Abstract
Rigidity sensing enables cells to respond to extracellular matrix stiffness and governs survival and apoptotic decisions. Tropomyosin 2.1 (Tpm2.1), a key actin-binding protein, is frequently downregulated in cancer. While re-expressing Tpm2.1 restores rigidity sensing and limits anchorage-independent growth, its ability to fully trigger anoikis, apoptosis induced by loss of matrix attachment, remains unclear. Here, we show that although Tpm2.1 overexpression reduces Akt activation and suppresses survival signaling in suspended breast cancer cells, most cells remain viable without matrix attachment. Time-resolved transcriptomic profiling of apoptotic and non-apoptotic suspended cells revealed compensatory survival programs, including reactivation of PI3K–Akt signaling and upregulation of intercellular adhesion molecules such as ICAM1. Importantly, anoikis resistance can be overcome. Pharmacological inhibition of Akt, EphA2, or ICAM1 in Tpm2.1-expressing cells significantly increases apoptosis under suspension conditions. These findings highlight that while rigidity sensing restoration alone may be insufficient to induce anoikis, it sensitizes cancer cells to targeted therapies, revealing actionable vulnerabilities to eliminate detached tumor cells.
Similar content being viewed by others
Introduction
Anchorage-independent growth is a hallmark of metastatic cancer, allowing tumor cells to survive and proliferate without attachment to the extracellular matrix (ECM)1,2,3,4,5. This capacity is closely linked to defects in rigidity sensing, a process by which cells detect and respond to matrix stiffness to regulate survival, proliferation, and apoptosis6,7,8. In normal tissues, detachment from the ECM activates anoikis, a specialized form of programmed cell death that prevents inappropriate survival and growth in suspension9,10,11,12. Metastatic cells evade anoikis, enabling dissemination through circulation and colonization of distant sites13.
Rigidity sensing is mediated by focal adhesions, macromolecular complexes that connect integrin clusters at the membrane to the actomyosin cytoskeleton14,15,16,17. Cells probe matrix stiffness by generating contractile forces through sarcomere-like units composed of actin, non-muscle myosin IIA, Tropomyosin 2.1 (Tpm2.1), α-actinin 4, tropomodulin, and focal adhesion proteins18. On rigid ECM, this resistance promotes focal adhesion maturation and stabilization of cytoskeletal structures, supporting spreading and survival. On soft ECM, insufficient resistance leads to focal adhesion disassembly and apoptosis19,20,21,22,23. Tpm2.1, a key structural component of the rigidity-sensing machinery, is frequently downregulated in cancer24,25. Its re-expression has been shown to restore rigidity sensing, suppress anchorage-independent growth, and induce anoikis7,12,26. However, the downstream effectors that determine whether rigidity sensing leads to survival or cell death remain poorly defined.
While it has been broadly assumed that restoring mechanosensitivity is sufficient to trigger anoikis7,27,28,29,30, this notion has not been rigorously tested in metastatic breast cancer cells. In this study, we show that Tpm2.1 overexpression increases apoptosis in suspended MDA-MB-231 cells, yet a majority of cells remain viable. This unexpected persistence of survival prompted a systematic investigation into the gene expression programs that distinguish apoptotic from non-apoptotic states in rigidity-restored cells.
Using time-resolved transcriptomic profiling, we compared apoptotic and surviving fractions of Tpm2.1-expressing cells across a 4-day suspension time course. We find that survival is driven by a dynamic and coordinated transcriptional program involving early activation of inflammatory signaling, progressive upregulation of PI3K-Akt and MAPK pathways, and persistent expression of cell adhesion molecules such as ICAM1. Mechanistically, we demonstrate that targeting these compensatory pathways through inhibition of ICAM1, phospho-Akt, or EphA2 substantially increases apoptosis in Tpm2.1-expressing cells. These findings define rigidity sensing as a priming signal rather than a binary switch and identify actionable targets that convert mechanosensitive survival states into apoptotic ones.
Results
Tpm2.1 expression promotes anoikis and inhibits migration in anoikis-resistant MDA-MB-231 breast cancer cells
Previous studies have shown that MDA-MB-231 cells, a metastatic breast cancer cell line, exhibit aberrant matrix rigidity sensing due to low expression of the cytoskeletal protein Tpm2.1, allowing them to survive and grow on soft agar7,12. Re-expression of Tpm2.1 has been reported to restore rigidity sensing and inhibit anchorage-independent growth.
Growth in non-adherent conditions is an established assay for anchorage independent growth31 and mimics the behavior of circulating tumor cells32. Therefore, to assess whether Tpm2.1 overexpression promotes apoptosis in suspension, we cultured wild-type MDA-MB-231 cells (MDA-WT), empty vector controls (MDA-EV), and Tpm2.1-expressing cells (MDA-Tpm2.1)33 on ultra-low attachment (ULA) plates (see methods), which prevents cells from attaching to the substrate and maintain cells in suspension33. To study how apoptosis progresses after detachment, we performed a time-course experiment in both adherent and suspension conditions at 5 h, 1 day, 2 days, and 4 days (Fig. 1A). Apoptosis assays revealed that Tpm2.1 expression significantly increased anoikis compared to both MDA-WT and MDA-EV cells, with elevated apoptosis observed as early as day 1 and peaking by day 4 (Fig. 1D, E). These results indicate that Tpm2.1 re-expression reactivates cell death programs normally suppressed in metastatic cells. However, approximately 70% of MDA-Tpm2.1 cells remained viable in suspension by day 4, suggesting the presence of a subpopulation that resists apoptosis despite restored rigidity sensing.
To determine whether these surviving cells continued to proliferate in suspension, we monitored cell growth over time (Fig. 1B, C). MDA-EV cells showed robust proliferation, with a 2.5-fold increase in cell numbers by day 5 and a 4.4-fold increase by day 8. In contrast, MDA-Tpm2.1 cells exhibited significantly reduced growth under the same conditions, with only a 1.2-fold increase by day 5 and 1.6-fold by day 8. This reduction in growth indicates that, beyond promoting apoptosis, Tpm2.1 limits the ability of cancer cells to proliferate in suspension.
Similarly, soft agar assays showed that MDA-WT and MDA-EV cells formed significantly more colonies than MDA-Tpm2.1 cells (Fig. S1A, B), consistent with previous reports that Tpm2.1 expression suppresses anchorage-independent growth7,26.
RNA-seq and western blot data show that Tpm2.1 expression levels are comparable in both MDA-WT and MDA-EV cells (Fig. S1C, D). Additionally, the levels of apoptosis under suspension conditions were similar between MDA-WT and MDA-EV cells (Fig. 1E). Therefore, we used MDA-EV cells as a control for all subsequent experiments.
