Abstract
Co-infections involving human metapneumovirus (hMPV), respiratory syncytial virus (RSV), and influenza A virus (IAV) often exacerbate disease severity in vulnerable populations. Here, we employed a structure-based immunoinformatics approach to design a multi-epitope subunit vaccine targeting these pathogens. The construct incorporated two epitopes each for cytotoxic T lymphocytes (CTLs), helper T lymphocytes (HTLs), and B cells, derived from the fusion proteins of hMPV and RSV, as well as the neuraminidase protein of IAV. These epitopes were linked with an adjuvant and optimized spacers to enhance immunogenicity and structural stability. Structural modeling confirmed correct folding, and molecular docking predicted a stable interaction with Toll-Like Receptor 4 (TLR4) − 277.43 kcal/mol. Molecular dynamics simulations indicated a compact and stable complex with restricted conformational motions, while MM/GBSA analysis yielded a favorable binding free energy (–121.72 kcal/mol) dominated by electrostatic and van der Waals interactions. Immune simulations predicted strong humoral and cellular responses, including high antibody titers, IFN-γ and IL-2 production, and durable memory formation. Codon optimization achieved a codon adaptation index (CAI) of 0.98 and a GC content of 51.24%, suggesting efficient expression in Escherichia coli. These findings highlight the construct as a structurally stable, immunogenic, and expression-ready vaccine candidate, warranting experimental validation against hMPV, RSV, and IAV.
Similar content being viewed by others
Introduction
Co-infections are characterized by the simultaneous invasion and replication of multiple distinct pathogenic organisms within a single host, which can modulate host-pathogen interactions, alter immune responses, and complicate both the clinical and therapeutic management of infectious diseases1,2,3. Viral co-infections often involve complex interactions such as synergy, interference, mutual dependence, or competitive exclusion. Among these, interference between one virus suppresses the replication of another is commonly observed4,5.
Epidemiological studies report that co-infection with multiple respiratory viruses occurs in approximately 10–20% of clinical cases, highlighting its clinical significance6,7. However, the exact impact of viral co-infection on disease progression remains debated, as some studies report exacerbated severity or altered clinical course5,8, while others find no significant differences6.
Respiratory viral infections represent a major global health burden. Influenza alone causes nearly one billion cases yearly, leading to 3–5 million hospitalizations and an estimated 290,000–650,000 deaths, particularly among vulnerable populations such as children and the elderly9. Beyond mortality, these infections can also have a financial impact on patients, families, and the healthcare system10. Acute respiratory infections represent one of the leading causes of mortality among children worldwide11,12, and a large fraction of these infections is attributable to viral pathogens, especially human metapneumovirus (hMPV), respiratory syncytial virus (RSV), and influenza viruses13.
Human respiratory syncytial virus (hRSV) is an enveloped, negative-sense, single-stranded RNA virus belonging to the Paramyxoviridae family and the Pneumovirus genus. Its size ranges from 100 to 200 nm in diameter and exhibits either spherical or filamentous morphology14. Its single-stranded RNA genome is encapsidated and linked with a lipid bilayer that integrates the surface glycoproteins G (responsible for receptor binding) and F (facilitating membrane fusion). The virus makes several proteins from its genome. The hRSV genome encodes multiple proteins, including nonstructural proteins NS1, NS2, and M2, a phosphoprotein (P), and the RNA-dependent RNA polymerase (L), all important for viral replication and transcription15. hRSV is one of the leading viral pathogens globally, mostly responsible for acute lower respiratory tract infections in infants, young children, and the elderly16. It infects both the upper and lower parts of the airway. It can block airflow, cause bronchiolitis, and lead to apnea, pneumonia, or even respiratory failure16.
Similarly, human metapneumovirus (hMPV), genetically divided into two subgroups A and B, is a major cause of respiratory tract infections across age groups and often co-circulates with RSV and influenza. Co-infection involving hMPV, RSV, and IAV has been reported to elicit more complex and severe immune responses than single virus infections17. These viruses may interact competitively or synergistically, disrupting host immune regulation and resulting in delayed viral clearance, prolonged disease duration, and a heightened risk of complications, particularly in immunocompromised individuals18.
Both cellular and humoral immune responses are essential for antiviral defense. CD8+ cytotoxic T lymphocytes (CTLs) play a pivotal role in clearing infected cells, while CD4+ helper T lymphocytes (HTLs) assist in antibody production and coordinate adaptive immunity. CD4+ T cells also contribute to antiviral defense by secreting interferon-gamma (IFN-γ), a key cytokine in the antiviral response. These mechanisms highlight the importance of eliciting both arms of the immune system in vaccine development19,20.
Immunoinformatics, an interdisciplinary field combining immunology and computational biology, enables the prediction and modeling of immune responses using in silico tools. It offers a rapid and cost-effective alternative to conventional laboratory methods for identifying vaccine targets. Notably, this approach proved highly valuable during the COVID-19 pandemic, as it accelerated the development of subunit vaccines against SARS-CoV-221,22.
In the context of viral co-infections, immunoinformatics enables the identification of conserved antigenic epitopes, prediction of CTL, HTL, and B-cell responses, and rational design of multi-epitope subunit vaccines (MESVs)16,17. Molecular-level simulations of host-pathogen interactions further enhance our understanding of immune complexity, supporting the development of targeted immunotherapies and multi-pathogen vaccine strategies17.
In this study, an immunoinformatics-based strategy was employed to design multi-epitope subunit vaccines (MESVs) targeting conserved regions of fusion and neuraminidase proteins from RSV, hMPV, and IAV. Neuraminidase (NA) plays a key role in the release of influenza virions by cleaving sialic acid residues, and immune responses directed against this protein can lessen viral shedding and disease impact while offering broader protection that is less influenced by antigenic drift23. Current research and health authorities highlight NA-based antigens as valuable candidates for next-generation “universal” influenza vaccines because of their capacity to induce cross-protective neuraminidase-inhibiting antibodies24. The fusion (F) protein of hMPV and RSV is a highly conserved antigen crucial for viral entry and a dominant target of neutralizing antibodies, making it ideal for vaccine development. Stabilizing the F protein in its prefusion conformation significantly enhances immunogenicity, enabling potent and cross-protective immune responses observed in both clinical and preclinical studies25. Computational tools were used to predict CTL, HTL, and B-cell epitopes, which were linked with appropriate adjuvants and spacers. Structural modeling, molecular dynamics simulations, and TLR-4 docking analyses validated the stability, immunogenic potential, and receptor-binding affinity of the designed construct. Compared to conventional vaccine approaches, this in silico strategy offers enhanced safety, precision, and cost-effectiveness for combating viral co-infections26.
Methodology
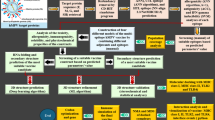
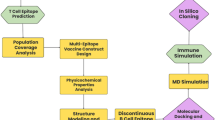
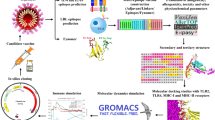
Figure 1 illustrates the overall computational workflow employed to design and evaluate the chimeric multi-epitope vaccine construct against hMPV, RSV, and IAV. The process began with the retrieval of complete proteomes of the target viruses from the NCBI database, followed by immunoinformatics-based prediction of conserved CTL, HTL, and B-cell epitopes using NetCTL, IEDB, and ABCPred servers. The shortlisted epitopes were evaluated for key physicochemical properties such as molecular weight, half-life, instability index, aliphatic index, and GRAVY score to ensure stability and immunogenic potential. These epitopes were then assembled into a chimeric multi-epitope subunit vaccine (CMESV) construct, incorporating suitable linkers and an adjuvant sequence to enhance immune responsiveness. The tertiary structure of the vaccine model was built and validated through ProSA-web, Ramachandran plot analysis, and iMODS-based structural refinement. The final construct was subjected to molecular docking with TLR4 to assess receptor binding interactions, followed by molecular dynamics (MD) simulations for conformational stability, including PCA, RMSD, RMSF, and radius of gyration (Rg) analyses.
