Additionally, segment-specific analysis of the gastrointestinal tract identified DEGs in different regions: 146 in the abomasum, 302 in the ileum, 584 in the jejunum, and 332 in the duodenum. The findings highlight genes and pathways contributing to GIN resistance and tissue-specific response mechanisms. These insights can support the selection and breeding of sheep with enhanced resistance to H. contortus, offering a genetic approach to combating GIN infections.
Similar content being viewed by others
Introduction
In South Africa, sheep production plays a crucial role in meeting the economic and socio-economic needs of rural communities by providing income and employment, as well as manure for crop farming, and contributing to agricultural diversification and sustainable farming practices1. Gastrointestinal nematode (GIN) parasitic infections affect a majority of livestock species2 3, causing significant economic losses in developing and developed countries2 3. Parasitic nematodes infecting sheep impose severe constraints on health, welfare and productivity, causing substantial economic losses2,4. These parasites reduce animal productivity through direct feeding or damaging the absorptive lining of the gastrointestinal tract, resulting in inefficient digestion and nutrient absorption by the host5, which affects the health of animals, including weight loss, slow growth, lesser wool, meat and milk production; treatment expenses and death of animals6.
Haemonchus contortus, often known as the barber pole worm, is regarded as the most significant GIN parasite in tropical and subtropical regions, leading to substantial profit losses in sheep production. Haemonchus contortus is a blood-feeding nematode that draws blood from the capillaries in the stomach mucosa and is of particular concern due to its pathogenicity7,8 and lives and moves freely in the abomasum9. Animals, especially lambs or kids and lactating ewes, are found dead because of hyperacute haemonchosis, and those that survived show signs of severe anaemia10, falling wool or fleece and dark, firm and scant-coloured faces due to occult blood content11. Animals infected by H. contortus are also characterised by poor carcass quality, which results in economic loss to farmers and the livestock industry12,13.
Animals that are infested with haemonchosis are characterised by changes in the mucous membrane of the eyes, from normal red-pink colour to white (pale) colour10, weight loss and unenergetic or weaker due to blood loss11,14. This infestation causes animals to have low appetites, and this triggers the deficiency of plasma or blood protein due to an increased demand for protein and a decrease in the nutrient supply14. These clinical signs depend on the quantity and burden of parasites present in an animal. Clinical signs such as weight loss due to diarrhea, slow growth, lesser meat, wool and milk production and death of animals6 are mostly used by farmers as indicator traits. These clinical signs had been diagnosed using traditional methods such as faecal egg count (FEC), body condition scores (BCS) and FAMACHA scores4. FEC generally requires laboratory analysis, whereas BCS and FAMACHA can be applied by farmers with basic training, eliminating the need for veterinary experts. However, these diagnostic methods are time-consuming and may vary in accuracy depending on the assessor’s experience15.
Anthelmintic drugs remain an important tool for chemical control against GIN, but their effectiveness can vary due to an increase in anthelmintic resistance16. In addition, the use of anthelmintic drugs harms the development of natural immunity in animals17. The adverse effects of anthelmintic treatments for GINs highlight the need to start effective and sustainable control strategies, including enhancing genetically resistant livestock populations18. Immunological capacity with lower nematodes burden and concentration of nematodes eggs in faeces is associated with resistance19. The defence mechanisms that influence tolerance include immune response mechanisms (e.g. antibody response), a rise in eosinophil levels in the blood, and pepsinogen, which is a substance released by the stomach lining that is transformed into the enzyme pepsin by gastric acid4. Both nonspecific and specific immune responses help repair tissue damage inflicted by parasites while tolerating the infection to persist without directly suppressing the parasite20,21. Additionally, mechanisms that differ between individuals and populations21 include developing the host’s ability to resist parasitic infections by triggering an immune response aimed at reducing parasite fertility22. These mechanisms are frequently shaped by environmental or ecological factors and immunological, physiological and behavioural characteristics that affect the host condition and parasite burden4,21.
This study focused on the abomasum, duodenum, jejunum, and ileum, which play key roles in nutrient digestion, absorption, and immune defense23. The abomasum initiates protein digestion, while the small intestine continues nutrient processing, regulates stomach emptying, and contributes to mucosal immunity by balancing tolerance to commensals with protection against pathogens24,25,26,26,27.
An understanding of the gene expression effects and functions of different genes related to disease/parasite resistance and susceptibility will contribute to valuable information for breeding resistant sheep18. RNA-Seq has been utilised to better our understanding of the dynamics of metabolic processes in cells and tissues and how changes in the transcriptome profiles affect health and diseases28. Genome and transcriptome profiling have been used in the analysis of livestock suffering from GIN infection, identifying differentially expressed genes and pathways linked to resistance or susceptibility to GIN3,29.
RNA-Seq was utilised to profile the differentially expressed genes in Dohne Merino sheep naturally infected with H. contortus, aiming to identify genes and pathways involved in determining resistance or susceptibility to the parasite. For this purpose, Dohne Merino sheep that differ in resistance to H. contortus were selected from Wauldby Farm, in the Eastern Cape Province. This study specifically focused on four gastrointestinal tract segments, abomasum, duodenum, jejunum and ileum, that contribute to the breakdown and absorption of nutrients24.