Given the highly metastatic nature of MDA-MB-231 cells34,35, we next assessed whether Tpm2.1 expression also affected cell migration. Transwell migration assays revealed significantly reduced motility in MDA-Tpm2.1 cells compared to MDA-EV controls (Fig. 1F, G).
Together, these results demonstrate that Tpm2.1 re-expression induces apoptosis, impairs proliferation under non-adherent conditions, and inhibits migration in metastatic breast cancer cells. These findings establish Tpm2.1 as a key suppressor of anchorage-independent survival and metastatic behavior. However, the persistence of viable cells in suspension points to the existence of compensatory survival mechanisms that bypass canonical anoikis signaling.
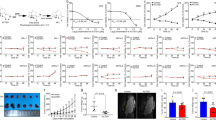
Tpm2.1 expression promotes anoikis in anoikis-resistant MDA-MB-231 breast cancer cells. (A) Schematic of MDA-EV and MDA-Tpm2.1 cells cultured under adherent (tissue culture plate) and suspension (low-attachment plate) conditions. (B) Representative phase-contrast images of MDA-WT, MDA-EV and MDA-Tpm2.1 cells after 8 days in suspension culture. Scale bar = 300 μm (C) Mean ± SD of fold change in cell density for MDA-Tpm2.1 and MDA-EV cells cultured in suspension for 5 and 8 days, measured from three technical replicates Statistical significance was determined using two-way ANOVA followed by Tukey’s multiple comparisons test. (D) Pseudocolor flow cytometry plots of Annexin V and propidium iodide (PI) staining in cells cultured in suspension for 4 days show increased apoptosis in MDA-Tpm2.1 cells. Quadrants reflect populations enriched for live(Q4), early apoptotic(Q3), and late apoptotic(Q2) or necrotic(Q1) cells based on Annexin V and PI signal intensities. (E) Mean ± SEM of total apoptotic cells (%) in attached (1 day) and suspension at 5 h, 1 day, 2 days, 4 days, and 8 days. Data represents two independent biological replicates with 2-3 technical replicate wells per condition. Statistical significance was determined using two-way ANOVA followed by Tukey’s multiple comparisons test. (F) Representative phase images of MDA-EV and MDA-Tpm2.1 cells migrated to the bottom well after 48 h in the trans-well chamber. Scale bar = 300 μm (G) Mean ± SD of the total number of migrated cells. Statistical significance was determined using unpaired t-test followed by Welch’s correction. (**) for p value < 0.01, (***) for p value < 0.001, (****) for p value <0.0001.
Transcriptomic profiling reveals distinct gene expression programs in apoptotic and non-apoptotic Tpm2.1-expressing cells
Tpm2.1 expression has been identified as a key regulator of mechanosensing in MDA-MB-231 cells. However, as shown above, despite Tpm2.1 overexpression, a substantial fraction of cells remains viable under non-adherent conditions, indicating incomplete restoration of anoikis sensitivity. To investigate how Tpm2.1 influences the transition to distinct physiological states, we examined the gene expression programs associated with apoptotic versus non-apoptotic responses during suspension culture.
At each time point, MDA-Tpm2.1 cells were separated into two subpopulations: Annexin V-negative (non-apoptotic, Tpm_NoAp) and Annexin V-positive (apoptotic, Tpm_Apop) cells. Cells were sorted using magnetic separation with an Annexin V Microbead Kit36, which isolates apoptotic cells by binding Annexin V-labeled microbeads followed by magnetic column selection (Fig. 2A).
We then performed time-resolved RNA sequencing of MDA-EV, MDA-Tpm2.1, Tpm_NoAp, and Tpm_Apop cells. Transcriptomic analysis revealed widespread gene expression changes over time and across conditions. A Spearman correlation heatmap (Fig. 2B) showed that the most distinct transcriptional divergence emerged at day 4, when Tpm_Apop cells displayed a gene expression profile markedly different from all other samples. Early time points (rigid and 5-hour suspension) showed minimal differences between MDA-EV and MDA-Tpm2.1 cells, suggesting that short-term detachment does not yet invoke a Tpm2.1-specific response. In contrast, longer suspension periods (≥ 1 day) revealed progressive separation of Tpm2.1-expressing cells from controls, with the strongest effect observed in the apoptotic subpopulation at day 4.
To confirm that these differences were not due to variation in Tpm2.1 expression itself, we measured Tpm2.1 transcript levels across all conditions and found them to be stable (Fig. S1E). Differential gene expression analysis, performed using the Novomagic platform (see “Methods”), revealed thousands of significantly altered genes when comparing Tpm_Apop to MDA-EV cells. By contrast, the Tpm_NoAp population showed only a few hundred differentially expressed genes relative to MDA-EV (Fig. 2C, D), suggesting that the apoptotic state in Tpm2.1-overexpressing cells is accompanied by a dramatic and distinct transcriptional shift.
Transcriptomic profiling of apoptotic and non-apoptotic subpopulations in Tpm2.1-expressing MDA-MB-231 cells under suspension conditions. (A) Schematic of the experimental design. MDA-MB-231 cells stably expressing an empty vector (MDA-EV) or Tpm2.1 (MDA-Tpm2.1) were cultured under adherent or suspension conditions. At each suspension time point (5 h, 1 day, 2 days, and 4 days), MDA-Tpm2.1 cells were separated into apoptotic (Annexin V-positive) and non-apoptotic (Annexin V-negative) subpopulations using Annexin V-conjugated magnetic microbeads, followed by transcriptomic profiling. (B) Spearman correlation heatmap of all RNA-seq samples across time points and conditions. Correlation strength is color-coded, with red indicating strong positive correlation and blue indicating strong negative correlation. The transcriptional profile of apoptotic Tpm2.1-expressing cells at day 4 (Tpm_Apop_4d) is most distinct from all other conditions. (C) Volcano plots showing differentially expressed genes between Tpm_NoAp and Tpm_Apop populations relative to MDA-EV at day 4. Red and green dots represent significantly upregulated and downregulated genes, respectively (adjusted p < 0.05, |log2FoldChange| > 1). (D) Venn diagram showing the number and overlap of differentially expressed genes across multiple comparisons: Tpm_NoAp vs. MDA-EV, Tpm_Apop vs. MDA-EV, MDA-Tpm2.1 (bulk) vs. MDA-EV, and MDA-EV under rigid conditions vs. day 4 suspension. Differential expression analysis was performed using Novogene’s Novomagic platform, employing the DESeq2 pipeline [adjusted p-value (padj) < 0.05 and |log2FoldChange| > 1] as significance thresholds.
