Abstract
Acne, the most prevalent inflammatory skin condition globally, lacks comprehensive understanding of its cellular and molecular mechanisms. This research integrates single-cell transcriptomics with spatial immunofluorescence to detail cellular composition and functional states in healthy skin, mild acne, and severe acne. Findings indicate that keratinocytes in mild acne secrete antimicrobial peptides and maintain barrier defense, but transition to abnormal differentiation and pathological keratinization in severe acne. Natural killer (NK) cells initially amplify inflammation and activate the immune response, but later become exhausted and apoptotic. This interaction creates a “dual immune-barrier imbalance,” driving acne progression. Functional enrichment and pseudo-time analyses demonstrated a strong association between disease severity and TNF, and chemokine signaling pathways, as well as NK cell-mediated cytotoxicity and Wnt/p53-apoptosis pathways. These findings were corroborated by immunofluorescence validation. This study introduces an innovative pathological model that establishes theoretical foundations for early diagnosis and precision interventions, while also outlining future therapeutic strategies centered on “immune-barrier co-regulation.” The findings provide essential insights into the pathogenesis of acne and open avenues for investigating novel intervention targets.
Similar content being viewed by others
Introduction
Acne is a highly prevalent and intricate chronic inflammatory dermatological condition, affecting approximately 85% of adolescents1. Its clinical manifestations range from comedones and papules to pustules, nodules, and cysts, frequently accompanied by cutaneous lesions and psychological burdens such as anxiety and depression, which significantly diminish quality of life2. Although antimicrobial agents, systemic hormones, and retinoids may offer short-term symptom relief, they are characterized by high recurrence rates, limited efficacy duration, and are often associated with drug resistance and adverse reactions, underscoring significant limitations in long-term management3,4. The pathogenesis of acne involves the synergistic interaction of multiple factors, including aberrant sebum production, follicular keratinization disorders, colonization by Propionibacterium acnes, and immune-inflammatory responses5,6. However, the key regulatory mechanisms at the cellular and molecular levels remain inadequately understood, impeding a deeper comprehension of its pathological essence and the development of precise therapeutic strategies. Consequently, a systematic elucidation of the fundamental pathogenic processes and molecular networks underlying acne is essential. Such an approach not only enhances our understanding of the pathological mechanisms but also establishes a scientific basis for identifying novel therapeutic targets and developing personalized, long-term effective treatments. This endeavor constitutes a pivotal challenge necessitating urgent advancements within the field of dermatology.
In recent years, the pathogenesis of acne has been recognized to involve abnormal follicular keratinization, excessive sebum secretion, Propionibacterium acnes colonization, and host immune imbalance as primary drivers7. However, most studies have predominantly utilized histopathology or bulk omics approaches, which pose challenges in elucidating single-cell-level heterogeneity and dynamic changes8. The advent of single-cell transcriptomics (scRNA-seq) technology has marked significant advancements in the study of the skin immune microenvironment9. Yang et al. employed scRNA-seq to analyze lesion tissue from six acne patients, identifying eight distinct cell subpopulations10. Their findings highlighted significant differences in immune cell populations between lesional and non-lesional sites, as well as the role of specific ligand-receptor pairs (e.g., SPP1 and IL6) in acne pathogenesis. Deng et al. utilized scRNA-seq and spatial transcriptomics to uncover substantial alterations in intercellular communication within early-stage acne lesions, revealing abnormalities in multiple signaling pathways11. Despite these advancements, the application of scRNA-seq in acne research remains extremely limited, with the intricate characteristics, functional states, and dynamic changes still requiring further exploration. The holistic concept of the “skin microenvironment” has gained significant attention, highlighting the dynamic interactions and ecological balance among various components, including immune cells, sebaceous glands, keratinocytes, and the skin microbiome12,13,14. Acne development arises not from abnormalities in a single cell type or pathway but from the disruption of this skin ecological microenvironment15. Comprehensive single-cell omics research not only uncovers the fundamental pathogenic mechanisms of acne and elucidates the intricate interactions between the immune microenvironment and other skin components but also provides a theoretical foundation for “resetting the skin ecological microenvironment.” This research holds critical and urgent scientific importance, driving the advancement of precise and sustainable therapeutic strategies.
In this study, we conducted a systematic analysis of the microenvironmental heterogeneity of healthy skin, mild acne, and severe acne tissues at the single-cell level, marking the first such investigation (Fig. 1A). Our findings highlighted significant variations in the proportions of NK cells and keratinocytes across different stages of the disease. Utilizing healthy samples as a baseline, we performed a longitudinal analysis of the heterogeneity of NK cells and keratinocytes as acne progressed from mild to severe stages. This analysis led to the identification of specific marker genes and functional states associated with acne. The results indicate that keratinocytes transition from a steady-state defense role to pathological differentiation as the disease progresses, while NK cells shift from immune activation to functional exhaustion. These observations suggest that an imbalance in cellular function is pivotal in the progression of acne. The findings not only enhance our understanding of acne pathogenesis but also establish a critical foundation for exploring potential intervention strategies.
Single-cell transcriptomic analysis reveals dynamic remodeling of immune-epithelial structures during acne progression. (A) Workflow for single-cell RNA sequencing from skin biopsies, including tissue dissociation, single-cell suspension preparation, library preparation, and sequencing. (B) UMAP plot showing 19 transcriptionally distinct clusters across all samples, each labeled with a unique color. (C) UMAP plot colored by major cell lineages, including keratinocytes, NK cells, fibroblasts, monocytes/macrophages, endothelial cells, B cells, and neutrophils. (D) Cell type composition cross-section illustrating cellular composition across NC, MA, and SA, highlighting NK cell enrichment in MA and keratinocyte expansion in SA. (E) Stage-specific UMAP plots (NC, MA, SA) revealing cellular trajectories from NK cell-driven immune activation to keratinocyte-driven pathological remodeling.