Materials and methods
Ethics statement
Ethical approval
The animal procedures described in this study were conducted under the approval of the Animal Research Ethics Committee of the College of Agriculture and Environmental Sciences, University of South Africa (UNISA), with the reference number: 2019/CAES/037.
Compliance with guidelines and ARRIVE
All methods were carried out in accordance with the relevant guidelines and regulations. The reporting of animal experiments in this study complies with the ARRIVE guidelines (Animal Research: Reporting of In Vivo Experiments).
Dohne Merino sheep population selection and egg identification with worm counts analysis
Six adult ewes (aged 7–9 years) were categorised as H. contortus resistant (n = 3) or susceptible (n = 3) based on Estimated Breeding Values (EBVs) derived from faecal egg count (FEC) phenotypes. EBVs were calculated using the animal model with best linear unbiased prediction (BLUP), incorporating phenotypic faecal egg count records, pedigree information, and fixed effects such as sex, birth type, age, and contemporary group. Variance components were estimated by restricted maximum likelihood (REML) using the BLUPF90 software suite, and EBV accuracies were derived from the prediction error variance(30,31,32,33,34. Based on observed Egg identification (Fig. 1), actively searching for worms within each animal in the field, evidence of worm infestation was found in four animals, two susceptible and two resistant (Table 1), and only 2 resistant and 2 susceptible samples were used.
Haemonchus contortus eggs identified using McMaster floatation technique.
The sheep were sampled from the Wauldby farm in the Stutterheim district of the Eastern Cape Province, South Africa. The farm is located at GPS coordinates 27° 37’ East, 32° 35’ South. It is a region characterised by high summer rainfall (~ 800 mm annually), has mild winters and does not experience severe frost.
Faecal pellet samples from six ewes were analysed for parasite egg counts using the modified quantitative McMaster floatation technique at the Agricultural Research Council, Parasitology laboratory. Briefly, two grams of faeces were homogenised in 56 mL of 40% sucrose solution using a mechanical blender. A few drops of propylene were added to the mixture to enhance egg visibility. The mixture was aliquoted using a transfer pipette into two chambers of the McMaster slide (need to add info) and left on the bench for 5 min to ensure parasite eggs floated to the surface. Eggs within the grid area of both chambers were counted under the compound microscope at 10 × magnification. Finally, total eggs per gram (EPG) of faeces were calculated as:
Tissue samples collection
All six Dohne Merino sheep were humanely slaughtered at the Tandala Farm Abattoir, Sutterheim, Eastern Cape. Post-slaughtering, the intestines and abomasum were aseptically excised. Tissue Sects. (1.5—2.0 cm) were collected from each segment (i.e., abomasum, duodenum, ileum and jejunum) and placed into sterile collection tubes containing RNAlater® (Ambion, Austin, Texas, United States) solution to prevent RNA degradation. Samples were transported within 24 h to the Agricultural Research Council-Biotechnology Platform (ARC-BTP) and preserved at -800C until future analysis.
RNA isolation from abomasum and small intestine tissues
Tissue samples weighing 30 mg were taken from the duodenum, ileum, jejunum and abomasum and disrupted by homogenising using Geno/Grinder® 2010 (SPEX Sample Prep LLC, New Jersey, United States), and total RNA was isolated from each section using the Qiagen Mini Kit (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. The RNA samples were treated with RNase-free DNase I (Qiagen, Germany) to eliminate genomic DNA contamination, and the RNA was subsequently eluted with 50 μl of RNase-free water. The integrity and concentration of the RNA were evaluated employing Qubit Fluorometric Quantitation (Thermo Fisher Scientific, Waltham, MA, USA) and through absorbance measurements at 260 nm and determining the A260/A280 ratio using NanoDrop® ND-1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). Gel electrophoresis was performed to confirm the absence of RNA degradation, utilising one μl of RNA on a 1% TAE agarose gel. The eluted RNA samples were preserved at − 80 °C for subsequent analysis.
Library preparation and RNA sequencing using Illumina HiSeq 2500
The TrueSeq RNA Sample Preparation kit (Illumina, San Diego, CA) was utilised to create mRNA library preparations using 4 μg of total RNA for each sample, according to the manufacturer’s protocol. The sample library size, quantity and quality control analysis were done using The LabChip® GX Touch™ by PerkinElmer. Sequencing was performed on the Illumina HiSeq 2500 at the Agricultural Research Council-Biotechnology Platform in South Africa to produce 125 bp paired-end data.
Data quality control and bioinformatics analysis
The integrity of all generated raw reads was checked utilising FastQC (Babraham Institute, Cambridge, UK). All raw reads were trimmed to eliminate adapter contaminants and filtered for poor-quality reads shorter than 13 bp using Trimmomatic, a quality control tool. After the filtration by Trimmomatic, FastQC was employed once more to assess the quality of the paired-end reads, ensuring that only high-quality reads longer than 20 bp on both ends of the paired-end format were utilised for subsequent downstream analysis.
All clean paired-end reads from every sample were mapped to the reference sheep genome (Oar version 4.0) using HISAT235. Samtools 1.2.1 was utilised to sort and convert the SAM files into BAM files. Quality-controlled reads were then assembled into transcripts using StringTie2, and the transcript was merged using StringTie—merge. Furthermore, the gffcompare programme that is based on the CuffCompare (http://cole-trapnell-lab.github.io/cufflinks/cuffcompare) utility, which is part of the Cufflinks was employed to compare the transcript with the reference annotation. The abundance of the transcript was reported as fragmentation per kilobase of transcript per million mapped fragments (FPKM). StringTie was utilised to estimate the transcript abundances and generate count tables for Ballgown analysis (https://ccb.jhu.edu/software/stringtie/).
