Abstract
Malignant melanoma, a rare and aggressive skin cancer, arises from the transformation of cutaneous melanocytes and is associated with a poor prognosis. Evodiamine (EVO), a bioactive compound derived from traditional Chinese herbal medicine, has demonstrated significant inhibitory effects on various tumor cells. In this study, we aimed to investigate the potential of EVO in regulating melanoma immunity and elucidate its underlying mechanisms. Experimental results revealed that the IC50 value of EVO in B16-F10 cells for 24, 48, and 72 h were 11.73, 5.083, and 4.604 µM, respectively. EVO inhibited the proliferation, invasion, and metastasis of B16-F10 cells by more than 50%, while promoting apoptosis of higher concentration of EVO. EVO also significantly suppressed tumor growth by more than 80% and reduced spleen damage in tumor-bearing mice. Treatment with EVO led to a marked increase in T-cell subsets in the spleen, bone marrow, and tumors, with immunohistochemical (IHC) staining in particular showing about 50% higher. Furthermore, EVO inhibited the expression of programmed cell death ligand 1 (PD-L1) and the PI3K/AKT signaling pathway-related proteins in both B16-F10 cells and tumors. These findings suggest that EVO exerts antitumor effects by enhancing the tumor immune microenvironment and indicates its potential as a therapeutic agent for melanoma.
Similar content being viewed by others
Introduction
Cancer remains a major global health issue, significantly impacting human quality of life. Malignant melanoma, originating from or developing in melanocytes, is a particularly dangerous form of cancer1,2 with a high malignancy rate and early systemic metastasis. It is associated with a poor prognosis, with a five-year survival rate of less than 30%3,4. Conventional treatments for malignant melanoma include surgical resection, chemotherapy, immunotherapy, and targeted therapy5. Despite recent advances in immunotherapy using programmed cell death protein 1 (PD-1) / programmed cell death ligand 1 (PD-L1) inhibitors, over 50% of patients experience disease progression following an initial response6,7,8. This underscores the urgent need for novel immunosuppressants or immune adjuvants to enhance the immune microenvironment.
Evodiamine (EVO), a natural alkaloid extracted from the fruit of Evodia rutaecarpa, is a traditional Chinese medicinal ingredient9. EVO exhibits a range of functions, including anti-inflammatory and antidiarrheal effects, as well as treatment for headaches and skin diseases10. It also inhibits tumor cell proliferation, invasion, and migration, while promoting apoptosis through various pathways11,12,13, showing promising antitumor effects. Above all, some studies have highlighted the potential of EVO for the treatment of human melanoma A-375 cells. On the one hand, EVO induces intrinsic apoptosis, necroptosis and cell cycle arrest via ROS activation14,15,16. On the other hand, EVO initiates caspase-mediated a typical apoptosis pathway at the early stage and caspase-independent pathways such as MAPKs-mediated necrosis at the later stage17,18,19,20.
PD-1 and its primary ligand, PD-L1, are critical immunosuppressive checkpoints. Abnormal expression of the PD-1/PD-L1 pathway is well-documented in malignant melanoma21,22. PD-1 is typically expressed on T cells, while PD-L1 is predominantly on tumor cells23. The binding of PD-1 to PD-L1 inhibits T cell-mediated tumor cell killing, leading to enhanced tumor immune escape and progression24. Blocking this interaction can restore T cell activity and improve the tumor microenvironment (TME)25,26. Currently, PD-1/PD-L1 inhibitors are used to treat various tumors, including melanoma, non-small cell lung cancer (NSCLC), head and neck squamous cell carcinoma (HNSCC), and colorectal cancer (CRC)27,28,29,30,31. Previous studies have shown that EVO can suppress NSCLC and prostate cancer by repressing PD-L1 expressions32,33. In melanoma, it’s worth noting that nivolumab, a human anti-PD-1 monoclonal antibody, was FDA-approved in 2014 for the treatment of patients with refractor melanoma34 and in 2017 for the adjuvant treatment of melanoma patients with involvement of lymph nodes or metastatic melanoma patients who had complete resection35. Further, in 2022 the immune checkpoint inhibitors (ICIs)combination therapy of nivolumab and relatlimab (anti- Lymphocyte-activation gene 3) was FDA-approved for unresectable or metastatic melanoma36. Thus, we are pinning our hopes on EVO as a new ICI against melanoma.
PI3K/AKT signaling is a crucial component of many biological activities37 including cell proliferation, metabolism, motility, gene transcription, and protein synthesis38. Overactivation of this pathway in tumor cells promotes tumor growth and development39. Meanwhile, the antitumor mechanism of EVO has been linked to the PI3K/AKT signaling pathway40,41. EVO has been shown to inhibit IRS4/PI3K/AKT or PI3K/AKT/caspase signaling pathways in A375 cells41,42. Previous research has shown that transcription factors such as HIF-1α, NF-κB, AP-1, and STATs regulate PD-L1 expression by binding to its promoter, while hypoxia, cytokines, and EGF signaling pathways control PD-L1 expression at the transcriptional level. However, the MAPK and PI3K/AKT pathways are particularly influential in regulating PD-L1 expression43,44. For instance, in breast cancer, HMGB1-RAGE increases PD-L1 expression via the PI3K/AKT pathway45, and in hepatocellular carcinoma (HCC), blocking the PI3K/AKT/GSK3/PD-L1 pathway enhances liver immune surveillance46. Previous studies have also linked PD-L1 expression to the PI3K/AKT pathway in melanoma47,48,49.
With the advent of PD-1-based immunotherapies, which are vastly more effective compared to previous standard-of-care treatments in the advanced setting stage of melanoma50. Although previous studies have demonstrated the potential of EVO in treating melanoma, its mechanism in improving the TME and immunomodulation remains unexplored. In this study, we aimed to investigate the potential of EVO in regulating melanoma immunity and elucidate its underlying mechanisms. And our results suggest that EVO may inhibit tumor immune evasion and serve as a promising adjuvant for tumor immunotherapy. We believe that EVO hold great promise to enhance immunomodulation for lasting therapeutic response and ultimately a cancer cure.