Results analysis
Single-Cell transcriptome profiling of the cellular landscape in acne lesions
To systematically characterize the cellular ecology of acne lesions, single-cell transcriptomic sequencing was conducted on six skin samples from patients with NC, MA, and SA. Utilizing dimensionality reduction and clustering analysis via Seurat, seven major cell types and 19 subpopulations were identified, including keratinocytes, NK cells, fibroblasts, monocytes/macrophages, B cells, endothelial cells, and neutrophils (Fig. 1BC). The progression of acne was associated with systematic remodeling of cellular composition and function16,17. The results (Fig. 1D) indicated a significantly elevated proportion of NK cells in mild lesions, suggesting their role in enhancing inflammation through immune activation and chemotactic signaling during the early stages of acne. In severe lesions, there was a notable expansion and abnormal differentiation of keratinocytes. Although NK cells were still present, they predominantly exhibited signs of functional exhaustion and apoptotic tendencies. Further cell trajectory analysis elucidated this dynamic succession process (Fig. 1E). The maintenance of barrier homeostasis in healthy skin is predominantly dependent on keratinocytes, with a minimal presence of immune cells18. However, in mild lesions, there is a notable infiltration of NK cells, which significantly enhance inflammatory signaling. As the condition progresses to a severe stage, there is a transition in the cellular environment from one dominated by immune cells to one characterized by the pathological expansion of structural cells. This stage is marked by aberrant keratinocyte differentiation and disorganization of the stratum, accompanied by a progressive decline in NK cell functionality. Keratinocytes and NK cells emerge as the most dynamically active cell populations, acting as central drivers of disease progression. This finding offers a comprehensive framework for understanding the pathogenesis and for identifying precise therapeutic targets.
Dynamic changes in keratinocytes
Single-cell transcriptomic analysis has uncovered substantial heterogeneity and dynamic evolution of keratinocytes during acne development19. Through unsupervised clustering, 15 distinct functional subgroups of keratinocytes were identified (Fig. 2AB), exhibiting significant compositional differences across samples at various developmental stages. In NC samples, subgroups 0, 1, and 12 were predominantly enriched, indicative of differentiated and mature populations that maintain barrier function under homeostatic conditions (Fig. 2C). In contrast, MA samples specifically showed enrichment of subgroups 5 and 8, characterized by elevated expression of the defensin gene DEFB4A (Fig. 2DE). This suggests that keratinocytes activate the immune barrier via antimicrobial peptide secretion during the mild stage, thereby adopting an “immune defense” phenotype. In SA samples, there was a marked expansion of subgroups 2, 3, 4, and 14, accompanied by abnormally high expression of structural proteins such as KRT1, KRT10, and KRT2 (Fig. 2F-I). This indicates a tendency towards pathological differentiation and epidermal structural instability in the severe stage. The findings indicate a biphasic evolutionary pattern in the progression of KC, marked by a transition from an “immunodefense-dominant” phase to a “pathological keratinization” phase.
Single-cell transcriptomics reveals keratinocyte heterogeneity and dynamic succession during acne progression. (A) UMAP of all keratinocytes, showing 15 transcriptionally defined keratinocyte subclusters (C0-C14). Numeric labels 0–14 on the UMAP correspond directly to these subcluster IDs. (B) Stage-stratified UMAPs for NC, MA and SA, illustrating the distribution of keratinocyte subclusters C0-C14 in each clinical stage. (C) Proportional composition of each subpopulation across different stage samples. (D) Venn diagram summarizing the overlap of marker genes between keratinocyte subclusters C5 and C8. (E) Spatial distribution and violin plot of antimicrobial peptide gene DEFB4A in UMAP. (F) Venn diagram showing the overlap of marker genes among keratinocyte subclusters C2, C3, C4 and C14. (G–I) Spatial distribution and violin plots of structural protein genes KRT1, KRT10, and KRT2.
Further analysis of differentiation trajectories offers dynamic validation for the previously mentioned conclusions (Fig. 3A-C). The differentiation process of KC can be delineated into two principal pathways (Fig. 3B). To further elucidate the dynamic biological changes accompanying KCs differentiation, we integrated GO enrichment results of upregulated and downregulated genes along pseudotime. Based on the GO enrichment results of both upregulated and downregulated genes (Figure S1A, S1B), the differentiation of KCs follows a continuous dynamic process that progresses from active proliferation to protein homeostasis regulation and finally to structural and functional specialization. The initial state S1 is characterized by the upregulation of translation and protein synthesis pathways, accompanied by the downregulation of stress response, migration, and signaling processes, indicating a metabolically active preparatory phase for cellular homeostasis. Upon entering S2, the upregulation of protein folding and chaperone-related pathways suggests that cells undergo prominent protein homeostasis regulation and stress adaptation during the early differentiation phase, representing a critical transition point toward multiple differentiation trajectories. Subsequently, differentiation bifurcates into distinct paths:
Single-cell differentiation trajectory inference reveals dynamic remodeling of keratinocyte states during acne progression. (A) Pseudo-time differentiation trajectories of keratinocytes in NC, MA, and SA samples. (B) Two pathways of KC differentiation. (C) Proportional composition of five differentiation states (S1–S5) in NC, MA, and SA samples. (D) Distribution of subpopulations along pseudo-time trajectories. (E) State-specific trajectory trees for representative subpopulations (C2, C3, C4, C14).
Along the S2→S3/S4 branch, upregulated genes are enriched in epidermal development, keratinization, and cytoskeletal organization, while downregulated pathways involve signaling and migration, reflecting the gradual loss of cellular plasticity and progression toward morphological maturation and barrier formation.