Differential gene expression analysis of Dohne Merino sheep naturally infected with Haemonchus contortus
All sequence data from 3 biological replicates from animal segments (the abomasum, jejunum, ileum and duodenum) were pooled and grouped as resistant and susceptible. To estimate DEGs, gene count data generated using PrepDE were used using Bioconductor package DESeq2 within R version 3.2.2 (Bioconductor, Buffalo, USA). For the Benjamini–Hochberg false discovery rate (FDR), multiple corrections were applied. Transcripts with an FDR < 0.05 and absolute log2fold change values > ± 1 were deemed differentially expressed genes.
Differential gene expression analysis of the abomasum, jejunum, ileum and duodenum
All sequence data from biological replicates of the abomasum, jejunum, ileum and duodenum were analysed separately. Gene count data generated by PrepDE were used to estimate DEGs, and they were analysed using DESeq2, a Bioconductor package within R version 3.2.2 (Bioconductor, Buffalo, USA). Multiple corrections were implemented for the false discovery rate (FDR). Transcripts with an FDR < 0.05 and overall log2 fold change values > ± 1 were identified as differentially expressed genes.
Functional annotation analysis
Bioinformatics resource tools were utilised for gene annotation and pathway analysis to gain insight into the functions of the differentially expressed genes (DEGs) in resistant and susceptible animals. The Database for Annotation, Visualization and Integrated Discovery (David) v6.8 online package was used for GO analysis of DEGs36(http://david.abcc.ncifcrf.gov/). To understand the high-level functions of genes, annotations for biological processes, cellular components, and molecular functions were categorised based on Gene Ontology (GO) using the WEGO v2.0 online tool (). GO analysis was used to analyse the roles of the DEGs and to describe the properties of genes in any organism. KEGG (Kyoto Encyclopaedia of Genes and Genomes) pathways are specifically employed to analyse extensive molecular datasets generated by genome sequencing and various high-throughput experimental techniques38 and to perform pathways enrichment analysis of DEGs. Therefore, the KEGG pathways database was utilised through the KEGG mapper within the Kyoto Encyclopaedia of Genes and Genomes databases (KEGG, http://www.genome.jp/kegg) to understand the specific function of the DEGs in certain biological processes39.
Results
Haemonchus contortus egg count and identified using McMaster floatation technique
Based on the six selected animals, three were classified as resistant and three as susceptible using phenotypic diagnostics based on estimated breeding values (EBVs). Egg identification (Fig. 1) revealed that only two animals with high EBVs had detectable eggs among the six sheep examined. Additionally, after actively searching for worms within each animal in the field, evidence of worm infestation was found in four animals, two susceptible and two resistant (Table 1).
Although one animal was initially classified as susceptible based on phenotypic evaluation, no worms or eggs were detected after slaughter. As a result, this animal was designated as a non-infected control for the susceptible group. Similarly, an animal classified as resistant but showing no signs of worm infestation was used as a non-infected control for the resistant group.
RNA sequencing and read mapping to the reference genome
Sheep infected with H. contortus
The average number of reads was 6,691,243, with each read having a length of 125 bp in a paired-end format from 24 tissue samples of six Dohne Merino sheep were generated using HiSeq 25,000. Moreover, for each segment, the average was 6 626 595,5 reads for abomasum, 6 883 121,7 reads for jejunum, 7 406 079,3 reads for ileum, and 5 849 176,5 reads for duodenum, respectively. The alignment rate of the reads for each segment was above 90% to the ovine reference genome (Ovis aries v4.0) (ftp:// ftp.ncbi.nlm.nih.gov/genomes/all/GCF/000/298/735/GCF_000298735.2_Oar_v4.0/) and the overall reads alignment was 94.44% (Table 2).
Differentially expressed genes
Differentially expressed genes in sheep infected with H. contortus
A total of 38 156 DEGs were identified, showing variation in expression between resistant and susceptible animals (Fig. 2A). Of 38 156 DEGs, only 34 genes differ significantly (FDR < 0.05 and log2fold change value above 1.0), among these, 14 genes were found to be upregulated (e.g., TRNAA-AGC, LOC105611628) and 20 genes downregulated (e.g., ART1, APLF, TEP1, LOC101113750) (Table 3; Fig. 2A).
Volcano plot showing genes with significant differences in the relative abundance of study animals. The dot colour denotes significantly differentially expressed (red dot) at FDR < 0.05, non-significant (black dot), down-regulated (< 0) and up-regulated (> 0) with (A) Animals infected with H, contortus (B) ileum, (C) abomasum, (D) jejunum and (E) duodenum.
Tissue-specific differentially expressed genes
A total of 146 genes were identified as differentially expressed (FDR < 0.05 and log2fold change value above 2) between the abomasum of resistant and susceptible animals (Fig. 2C). Of these, 58 were upregulated and 88 downregulated, and most were classified as unpredicted genes and were not included for further analysis (Table 4). Seven labelled genes were YES1, EPOR, RPSA, CHD9, PPP4R1, EVA1B and LOC114116061, and all these genes were downregulated.
