Abstract
T-cell acute lymphoblastic leukemia (T-ALL) is an aggressive hematologic malignancy arising from abnormal T-cell differentiation. Conventional treatment strategies remain suboptimal, and no effective targeted therapy is currently available. Here, we found that ASS1 is significantly upregulated in T-ALL and is associated with poor prognosis. Inhibition of ASS1 expression markedly suppresses T-ALL progression both in vitro and in vivo. Mechanistically, RNA-seq revealed that ASS1 depletion led to a significant downregulation of one-carbon metabolism and ribose synthesis pathways, and the mTORC1/c-Myc signaling pathway. Experimental validation confirmed that mTORC1/c-Myc signaling was significantly downregulated upon ASS1 blockage. Furthermore, the intracellular arginine level was significantly reduced following ASS1 knockdown, while supplementation with arginine enhanced mTORC1/c-Myc pathway activity. Notably, restoration of c-Myc effectively rescued T-ALL cells from the suppressive effects induced by ASS1 depletion. Collectively, our findings demonstrate that ASS1 supports mTORC1/c-Myc pathway activity by regulating arginine availability, thereby promoting the survival of T-ALL cells and leukemia progression.
Similar content being viewed by others
Introduction
T-cell acute lymphoblastic leukemia (T-ALL) is an aggressive hematologic malignancy characterized by malignant proliferation and differentiation arrest of hematopoietic precursors1,2. Its marked heterogeneity impedes the development of effective therapies3,4,5. Therefore, identifying novel therapeutic targets and elucidating their key mechanistic roles in T-ALL is of critical importance for overcoming the clinical challenges in T-ALL treatment.
Metabolic reprogramming enables tumor cells to adopt distinct metabolic dependencies for survival and proliferation6,7. Argininosuccinate synthase 1 (ASS1) is a rate-limiting metabolic enzyme that catalyzes the ATP-dependent condensation of citrulline and aspartate to form argininosuccinate in the urea cycle and arginine biosynthesis pathways8. In over 70% of tumors, transcriptional suppression of ASS1 induces arginine auxotrophy, thereby establishing the mechanistic basis for arginine-deprivation therapy9,10. However, ASS1 is frequently upregulated across diverse malignancies and functions as an oncogene; in ovarian cancer, for example, elevated ASS1 expression antagonizes apoptotic pathways via the AMPK/CPT1A signaling axis11. Although arginine deprivation therapy represents an effective strategy for ASS1-deficient tumors, this approach typically proves ineffective against malignancies with ASS1 upregulation12,13,14. Therefore, elucidating the tumor-promoting mechanisms of ASS1 in ASS1-positive neoplasms carries substantial clinical implications.
Here, we report for the first time that ASS1 exhibits significant upregulation in T-ALL. ASS1 knockdown significantly suppresses T-ALL cell survival in vitro and inhibits leukemia progression in vivo. Mechanistic studies revealed that ASS1 knockdown suppresses mTORC1/c-Myc pathway and inhibits T-ALL cell survival and progression. These findings reveal the importance of ASS1 in T-ALL progression and provide a novel therapeutic target for T-ALL therapy.
Results
Elevated ASS1 expression in T-ALL correlates with adverse prognosis
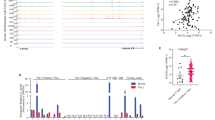
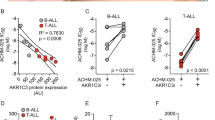
T-ALL cells harbor numerous genetic alterations, with specific gain-of-function mutations driving oncogenic survival and proliferation pathways that may serve as promising therapeutic targets2,15. To systematically identify aberrantly upregulated genes in T-ALL, we analyzed a T-ALL RNA-seq dataset and found that ASS1 was markedly elevated in leukemic blasts (Fig. 1A). We further investigated ASS1 expression across multiple leukemia cell lines and revealed its highest mean expression level in T-ALL. (Fig. 1B). To validate these observations, we quantified both mRNA and protein levels of ASS1 in T-ALL cell lines (Jurkat, MOLT-4, KE-37, CUTLL1, Loucy, CCRF-CEM) and normal T cells. Consistently, ASS1 exhibited marked upregulation in all malignant cell lines compared to normal T cells (Fig. 1C, D). Importantly, ALL patients with higher ASS1 expressions had poor survival outcomes than patients with lower ASS1 expressions (Fig. 1E). Collectively, these findings demonstrate significant ASS1 upregulation in T-ALL and with strong clinical prognostic relevance, implicating it as a potential biomarker for T-ALL patient prognosis.
