Abstract
Cisplatin resistance in tongue squamous cell carcinoma (TSCC) correlates with poor prognosis, where natural killer (NK) cells in the tumor microenvironment (TME) play a crucial role. This study investigated the mechanism by which exosomes from cisplatin-resistant TSCC cells suppress NK cell function. We found that exosomal long non-coding RNA SNHG26, highly enriched in cisplatin-resistant TSCC cells and their exosomes, was transferred to NK cells. Within NK cells, SNHG26 acted as a scaffold promoting WWP2-mediated ubiquitination and degradation of the transcription factor SOX2, thereby inhibiting HLA-DRA transcription and subsequent IL-2/JAK-STAT5 signaling. Concurrently, SNHG26 competitively bound miR-515-5p, relieving its suppression of TGFB1 mRNA and activating the TGF-β1/Smad2 pathway. These dual mechanisms significantly impaired NK cell proliferation, activation, and cytotoxicity. SNHG26 depletion reversed NK cell suppression and cisplatin resistance in vitro and in vivo. Thus, our study identifies exosomal SNHG26 as a key mediator of cisplatin resistance and NK cell dysfunction in TSCC, suggesting its potential as a promising therapeutic target.
Similar content being viewed by others
Introduction
Tongue squamous cell carcinomas (TSCC) are malignant neoplasms derived from epithelial tissues, primarily affecting the anterior two-thirds of the tongue. This classification holds significant relevance within the realm of head and neck cancers1. Based on statistical data, the global incidence of TSCC amounts to ~180,000 new cases annually2,3. In contrast to lung cancer and breast cancer, TSCC exhibits a relatively low incidence rate and has not garnered significant attention from both societal and oncological spheres. However, it is important to acknowledge that the initial manifestations of TSCC are inconspicuous and susceptible to being disregarded. This carcinoma exhibits a propensity for local infiltration and dissemination of cancer cells to adjacent lymph nodes, attributable to the tongue’s frequent movement and its abundant local vascular and lymphatic networks. Consequently, by the time the disease is detected, a majority of cases have already progressed to intermediate or advanced stages4.
The advancement of medical research has led to the evolution of treatment for TSCC, transitioning from solely surgical intervention to a comprehensive sequential approach encompassing preoperative induction chemotherapy, surgery, and postoperative radiotherapy and chemotherapy5. Despite these advancements, the prognosis for patients remains suboptimal, with a 5-year overall survival rate of ~50%6. Chemotherapy resistance in the clinical setting is a primary determinant of unfavorable prognosis among patients7. Cisplatin (CDDP), being the foremost and fundamental drug employed in the treatment of TSCC, exerts a potent and widespread anticancer effect by directly impairing the DNA of malignant cells5. Nevertheless, the occurrence of cisplatin resistance in TSCC is frequent during clinical administration, resulting in a mere 27.7% rate of pathological effectiveness8. The underlying reasons and mechanisms for this phenomenon remain elusive. Therefore, investigating the mechanisms associated with cisplatin resistance in TSCC and developing effective interventions holds significant clinical importance.
Previous research has primarily concentrated on investigating the genetic and molecular alterations within tumor cells; however, the clinical application of these findings has encountered obstacles. This is attributed to the intricate interplay between tumor cells and their surrounding “soil” known as the tumor microenvironment (TME), resulting in the formation of a cohesive entity9. Solid tumor tissues harbor tumor cells within a dense, hypoxic, and acidic microenvironment, which is characterized by immunosuppression. This microenvironment fosters uncontrolled tumor growth, resistance to apoptosis, angiogenesis, invasion, and metastasis, ultimately exacerbating patient prognosis10. In the context of TSCC and other head and neck cancer tissues, the proportion of immunosuppressive cells within the TME tends to increase as malignancy progresses, whereas the proportion of immune-promoting cells tends to decrease (Fig. S1)11. Natural killer (NK) cells, which constitute the fundamental component of the innate immune system, possess the capability to directly induce apoptosis in tumor cells and eliminate them, irrespective of the presence of complement and antibodies. With regard to cytotoxicity, NK cells occupy the second rank, surpassed only by CD8 + T cells. Additionally, NK cells play a crucial role in immunomodulation by releasing cytokines that activate Dendritic cells (DC), tumor-associated macrophages (TAM), and CD8 + T cells, thereby regulating the adaptive immune system (Fig. S2)12. The utilization of NK cell-based chimeric antigen receptor therapy (CAR-NK) has demonstrated remarkable effectiveness and safety in combating cancer, thus emerging as a promising avenue for advancing immunotherapy13.
Exosomes, which typically range in diameter from 20 to 200 nm, are secreted by diverse cell types within the human body. These diminutive vesicles serve as crucial mediators of intercellular communication, harboring a diverse array of functional biomolecules such as DNA, RNA, and proteins within the originating cells14. Research has demonstrated that tumor cells possess the capability to generate exosomes in substantial amounts, which they employ as a means of conveying diverse effector molecules to adjacent or remote cells, thereby establishing a microenvironment conducive to their proliferation and metastasis. Furthermore, these exosomes can also transmit signal factors that impede immune response, thereby diminishing the immunity of individuals afflicted with tumors, particularly those in advanced stages15. The involvement of exosomes derived from tumor cells is crucial in facilitating tumor immune evasion and tolerance.
Long non-coding RNA (lncRNA) has been extensively associated with the initiation, progression, and resistance to chemotherapy in tumors16. Consequently, it holds significant promise for utilization in cancer diagnosis, prognosis assessment, and targeted therapeutic interventions17. Exosomes transport a considerable quantity of lncRNAs derived from tumor cells18, which is anticipated to serve as a pivotal factor in investigating the interplay between tumor cells and the immune microenvironment of TSCC. The small nucleolar RNA host genes (SNHGs) family are a class of lncRNAs that originate from small nucleolar RNA of eukaryotic cells and participate in the regulation of cell proliferation and differentiation19. In our previous study, it was established that SNHG26 exhibits a notably elevated expression in TSCC cells that are resistant to cisplatin. Moreover, it was found that SNHG26 facilitates the proliferation and epithelial-mesenchymal transition (EMT) of TSCC cells by specifically targeting PGK1 ubiquitination and activating the Akt/mTOR signaling pathway. Consequently, this mechanism ultimately leads to an augmentation in cisplatin resistance20. There are numerous potential mechanisms through which SNHG26 targets cells, warranting further study (Fig. S3).
The objective of this study is to examine the function and mechanism of exosomes released by cisplatin-resistant TSCC (TSCC/CDDP) cells in relation to the immunosuppressive effects on NK cells. This study employed clinical samples, patient-derived organoids (PDO), TSCC/NK cell co-culture system, patient-derived tumor xenograft (PDX) models, and humanized mice as research subjects, and utilized drug sensitivity testing, single cell and transcriptome sequencing analysis, gene regulation, DNA/RNA/protein interaction experiments, and other methodologies to corroborate the notion that the TSCC/CDDP cells can impede the anticancer efficacy of microenvironment NK cells via the exosome SNHG26, at clinical, in vivo, in vitro, and molecular levels.
Results
The tissues of TSCC that exhibited resistance to cisplatin demonstrated a notable reduction in both the content and activity of NK cells, in contrast to the sensitive tissues
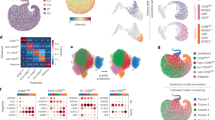
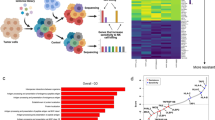
To accurately differentiate between cisplatin-sensitive and cisplatin-resistant tissues of TSCC, we developed PDOs (Fig. 1A) and PDX models (Fig. 1B) using freshly collected TSCC tissues obtained from clinical settings. In the present study, the categorization of tissue as drug resistant is contingent upon the inhibition of cisplatin on tumor activity being below 30%. Following cisplatin sensitivity assessment using PDO and PDX models, scRNA-seq of the corresponding tumor tissues revealed notable reduced NK cell abundance (defined by CD56+/NKG2D+populations) in cisplatin-resistant TSCC specimens compared to cisplatin-sensitive controls (Fig. 1C). Subsequently, flow cytometry was employed to validate the abundance of NK cells in both categories of tissues, and the findings substantiated a noteworthy decrease in NK cells abundance within cisplatin-resistant TSCC tissues (n = 10) compared to cisplatin-sensitive tissues (n = 10) (Fig. 1D). Furthermore, through the utilization of qRT-PCR and WB techniques, we discovered that the expression levels of the activating molecules CD69, NKG2D, and NKp46 were comparatively diminished, while the inhibitory molecules CD94, NKG2A, and CD158a were relatively elevated on the surface of NK cells within the resistant tissues, in contrast to the cisplatin-sensitive tissues (Fig. 1E).