Distinct transcriptional programs underlie survival and death in suspended MDA-MB-231 cells
To identify the signaling pathways associated with anoikis resistance or susceptibility, we performed Kyoto Encyclopedia of Genes and Genomes (KEGG)37,38,39,40 pathway enrichment analysis across all time points (5 h, 1d, 2d, and 4d) in MDA-EV, Tpm_NoAp, and Tpm_Apop. Despite all groups experiencing suspension stress, the three cell populations activated markedly different transcriptional programs that corresponded to their fate (Fig. 3A). The complete KEGG pathway enrichment results for all groups and time points are provided in the Supplementary Information (see also Data Availability).
MDA-EV cells, which proliferate in suspension, rapidly adapted to detachment by upregulating steroid hormone biosynthesis, likely reflecting lipid-based compensatory signaling41,42. This early adaptation was accompanied by suppression of cell-cell communication pathways (e.g., Hippo, gap junctions, tight junctions), suggestive of disengagement from multicellular coordination43,44,45. Over time, MDA-EV cells progressively reactivated ECM-receptor interaction46, focal adhesion47, and the PI3K-Akt48 signaling pathway, supporting anchorage-independent growth49,50,51. Notably, this occurred in parallel with downregulation of p53, DNA replication, and cell cycle pathways, indicating a decoupling from canonical adhesion-mediated checkpoint control52.
Tpm2.1-overexpressing cells that resisted anoikis (Tpm_NoAp) followed a markedly different trajectory. While they also suppressed cell cycle, p53, and Hippo pathways early on, they uniquely mounted a robust inflammatory and immune-related transcriptional response. These include upregulation of TNF, IL-17, NF-κB, and Toll-like receptor (TLR) signaling as early as 5 h post-detachment. Concomitantly, both Tpm_NoAp and Tpm_Apop cells showed early suppression of the MAPK pathway, known to modulate apoptosis53,54. However, MAPK signaling was selectively reactivated in Tpm_NoAp cells by day 4, potentially contributing to longer-term survival55.
A defining feature of the Tpm_NoAp population was the persistent upregulation of the cell adhesion molecules (CAMs) pathway, including genes such as ICAM1, ICAM2, CDH15, F11R, HLA-A, and HLA-B. These promote cell-cell adhesion and immune-related signaling and may collectively support multicellular clustering, enhancing resistance to anoikis56. This was further complemented by delayed activation of the PI3K-Akt pathway at 4 days, including elevated expression of ITGA5, ITGA11, ITGB4, CDKN1A, NTRK1, and LAMB3. Genes which mediate adhesion, survival, and tumor progression57,58,59. Together, these data suggest that anoikis resistance in Tpm_NoAp cells is mediated by an integrated transcriptional program that includes early immune priming, reinforced cell–cell connectivity, and late pro-survival signaling.
In contrast, Tpm_Apop cells failed to sustain these compensatory responses. Although they transiently activated inflammatory and steroid biosynthesis pathways, they did not reactivate PI3K-Akt or MAPK signaling at later stages. Instead, they showed persistent downregulation of focal adhesion, cell cycle, DNA repair, and RNA processing pathways. Notably, only Tpm_Apop cells upregulated the TLR pathway at day 4, potentially indicating activation of innate immune-like stress signals linked to cell death60,61. These features suggest a transcriptional trajectory consistent with irreversible commitment to apoptosis.
To further investigate the differential regulation of pro-survival circuits, we focused on the CAM and PI3K-Akt pathways, which emerged as central to the divergence between Tpm_NoAp and Tpm_Apop cells.
Analysis of the CAM pathway (Fig. 3B) revealed elevated expression of intercellular adhesion genes, including F11R, HLA-A, HLA-B, ICAM1, ICAM2, and CDH15, specifically in the Tpm_NoAp population by day 4. These genes support immune signaling and promote tight cell-to-cell adhesion62,63. Their selective upregulation in non-apoptotic cells, but not in the apoptotic population, suggests that collective cohesion may act as a compensatory survival mechanism in the absence of extracellular matrix attachment.
In parallel, expression profiling of the PI3K-Akt pathway (Fig. 3C) showed upregulation of integrin subunits (ITGA5, ITGA11, ITGB4) and additional adhesion and survival-related genes, including TNXB, CSF3, CDKN1A, NTRK1, and LAMB3, in MDA-EV and Tpm_NoAp cells. These genes were not enriched in the apoptotic group, reinforcing the idea that the PI3K–Akt axis is selectively activated in populations capable of long-term survival in suspension.
Together, these pathway-level analyses underscore how non-apoptotic cells engage both adhesive and signaling-based mechanisms to maintain viability after detachment, while apoptotic cells fail to activate these compensatory programs.
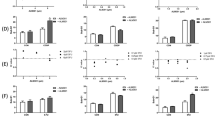
KEGG pathway enrichment analyses for MDA-MB-231 cancer cells induced by Tpm2.1 overexpression and suspension growth conditions. (A) KEGG pathway enrichment analysis of upregulated and downregulated genes in MDA-EV, Tpm_NoAp, and Tpm_Apop cells after 5 h and 4 days in suspension, relative to their respective cell populations cultured under attached (rigid) conditions used as controls (MDA-EV, and MDA-Tpm2.1, respectively). Red bars represent upregulated genes. Green bars represent downregulated genes. All pathways have adjusted p < 0.05, |log2FoldChange| > 1. (B,C) Heatmaps of differentially expressed genes from KEGG analysis for the Cell Adhesion Molecules (CAM) pathway (B) and the PI3K-Akt signaling pathway (C). Expression levels are color-coded, with red indicating high expression and green indicating low expression. Heatmaps were generated using an R script, with adjusted p-value (padj) < 0.05 and |log2FoldChange| > 1, applying Pearson correlation and the centroid method for clustering.
Cell-cell interactions contribute to anoikis resistance in Tpm2.1-expressing cells
The pathway enrichment analysis led us to hypothesize that upregulation of cell-cell interactions may promote survival in Tpm_NoAp cells during anchorage-independent growth.
We therefore seeded cells at low (1 × 103 cells/well), medium (5 × 104 cells/well), and high (1 × 106 cells/well) densities in low-attachment plates and measured apoptosis after 4 days in suspension (Fig. 4A, B). MDA-Tpm2.1 cells exhibited significantly higher apoptosis levels at low cell density, which was markedly reduced at higher densities. This indicates that cell-cell communication supports survival. In contrast, MDA-EV cells showed no appreciable change in apoptosis across different seeding densities.