In the ileum of resistant and susceptible animals, a total of 302 genes were differentially expressed (FDR < 0.05 and log2 fold change value greater than 2) (Fig. 2B), with 161 genes upregulated and 141 downregulated (Table 5). Six (LOC101122805, MGAT4A, DLG1, LOC10119675, NDUFA5 and LOC101123419) of the eight highly expressed DEGs were upregulated with two (COL4A3 and C2H2orf76) DGEs being downregulated.
Across the jejunum of resistant and susceptible animals, a total of 584 genes were identified as differentially expressed (FDR < 0.05 and log2 fold change value exceeding 2) (Fig. 2D). Of these, 303 genes were upregulated (including SLCO4C1, TTYH3, RFWD2, CA12, PAX4 and CXCL9 ) and 281 were downregulated (included SUGCT and LOC101109633) (Table 6).
Within the duodenum of resistant and susceptible animals, a total of 332 genes were differentially expressed (FDR < 0.05 and a log2 fold change value exceeding 2) (Fig. 2E). Among these, 206 genes were upregulated, whereas 126 were downregulated. For the ten most highly significant DEGs, only six were labelled, and four were unpredicted DEGs. Only four (GALNT1, SEPP1, SEMA6C and TNFRSF1B) DEGs were upregulated with two (TRNAC-ACA and COPB1) downregulated (Table 7; Fig. 2E).
Of all DEGs that were identified and predicted, only one (GAL3ST3) was common in all the segments of the small intestine, and one (KLHDC3) gene was expressed among three segments: abomasum, ileum and jejunum. Two genes (MESD and POLK) genes were co-expressed in the abomasum and duodenum. GLTPD2, VWA1 and ECH1 were three DEGs that were common in the ileum and duodenum. Six (TATDN2, RFC1, TRNAE−UUC, MFGE8, TRNAC-GCA and GNG4) genes were shared between abomasum and jejunum. One gene (CUBN) was common in the abomasum and ileum, while 14 genes (LOC105604103, BIRC6, CXCL11, TRNAW-CCA, OTOP2, CDK10, FABP1, PCBP4, LOC106990526, LOC101120782, SPCS1, LOC101107261, EDEM3 and GPR1) were co-expressed in the ileum and jejunum; and 12 DEGs (ALDH8A1, SHISA4, LOC105605834, CCDC85C, LOC106991822, UBR5, LOC105604756, GIT2, WDR55, SLC35D3, LOC114108816 and NUDT22) were common in jejunum and duodenum (Fig. 3). Genes that were only expressed in the ileum were 161, 376 in the jejunum, 199 in the duodenum with 76 exclusively in abomasum as illustrated in Fig. 3.
Venn diagram plot showing the number of differentials expressed genes with significant differences at a false discovery rate (FDR) cut-off < 0.05 for all DEGs of each segment using freely online Venny 2.1 (Oliveros, J.C, 2007–2015) (https://bioinfogp.cnb.csic.es/tools/venny/index.html).
Functional annotation of differentially expressed genes
The GO analysis identified significant genes involved in biological processes, including immune system functions (genes involved SIVA1, APLF, DUSP1 and PGF) and response to stimulus (a gene involved is PGF) were also identified (Fig. 4A). Genes associated with the immune system, and response to stimulus were identified (Fig. 4B). A total of 602 genes were annotated and the high number of annotated genes were from jejunum under all categories. The high number of DEGs was associated with the cellular function category. Moreover, the GO analysis highlighted significant genes involved in various biological processes across all segments, including immune system functions and responses to stimuli. For abomasum, only two genes were responsible for immune response and 12 for response to the stimulus. For biological processes in the ileum, GO analysis revealed five significant genes associated with immune system processes and 15 genes for response to the stimulus. In the jejunum, the GO analysis revealed eight significant genes for immune system processes in addition to 51 genes that were associated with response to the stimulus. In the duodenum, four genes related to immune system processes and twenty genes associated with responses to stimuli were detected. In total, 14 GO terms were found with the most significantly enriched categories of GO functional annotations, as shown in Fig. 4. Most GO terms were associated with BPs, CC and MFs, including cell part (GO:0,044,464), cell (GO:005,623) developmental process (GO:0,032,502), organelle (GO:0,043,226), cellular process (GO:0,009,987) and binding (GO:0,005,488) (Fig. 4C).
GO analysis describing the properties of DEGs of the (A) gastrointestinal tract, (B) abomasum, duodenum, ileum and jejunum of resistant and susceptible sheep, (C)The -log10 of P-values obtained from all datasets of major functional annotation clusters in descending order.
Biological pathways analysis of the DEGs
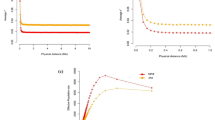
The KEGG pathway enrichment analysis was conducted to gain insight into the biological pathways that were involved in resistant and susceptible animals identified three significantly fold enriched KEGG pathways that were identified using count = 2 and EASE = 0.5 (Table 8). These pathways included the Rap1 signaling pathway, the PI3K-Akt signaling pathway (Fig. 5A), and cancer-related pathways. More than 10 KEGG pathways were identified using count = 2 and EASE = 0.5 in all segments that were significantly fold enriched that are involved in the segments of the resistant and susceptible animals. The enriched pathways include Protein processing in the endoplasmic reticulum pathway, which was enhanced in both duodenum and ileum and Phagosome (Fig. 5B) in both jejunum and ileum, with Arachidonic acid metabolism (Fig. 5C), Metabolic pathways in duodenum and jejunum and PI3K-Akt signaling pathway were notably enhanced in the abomasum (Fig. 5A).