In contrast, the independent S1→S5 trajectory is characterized by the upregulation of genes associated with epidermal barrier establishment and immune defense, along with suppression of proliferation- and stress-related genes, indicating a direct differentiation toward functional keratinocytes with antimicrobial and barrier properties. Overall, the evolution of KCs reveals a unidirectional differentiation trend from metabolic activity to functional maturation, with a pivotal regulatory transition in protein homeostasis and two distinct terminal outcomes-a structural remodeling type and an immune-barrier type of keratinocyte specialization. As illustrated in Fig. 3D-E, KCs in MA samples predominantly expanded along the immune activation pathway, demonstrating enhanced defensive capabilities. In contrast, KCs in SA samples primarily remained in low-differentiation or abnormally differentiated states, which not only impaired barrier function but also contributed to the disruption of epidermal stratification. This altered differentiation trajectory suggests that the reorientation of KC functions and the imbalance of their states are central drivers in the progression of acne from mild to severe forms. The transition from immune barrier function to the amplification of pathological keratinization represents a crucial mechanism in the cellular ecological restructuring associated with acne.
Dynamic changes in NK cells
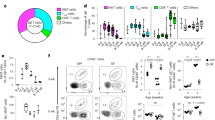
The analysis of single-cell transcriptomics has uncovered substantial heterogeneity and dynamic progression in NK cell populations throughout the course of acne20. Through unsupervised clustering, nine distinct NK cell subpopulations were identified, each displaying varied distribution across NC, MA, and SA samples (Fig. 4AB). In NC samples, NK cells were found to be limited in number and primarily exhibited a quiescent phenotype (Fig. 4C). Conversely, MA samples demonstrated a specific enrichment of subpopulations 0 and 8, which were characterized by a high expression of genes associated with defense and inflammatory responses, such as KRT6A, KRT16, KRT17, S100A7, S100A8, S100A9, and SERPINB4 (Fig. 4D-K). This suggests that in the mild disease phase, NK cells display a unique inflammatory-activation profile, characterized by the significant upregulation of KRT6A, KRT16, KRT17, S100A7, S100A8, S100A9, and SERPINB4, which together enhance epithelial stress responses and local inflammatory signaling (Fig. 4E-G). In contrast, SA samples predominantly featured NK cells enriched in Subpopulation 4 (Fig. 4HK), which exhibited transcriptional signatures suggestive of apoptosis and functional exhaustion, indicating a progressive decline in immune activity.
Single-cell transcriptomics reveal NK cell subpopulation heterogeneity and their dynamic changes in the acne process. (A) UMAP plot showing unsupervised clustering of all NK cells, identifying 9 functional subpopulations. (B) Distribution of NK cells across different staging samples. (C) Proportional composition of each NK cell subset in NC, MA, and SA samples. (D) Venn diagram revealing the intersection and differences in NK cell-specific genes under different conditions. (E–J) Spatial distribution and violin plots of epithelial barrier protein genes KRT6A, KRT16, KRT17, and inflammation/injury-associated molecules S100A7, S100A8, S100A9, and SERPINB4.
The application of pseudo-temporal trajectory inference has elucidated the functional transitions of NK cells throughout the progression of acne (Fig. 5A-D). Two principal differentiation trajectories for NK cells were identified (Fig. 5B). GO enrichment analysis across NK cell states revealed three major differentiation trajectories in acne skin, each associated with distinct functional states (Figure S1C, S1D). The first trajectory (S1→S2) represents the transition from a resting to a metabolically activated state, enriched in pathways related to energy metabolism, protein synthesis, and signal perception. This phase likely reflects the rapid recruitment and activation of NK cells in response to sebaceous gland inflammation and microbial stimulation. The second trajectory (S1→S3→S4→S5/S6) indicates a progressive differentiation toward classical cytotoxic effector NK cells, characterized by enrichment in pathways associated with degranulation, migration, and cytotoxic processes, suggesting their involvement in target cell clearance and amplification of local inflammation. The third trajectory (S1→S3→S7→S8→S11) corresponds to the differentiation toward tissue-resident and immunoregulatory NK cells, enriched in cytokine signaling, immune regulation, and tissue homeostasis pathways. These cells are likely involved in limiting excessive inflammation and promoting post-inflammatory tissue repair. In samples from MA, NK cells predominantly expand along the immune activation trajectory, thereby contributing to inflammatory amplification and immune barrier stress (Fig. 5E). Conversely, samples from SA are characterized by NK cells residing within the apoptotic subpopulation, which is indicative of immune effector exhaustion and subsequent cell death (Fig. 5F). These observations suggest that NK cells serve as amplifiers of inflammatory processes in the early stages of acne, ultimately evolving into a hallmark population indicative of immune failure in the later stages. Furthermore, this transition suggests that the preservation of natural killer (NK) cell effector molecules, such as NKG7 and GNLY, may be a critical strategy for preventing NK cell dysfunction during the course of the disease21,22.
Single-cell differentiation trajectory inference reveals bifurcated fate and state transitions of NK cells during acne progression. (A) Overall differentiation trajectories of NK cells. (B) Two primary NK cell differentiation pathways: activation-directed branch and normal homeostasis branch. (C) Trajectory distribution in NC, MA, and SA samples, showing significant differences in NK cell differentiation direction across disease stages. (D) Proportional composition of 11 differentiation states (S1–S11) in NC, MA, and SA samples. (D) Pseudo-time trajectories of NK cell subpopulations (0–8). (E–F) State-specific trajectory trees for representative NK cell subpopulations (N0 and N8).