Among the upregulated cell adhesion molecules, ICAM-1 showed the highest fold change in non-apoptotic MDA-Tpm2.1 cells under suspension conditions compared to adherent conditions (Fig. 3B). ICAM-1 is known to be overexpressed in circulating tumor cells and has been implicated in promoting breast cancer metastasis64. We therefore investigated whether ICAM-1 contributes functionally to the survival of MDA-Tpm2.1 cells in suspension. Inhibition of ICAM-1 led to increased apoptosis in MDA-Tpm2.1 cells at both 1 and 4 days in suspension. Under adherent conditions, ICAM-1 inhibition had no significant effect on day 1, though a modest increase in apoptosis was observed by day 4 (Fig. 4C, D).
Collectively, these findings suggest that enhanced cell-cell adhesion contributes to anoikis resistance in Tpm2.1-expressing cells, and that disrupting this interaction, particularly via ICAM-1 inhibition, can sensitize these cells to undergo apoptosis under anchorage-independent conditions.
Change in cell-cell interaction affects anoikis in MDA-Tpm2.1 cells. (A) Phase-contrast images of MDA-EV and MDA-Tpm2.1 cells cultured at three different cell densities for 4 days in suspension. MDA-EV cells are indicated by arrows, MDA-Tpm2.1 cell aggregates are highlighted by circles. Scale bar = 750 μm (B) Mean ± SD of total apoptotic cells (Annexin V+ only, PI+ only, and Annexin V+/PI+; %) in suspension cultures at different cell densities for 4 days. Statistical significance was determined using two-way ANOVA followed by Šídák’s multiple comparisons test. (C,D) Mean ± SD of total apoptotic cells (Annexin V+ only, PI+ only, and Annexin V+/PI+; %) in MDA-EV (C) and MDA-Tpm2.1 (D) cells treated with ICAM-1 inhibitor (ICAM-1-IN-1, 100nM) for 1 day or 4 days under suspension and attached conditions. The ICAM-1 inhibitor was added at the time of seeding. Statistical significance was determined using unpaired t-test followed by Welch’s correction and Holm-Šídák adjustment for multiple comparisons (α = 0.05). (**) for p value < 0.01, (***) for p value < 0.001, (****) for p value <0.0001.
Tpm2.1-mediated suppression of Akt and EphA2 signaling promotes anoikis in breast cancer cells
A second key distinction identified in our pathway enrichment analysis was the selective upregulation of the PI3K-Akt signaling pathway in MDA-EV and Tpm_NoAp cells, but not in Tpm_Apop cells, after 4 days in suspension. Activation of this pathway occurs through phosphorylation of Akt at Threonine 308 by PDK1 (3-phosphoinositide-dependent kinase-1) and at Serine 473 by mTOR, both of which are essential for full Akt activation65,66. Therefore, we sought to compare the levels of phosphorylated Akt and total Akt in MDA-EV and MDA-Tpm2.1 cells, under both suspension and adherent conditions.
Western blot analysis revealed that phospho-Akt (Thr308) levels were reduced in MDA-Tpm2.1 cells compared to MDA-EV cells during suspension culture, while no such difference was observed under adherent conditions. Total Akt expression decreased in both cell types in suspension relative to adhesion, but levels were comparable between MDA-EV and MDA-Tpm2.1 cells under each condition. These results indicate that Tpm2.1 suppresses Akt activation specifically in suspension, rather than altering its basal expression (Fig. 5A–C).
To assess the functional role of Akt signaling in anoikis resistance, we treated the cells with Afuresertib Hydrochloride, an Akt kinase inhibitor, for 4 days, and then measured apoptosis. Since our pathway enrichment analysis revealed that this pathway was upregulated after 4 days in suspension, we sought to determine whether the timing of Akt inhibition influences apoptotic outcomes. Therefore, the inhibitor was added at three time-points: during cell seeding (0 h), 1 day, and 2 days post-seeding. Both MDA-EV and MDA-Tpm2.1 cells exhibited increased apoptosis upon Akt inhibition. Notably, MDA-EV cells showed a more pronounced apoptotic response when the inhibitor was added at the time of seeding (Fig. 5D). In contrast, MDA-Tpm2.1 cells displayed similar levels of apoptosis regardless of the timing of inhibitor administration (Fig. 5E).
To further understand the mechanisms by which the Akt pathway supports anoikis resistance, we examined its downstream effectors. Heatmap analysis of the PI3K-Akt pathway revealed elevated expression of EphA2 in MDA-EV cells and in the non-apoptotic population of MDA-Tpm2.1 cells cultured in suspension for 4 days, compared to the apoptotic fraction of MDA-Tpm2.1 cells (Fig. 3C). Based on this observation, we examined EphA2 protein levels in MDA-EV and MDA-Tpm2.1 cells under both attached and suspension conditions. EphA2 expression decreased in suspension relative to attached conditions in both cell types and was further reduced in MDA-Tpm2.1 cells compared to MDA-EV cells under suspension conditions (Fig. 5F,G).
EphA2 can also be activated via a non-canonical mechanism involving Akt-mediated phosphorylation at serine 897 (Ser897). Western blot analysis revealed reduced levels of phospho-EphA2 (Ser897) in MDA-Tpm2.1 cells relative to MDA-EV cells under both attached and suspension conditions (Fig. S2). Supporting a pro-survival role for EphA2 signaling, treatment with an EphA2 kinase inhibitor significantly increased apoptosis in MDA-EV cells in suspension, to levels comparable to those observed in MDA-Tpm2.1 cells (Fig. 5H).
Together, these results suggest that Akt and EphA2 signaling promote anoikis resistance in breast cancer cells under anchorage-independent conditions and that suppression of these pathways by Tpm2.1 enhances apoptotic sensitivity in suspension.
Tpm2.1 expression downregulates the Akt and EphA2 pathways in MDA cells under suspension growth conditions. (A) Western blot showing the expression of phospho-Akt (T308) and total Akt in MDA-EV and MDA-Tpm2.1 cells cultured under suspension and attached conditions for 5 days. (B,C) Quantification of phospho-Akt (T308), N=3 (B), and total Akt levels, N=2 (C), in MDA-EV and MDA-Tpm2.1 cells under suspension and attached conditions for 5 days, presented as Mean ± SD. Statistical significance was determined using two-way ANOVA. (D,E) Quantification of total apoptotic cells (%) presented as Mean ± SD for MDA-EV (D) and MDA-Tpm2.1 (E) cells after treatment with Akt inhibitor (5 µM) for 4 days under suspension and attached conditions. The Akt inhibitor was added at the time of seeding (0 h), 1 day, and 2 days after seeding, as indicated on the X-axis of panels D and E. Statistical significance was determined using two-way ANOVA followed by Šídák’s multiple comparisons test. (F) Western blot showing the expression of EphA2 in MDA-EV and MDA-Tpm2.1 cells cultured under suspension and attached conditions for 5 days. (G) Quantification of EphA2 levels in MDA-EV and MDA-Tpm2.1 cells under suspension and attached conditions for 5 days. N=1. (H) Quantification of total apoptotic cells (%) presented as Mean ± SD for MDA-EV and MDA-Tpm2.1 cells after treatment with EphA2 kinase inhibitor (1.5nM) for 4 days under suspension and attached conditions. Statistical significance was determined using an unpaired t-test followed by Welch’s correction and Holm-Šídák adjustment for multiple comparisons (α = 0.05). Uncropped versions of the Western blots are provided in Supplementary Figures 3 and 4. (*) for p value <0.05, (**) for p value < 0.01, (***) for p value < 0.001, (****) for p value <0.0001.