Materials and methods
Cell culture and reagents
The C57BL/6 mouse melanoma cell line, B16-F10, was obtained from the American Type Culture Collection (ATCC, https://www.atcc.org/products/crl-6475). Cells were maintained in RPMI-1640 medium (Meilunbio, China) supplemented with 10% fetal bovine serum (FBS, ExCell Bio, USA) and 1% penicillin-streptomycin (HyClone, USA) and were cultured in a humidified CO2 incubator at 37℃. The mouse fibroblast cell line L929 was sourced from previously preserved stocks in our laboratory, with the same culture conditions as B16-F10 cells. Dimethyl sulfoxide (DMSO) was obtained from Sigma-Aldrich Chemical Co. (St. Louis, MO, USA). The AKT activator SC79 was purchased from MedChemExpress Co., USA, and evodiamine (≥ 98%) was obtained from Shaanxi Biaopu Pharmaceutical Technology Co., China.
Cell counting Kit-8 (CCK-8) assay
The CCK-8 assay was employed to assess cell viability. B16-F10 cells growing to log phase were seeded in 96-well plates at a density of 5 × 103 cells per well. After overnight cell culture, cells were treated with medium containing EVO at concentrations of 0, 1, 2, 4, 6, and 8 µM for 24, 48, and 72 h, respectively. Following treatment, 10 µL of CCK-8 solution (Yeasen, China) was added to each well and incubated at 37℃ in a 5% CO2 atmosphere for 2 h. Absorbance was measured at 450 nm by microplate reader (Tecan, Switzerland) to determine cell viability51.
Colony formation assay
B16-F10 cells were pre-treated with EVO at concentrations of 2, 4, and 6 µM for 48 h. The treated cells were then seeded in 6-well plates at a density of 1 × 103 cells per well in RPMI-1640 medium with 10% FBS. After 10 days of cell culture, colonies were visible. The colonies were fixed with 10% formaldehyde (Solarbio, China) for 15 min and stained with Giemsa (Solarbio, China) stain for 30 s. After washing with phosphate buffered saline (PBS, Meilunbio, China) and air-drying, colonies were photographed under natural light. Colony counts and calculations were performed using ImageJ software52. The clone formation rate is calculated as follows:
Apoptosis assay
B16-F10 cells were seeded in 6-well plates at a density of 2.5 × 105 cells per well and treated with EVO at concentrations of 2, 4, and 6 µM for 48 h. Cells were then digested with EDTA-free trypsin (Meilunbio, China) and washed two times with pre-cooled PBS. To determine the proportion of apoptotic cells, an Annexin V-FITC/PI apoptosis detection kit (Meilunbio, China) was used according to the manufacturer’s instructions. The specific experimental procedure is as follows. Cells were resuspended with pre-cooled 1×Binding Buffer and adjusted to a concentration of 1 × 106 cell/mL. Take 100 µL of cell suspension and add 5 µL each of Annexin V-FITC and PI staining solution and mix well, and then incubate for 10 min at room temperature and protected from light. After that, 400 µL of 1×Binding Buffer was added and mixed well. Samples were analyzed using a flow cytometer (BD Biosciences, USA), and data were analyzed with Cell Quest software53.
Transwell assay
An invasion assay was conducted using a Boyden chamber (Costar, USA). Matrigel® Basement Membrane Matrix (BD Biosciences, USA) was diluted in serum-free medium to the desired concentration. Fifty microliters of Matrigel were added to each upper chamber and incubated at 37℃ overnight to allow for gelling. Different concentrations of EVO-treated B16-F10 cells were suspended in serum-free RPMI-1640 medium and added to the upper wells of the chambers at a density of 2 × 10⁵ cells per well. In the lower chamber, 600 µL of RPMI-1640 medium supplemented with 20% FBS was added. After incubation at 37℃ in a 5% CO2 atmosphere for 24 h, non-migrating cells on the top of the upper wells were removed with cotton swabs. Migrating cells were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet (Solarbio, China), rinsed in pure water and air-dried. The cells were then observed and photographed under an inverted optical fluorescence microscope and the number of cells was counted using ImageJ software54.
Wound healing assay
B16-F10 cells were seeded in 6-well plates at a density of 5 × 10⁵ cells per well and cultured in RPMI-1640 medium with 10% FBS until a fully confluent monolayer was achieved. Once the cells reached 80% confluence, a wound was created by scratching the monolayer with a pipette tip. Each well was washed with PBS to remove free cells and then incubated with medium containing different concentrations of EVO at 37 °C in a 5% CO2 atmosphere. Wound edges were photographed at 0, 12, 24, and 48 h post-injury using microscopy, and the relative area of wound healing was calculated using ImageJ software51.
In vivo assay
Six-week-old male C57BL/6 mice (body weight 18–20 g) were purchased from the Experimental Animal Center of Lanzhou University. After one week of adaptive feeding, 2 × 10⁵ B16-F10 cells suspended in 200 µL of PBS were injected subcutaneously into the backs of the mice to establish the homograft mouse model55. When the tumor volume reached 50 mm³56, the mice were randomly divided into two groups (4 mice per group) and treated with 200 µL solutions of EVO (10 mg/kg) or saline via intragastric administration every one day for 7 days. No anesthesia was used during modeling and gavage. During the treatment period, tumor volume was measured and recorded every two days. The formula for calculating tumor volume is as follows: tumor volume (V) mm3 = (length) × (width) 2/2. After treatment, all mice were euthanized by injection of pentobarbital sodium with cervical dislocation, and the tumor tissue, heart, liver, spleen, lungs, and kidneys were excised for further analysis.
T-cell detection by flow cytometry
After euthanizing the mice, single-cell suspensions of bone marrow and spleen tissues were prepared using cell strainers (70 μm). The tissues were mixed with PBS and filtered through a 100 μm mesh filter, then centrifuged at 198 g for 7 min at 4℃. The supernatant was discarded. Erythrocytes were removed using red blood cell lysis buffer (MultiSciences, China) for 15 min at room temperature. The cells were then centrifuged at 198 g for 10 min at 4℃, and the supernatant was discarded. Cells were washed once with PBS and resuspended in cell staining buffer (MultiSciences, China) at a concentration of 1 × 10⁶ cells/mL. Cells (1 × 10⁵) were resuspended in a cell staining buffer containing surface antibodies: anti-FITC-CD3, anti-APC-CD4, and anti-PE-CD8 (MultiSciences, China, Lot: AM003E0201, AM00405, AM008A0204, 1:8). The cells were incubated in the dark at room temperature for 15 min. After incubation, 1 mL of cell staining buffer was added, and the tube was centrifuged at 300 g for 10 min. The supernatant was discarded, and the cells were resuspended in 500 µL of cell staining buffer. T-cell subpopulations were analyzed using a flow cytometer52.