Overview of the computational-based steps used to design a chimeric multi-epitope vaccine against hMPV, RSV, and IAV.
Proteins sequences retrieval
The full-length proteomes of Human Respiratory Syncytial Virus (RSV), Human Metapneumovirus (hMPV), and Influenza A Virus (IAV), comprising nine, ten, and fourteen proteins, respectively were retrieved in FASTA format from the NCBI database (https://www.ncbi.nlm.nih.gov/) using the accession numbers of RSV (KF826817.1, KF826821.1 and KF530269.1), hMPV (KJ627434.1, KJ627437.1 and KJ627436.1), and IAV (CY146835.1 and EF190976.1) and saved for subsequent analysis to identify highly antigenic proteins for chimeric multiepitope subunit vaccine development. Individual protein accession numbers used in this study are listed in Supplementary Fig. S (1–3) and Table S (1–3).
Prediction of antigenic and non-toxic proteins
The complete proteome of the Human respiratory syncytial virus (RSV), Human Metaneumovirus (hMPV), and Influenza A virus (IAV) (Nine, ten, and fourteen proteins respectively) were submitted to the VaxiJen v2.0 server (http://www.ddgpharmfac.net/vaxijen/VaxiJen/VaxiJen.html) to find out their antigenicity. The fusion (F) proteins of RSV and hMPV were selected because they are highly conserved, surface-exposed proteins essential for viral entry and major targets of neutralizing antibodies and T-cell responses. Similarly, the neuraminidase (NA) of IAV plays a vital role in viral release and cross-protective immunity. These proteins showed higher antigenicity scores (> 0.5) compared to other viral proteins, showing their better potential to induce strong and broad immune responses, thus making them ideal candidates for chimeric vaccine design. The ToxDL server (http://www.csbio.sjtu.edu.cn/bioinf/ToxDL/index.html) was utilized to evaluate the potential toxicity of the protein. This tool applies an interpretable deep learning based algorithm that classifies proteins into two categories, i.e., toxic or non-toxic, based on their sequence features.
Identification of the conserved regions in antigenic proteins
The highly antigenic fusion protein of RSV, hMPV, and neuraminidase of IAV were fed to Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/) online server for the identification of the conserved regions. The predicted epitopes for B and T cells were investigated in the aligned retrieved fusion protein of RSV, hMPV, and neuraminidase of IAV for conservation. The highly conserved region was chosen for the selection of each CTL, HTL, and BCL epitope.
Prediction and screening of cytotoxic T-lymphocytes (CTL) epitopes
CTL epitopes help the immune system remove virus-infected cells and lower the viral load. To find these, we used the NetCTL 1.2 server (https://services.healthtech.dtu.dk/services/NetCTL-1.2/) to predict epitopes from the fusion proteins of RSV and hMPV, and the neuraminidase of IAV. The aforementioned tool checks MHC class I binding, proteasome cleavage sites, and TAP transport. We selected epitopes with a score above 0.75.
Prediction of HTL epitopes
We used the Immune Epitope Database (IEDB) (http://tools.iedb.org/mhcii/) to find helper T cell (HTL) epitopes from the fusion proteins of RSV and hMPV, and the neuraminidase of IAV. Epitopes with the lowest percentile ranks showing strong MHC class II binding were chosen for the vaccine design.
Prediction of B-cell epitopes
B-cell epitopes were predicted using the ABCPred server (http://crdd.osdd.net/raghava/abcpred/), which utilizes a recurrent neural network-based algorithm to identify sequences likely to elicit antibody responses27. Epitopes were selected based on high prediction scores, with a threshold value of 0.80 applied to ensure the inclusion of highly probable antigenic regions.
Multiple epitope vaccine designing and evaluation
The selected epitopes comprising cytotoxic T lymphocytes (CTLs), helper T lymphocytes (HTLs), and B-cell epitopes derived from the fusion proteins of RSV and hMPV, as well as the neuraminidase of IAV, were assembled into a chimeric multi-epitope subunit vaccine construct. To ensure proper immunogenic presentation and structural flexibility, AAY, GPGPG, and KK linkers were employed to join CTL, HTL, and B-cell epitopes, respectively. Moreover, the human-derived β-defensin-2 was selected as an adjuvant for its potent antimicrobial and immunomodulatory properties, acting as a TLR4 agonist that promotes dendritic cell maturation and cytokine secretion crucial for adaptive immune activation. Its incorporation enhances antigen presentation and stimulates robust humoral and cellular immune responses. To further optimize immune stimulation and structural stability, β-defensin-2 was conjugated to the N-terminus of the vaccine construct via an EAAK linker, ensuring spatial separation and efficient epitope exposure. This strategic design strengthens the overall immunogenic efficacy of the multi-epitope vaccine against viral co-infections28.
Antigenicity and allergenicity of construct
We used the VaxiJen v2.0 server (https://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html) to test if the vaccine is likely to activate an immune response. A score above 0.4 was considered antigenic. We also checked allergenicity using AllerTOP v2.0 (https://www.ddg-pharmfac.net/AllerTOP/) to make sure the vaccine was safe and not likely to cause allergies.
Physicochemical properties of the construct
We used the ProtParam tool (https://web.expasy.org/protparam/) on the ExPASy server to analyze the vaccine’s properties, including amino acid content, molecular weight, pI, stability, half-life, aliphatic index, and GRAVY score.
3-D structure prediction and validation
The accurate 3D structure of the proteins is crucial for vaccine development, as it enables precise epitope mapping, stable construct modeling, and effective prediction of antigen-receptor interactions essential for immune activation. Thus, we used the Robetta server (https://robetta.bakerlab.org/) to model the 3D structure of the vaccine using its amino acid sequence in FASTA format. The chosen model was checked with ProSA-Web (https://prosa.services.came.sbg.ac.at/prosa.php) and PROCHECK (https://saves.mbi.ucla.edu/).
Population coverage analysis
The population coverage of the selected epitopes was evaluated using the IEDB Population Coverage tool (https://tools.iedb.org/population/). Predicted HLA-binding epitopes were assessed to estimate their potential recognition among global and regional human populations. The analysis incorporated the frequency distribution of HLA class I and class II alleles across diverse ethnic groups. Overall and overlapping population coverages were computed to determine the vaccine construct’s ability to induce broad immune responses worldwide.
Molecular docking with TLR4
Effective stimulation of the human immune response requires the vaccine to interact with host immune receptors. Molecular docking serves as a crucial in silico approach to predict the binding affinity and three-dimensional (3-D) orientation between a ligand and its target receptor. For this purpose, the 3-D structure of Toll-like receptor 4 (TLR-4; PDB ID: 2Z63) was retrieved from the RCSB Protein Data Bank (https://www.rcsb.org/). Before docking, all heteroatoms and water molecules were removed using PyMOL to prepare the receptor structure. The refined TLR-4 structure and the modeled vaccine concept were submitted to the HDOCK server (http://hdock.phys.hust.edu.cn/) to evaluate their binding interactions. The top-ranked docked complex, determined based on docking score, was selected for further structural and interaction analysis by using online server (https://bio.tools/pdbsum_generate).