A Differentiating PDOs to distinguish tissues that are resistant or sensitive to cisplatin (Scale: 25 μm). B Differentiating between cisplatin resistant and sensitive tissues using the PDX models. C Single-cell profiling reveals altered NK cell distribution and abundance in cisplatin-resistant TSCC (Left: UMAP split by CDDP-res and CDDP-sen and projection of single cells colored by cell type; Right: Proportional changes of major immune cell types between groups). D Flow cytometry analysis of NK cells content in TSCC tissues (n = 10). E The identification of surface markers of NK cells in TSCC through the utilization of qRT-PCR and WB techniques. The CDDP-sen referred to the cisplatin-sensitive tissues of TSCC, while the CDDP-res denoted the cisplatin resistant tissue of TSCC. NC negative control. Compare each group with the NC group when there is no special identification on the statistical chart; *p < 0.05, **p < 0.01, ***p < 0.001. The figures were generated using R language (version 4.2.0), GraphPad Prism (version 9.0) and Figdraw.
Exosomes derived from cisplatin-resistant TSCC cells can significantly inhibit the activity and toxicity of NK cells
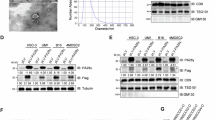
In order to investigate the role of exosomes in the inhibitory effect of cisplatin-resistant TSCC cells on NK cells, we established a co-culture system involving the aforementioned cell types. Our results indicated that cisplatin-resistant cells (CAL27/CDDP) exerted a pronounced inhibitory effect on the proliferation and cytotoxicity of human NK-92 cells, as compared to cisplatin-sensitive cells (CAL27). Furthermore, the addition of an exosome inhibitor (GW4869) to cisplatin-resistant cells effectively reversed the inhibitory effect on NK-92 cells, implicating the involvement of exosomes in this mechanism (Fig. 2A). The supernatants derived from TSCC cells were subjected to ultracentrifugation. Subsequent analysis using transmission electron microscopy revealed the presence of vesicle-like structures ranging in diameter from 30 to 313 nm. Notably, phospholipid bilayers were observed at the periphery of these vesicles. Furthermore, WB experiments demonstrated the expression of surface-specific markers (CD9 and TSG101) in the extracted samples, while the absence of cell content-specific markers (Calnexin) was observed (Fig. 2B). Following that, exosomes were prelabeled with Dil red fluorescent dye and subsequently incubated with NK cells. The findings obtained from fluorescence microscopy demonstrated the presence of red fluorescence in NK cells following a 12-hour incubation period, with a notable increase in red fluorescence observed after 24 h of incubation (Fig. 2C). These results indicated the potential uptake of exosomes originating from TSCC cells by NK cells.
A o-culture experiments were conducted to assess the impact of cisplatin-resistant and cisplatin-sensitive TSCC cells on the proliferation and toxicity of NK-92 cells. B Transmission electron microscopy image, particle size analysis and WB detection of characteristic proteins of exosomes derived from TSCC cells (Scale: 100 nm). C The process of extracellular vesicle uptake by NK cells was observed using fluorescence microscopy (Scale: 100 μm). D The activity of NK cells was assessed by CCK-8 detection. E The LDH assay was employed to assess the cytotoxicity of NK cells. F The quantification of Perforin and Granzyme B expression levels in NK cells through the utilization of a qRT-PCR assay. G The concentration of IFN-γ in NK cells was quantified using the ELISA technique. H The detection of Perforin and Granzyme B expression in NK cells was performed using WB analysis. exo exosome, NC negative control. Compare each group with the NC group when there is no special identification on the statistical chart; *p < 0.05, **p < 0.01, ***p < 0.001. The figures were generated using GraphPad Prism (version 9.0) and Figdraw.
Furthermore, co-culturing of exosomes obtained from cisplatin-sensitive (CAL27, HSC3) and cisplatin-resistant (CAL27/CDDP and HSC3/CDDP) TSCC cells with NK-92 cells was conducted. The activity of NK-92 cells was assessed using the CCK-8 assay, while the toxicity of NK-92 cells was evaluated using the LDH assay. Compared to exosomes from cisplatin-sensitive cells, exosomes derived from cisplatin-resistant TSCC cells significantly impeded the proliferation and cytotoxicity of NK cells (Fig. 2D, E). Moreover, we employed qRT-PCR, WB, and ELISA techniques to ascertain the presence of crucial cytotoxic factors, namely Perforin, Granzyme B, and IFN-γ, released by NK cells. The findings of this study demonstrated that exosomes originating from TSCC cells resistant to cisplatin effectively suppressed the generation of NK cytotoxic factors (Fig. 2F–H).
The inhibitory effect of exosomes derived from cisplatin-resistant TSCC cells on NK cells is mediated by SNHG26
To identify exosomal lncRNAs driving NK cell suppression, we conducted RNA-seq on exosomes from cisplatin-resistant versus sensitive TSCC tissues. Differential analysis (|log₂FC|>2, FDR <0.05) revealed multiple upregulated lncRNAs, with SNHG26 ranking among the top three most enriched transcripts (Fig. S4). This was cross-validated by our prior cell-level sequencing data20, which consistently showed SNHG26 upregulation in resistant cells. Given its persistent overexpression in both cellular and exosomal compartments, coupled with its established role in promoting cisplatin resistance, we selected SNHG26 for further functional investigation.
To further explore the role of SHNG26 in this study, the exosomes derived from TSCC cells that are sensitive and resistant to cisplatin were subjected to co-cultivation with NK cells. The utilization of FISH and qRT-PCR techniques demonstrated a noteworthy enhancement in the expression of SNHG26 in recipient NK cells upon the administration of exosomes originating from cisplatin-resistant TSCC cells (Fig. 3A). To ascertain the regulatory capacity of SNHG26 on NK cells in TSCC tissues, the researcher employed the TCGA-HNSC database for bioinformatics analysis. The findings demonstrated a significant modulation of immune-related functions in TSCC patients by SNHG26, exhibiting a negative correlation with the extent of infiltration and enrichment of NK cells within the TME (Fig. 3B). Furthermore, we conducted an investigation into the expression of SNHG26 and the NK cells marker CD56 in TSCC tissue microarray (HN054Oc01) using FISH in conjunction with mIF. The findings revealed a significant negative correlation between the expression of SNHG26 and CD56 (Fig. 3C). In addition, siRNA was employed to downregulate the expression of SNHG26. Subsequently, the activity of NK cells was assessed using the CCK-8 assay, while the toxicity of NK cells was measured through the LDH assay. Moreover, the expression levels of relevant cytokines were determined using qRT-PCR and WB techniques. The findings revealed that the inhibition of SNHG26 in NK cells effectively counteracted the notable suppressive impact exerted by exosomes derived from cisplatin-resistant TSCC cells on NK cell activity and toxicity, as depicted in Fig. 3D–G.
A he impact of exosomes on the expression of SNHG26 in NK cells was assessed through the utilization of FISH and qRT-PCR in TSCC (Scale: 2.5 μm). B The TCGA-HNSC database was employed to examine the impact of SNHG26 on NK cells within the microenvironment of TSCC. C The detection of SNHG26 and CD56 expression in TSCC tissue microarray was performed using a combination of FISH and mIF techniques (n = 45). D The activity of NK cells was assessed by CCK-8 detection. E The LDH assay was employed to assess the cytotoxicity of NK cells. F The quantification of Perforin and Granzyme B expression levels in NK cells through the utilization of a qRT-PCR assay. G The detection of Perforin and Granzyme B expression in NK cells was performed using WB analysis. exo exosome, NC negative control, si- siRNA. Compare each group with the NC group when there is no special identification on the statistical chart; *p < 0.05, **p < 0.01, ***p < 0.001. The figures were generated using R language (version 4.2.0) and GraphPad Prism (version 9.0).