Discussion
Resistance to anoikis is a hallmark of metastatic cancer cells, allowing them to survive outside their native tissue environment13. While prior studies have shown that restoring the actin-binding protein Tropomyosin 2.1 (Tpm2.1) reactivates cytoskeletal tension and impairs anchorage-independent growth7,12,67, whether this restoration is sufficient to eliminate survival capacity has remained unclear. Here, we demonstrate that although Tpm2.1 re-expression sensitizes metastatic breast cancer cells to apoptosis in suspension, the majority of cells remain viable by activating compensatory survival pathways. Importantly, we show that targeting key players of these pathways, specifically Akt, EphA2, and ICAM1, substantially increases anoikis and dramatically reduces the fraction of surviving cells. These findings not only clarify how anoikis resistance persists despite restored rigidity sensing but also uncover actionable strategies to eliminate resistant cells by disrupting survival adaptations. A summary schematic of these divergent cellular fates is presented in Fig. 6.
Our results reveal that restored rigidity sensing alone is insufficient to enforce uniform apoptotic commitment in matrix-deprived breast cancer cells. While Tpm2.1 expression disrupts canonical oncogenic signaling, a large proportion of cells engage adaptive mechanisms that support their survival. These include early activation of immune pathways, enhanced intercellular adhesion, and delayed upregulation of PI3K-Akt signaling. Together, these features suggest that rigidity sensing functions as a priming signal, lowering the threshold for apoptosis but not fully eliminating survival potential unless these secondary programs are blocked. This creates a therapeutic opportunity: combining cytoskeletal reprogramming with targeted inhibition of survival circuits can effectively eradicate otherwise persistent cell populations.
Conversely, cells that undergo anoikis fail to mount these compensatory responses. They remain transcriptionally repressed in key survival and adhesion modules, and uniquely upregulate innate immune signatures at later timepoints, potentially reflecting terminal stress signaling rather than adaptation. These divergent transcriptional trajectories highlight how survival under detachment stress is not solely dictated by mechanical sensing, but by the cell’s ability to engage dynamic gene regulatory programs.
Mechanistically, we show that Tpm2.1 suppresses Akt phosphorylation in suspension without affecting total Akt levels. Pharmacological inhibition of Akt enhances apoptosis in both control and Tpm2.1-expressing cells, with the latter showing sustained vulnerability across the time course. This suggests that restored rigidity sensing increases reliance on residual Akt activity, rendering cells more sensitive to its disruption.
We further implicate EphA2, a downstream target of Akt, in detachment survival. EphA2 expression and phosphorylation (Ser897) were reduced in Tpm2.1-expressing cells, and its inhibition robustly sensitized both MDA-EV and Tpm2.1-overexpressing cells to anoikis. These findings position EphA2 as a potent regulator of anchorage-independent survival and a compelling candidate for therapeutic intervention.
A complementary mechanism of resistance is mediated by intercellular adhesion. Tpm2.1-expressing cells form larger multicellular clusters in suspension, and transcriptomic analyses revealed upregulation of adhesion molecules, particularly ICAM1, in surviving cells. Functional inhibition of ICAM1 modestly but consistently increased apoptosis, supporting a model in which physical cell-cell interactions promote collective resilience.
In summary, our study provides a mechanistic framework for how rigidity sensing intersects with inflammatory, adhesive, and kinase-mediated survival programs in metastatic breast cancer. While Tpm2.1 re-expression primes cells for anoikis by disrupting oncogenic signaling, full commitment to apoptosis requires concurrent disruption of the compensatory pathways that enable survival in suspension. By functionally validating Akt, EphA2, and ICAM1 as key effectors of this resilience, we identify combination strategies that markedly enhance apoptosis and reduce anchorage-independent viability. These findings establish a roadmap for targeting mechanosensitive survival states in metastatic cancer.
Illustrated Working Model. Schematic model depicting the distinct signaling pathways activated in breast cancer cells (MDA-EV) and Tpm2.1-expressing cells (MDA-Tpm2.1), following their separation into non-apoptotic (TpmNoAp) and apoptotic (TpmApop) subpopulations after 5 h and 4 days in suspension. The model highlights subpopulation-specific pathway adaptations and the associated cellular fates under anchorage-independent conditions. Created in https://BioRender.com.
Materials and methods
Cell culture
MDA-MB-231 cells (wild-type, control, and Tpm2.1 overexpressed) were a generous gift from Dr. H. Wolfenson (Technion, Israel) and were originally obtained from ATCC68. Tpm2.1-overexpressing cells (MDA-Tpm2.1) were generated by lentiviral overexpression of Tpm2.1 in MDA-MB-231 cells. All Cells were maintained in DMEM supplemented with 10% (v/v) fetal bovine serum (FBS), 1% penicillin–streptomycin, and 2 µg/mL puromycin at 37 °C in a humidified incubator with 5% CO2.
To assess growth under non-adherent conditions, cells were cultured in ultra-low attachment plates or flasks (Corning, Cat. No. 3471 and 3814). Growth under suspension conditions serves as an alternative assay for cellular transformation and correlates strongly with results from soft agar assays31. For adherent conditions, cells were cultured on standard tissue culture-treated plates or flasks.
Separation of apoptotic and non-apoptotic cells
MDA-Tpm2.1 cells were cultured under suspension conditions in ultra-low attachment plates for varying time points (5 h, 1 day, 2 days, and 4 days). Apoptotic and non-apoptotic populations were separated using the Annexin V MicroBead Kit (Miltenyi Biotec) according to the manufacturer’s protocol, which isolates cells based on Annexin V binding, a marker of early apoptosis36.