Immunohistochemical (IHC) and Hematoxylin-eosin (H&E) staining
Tumor tissues and spleens from mice were fixed with 4% paraformaldehyde for 24 h, embedded in paraffin, sectioned, and stained with hematoxylin and eosin (Yuanyebio, China). Paraffin-embedded tissue sections of tumors and spleens were deparaffinized and subjected to immunohistochemical staining for PD-L1 (Huabio, China, 1:500), CD4 (Huabio, China, 1:1000), or CD8 (Huabio, China, 1:1000) at 4℃ overnight. After staining, sections were treated with a biotin-conjugated secondary antibody and horseradish peroxidase-streptavidin at room temperature for 2 h. Sections were developed using DAB (ZSGB-bio, China) and counterstained with hematoxylin. Finally, sections were mounted with neutral resin. Pathological changes and positively stained areas were observed using an inverted microscope. IHC was quantitatively analyzed using ImageJ software, with mean optical density serving as the intensity of staining57.
Real-time quantitative reverse transcription polymerase chain reaction (qRT-PCR)
B16-F10 cells were cultured in complete medium with 2, 4, and 6 µM EVO for 48 h. Total RNA was isolated from the treated cells using TRIzol reagent (Invitrogen, USA). The cells were washed three times with pre-cooled PBS, after which 1 mL of Trizol was added and transferred to a 1.5 mL enzyme-free centrifuge tube after sufficient blowing. Add 200 µL of chloroform (Fuyu Chemical, China), shake well for 15 s, centrifuge at 4 °C, 12,000 g for 15 min and aspirate the supernatant into a new centrifuge tube. Add an equal volume of pre-cooled isopropanol (Solarbio, China) and allow to stand at room temperature for 10 min before centrifuging at 12,000 g for 10 min at 4 °C. Afterwards, the supernatant was discarded, 1 mL of 75% ethanol was added and gently mixed to remove impurities. The sample was centrifuged at 7500 g for 5 min at 4 °C. Discard the supernatant. After drying at 37℃ for 5 min, add appropriate amount of DEPC water (Solarbio, China) and mix well. The concentration of RNA was detected by UV5 Nano UV-visible spectrophotometer (MettlerToledo, Switzerland) and stored at -80℃ after dispensing. Two micrograms of RNA were converted into cDNA using the Takara Reverse Transcription Kit (Takara, Japan) to prepare a reverse transcription system on ice. The samples were inserted into the PCR machine and stored at -20 °C after completion. Reaction systems were formulated for qRT-PCR using SYBR Premix Ex Taq II (Takara, Japan) following the manufacturer’s protocol58. Relative expression of PD-L1 was quantified with GAPDH as the housekeeping gene. The PCR primers used were: GAPDH, 5′-AGGTCGGTGTGAACGGATTTG-3′ (sense) and 5′-GGGGTCGTTGATGGCAACA-3′ (antisense); PD-L1, 5′-GAGCCCATTAGCTCTGTGTGA-3′ (sense) and 5′-CTTGACATGTGACAGAACAAGGA-3′ (antisense).
Western blotting analysis
Total protein was extracted from treated B16-F10 cells and tumor tissues using RIPA lysis buffer (Thermo Fisher, USA). Protein concentration was determined using a BCA kit (Beyotime, China). Afterwards protein denaturation using alternating hot and cold water baths. Protein samples, with 10 µg of protein per well, were separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE, Solarbio, China) and then transferred to polyvinylidene fluoride (PVDF, Millipore, USA) membranes. The membranes were blocked with 5% skim milk (Solarbio, China) in TBST (Solarbio, China) at room temperature for 2 h and incubated with primary antibodies at 4℃ overnight. Subsequently, membranes were incubated with secondary antibodies at room temperature for 2 h. Proteins were visualized using an ultrasensitive chemiluminescence reagent (Yeasen, China) and a fully automated chemiluminescence immunoassay analyzer (Clinx, China), and grayscale values were quantified using ImageJ software58. The following antibodies were used: PI3K (ImmunoWay, USA, 1:1000), P-PI3K (ImmunoWay, USA, 1:1000), AKT (ImmunoWay, USA, 1:1000), P-AKT (ImmunoWay, USA, 1:1000), goat anti-rabbit secondary antibody (ImmunoWay, USA, 1:1000), and PD-L1 (Huabio, China, 1:500). GAPDH (ImmunoWay, USA, 1:1000) was used as an internal control.
Statistical analysis
All experiments were performed in triplicate. Data are presented as mean ± standard deviation. Statistical analyses were conducted using IBM SPSS Statistics for Windows, version 20.0 (IBM Corp., Armonk, NY, USA) and GraphPad Prism software (version 7.0, GraphPad). Differences between groups were assessed using Student’s t-test and one-way analysis of variance (ANOVA) with Tukey’s multiple comparison test. A P-value of < 0.05 was considered statistically significant.
Results
EVO inhibits proliferation, invasion, and metastasis and promotes apoptosis of B16-F10 cells
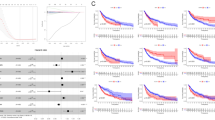
The effects of EVO on B16-F10 and L929 cell proliferation were assessed using CCK-8 assay. EVO was found to inhibit the proliferation of B16-F10 cells in a time-dependent and concentration-dependent manner compared to the untreated control group. The IC50 values at 24, 48, and 72 h were 11.73, 5.083, and 4.604 µM, respectively, as determined by the CCK-8 assay. L929 cell viability remained above 80% after treatment with various concentrations of EVO (Fig. 1A). Based on the IC50 value of EVO at 48 h, three concentrations—2, 4, and 6 µM—were selected for subsequent experiments. The results of colony formation assay showed that the clone formation rate of B16-F10 cells gradually decreased with the increase of EVO concentration (P < 0.05), especially in the 4 µM and 6 µM groups (P < 0.001), the clone formation rate was lower than 30% (Fig. 1B). The percentage of apoptotic B16-F10 cells treated with EVO was significantly higher compared to the control group, especially the apoptotic rate of B16-F10 cells was greater than 15% after 4 µM or 6 µM EVO action (P < 0.05) (Fig. 1C). To evaluate the effects of EVO on tumor metastasis, the ability of EVO to inhibit tumor migration and invasion was analyzed. Results from the transwell assay demonstrated that EVO significantly reduced the invasive capability of B16-F10 cells compared to the control group (P < 0.001) (Fig. 1D). Additionally, the wound healing assay showed that the wound healing area of control B16-F10 cells was as high as 70% after 48 h compared with the initial scratch area. However, the wound healing area of EVO-treated B16-F10 cells was even smaller, especially after 4 µM and 6 µM EVO, which was less than 50% of the area of the control group (P < 0.001), indicating that EVO substantially inhibited cell migration (Fig. 1E).