The activity and cytotoxicity of NK cells were suppressed by SNHG26 through its targeting of HLA-DRA/ (IL-2/JAK-) STAT5 and TGF-β1/Smad2 pathways within NK cells
In this study, a plasmid was employed to induce the overexpression of SNHG26 in NK cells. Normal NK cells were used as a control group for the purpose of identifying differentially expressed genes through NGS (Fig. 4A). Additionally, sequencing data of NK cells from the GEO database, specifically comparing high-activity group versus low-activity group, were extracted and analyzed to identify the differentially expressed genes between these two groups (Fig. 4B). Furthermore, the sequencing data of TSCC tissues obtained from the TCGA database were extracted and organized. Subsequently, the core genomic modules associated with NK cells activity were analyzed using WCGNA (Fig. 4C). By integrating the aforementioned three datasets, we identified the potential target genes of SNHG26 in NK cells, namely HLA-DRA and TGFB1 (Fig. 4D). We subsequently conducted qRT-PCR and WB analyses to confirm the inhibitory effect of SNHG26 on HLA-DRA expression and the promotive effect on TGFB1 expression at both the RNA and protein levels. Moreover, SNHG26 overexpression led to increased phosphorylation of Smad2, indicating enhanced activation of the TGF-β1/Smad2 signaling axis. Furthermore, our study revealed that the upregulation of HLA-DRA resulted in enhanced phosphorylation of STAT5. Additionally, we observed that HLA-DRA played a crucial role in mediating the inhibitory impact of SNHG26 on STAT5 phosphorylation(Fig. 4E–G and Figs. S5–S7). Through the utilization of CCK-8 and LDH assays, it was determined that the upregulation of HLA-DRA and the downregulation of TGFB1 effectively counteracted the suppressive impact exerted by SNHG26 on the proliferation and cytotoxicity of NK cells(Fig. 4H, I).
A Identification of differentially expressed genes in NK cells exhibiting overexpression of SNHG26 compared to normal NK cells. B Genes exhibiting differential expression in highly active NK cells compared to their low active counterparts (GSE12198). C Core gene module associated with the activation of NK cells in tissues of TSCC (TCGA-HNSC). D Identifying candidate target genes through intersection analysis. E The impact of SNHG26 on the expression levels of HLA-DRA and TGFB1 as determined through qRT-PCR. F The impact of SNHG26 detection through WB on the protein levels of HLA-DRA and TGF-β1/Smad2. G The impact of HLA-DRA and SNHG26 on the (IL-2/JAK-) STAT5 as determined through Western blot detection. H, I The CCK-8 and LDH assays were employed to ascertain whether the modulation of HLA-DRA and TGFB1 in NK cells could potentially counteract the impact of SNHG26 on NK cell proliferation and cytotoxicity. NC negative control, si- siRNA, p- overexpression plasmid, ph- phosphorylated. Compare each group with the NC group when there is no special identification on the statistical chart; *p < 0.05, **p < 0.01, ***p < 0.001. The figures were generated using R language (version 4.2.0) and GraphPad Prism (version 9.0).
SNHG26 could inhibit HLA-DRA transcription by promoting ubiquitination of SOX2
By utilizing the database available at catRAPID (http://service.tartaglialab.com/page/catrapid_group), we conducted an analysis to predict the potential binding protein of SNHG26. Additionally, we explored the potential transcription factors of HLA-DRA through the resource provided at JASPAR (https://jaspar.genereg.net/). Upon intersecting the results from both analyses, we discovered that SOX2 could potentially serve as a crucial molecule in mediating the regulatory effects of SNHG26 on HLA-DRA (Fig. 5A). Following qRT-PCR and WB experiments, it was determined that SNHG26 does not exert an influence on the RNA expression of SOX2. However, it does inhibit the protein level expression of SOX2. Conversely, SOX2 was found to enhance the RNA and protein expression of HLA-DRA. Furthermore, the overexpression of SOX2 was observed to counteract the inhibitory impact of SNHG26 on HLA-DRA (Fig. 5B, C). These experiments provide compelling evidence that SOX2 plays a central role in mediating the regulatory effects of SNHG26 on HLA-DRA.
A Bioinformatics analysis was employed to investigate the potential interaction between the target protein SOX2 and the genes SNHG26 and HLA-DRA. B, C The regulation of HLA-DRA by SNHG26 mediated by SOX2 was verified through qRT-PCR and WB experiments. D The ChIP assay was employed to authenticate the interaction between SOX2 and the promoter region of HLA-DRA. E The binding of SOX2 to the HLA-DRA promoter region was confirmed through the utilization of a dual-luciferase reporter assay. F The upregulation of SNHG26 facilitated the CHX-induced suppression of SOX2 production. G The inhibition of SOX2 degradation induced by MG132 can be enhanced through the knockdown of SNHG26. H Based on the bioinformatics analysis, it was indicated that there exists a potential for interaction among SNHG26, SOX2, and WWP2. I The binding of SNHG26, SOX2, and WWP2 in NK cytoplasm was confirmed by employing FISH in conjunction with mIF experiments (Pearson’s R value >0.8). J RNA pull-down and RIP experiments, conducted subsequent to nucleocytoplasmic separation, demonstrated the ability of SNHG26 to interact with SOX2 and WWP2 within the cytoplasm of NK cells. K The verification of the interaction between SOX2 and WWP2 in NK cytoplasm was conducted through co-IP, and the confirmation of the augmented impact of SNHG26 on this interaction was also established. L The co-IP experiment provided evidence that SNHG26 and WWP2 have the ability to enhance the ubiquitination level of SOX2. NC negative control, si- siRNA, p- overexpression plasmid. Compare each group with the NC group when there is no special identification on the statistical chart; *p < 0.05, **p < 0.01, ***p < 0.001. The figures were generated using R language (version 4.2.0) and GraphPad Prism (version 9.0).
In order to examine the involvement of SOX2 in the transcriptional regulation of HLA-DRA, we initially validated the binding ability of SOX2 to the promoter region of HLA-DRA (ACAATGG) through ChiP assay (Fig. 5D). The promoter region of HLA-DRA was cloned into the pGL3 enhancer vector, followed by Luciferase assays. Our findings indicate a significant increase in promoter activity of HLA-DRA upon overexpression of SOX2, while its activity decreased upon knockdown of SOX2. However, there was no increase in the activity of the mutated HLA-DRA promoter upon overexpression of SOX2 (Fig. 5E). These results strongly suggest that SOX2 has the ability to bind to the promoter region of HLA-DRA and initiate its transcription.
To investigate the mechanism by which SNHG26 reduces SOX2 protein levels, we treated NK-92 cells overexpressing SNHG26 with the protein synthesis inhibitor cycloheximide (CHX, 100 μg/mL). WB analysis revealed that SNHG26 overexpression accelerated the rate of SOX2 protein degradation. Furthermore, this degradation was effectively blocked by cotreatment with the proteasome inhibitor MG132 (10 μM), suggesting that SNHG26 promotes SOX2 degradation primarily via the proteasomal pathway (Fig. 5F, G). The molecular docking platform (https://www.dockeasy.cn/) was employed to predict WWP2 as a potential E3 ubiquitin ligase of SOX2. Additionally, the analysis results from the RPISeq database (http://pridb.gdcb.iastate.edu/R-PISeq/) demonstrate the simultaneous binding of SNHG26 to both SOX2 and WWP2 (Fig. 5H). The confirmation of the co-localization of SNHG26, SOX2, and WWP2 in the cytoplasm of NK-92 cells was achieved through the utilization of FISH and mIF techniques (Fig. 5I). Consequently, we postulated that SNHG26 facilitates the ubiquitination of SOX2 prior to its nuclear translocation through the involvement of WWP2. Initially, we substantiated the binding capabilities of SHNG26 to SOX2 and WWP2 through the utilization of RIP and RNA pull-down assays, respectively (Fig. 5J). The subsequent co-IP assay findings revealed that the overexpression of SNHG26 significantly augmented the binding of SOX2 and WWP2 (Fig. 5K). Following the co-IP assay involving ubiquitin molecules, it was observed that the overexpression of SNHG26 facilitated the WWP2-mediated ubiquitination of SOX2 (Fig. 5L). The data presented in this study indicate that SNHG26 may function as a structural framework for the recruitment of WWP2 to SOX2, consequently promoting the WWP2-mediated process of ubiquitination degradation of SOX2.
SNHG26 could promote TGFB1 expression by competitive binding to miR-515-5p
The upregulation of TGFB1 expression in the cytoplasm by SNHG26 aligns with the prevailing focus on competing endogenous RNA (ceRNA) investigations. To identify potential miRNA interactions, we employed miRDB (https://mirdb.org/) for SNHG26 and ENCORI (https://starbase.sysu.edu.cn/) for TGFB1. By intersecting the results, we identified miRNA-515-5p as a promising candidate molecule (Fig. 6A). The findings from the qRT-PCR and WB analyses revealed that SNHG26 exerted an inhibitory effect on the expression of miR-515-5p. Furthermore, it was observed that miR-515-5p effectively suppressed the expression of TGFB1 at both the RNA and protein levels (Fig. 6B). Subsequently, a dual-luciferase reporter assay was conducted to authenticate the reciprocal interaction involving SNHG26/miR-515-5p/TGFB1. The luciferase reporter plasmid was utilized to incorporate the sequences of SNHG26-Wt, SNHG26-Mut, TGFB1-Wt, or TGFB1-Mut. The findings revealed that solely the miR-515-5p mimic successfully suppressed the luciferase activity in the Wt group in comparison to the remaining groups. These outcomes suggest a direct interaction between SNHG26 and miR-515-5p, as well as between miR-515-5p and TGFB1 (Fig. 6C). The impact of cotransfection on TGFB1 expression was examined at both the RNA and protein levels. Our findings demonstrated that the introduction of the miR-515-5p inhibitor successfully counteracted the downregulation of TGFB1 RNA and protein expression induced by si-SNHG26. Conversely, cotransfection with the miR-515-5p mimic effectively reversed the upregulation of TGFB1 RNA and protein expression facilitated by the p-SNHG26 (Fig. 6D). The findings of this study indicate that SNHG26 exerts a promotive effect on TGFB1 expression at both the RNA and protein levels through its competitive binding to miR-515-5p.