Apoptosis measurement
Cells were cultured under adherent or suspension conditions for the indicated time periods. Adherent cells were harvested by trypsinization. All cells were collected by centrifugation, washed with DPBS, and resuspended in 100 µL Annexin V binding buffer. Apoptosis was assessed using Alexa Fluor™ 647-conjugated Annexin V (BD Pharmingen, Cat. No. 567356) and propidium iodide (PI; Invitrogen, Cat. No. BMS500F1-300), following the manufacturers’ protocols. Annexin V- and PI-positive populations were quantified by flow cytometry (BD FACSymphony A5) and analyzed using FlowJo software.
Cell growth assay
Cells (5 × 104 cells per well) were seeded in 6-well ultra-low attachment plates and cultured for 5 or 8 days. Cell viability was assessed using Trypan Blue exclusion, and live/dead cell counts were obtained using a DeNovix CellDrop FL cell counter. Fold change in cell number was calculated by dividing the number of live cells on Day 5 or Day 8 by the initial seeding density.
Transwell-migration assay
MDA-EV and MDA-Tpm2.1 cells (1 × 105 per well) were seeded in serum-free medium into the upper chamber of Transwell inserts (8 μm pore size; Corning Costar, catalog no. 3464) and incubated for 48 h at 37 °C in a humidified incubator with 5% CO2. The lower chamber contained 500 µl of medium supplemented with 10% fetal bovine serum (FBS) as a chemoattractant. After incubation, the inserts were removed, and cells that had migrated to the lower surface of the membrane were fixed with 4% paraformaldehyde and stained with Hoechst dye. Phase-contrast and DAPI fluorescence images were acquired, and migrated cells were quantified using ImageJ (NIH).
Inhibitor treatments
Cells were treated with the following small-molecule inhibitors: an Akt kinase inhibitor (Afuresertib Hydrochloride, 5 µM; MedChemExpress, Cat. No. HY-15727 A/CS-3385), an ICAM-1 inhibitor (ICAM-1-IN-1, 100 nM; MedChemExpress, Cat. No. HY-U00003), and EphA2 kinase inhibitor (ALW-II-41-27, 1.5nM; MedChemExpress, Cat. No. HY18007), as indicated in each experiment.
Western blot
Cells were cultured in suspension (ultra-low attachment plates) or adherent (tissue culture plates) conditions for 5 days. Total protein was extracted using RIPA buffer (Cell Signaling Technology, Cat. No. 9806 S) supplemented with protease inhibitors (Sigma-Aldrich, Cat. No. 11697498001) and incubated on ice for 45 min. Lysates were mixed with Laemmli buffer, separated on 4–20% gradient SDS–PAGE gels (Bio-Rad, Cat. No. 4561096) at 120 V for 90 min, and transferred to nitrocellulose membranes using the Trans-Blot Turbo Transfer System (Bio-Rad).
Membranes were blocked in 5% BSA in 1× TBST for 1 h at room temperature, followed by overnight incubation at 4 °C with primary antibodies. After washing, membranes were incubated with HRP-conjugated secondary antibodies for 1 h at room temperature. The following primary antibodies were used: Anti-Tropomyosin (clone TM311, Sigma-Aldrich, T2780; 1:1000), Akt (pan) (Cell Signaling Technology, C67E7; 1:1000), phospho-Akt (T308) (Cell Signaling Technology, 244F9; 1:500), EphA2 (D4A2) (Cell Signaling Technology, 6997 S; 1:1000), phoshpho-EphA2(S897) (Cell Signaling Technology, 6347 S; 1:1000) and Cyclophilin B (Cell Signaling Technology, 43603 S; 1:1000), used as a loading control. Secondary antibodies included HRP-conjugated anti-mouse IgG (EMD Millipore, AP160P) and anti-rabbit IgG (Santa Cruz Biotechnology, sc-2357).
RNA isolation
Total RNA was isolated using the RNeasy Mini Kit (Qiagen®, Germantown, MD, USA; Cat. #74104) according to the manufacturer’s protocol. RNA concentration and purity were assessed using a NanoDrop Spectrophotometer (Thermo Fisher Scientific). A total of 28 RNA samples, representing 14 experimental conditions with biological duplicates, were submitted to Novogene Co. Ltd. (Nanjing, China) for library preparation and high-throughput sequencing.
RNA sequencing and analysis
Library preparation for transcriptome sequencing- non strand specific library
Messenger RNA was purified from total RNA using poly-T oligo-attached magnetic beads. After fragmentation, the first strand cDNA was synthesized using random hexamer primers followed by the second strand cDNA synthesis. The library was ready after end repair, A-tailing, adapter ligation, size selection, amplification, and purification. The library was checked with Qubit and real-time PCR for quantification and bioanalyzer for size distribution detection.
Clustering and sequencing
After library quality control, different libraries were pooled based on the effective concentration and targeted data amount, then subjected to Illumina sequencing on the NovaSeq6000 Platform, using paired-end 150 bp read length. The basic principle of sequencing is “Sequencing by Synthesis”, where fluorescently labeled dNTPs, DNA polymerase, and adapter primers are added to the sequencing flow cell for amplification. As each sequencing cluster extends its complementary strand, the addition of each fluorescently labeled dNTP releases a corresponding fluorescence signal. The sequencer captures these fluorescence signals and converts them into sequencing peaks through computer software, thereby obtaining the sequence information of the target fragment.
RNA analysis
Differential expression analysis was performed using Novogene’s Novomagic platform, employing the DESeq2 pipeline with adjusted p-value (padj) < 0.05 and |log2FoldChange| > 1 as significance thresholds.
Heatmap produced by an R script, with padj < 0.05 and |log2Fold|>1, applying Spearman correlation and the centroid method for clustering.
Data availability
The RNA-seq data discussed in this publication have been deposited in NCBI’s Gene Expression Omnibus (Edgar et al., 2002) and are accessible through GEO Series accession number GSE304274 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE304274).
References
Freedman, V. H. & Shin, S. I. Cellular tumorigenicity in nude mice: correlation with cell growth in semi-solid medium. Cell 3, 355–359 (1974).
Pickup, M. W., Mouw, J. K. & Weaver, V. M. The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. 15, 1243–1253 (2014).
Sheetz, M. A Tale of two states: normal and Transformed, with and without rigidity sensing. Annu. Rev. Cell. Dev. Biol. 35, 169–190 (2019).
Welch, D. R. & Hurst, D. R. Defining the hallmarks of metastasis. Cancer Res. 79, 3011–3027 (2019).
Fares, J., Fares, M. Y., Khachfe, H. H., Salhab, H. A. & Fares, Y. Molecular principles of metastasis: a hallmark of cancer revisited. Signal. Transduct. Target. Therapy. 5, 28 (2020).