Functional enrichment analysis reveals key signaling pathways
Functional enrichment and cell communication analyses have elucidated the dynamic remodeling of signaling interactions between NK cells and keratinocytes throughout the progression of acne23,24. The results, as illustrated in Fig. 6, indicate that during the mild stage of acne (MA vs. NC), NK cells exhibit significant immune activation, as evidenced by the upregulation of NK cell-mediated cytotoxicity and cytokine-receptor interaction pathways (Fig. 6CD). Concurrently, activation of the tumor necrosis factor (TNF) and chemokine pathways, along with increased expression of genes such as DEFB4A and S100A8/9 (Fig. 6E-H). Further analysis of communication patterns reveals (Fig. 6A) that the interaction from NK0 to KC5 primarily relies on the HSP90AA1–FGFR3 and B2M–TFRC pathways, whereas the interaction from NK0 to KC8 is dependent on HSP90AA1–EGFR and B2M–TFRC. In contrast, the interaction from NK8 to KC5 establishes robust connections through B2M–TFRC, HSP90AA1–EGFR, and RTN4–GJB2 pathways. These signaling pathways synergistically enhance the activation of keratinocyte receptors, regulate iron metabolism, and strengthen barrier junctions, thereby accelerating local inflammatory amplification and establishing an “immune-barrier synergy” pattern25,26. NK cells progressively accumulate within the p53 and apoptosis pathways, exhibiting signs of functional exhaustion (Fig. 6CD). The keratinocyte pathway lineage shifts towards keratinization and Wnt signaling, accompanied by the abnormal overexpression of KRT1/KRT10, which indicates pathological keratinization and barrier dysfunction (Fig. 6E-H). Correspondingly, communication networks significantly contract (Fig. 6B), retaining only a limited number of pathways: NK0 to KC2 (HSP90AA1–EGFR), NK8 to KC4 (LAMA2–RPSA, B2M–TFRC), NK8 to KC2 (B2M–TFRC, HSP90AA1–FGFR3), and NK8 to KC14 (HSP90AA1–EGFR). Residual iron metabolism and growth factor signaling suggest a reduction in NK cell-mediated effects, whereas pathological proliferation of keratinocytes and matrix remodeling increasingly predominate. Collectively, these findings elucidate the progression of acne from an “NK-mediated inflammatory amplification” phase to a dual-driven “immune-structural imbalance” state. This state is characterized by “NK cell exhaustion and pathological keratinization of keratinocytes,” offering crucial molecular insights into the developmental mechanisms of the condition.
Functional enrichment and cell-cell communication analysis reveal dynamic remodeling of NK cell-keratinocyte interactions during acne progression. (A) String diagram of cell-cell communication between NK cell and keratinocyte subpopulations in mild acne lesions. (B) String diagram of cell-cell communication between NK cells and keratinocyte subpopulations in severe acne lesions. (C–D) Pathway enrichment analysis of N0 (C) and N8 (D) (MA vs. NC). (E–H) Pathway enrichment analysis of KC2(E), KC3(F), KC4(G), and KC14 (H) (MA vs. NC). The figure is from the KEGG database (www.kegg.jp/kegg/kegg1.html) [Map ID: e.g., hsa00190] and is used with permission from Kanehisa Laboratories.
Immunofluorescence verification
Immunofluorescence staining and quantitative analysis demonstrated a dynamic imbalance between NK cell effector molecules and keratinocyte differentiation markers during the progression of acne, both spatially and at the molecular level24,27. Based on its superior cell specificity, direct functional association, and technical reliability, defining NK cells using immunofluorescence with NKG7 is a rigorous and efficient strategy. NKG7 is stably and highly expressed in all functional NK cell subsets, effectively avoiding missed detections caused by heterogeneity or downregulation of traditional surface markers (such as CD56/CD16). Furthermore, as a key component of cytotoxic granules, its expression directly identifies effector cells with cytotoxic potential, achieving a unification of phenotype and function. In practice, the intracellular protein characteristics of NKG7 enable it to generate clear and stable signals in tissue samples28,29. Therefore, NKG7 is not only a feasible NK cell marker but also a superior functional definition standard compared to traditional markers. In the MA stage, there was a significant upregulation of KRT6A and NKG7 signaling (Fig. 7A), suggesting that NK cells were in a state of heightened immune activation. This activation contributed to the amplification of the local inflammatory response through the release of cytotoxic molecules and interactions with keratinocytes. Further quantitative analysis revealed that KRT6A expression peaked during the MA stage, whereas NKG7, despite being markedly elevated in MA, exhibited a significant decline in SA (Fig. 7BC). This pattern indicates that NK cells transition from an “early immune amplification” phase to a “late functional exhaustion” phase. In relation to keratinocyte differentiation markers, KRT10 and Involucrin demonstrated a significant increase as early as the MA stage, suggesting accelerated and aberrant barrier repair responses (Fig. 8A). As the condition progressed to the SA stage, these markers exhibited further upregulation, albeit with disrupted spatial distribution (Fig. 8BC), indicative of keratinocytes being arrested in abnormal differentiation and pathological keratinization. This phased transition from “immune activation to barrier remodeling” is consistent with single-cell transcriptomics data, which reveal an enrichment of chemokine/cytokine pathways in MA samples and enhanced keratinization/epidermal differentiation pathways in SA samples. This highlights the critical trajectory of acne progression from an inflammation-driven phase to one characterized by structural imbalance. Consequently, maintaining NK effector function while simultaneously correcting keratinocyte differentiation appears to offer greater biological plausibility and translational potential compared to anti-inflammatory therapy alone.
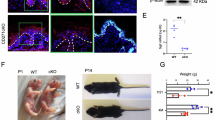
Immunofluorescence validation of keratinocyte immune activation and NK cell exhaustion during acne progression. (A) Immunofluorescence staining of KRT6A and NKG7 in NC, MA, and SA skin tissues. Scale bar, 50 μm. (B) Quantitative analysis of KRT6A fluorescence intensity across groups. (C) Quantitative analysis of NKG7 fluorescence intensity in each group. (Data are expressed as mean ± standard deviation, n = 3, *p < 0.05, **p < 0.01).