A By means of bioinformatics analysis, miRNAs exhibiting competitive binding associations with SNHG26 and TGFB1 were identified. B The impact of SNHG26 on the expression of miR-515-5p, as well as the influence of miR-515-5p on the expression of TGFB1, were assessed through qRT-PCR and WB techniques. C The luciferase reporter assay was conducted on NK-92 cells that were co-transfected with miR-515-5p mimics and luciferase reporters containing either the wild-type or mutant 3’ UTRs of SNHG26 or TGFB1. D The expression levels of TGFB1 RNA and protein were analyzed using qRT-PCR and WB techniques subsequent to cotransfection with si-SNHG26+inhibitor miR-515-5p or p-SNHG26+mimic miR-515-5p. E The molecular mechanism of SNHG26 in TSCC was depicted through a schematic illustration (The black dashed line represents the previously released content, while the purple dashed line represents the direct mechanism of action). NC negative control, si- siRNA, p- overexpression plasmid, wt wild, mut mutant. Compare each group with the NC group when there is no special identification on the statistical chart; *p < 0.05, **p < 0.01, ***p < 0.001. The figures were generated using GraphPad Prism (version 9.0) and Figdraw.
SNHG26 promoted TSCC growth by inhibiting NK cells in vivo
In order to investigate the impact of SNHG26 on NK cells within the TSCC microenvironment in vivo, a mouse model of PBMC was employed. While these mice possess immune system characteristics of human origin, the overall count of immune cells is relatively low, rendering them suitable for tumor xenograft experiments akin to other prevalent immunodeficient mouse models. Xenografts were produced through the subcutaneous injection of CAL27 cells that were stably infected with Lv-SHNG26 or Lv-Vector, followed by intraperitoneal administration of cisplatin (Fig. 7A–C). The tumor volume and mass of the OE-SNHG26 group exhibited a statistically significant increase compared to the Vector group (Fig. 7D). This finding implied that the overexpression of SNHG26 may facilitate the proliferation of TSCC, aligning with our prior observations in BALB/C mice20. Through the utilization of FISH and IF assays, our study revealed that the overexpression of SNHG26 in TSCC tissues leads to a significant reduction in the content of NK (CD56+) cells (Fig. 7E). Furthermore, our findings indicate that SNHG26 overexpression hampers the expression of HLA-DRA while simultaneously enhancing the expression of TGFB1, both at the RNA and protein levels (Fig. 7F). The findings of this study provided evidence that SNHG26 exerts a promotive effect on TSCC growth through the inhibition of NK cells in an in vivo.
A The construction process of PBMC humanized mouse model for TSCC (huPBMC-NSG). B Appearance of the huPBMC-NSG and identification of TSCC by HE staining (Scale: 1 mm). C Flow cytometry was used to identify human immune cell components (hCD45 + mCD45-) and determine the proportion of Marker cells within the human cell population. D The quantification of tumor tissue volume and weight in the Vector and OE-SNHG26 cohorts of huPBMC-NSG. E The FISH&mIF assays were employed to compare the expression levels of SNHG26 and CD56 in the Vector and OE-SNHG26 groups (Scale: 100 μm). F The expression levels of SNHG26, HLA-DRA, and TGFB1 were compared between the NC and OE groups using qRT-PCR or IF experiments (Scale: 20 μm). OE overexpression. Compare each group with the NC group when there is no special identification on the statistical chart; *p < 0.05, **p < 0.01, ***p < 0.001. The figures were generated using GraphPad Prism (version 9.0) and Figdraw.
Mendelian randomization (MR) was employed to examine the correlation between HLA-DR and TGF-β1 with oral cancer
MR is a data analysis methodology employed in epidemiological studies to assess causal inference. This technique utilizes genetic variants that exhibit robust associations with exposure factors, serving as instrumental variables to evaluate the causal association between said exposure factors and outcomes. Based on the objective of this study, we opted to select exposure and outcome factors that closely resemble the desired characteristics from the GWAS database (https://gwas.mrcieu.ac.uk/). Specifically, the exposure factor examined was the occurrence of oral cancer (ieu-b-4961), while the outcome factors investigated were the expression of HLA-DR in NK cells (ebi-a-GCST90001885) and the expression of TGF-β1 in the microenvironment (prot-c-2333-72-1). The results showed that oral cancer cells could decrease the expression of HLA-DR in NK cells, but p = 0.054 > 0.05, that is, there is no significant causal relationship between the two. This finding is consistent with the results of our cell experiments, which showed that TSCC cells did not significantly inhibit the expression of HLA-DRA in NK cells. Similarly, oral cancer cells can cause a decrease in the expression of TGF-β1 in the microenvironment, but p = 0.549, much greater than 0.05, and there is no causal relationship between the two (Table 1 and Fig. 8A–D). Due to the disparities observed between the GWAS data and the variables examined in this study, it is not possible to establish definitive conclusions. However, two indications emerge from the findings: firstly, normal TSCC cells do not appear to induce substantial alterations in HLA-DR and TGF-β1 within the microenvironment. Secondly, the increased expression of SNHG26 in drug-resistant TSCC cells and its subsequent regulation of HLA-DR and TGF-β1 exist “antecedents” to be explored.
A, B Forest plot of the causal effects of SNP associated with oral cancer on HLA-DR or TGFB1 expression. The significance of red lines are MR results of the MR-Egger test and the IVW method. C, D Scatter plots of genetic associations with oral cancer against the genetic associations with HLA-DR or TGFB1. The slopes of each line represent the causal association for each method. The figures were generated using the R language (version 4.2.0).
Discussion
Cisplatin resistance and an immunosuppressive microenvironment are recognized as major factors contributing to treatment failure in TSCC. Therefore, a deeper understanding of their underlying mechanisms is crucial for improving patient prognosis.
Our integrated multi-omics and functional analyses provide compelling evidence for selecting NK cells as the primary focus in studying cisplatin resistance mechanisms in TSCC. Initial clinical single-cell RNA sequencing of patient tissues revealed NK cells as the most significantly impaired immune population in cisplatin-resistant TSCC, showing both reduced infiltration abundance and suppressed activation status characterized by decreased expression of activating receptors alongside increased inhibitory markers. NK cells, which are derived from the bone marrow, are crucial immune cells that form an integral part of the innate immune system. Despite being the third largest category of lymphocytes, after T cells and B cells, NK cells have not received as much research focus. However, recent investigations have highlighted the significance of NK cells as key constituents of the tumor immune microenvironment, demonstrating their ability to exert anticancer effects (Fig. S1). NK cells exert their direct cytotoxic effects on tumor cells primarily through three mechanisms: (1) the exocytic release of granules containing perforin 1 and various members of the granzyme protease family; (2) the secretion of interferon gamma (IFNG) and tumor necrosis factor (TNF); (3) the engagement of death receptors such as Fas cell surface death receptor (FAS) and TNF receptor superfamily member 10b (TNFRSF10B) on the surface of target cells are key mechanisms involved in this process21. In recent times, CAR-NK therapy has emerged as a substitute for CAR-T therapy in the management of solid tumors22. This alternative exhibits several advantages: (1) it obviates the need for autologous NK cells, enabling the production of readily available CAR-NK cells on a substantial scale, thereby facilitating their administration to patients at any given moment23; (2) CAR-NK therapy demonstrates a low incidence of cytokine release syndrome and neurotoxicity24; (3) in addition to the CAR pathway, NK cells possess diverse mechanisms for effectively targeting and eradicating cancer cells25. In conclusion, NK cells play an important role and have great research potential in the TME.