Discher, D. E., Janmey, P. & Wang, Y. Tissue cells feel and respond to the stiffness of their substrate. Science 310, 1139–1143 (2005).
Yang, B. et al. Stopping transformed cancer cell growth by rigidity sensing. Nat. Mater. 19, 239–250 (2020).
Saraswathibhatla, A., Indana, D. & Chaudhuri, O. Cell-extracellular matrix mechanotransduction in 3D. Nat. Rev. Mol. Cell. Biol. 24, 495–516 (2023).
Meredith, J. E., Fazeli, B. & Schwartz, M. A. The extracellular matrix as a cell survival factor. Mol. Biol. Cell. 4, 953–961 (1993).
Frisch, S. M. & Francis, H. Disruption of epithelial cell-matrix interactions induces apoptosis. J. Cell. Biol. 124, 619–626 (1994).
Taddei, M., Giannoni, E., Fiaschi, T. & Chiarugi, P. Anoikis: an emerging hallmark in health and diseases. J. Pathol. 226, 380–393 (2012).
Wolfenson, H., Yang, B. & Sheetz, M. P. Steps in mechanotransduction pathways that control cell morphology. Annu. Rev. Physiol. 81, 585–605 (2019).
Dai, Y. et al. Anoikis resistance–protagonists of breast cancer cells survive and metastasize after ECM detachment. Cell. Commun. Signal. 21, 190 (2023).
Galbraith, C. G., Yamada, K. M. & Sheetz, M. P. The relationship between force and focal complex development. J. Cell. Biol. 159, 695–705 (2002).
Sergey, V., Plotnikov, Ana, M., Pasapera, B., Sabass, C. M. & Waterman Force fluctuations within focal adhesions mediate ECM-Rigidity sensing to guide directed cell migration. Cell 151, 1513–1527 (2012).
Elosegui-Artola, A. et al. Roca-Cusachs, rigidity sensing and adaptation through regulation of integrin types. Nat. Mater. 13, 631–637 (2014).
Iskratsch, T., Wolfenson, H. & Sheetz, M. P. Appreciating force and shape—the rise of mechanotransduction in cell biology. Nat. Rev. Mol. Cell. Biol. 15, 825–833 (2014).
Schiller, H. B. & Fässler, R. Mechanosensitivity and compositional dynamics of cell-matrix adhesions. EMBO Rep. 14, 509–519 (2013).
Vogel, V. & Sheetz, M. P. Cell fate regulation by coupling mechanical cycles to biochemical signaling pathways. Curr. Opin. Cell Biol. 21, 38–46 (2009).
Zhang, Y. H., Zhao, C. Q., Jiang, L. S. & Dai, L. Y. Substrate stiffness regulates apoptosis and the mRNA expression of extracellular matrix regulatory genes in the rat annular cells. Matrix Biol. 30, 135–144 (2011).
Meacci, G. et al. Sheetz, α-Actinin links extracellular matrix rigidity-sensing contractile units with periodic cell-edge Retractions. Mol. Biol. Cell. 27, 3471–3479 (2016).
Kumar, A., Placone, J. K. & Engler, A. J. Understanding the extracellular forces that determine cell fate and maintenance. Development 144, 4261–4270 (2017).
Zhou, D. W., Lee, T. T., Weng, S., Fu, J. & García, A. J. Effects of substrate stiffness and actomyosin contractility on coupling between force transmission and vinculin-paxillin recruitment at single focal adhesions. Mol. Biol. Cell. 28, 1901–1911 (2017).
Goldberg, G. S. et al. Global effects of anchorage on gene expression during mammary carcinoma cell growth reveal role of tumor necrosis Factor-related Apoptosis-inducing ligand in Anoikis1. Cancer Res. 61, 1334–1337 (2001).
Merten, O. W. Advances in cell culture: anchorage dependence. Philos. Trans. R Soc. Lond. B Biol. Sci. 370, 20140040 (2015).
Wolfenson, H. et al. Tropomyosin controls sarcomere-like contractions for rigidity sensing and suppressing growth on soft matrices. Nat. Cell. Biol. 18, 33–42 (2016).
Ma, Z., Myers, D. P., Wu, R. F. & Nwariaku, F. E. Terada, p66Shc mediates Anoikis through RhoA. J. Cell. Biol. 179, 23–31 (2007).
Singh, P., Ravanan, P. & Talwar, P. Death associated protein kinase 1 (DAPK1): A regulator of apoptosis and autophagy. Front. Mol. Neurosci. (2016).
Tijore, A. et al. Selective killing of transformed cells by mechanical stretch. Biomaterials 275, 120866 (2021).
Wu, Z. et al. TPM2 attenuates progression of prostate cancer by blocking PDLIM7-mediated nuclear translocation of YAP1. Cell. Bioscience. 13, 39 (2023).
Rotem, A. et al. Alternative to the soft-agar assay that permits high-throughput drug and genetic screens for cellular transformation. Proc. Natl. Acad. Sci. USA 112, 5708–5713 (2015).
Park, J. Y. et al. Development of suspension cell culture model to mimic Circulating tumor cells. Oncotarget 9, 622–640 (2018).
Izar, B. & Rotem, A. GILA, a replacement for the Soft-Agar assay that permits High-Throughput drug and genetic screens for cellular transformation. Curr Protoc. Mol. Biol 116, 282812–282821 (2016).
Yang, J. et al. Weinberg, Twist, a master regulator of Morphogenesis, plays an essential role in tumor metastasis. Cell 117, 927–939 (2004).
Minn, A. J. et al. Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 (2005).
Lobascio, A. M., Klinger, F. G. & De Felici, M. Isolation of apoptotic mouse fetal oocytes by AnnexinV assay. Int. J. Dev. Biol. 51 (2007).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M. Post-genome Informatics (OUP Oxford, 2000).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–d677 (2025).
Antalis, C. J., Uchida, A., Buhman, K. K. & Siddiqui, R. A. Migration of MDA-MB-231 breast cancer cells depends on the availability of exogenous lipids and cholesterol esterification. Clin. Exp. Metastasis. 28, 733–741 (2011).
Rodrigues dos Santos, C. et al. LDL-cholesterol signaling induces breast cancer proliferation and invasion. Lipids Health Dis. 13, 16 (2014).
Aasen, T., Mesnil, M., Naus, C. C., Lampe, P. D. & Laird, D. W. Gap junctions and cancer: communicating for 50 years. Nat. Rev. Cancer. 16, 775–788 (2016).
Kyuno, D. et al. Role of tight junctions in the epithelial-to-mesenchymal transition of cancer cells. Biochim. Et Biophys. Acta (BBA) - Biomembr. 1863, 183503 (2021).