Immunofluorescence validation of keratinocyte differentiation defects in acne progression. (A) Immunofluorescence staining for KRT10 and Involucrin in NC, MA, and SA skin tissues. Scale bar, 50 μm. (B) Quantitative analysis of KRT10 fluorescence intensity across groups. (C) Quantitative analysis of Involucrin fluorescence intensity in each group. (Data are expressed as mean ± standard deviation, n = 3, *p < 0.05, **p < 0.01).
Conclusion
This study presents a comprehensive framework for single-cell transcriptomic analysis and spatial validation, reconceptualizing acne as a condition characterized by a “dual immune-barrier imbalance” rather than merely an inflammatory disorder. By elucidating the dynamic progression of keratinocytes and NK cells across healthy, mild, and severe stages of acne, we demonstrate a sequential transition from immune activation to immune exhaustion. This transition is accompanied by a pathological shift from antimicrobial defense to aberrant keratinization. Our findings extend previous theories that predominantly focused on Cutibacterium acnes-induced inflammation and sebaceous gland hyperactivity, proposing instead that acne progression signifies a disruption of immune–epithelial homeostasis. Mechanistically, the enrichment of TNF and chemokine pathways in keratinocytes suggests that innate immune activation precedes epidermal remodeling. Concurrently, the accumulation of NK cells within p53 and apoptosis pathways indicates a gradual decline in cytotoxic function. This reciprocal dysregulation establishes a self-amplifying feedback loop, wherein keratinocyte-derived cytokines recruit and activate NK cells, and the activated NK cells subsequently induce keratinocyte stress and cornification, thereby perpetuating inflammation and structural degradation. From a translational perspective, these findings suggest that conventional anti-inflammatory or antimicrobial therapies may be inadequate for long-term management. Instead, dual-modulatory strategies that aim to preserve NK cell activity while normalizing keratinocyte differentiation could potentially offer more sustained remission. Furthermore, markers such as NKG7, KRT6A, and KRT10 may serve as stage-specific indicators for disease stratification and treatment evaluation. Although the limited cohort size and absence of mechanistic validation present limitations, future integration with organoid models, spatial multi-omics, and longitudinal analyses will further elucidate causal cellular interactions and facilitate predictive modeling for personalized acne management.
This study systematically delineated the cellular ecological restructuring of acne lesions by integrating single-cell transcriptomics with spatial immunofluorescence validation. The findings elucidate critical fate transitions of keratinocytes and NK cells throughout disease progression. Initially, keratinocytes exhibit antimicrobial and barrier defense functions, but as the disease advances to severe stages, they undergo a shift towards abnormal differentiation and pathological keratinization. In the early stages, NK cells contribute to inflammatory amplification; however, as the disease progresses, they increasingly experience effector exhaustion and apoptosis. The aberrant interaction between these two cell types emerges as the central mechanism driving the transition from mild to severe acne, delineating a pathological trajectory characterized by a “dual imbalance of immunity and barrier function.”
This discovery not only addresses a critical gap in our understanding of the dynamic mechanisms governing immune-barrier interactions in acne at a single-cell resolution, but also introduces a novel pathological model30. Specifically, it suggests that the dysregulation of keratinocyte function, in conjunction with the exhaustion of NK cells, collectively drives persistent inflammation and structural imbalance. This mechanism surpasses traditional frameworks that primarily focus on sebum secretion or bacterial colonization, underscoring the pivotal role of cellular functional evolution in the progression of acne. From a translational perspective, the study proposes that preserving NK cell functional activity while concurrently correcting keratinocyte differentiation may constitute a dual-target intervention strategy to arrest acne progression10,31,32. Moreover, the molecular signatures of keratinocytes and NK cells emerge as promising biomarkers for diagnostic staging and predicting treatment efficacy, thereby laying the groundwork for personalized therapeutic approaches.
In acne lesions, we identified a distinct NK-cell subset (C0 and C8) that expresses the keratin protein KRT6A. Given that KRT6A is a classical epithelial cytoskeletal protein, its presence in NK cells is highly unexpected and suggests potential non-canonical biological mechanisms33. Based on our current data, we considered two major possibilities.
The first is the passive acquisition hypothesis, in which NK cells obtain pre-existing KRT6A protein from apoptotic or necrotic follicular keratinocytes through cell fusion or efferocytosis. Although such phenomena have been documented in inflamed and injured tissues, they are typically accompanied by a marked increase in mitochondrial gene proportion (mt genes) and a higher number of detected genes (nFeature_RNA) in the phagocytic cells (Figure S1FG). In contrast, our analysis showed that the C0 and C8 NK subsets do not exhibit elevated mt-gene percentages or nFeature_RNA values compared with other NK subsets, and therefore lack the characteristic signatures of phagocytosis. These findings do not support the passive acquisition model.
Instead, our results are more consistent with a microenvironment-driven, active inflammatory reprogramming mechanism. Proinflammatory cytokines enriched in acne lesions (such as IFN-γ and IL-1β) may induce epigenetic remodeling in NK cells, altering their transcriptional program and releasing the repressive control over genes like KRT6A that are normally silenced in resting NK cells, thereby enabling de novo expression. Moreover, high-resolution immunofluorescence provided critical validation: the NK-cell marker NKG7 colocalized clearly with KRT6A and the nuclear dye DAPI at the single-cell level. This observation rules out the possibility of surface adherence or attached cellular debris and strongly supports KRT6A as an endogenously produced protein in NK cells.