ASS1 high expression in T-ALL correlates with adverse prognosis. A. mRNA expression levels of ASS1 in T-ALL RNA-seq databases (GEO, GSE26713). Dataset includes 7 normal bone marrow (BM) and 117 T-ALL samples. Y-axis represents the normalized expression values (log₂-transformed signal intensities) obtained from microarray analysis (RMA normalization). B. mRNA expression levels of ASS1 in ALL RNA-seq databases (GEO, GSE13425), n = 190. Y-axis represents the normalized expression values (log₂-transformed signal intensities) obtained from microarray analysis (RMA normalization). C The qPCR analysis of ASS1 expressions in T-ALL cell lines versus normal T cells. The Y-axis shows fold change relative to Normal-T1. D Protein expression levels of ASS1 in T-ALL cell lines and normal T cells. E Event-free survival in ALL patients stratified by ASS1 expression (n = 256, dbGaP, phs000463 and phs000464). Data are mean ± SD (Two-tailed unpaired student’s t-test, *** p < 0.001).
ASS1 Blockade inhibits T-ALL cell proliferation and survival
To investigate the role of ASS1 in T-ALL pathogenesis, we performed ASS1 knockdown in T-ALL cell lines exhibiting high ASS1 expression (Jurkat, MOLT-4, KE-37). Notably, ASS1 depletion profoundly inhibited proliferation across all tested cell lines (Fig. 2A-C), suggesting its critical role in maintaining leukemic cell survival and growth in vitro. To determine whether the inhibition of T-ALL cells was caused by the off-target effect of shRNA, we performed rescue experiments by overexpressing shRNA-resistant ASS1. The results showed that reintroduction of shRNA-resistant ASS1 rescued the proliferative defect of T-ALL cells, indicating that the growth inhibition of T-ALL cells caused by ASS1 knockdown was not due to shRNA off-target effects (Fig. 2D). Furthermore, we assessed apoptosis in T-ALL cells following ASS1 knockdown and showed significantly increased Annexin V/PI double-positive apoptotic cells in both Jurkat and MOLT-4 cell lines upon ASS1 depletion (Fig. 2E, F). Together, these results demonstrate that ASS1 knockdown effectively inhibits cellular proliferation and induces apoptosis in T-ALL cells.
ASS1 depletion suppresses T-ALL proliferation and induces apoptosis. A-C. ASS1 Protein expression and proliferation curves of (A) Jurkat cells, (B) MOLT-4 cells, (C) KE-37 cells following ASS1 knockdown. The knockdown efficiency of T-ALL cells was detected at 72 h after transfection with the shASS1 lentiviral vector, and cell counting was performed on Day 3, 5, and 7 after transfection. D. ASS1 Protein expression and proliferation curves of Jurkat ASS1 knockdown cells, and ASS1-knockdown & overexpression cells. E. Apoptosis analysis in Jurkat cells following ASS1 knockdown. Apoptosis was quantified at 72 h after transfection with ASS1-sh1 lentiviral vector. F. Apoptosis analysis in MOLT-4 cells following ASS1 knockdown. Apoptosis was detected 72 h after transfection with ASS1-sh1 lentiviral vector. Data are mean ± SD (Two-tailed unpaired Student’s t-test, * p < 0.05, ** p < 0.01, *** p < 0.001).
To investigate the impact of ASS1 knockdown on the progression of T-ALL in vivo, we established a doxycycline-inducible ASS1 knockdown Jurkat cell line and validated its efficiency in vitro, resulting in significant ASS1 decrease and growth suppression with doxycycline treatment (Fig. 3A, B). Further, we generated cell-derived xenograft (CDX) mouse models by using doxycycline-inducible ASS1-knockdown Jurkat cells. Mice received doxycycline from day 12 post-engraftment to day 27, and with longitudinal tumor burden monitored via bioluminescence imaging (Fig. 3C). Similar to the results in vitro, ASS1 knockdown suppressed T-ALL progression in vivo. (Fig. 3D, E). Bone marrow infiltration, which is frequently observed in the progression of T-ALL, is considered a major cause of relapse and refractory disease16. Then, we examined bone marrow invasion and found the invasion of T-ALL cells were dramatically decreased after doxycycline-induced ASS1 knockdown (Fig. 3F, G). Additionally, the survival time of T-ALL CDX models was significantly extended following doxycycline-induced ASS1 knockdown (Fig. 3H). These results demonstrated that ASS1 depletion attenuates T-ALL progression in vivo, underscoring the critical role of ASS1 in T-ALL progression.