Effects of EVO treatment on the mouse melanoma cell line B16-F10. (A) Changes in cell proliferation of B16-F10 and L929 cells were assessed using the CCK-8 assay after treatment with different concentrations of evodiamine for 24, 48, and 72 h (n = 5). (B) Proliferation ability of B16-F10 cells treated with EVO at concentrations of 2 µM, 4 µM, and 6 µM for 48 h was evaluated using the colony formation assay (n = 3). (C) Apoptosis in B16-F10 cells was detected using the Annexin V-FITC/PI staining method and flow cytometry. (D) Invasion ability of B16-F10 cells was assessed using the transwell assay (scale bar = 100 μm). (E) Migration ability of B16-F10 cells was evaluated using the wound healing assay (scale bar = 200 μm). Data are expressed as mean ± SD. * p < 0.05, ** p < 0.01, *** p < 0.001.
Tumor growth in melanoma-bearing mice was inhibited by EVO
Given that EVO demonstrated significant inhibitory effects on B16-F10 cell growth in vitro, we evaluated its antitumor efficacy in vivo using a homograft tumor model. Tumors were excised one day after the final EVO administration (Fig. 2A). The results revealed that EVO significantly reduced both the mass and volume of the tumor tissue (P < 0.001) (Fig. 2B and C), suggesting that EVO has an inhibitory effect on the growth of melanoma syngeneic subcutaneous graft tumors. Additionally, histopathological analysis of tumor tissues showed that EVO-treated tumors had smaller nuclei with a decreased nuclear/cytoplasmic ratio and exhibited signs of apoptosis. In contrast, tumor cells from the control group were numerous and dense, displaying atypical shapes, increased mitotic figures, and an abnormal nuclear/cytoplasmic ratio (Fig. 2D).
Effect of EVO treatment on tumor growth in the homograft mouse model inoculated subcutaneously with B16-F10 cells. (A–C) The homograft mouse model was randomly divided into two groups for intragastric administration: model group (saline) and EVO group (10 mg/kg). Tumors were collected upon sacrifice (A) and weighed (B) (n = 4). Tumor volume (C) was compared among groups throughout the experimental period (n = 4). (D) HE staining images of tumors were photographed (scale bar = 50 μm) (n = 4). Data are expressed as mean ± SD. *** p < 0.001.
Inhibition of splenomegaly and evaluation of EVO biosafety in vivo
To assess the safety of EVO in vivo, we performed histopathological analysis on the main organs (heart, liver, lungs, and kidneys) of both control and EVO-treated mice. H&E staining showed that all organs in both groups appeared structurally intact with well-defined margins. No tissue degeneration, necrosis, or other significant pathological changes were observed (Fig. 3A), indicating the biosafety of EVO. The spleen, a crucial secondary immune organ59, is often enlarged and structurally compromised in tumor-bearing mice60. We compared spleen sizes across groups and found that the spleens of EVO-treated mice were significantly smaller than those of the model group but still larger than those of the control group (Fig. 3C). For consistency, the first spleen from each group was used for flow cytometry to ensure tissue freshness. Further analysis of the spleen coefficient (spleen mass/mouse mass) demonstrated that the mean spleen coefficient of control group was 4.42, while that of the model group was 6.962, which was 1.57 times higher than that of normal mice (P < 0.001). EVO partially restored this measure in melanoma-bearing mice, which decreased the mean value to 5.184, 1.22 times of the control group (P < 0.001) (Fig. 3D). Additionally, pathological examination revealed that the white and red pulp margins in the EVO-treated group were well-defined, with regular structure and dense lymphocyte infiltration in the white pulp compared to the model group (Fig. 3B). These findings suggest that EVO can potentially enhance the TME without causing toxicity in vivo.
Effects of EVO on main organs, especially the spleen, in melanoma-bearing mice. (A) Main organs of melanoma-bearing mice were collected and subjected to H&E staining, compared with wild-type mice of the control group (scale bar = 50 μm) (n = 4). (B–D) The spleen from three groups of mice was collected (C), stained with H&E (scale bar: upper = 100 μm, lower = 50 μm) (B), and weighed (D) (n = 4). Data are expressed as mean ± SD. *** p < 0.001.
EVO-regulated T-cell infiltration in vivo
T lymphocytes, as one of the most important immune cells in TME, play an important role in the function of tumour immunity. In order to further investigate the effect of EVO on TME, the infiltration of CD4⁺ and CD8⁺ T cells in the spleen and bone marrow of mice from each group was quantified using flow cytometry. Quantitative analysis revealed that the number of CD4⁺ and CD8⁺ T cells in the bone marrow of EVO-treated mice was significantly higher compared to the model group (P < 0.01) (Fig. 4A). While the number of CD4⁺ T cells in the spleens of the EVO-treated group was increased relative to the model group (P < 0.05), the number of CD8⁺ T cells did not show a significant difference (Fig. 4B). Immunohistochemical staining of spleen and tumor tissues demonstrated more pronounced positive expression of CD4⁺ and CD8⁺ in the EVO-treated group compared to the model group (P < 0.01) (Fig. 4C and D). These results suggest that EVO enhances T-cell infiltration and regulates T-cell levels in melanoma-bearing mice.
Regulation of T-cells in melanoma-bearing mice by EVO. (A,B) Percentages of CD4+ T cells and CD8+ T cells in (A) bone marrow and (B) spleen were detected by flow cytometry (n = 2). (C,D) Tumor and spleen sections were analyzed by IHC for CD4 and CD8, with quantitative analysis (scale bar = 50 μm) (n = 4). Data are expressed as mean ± SD. * p < 0.05, ** p < 0.01, *** p < 0.001.