KEGG pathways significantly enriched using up- and down-regulated DEGs for segments of resistant and susceptible sheep. (A) PI3K-Akt signaling pathway significantly enriched in the abomasum (red squares with red stars) and sheep naturally infected with H. contortus (yellow squares with red stars), (B) Phagosome pathway significantly enriched in ileum and jejunum, highlighted in red (Ileum), yellow (Jejunum) and purple (genes shared on both segments), (C) Arachidonic acid metabolism, Metabolic pathways enriched in duodenum and jejunum [red star (jejunum) and blue squares with red stars (duodenum)] are genes involved.
Discussion
This study identified 1364 significantly differentially expressed genes (DEGs) across abomasum, duodenum, jejunum and ileum tissues of Dohne Merino sheep infected with resistance to Haemonchus contortus. Resistant and susceptible animals were classified based on (EBVs) derived from faecal egg count (FEC/EPG) data19. These DEGs and their associated pathways highlight a complex interplay of immune, metabolic and cellular processes that differentiate resistant from susceptible animals. The patterns observed are consistent with previous reports in sheep and goats bred for gastrointestinal nematode (GIN) resistance, such as Merino sheep selected for H. contortus resistance showing upregulation of immune-related genes40, Creole goats demonstrating early activation of Th1/Th2 pathways during infection41, and genome-wide studies in Corriedale and other breeds identifying immune-related QTLs linked to fecal egg count variation(42,43 underscoring the importance of transcriptomic signatures in explaining host–parasite interactions across small ruminant species.
At the gene level, several DEGs of interest were identified. YES1, a proto-oncogene implicated in cell growth and survival, was downregulated in the abomasum of resistant animals, suggesting reduced proliferative activity and a shift towards apoptosis as a protective response. Likewise, EPOR (erythropoietin receptor) was downregulated in resistant sheep, which may reflect tighter control of parasite-induced anaemia compared with susceptible animals, where erythropoietic pathways are often activated following blood loss44. Downregulation of RPSA in resistant sheep is also identified, as altered extracellular matrix interactions may impair parasite establishment in abomasal tissue. While these genes have not been consistently implicated in small ruminant nematode resistance, they suggest tissue-specific host adaptations that warrant further validation.
The stronger and more consistent signals across studies relate to immune signalling and tissue-defence pathways. Transcriptome analysis of Merino selection lines resistant to H. contortus identified upregulation of NOS2, TGFB2, IL2RA and TLR4 in resistant animals, implicating nitric oxide production, T-cell activation and innate antigen recognition44. Comparable RNA-seq studies in Creole goats revealed early activation of Th2/Th1-related pathways in resistant kids(41), while single-cell transcriptomic profiling in goats recently showed cell-type-specific activation of inflammatory programmes involving NF-κB and Th1/Th2 axes(45). Together, these results highlight that coordinated innate and adaptive responses in abomasal mucosa are central to limiting worm establishment and fecundity across both sheep and goats.
Chemokine signalling also emerged as a key resistance mechanism in this study. Notably, CXCL9 was upregulated in the jejunum of resistant animals, a chemokine known to recruit T cells and orchestrate inflammatory responses. Similar cytokine and chemokine-driven axes have been implicated in goat and sheep studies of resistance to GIN3,41. Functional annotation further supported an immune focus, with enriched GO terms and KEGG pathways centred on immune system processes, response to stimulus, apoptosis regulation and phagosome activity. The PI3K–Akt signalling pathway, enriched in the abomasum and jejunum, has been linked to macrophage activation and immune regulation during H. contortus infection(46). The phagosome pathway, enriched in the ileum and jejunum, also underscores the importance of antigen processing and innate immune cell activity in resistant animals.
The enrichment of antigen presentation and inflammatory mediator pathways further highlights how resistant sheep counteract nematode infection. Upregulation of TAP2 and MHC class I genes (OLA-I) in resistant animals suggests improved antigen presentation and cytotoxic responses, consistent with effective helminth control in goats and sheep41,47,48. Inflammatory mediator pathways were also characteristic of resistance. For example, the arachidonic acid metabolism pathway, enriched in duodenum and jejunum, involves PLA2 and LTA4H, enzymes central to leukotriene and eicosanoid production. These molecules govern leukocyte recruitment and inflammatory tone, and elevated eicosanoid and Th2 cytokine signalling (e.g. IL-5) have been reported in resistant Pelibuey and other sheep breeds following H. contortus challenge49,50. These findings confirm that local inflammatory mediator production is a conserved componcent of small ruminant resistance to nematodes.
Importantly, this transcriptomic evidence aligns with genetic studies of nematode resistance in small ruminants. Genomic scans and selection experiments demonstrate that resistance to GIN is heritable and polygenic, with immune-related loci consistently implicated42,43,48 Silva et al.51identified genomic regions associated with parasite resistance traits in sheep, particularly those influencing immune function and faecal egg counts, while Periasamy (2014) reported variation in immune pathway genes strongly associated with faecal egg counts. By mapping our DEGs onto these previously identified loci and pathways, we strengthen the biological interpretation of our findings: resistance in Dohne Merinos is not driven by a single gene effect, but rather by coordinated changes in chemokine/cytokine signalling, antigen presentation, inflammatory mediator synthesis, and regulation of cell growth/apoptosis.