ASS1 depletion impairs T-ALL progression in vivo. A-B. ASS1 Protein expression and cell proliferation of doxycycline-inducible ASS1-knockdown Jurkat cells. C. Experimental design for T-ALL cell-derived xenograft (CDX) model. Mice were injected with 3 × 10⁶ luciferase labeled Jurkat-tet-ASS1-sh1 cells via the tail vein on day 0. On the day 12–27, the mice were administered 1.5 ug/mL doxycycline in drinking water, and the progression of T - ALL in the mice was detected by bioluminescence imaging. D. Bioluminescence imaging in vivo of leukemic progression in mice (n = 4). E. Quantitative analysis of leukemic burden in mice. p values for shC vs. shASS1 on day 27. F. Flow cytometry analysis of T-ALL cell invasion to bone marrow by using anti-human CD7 antibody through flow cytometry. G. Statistical quantification of T-ALL cell invasion to bone marrow. H. Survival curves of T-ALL CDX (n = 5). Data are mean ± SD (two-tailed unpaired student’s t-test, * p < 0.05, ** p < 0.01).
ASS1 knockdown abrogates mTORC1/c-Myc axis in T-ALL
ASS1 has been identified as essential for T-ALL growth and progression. To investigate the underlying mechanism, we performed RNA-seq analysis. A heatmap revealed extensive transcriptional alterations in Jurkat cells following ASS1 knockdown (Fig. 4A and Supplementary Dataset File). Pathway analysis (KEGG database) of differentially expressed genes (DEGs) highlighted significant downregulation of the folate-mediated one-carbon metabolism pathway (e.g., AHCY, MAT1A) and the nucleotide sugar biosynthesis pathway (e.g., HK2, PGM2)17,18,19, demonstrating that ASS1 knockdown disrupts one-carbon metabolism and nucleotide synthesis in T-ALL cells (Fig. 4A, B). One-carbon metabolism and nucleotide synthesis are well-established pathways that promote tumor growth and proliferation20,21. Given the strong link between one-carbon metabolism and the mTORC1/c-Myc, we focused on examining the change of mTORC1 and c-Myc signaling after ASS1 knockdown22,23. GSEA analysis demonstrated marked reductions in Hallmark mTORC1 signaling (e.g., RRP9, HMGCS1), and Hallmark MYC targets pathways (e.g., MCM2, DEK) (Fig. 4C-F). Additionally, significant enrichment of cell cycle-related pathways (Hallmark E2F targets and Hallmark G2M checkpoint) suggested impaired proliferation in ASS1-depleted T-ALL cells (Fig. 4C, G). To further confirm the accuracy of RNA-seq data, we verified the RNA expression levels of the classical genes in the above pathway by qPCR. We found that their levels decreased significantly after ASS1 knockdown, which was consistent with the RNA-seq data (Fig. 4H). Collectively, our findings suggest that ASS1 facilitates T-cell acute lymphoblastic leukemia survival and growth through the mTORC1/c-Myc pathway.
Targeting ASS1 suppresses T-ALL progression through the mTORC1/c-Myc axis. A. Differentially expressed genes (DEGs) analysis in ASS1-knockdown Jurkat cells. Color scale represents Z-scores of gene expression. B. KEGG pathway analysis of altered signaling in ASS1-knockdown Jurkat cells. C. GSEA analysis of pathways modulated by ASS1 knockdown. D-G. GSEA plots for enriched pathways: (D) mTORC1 signaling, (E) c-Myc targets V1, (F) c-Myc targets V2, (G) G2M checkpoint. H. The mRNA expression level of the described gene. 48 h after transfecting the ASS1 knockdown virus, total RNA from T-ALL cells was extracted, and mRNA quantification was performed using qPCR. Data are mean ± SD (Two-tailed unpaired Student’s t-test, * p < 0.05, ** p < 0.01, *** p < 0.001).
ASS1 Inhibition disrupts mTORC1/c-Myc signaling via arginine depletion
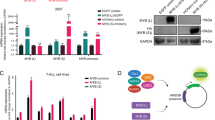
To validate the regulatory role of ASS1 in the mTORC1/c-Myc pathway, we quantified c-Myc protein expression The c-Myc protein levels were significantly upregulated in T-ALL cells compared to normal T cells (Fig. 5A). Furthermore, we performed a correlation analysis of ASS1 and c-Myc expression levels in T-ALL cell lines, revealing a positive correlation between ASS1 and c-Myc expression in T-ALL (Fig. 5B). To investigate whether ASS1 regulates the mTORC1/c-Myc pathway, we knocked down ASS1 and then detected the activity of mTORC1 and the protein level of c-Myc. We found that A significant reduction in both S6 phosphorylation (a reliable marker of mTORC1 activation24 and c-Myc expression was observed upon ASS1 depletion (Fig. 5C-E). Through our RNA-seq and qPCR results, we found that the mRNA level of c-Myc was significantly reduced. Therefore, we speculated that c-Myc is regulated at the transcriptional level (Fig. 4A, H). As mTORC1 activation requires amino acid sufficiency (including arginine)25, we hypothesized ASS1 depletion impairs mTORC1 signaling via arginine depletion. Therefore, we measured the arginine levels in T-ALL cells after ASS1 knockdown and found that arginine levels were markedly decreased (Fig. 5F). To verify the effects of arginine on mTORC1/c-Myc pathway, we treated T-ALL cells with culture media with or without arginine and found that arginine supplementation restored mTORC1 activity and c-Myc expression (Fig. 5G). Additionally, the proliferation ability of T-ALL cell lines was significantly enhanced after adding arginine (Fig. 5H). Based on the above results, ASS1 sustains the activity of the mTORC1/c-Myc pathway by maintaining the intracellular arginine level.