The initial observation that cisplatin-resistant TSCC cells suppressed NK cell function in transwell systems suggested non-contact mediated communication, consistent with known tumor-stromal interaction paradigms26. This led us to hypothesize exosomal involvement, as tumor-derived exosomes are well-documented mediators of immunosuppressive signaling in the tumor microenvironment27. We specifically tested this using GW4869, an established inhibitor of exosome biogenesis, which significantly reversed NK cell suppression—demonstrating exosome-dependency. The sufficiency of purified resistant cell exosomes to suppress NK function independently confirms their mechanistic role. This systematic verification provided rationale for focusing on exosomal cargo, ultimately suggesting that SNHG26 functions as a key mediator of NK cell dysfunction in cisplatin-resistant TSCC, expanding our understanding of lncRNA-mediated resistance mechanisms.
HLA-DR, a major histocompatibility complex class II (MHC-II) molecule, consists of two subunits, namely the α-subunit and β-subunit. The α-subunit is encoded by the HLA-DRA gene. Previous research has shown that the expression of HLA-DRA on the membrane of NK cells serves as a crucial indicator of their proliferation and activation. This expression enables the presentation of processed antigen fragments to CD4+ and CD8 + T cells during the initial phase of the immune response28. The activated CD4+ and CD8 + T cells, in turn, stimulate the proliferation and activation of NK cells through the secretion of IL-229. In the present study, it was observed that augmented HLA-DRA expression facilitated the proliferation and activation of NK cells, while also inducing a notable increase in the phosphorylation of STAT5 within the IL-2/JAK-STAT signaling axis. These findings imply that HLA-DRA not only promotes NK cells proliferation and activation through its antigen-presenting function, but also by directly enhancing the IL-2-mediated NK activation pathway (IL-2/JAK-STAT5).
In the context of the TME, TGF-β1 plays a pivotal role in immunosuppression, regulating the production and functionality of diverse immune cell types. Specifically, it governs adaptive immunity by directly facilitating the proliferation of Treg cells while concurrently impeding the generation and efficacy of effector T cells and antigen-presenting dendritic cells. Likewise, TGF-β1 exercises control over the innate immune system by suppressing NK cells and modulating the intricate behavior of macrophages and neutrophils, thereby establishing a network of negative immune regulation30. Previous research has demonstrated that the presence of tumor cells can impede the anticancer capabilities of NK cells through the secretion of TGF-β131. It is noteworthy to mention that NK cells possess the ability to internally generate and release TGF-β1, thereby exerting regulatory control over their own functioning as well as that of adjacent immune cells32. The TGF-β receptor/Smad pathway plays a crucial role in the transduction of TGF-β33. Our study demonstrated that the upregulation of TGFB1 in NK cells mediates the inhibition of their proliferation and activation induced by SNHG26. Additionally, the phosphorylation level of Smad2 within the TGF-β receptor/Smad signaling axis is enhanced. These findings imply that SNHG26 exerts its inhibitory effects on NK cell proliferation and activation through the activation of the TGF-β1/Smad2 pathway.
Our integration of multi-level sequencing data—from primary tumor exosomes to drug-resistant cell lines—provides evidence for SNHG26 as a master regulator of cisplatin resistance in TSCC. The consistent overexpression of SNHG26 across cellular and exosomal contexts suggests an active packaging mechanism that may facilitate intercellular communication within the TME. SNHG26, an lncRNA, is synthesized in the nucleus and has the ability to translocate to the cytoplasm, thereby being present in both cellular compartments. Previous research indicates the existence of five potential regulatory mechanisms of SNHG26 in NK34. In the present study, it was observed that exosomes originating from cisplatin-resistant TSCC cells predominantly augmented the cytoplasmic expression of SNHG26 in NK cells, thereby directing our attention towards investigating the cytoplasmic mechanism of action. In the current investigation, the correlation between SNHG26 and TGFB1 expression aligns with the regulatory pattern of ceRNA. However, the observed regulation pattern does not conform to the expected trend for HLA-DRA. Previous research has demonstrated that lncRNA can serve as a “structural scaffold” within the cytoplasm and facilitate post-translational modifications of proteins35. Therefore, we have opted to explore this aspect as a potential breakthrough in our study.
The upregulation of SOX2, functioning as a transcription factor, has been observed to facilitate the advancement of cancer and confer resistance to therapeutic agents36. Nevertheless, the regulatory function of SOX2 in NK cells and the correlation between SOX2 and HLA-DR have yet to be documented in existing literature. WWP2, a significant E3 ubiquitin ligase, has been empirically linked to the onset, progression, and chemoresistance of diverse malignancies due to its aberrantly elevated expression37. A study provided confirmation that WWP2 functions as a SOX2-specific E3 ubiquitin ligase in glioblastoma stem cells38. This study elucidated the transcriptional-enhancing impact of SOX2 on HLA-DRA and unveiled the potential of lncRNA as a “scaffold” to facilitate the interaction between WWP2 and SOX2. MiR-515-5p has been widely studied and identified as a classic tumor suppressor. It has been shown that circANKS1B can promote metastasis and cisplatin resistance of oral squamous cell carcinoma by promoting TGFB1 expression through sponge miR-515-5p39. In this study, we explored the ceRNA regulatory axis of SNHG26/miR-515-5p/TGFB1.
Although direct experimental validation of the crosstalk between these two axes is beyond the scope of this study, our integrated bioinformatic analysis of TCGA-OSCC and STRING databases suggests potential functional coordination between them, which represents an exciting avenue for future investigation. Detailed interaction networks are mapped in Fig. S8.
Despite the compelling evidence presented herein, our study has several limitations that warrant acknowledgment. First, while we demonstrate that exosomal SNHG26 is sufficient to suppress NK cell function and that its knockdown reverses this effect, the clinical translation of targeting SNHG26 requires further validation in larger, independent patient cohorts. Second, our mechanistic insights are primarily derived from the NK-92 cell line; although this is a standard model, future studies should confirm these pathways in primary human NK cells cocultured with patient-derived TSCC exosomes to enhance physiological relevance. Third, our in vivo findings are based on a humanized mouse model with a reconstituted immune system. This system may not fully recapitulate the complex interplay between SNHG26-laden exosomes and all immune cell populations (e.g., T cells and macrophages) within a native human tumor microenvironment. Lastly, although we identified two parallel signaling pathways (HLA-DRA/STAT5 and TGFB1/Smad2), the potential crosstalk between these axes within NK cells remains an open question.
These limitations, however, clearly delineate valuable directions for future research. To comprehensively map the expression and transfer of SNHG26 across all immune cell subsets within the TME, future studies should employ single-cell RNA sequencing of TSCC samples, with particular focus on T cells and macrophages. Additionally, generating NK cell-specific Sox2 knockout mice would be a powerful tool to definitively validate the in vivo role of this axis. More broadly, these limitations highlight the need to explore the therapeutic efficacy of SNHG26 inhibition in immunocompetent models and to validate our findings in primary patient-derived systems.
Methods
Cell lines
The human TSCC cell lines CAL27 and HSC3 were obtained from the American Type Culture Collection (ATCC). Subsequently, the corresponding cisplatin-resistant cell lines CAL27/CDDP and HSC3/CDDP were established using the drug concentration gradient method40. The aforementioned cell lines were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM), supplemented with 10% fetal bovine serum (FBS). NK-92 cells were also purchased from ATCC, and they were cultured in RPMI-1640 medium supplemented with 20% fetal bovine serum (FBS), 100 international units per milliliter (IU/mL) of recombinant human interleukin-2 (IL-2), 100 IU/mL of IL-15, and 50 IU/mL of IL-21. All cell lines mentioned above were cultured at 37 °C in a 5% CO2 incubator and authenticated through short tandem repeat (STR) analysis.
Clinical tissues
TSCC tissues were procured from patients diagnosed with TSCC who had undergone surgical procedures at the First Affiliated Hospital of Nanchang University in Nanchang, China, spanning the period from September 2020 to June 2022. The diagnosis of TSCC was established through histopathological examination, and no preoperative treatment was administered to the patients. Written informed consent was obtained from all participants, and the study was approved by the Human Ethics Committee of the First Affiliated Hospital of Nanchang University under the reference number 2023CDYFYYLK05-011.
The establishment of patient-derived organoids (PDO) for TSCC
The TSCC tissues underwent two washes with phosphate buffer solution (PBS) supplemented with compound antibiotics. Following this, the enzymatic complex solution was added in the appropriate quantity, and the tissues were finely fragmented using an electric homogenizer. The resulting homogenized tissue fragments were then transferred to a centrifuge tube and subjected to enzymatic digestion at 200 rpm for 1 h on a shaking table set at 37 °C. Subsequently, the enzymatically hydrolyzed cells were filtered through a 70-μm cell screen to obtain suspensions consisting of either single cells or oligocytotic cells. The cell suspension was subjected to two rounds of centrifugation at 200 × g, resulting in the separation of the tumor cell layers. The isolated individual cells were then introduced into a 3D culture matrix and placed in a specialized cell incubator with a low temperature setting. Following a 30-min incubation at 37 °C, the formation of a solid cell layer prompted the addition of the complete PDO culture base, consisting of advanced DMEM/F12 supplemented with various additives, as outlined by ref. 41. The growth of the organoids was subsequently monitored under a microscope at regular intervals.