Codon optimization and vaccine in silico cloning
Codon optimization is a computer-based method used to improve protein expression by adjusting codons to match the host organism. We used the JCat tool (http://www.jcat.de/) to convert the vaccine’s amino acid sequence into a DNA sequence optimized for E. coli K12. JCat also gave CAI and GC content values to check if the gene would express well. NdeI and XhoI sites were added to the ends of the sequence for cloning into the pET-28a(+) vector. The cloning setup was checked using SnapGene.
Immune simulation
The C-immSim online server (https://kraken.iac.rm.cnr.it/C-IMMSIM/) was used for immune simulation to assess the human immune response to the constructed vaccine. The aforementioned server evaluated the antigenicity of the developed vaccine and its capacity to elicit immunological responses upon administration. The server evaluated the quantity of immune cells like helper T-cells 1 and 2 (Th1 and Th2), respectively. Additionally, the server measured various other immunological responses, including the production of antibodies, cytokines, and interferon, in response to administered vaccines.
Molecular dynamics simulation of the constructed vaccine and human TLR4 complex
Molecular dynamics (MD) simulations were performed using AMBER v2429 to assess the structural stability and conformational dynamics of the chimeric multi-epitope vaccine in complex with human Toll-Like Receptor 4 (TLR4; PDB ID: 2Z63). The initial complex was prepared in PyMOL by removing crystallographic water molecules and heteroatoms. The ff19SB force field was used for protein parametrization30. The system was solvated in an octahedral box of OPC water molecules with a 12.0 Å buffer and neutralized with Na+ and Cl− ions. NaCl was added to a final concentration of 0.15 M to mimic physiological conditions31. Energy minimization was carried out in two stages: 10,000 steps with positional restraints (10 kcal/mol·Å²) on protein heavy atoms, followed by 50,000 steps without restraints. The system was gradually heated from 0 K to 300 K over 500 ps under constant volume (NVT), using a Langevin thermostat (collision frequency of 2 ps− 1). This was followed by 1 ns of equilibration at 1 atm under constant pressure (NPT), using the Berendsen barostat, during which positional restraints were progressively removed. A 100 ns production run was performed at 300 K and 1 atm using periodic boundary conditions. Long-range electrostatics were handled with the particle mesh Ewald method. SHAKE constraints were applied to hydrogen-containing bonds, allowing a 2 fs timestep. Snapshots were saved every 10 ps for analysis. Analysis was performed using CPPTRAJ32. Root mean square deviation (RMSD) was calculated to assess the global stability of the complex over time. Root mean square fluctuation (RMSF) was calculated to examine residue-level flexibility. The radius of gyration (Rg) was calculated to monitor overall structural compactness. Intramolecular hydrogen bonds were analyzed to evaluate the complex stability. Principal component analysis (PCA) was conducted to identify dominant motions during the simulation. The binding free energy between the vaccine and TLR4 was estimated using the MM/GBSA method from extracted snapshots of the production phase. VMD and PyMOL were used for structural visualization.
Results
Vaccines provoke the immune system and play a crucial role in fighting against different diseases. Small antigenic determinants can efficiently set off immune reactions rather than entire virus proteins33. By determining possible immunogenic sites, immunoinformatics provides a fast and accurate method of vaccine design34. The fusion protein of RSV and hMPV and neuraminidase of IAV strains were analyzed using an immunoinformatics method in order to find immunogenic, non-allergenic CTL, HTL, and B-cell epitopes. Broad protection against RSV, hMPV, and IAV is sought by designing chimeric multi-epitope subunit vaccines from these epitopes35.
Identification of antigenic proteins and conserved regions in RSV, hMPV, and IAV for chimeric multiepitope vaccine
Among the seven proteins of RSV and hMPV and fourteen proteins of IAV analyzed for antigenicity, the fusion proteins of RSV and hMPV, along with the neuraminidase protein of IAV, showed the highest antigenicity scores (0.5219 for RSV, 0.4775 for hMPV, and 0.5024 for IAV). Thus, these proteins were selected for the construction of the chimeric multi-epitope subunit vaccine (Supplementary Table S4). The fusion and neuraminidase proteins of the selected viruses presented highly conserved regions across multiple isolates and strains (Supplementary Figs. S1–S3). These conserved regions served as the basis for identifying HTL, CTL, and BCL epitopes incorporated into the final vaccine construct. Targeting such conserved viral regions reduces the likelihood of immune escape and enhances vaccine efficacy against diverse viral strains.
Cytotoxic T cell epitope prediction and screening
CTL epitopes play a critical role in recognizing virus-infected cells and eliciting cellular immune responses. From the fusion and neuraminidase proteins of hMPV, RSV, and IAV, a total of 531, 566, and 461 CTL epitopes were predicted, respectively. The top ten epitopes with the highest prediction scores were shortlisted from each protein (Supplementary Tables S5–S7). Among these, two highly antigenic, non-allergenic, non-toxic, and conserved epitopes from each viral protein were selected for incorporation into the final chimeric multi-epitope vaccine construct (Table 1).
Helper T lymphocytes (HTL) epitopes prediction
HTLs play a vital role in the coordination of immune responses by the secretion of cytokines and the activation of other immune cells. Accordingly, a total of ten top-ranked HTL epitopes were selected from the fusion proteins of hMPV, RSV, and the neuraminidase protein of IAV (Supplementary Table S3). Among these, two epitopes from each virus exhibited high antigenicity, non-allergenicity, and non-toxicity and were selected for inclusion in the final chimeric multi-epitope vaccine construct (Table 2).
B-Cell epitopes prediction
B lymphocytes play a crucial role in adaptive immunity due to the production of antibodies that neutralize pathogens, thus offering long-term protection. A total of 21, 23, and 17 B-cell epitopes were identified from the fusion proteins of hMPV and RSV and the neuraminidase protein of IAV, respectively (Supplementary Table S4). Ten top-scoring epitopes from each protein were shortlisted, and two highly antigenic, non-allergenic, non-toxic, and conserved epitopes from each virus were selected for inclusion in the final chimeric multi-epitope vaccine construct (Table 3).
Chimeric multi-epitope vaccine construction
To assemble the final chimeric multi-epitope subunit vaccine (MESV), selected immunodominant epitopes were used. These epitopes were derived from the fusion proteins of hMPV and RSV, as well as the neuraminidase of IAV. They were joined using specific linkers to ensure structural flexibility and optimal immune presentation (Fig. 2A). Cytotoxic T lymphocyte (CTL) epitopes were linked via AAY linkers, helper T lymphocyte (HTL) epitopes via GPGPG linkers, and B-cell epitopes via KK linkers. The AAY linker facilitates efficient proteasomal processing and MHC class I presentation. GPGPG linkers enhance the presentation and immunogenicity of HTL and B-cell epitopes. KK linkers maintain structural separation, minimizing epitope interference and boosting immunogenicity. An immunostimulatory adjuvant, human β-defensin 2, was conjugated to the N-terminus of the construct using an EAAK linker to further enhance immune activation. Human β-defensin 2 is part of the body’s natural defense system and can activate specific immune responses against antigens. The finalized MESV construct comprised 442 amino acids in total. The specific epitopes incorporated into the vaccine design are detailed in Tables 1, 2 and 3.