ASS1 couples arginine metabolism to mTORC1/c-Myc activation. A. c-Myc expression levels in T-ALL cell lines versus normal T cells. B. Correlation analysis of ASS1 and c-Myc protein expression in T-ALL and normal T cells. C-E. Protein levels of ASS1, p-S6, and c-Myc in ASS1-knockdown (C) Jurkat, (D) MOLT-4, and (E) KE-37 cells. T-ALL cells were collected at 72 h after transfection using lentivirus vectors, using non-target shRNA (shC) as negative control. F. Intracellular arginine quantification in ASS1-knockdown Jurkat cells. Jurkat cells were first transfected with the lentiviral vector carrying shASS1. After 72 h, the cells were cultured in arginine-free medium for 24 h, and then the intracellular arginine levels were quantified. G. mTORC1/c-Myc pathway activity and (H) cell proliferation of shC control and ASS1-knockdown Jurkat cells with or without arginine (500 µM) supplementation for 48 h. Data are mean ± SD (Two-tailed unpaired Student’s t-test, ** p < 0.01).
To determine if the mTORC1/c-Myc pathway is the main pathway through which ASS1 maintains the survival and proliferation of T-ALL cells, we knocked down ASS1 and performed c-Myc rescue experiments. The results showed that the c-Myc overexpression reversed ASS1 depletion-induced proliferation defects (Fig. 6A-C). Collectively, ASS1 sustains mTORC1/c-Myc activity through arginine provision, thereby driving T-ALL survival, proliferation, and disease progression (Fig. 6D).
c-Myc overexpression rescues proliferation defects in ASS1-deficient T-ALL cells. A-C Protein expression and proliferation analysis in ASS1-knockdown T-ALL cells with c-Myc overexpression: (A) Jurkat, (B) MOLT-4, (C) KE-37. T-ALL cells were collected at 72 h after transfection using lentivirus vectors. The y-axis represents fold change relative to Day 3 controls. D Schematic diagram illustrating the mechanism of ASS1-mediated regulation through arginine-mediated mTORC1/c-Myc signaling. Data are mean ± SD (two-tailed unpaired student’s t-test, * p < 0.05, ** p < 0.01).
Discussion
Due to the lack of mature targeted therapeutic drugs, the refractoriness and recurrence of T-ALL have always been difficult problems faced in the clinic26. Therefore, there is an urgent need to develop new therapeutic targets. Our study demonstrated for the first time that the expression of ASS1 is upregulated in T-ALL. ASS1 promotes the survival and in vivo progression of T-ALL. Mechanistically, it was demonstrated that ASS1 promotes the activity of the mTORC1/c-Myc pathway by maintaining intracellular arginine levels. We innovatively proposed that ASS1 is a potential therapeutic target for T-ALL, which has important theoretical and clinical reference significance.
Argininosuccinate synthase 1 (ASS1) is a key rate-limiting enzyme in the urea cycle and can catalyze the formation of argininosuccinate from citrulline and aspartate27. Under the action of argininosuccinate lyase (ASL), arginine is produced28. In various tumors, the loss of ASS1 can promote tumor progression, such as in myxofibrosarcomas, breast cancer, malignant melanoma, liver cancer, prostate cancer, pancreatic cancer, renal cancer, and osteosarcomas9,29,30. This is mainly because ASS1 can catalyze the combination of aspartate and citrulline to argininosuccinate, resulting in the consumption of aspartate, which is essential for tumor proliferation by synthesizing pyrimidine nucleotides9. Interestingly, many studies in recent years have shown that ASS1 is beneficial for the survival of tumor cells. Feng et al. pointed out that ASS1 is significantly highly expressed in ovarian carcinoma and can enhance the resistance of ovarian cancer to anoikis through AMPK/CPT1A - mediated fatty acid metabolism11. Hu et al. showed that ASS1 can promote the tolerance of NSCLC to ferroptosis by mediating the reductive carboxylation of cytoplasmic glutamine31. This indicates that ASS1 plays an important role in drug resistance and promotes survival in certain tumors. In this study, our results reveal that the expression of ASS1 was significantly regulated in T-ALL. Moreover, knocking down ASS1 could significantly inhibit the survival of T-ALL in vitro and the progression of leukemia in vivo, which is similar to the previous studies showing that ASS1 promotes tumor survival.