To assess cisplatin sensitivity in tumor organoids, we established a standardized 96-well plate-based drug screening platform. Organoids were exposed to clinically relevant cisplatin concentrations (1 μg/ml, corresponding to plasma Cmax levels observed in TSCC patients receiving standard regimens) for 24 h. Following drug removal through PBS washing cycles (3 × 5 min), organoids underwent 24 h recovery culture in fresh complete medium supplemented with 10% Matrigel. Cellular viability was quantified using the CellTiter-Glo® 3D Assay System (Promega) per manufacturer’s protocol: Briefly, 100 μl reagent was added per well, followed by 5 min orbital shaking (500 rpm) and 30 min dark incubation. Luminescent signals were acquired using a GloMax® Discover Microplate Luminometer (Promega) with integration time optimized at 1 s/well.
The treatment-to-control (T/C) ratio was calculated using the Eq. 1:
where RLU represents relative luminescence units. Cisplatin resistance was defined as a T/C ratio >30%, consistent with NCCN-guided in vitro resistance thresholds for platinum agents.
The construction of patient-derived xenograft (PDX) models for TSCC
The construction of the TSCC PDX model was carried out in accordance with our prior investigation42. The drug sensitivity of the tumor tissue was assessed upon its transplantation into the P3 generation PDX model. Drug response was quantified using the tumor growth inhibition (TGI) metric, which was calculated as Eq. 2:
In alignment with RECIST 1.1 guidelines for antitumor efficacy evaluation, a threshold of TGI >30% was defined as clinically significant drug sensitivity.
The experimental protocol underwent review and supervision by the Animal Welfare Ethics Committee of the First Affiliated Hospital of Nanchang University (CDYFY-IACACC-202304QR015). For all surgical procedures, mice were anesthetized using inhalation of isoflurane (3–4% for induction and 1.5–2% for maintenance in 100% oxygen) delivered via a precision vaporizer. Depth of anesthesia was confirmed by the absence of a pedal reflex. Body temperature was maintained at 37 °C using a heating pad throughout the procedure. For endpoint euthanasia, mice were first deeply anesthetized with isoflurane. Euthanasia was then performed by cardiac puncture followed by cervical dislocation as a secondary method to ensure death. All procedures were carried out in strict accordance with the AVMA Guidelines for the Euthanasia of Animals (2020) to minimize suffering.
Peripheral blood mononuclear cells (PBMC) humanized mouse model for TSCC
The second layer of peripheral blood mononuclear cells (PBMCs) with a milky white appearance was isolated using Ficoll density gradient centrifugation at a speed of 1750 rpm and a temperature of 20 °C for a duration of 20 min. Subsequently, a volume of red blood cell lysate three to five times greater than the PBMC layer was added and allowed to lyse at a temperature of 4 °C for a duration of 5 min. The resulting cells were then resuspended in a volume of buffer three to five times greater than the initial PBMC layer. Platelets were subsequently removed through low-speed centrifugation at 750 rpm for a duration of 10 min at a temperature of 4 °C. Freshly purified PBMCs were resuspended in PBS at a concentration of 5 × 107 cells/mL and maintained on ice. Subsequently, the PBMCs were injected into hIL-15 NSG mice via the tail vein at a dosage of 1 × 107 cells per mouse, with the aim of establishing a humanized mouse model43. On the second day following the implantation of PBMC into NSG mice, CAL27 cells in logarithmic growth phase were enzymatically digested using 0.05% trypsin. Subsequently, the cells were treated with sterile PBS and Matrigel matrix gel at a concentration of 1:1. The resulting cell suspension was then resuspended to a concentration of 2 × 106 cells per 100 μl and injected subcutaneously into the dorsal region of the mice, with a volume of 500 μl per mouse.
The experimental protocol underwent review and supervision by the Animal Welfare Ethics Committee of the First Affiliated Hospital of Nanchang University (CDYFY-IACACC-202304QR015). During tumor cell inoculation and for terminal procedures, mice were anesthetized by intraperitoneal injection of a ketamine (80 mg/kg) and xylazine (10 mg/kg) mixture. Anesthesia depth was monitored by the loss of pedal reflex. Euthanasia was performed when tumor volume exceeded 1500 mm³ or if mice showed any signs of distress (e.g., lethargy, >20% weight loss). Mice were first deeply anesthetized with the ketamine/xylazine mixture. Euthanasia was then conducted by CO₂ asphyxiation in a pre-filled chamber at a flow rate of 30–50% of the chamber volume per minute, followed by cervical dislocation for confirmation. These methods are consistent with the AVMA Guidelines and were approved by the institutional animal ethics committee.
Extraction and identification of exosomes from TSCC cells
The supernatants from the culture of TSCC cells were collected, and the isolation of exosomes was performed using differential centrifugation, following the previously described method44. Purified exosomes were promptly mixed with a solution containing 4% glutaraldehyde and 1% osmium tetroxide. Subsequently, the resulting suspension was applied onto a Formvar/carbon-coated electron microscopy grid and left undisturbed for a duration of 5 min. Following this, the cells were subjected to staining using a 10% uranyl acetate solution for a period of 5 min, after which they were examined using transmission electron microscopy (TEM). Nanoparticle tracking analysis (NTA) was conducted in order to ascertain the distribution, dimensions, and quantity of exosomes. Western blotting (WB) was employed to identify the protein constituents within the extracts, utilizing CD9, TSG101, and Calnexin as the selected indicators.
Extraction of exosomes from TSCC tissues
Fresh tumor tissues were enzymatically digested using collagenase IV (2 mg/mL) and hyaluronidase (100 U/mL), followed by sequential differential centrifugation at 300×g (10 min), 2000×g (20 min), and 110,000×g (70 min) to isolate exosomes. Nanoparticle tracking analysis (NTA, Malvern Panalytical) confirmed vesicle size distribution (30–150 nm), while transmission electron microscopy (Hitachi HT7800) verified morphological integrity. Exosomal RNA was extracted using TRIzol LS reagent, with RNA integrity assessed by Agilent 2100 Bioanalyzer (RIN >7.0). Libraries were prepared with NEBNext Small RNA Library Prep Kit and sequenced on Illumina NovaSeq platform (2 × 150 bp paired-end), yielding ≥10 million clean reads per sample. Raw data underwent quality control via FastQC and adapter trimming using Trimmomatic. Differentially expressed miRNAs were identified with |log2FC|≥2 and FDR <0.05 (Benjamini–Hochberg correction), consistent with rigorous thresholds applied in comparable studies.
Cell counting kit-8 (CCK-8) and lactate dehydrogenase (LDH) assays
The CCK-8 kit (Servicebio, China) and LDH kit (Servicebio, China) were employed in accordance with the manufacturer’s instructions to assess the proliferation or toxicity of NK cells or TSCC cells.
Quantitative real-time polymerase chain reaction (qRT-PCR), Western blotting (WB) and Immunohistochemistry (IHC)/Immunofluorescence (IF)
The methods of qRT-PCR, WB and IHC/IF were conducted in accordance with previous descriptions20. The primers and antibodies utilized in this study are documented in Tables S1 and S2.
Fluorescence in situ hybridization (FISH)
The methodology of FISH was conducted in accordance with the procedures outlined in the preceding study20. The specific sequences of the SNHG26 fluorescent probes employed in this investigation can be found in Table S3.
Enzyme-linked immunosorbent assay (ELISA)
The concentrations of IFN-γ produced by NK cells were measured by IFN-γ ELISA kits (eBioscience, USA) in accordance with the manufacturer’s guidelines.
RNA immunoprecipitation (RIP)
The Magna RIP RNA-binding protein immunoprecipitation kit (Millipore, USA) was utilized to conduct RIP assays, following the guidelines provided by the manufacturer. Table S2 presents the anti-argonaute 2 (AGO2) and IgG antibodies employed in this study.
RNA pulldown assay
The RNA pulldown assay employed in this study was conducted according to the methodology outlined by ref. 45. The sequences of the SNHG26 probe for RNA pulldown are listed in Table S4.
Transfection experiment
All lentiviral vectors used in this study were purchased from Genechem (Shanghai, China). The SNHG26 siRNA target sequences used in this study are listed in Table S5.