3-D structure prediction and validation
Five 3D structural models of the designed chimeric vaccine were generated, and the first model demonstrated the most stable and well-defined conformation (Fig. 2A, C). This model was selected for subsequent structural and immunological analyses. Each epitope, including CTL, HTL, and B-cell epitopes, is represented in distinct colors and depicted as a ribbon. A surface shade has been applied for enhanced depiction (Fig. 2C). The selected model was validated by ProSA-web (Fig. 2D), ERRAT, and Ramachandran plot (Fig. 2B). The ProSA-web analysis revealed a score of − 8.3 for the 3D model of the proposed vaccine design. The overall quality factor reported by ERRAT was 96.31%, indicating high model accuracy (Supplementary Fig. S4). The Ramachandran plot study indicated that the majority of amino acids (98.4%) are located in the favored zone, whilst a minimal proportion (1.6%) is found in the prohibited region (Fig. 2B). The aforementioned results demonstrated that the quality of the chosen vaccine construct designs is appropriate and suitable for the next steps, including molecular docking, modelling, and free energy calculations.
3-D structure modeling and validation of the chimeric multi-epitope vaccine. (A) Chosen epitopes and linkers used in the chimeric subunit vaccine design, (B) Ramachandran plot analysis showing the distribution of backbone dihedral angles, indicating model stereochemical quality. (C) Predicted 3-D structure of the proposed chimeric multiepitope vaccine, (D) Z-score analysis from ProSA-web, representing the overall model quality compared to experimentally determined structures.
Physicochemical properties, allergenicity, and antigenicity of the vaccine construct
The antigenic potential of the designed vaccine construct was evaluated using the VaxiJen v2.0 server. It yielded a score of 0.50, which exceeds the threshold value of 0.4 and indicates a strong capacity to produce an immune response. Allergenicity assessment was conducted using the AllerTOP v2.0 server. The construct was predicted to be non-allergenic, suggesting its potential safety in immunization applications. We used the ProtParam tool to check properties like molecular weight, half-life, aliphatic index, pI, stability, and GRAVY score. The vaccine’s molecular weight is 46,715.05 Da. Its predicted half-life is 30 h in human cells, 20 h in yeast, and over 10 h in E. coli, showing good stability. The instability index is 28.79, which means the construct is stable. An aliphatic index of 74.28 suggested good thermostability, while a theoretical pI supported its acidic nature. The GRAVY score of -0.276 reflected the overall hydrophilic character of the vaccine, favoring solubility and biological interaction potential (Table 4).
Molecular docking with TLR4 receptor
An effective immunological response from a vaccine depends on its high binding affinity to the host immune receptors. Toll-like receptors (TLRs) play a crucial role in controlling inflammatory pathways. They help regulate immune responses to infections. TLRs recognize pathogen-associated molecular patterns (PAMPs). This recognition initiates intracellular signaling pathways and modulates gene expression. As a result, innate and adaptive immune responses are activated36. Toll-like receptors play a critical role in recognizing many viral components, including nucleic acids and envelope glycoproteins. This recognition initiates a cascade of events that leads to the synthesis of type I interferons (IFN-I), inflammatory cytokines, and chemokines. TLRs help activate and mature dendritic cells, linking the body’s early (innate) and later (adaptive) immune responses. We used the HDOCK server to test how the chimeric vaccine binds to the host’s immune receptors. The docking analysis assessed the binding affinity of the vaccine construct with human Toll-like receptor 4 (TLR-4; PDB ID: 2Z63). The resulting docking score was − 277.43 kcal/mol, suggesting a strong and stable interaction between the vaccine and TLR-4, which may support effective immune activation (Fig. 3). As per the PDBsum server, TLR-4 (Chain A) and the chimeric constructed vaccine (Chain B) have 9 hydrogen bonds: Thr413-Ser261, Gln484-Arg164, Glu485-Arg164 (two hydrogen bonds), Asn481-Arg311, Gln505-Arg311, Arg311-Gln507, Gln507-Arg327, and Gln507-Asp307. Additionally, four salt bridges; Glu439-Arg248, Glu485-Arg164, Glu509-Arg327, and Arg460-Asp307, and 183 non-bonded interactions were found in the constructed vaccine and TLR4 complex (Fig. 3).
Binding interaction between the docked vaccine and TLR-4 receptor.
Population coverage analysis
Population coverage analysis was performed using the IEDB Population Coverage tool. The objective was to identify the most representative epitopes across distinct immune recognition strategies. The analysis utilized curated data from the IEDB database. It accounted for regional variations in the global distribution of HLA and MHC alleles. These variations arise from complex environmental and genetic influences. Population coverage is a crucial factor in assessing the potential efficacy and broad applicability of a vaccine candidate. The IEDB analysis predicted a global coverage of 42% for MHC class I T-cell epitopes. For MHC class II T-cell epitopes, the predicted coverage was 81%. When both classes were considered together, the overall coverage increased to 91%. Figure 4 shows the regions with the highest combined epitope coverage worldwide.
The estimated population coverage analysis of individual and combined MHC-I and MHC-II epitopes.
Normal mode analysis of the constructed vaccine with TLR-4
The iMODS analysis revealed reduced structural distortions and high conformational stability of the vaccine–receptor complex, indicating a stable interaction and favorable structural dynamics between the vaccine construct and the immune receptor (Fig. 4). Molecular simulations estimated the flexibility and motion of atoms within the antigen–receptor interface. Figure 5A displays the deformability profile of the MEVC-TLR4 complex, showing a localized peak within a flexible region of the vaccine build. B-factor analysis, as shown in (Fig. 5B), compares the intrinsic mobility of residues derived from both the PDB and NMA models. The eigenvalue, depicted in (Fig. 5C), reflects the stiffness of the structure; a lower value indicates greater flexibility. (Fig. 5D) presents the variance associated with individual and cumulative motions, represented in purple and green, respectively. The covariance matrix (Fig. 5E) shows the dynamic correlations between residue pairs, where red denotes correlated motion, blue indicates anti-correlated motion, and white represents uncorrelated motion. Figure 5F represents the elastic network model, resembling a spring map, which visualizes the strength of interatomic interactions within the complex. Lastly, Fig. 5G summarizes the overall results of the iMODS analysis, confirming the structural integrity and potential functional stability of the vaccine TLR4 complex.
Normal mode analysis (NMA) of the chimeric multi-epitope vaccine construct in complex with TLR-4. (A) Deformability plot showing flexible regions within the complex. (B) B-factor plot comparing experimental and predicted residue fluctuations. (C) Eigenvalue representing the overall structural stiffness. (D) Variance plot showing individual and cumulative variances of the modes. (E) The covariance matrix of residue movements indicates correlated (red), uncorrelated (white), and anti-correlated (blue) motions. (F) Elastic network model illustrating inter-residue connections as springs. (G) Summary plot of the overall mobility and interaction dynamics derived from iMODS.
In silico cloning of chimeric multi-epitope subunit vaccine in the pET-28a(+) expression vector
The codon optimization of the 1,326-nucleotide chimeric multi-epitope subunit vaccine for E. coli K12 resulted in improved expression potential. The optimized sequence showed a high Codon Adaptation Index (CAI) and balanced GC content, indicating efficient transcription and translation compatibility within the selected host system. The optimized sequence yielded a CAI value of 1.0 and a GC content of 51.24%, both of which suggest high compatibility and potential for robust expression in the E. coli system. Subsequently, SnapGene software (v3.3.4) was used to simulate the cloning strategy. ScaI and BmtI restriction sites were added to the sequence ends to help insert it into the E. coli pET-28a(+) vector. The software confirmed the absence of internal restriction sites that could interfere with the cloning process. As illustrated in (Fig. 6), the optimized gene sequence was successfully integrated into the vector map, validating the construct’s readiness for downstream expression and purification.