Arginine is an amino acid with diverse functions produced in the urea cycle and is the substrate downstream of the rate-limiting enzyme ASS132. The intracellular arginine level is regulated by multiple proteins, such as arginine synthesis-related enzymes (ASS1, ASL), arginine transporters (SLC7A1), and arginine decomposition-related enzymes (ARG1/2, NOS). Our results show that knocking down ASS1 can significantly reduce the arginine level in T-ALL cells, which is consistent with previous studies.
Arginine is a semi-essential amino acid with diverse functions. Apart from being a basic component of proteins, it is also a precursor of polyamines, creatinine, and nitric oxide, which can promote the survival and proliferation of tumor cells7. mTORC1 is a major growth regulatory factor that is activated by nutritional factors at the lysosome. Chantranupong et al. reported that arginine can activate the mTORC1 pathway through the CASTOR protein25,33. Our results indicate that knocking down ASS1 can reduce the intracellular arginine level and decrease the activity of the mTORC1/c-Myc pathway. Additionally, the results also demonstrate that arginine can directly affect the activity of the mTORC1/c-Myc pathway. These results are consistent with the previous research conclusions. Subsequently, we also performed a rescue experiment with overexpression of c-Myc to determine whether mTORC1/c-Myc is the main pathway downstream of ASS1 that affects the survival and proliferation of T-ALL. The results showed that after overexpression of c-Myc, the inhibition of T-ALL cells caused by ASS1 knockdown was rescued.
This study elucidated that ASS1 could serve as a potential target for T-ALL therapy and illustrated the key mechanism by which blocking ASS1 inhibits the progression of T-ALL. However, this study only verified the findings in T-ALL cell lines and CDX, but not in primary T-ALL leukemia cells and patient-derived tumor xenograft (PDX), which is the limitation of this study.
In summary, this study for the first time proposes the importance of ASS1 in maintaining the survival of T-ALL cells and the progression of leukemia in animal models. Moreover, it demonstrates the mechanism by which ASS1 maintains the activity of the mTORC1/c-Myc pathway by regulating the intracellular arginine level, providing a new theoretical basis for the targeted therapy of T-ALL.
Methods
Ethics statement
Human T cells were acquired from Changsha Economic Development under the protocol JH-WBC-20,230,417, ethical document number K202304-023.
All the mice were maintained in pathogen-free facilities and used strictly in accordance with the protocols approved by Hainan University Institutional Animal Use and Care Committee under animal license ID: HNUAUCC-2024-00183. All animal care and use of experimental procedures were provided by the Hainan University Institutional Animal Use and Care Committee and conducted as per the ARRIVE guidelines.
Cell culture
Jurkat and HEK-293T were purchased from the Chinese Academy of Sciences Cell Bank (CASCB). MOLT-4, CCRF-CEM and Loucy were purchased from the American Type Tissue Culture Collection (ATCC). KE-37 cells were purchased from the Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures (DSMZ). CUTLL-1 cells were purchased from Sigma-Aldrich (SCC286). The Jurkat, MOLT-4, KE-37, CUTLL-1, Loucy, and CCRF-CEM were cultured in RPMI-1640 (Gibco, C11875500BT) with 10% FBS (Gibco, 10099-141), HEK-293T were cultured in DMEM (Gibco, C11995500BT) with 10% FBS (Gibco, 10099-141). All these cells were confirmed by STR profiling analysis and were cultured in 37 ℃ cell incubator with 5% CO2.
Plasmids construction
Gene knockdown was achieved using short hairpin RNA (shRNA) constructs cloned into the pLKO.1 lentiviral vector. The shRNA sequences were designed as follows: ASS1-sh1: 5’- GGAACGATCAGGTCCGGTTTGCTCGAGCAAACCGGA, CCTGATCGTTCCttttt-3’. ASS1-sh2: 5’-GCTATGACGTCATTGCCTATCCTCGA, GGATAGGCAATGACGTCATAGCttttt-3’. ASS1-shUTR: 5'-CCCAAGTACAGGCGC, TAATTGCTCGAGCAATTAGCGCCTGTACTTGGGttttt-3'. The shRNA fragments into the Age1 and EcoR1 restriction sites of the pLKO.1 puro plasmid (Addgene, 8453). Similarly, gene overexpression was achieved by PCR amplification of the target gene’s coding sequence (CDS), followed by molecular cloning into the pLVX-M-puro plasmid (Addgene, 125839).
Lentiviral vector Preparation
For lentiviral production, HEK-293T cells were co-transfected with the pLKO.1 puro or pLVX-M-puro plasmid, along with the packaging plasmids psPAX2 (Addgene, 12260) and pMD2.G (Addgene, 12259), using a standard calcium phosphate transfection protocol. Viral supernatants were harvested 72 h post-transfection and concentrated via ultracentrifugation at 25,000 × g for 3 h. The concentrated lentiviral particles were then used to transduce T-cell acute lymphoblastic leukemia (T-ALL) cells. At 72 h post-transduction, gene expression levels were assessed by quantitative RT-PCR or Western blotting to confirm knockdown efficiency.