Overall, this study emphasises the parallels between sheep and goats in their genetic and transcriptomic responses to nematode infection. The integration of transcriptomic signatures with candidate gene and genome-wide selection evidence highlights both shared and breed-specific mechanisms of resilience. This provides strong justification for breeding strategies that combine traditional FEC/EPG and EBV selection with genomic and transcriptomic insights to accelerate progress towards sustainable control of H. contortus in small ruminant production systems.
Conclusions
Using RNA sequencing, this study identified key genes and pathways associated with resistance and susceptibility to natural H. contortus infections in Dohne Merino sheep. Resistant animals exhibited strong immune responses, apoptosis regulation, and tissue repair mechanisms, while susceptible animals relied more on DNA repair and alternative defense strategies. Specifically, resistant animals prioritised apoptosis, immune activation (e.g., CXCL9, DLG1), and antigen presentation (TAP2, OLA-I), whereas susceptible individuals showed increased reliance on DNA repair (APLF) and erythropoiesis (EPOR), predisposing them to anaemia. Enrichment of pathways such as PI3K–Akt and arachidonic acid metabolism highlights the interplay between immunity and metabolism in parasite defense. These findings provide new insights into host–parasite interactions with direct applications for breeding programs aimed at enhancing resistance and reducing economic losses caused by H. contortus.
The results further demonstrate that all four gastrointestinal tract segments: abomasum, duodenum, jejunum, and ileum, show diverse defense mechanisms against parasite infection. These include pathways that inhibit or limit infection, stimulate immune responses, regulate stimulus–response functions, and activate tumour suppressors and apoptosis. Collectively, the genes and pathways identified here provide valuable targets for breeding strategies to develop Dohne Merino sheep and other sheep breeds that are more resilient to H. contortus, thereby improving productivity and reducing the economic burden of parasitic infections.
Data availability
The RNA sequence data have been submitted to the NCBI Short Reads Archive (SRA) under the accession number PRJNA1044530 (https://www.ncbi.nlm.nih.gov/bioproject/?term = PRJNA1044530).
References
Nyam, Y. S., Matthews, N. & Bahta, Y. T. Improving livelihood of smallholder farmers through region specific strategies: A case study of South African sheep production. Agrekon https://doi.org/10.1080/03031853.2019.1639205 (2019).
Sargison, N. D. Pharmaceutical treatments of gastrointestinal nematode infections of sheep-future of anthelmintic drugs. Vet. Parasitol 189, 79–84. https://doi.org/10.1016/j.vetpar.2012.03.035 (2012).
Bhuiyan, A. A. et al. Exploring the genetic resistance to gastrointestinal nematodes infection in goat using RNA-sequencing. Int. J. Mol. Sci. https://doi.org/10.3390/ijms18040751 (2017).
Zvinorova, P. I. et al. Breeding for resistance to gastrointestinal nematodes—The potential in low-input/output small ruminant production systems. Vet. Parasitol. 225, 19–28. https://doi.org/10.1016/j.vetpar.2016.05.015 (2016).
Sargison, N. D. Pharmaceutical treatments of gastrointestinal nematode infections of sheep-future of anthelmintic drugs. Vet. Parasitol. 189, 79–84. https://doi.org/10.1016/j.vetpar.2012.03.035 (2012).
Mphahlele, M., Thekisoe, O. M. M., Tsotetsi-Khambule, A. M., Moerane, R. & Mashiloane, M. L. Risk factors associated with occurrence of anthelmintic resistance in sheep of resource-poor farmers in Limpopo province South Africa. Trop. Anim. Health Prod. https://doi.org/10.1007/s11250-018-1724-2 (2018).
De Wolf, B. M. et al. The effect of vitamin E supplementation on an experimental Haemonchus contortus infection in lambs. Vet. Parasitol. 205, 140–149. https://doi.org/10.1016/j.vetpar.2014.07.013 (2014).
Mravčáková, D. et al. Effect of artemisia absinthium and malva sylvestris on antioxidant parameters and abomasal histopathology in lambs experimentally infected with Haemonchus contortus. Animals 11, 1–13. https://doi.org/10.3390/ani11020462 (2021).
Naeem, M., Iqbal, Z. & Roohi, N. Ovine haemonchosis: A review. Trop anim health prod. Trop. Anim. Health Prod. https://doi.org/10.1007/s11250-020-02439-8 (2021).
Besier, R. B., Kahn, L. P., Sargison, N. D. & Van Wyk, J. A. The pathophysiology, ecology and epidemiology of Haemonchus contortus infection in small ruminants. Adv. Parasitol. https://doi.org/10.1016/bs.apar.2016.02.022 (2016).
Besier, R. B., Kahn, L. P., Sargison, N. D. & Van, W. J. A. Diagnosis, treatment and management of Haemonchus contortus in small ruminants. Haemonchus contortus Haemonchosis Past Present Future Trends https://doi.org/10.1016/bs.apar.2016.02.024 (2016).