In silico cloning of the codon-optimized chimeric vaccine into the pET-28a(+) vector. The diagram shows the successful insertion of the gene using Sca I and Bmt I sites, confirming it can be expressed in E. coli.
Immune response simulation analysis
Vaccines provoke the immune system of the host without causing disease. The immune system can be stimulated by vaccines through different mechanisms. Co-administration of protein-based antigens with immunologic adjuvants enhances local innate immune activation, leading to the functioning of antigen-presenting cells and the release of pro-inflammatory cytokines by macrophages. These macrophages, often referred to as dendritic cells, internalize the antigen and present it on their surface via major histocompatibility complex (MHC) molecules. T cells recognize the MHC-antigen complex through their T-cell receptors, thereby activating adaptive immunity and generating memory T cells. In contrast, polysaccharide antigens typically induce antibody responses without T-cell involvement. These antigens bind directly to mature B cells, initiating their differentiation into antibody-producing plasma cells. Immune simulation was conducted to assess the immunogenicity of the constructed vaccine. In the C-ImmSim simulation, three injections were administered at time steps 1, 84, and 168, corresponding biologically to days 0, 28, and 56, respectively, assuming each time step represents 8 h of real time. This simulation evaluated the immune system’s response, particularly antibody production, following the administration of the constructs. Notably, the immune system exhibited a robust response, with significantly elevated antibody production observed after each vaccine injection (Fig. 7). The chimeric multiepitope subunit vaccine construct was administered, and antigen titers were initially low until the seventh day. Subsequently, they progressively increased, reaching their maximum on the 12th day. Nevertheless, a rise in antibody titers was observed beginning on the 15th day. The activation of supplementary immune system components was accompanied by the complete neutralisation of antigens by the 17th day. The combined IgM and IgG titers were 4.5 × 107, and the combined IgG1 + IgG2 and IgG1 titers were also elevated post-vaccination (Fig. 7A). The evaluation of interleukin (IL) and cytokine responses, as shown in (Fig. 7B), indicates a notable increase in the levels of IFN-γ and IL-2. The results demonstrate a reliable and strong immune response subsequent to the administration of the vaccine. The cellular immune response upon re-exposure to pathogens was significantly robust, marked by the development of memory cells. T cell populations were found to be greater than 1500 cells/mm³, with peak concentrations of phagocytic natural killer cells, dendritic cells, and phagocytic macrophages reported at levels exceeding 380 cells/mm3 and 200 cells/mm3, respectively (Fig. 7B–F).
In silico immune simulation results for the constructed multi-epitope vaccine, generated using the C-ImmSim server. (A) Primary, secondary, and tertiary immune responses following the 3-D vaccine show elevated levels of IgM, IgG1, and IgG2 antibodies. (B) B cell population dynamics, including memory B cell generation and clonal expansion. (C) T-helper (CD4+) and cytotoxic T lymphocyte (CD8+) response profiles, including active and memory cell populations. (D) Cytokine expression levels (e.g., IFN-γ, IL-2) indicate activation of cell-mediated immunity. (E) Macrophage and dendritic cell activation over time, reflecting innate immune involvement. (F) Macrophage population in response to the vaccine. The simulation supports the immunogenic potential of the vaccine construct in eliciting strong, long-lasting humoral and cellular immune responses.
Molecular synamics simulation analysis of the TLR4-vaccine complex
To evaluate the structural stability and conformational behavior of the TLR4-vaccine complex, several parameters were monitored across the 100 ns production phase (Fig. 8A-D) and (Table S8). The RMSD profile (Fig. 8A) was used to assess global stability over time. The system showed a gradual rise in RMSD during the first 60 ns, reflecting initial structural adjustments. After ~ 65 ns, RMSD values stabilized around 5–6 Å, indicating that the complex reached equilibrium. Minor fluctuations observed after 70 ns suggest localized rearrangements without global instability. RMSF analysis (Fig. 8B) captured residue-wise flexibility across the complex. The TLR4 region (residues 1–569) exhibited limited fluctuations, mostly under 3 Å, indicating a stable backbone throughout the simulation. In contrast, the vaccine region (residues 570–1010) showed increased flexibility, with peaks exceeding 9 Å at certain loop regions, likely reflecting inherent flexibility in the designed construct. The Rg (Fig. 8C) was calculated to monitor the compactness of the TLR4-vaccine complex. The Rg remained steady around 39–40 Å during the initial 70 ns, suggesting stable folding. A gradual increase after 75 ns, reaching ~ 41.5 Å, indicates slight structural expansion, possibly due to flexible segments in the vaccine construct. No abrupt changes were observed, supporting overall structural retention. A contact map analysis (Fig. 8D) tracked native and non-native contacts, minimum/maximum distances, and contact stability across the trajectory. Native contacts remained consistently populated throughout, while non-native contacts fluctuated without showing dominance, suggesting preserved structural integrity. Maximum distance trends remained stable, with no significant expansion or dissociation events during the simulation. Collectively, RMSD stabilization after 65 ns, stable Rg, maintained native contacts, and localized flexibility observed in RMSF profiles indicate that the TLR4-vaccine complex remained structurally stable under simulated conditions, with expected mobility confined to vaccine regions.
Structural stability and conformational analysis of the TLR4-vaccine complex during 100 ns molecular dynamics simulation. (A) Root mean square deviation (RMSD) of backbone atoms, showing system equilibration after ~ 65 ns. (B) Root mean square fluctuation (RMSF) per residue, highlighting localized flexibility in the vaccine region (residues 570–1010). (C) Radius of gyration (Rg), indicating overall compactness with slight expansion after 75 ns. (D) Contact analysis, showing stable native contacts and consistent minimum distances, with no significant unfolding events detected throughout the simulation.
Conformational dynamics and principal component analysis
Principal component analysis (PCA) was conducted to investigate large-scale conformational changes in the TLR4-vaccine complex over the course of the simulation (Fig. 9A). The trajectory was projected along the first two principal components (PC1 and PC2), which captured the dominant motions within the system. The PCA plot revealed three distinct conformational states sampled during the simulation. The first state (A) was identified at ~ 28.3 ns, the second state (B) at ~ 49.2 ns, and the third state (C) at ~ 94.8 ns. The transitions between these states suggest gradual exploration of the conformational space rather than abrupt structural shifts. The presence of distinct clusters indicates that the complex underwent coordinated motions but remained within a restricted conformational landscape, supporting the structural stability observed in RMSD and Rg analyses. To visualize structural changes, representative snapshots corresponding to the early (blue) and late (red) stages of the simulation were superimposed (Fig. 9B). This comparison showed minimal deviation in the TLR4 region, consistent with its structural rigidity, while subtle shifts were observed in the vaccine region, reflecting expected flexibility. The PCA confirmed that the TLR4-vaccine complex explored a limited conformational space during the simulation, without significant structural rearrangement, supporting the stability of the complex under simulated conditions.