Quantitative PCR
Total RNA was isolated from cells using a commercial RNA extraction kit, and cDNA was synthesized from 1 µg RNA with a reverse transcription kit. qPCR reactions were performed in triplicate using SYBR Green master mix under the following conditions: 95 °C for 2 min (initial denaturation), followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Melt curve analysis (60–95 °C) confirmed amplicon specificity. Relative gene expression was calculated via the ΔΔCt method normalized to GAPDH.
Western Immunoblotting
Cells were lysed in RIPA buffer containing protease inhibitors. Protein concentrations were determined by BCA assay. Equal amounts (10–20 µg) of protein lysates were separated by SDS-PAGE (8–12% gels) and transferred to PVDF membranes. Membranes were blocked with 5% non-fat milk in TBST (1 h, RT) and incubated overnight at 4 °C with primary antibodies (1:1000), including ASS1 (CST, 70720 S), c-Myc (CST, 18583 S), p-S6 (ABclonal, AP0537), S6 (Proteintech, 66886-1-Ig), β-Tubulin (ABclonal, A12289). Then, incubate the membrane with an appropriate secondary antibody. Detect the target protein bands using the ChemiDoc imaging system (Bio-Rad).
Flow cytometry
Bone marrow cells were harvested, washed thrice with a washing buffer (PBS supplemented with 5% FBS), and subsequently aliquoted at a density of 1 × 10⁶ cells per sample. The cells were incubated with an anti-human CD7 antibody (BD Biosciences, 562635) in the dark at 4 °C for 30 min. To eliminate unbound antibodies, the cells were subjected to three additional washes with the washing buffer. Prior to analysis, the cells were resuspended in PBS containing 7-AAD viability dye (at a dilution of 1:50) to exclude non-viable cells. For apoptosis assessment, T-ALL cells were stained with Annexin V-FITC (1:20) and PI staining solution (1:10) from an apoptosis detection kit (Yeasen, 40302ES50). Samples were promptly analyzed using a CytoFLEX S flow cytometer (Beckman Coulter).
L-arginine levels analysis
L-Arginine levels were quantified using a colorimetric assay kit (Elabscience, E-BC-K850-M). In brief, 3 × 10⁶ cells were homogenized in 200 µL of 0.9% NaCl solution. Subsequently, the homogenate was centrifuged at 12,000 × g for 10 min. Then, 150 µL of the supernatant was mixed with one-fourth of its volume of Reagent 4. After incubation at 37 °C for 20 min, the samples were centrifuged again at 12,000 × g for 10 min, followed by filtration through a 10 kDa ultrafiltration unit. The filtrate was then reacted with 60 µL of Reagent 3 at 60 °C for 2 h. Next, 400 µL of the chromogenic reagent was added. The mixture was boiled at 98 °C for 30 min, cooled down, and 200 µL aliquots were taken to measure the absorbance at 520 nm using a microplate reader. The concentrations were calculated based on a standard curve.
RNA-seq analysis
For sample preparation, Jurkat cells were transfected with either ASS1 knockdown lentiviral vectors or non-target control lentiviral vectors. After 72 h, total RNA was extracted, and mRNA libraries (300–400 bp) were generated using the Ultima Dual-Mode RNA Library Prep Kit for Illumina (Yeasen, 12252ES24). Paired-end sequencing (150 bp) was conducted on a Illumina NovaSeq 6000 platform.
Reads were aligned to the hg38 genome using STAR. Counts for each gene were obtained using featureCounts, and gene expression levels were quantified with DESeq2 to calculate normalized counts. Differentially expressed genes (DEGs) were identified based on p < 0.05 and |log₂ Fold Change| > 2. Functional enrichment analyses, including KEGG pathway and Gene Set Enrichment Analysis (GSEA), were conducted using the clusterProfiler package.
Mouse models
The 6 to 8-week-old NCG mice (Strain NO. T001475, NOD/ShiLtJGpt-Prkdcem26Cd52Il2rgem26Cd22/Gpt) were purchased from GemPharmatech (Nanjing, China). For establishing Jurkat cells-derived xenograft (CDX), each NCG mouse was injected with 3e6 Jurkat-tet-shASS1 cells via the tail vein. From day 12 to day 27, Doxycycline (1.5 ug/ml) was added to the drinking water of the ASS1 knockdown CDX group. The imaging of CDX was acquired with an in vivo imaging instrument (Spectral Instruments Imaging, AMI HTX) according to its protocol. When the experimental endpoint was reached, all mice were euthanized. Specifically, all mice were induced into a pain-free state by inhalation anesthesia (isoflurane, 5%) before performing cervical dislocation. Subsequently, bone marrow cells were obtained from the femurs of mice for T-ALL infiltration detection.