Functionally, such inflammation-induced reprogramming may have important adaptive significance. KRT6A is a stress-responsive keratin that enhances cytoskeletal stability during epithelial tissue repair. By analogy, in the highly inflamed and injury-prone microenvironment of acne lesions, KRT6A expression may help NK cells maintain structural integrity, thereby facilitating their migration, tissue residency, or cytotoxic function under high-stress conditions. This finding highlights an unexpected degree of plasticity in NK cells within chronic inflammatory environments and provides new insights into the adaptive remodeling of innate lymphocytes in cutaneous inflammatory disease.
It is important to note that although our transcriptomic analysis and high-magnification immunofluorescence observations both demonstrate spatial overlap between KRT6A and NKG7 in regions enriched with NK cells, this finding does not eliminate the possibility that KRT6A signals may originate from adjacent keratinocytes. The presence of a large number of keratinocytes with high KRT6A expression in the epidermis could contribute to significant background staining, potentially masking the relatively weak intrinsic expression within NK cells. Consequently, caution is warranted when interpreting intrinsic KRT6A expression in NK cells. Future studies should consider incorporating additional NK-specific markers, such as CD56 or NKp46, for multicolor immunofluorescence or employing spatial transcriptomics techniques to more accurately differentiate genuine NK cell expression from keratinocyte background signals.
Consequently, this study enhances our comprehension of acne pathogenesis and offers novel theoretical foundations for the development of precision intervention strategies. Future investigations into comprehensive treatment models grounded in “immune-barrier synergistic regulation” are promising for transitioning acne management from empirical approaches to mechanism-driven precision medicine.
Materials and methods
Ethics
This study was approved by the Ethics Committee of China-Japan Union Hospital of Jilin University (Ethics Approval No.: 2024112111). Written informed consent was obtained from all participants. All methods were conducted in accordance with relevant guidelines and regulations.
Study population and sample collection
In this study, six participants were recruited from the Department of Dermatology at the China-Japan Union Hospital of Jilin University (Table 1). These participants were categorized into three distinct groups: healthy control (NC, n = 2), mild acne (MA, n = 2), and severe acne (SA, n = 2). The recruitment process adhered to the Global Acne Grading System (GAGS) to ensure standardized classification. Mild acne was defined by the presence of papules and comedones, with a total GAGS score of less than 18 and a lesion duration of less than 24 h. In contrast, severe acne was characterized by the presence of pustules, nodules, or cystic lesions, with a GAGS score exceeding 30 and a lesion duration of more than 72 h. All participants were between the ages of 18 and 30 years, had not received systemic antibiotics, glucocorticoids, or immunomodulators within four weeks prior to sampling, and were free from other inflammatory or autoimmune skin conditions. Individuals with endocrine disorders, systemic infections, or pregnancy were excluded from the study. Skin biopsies, measuring 2–4 mm, were collected under sterile conditions and immediately processed for single-cell RNA sequencing and immunofluorescence validation. To reduce temporal bias, samples of mild acne were collected within 24 h of papule formation, while samples of severe acne were obtained within 72 h following the development of pustules or nodules. Despite the limited cohort size, each sample yielded over 5,000 high-quality single-cell transcriptomes, facilitating a robust analysis of cellular heterogeneity and signaling dynamics.
To reduce inter-individual variability and mitigate batch effects due to the limited number of donors, all samples were processed under uniform experimental and sequencing conditions at the same facility. Tissue dissociation, enzymatic digestion, and single-cell library construction were conducted concurrently using standardized reagents and protocols. During data processing, donor-specific batch effects were addressed using the Seurat v5 integration workflow, which employs canonical correlation analysis (CCA) and mutual nearest neighbor (MNN) anchoring. This methodology aligns shared cellular states across individuals, facilitating the identification of biologically relevant transcriptional differences rather than those specific to individual donors. Furthermore, clustering and differential expression analyses were conducted post-integration to ensure that the observed variations were indicative of disease-associated changes rather than artifacts arising from inter-donor variability.
Single-Cell suspension Preparation and library sequencing
Samples intended for single-cell sequencing were initially placed in ice-cold Hank’s Balanced Salt Solution (HBSS) supplemented with 1% bovine serum albumin (BSA). Subsequent to mechanical disruption, the samples were treated with a combination of collagenase I/IV (1–2 mg/mL), Dispase (1 U/mL), and DNase I (50 U/mL), and subjected to gentle enzymatic digestion at 37 °C for a duration of 20–40 minutes. The resulting digested material was passed through a 70 µm filter to eliminate debris/aggregates removal, and cell viability was assessed to ensure a viability rate of at least 80%. The single-cell suspensions were then prepared for library construction using the 10x Genomics Chromium platform (Single Cell 3’ v3.1) and sequenced on the Illumina NovaSeq 6000 platform, with a target sequencing depth of no less than 50,000 reads per cell.
Data processing and quality control
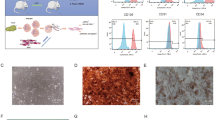
The raw data were aligned to the hg19 reference genome utilizing Cell Ranger (version 7.0, 10x Genomics). We performed single-cell RNA sequencing data analysis using Seurat (v5.0). Initial quality control retained cells with 500–4000 detected genes and ≤ 10% mitochondrial gene content. Doublets were removed using function DoubletFinder with a 5% expected rate and a pK value of 0.09, which was empirically optimized via artificial doublet simulations. To partition the cells into distinct clusters, graph-based clustering was performed on the constructed shared nearest-neighbor graph using the FindClusters function. A resolution parameter of 0.5 was applied to control the granularity of the downstream clustering output. Cell types were defined for the resulting clusters by integrating computational predictions with biological knowledge (Figure S1E). The SingleR package, using the HumanPrimaryCellAtlasData reference, provided an automated preliminary annotation. These predictions were then evaluated and refined based on the expression of canonical marker genes. This integrated approach ensured that the final cell type labels were robust and biologically meaningful.