(A) Principal component analysis (PCA) projection of the TLR4-vaccine complex along PC1 and PC2 during the 100 ns MD simulation. Three distinct conformational states (A: 28.3 ns, B: 49.2 ns, C: 94.8 ns) are indicated. (B) Structural superposition of the complex at 0 ns (blue) and 100 ns (red), highlighting the overall structural preservation, with minor adjustments localized to the vaccine region.
Binding free energy analysis of the TLR4-vaccine complex
The binding free energy of the TLR4-vaccine complex was estimated using the Molecular Mechanics Generalized Born Surface Area (MM/GBSA) method based on snapshots from the production phase. The calculated energy components are summarized in Table 5. The total binding free energy (ΔGTOTAL) was estimated as − 121.72 kcal/mol, indicating a stable and favorable interaction between the vaccine construct and TLR4. The major contribution came from electrostatic interactions (ΔEEL: − 378.08 kcal/mol) and van der Waals forces (ΔEVDW: − 64.00 kcal/mol). The non-polar solvation term (ΔESASA: − 8.85 kcal/mol) contributed marginally to stabilization. In contrast, the polar solvation energy (ΔEGB: 329.22 kcal/mol) opposed binding, as expected due to desolvation penalties upon complex formation. Despite this, the gas-phase interaction energy (ΔGGAS: − 442.09 kcal/mol) effectively compensated, resulting in an overall favorable binding profile. The low standard error values across all components confirmed the stability and consistency of the calculated energy terms throughout the sampled frames. Overall, MM/GBSA analysis supports strong binding affinity between the multi-epitope vaccine construct and TLR4, driven primarily by electrostatic and van der Waals interactions.
Discussion
The growing burden of multidrug-resistant pathogens highlights the urgent need for novel preventive strategies, including effective vaccines37. In this context, the use of predicted epitopes enables the rational design of multi-epitope vaccines that address key limitations of conventional single-peptide vaccines, such as limited immune coverage and suboptimal immunogenicity38. Combining multiple B-cell and T-cell epitopes within a single vaccine construct is envisioned to promote a broader engagement of the adaptive immune system, encompassing both humoral and cellular responses39,40. In contrast to currently licensed monovalent or bivalent respiratory vaccines, which are based on whole antigens or inactivated pathogens and often drive immunity toward a limited set of dominant surface epitopes, multi-epitope strategies seek to focus immune recognition on conserved regions. Such a design may offer advantages in terms of cross-reactivity and reduced vulnerability to antigenic drift. The success of such an approach relies on careful epitope selection, sufficient HLA coverage, and appropriate immunostimulatory support to ensure effective immune activation in the absence of native protein structure41,42,43. The approach enables modular updating and recombinant scalability, while the absence of native antigen architecture requires optimized antigen presentation and adjuvant-mediated immune activation to achieve immunogenicity comparable to conventional vaccine platforms41.
In this study, a structure-based immunoinformatics framework was used to design a multi-epitope vaccine targeting Human Metapneumovirus, Respiratory Syncytial Virus, and Influenza A Virus. The final construct incorporated cytotoxic T-lymphocyte (CTL), helper T-lymphocyte (HTL), and B-cell epitopes with high predicted antigenicity, strong MHC binding affinity, and broad population coverage, suggesting the potential to induce coordinated cellular and humoral immunity44. The inclusion of an N-terminal adjuvant significantly enhanced the predicted immune profile, as evidenced by elevated IFN-γ, IgG, and memory T-cell responses in immune simulations38,39,40. Flexible linkers further improved epitope accessibility and construct stability, as supported by secondary and tertiary structural analyses41. The vaccine also exhibited favorable physicochemical properties, including predicted stability, non-allergenicity, and solubility, indicating feasibility for recombinant production and downstream experimental validation.
Accurate structural modeling is critical for predicting immune recognition and receptor binding in multi-epitope vaccines8. However, the chimeric construct lacks a close structural homolog, classical homology-based modeling is inherently unreliable. When sequence identity falls below the 30% “twilight zone,” alignment errors increase sharply, and both core folds and loop regions may be mis-predicted45. Recent benchmarks confirm that under these conditions, homology models are unsuitable for atomic-level interpretation, necessitating the use of threading or hybrid de novo approaches46. In this study, the refined tertiary structure showed acceptable stereochemical quality, supporting the likelihood that CTL, HTL, and B-cell epitopes remain accessible for immune recognition.
To estimate innate immune activation potential, the interaction between the vaccine construct and human Toll-like receptor 4 (TLR4) was evaluated. Docking analysis indicated that the vaccine engages TLR4 in a manner consistent with known ligand–receptor recognition mechanisms44, which is biologically relevant because TLR4 activation drives dendritic cell maturation, cytokine secretion, and T-cell priming. Because docking provides only a static view, molecular dynamics (MD) simulations were used to evaluate the stability and behavior of the vaccine–TLR4 complex under near-physiological conditions.
The MD trajectory demonstrated that the complex stabilized after ~ 65 ns, with no major structural rearrangements thereafter8. The consistent radius of gyration indicated maintenance of compactness, while RMSF analysis showed rigidity of the TLR4 chain and expected flexibility within vaccine linker and loop regions. Contact analysis confirmed preservation of native interactions, and principal component analysis identified only three dominant conformational states, indicating restricted and coordinated motion within a stable conformational landscape34. Together, these results support the structural robustness of the vaccine TLR4 interaction.
Binding free energy calculations using MM/GBSA further supported these findings. Although MM/GBSA is known to overestimate absolute affinities due to simplified solvation and entropy treatment, its use across multiple MD snapshots improves the reliability of relative binding trends47. The total binding energy was favorable, driven primarily by electrostatic and van der Waals interactions, despite counteracting polar solvation effects, consistent with known mechanisms of TLR4–peptide recognition46,47. These results indicate that the vaccine forms a stable and energetically favorable complex with TLR4, supporting its potential to function as an immunostimulatory agent.
Population-coverage analysis using IEDB revealed high global and regional HLA representation driven by prevalent alleles such as HLA-A02:01, HLA-A24:02, and HLA-B07:02 (MHC-I), together with HLA-DRB104:01, DRB107:01, and DRB1*15:01 (MHC-II), which dominate antigen presentation across Europe, East Asia, South Asia, and the Americas48. However, reduced representation of certain HLA-B and HLA-DR alleles common in West and East African and indigenous populations suggests a potential coverage gap, highlighting the need for additional epitopes to improve global equity49.
The immunogenic potential of the construct was further evaluated using C-ImmSim immune simulations. The predicted induction of class-switched antibodies and Th1-associated cytokines, including IFN-γ and IL-2, indicates activation of immune pathways associated with viral clearance and memory formation50,51. These findings suggest that the construct promotes a balanced humoral and cellular response, which is essential for durable protection against viral infections. However, molecular-level simulations do not capture tissue-specific immunity. MD modeling evaluates epitope accessibility and receptor binding but cannot distinguish systemic IgG responses from mucosal IgA-dominated immunity, which is critical for respiratory pathogens8. Likewise, C-ImmSim provides qualitative and semi-quantitative insights into immune dynamics but cannot fully represent host-pathogen interactions, immune evasion, or organ-specific responses52,53. Therefore, in vivo and ex vivo validation remains essential.
From a translational perspective, favorable codon adaptation, in-silico cloning, and predicted solubility suggest feasibility for recombinant expression, particularly in E. coli. Nevertheless, computational solubility correlates only moderately with experimental outcomes and is influenced by host-specific folding, stress responses, and post-translational effects54,55. Alternative expression systems may therefore be required. Despite these limitations, similar computational pipelines have successfully produced viable vaccine candidates, validating this approach as a powerful early-stage screening strategy56,57.