EVO regulates PD-L1 expression through the PI3K/AKT signaling pathway
PD-L1 plays a crucial role in tumor immune evasion. To investigate the effect of EVO on PD-L1 expression, we analyzed tumor tissues from various mouse groups. Results indicated that PD-L1 expression was significantly reduced in the tumors of the EVO-treated group compared to the model group, at both the mRNA and protein levels (P < 0.01) (Fig. 5A and B). Immunohistochemical staining also revealed significantly less yellow-brown positive PD-L1 expression in the EVO-treated tumors compared to the model group (Fig. 5C), confirming that EVO reduces PD-L1 expression in melanoma. To elucidate the mechanism underlying the regulation of EVO of PD-L1 expression, we examined the PI3K/AKT signaling pathway using western blotting. We observed that with increasing concentrations of EVO, there was a decrease in PI3K phosphorylation, AKT phosphorylation, and PD-L1 expression in B16-F10 cells (P < 0.01). Specifically, higher EVO concentrations were associated with lower expression levels of these proteins (Fig. 5D). Additionally, EVO treatment led to decreased expression of these proteins in homograft tumors (P < 0.01) (Fig. 5E). Further investigation involved co-treating B16-F10 cells with the AKT activator SC79 and EVO to assess changes in the PI3K/AKT signaling pathway and PD-L1 expression by Western blotting. The experiment was set up as four groups. The control group was incubated normally for 48 h without any treatment. The EVO group was treated with complete medium containing 4 µM EVO for 48 h. The SC79 group B16-F10 cells were treated with complete medium containing 20 µM AKT agonist SC79. The SC79 group was pretreated with complete medium containing 20 µM SC79 for 2 h. After that, the original medium was discarded, and fresh complete medium was added to continue incubation for 48 h. The EVO + SC79 group was pretreated with SC79 for 2 h, and then treated with complete medium containing 4 µM EVO for 48 h. The results showed that the expression of both P-AKT and PD-L1 was significantly up-regulated in B16-F10 cells pretreated with SC79 only compared to the control group (P < 0.01). EVO alone significantly reduced PI3K phosphorylation, AKT phosphorylation, and PD-L1 expression (P < 0.05). However, co-treatment with SC79 and EVO resulted in a notable rebound in PI3K phosphorylation, AKT phosphorylation, and PD-L1 expression compared to EVO alone (P < 0.05) (Fig. 5F). These findings suggest that the regulation of PD-L1 expression by EVO is linked to the PI3K/AKT signaling pathway.
Regulation of PD-L1 expression by EVO through the PI3K/AKT signaling pathway. (A–C) Tumor tissues from tumor-bearing mice were harvested, and RNA and protein were extracted. PD-L1 expression was analyzed by (A) qRT-PCR (n = 4), (B) Western blotting (n = 5), and (C) IHC (scale bar = 40 μm) (n = 4). (D) B16-F10 cells were treated with EVO at different concentrations (2, 4, 6 µM) for 48 h, and the expressions of PI3K, P-PI3K, AKT, P-AKT, and PD-L1 were detected by Western blotting (n = 4). (E) Similar experiments were performed with tumors isolated from the subcutaneous mouse model (n = 4). (F) Expressions of PI3K, P-PI3K, AKT, P-AKT, and PD-L1 in B16-F10 cells treated with EVO combined with SC79 were detected by western blotting (n = 4). Data are expressed as mean ± SD. * p < 0.05, ** p < 0.01, *** p < 0.001.
Discussion
In the early 20th century, the concept of ‘Cancer Immunoediting’ was defined by numerous scholars, describing it as a dynamic process comprising three stages: elimination, equilibrium, and escape61,62. This discovery paved the way for immunotherapy as a promising approach to controlling tumors. In recognition of their pioneering work on utilizing the immune system to combat cancer, two scientists were awarded the 2018 Nobel Prize in Physiology/Medicine56. Recently, immunotherapy has provided survival prospects for several patients with tumors unresponsive to traditional treatments, with melanoma notably benefiting due to its established immunogenicity63. Inhibitors targeting PD-1/PD-L1 and CTLA-4 have shown potential in melanoma treatment64. However, due to the complexity and variability of the TME, these drugs often exhibit limited and inconsistent efficacy in most patients65,66. Consequently, the development of adjuvants with immunotherapeutic potential is crucial.
Traditional Chinese medicine (TCM) has recently attracted attention for its low systemic side effects and notable antitumor properties. However, EVO has poor solubility and bioavailability, so recently EVO has been loaded into delivery vehicles to improve its antimelanoma effect, such as ethosomes delivery system and zeolitic imidazolate framework-867,68. Despite this, research on EVO has largely focused on its direct effects on tumor cells, with limited exploration of its potential as an immune adjuvant. This study demonstrates that EVO not only inhibits the proliferation, invasion, and metastasis of the mouse melanoma cell line B16-F10 in a concentration-dependent manner, and the inhibition of 4µM and 6µM EVO reached more than 50% (P < 0.001), but also significantly suppresses the growth of melanoma subcutaneous transplants (P < 0.001), indicating its superior efficacy against melanoma. Moreover, the IC50 values of different action times showed that when the action time of EVO was longer than 24 h, the IC50 values were all within 10 µM, which further indicated that a small dose of EVO could inhibit the proliferation of melanoma cells. The results of the in vitro experiments were generally consistent with previous studies, but Liu et al. used a larger concentration of 10 µM EVO and obtained a higher apoptosis rate16. The antitumor effect of this in vivo experiment was significantly enhanced compared to the in vivo effect of EVO treatment alone by Zhou et al.67. This may be due to the difference in the duration and frequency of administration.
To assess the biological safety and tumor immunomodulatory capacity of EVO in vivo, splenic damage was evaluated in treated mice. EVO significantly reduced tumor-induced splenomegaly in mice (P < 0.001). Given the role of the spleen in immune cell production, it was hypothesized that EVO might influence tumor immunity. T cells, crucial for combating tumor cells within the TME69,70, include CD4+ helper T cells and CD8+ cytotoxic T cells71. CD4+ T cells exert their anti-tumor function by assisting CD8+ T cells and B cells and through direct cytotoxicity. CD8+ T cells exert anti-tumor functions mainly by releasing cytotoxins such as perforin and granzyme B into tumor cells, or by inducing apoptosis through FAS-FASL interactions72,73. Moreover, ICIs induce cancer-cell killing by inducing CD8+ T cells after targeting the dysfunctional immune system74. Flow cytometry revealed significantly higher proportions of CD4+ and CD8+ T cell subsets in the bone marrow and CD4+ T cells in the spleen of EVO-treated mice (P < 0.01). Immunohistochemistry also showed elevated expression of CD4+ and CD8+ cells in the spleen and tumor tissues of EVO-treated mice (P < 0.01). However, discrepancies between flow cytometry and immunohistochemical results, particularly the higher CD8 expression in immunohistochemical sections compared to flow cytometry, were noted. This discrepancy may be attributed to fluorescence compensation issues during flow cytometry. Despite this, both methods suggest that EVO has the potential to regulate T cells in tumor-bearing mice.
