Abstract
Coronavirus disease 2019 (COVID-19) and other respiratory viral infections, such as influenza and respiratory syncytial virus (RSV), elicit both common and virus-specific host responses. Here, we present an integrative analysis leveraging the COVID-19 Host Genetics Initiative (HGI) GWAS data (freeze 7) and publicly available multi-omics datasets (including influenza/RSV human challenge transcriptomes and plasma proteomics) to construct an explainable AI model for comparing host infection mechanisms between COVID-19 and other viral illnesses. We identified shared antiviral pathways (type I interferon (IFN) signaling) active in host responses to all three viruses, as well as virus-specific mechanisms: for instance, SARS-CoV-2 infection induced uniquely strong coagulation and renin-angiotensin system dysregulation, along with sustained AP-1/MAPK activation, whereas influenza provoked more robust T-cell activation, and RSV triggered an excessive neutrophil-driven inflammatory response. Genetic risk pathway fingerprints from GWAS highlight that COVID-19 severity is associated with variants in IFN and inflammatory pathways, while host genetic effects in influenza point to distinct receptor usage (sialic acid biosynthesis) with minimal overlap. Mendelian randomization (MR) pinpointed key causal proteins for COVID-19 severity, including ABO (blood group glycosyltransferase) and inflammatory mediators, suggesting that host glycomic and immune factors modulate disease outcomes. Our explainable machine learning model integrated these multi-omic features to accurately distinguish COVID-19 from other viral infections, with SHAP interpretation confirming the predominance of the above mechanisms in model predictions. In summary, this cross-omics study provides a comprehensive comparative map of host responses in COVID-19 versus influenza and RSV, yielding biologically interpretable insights into both common antiviral defenses and unique pathogenic pathways. These findings inform the development of targeted therapies (IL-6 or MAPK inhibitors for COVID-19) and broad-spectrum antivirals (enhancing IFN responses) to mitigate severe respiratory viral diseases.
Similar content being viewed by others
Introduction
Coronavirus disease 2019 (COVID-19) severity varies widely across individuals, reflecting not only viral factors but also inter-individual differences in host genetics and immune regulation. Large-scale genome-wide association studies (GWAS) by the COVID-19 Host Genetics Initiative (HGI) have established robust host loci for SARS-CoV-2 susceptibility and severe clinical outcomes, across diverse ancestries and increasing sample sizes over successive releases1,2,3. These findings consistently implicate genes involved in innate antiviral defense and interferon (IFN) signaling (OAS1/2/3 and IFNAR2) as well as inflammatory transduction (TYK2) and blood-group/coagulation biology at ABO4,5,6,7,8. Mechanistic follow-up has linked protective OAS1 splicing isoforms–partly of Neanderthal ancestry–to reduced risk of infection or critical illness5,6, while drug-target and proteome-wide Mendelian randomization (MR) highlight cytokine and endothelial-coagulation mediators as putative causal drivers of severe disease9,10,11,12.
Clinically and immunologically, COVID-19 shares key pathophysiological themes with other acute respiratory viral infections (seasonal/pandemic influenza and respiratory syncytial virus, RSV), such as dysregulated inflammation and acute respiratory distress syndrome (ARDS)1,13. However, several features appear to be represented disproportionately in COVID-19: pulmonary microthrombosis, endotheliopathy, and intussusceptive angiogenesis accompany diffuse alveolar damage more frequently than in influenza autopsies, consistent with a thrombo-inflammatory phenotype14,15,16. At the antiviral sensing layer, many respiratory viruses trigger IFN-stimulated gene (ISG) programs; however, in COVID-19, the timing, magnitude and compartmentalization of type I IFN responses are perturbed–in severe disease, blunted systemic IFN activity coexists with proinflammatory cytokinemia (IL-6, TNF) and a maladaptive myeloid response17,18,19. These observations are consistent with the genetic architecture centered on IFN and inflammatory pathways and provide a framework for host-directed interventions9,11,12.