Overall, the integration of epitope selection, structural modeling, molecular docking, molecular dynamics simulation, immune simulation, and population-coverage analysis provides a comprehensive in-silico validation of this multi-epitope vaccine candidate. By targeting conserved regions of the fusion proteins of hMPV and RSV and conserved neuraminidase regions of IAV, the construct is designed to elicit both neutralizing antibodies and cross-reactive T-cell responses, potentially enabling broader and more durable protection than single-pathogen designs53,54,55,56,57. These results provide a strong rationale for advancing this candidate to experimental validation.
Conclusion
Emerging respiratory viral co-infections caused by hMPV, RSV, and IAV remain a significant public health concern, especially in vulnerable populations. In this study, a multi-epitope vaccine was designed using a structure-based immunoinformatics approach to target these pathogens. The designed construct incorporated carefully selected CTL, HTL, and B-cell epitopes, along with an appropriate adjuvant, aiming to stimulate both innate and adaptive immunity. Structural modeling and molecular docking confirmed favorable binding to human TLR4, while molecular dynamics simulations demonstrated the stability of the vaccine-receptor complex under physiological conditions. Binding free energy analysis supported a strong interaction, primarily driven by electrostatic and van der Waals forces. In silico immune simulation predicted robust immune activation, and codon optimization confirmed the construct’s suitability for expression in Escherichia coli. These results support the designed multi-epitope vaccine as a promising candidate for experimental validation and further development.
Limitations and future directions
Designing a multi-epitope vaccine targeting multiple pathogens entails several significant challenges. One of the primary concerns is achieving an optimal balance of immunogenicity among diverse antigenic components to prevent potential immune interference. Additionally, ensuring broad human leukocyte antigen (HLA) coverage while minimizing the risk of autoimmune cross-reactivity is critical for achieving population-wide efficacy and safety. Furthermore, the development of a stable vaccine formulation that maintains consistent epitope expression and minimizes antigen degradation or loss during manufacturing and storage remains a considerable technical hurdle. While our study provides an in-silico assessment of the designed multi-epitope vaccine, experimental validation remains essential to confirm its immunogenicity and safety. The structural stability, binding affinity, and immune stimulation potential demonstrated computationally should now be verified through in vitro assays, such as protein expression, receptor binding, and cytokine profiling. Further in vivo studies will be required to evaluate immunogenicity, protective efficacy, and safety under physiological conditions. Experimental validation will be essential to confirm the immunogenicity and protective efficacy of the designed vaccine. Nonetheless, this computational framework provides an efficient strategy for accelerating early-stage vaccine discovery and candidate selection.
Data availability
All data is publicly available on the FDA website, the original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
References
Babawale, P. I. & Guerrero-Plata, A. Respiratory viral coinfections: Insights into epidemiology, immune response, pathology, and clinical outcomes. Pathogens 13, 316 (2024).
Salas-Benito, J. S. & De Nova-Ocampo, M. Viral interference and persistence in mosquito-borne flaviviruses. J. Immunol. Res. 2015, 873404 (2015).
Du, Y., Wang, C. & Zhang, Y. Viral coinfections. Viruses 14, 2645 (2022).
Chin, T., Foxman, E. F., Watkins, T. A. & Lipsitch, M. Considerations for viral co-infection studies in human populations. MBio 15, e00658–e00624 (2024).
Chu, C. J. & Lee, S. D. Hepatitis B virus/hepatitis C virus coinfection: Epidemiology, clinical features, viral interactions and treatment. J. Gastroenterol. Hepatol. 23, 512–520 (2008).
Piret, J. & Boivin, G. Viral interference between respiratory viruses. Emerg. Infect. Dis. 28, 273 (2022).
Liu, B. et al. Developing a new sepsis screening tool based on lymphocyte count, international normalized ratio and procalcitonin (LIP score). Sci. Rep. 12, 20002 (2022).
Zeng, H. et al. A signal enhanced lateral flow immunoassay based on core–shell quantum dots labeled antibody and antigen for sensitive detection of CP4-EPSPS protein. Microchem. J. 209, 112758 (2025).
Falsey, A. R. Neurologic complications of influenza and potential protective vaccine effects. Influenza Other Respir Virus. 19, e70071. (2025). https://doi.org/10.1111/irv.70071
Tregoning, J. S. & Schwarze, J. Respiratory viral infections in infants: Causes, clinical symptoms, virology, and immunology. Clin. Microbiol. Rev. 23, 74–98 (2010).
Wang, L. et al. Cigarette smoke extract-treated mouse airway epithelial cells-derived exosomal LncRNA MEG3 promotes M1 macrophage polarization and pyroptosis in chronic obstructive pulmonary disease by upregulating TREM-1 via m6A methylation. Immune Netw. 24, e3 (2024).
Langley, J. M. et al. Incidence of respiratory syncytial virus lower respiratory tract infections during the first 2 years of life: A prospective study across diverse global settings. J. Infect. Dis. 226, 374–385. https://doi.org/10.1093/infdis/jiac227 (2022).
Esper, F., Boucher, D., Weibel, C., Martinello, R. A. & Kahn, J. S. Human metapneumovirus infection in the United States: Clinical manifestations associated with a newly emerging respiratory infection in children. Pediatrics 111, 1407–1410 (2003).
Arnott, A. et al. A study of the genetic variability of human respiratory syncytial virus (HRSV) in Cambodia reveals the existence of a new HRSV group B genotype. J. Clin. Microbiol. 49, 3504–3513 (2011).
Bohmwald, K. et al. Human respiratory syncytial virus: infection and pathology. In Seminars in Respiratory and Critical Care Medicine, 522–537 (Thieme Medical, 2016).
Dawre, S. & Maru, S. Human respiratory viral infections: Current status and future prospects of nanotechnology-based approaches for prophylaxis and treatment. Life Sci. 278, 119561 (2021).
Martin, E. T., Kuypers, J., Wald, A. & Englund, J. A. Multiple versus single virus respiratory infections: Viral load and clinical disease severity in hospitalized children. Influenza Other Respir Virus. 6, 71–77 (2012).
Groen, K. Human metapneumovirus: Evolution and interaction with the innate immune system (2023).
Yang, X., Chen, M., Cao, L. & Zhao, M. Bibliometric analysis of scientific papers on adverse reactions to COVID-19 vaccines published between 2019 and 2023. Hum. Vaccines Immunother. 19, 2270194 (2023).
Shchelkunova, G. A. & Shchelkunov, S. N. Smallpox, monkeypox and other human orthopoxvirus infections. Viruses 15, 103 (2022).
Zhang, J. et al. Predicting in vitro and in vivo anti-SARS-CoV-2 activities of antivirals by intracellular bioavailability and biochemical activity. ACS Omega. 7, 45023–45035 (2022).
Zhang, H. Q. et al. An analysis of reported cases of hemophagocytic lymphohistiocytosis (HLH) after COVID-19 vaccination. Hum. Vaccines Immunother. 19, 2263229 (2023).
Abbadi, N. & Mousa, J. J. Broadly protective neuraminidase-based influenza vaccines and monoclonal antibodies: Target epitopes and mechanisms of action. Viruses 15, 200 (2023).
Cortés, G. et al. Boosting neuraminidase immunity in the presence of hemagglutinin with the next generation of influenza vaccines. npj Vaccines. 9, 228 (2024).