Future research should further investigate how EVO regulates T cells and its impact on other immune cells within the TME. The activation of T cells within the TME is often hindered by the PD-1/PD-L1 signaling pathway, facilitating tumor immune escape75. PD-L1 is frequently overexpressed in melanoma and is associated with poor prognosis76. EVO significantly reduced PD-L1 expression in B16-F10 cells and melanoma-bearing mice, alongside a notable decrease in factors related to the PI3K/AKT signaling pathway (P < 0.01). The results of immunoblotting of the PI3K/AKT signaling pathway were consistent with previous studies41,42. This effect was partially reversed by co-treatment with the AKT agonist SC79 (P < 0.05), suggesting that EVO’s reduction of PD-L1 expression is mediated through the PI3K/AKT pathway. Unlike previous studies, this paper for the first time links PD-L1 regulation mediated by the PI3K/AKT signaling pathway to immune modulation in TME of melanoma by EVO. The preliminary results showed that the PI3K/AKT signaling pathway played an important role in the inhibition of PD-L1 expression in melanoma by EVO. But its more in-depth and specific regulatory mechanisms need to be further investigated.
In conclusion, EVO showed excellent ability to inhibit melanoma growth in vivo and in vitro, with an inhibition rate of more than 50%, and its mechanism of action may be related to the down-regulation of PD-L1 expression through the PI3K/AKT signaling pathway, increasing T-cell infiltration, and attenuating splenic injury. However, it was important to note that the efficacy shown by EVO in melanoma treatment may hold significant promise as a novel immunotherapeutic agent targeting PD-1/PD-L1 signaling in tumor immunosuppressive contexts.
Data availability
These data are included in this article. The data supporting the findings of this study are available from the corresponding author, [Jing Wang], upon reasonable request.
References
Centeno, P. P., Pavet, V. & Marais, R. The journey from melanocytes to melanoma. Nat. Rev. Cancer. 23(6), 372–390 (2023).
Brombin, A. & Patton, E. E. Melanocyte lineage dynamics in development, growth and disease. Dev. (Cambridge England) ;151(15). (2024).
Miller, K. D. et al. Cancer statistics for adolescents and young adults, 2020. CA Cancer J. Clin. 70(6), 443–459 (2020).
Song, M., Liu, C., Chen, S. & Zhang, W. Nanocarrier-Based drug delivery for melanoma therapeutics. Int. J. Mol. Sci. ;22(4). (2021).
Davis, L. E., Shalin, S. C. & Tackett, A. J. Current state of melanoma diagnosis and treatment. Cancer Biol. Ther. 20(11), 1366–1379 (2019).
Schadendorf, D. et al. Melanoma Lancet ;392(10151):971–984. (2018).
Tseng, Y-J., Lee, C-H., Chen, W-Y., Yang, J-L. & Tzeng, H-T. Inhibition of PAI-1 blocks PD-L1 endocytosis and improves the response of melanoma cells to immune checkpoint Blockade. J. Invest. Dermatology. 141(11), 2690–8e6 (2021).
Wang, R., Chen, Y., Xie, Y., Ma, X. & Liu, Y. Deciphering and overcoming Anti-PD-1 resistance in melanoma: A comprehensive review of mechanisms, biomarker developments, and therapeutic strategies. Int. Immunopharmacol. 132, 111989 (2024).
Panda, M., Tripathi, S. K., Zengin, G. & Biswal, B. K. Evodiamine as an anticancer agent: a comprehensive review on its therapeutic application, pharmacokinetic, toxicity, and metabolism in various cancers. Cell. Biol. Toxicol. (2022).
Li, M. & Wang, C. Traditional uses, phytochemistry, pharmacology, pharmacokinetics and toxicology of the fruit of Tetradium ruticarpum: A review. J. Ethnopharmacol. 263, 113231 (2020).
Yun, U. J., Bae, S. J., Song, Y. R. & Kim, Y. W. A critical YAP in malignancy of HCC is regulated by Evodiamine. Int. J. Mol. Sci. ;23(3). (2022).
Panda, M., Tripathi, S. K., Zengin, G. & Biswal, B. K. Evodiamine as an anticancer agent: a comprehensive review on its therapeutic application, pharmacokinetic, toxicity, and metabolism in various cancers. Cell Biol. Toxicol. 39(1), 1–31 (2023).
Liu, J. et al. Evodiamine inhibits proliferation and induces apoptosis of nasopharyngeal carcinoma cells via the SRC/ERBB2-mediated MAPK/ERK signaling pathway. J. Transl Med. 22(1), 859 (2024).
Yang, J., Wu, L. J., Tashiro, S., Onodera, S. & Ikejima, T. Nitric oxide activated by p38 and NF-kappaB facilitates apoptosis and cell cycle arrest under oxidative stress in evodiamine-treated human melanoma A375-S2 cells. Free Radic Res. 42(1), 1–11 (2008).
Yang, J., Wu, L. J., Tashino, S., Onodera, S. & Ikejima, T. Critical roles of reactive oxygen species in mitochondrial permeability transition in mediating evodiamine-induced human melanoma A375-S2 cell apoptosis. Free Radic Res. 41(10), 1099–1108 (2007).
Liu, N., Li, Y., Chen, G. & Ge, K. Evodiamine induces reactive oxygen species-dependent apoptosis and necroptosis in human melanoma A-375 cells. Oncol. Lett. 20(4), 121 (2020).
Zhang, Y., Wu, L. J., Tashiro, S., Onodera, S. & Ikejima, T. Evodiamine induces A375-S2 cell death through two different pathways. Yao Xue Xue Bao = Acta Pharm. Sinica. 38(9), 650–653 (2003).
Zhang, Y., Wu, L. J., Tashiro, S., Onodera, S. & Ikejima, T. Evodiamine induces tumor cell death through different pathways: apoptosis and necrosis. Acta Pharmacol. Sin. 25(1), 83–89 (2004).
Fei, X. F. et al. Evodiamine, a constituent of evodiae fructus, induces anti-proliferating effects in tumor cells. Cancer Sci. 94(1), 92–98 (2003).
Zhang, Y., Wu, L. J., Tashiro, S., Onodera, S. & Ikejima, T. Intracellular regulation of evodiamine-induced A375-S2 cell death. Biol. Pharm. Bull. 26(11), 1543–1547 (2003).
Wu, M. et al. Improvement of the anticancer efficacy of PD-1/PD-L1 Blockade via combination therapy and PD-L1 regulation. J. Hematol. Oncol. 15(1), 24 (2022).
Liu, D., Gao, S., Zhai, Y., Yang, X. & Zhai, G. Research progress of tumor targeted drug delivery based on PD-1/PD-L1. Int. J. Pharm. 616, 121527 (2022).