Statistical analysis
The intergroup comparisons were assessed using two-tailed unpaired Student’s t-tests. Unless explicitly stated, three independent replicates have been performed unless otherwise specified, and data are represented as means ± SD. Analytical computations were executed through GraphPad Prism software (v8.4.3). The survival curves were tested with the log-rank test. * p < 0.05, ** p < 0.01, *** p < 0.001, ns = no significance.
Data availability
The RNA-seq sequence data have been deposited in the Genome Sequence Archive (GSA) for human under the accession number HRA010986. The other published data utilized in this paper are listed as follows: The microarray data were obtained from the Gene Expression Omnibus (GEO) (GSE26713, GSE13425), and the survival data were retrieved from TARGET (phs000463, phs000464). The data that supports the findings of this study are available on request from the corresponding author upon reasonable request.
References
Balducci, E. et al. Genomic imbalance analysis provides new insight into prognostic factors in adult and pediatric T-ALL. Blood 144, 988–1000. https://doi.org/10.1182/blood.2023022154 (2024).
Simonin, M. et al. NGS-based stratification refines the risk stratification in T-ALL and identifies a very-high-risk subgroup of patients. Blood 144, 1570–1580. https://doi.org/10.1182/blood.2023023754 (2024).
Si Lim, S. J., Ford, J. B. & Hermiston, M. L. How I treat newly diagnosed and refractory T-cell acute lymphoblastic lymphoma in children and young adults. Blood 141, 3019–3030. https://doi.org/10.1182/blood.2022016503 (2023).
Anand, P. et al. Single-cell RNA-seq reveals developmental plasticity with coexisting oncogenic States and immune evasion programs in ETP-ALL. Blood 137, 2463–2480. https://doi.org/10.1182/blood.2019004547 (2021).
Gocho, Y. et al. Network-based systems Pharmacology reveals heterogeneity in LCK and BCL2 signaling and therapeutic sensitivity of T-cell acute lymphoblastic leukemia. Nat. Cancer. 2, 284–299. https://doi.org/10.1038/s43018-020-00167-4 (2021).
Zhang, Z. J. et al. ATF4 renders human T-cell acute lymphoblastic leukemia cell resistance to FGFR1 inhibitors through amino acid metabolic reprogramming. Acta Pharmacol. Sin. 44, 2282–2295. https://doi.org/10.1038/s41401-023-01108-4 (2023).
Mossmann, D. et al. Arginine reprograms metabolism in liver cancer via RBM39. Cell 186, 5068–5083 e5023, https://doi.org/10.1016/j.cell.2023.09.011 (2023).
Keshet, R. et al. Targeting purine synthesis in ASS1-expressing tumors enhances the response to immune checkpoint inhibitors. Nat. Cancer. 1, 894–908. https://doi.org/10.1038/s43018-020-0106-7 (2020).
Rabinovich, S. et al. Diversion of aspartate in ASS1-deficient tumours fosters de Novo pyrimidine synthesis. Nature 527, 379–383. https://doi.org/10.1038/nature15529 (2015).
Gupta, S. et al. Argininosuccinate Synthetase-1 (ASS1) loss in High-Grade neuroendocrine carcinomas of the urinary bladder: implications for targeted therapy with ADI-PEG 20. Endocr. Pathol. 29, 236–241. https://doi.org/10.1007/s12022-018-9516-9 (2018).
Feng, X. et al. ASS1 enhances Anoikis resistance via AMPK/CPT1A-mediated fatty acid metabolism in ovarian cancer. Cancer Lett. 217082 https://doi.org/10.1016/j.canlet.2024.217082 (2024).
Szlosarek, P. W. et al. Arginine deprivation with pegylated arginine deiminase in patients with argininosuccinate synthetase 1-Deficient malignant pleural mesothelioma: A randomized clinical trial. JAMA Oncol. 3, 58–66. https://doi.org/10.1001/jamaoncol.2016.3049 (2017).
Chu, C. Y. et al. Genome-wide CRISPR/Cas9 knockout screening uncovers a novel inflammatory pathway critical for resistance to arginine-deprivation therapy. Theranostics 11, 3624–3641. https://doi.org/10.7150/thno.51795 (2021).
Burki, T. K. Arginine deprivation for ASS1-deficient mesothelioma. Lancet Oncol. 17, e423. https://doi.org/10.1016/S1470-2045(16)30446-6 (2016).
Weng, A. P. et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306, 269–271. https://doi.org/10.1126/science.1102160 (2004).