Tariq, K. A. Anthelmintics and emergence of anthelmintic resistant nematodes in sheep : Need of an integrated nematode management. Int. J. Vet. Sci. Anim. Husb. 2, 13–19 (2017).
Arsenopoulos, K. V., Fthenakis, G. C., Katsarou, E. I. & Papadopoulos, E. Haemonchosis: A challenging parasitic infection of sheep and goats. Animals 11, 1–29. https://doi.org/10.3390/ani11020363 (2021).
Abdo, B., Tesfaye, W. & Shibbiru, T. Prevalence of Haemonchosis and associated risk factors in small ruminants slaughtered at Bishoftu ELFORA export abattoir Ethiopia. J. Nat. Sci. Res. 7, 48–52 (2017).
Halvarsson, P. & Höglund, J. Sheep Nemabiome diversity and its response to anthelmintic treatment in Swedish sheep herds. Parasit. Vectors. 1, 1 (2020).
Van Zyl, E. A. et al. The use of Lespedeza cuneata for natural control of gastrointestinal nematodes in Merino sheep. Onderstepoort J. Vet. Res. 84, 1–7. https://doi.org/10.4102/ojvr.v84i1.1259 (2017).
Ketzis, J. K. et al. Evaluation of efficacy expectations for novel and non-chemical helminth control strategies in ruminants. Vet. Parasitol. 139, 321–335. https://doi.org/10.1016/j.vetpar.2006.04.022 (2006).
Mandal, M., Mishra, C. & Dash, S. K. Genomic insight to the disease resistance in goat. Pharma Innovation 7, 98–103 (2018).
Greer, A. W., McKenzie, J. L., McAnulty, R. W., Huntley, J. F. & McNeilly, T. N. Immune development and performance characteristics of Romney sheep selected for either resistance or resilience to gastrointestinal nematodes. Vet. Parasitol. 250, 60–67. https://doi.org/10.1016/j.vetpar.2017.12.013 (2018).
Read, A. F., Graham, A. L. & Råberg, L. Animal defenses against infectious agents: Is damage control more important than pathogen control?. Plos Biol. 6, 2638–2641. https://doi.org/10.1371/journal.pbio.1000004 (2008).
Klemme, I. & Karvonen, A. Vertebrate defense against parasites: Interactions between avoidance, resistance, and tolerance. Ecol. Evol. 7, 561–571. https://doi.org/10.1002/ece3.2645 (2017).
Guo, Z. et al. Possible mechanisms of host resistance to Haemonchus contortus infection in sheep breeds native to the Canary Islands. Sci. Rep. 6, 1–14. https://doi.org/10.1038/srep26200 (2016).
Jia, J. et al. Effect of high proportion concentrate dietary on Yak jejunal structure, physiological function and protein composition during cold season. Sci. Rep. 11, 1–9. https://doi.org/10.1038/s41598-021-84991-3 (2021).
Gaowa, N., Li, W., Murphy, B. & Cox, M. S. The effects of artificially dosed adult rumen contents on abomasum transcriptome and associated microbial community structure in calves. Genes https://doi.org/10.3390/genes12030424 (2021).
Ma, T. et al. Altered mucosa-associated microbiota in the ileum and colon of neonatal calves in response to delayed first colostrum feeding. J. Dairy Sci. 102, 7073–7086. https://doi.org/10.3168/jds.2018-16130 (2019).
Mohammad, H. J., Ali, A. K. & Al-Ali, Z. A. J. R. Histomorphologal and histochemical structure in the duodenum of sheep (Ovis aries) and rabbit (Oryctolagus cuniculus); a comparative study. Online J. Anim. Feed Res. 10, 251–257 (2020).
Jia, J. et al. Effect of high proportion concentrate dietary on Yak jejunal structure, physiological function and protein composition during cold season. Sci Rep. 11, 1–9. https://doi.org/10.1038/s41598-021-84991-3 (2021).
Manzoni, C. et al. OUP accepted manuscript. Brief Bioinform. https://doi.org/10.1093/bib/bbw114 (2016).
Malatji, D. P., van Marle-Koster, E. & Muchadeyi, F. C. Gene expression profiles of the small intestine of village chickens from an Ascaridia galli infested environment. Vet. Parasitol. X 2, 100012. https://doi.org/10.1016/j.vpoa.2019.100012 (2019).
Swanepoel JW. A genetic evaluation of the Dohne Merino by Table of contents. 2006;
Dlamini NM, Visser C, Soma P, Muchadeyi FC. Genetic diversity and flock clustering of a South African Dohne Merino flock selected for resistance to Haemonchus contortus https://www.researchgate.net/publication/332877408
Dlamini, N. M., Visser, C., Snyman, M. A., Soma, P. & Muchadeyi, F. C. Genomic evaluation of resistance to Haemonchus contortus in a South African Dohne Merino flock. Small Rumin. Res. 175, 117–125. https://doi.org/10.1016/j.smallrumres.2019.04.020 (2019).
Nel, C. et al. Including genomic information in the genetic evaluation of production and reproduction traits in South African Merino sheep. J. Anim. Breed. Genet. 141, 65–82. https://doi.org/10.1111/jbg.12826 (2024).
Wicki, M. et al. Combined genomic evaluation of Merino and Dohne Merino Australian sheep populations. Genet. Sel. Evol. https://doi.org/10.1186/s12711-024-00934-2 (2024).