Budimir, N., Thomas, G. D., Dolina, J. S. & Salek-Ardakani, S. Reversing T-cell exhaustion in cancer: lessons learned from PD-1/PD-L1 immune checkpoint blockade. Cancer Immunol. Res. 10(2), 146–153 (2022).
Sharpe, A. H. & Pauken, K. E. The diverse functions of the PD1 inhibitory pathway. Nat. Rev. Immunol. 18(3), 153–167 (2018).
Lai, J., Beavis, P. A., Li, J. & Darcy, P. K. Augmenting adoptive T-cell immunotherapy by targeting the PD-1/PD-L1 Axis. Cancer Res. 81(23), 5803–5805 (2021).
Acúrcio, R. C. et al. Therapeutic targeting of PD-1/PD-L1 blockade by novel small-molecule inhibitors recruits cytotoxic T cells into solid tumor microenvironment. J. Immunother Cancer ;10(7). (2022).
Liu, Y., Zhang, X., Wang, G. & Cui, X. Triple combination therapy with PD-1/PD-L1, BRAF, and MEK inhibitor for stage III-IV melanoma: A systematic review and Meta-Analysis. Front. Oncol. 11, 693655 (2021).
Genova, C. et al. Therapeutic implications of tumor microenvironment in lung cancer: focus on immune checkpoint Blockade. Front. Immunol. 12, 799455 (2021).
Vathiotis, I. A., Johnson, J. M. & Argiris, A. Enhancing programmed cell death protein 1 axis Inhibition in head and neck squamous cell carcinoma: combination immunotherapy. Cancer Treat. Rev. 97, 102192 (2021).
Zhang, X. et al. Clinical benefits of PD-1/PD-L1 inhibitors in patients with metastatic colorectal cancer: a systematic review and meta-analysis. World J. Surg. Oncol. 20(1), 93 (2022).
Mo, D. C. et al. PD-1/PD-L1 inhibitor plus chemotherapy versus standard of care in the first-line treatment for recurrent or metastatic head and neck squamous cell carcinoma. European archives of oto-rhino-laryngology: official journal of the European federation of Oto-Rhino-Laryngological societies (EUFOS) : affiliated with the German society for Oto-Rhino-Laryngology. Head Neck Surg. 280(1), 1–9 (2023).
Yu, Y., Huang, X., Liang, C. & Zhang, P. Evodiamine impairs HIF1A histone lactylation to inhibit Sema3A-mediated angiogenesis and PD-L1 by inducing ferroptosis in prostate cancer. Eur. J. Pharmacol. 957, 176007 (2023).
Jiang, Z. B. et al. Evodiamine suppresses non-small cell lung cancer by elevating CD8(+) T cells and downregulating the MUC1-C/PD-L1 axis. J. Exp. Clin. Cancer Res. 39(1), 249 (2020).
Korman, A. J., Garrett-Thomson, S. C. & Lonberg, N. The foundations of immune checkpoint Blockade and the ipilimumab approval decennial. Nat. Rev. Drug Discovery. 21(7), 509–528 (2022).
Administration USFaD. FDA grants regular approval to nivolumab for adjuvant treatment of melanoma. United States (2017).
Tawbi, H. A. et al. Relatlimab and nivolumab versus nivolumab in untreated advanced melanoma. N Engl. J. Med. 386(1), 24–34 (2022).
Yu, L., Wei, J. & Liu, P. Attacking the PI3K/Akt/mTOR signaling pathway for targeted therapeutic treatment in human cancer. Semin Cancer Biol. 85, 69–94 (2022).
Chen, K. et al. The role of the PI3K/AKT signalling pathway in the corneal epithelium: recent updates. Cell. Death Dis. 13(5), 513 (2022).
Xue, C., Li, G., Lu, J. & Li, L. Crosstalk between circrnas and the PI3K/AKT signaling pathway in cancer progression. Signal. Transduct. Target. Ther. 6(1), 400 (2021).
Lei, Y. et al. Evodiamine as the active compound of evodiae fructus to inhibit proliferation and migration of prostate cancer through PI3K/AKT/NF-B signaling pathway. Dis. Markers. 2022, 4399334 (2022).
Guo, X. et al. Evodiamine inhibits growth of vemurafenib drug-resistant melanoma via suppressing IRS4/PI3K/AKT signaling pathway. J. Nat. Med. 78(2), 342–354 (2024).
Wang, C., Li, S. & Wang, M. W. Evodiamine-induced human melanoma A375-S2 cell death was mediated by PI3K/Akt/caspase and Fas-L/NF-kappaB signaling pathways and augmented by ubiquitin-proteasome inhibition. Toxicol. Vitro. 24(3), 898–904 (2010).
Wu, Y., Chen, W., Xu, Z. P. & Gu, W. PD-L1 distribution and perspective for cancer immunotherapy-blockade, knockdown, or inhibition. Front. Immunol. 10, 2022 (2019).
Pang, K. et al. Research progress of therapeutic effects and drug resistance of immunotherapy based on PD-1/PD-L1 blockade. Drug Resist. Updates. 66, 100907 (2023).
Amornsupak, K. et al. HMGB1 mediates invasion and PD-L1 expression through RAGE-PI3K/AKT signaling pathway in MDA-MB-231 breast cancer cells. BMC Cancer. 22(1), 578 (2022).
Wu, M., Xia, X., Hu, J., Fowlkes, N. W. & Li, S. WSX1 act as a tumor suppressor in hepatocellular carcinoma by downregulating neoplastic PD-L1 expression. Nat. Commun. 12(1), 3500 (2021).
Zhang, M. et al. Targeting inhibition of accumulation and function of myeloid-derived suppressor cells by artemisinin via PI3K/AKT, mTOR, and MAPK pathways enhances anti-PD-L1 immunotherapy in melanoma and liver tumors. J. Immunol. Res. 2022, 2253436 (2022).
Tonk, M. et al. The Drosophila melanogaster antimicrobial peptides Mtk-1 and Mtk-2 are active against the malarial parasite Plasmodium falciparum. Parasitol. Res. 118(6), 1993–1998 (2019).
Gong, Q. et al. Suppression of stemness and enhancement of chemosensibility in the resistant melanoma were induced by Astragalus polysaccharide through PD-L1 downregulation. Eur. J. Pharmacol. 916, 174726 (2022).
Sussman, T. A. & Ott, P. A. Adjuvant immunotherapy for melanoma patients: progress and opportunities. ESMO Open. 9(5), 102962 (2024).