The gene expression profiles of NK and KC cells were preprocessed through a standardized workflow, which included data normalization using the NormalizeData algorithm, identification of highly variable genes via the FindVariableFeatures method, and data scaling with the ScaleData function, with all parameters consistent with the previously described single-cell gene expression preprocessing steps. Utilizing the FindClusters function with a resolution parameter of 0.5 to delineate cell subpopulations.
Differential gene and functional enrichment analysis
Differentially expressed genes were identified through the application of the Wilcoxon signed-rank test, employing a threshold of |log2FC| > 0.25 and an adjusted p-value of less than 0.05. Functional enrichment analysis was conducted using clusterProfiler (version 4.6) for Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis. The results were visualized using bubble plots and bar charts34,35,36. Subsequently, key differentially regulated pathways in keratinocytes and natural killer (NK) cells were further analyzed for pseudo-time and intercellular communication studies.
Trajectory inference and cell communication analysis
Single-cell trajectory analysis was conducted utilizing Monocle2 (version 2.28.0). To construct pseudo-temporal trajectories, the top 3,000 temporally differentiated genes (TDGs) with a q-value less than 0.01 were selected. Intercellular communication networks were inferred using iTALK (version 1.5.0), with particular emphasis on interactions between keratinocytes and immune cells, as well as between natural killer (NK) cells and macrophages.
Immunofluorescence assay
Each group contains 2 independent case samples, with 3 random fields of view or sections taken from each case for quantitative analysis. The samples were initially fixed in 4% paraformaldehyde for a duration of 4 to 12 h, followed by dehydration using 30% sucrose. Subsequently, the samples were embedded and sectioned into cryosections with a thickness of 8 to 10 μm. The sections underwent a blocking process with 5% goat serum and were then incubated with primary antibodies, including Involucrin, KRT10, KRT6A, and NKG7. On the following day, the sections were incubated with Alexa Fluor-conjugated secondary antibodies, and counterstaining was performed using DAPI. The primary antibodies employed were KRT6A (Cat: ab238013), KRT10 (Cat: ab76318), Involucrin (Cat: MA5-11803), NKG7 (Cat: PA5-144131), and KRT14 (Cat: ab7800). All antibodies were validated by the manufacturers for immunofluorescence (IF) on human tissue. Additionally, single-antibody and no-primary controls were implemented to eliminate the possibility of nonspecific staining. Imaging was conducted utilizing a laser confocal microscope equipped with objective lenses ranging from 40× to 63× magnification. Image quantification was subsequently carried out using ImageJ and QuPath software.
Statistical analysis
All data were analyzed using R (v4.3) and GraphPad Prism (v9.0). Independent samples t-tests were performed for comparisons between two groups, while one-way analysis of variance (ANOVA) with FDR correction was used for comparisons among multiple groups. Differences were considered statistically significant at p < 0.05.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2025) in National Genomics Data Center (Nucleic Acids Res 2025), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA-Human: HRA008189) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa-human.
References
Li, D. G. et al. Evaluation of Point-of-Care decision support for adult acne treatment by primary care clinicians. Jama Dermatology. 156, 538–544. https://doi.org/10.1001/jamadermatol.2020.0135 (2020).
Bilal, H. et al. Stabilization of acne Vulgaris-Associated microbial dysbiosis with 2% supramolecular Salicylic acid. Pharmaceuticals 16 https://doi.org/10.3390/ph16010087 (2023).
Wi, Y. M. et al. Use of antibiotics within the last 14 days of life in Korean patients: A nationwide study. J. Korean Med. Sci. 38 https://doi.org/10.3346/jkms.2023.38.e66 (2023).
Yu, Y. et al. Suppression of Cutibacterium acnes-Mediated inflammatory reactions by fibroblast growth factor 21 in skin. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms23073589 (2022).
Al-Mendalawi, M. D. Evaluation of serum levels of interleukins 6, 8, 17 and 22 in acne vulgaris: A Cross-Sectional study. Indian J. Dermatol. 69, 91–91. https://doi.org/10.4103/ijd.ijd_625_23 (2024).
Perche, P. O. et al. Prescribing trends for acne vulgaris visits in the united States. Antibiotics-Basel 12 https://doi.org/10.3390/antibiotics12020269 (2023).
Zhang, J. et al. Acnes qPCR-Based antibiotics resistance assay (ACQUIRE) reveals widespread macrolide resistance in acne patients and can eliminate macrolide misuse in acne treatment. Front. Public. Health. 10 https://doi.org/10.3389/fpubh.2022.787299 (2022).
Yan, P. et al. Construction of the prognostic enhancer RNA regulatory network in osteosarcoma. Transl Oncol. 25 https://doi.org/10.1016/j.tranon.2022.101499 (2022).
Qin, Y. et al. Cuproptosis correlates with immunosuppressive tumor microenvironment based on pan-cancer multiomics and single-cell sequencing analysis. Mol. Cancer. 22 https://doi.org/10.1186/s12943-023-01752-8 (2023).
Yang, X., Wang, X. & Yang, J. Single-cell analysis reveals cellular heterogeneity, gene expression profiles, and pathway dynamics in acne vulgaris. Arch. Dermatol. Res. 317 https://doi.org/10.1007/s00403-025-03894-9 (2025).
Deng, M. et al. Analysis of intracellular communication reveals consistent gene changes associated with early-stage acne skin. Cell. Communication Signal. 22 https://doi.org/10.1186/s12964-024-01725-4 (2024).
Parrado, C. et al. Environmental stressors on skin Aging. Mechanistic insights. Front. Pharmacol. 10 https://doi.org/10.3389/fphar.2019.00759 (2019).