Hawkins, E. D. et al. T-cell acute leukaemia exhibits dynamic interactions with bone marrow microenvironments. Nature 538, 518–522. https://doi.org/10.1038/nature19801 (2016).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Yang, M. & Vousden, K. H. Serine and one-carbon metabolism in cancer. Nat. Rev. Cancer. 16, 650–662. https://doi.org/10.1038/nrc.2016.81 (2016).
Lee, S. E. et al. Unraveling the role of the mitochondrial one-carbon pathway in undifferentiated thyroid cancer by multi-omics analyses. Nat. Commun. 15, 1163. https://doi.org/10.1038/s41467-024-45366-0 (2024).
Zeng, J. D., Wu, W. K. K., Wang, H. Y. & Li, X. X. Serine and one-carbon metabolism, a Bridge that links mTOR signaling and DNA methylation in cancer. Pharmacol. Res. 149, 104352. https://doi.org/10.1016/j.phrs.2019.104352 (2019).
Ye, M. et al. Therapeutic targeting nudix hydrolase 1 creates a MYC-driven metabolic vulnerability. Nat. Commun. 15, 2377. https://doi.org/10.1038/s41467-024-46572-6 (2024).
Sun, S. et al. Constitutive activation of mTORC1 in endothelial cells leads to the development and progression of lymphangiosarcoma through VEGF autocrine signaling. Cancer Cell. 28, 758–772. https://doi.org/10.1016/j.ccell.2015.10.004 (2015).
Chantranupong, L. et al. The CASTOR proteins are arginine sensors for the mTORC1 pathway. Cell 165, 153–164. https://doi.org/10.1016/j.cell.2016.02.035 (2016).
Shi, J. et al. CAR T cells targeting CD99 as an approach to eradicate T-cell acute lymphoblastic leukemia without normal blood cells toxicity. J. Hematol. Oncol. 14, 162. https://doi.org/10.1186/s13045-021-01178-z (2021).
Zhang, R. et al. Regulation of Urea cycle by reversible high-stoichiometry lysine succinylation. Nat. Metab. 6, 550–566. https://doi.org/10.1038/s42255-024-01005-y (2024).
Gurung, S. et al. mRNA therapy corrects defective glutathione metabolism and restores ureagenesis in preclinical argininosuccinic aciduria. Sci. Transl Med. 16, eadh1334. https://doi.org/10.1126/scitranslmed.adh1334 (2024).
Huang, H. Y. et al. ASS1 as a novel tumor suppressor gene in myxofibrosarcomas: aberrant loss via epigenetic DNA methylation confers aggressive phenotypes, negative prognostic impact, and therapeutic relevance. Clin. Cancer Res. 19, 2861–2872. https://doi.org/10.1158/1078-0432.CCR-12-2641 (2013).
Qiu, F. et al. Arginine starvation impairs mitochondrial respiratory function in ASS1-deficient breast cancer cells. Sci. Signal. 7, ra31. https://doi.org/10.1126/scisignal.2004761 (2014).
Hu, Q. et al. ASS1-Mediated reductive carboxylation of cytosolic glutamine confers ferroptosis resistance in cancer cells. Cancer Res. 83, 1646–1665. https://doi.org/10.1158/0008-5472.CAN-22-1999 (2023).
Gai, X. et al. Oncogenic KRAS induces arginine auxotrophy and confers a therapeutic vulnerability to SLC7A1 Inhibition in Non-Small cell lung cancer. Cancer Res. 84, 1963–1977. https://doi.org/10.1158/0008-5472.CAN-23-2095 (2024).
Saxton, R. A., Chantranupong, L., Knockenhauer, K. E., Schwartz, T. U. & Sabatini, D. M. Mechanism of arginine sensing by CASTOR1 upstream of mTORC1. Nature 536, 229–233. https://doi.org/10.1038/nature19079 (2016).
Acknowledgements
We are obliged to Jinqiang Liang (School of Pharmaceutical Sciences, Hainan University) for their assistance in the CDX experiments.
Funding
This work was funded by the the Hainan Province Postdoctoral Research Project Funding QRCTJ2024-6-80 to Zijian Zhang, the Natural Science Foundation Youth Category A of Hubei Province JCZRJQ202500008 and the Natural Science Foundation of Chongqing CSTB2024NSCQ-MSX0066 to Haichuan Zhu.
Author information
Authors and Affiliations
Contributions
H.Z., and J.L. designed the experimental plans; Z.Z., X.L., J.L., Y.Z., Y.W., Q.Y. performed the experiments. Q.W., and J.D. performed the bioinformatic analysis. Z.Z., and H.Z. analyzed the data and drafted the manuscript. H.Z., and J.L. designed and supervised the study. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Z., Liu, X., Wu, Q. et al. ASS1 facilitates T-ALL progression via the arginine-mediated mTORC1/c-Myc signaling pathway. Sci Rep 15, 43783 (2025). https://doi.org/10.1038/s41598-025-27576-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-27576-8