Pang, Y. et al. Efficacy, immunogenicity and safety of respiratory syncytial virus prefusion F vaccine: Systematic review and meta-analysis. BMC Public Health 24, 1244 (2024).
Murhekar, M. V. et al. SARS-CoV-2 seroprevalence among the general population and healthcare workers in India, December 2020–January 2021. Int. J. Infect. Dis. 108, 145–155 (2021).
Saha, S. & Raghava, G. P. S. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Prot. Struct. Funct. Bioinform. 65, 40–48 (2006).
Kim, J., Yang, Y. L., Jang, S. H. & Jang, Y. S. Human β-defensin 2 plays a regulatory role in innate antiviral immunity and is capable of potentiating the induction of antigen-specific immunity. Virol. J. 15, 1–12 (2018).
Case, D. et al. AMBER University of California, San Francisco, Ta [Google Scholar]. (2018). (2018).
Tian, C. et al. Wu, ff19SB: amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J. Chem. Theory Comput. 16, 528–552 (2019).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Roe, D. R. & Cheatham, T. E. III PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Ma, S. et al. Development of a novel multi-epitope mRNA vaccine candidate to combat HMPV virus. Hum. Vaccines Immunother. 20, 2293300 (2024).
Alnajran, H. et al. Design of a peptide-based vaccine against human respiratory syncytial virus using a reverse vaccinology approach: evaluation of immunogenicity, antigenicity, allergenicity, and toxicity. Front. Immunol. 16, 1546254 (2025).
Maleki, A., Russo, G., Parasiliti Palumbo, G. A. & Pappalardo, F. silico design of recombinant multi-epitope vaccine against influenza A virus. BMC Bioinform. 22, 617 (2021).
Kawai, T. & Akira, S. Pathogen recognition with Toll-like receptors. Curr. Opin. Immunol. 17, 338–344 (2005).
Cabrera, A., Mason, E., Mullins, L. P. & Sadarangani, M. Antimicrobial resistance and vaccines in Enterobacteriaceae including extraintestinal pathogenic Escherichia coli and Klebsiella pneumoniae. npj Antimicrob. Resist. 3, 1–14 (2025).
Shahraki, P. K., Kiani, R., Siavash, M. & Bemani, P. Design of a multi-epitope vaccine against Staphylococcus aureus lukotoxin ED using in silico approaches. Sci. Rep. 15, 14517 (2025).
Akurut, E. et al. Design of a multi-epitope vaccine against drug-resistant mycobacterium tuberculosis and mycobacterium bovis using reverse vaccinology. Sci. Rep. 15, 27298 (2025).
Alshabrmi, F. M. et al. An in-silico investigation to design a multi-epitopes vaccine against multi-drug resistant Hafnia alvei. Vaccines 10, 1127 (2022).
Mortazavi, B., Molaei, A. & Fard, N. A. Multi-epitope vaccines, from design to expression; an in silico approach. Hum. Immunol. 85, 110804 (2024).
Biswas, R. et al. Designing multi-epitope vaccine against human cytomegalovirus integrating pan-genome and reverse vaccinology pipelines. Biologicals 87, 101782 (2024).
Yang, Z., Bogdan, P. & Nazarian, S. An in silico deep learning approach to multi-epitope vaccine design: A SARS-CoV-2 case study. Sci. Rep. 11, 3238 (2021).
Guo, W. et al. silico design of a multi-epitope vaccine against Mycobacterium avium subspecies paratuberculosis. Front. Immunol. 16, 1505313 (2025).
Biswas, R. & Anbarasu, A. Identification of novel zinc-binding inhibitors against key microbial metallohydrolase DapE in Klebsiella pneumoniae: An integrated ligand-based virtual screening, molecular docking, molecular dynamics, and MM/PBSA approach. Integr. Biology. 17, zyaf018 (2025).
Yuan, R. et al. Casp16 protein monomer structure prediction assessment. Struct. Funct. Bioinform. 94, 86–105 (2025).
Genheden, S. & Ryde, U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin. Drug Discov. 10, 449–461 (2015).
Reynisson, B. et al. Improved prediction of MHC II antigen presentation through integration and motif deconvolution of mass spectrometry MHC eluted ligand data. J. Proteome Res. 19, 2304–2315 (2020).
Cao, Y. et al. Imprinted SARS-CoV-2 humoral immunity induces convergent Omicron RBD evolution. Nature 614, 521–529 (2023).
Al-Dhabi, N. A., Valan, M. & Arasu Environmentally-friendly green approach for the production of zinc oxide nanoparticles and their anti-fungal, ovicidal, and larvicidal properties. Nanomaterials 8, 500 (2018).
Roy, A. et al. Integrating pan-genome and reverse vaccinology to design multi-epitope vaccine against Herpes simplex virus type-1. 3 Biotech. 14, 176 (2024).
Sgroi, G. Mycobacterium tuberculosis-immune system dynamics through agent-based modeling methodology: An important step in accelerating vaccine discovery (2022).
Rapin, N., Lund, O., Bernaschi, M. & Castiglione, F. Computational immunology meets bioinformatics: the use of prediction tools for molecular binding in the simulation of the immune system. PLoS One. 5, e9862 (2010).
Prabakaran, R. et al. Effect of charged mutation on aggregation of a pentapeptide: Insights from molecular dynamics simulations. Proteins Struct. Funct. Bioinform. 90, 405–417 (2022).
Hebditch, M., Carballo-Amador, M. A., Charonis, S., Curtis, R. & Warwicker, J. Protein–Sol: A web tool for predicting protein solubility from sequence. Bioinformatics 33, 3098–3100 (2017).
Wang, S. et al. Viral expression of NE/PPE enhances anti-colorectal cancer efficacy of oncolytic adenovirus by promoting TAM M1 polarization to reverse insufficient effector memory/effector CD8 + T cell infiltration. J. Exp. Clin. Cancer Res. 44, 97 (2025).
Oyarzún, P. & Kobe, B. Recombinant and epitope-based vaccines on the road to the market and implications for vaccine design and production. Hum. Vaccines Immunother. 12, 763–767 (2016).
Funding
The authors acknowledge financial support from the National Natural Science Foundation of China (No. 32300048), Guangdong Basic and Applied Basic Research Foundation (No. 2022A1515110158, 2024A1515012577), National Foreign Expert Individual Program (No. Y20240195), the Construction Project of Nano Technology and Application Engineering Research Center of Guangdong Medical University (No. 4SG24179G), the Postdoctoral funding project of Guangdong Medical University (3007/2BH24007), Dongguan Science and Technology of Social Development Program (20231800936272).
Author information
Authors and Affiliations
Contributions
L.L.: Supervision, Funding acquisition, Writing—review and editing. Y.C. and S.W.: Project administration, Writing—review and editing. C.W. and J.X.: Investigation, Methodology. A.S.: Software. X.X.: Data curation. J.T.: Investigation. Y.Q. and Y.Z.: Methodology, Software. A.J.: Software, Visualization, Writing—original draft. T.Y.: Writing—review and editing. S.U.: Data curation, Formal analysis, Writing—original draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, L., Chen, Y., Wu, S. et al. Immunoinformatics based designing of a broad-spectrum multi-epitope vaccine against co-infection of human metapneumovirus, respiratory syncytial virus, and influenza A virus. Sci Rep 16, 10244 (2026). https://doi.org/10.1038/s41598-026-40812-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-40812-z