Cibrian, D., de la Fuente, H. & Sanchez-Madrid, F. Metabolic pathways that control skin homeostasis and inflammation. Trends Mol. Med. 26, 975–986. https://doi.org/10.1016/j.molmed.2020.04.004 (2020).
Ahn, S. et al. Biomimetic and Estrogenic fibers promote tissue repair in mice and human skin via Estrogen receptor β. Biomaterials 255 https://doi.org/10.1016/j.biomaterials.2020.120149 (2020).
Ivanova, K., Ramon, E., Ivanova, A., Sanchez-Gomez, S. & Tzanov, T. Bio-Based Nano-Enabled Cosmetic Formulations for the Treatment of Cutibacterium acnes-Associated Skin Infections. Antioxidants 12, (2023). https://doi.org/10.3390/antiox12020432
Popovich, N. G. Topical antibiotic therapy for acne. Am. Pharm. NS21, 55–58. https://doi.org/10.1016/s0160-3450(16)31328-9 (1981).
Peng, Y. M. et al. Application of Zizao Yangrong granules for treating targeted Drugs-Related skin xerosis: A randomized Double-Blinded controlled study. Integr. Cancer Ther. 19 https://doi.org/10.1177/1534735420924832 (2020).
Sumpter, T. L., Balmert, S. C. & Kaplan, D. H. Cutaneous immune responses mediated by dendritic cells and mast cells. Jci Insight. 4 https://doi.org/10.1172/jci.insight.123947 (2019).
Li, J. et al. Single-cell RNA-seq reveals actinic keratosis-specific keratinocyte subgroups and their crosstalk with secretory-papillary fibroblasts. J. Eur. Acad. Dermatol. Venereol. 37, 2273–2283. https://doi.org/10.1111/jdv.19289 (2023).
Gunaydin, S. & Tezcan, I. Evaluation of peripheral lymphocyte subsets in acne vulgaris patients before and after systemic isotretinoin treatment. Indian J. Pharmacol. 54, 338–344. https://doi.org/10.4103/ijp.ijp_695_21 (2022).
Myers, J. A. et al. Balanced engagement of activating and inhibitory receptors mitigates human NK cell exhaustion. Jci Insight. 7 https://doi.org/10.1172/jci.insight.150079 (2022).
Alvarez, M. et al. Regulation of murine NK cell exhaustion through the activation of the DNA damage repair pathway. Jci Insight. 4 https://doi.org/10.1172/jci.insight.127729 (2019).
Zhao, C. X., Wang, S. L., Li, H. X. & Li, X. Integration of Single-Cell transcriptomics data reveal differences in cell composition and communication in acne. Clin. Cosmet. Invest. Dermatology. 16, 3413–3426. https://doi.org/10.2147/ccid.S436776 (2023).
Huang, L. et al. Association of different cell types and inflammation in early acne vulgaris. Front. Immunol. 15 https://doi.org/10.3389/fimmu.2024.1275269 (2024).
Kim, E. O. et al. The Secretion of Inflammatory Cytokines Triggered by TLR2 Through Calcium-Dependent and Calcium-Independent Pathways in Keratinocytes. Mediators Inflamm. (2024). https://doi.org/10.1155/mi/8892514 (2024).
Bolla, B. S. et al. Cutibacterium acnes regulates the epidermal barrier properties of HPV-KER human immortalized keratinocyte cultures. Sci. Rep. 10 https://doi.org/10.1038/s41598-020-69677-6 (2020).
Petty, A. J. et al. Insights into keratinocyte and Immunologic landscape in cutaneous Graft-Versus-Host disease through Single-Cell transcriptomics. Jid Innovations. 5 https://doi.org/10.1016/j.xjidi.2025.100373 (2025).
Li, X. Y. et al. NKG7 is required for optimal antitumor T-cell immunity. Cancer Immunol. Res. 10, 154–161. https://doi.org/10.1158/2326-6066.Cir-20-0649 (2022).
Ng, S. S. et al. The NK cell granule protein NKG7 regulates cytotoxic granule exocytosis and inflammation. Nat. Immunol. 21, 1205–. https://doi.org/10.1038/s41590-020-0758-6 (2020).
Firlej, E., Kowalska, W., Szymaszek, K., Rolinski, J. & Bartosinska, J. The role of skin immune system in acne. J. Clin. Med. 11 https://doi.org/10.3390/jcm11061579 (2022).
Esen, F. et al. Functional and phenotypic changes in natural killer cells expressing immune checkpoint receptors PD-1, CTLA-4, LAG-3, and TIGIT in non-small cell lung cancer: the comparative analysis of tumor microenvironment, peripheral venous blood, and tumor-draining veins. Immunol. Res. 73 https://doi.org/10.1007/s12026-024-09573-7 (2025).
Kamata, M. & Tada, Y. Crosstalk: keratinocytes and immune cells in psoriasis. Front. Immunol. 14 https://doi.org/10.3389/fimmu.2023.1286344 (2023).
Liu, J. Y. et al. Epigenetic regulation of CD38/CD48 by KDM6A mediates NK cell response in multiple myeloma. Nat. Commun. 15 https://doi.org/10.1038/s41467-024-45561-z (2024).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2024).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M. & Goto, S. K. E. G. G. Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Acknowledgements
This work was supported by the Natural Science Foundation of Jilin Province (YDZJ202301ZYTS506).
Author information
Authors and Affiliations
Contributions
W.XH: Writing – original draft. G.CG: Methodology, Data curation. Y.DD and P.JH: Investigation. J.RH: Writing – review & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, X., Gao, C., Yang, D. et al. Single-cell transcriptomics and spatial validation reveal dysfunction of keratinocytes and NK cells driving acne development. Sci Rep 16, 4227 (2026). https://doi.org/10.1038/s41598-025-34373-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34373-w