Pertea M, Kim D, Pertea G, Leek JT, Steven L. HHS Public Access. 2017;11: 1650–67. https://doi.org/10.1038/nprot.2016.095.Transcript-level
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13. https://doi.org/10.1093/nar/gkn923 (2009).
Ye, J. et al. WEGO: A web tool for plotting GO annotations. Nucleic Acids Res. 34, 293–297. https://doi.org/10.1093/nar/gkl031 (2006).
Chen, L. et al. Gene ontology and KEGG pathway enrichment analysis of a drug target-based classification system. PLoS ONE 10, 1–12. https://doi.org/10.1371/journal.pone.0126492 (2015).
Li, B. et al. Transcriptome analysis of adipose tissues from two fat-tailed sheep breeds reveals key genes involved in fat deposition. BMC Genomics https://doi.org/10.1186/s12864-018-4747-1 (2018).
Zhang, R. et al. Transcriptome analysis unraveled potential mechanisms of resistance to Haemonchus contortus infection in Merino sheep populations bred for parasite resistance. Vet. Res. https://doi.org/10.1186/s13567-019-0622-6 (2019).
Aboshady, H. M. et al. Dynamic transcriptomic changes of goat abomasal mucosa in response to Haemonchus contortus infection. Vet. Res. 51, 1–12. https://doi.org/10.1186/s13567-020-00768-y (2020).
McRae, K. M., McEwan, J. C., Dodds, K. G. & Gemmell, N. J. Signatures of selection in sheep bred for resistance or susceptibility to gastrointestinal nematodes. BMC Genomics https://doi.org/10.1186/1471-2164-15-637 (2014).
Benavides, M. V., Sonstegard, T. S. & Van Tassell, C. Genomic regions associated with sheep resistance to gastrointestinal nematodes. Trends Parasitol. https://doi.org/10.1016/j.pt.2016.03.007 (2016).
Lacroux, C. et al. Haemonchus contortus (Nematoda: Trichostrongylidae) infection in lambs elicits an unequivocal Th2 immune response. Vet. Res. 37, 607–622. https://doi.org/10.1051/vetres:2006022 (2006).
Wang, W. et al. Single-cell transcriptomic profiling unveils dynamic immune cell responses during Haemonchus contortus infection. Cells https://doi.org/10.3390/cells13100842 (2024).
Liu, J. et al. From innate to adaptive immunity: Abomasal transcriptomic responses of merino sheep to Haemonchus contortus infection. Mol. Biochem. Parasitol. 246, 6–11. https://doi.org/10.1016/j.molbiopara.2021.111424 (2021).
Gan, Q. F., Li, Y. R., Lund, M., Su, G. S. & Liang, X. W. Genome-wide association study identifies loci linked to serum electrolyte traits in Chinese Holstein cattle. Anim. Genet. Blackwell Publ. Ltd 50, 744–748. https://doi.org/10.1111/age.12851 (2019).
Benavides, M. V., Souza, C. J. H., Smith, W. D. & Moraes, J. C. F. Evaluation of protection in grazing lambs immunised with different doses of Haemonchus contortus gut membrane glycoproteins in Southern Brazil. Vet. Parasitol. https://doi.org/10.1016/j.vetpar.2021.109360 (2021).
Flay, K. J., Hill, F. I. & Muguiro, D. H. A review: Haemonchus contortus infection in pasture-based sheep production systems, with a focus on the pathogenesis of anaemia and changes in haematological parameters. Animals https://doi.org/10.3390/ani12101238 (2022).
Torres-Acosta, J. F. J. et al. Nutritional manipulation of sheep and goats for the control of gastrointestinal nematodes under hot humid and subhumid tropical conditions. Small Rumin. Res. 103, 28–40. https://doi.org/10.1016/j.smallrumres.2011.10.016 (2012).
Silva,M.V.G.B., et al. Identificationof quantitative trait loci affecting resistance to gastrointestinal parasites in a Red Maasai × Dorper backcross population. BMCGenomics. 13, 472. (2012).
Acknowledgements
The research received joint funding from the Agricultural Research Council—Biotechnology Platform and the National Research Foundation (SFH170627245382). Wauldby Farm (Robbie Blaine) for supplying us with the Dohne Merino sheep and the Tandala abattoir for slaughtering the animals for us to collect samples.
Author information
Authors and Affiliations
Contributions
T.M. performed data collection, lab work, statistical analyses and drafted the manuscript. F.C. was the project leader and engaged in the structure of the manuscript. R.E., D.P., M.A. and P. were responsible for experimental and project design and also participated in the coordination and preparation of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
Gastrointestinal tissues were collected from sheep carcasses that were slaughtered at Tandala Farm Abattoir, which is a commercial abattoir. The animal procedures described in this study were conducted under the approval of the Animal Research Ethics Committee of the College of Agriculture and Environmental Sciences, University of South Africa (UNISA), with the reference number: 2019/CAES/037. The study was reported in accordance with ARRIVE guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ramantswana, T.M., Malatji, D.P., Pierneef, R.E. et al. Differential gene expression analysis of Dohne Merino sheep naturally infected with Haemonchus contortus. Sci Rep 15, 41843 (2025). https://doi.org/10.1038/s41598-025-25782-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25782-y