Xiang, S. et al. The dietary flavonoid isoliquiritigenin induced apoptosis and suppressed metastasis in melanoma cells: an in vitro and in vivo study. Life Sci. 264, 118598 (2021).
Peng, D. et al. Melanoma suppression by quercein is correlated with RIG-I and type I interferon signaling. Biomed. Pharmacother. 125, 109984 (2020).
Zhu, P. et al. Arnicolide D exerts anti-melanoma effects and inhibits the NF-κB pathway. Phytomedicine 64, 153065 (2019).
Lin, X., Zhong, L., Wang, N., Chu, X. & Liu, B. Hsa_circ_0103232 promotes melanoma cells proliferation and invasion via targeting miR-661/RAB3D. Cell cycle (Georgetown. Tex) 21(17), 1811–1826 (2022).
Li, J-K. et al. A two-herb formula inhibits STAT3 signaling and exerts anti-melanoma effects in cell and animal models. J. Ethnopharmacol. ;268. (2021).
Smyth, M. J. & Teng, M. W. 2018 Nobel prize in physiology or medicine. Clin. Transl Immunol. 7(10), e1041 (2018).
Zheng, Y. et al. Manganese-enriched photonic/catalytic nanomedicine augments synergistic anti-TNBC photothermal/nanocatalytic/immuno-therapy via activating cGAS-STING pathway. Biomaterials ;293. (2023).
Liu, W. B. et al. Cucurbitacin E inhibits cellular proliferation and induces apoptosis in melanoma by suppressing HSDL2 expression. Chin. Med. 17(1), 28 (2022).
Wu, C. et al. Spleen mediates a distinct hematopoietic progenitor response supporting tumor-promoting myelopoiesis. J. Clin. Invest. 128(8), 3425–3438 (2018).
Lewis, S. M., Williams, A. & Eisenbarth, S. C. Structure and function of the immune system in the spleen. Sci. Immunol. ;4(33). (2019).
Dunn, G. P., Old, L. J. & Schreiber, R. D. The immunobiology of cancer immunosurveillance and immunoediting. Immunity 21(2), 137–148 (2004).
Reiman, J. M., Kmieciak, M., Manjili, M. H. & Knutson, K. L. Tumor immunoediting and immunosculpting pathways to cancer progression. Semin Cancer Biol. 17(4), 275–287 (2007).
Marzagalli, M., Ebelt, N. D. & Manuel, E. R. Unraveling the crosstalk between melanoma and immune cells in the tumor microenvironment. Semin Cancer Biol. 59, 236–250 (2019).
Zhang, J. et al. Targeted therapy, immunotherapy, and small molecules and peptidomimetics as emerging immunoregulatory agents for melanoma. Cancer Lett. 586, 216633 (2024).
de Azevedo, R. A. et al. MIF Inhibition as a strategy for overcoming resistance to immune checkpoint Blockade therapy in melanoma. Oncoimmunology 9(1), 1846915 (2020).
Aroldi, F. & Middleton, M. R. Long-Term outcomes of immune checkpoint Inhibition in metastatic melanoma. Am. J. Clin. Dermatol. 23(3), 331–338 (2022).
Zhou, Q. et al. Evodiamine encapsulated by hyaluronic acid modified zeolitic imidazolate framework-8 for tumor targeted therapy. Drug Deliv Transl Res. (2024).
Lin, H., Lin, L., Choi, Y. & Michniak-Kohn, B. Development and in-vitro evaluation of co-loaded berberine chloride and evodiamine ethosomes for treatment of melanoma. Int. J. Pharm. 581, 119278 (2020).
Waibl Polania, J., Lerner, E. C., Wilkinson, D. S., Hoyt-Miggelbrink, A. & Fecci, P. E. Pushing past the blockade: advancements in T cell-based cancer immunotherapies. Front. Immunol. 12, 777073 (2021).
Heidari-Foroozan, M., Rezalotfi, A. & Rezaei, N. The molecular landscape of T cell exhaustion in the tumor microenvironment and reinvigoration strategies. Int. Rev. Immunol. 43(6), 419–440 (2024).
Sun, L., Su, Y., Jiao, A., Wang, X. & Zhang, B. T cells in health and disease. Signal. Transduct. Target. Ther. 8(1), 235 (2023).
Peng, P. et al. Th1-Dominant CD4(+) T cells orchestrate endogenous systematic antitumor immune memory after Cryo-Thermal therapy. Front. Immunol. 13, 944115 (2022).
Martínez-Lostao, L., Anel, A. & Pardo, J. How do cytotoxic lymphocytes kill cancer cells?? Clin. Cancer Res. 21(22), 5047–5056 (2015).
Carlino, M. S., Larkin, J. & Long, G. V. Immune checkpoint inhibitors in melanoma. Lancet 398(10304), 1002–1014 (2021).
Dammeijer, F. et al. The PD-1/PD-L1-checkpoint restrains T cell immunity in tumor-draining lymph nodes. Cancer Cell. ;38(5). (2020).
Chuang, I. C. & Jang, C-S. Appraisal of clinicopathological prognosticators in advanced acral lentiginous melanoma with characterization of PD-L1 and CD8/CD4 immunoprofiles. Jpn J. Clin. Oncol. 52(9), 975–981 (2022).
Acknowledgements
We acknowledge the support of the Scientific and Technological Foundation of Gansu Province (20JR10FA670) and the Research Supporting Fund of the School/Hospital of Stomatology, Lanzhou University (lzukqky-2022-p04).
Author information
Authors and Affiliations
Contributions
J.L. and L.J. conceived and designed the study, performed statistical analyses, interpreted the data, drafted and revised the manuscript, and contributed to the language polishing of manuscript. Q.M., Z.Z., S.Z., and J.Q. performed some experiments. J.W. and Y.P. contributed to conception, study design, and interpretation of the data, and revised the manuscript. All authors have read, edited, and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics declarations
This study complied with relevant guidelines and regulations. This study was reviewed and approved by the Ethical Research Committee of the Lanzhou University School of Stomatology (approval number: LZUKQ-2021-024). The study is reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, J., Jiang, L., Ma, Q. et al. Evodiamine inhibits programmed cell death ligand 1 expression via the PI3K/AKT signaling pathway to regulate antitumor immunity in melanoma. Sci Rep 15, 6649 (2025). https://doi.org/10.1038/s41598-025-91137-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-91137-2
Keywords
This article is cited by
-
Natural compounds as immune checkpoint inhibitors in melanoma: a systematic review
Naunyn-Schmiedeberg's Archives of Pharmacology (2026)