Baroja, I., Kyriakidis, N. C., Halder, G. & Moya, I. M. Expected and unexpected effects after systemic Inhibition of Hippo transcriptional output in cancer. Nat. Commun. 15, 2700 (2024).
Jinka, R., Kapoor, R., G Sistla, P., A Raj, T. & Pande, G. Alterations in Cell-Extracellular matrix interactions during progression of cancers. Int. J. Cell. Biol. 2012, 219196 (2012).
Xu, L. H. et al. The focal adhesion kinase suppresses transformation-associated, anchorage-independent apoptosis in human breast cancer cells. Involvement of death receptor-related signaling pathways. J. Biol. Chem. 275, 30597–30604 (2000).
Akca, H., Demiray, A., Tokgun, O. & Yokota, J. Invasiveness and anchorage independent growth ability augmented by PTEN inactivation through the PI3K/AKT/NFkB pathway in lung cancer cells. Lung Cancer. 73, 302–309 (2011).
Ivetac, I. et al. Regulation of PI(3)K/Akt signalling and cellular transformation by inositol polyphosphate 4-phosphatase‐1. EMBO Rep. 10, 487–493 (2009).
Tsirtsaki, K. & Gkretsi, V. The focal adhesion protein Integrin-Linked kinase (ILK) as an important player in breast cancer pathogenesis. Cell. Adh Migr. 14, 204–213 (2020).
Huh, H. D. et al. Reprogramming anchorage dependency by adherent-to-suspension transition promotes metastatic dissemination. Mol. Cancer. 22, 63 (2023).
Paoli, P., Giannoni, E. & Chiarugi, P. Anoikis molecular pathways and its role in cancer progression. Biochim. Et Biophys. Acta (BBA) - Mol. Cell. Res. 1833, 3481–3498 (2013).
Zhang, W. & Liu, H. T. MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Cell Res. 12, 9–18 (2002).
Yue, J. & López, J. M. Understanding MAPK signaling pathways in apoptosis. Int. J. Mol. Sci. 21 (2020).
Renshaw, M. W., Ren, X. D. & Schwartz, M. A. Growth factor activation of MAP kinase requires cell adhesion. Embo j. 16, 5592–5599 (1997).
Zhong, X. & Rescorla, F. J. Cell surface adhesion molecules and adhesion-initiated signaling: Understanding of Anoikis resistance mechanisms and therapeutic opportunities. Cell. Signal. 24, 393–401 (2012).
Zhang, H., Zhang, L. & Lu, M. Inhibition of integrin subunit alpha 11 restrains gastric cancer progression through phosphatidylinositol 3-kinase/Akt pathway. Bioengineered 12, 11909–11921 (2021).
Huang, G. et al. The mechanism of ITGB4 in tumor migration and invasion. Front. Oncol. 14 (2024).
Li, H. et al. A novel oncogenic and Drug-Sensitive KIF5B-NTRK1 fusion in lung adenocarcinoma. Curr. Oncol. 31, 6621–6631 (2024).
Aliprantis, A. O., Yang, R. B., Weiss, D. S., Godowski, P. & Zychlinsky, A. The apoptotic signaling pathway activated by Toll-like receptor-2. Embo j. 19, 3325–3336 (2000).
Yi, H., Patel, A. K., Sodhi, C. P., Hackam, D. J. & Hackam, A. S. Novel role for the innate immune receptor Toll-Like receptor 4 (TLR4) in the regulation of the Wnt signaling pathway and photoreceptor apoptosis. PLOS ONE. 7, e36560 (2012).
Czubak-Prowizor, K., Babinska, A. & Swiatkowska, M. The F11 receptor (F11R)/Junctional adhesion Molecule-A (JAM-A) (F11R/JAM-A) in cancer progression. Mol. Cell. Biochem. 477, 79–98 (2022).
Guerra-Espinosa, C., Jiménez-Fernández, M., Sánchez-Madrid, F. & Serrador, J. M. ICAMs in Immunity, intercellular adhesion and communication. Cells 13 (2024).
Taftaf, R. et al. ICAM1 initiates CTC cluster formation and trans-endothelial migration in lung metastasis of breast cancer. Nat. Commun. 12, 4867 (2021).
Manning, B. D., Toker, A. & Signaling, A. K. T. P. K. B. Navigating Netw. Cell 169, 381–405 (2017).
He, Y. et al. Targeting PI3K/Akt signal transduction for cancer therapy. Signal. Transduct. Target. Therapy. 6, 425 (2021).
Desouza-Armstrong, M., Gunning, P. W. & Stehn, J. R. Tumor suppressor Tropomyosin Tpm2.1 regulates sensitivity to apoptosis beyond Anoikis characterized by changes in the levels of intrinsic apoptosis proteins. Cytoskeleton 74, 233–248 (2017).
Shi, L., Nadjar-Boger, E., Jafarinia, H., Carlier, A. & Wolfenson, H. YAP mediates apoptosis through failed integrin adhesion reinforcement. Cell Rep. 43 (2024).
Acknowledgements
The authors would like to thank Dr. H. Wolfenson (Technion, Israel), Dr. Elisabeth Nadjar-Boger, Lidan Shi, and Malak Amer for kindly providing the MDA-MB-231 cell lines and for their helpful discussions. We are also grateful to Emma Pfortmiller and Dr. Michelle Ward (UTMB, USA) for their assistance in generating heatmaps and for their valuable input. Special thanks to Meredith Weglarz and the UTMB Flow Cytometry & Cell Sorting Core for their support with cytometry measurements and insightful discussions.
Funding
This research was supported by UTMB startup funds and the Cancer Prevention & Research Institute of Texas (CPRIT) grant ID RR210018 to GN, a CPRIT scholar. Additional support was provided through postdoctoral fellowships from the Council for Higher Education of Israel and Bar-Ilan University, Israel, to AGV.
Author information
Authors and Affiliations
Contributions
A.G.V. and N.D. conceptualization, data curation, formal analysis, investigation, methodology, writing (original draft). A.G.V. conducted the separation experiments and RNA isolation. N.D. conducted the Western blot experiments, Inhibitor treatments, and apoptosis measurements. G.N. conceptualization, project administration, supervision, writing (review and editing). We wish to honor our late co-author, Dr. Mike P. Sheetz, whose foundational contributions and mentorship were integral to this work. He unfortunately passed away before reviewing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vivante, A.G., Dwivedi, N., Sheetz, M.P. et al. Targeting the Akt–EphA2 axis and cell–cell adhesion enhances anoikis sensitivity in cancer cells. Sci Rep 16, 6197 (2026). https://doi.org/10.1038/s41598-026-37327-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37327-y