Flow cytometry analysis
Flow cytometry was conducted using a LSR-II flow cytometer (BD Biosciences, USA). The obtained data were subsequently subjected to analysis using Flow Jo 7.6.5 software. Initially, a cell suspension of 1 × 105 cells in 20 μl was aliquoted into a 1.5 ml EP tube, to which 2 μl of IgG was added per tube. Subsequently, the mixture was incubated in the dark at 4 °C for 30 min. Following this, 1 ml of FACS solution was added, mixed thoroughly, and centrifuged at 500 rcf for 5 min. After discarding the supernatant, the cell suspension was adjusted to 20 μl, and the corresponding flow cytometry antibodies were added. The mixture was then incubated at 4 °C in the dark for another 30 min, with vortex mixing every 10 min. After 30 min, the suspension was resuspended in 1 ml of PBS, centrifuged at 500 rcf for 5 min, and the supernatant was discarded. Finally, the cells were resuspended in 200 μl of PBS and transferred to a flow cytometry tube for analysis. Firstly, cell debris was removed using the FSC-SSC plot. Lymphocyte populations were gated based on cell size and complexity, designated as Gate 1. Within Gate 1, cells that were strongly positive for both CD45 and FSC, representing a highly pure lymphocyte population, were further gated and designated as Gate 2. Subsequently, the CD3-negative and CD56-positive lymphocyte population within Gate 2 was identified as the desired NK cells (Fig. S9).
Luciferase reporter assay
The wild-type TGFB1 3’UTR and SNHG26 sequences were inserted into the pLG3 plasmid. Mutant TGFB1 3’UTR and SNHG26 pLG3 plasmids were created using mutagenesis kits (Qiagen, USA) following the manufacturer’s instructions. NK-92 cells were cultured in 96-well plates and co-transfected with a luciferase reporter vector and miR-515-5p mimic or used as a negative control with Liposome 2000 transfection reagent (Thermo Fisher, USA). After 48 h, the activities of firefly and reninase were measured using a dual-luciferase reporter assay system (Promega, USA).
Immunoprecipitation (IP) assay
In order to assess the impact of SNHG26 on the ubiquitylation process of SOX2, the NK-92 cells were subjected to transfection with the designated vectors and subsequently exposed to 10 μM MG132 for a duration of 6 h. Following this, the NK-92 cell lysates were subjected to incubation with an anti-FLAG antibody at a temperature of 4 °C. On the subsequent day, the protein complexes were captured through the utilization of protein A/G agarose beads (Thermo Fisher, USA). The levels of expression pertaining to SOX2-(HA-Ub)n within the immune complexes were subsequently determined through the employment of Western Blot analysis, utilizing an anti-HA antibody.
Co-immunoprecipitation (Co-IP) assay
Prior to conducting the experiments, NK-92 cells were transfected with either SNHG26 or Mock. Following a 48-h incubation period, NK-92 cell lysates were subjected to incubation with anti-SOX2/anti-WWP2 antibody. Subsequently, immune complexes were isolated using Protein A/G agarose beads (Thermo Fisher, USA). The expression levels of SOX2 and WWP2 in the immune complexes were assessed through Western blot analysis.
Chromatin immunoprecipitation (Chip) assay
The experiment utilized a Chip kit (MilliporeSigma, USA) and was conducted in accordance with the provided instructions. Primer sets used for HLA-DRA promoter (HLA-DRA-p) are listed in Table S1.
Single-cell RNA sequencing (scRNA-seq)
The 10×Chromium single-cell platform (10X Genomics) was utilized to conduct scRNA-seq. The cells were rinsed with a phosphate buffer solution containing 0.04% weight/volume bovine serum albumin (BSA, Sangon). Subsequently, the cells were enumerated using the CountessII Automated Cell Counter and the concentration was standardized to 1 × 106 cells per milliliter. The cDNA libraries were constructed using the 10× Chromium TM Single cell 3’ Library Kit following the manufacturer’s original protocol available on the 10x Genomics website. Cell Ranger 1.3 (http://10xgenomics.com) was employed to process the output of Chromium single cell 3’RNA-seq. The initial clustering was performed using the R package “Seurat”(Version 5.1.0), and the clustering was visualized using Loupe Cell Browser 3.1.0. The R package “SingleR” (Version 1.0.1) was utilized to identify the predominant cell types. The sequencing data were uploaded to the GEO database (GSE296834).
Next generation sequencing (NGS)
Total RNA was extracted from NK cells using TRIzol (Invitrogen, USA) and subsequently enriched for eukaryotic RNA through the use of magnetic beads with Oligo (dT). The interrupted mRNA served as a template for the synthesis of one-strand cDNA, employing a six-base random primer. Subsequently, a two-strand reaction system was prepared to synthesize two-strand cDNA, and the resulting double-strand cDNA was purified using a kit. The purified double-stranded cDNA underwent end-trimming, A-tail addition, and sequencing adapter ligation, followed by fragment size selection and, finally, PCR amplification. The library was assessed using the Agilent 2100 Bioanalyzer and sequenced using the Illumina HiSeqTM 2500 platform, resulting in the generation of 125 bp paired-end data. The differential expression of RNA (referred to as differentially expressed genes [DEGs]) was determined using the edgeR R package, with a significance threshold of |logFC|≥2, p < 0.05. The sequencing data were uploaded to the GEO database (GSE298791).
Bioinformatics analysis
The microarray data for high-activity NK cells and low-activity NK cells were retrieved and extracted from the GEO database (GSE12198). The downloaded data was in MINiML format. The differential expression of mRNAs was analyzed using the limma package of R software (version: 3.40.2). Adjusted p-values were utilized in GEO to correct for false positive results. A threshold for differential expression of mRNAs was defined as “adjusted p < 0.05 and log2 (fold change) >1 or log2 (fold change) <−1”. The sequencing and TME data of OSCC samples were obtained from the TGCA-OSCC database, and subsequently subjected to Weighted gene co-expression network analysis (WGCNA). Initially, the expression information of all genes in the samples was extracted and subjected to weighted correlation analysis. Subsequently, hierarchical clustering analysis was conducted based on the weighted correlation, and the resulting clusters were segmented according to predefined criteria to identify distinct gene modules. Subsequently, an examination was conducted on the interconnection between the models, whereby the interaction network of diverse models was observed from a systemic perspective. This analysis led to the derivation of the Topological overlap matrix (TOM) and the creation of a correlation map. Ultimately, the genomic modules were thoroughly scrutinized, resulting in the identification of the module exhibiting the highest association with NK cells function. Consequently, the internal gene network within this module was extracted for further investigations. The evaluation of the combination of SNHG26 and SOX2/WWP2 was conducted using the bioinformatic database RPISeq, accessible at http://pridb.gdcb.iastate.edu/RPISeq/.
Mendelian randomization (MR) analysis
The inverse-variance weighted (IVW), weighted median, and MR-Egger regression methods were used in a two-sample MR analysis. The publicly accessible summary statistics datasets of genome-wide association studies (GWAS) were utilized in this study. The Instrumental variable was chosen based on the following selection criteria: (1) Potential instrumental variables (IVs) were identified by selecting single nucleotide polymorphisms (SNPs) associated with each genus using locus wide significance thresholds (p < 5.0 × 10−6); (2) To ensure independence among the genetic variants, the SNPs were additionally subjected to linkage disequilibrium (LD) splicing, with a distance threshold of 10,000 kb and an R2 value below 0.001; (3)To ensure the alignment of the effects of SNPs on both exposure and diseases, any palindromic SNPs exhibiting minor allele frequencies exceeding 0.3 were eliminated. All MR analyses were performed in the MR Base platform (App version: 1.4.3 8a77eb, R version: 4.0.3).
Statistical analysis
Statistical analysis was conducted using SPSS version 22.0, with the results being analyzed through analysis of variance (ANOVA). The significant differences between the groups were evaluated using the LSD test for multiple comparisons. All data were presented as the mean ± standard deviation (SD), and a significance level of p < 0.05 was established.
Data availability
The datasets generated during this study are included in this article, its supplementary information files, and the GEO repository under accession numbers (GSE296834, GSE298791, and GSE207675). Additional data were available from the corresponding author upon reasonable request.
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 72, 7–33 (2022).
Sung, H. et al. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Xie, L. & Shang, Z. Changing trend of oral cancer disease burden in china from 1990 to 2019 and the forecast for the next 20 years. Oral. Dis. 30, 195–206 (2022).
Walsh, T. et al. Clinical assessment for the detection of oral cavity cancer and potentially malignant disorders in apparently healthy adults. Cochrane Database Syst. Rev. 12, D10173 (2021).
Caudell, J. J. et al. Nccn guidelines(r) insights: head and neck cancers, version 1.2022. J. Natl Compr. Cancer Netw. 20, 224–234 (2022).
Subramaniam, N. et al. Geographical heterogeneity in the American joint committee on cancer oral cancer staging and prognostic implications. Oral. Oncol. 113, 105122 (2021).
de Oliveira, T. B., Marta, G. N., de Castro, J. G. & Kowalski, L. P. Induction chemotherapy for advanced oral cavity cancer. Curr. Oncol. Rep. 23, 129 (2021).
Zhong, L. P. et al. Randomized phase iii trial of induction chemotherapy with docetaxel, cisplatin, and fluorouracil followed by surgery versus up-front surgery in locally advanced resectable oral squamous cell carcinoma. J. Clin. Oncol. 31, 744–751 (2013).
Klemm, F. et al. Interrogation of the microenvironmental landscape in brain tumors reveals disease-specific alterations of immune cells. Cell 181, 1643–1660 (2020).
Tiwari, A., Trivedi, R. & Lin, S. Y. Tumor microenvironment: barrier or opportunity towards effective cancer therapy. J. Biomed. Sci. 29, 83 (2022).
Watermann, C. et al. Recurrent hnscc harbor an immunosuppressive tumor immune microenvironment suggesting successful tumor immune evasion. Clin. Cancer Res. 27, 632–644 (2021).
Zhou, Y., Cheng, L., Liu, L. & Li, X. Nk cells are never alone: crosstalk and communication in tumour microenvironments. Mol. Cancer 22, 34 (2023).
Li, Y. et al. Kir-based inhibitory cars overcome car-nk cell trogocytosis-mediated fratricide and tumor escape. Nat. Med. 28, 2133–2144 (2022).
Tenchov, R. et al. Exosomes horizontal line nature’s lipid nanoparticles, a rising star in drug delivery and diagnostics. Acs Nano 16, 17802–17846 (2022).
Clancy, J. W. & D’Souza-Schorey, C. Tumor-derived extracellular vesicles: multifunctional entities in the tumor microenvironment. Annu. Rev. Pathol. Mech. Dis. 18, 205–229 (2023).
Yang, Z., Ma, R., Li, J. & Zhao, L. Noncoding rnas in esophageal cancer: a glimpse into implications for therapy resistance. Pharmacol. Res. 188, 106678 (2023).
Adnane, S., Marino, A. & Leucci, E. LncRNAs in human cancers: signal from noise. Trends Cell Biol. 32, 565–573 (2022).
Yu, Z. et al. Exosomal loc85009 inhibits docetaxel resistance in lung adenocarcinoma through regulating atg5-induced autophagy. Drug Resist. Update 67, 100915 (2023).
Saeinasab, M., Atlasi, Y. & M, M. M. Functional role of lncrnas in gastrointestinal malignancies: the peculiar case of small nucleolar rna host gene family. FEBS J. 291, 1353–1385 (2024).
Jiang, Q. et al. LncRNA SNHG26 promoted the growth, metastasis, and cisplatin resistance of tongue squamous cell carcinoma through pgk1/akt/mtor signal pathway. Mol. Ther.-Oncolytics. 24, 355–370 (2022).
Tong, L. et al. Nk cells and solid tumors: therapeutic potential and persisting obstacles. Mol. Cancer 21, 206 (2022).
Pan, K. et al. CAR race to cancer immunotherapy: from CAR T, CAR NK to CAR macrophage therapy. J. Exp. Clin. Cancer Res. 41, 119 (2022).
Liu, E. et al. Use of CAR-transduced natural killer cells in CD19-positive lymphoid tumors. N. Engl. J. Med. 382, 545–553 (2020).
Klingemann, H. Are natural killer cells superior CAR drivers?. Oncoimmunology 3, e28147 (2014).
Pende, D. et al. Killer Ig-like receptors (KIRs): their role in nk cell modulation and developments leading to their clinical exploitation. Front. Immunol. 10, 1179 (2019).
Xavier, C. P. R. et al. The role of extracellular vesicles in the hallmarks of cancer and drug resistance. Cells 9, 1141 (2020).
Daßler-Plenker, J., Küttner, V. & Egeblad, M. Communication in tiny packages: exosomes as means of tumor-stroma communication. Biochim. Biophys. Acta Rev. Cancer 1873, 188340 (2020).
Erokhina, S. A. et al. HLA-DR-expressing nk cells: effective killers suspected for antigen presentation. J. Leukoc. Biol. 109, 327–337 (2021).
Whyte, C. E. et al. Context-dependent effects of IL-2 rewire immunity into distinct cellular circuits. J. Exp. Med. 219, e20212391 (2022).
Tschernia, N. P. & Gulley, J. L. Tumor in the crossfire: inhibiting TGF-β to enhance cancer immunotherapy. Biodrugs 36, 153–180 (2022).
Regis, S., Dondero, A., Caliendo, F., Bottino, C. & Castriconi, R. Nk cell function regulation by TGF-beta-induced epigenetic mechanisms. Front. Immunol. 11, 311 (2020).
Horwitz, D. A., Gray, J. D. & Ohtsuka, K. Role of nk cells and tgf-beta in the regulation of t-cell-dependent antibody production in health and autoimmune disease. Microbes Infect. 1, 1305–1311 (1999).
Ahmadi, A., Najafi, M., Farhood, B. & Mortezaee, K. Transforming growth factor-beta signaling: tumorigenesis and targeting for cancer therapy. J. Cell. Physiol. 234, 12173–12187 (2019).
Zimta, A. A. et al. An emerging class of long non-coding RNA with oncogenic role arises from the snorna host genes. Front. Oncol. 10, 389 (2020).
Xin, L. et al. Transfer of lncRNA CRNDE in TAM-derived exosomes is linked with cisplatin resistance in gastric cancer. EMBO Rep. 22, e52124 (2021).
Novak, D. et al. Sox2 in development and cancer biology. Semin. Cancer Biol. 67, 74–82 (2020).
Zhang, R., Zhang, J., Luo, W., Luo, Z. & Shi, S. Wwp2 is one promising novel oncogene. Pathol. Oncol. Res. 25, 443–446 (2019).
Mahlokozera, T. et al. Competitive binding of E3 ligases TRIM26 and WWP2 controls SOX2 in glioblastoma. Nat. Commun. 12, 6321 (2021).
Yan, J. & Xu, H. Regulation of transforming growth factor-beta1 by circanks1b/mir-515-5p affects the metastatic potential and cisplatin resistance in oral squamous cell carcinoma. Bioengineered 12, 12420–12430 (2021).
Chen, W. et al. Decreased expression of mitochondrial miR-5787 contributes to chemoresistance by reprogramming glucose metabolism and inhibiting MT-CO3 translation. Theranostics 9, 5739–5754 (2019).
Sachs, N. et al. A living biobank of breast cancer organoids captures disease heterogeneity. Cell 172, 373–386 (2018).
He, F. et al. Establishment and identification of patient-derived xenograft model for oral squamous cell carcinoma. J. Oncol. 2022, 3135470 (2022).
Ye, C. et al. A rapid, sensitive, and reproducible in vivo pbmc humanized murine model for determining therapeutic-related cytokine release syndrome. FASEB J. 34, 12963–12975 (2020).
Iwai, K., Minamisawa, T., Suga, K., Yajima, Y. & Shiba, K. Isolation of human salivary extracellular vesicles by iodixanol density gradient ultracentrifugation and their characterizations. J. Extracell. Vesicles 5, 30829 (2016).
Zhang, P. F. et al. Cancer cell-derived exosomal circUHRF1 induces natural killer cell exhaustion and may cause resistance to anti-pd1 therapy in hepatocellular carcinoma. Mol. Cancer 19, 110 (2020).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (No. 82403716), Jiangxi Natural Science Foundation (No. 20232BAB216073, 20242BAB25514), and Key Projects of Jiangxi Administration of Traditional Chinese Medicine (No. GZY-KJS-2023-028). The authors wish to thank the First Affiliated Hospital of Nanchang University for its help in the detection of experimental samples. The article received language editing support from HOME for Researchers.
Author information
Authors and Affiliations
Contributions
Q.J. and J.Q. were responsible for the conception, design, and study supervision. Y.X. and F.H. were responsible for the development of the methodology, analysis, and experiments. C.L. performed the statistical and bioinformatic analyses. Q.J. were responsible for tissue and clinical data collection. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jiang, Q., Xin, Y., He, F. et al. Exosomal SNHG26 mediates immunosuppression by impairing NK cells in tongue cancer. npj Precis. Onc. 9, 402 (2025). https://doi.org/10.1038/s41698-025-01185-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01185-0