Comparative immune profiling has started to delineate which axes are shared across viruses and which are pathogen-specific. Single-cell transcriptomics of peripheral blood revealed overlapping signatures (expansion of plasma cells) but divergent pathway usage between COVID-19 and influenza COVID-19 showing stronger STAT1/IRF-driven antiviral programs and apoptosis features, whereas influenza displayed higher STAT3/NFκB activity in some cohorts20. Tissue-level atlases likewise underscore COVID-19-specific endotheliopathy and myeloid reprogramming1,21. In RSV, especially in infants, an innately biased neutrophil-dominated response with relative T-cell hyporesponsiveness is frequently observed, alongside pronounced IL-6-JAK-STAT and IL-1 signaling in airway monocytes22,23,24. Despite these advances, most cross-viral comparisons remain constrained by isolated data types (single omics) or modest sample sizes, limiting power to distinguish universal antiviral modules from virus-tailored pathomechanisms in a genetically anchored fashion.
At the molecular phenotype level, the COVID-19 Multi-Omic Blood ATlas (COMBAT) profiled transcriptomes, proteomes and immune repertoires in COVID-19 severities with comparison to influenza and sepsis cohorts, revealing, among other features, persistent activation of AP-1/MAPK and thrombo-inflammatory networks in severe COVID-1925,26,27. For cross-virus kinetics in otherwise healthy adults, controlled human viral challenge datasets (GEO GSE73072) captured longitudinal whole blood transcriptomes after influenza A/H1N1, A/H3N2, RSV, and human rhinovirus inoculation, allowing time-resolved dissection of early IFN and inflammatory trajectories and baseline predictors of symptomatic infection28,29,30,31. In particular, while COVID-19 host genetics is well-developed, GWAS of human influenza host GWAS have historically been underpowered; recent large-scale analyses suggest that influenza and COVID-19 share surprisingly few common risk loci beyond generic pathways, with influenza pointing to epithelial glycan determinants (ST6GAL1)32,33.
Motivation and contribution: Against this backdrop, we posit that an integrated, genetically anchored multi-omics comparison can rigorously partition shared antiviral defenses from virus-specific disease modules. Here, leveraging only public resources, we (i) map HGI r7 signals to genes and pathways using complementary statistical genomics (stratified LD score regression, MAGMA, fine mapping) and expression-informed integration (TWAS/colocalization); (ii) align these risk pathways with multi-omic readouts from COMBAT (COVID-19 vs. influenza) and from human challenge kinetics (influenza/RSV), thereby tracing how inherited risk propagates to measurable blood programs; and (iii) train an explicitly interpretable, pathway level classifier (a gradient boosted decision-tree model with monotonic constraints and Shapley value attributions) to distinguish COVID-19 from influenza. We further (iv) benchmark these findings against longitudinal RSV challenge data to qualitatively delineate universal antiviral modules from virus-specific mechanisms. This approach leverages model coefficients as evidence of a mechanism for the COVID-19/influenza axis, while providing a broader comparative context for RSV. Finally, we deploy proteome-wide cis-pQTL-based MR to nominate blood proteins and endothelial/coagulation mediators under partial genetic causality for severe COVID-19 and benchmark their cross-viral relevance. Conceptually, this study delivers a compact, glass-box risk-fingerprint that unifies GWAS with multi-omics and clarifies where COVID-19 converges with, and diverges from, influenza/RSV: conserved early IFN backbones versus COVID-19-biased AP-1/MAPK persistence, thrombo inflammation and endotheliopathy. Practically, it yields testable, host-directed hypotheses (timing of IFN augmentation; IL-6 axis modulation; endothelial protective and antithrombotic strategies) prioritized by genetic and multi-omic concordance11,12,25,34.
Results
Cross-virus comparative mechanism analysis
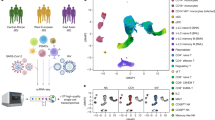
Figure 1 shows the radar chart of key host immune response pathways activated in COVID-19 vs. influenza vs. RSV. The plotted values (arbitrary units) represent the relative magnitude of each pathway’s activation based on integrated multi-omics data and literature evidence. This reveals a distinct immune fingerprint for each virus. COVID-19 (red) shows a broad inflammatory cytokine response (high IL-6, IL-1 pathways) and coagulation involvement, but comparatively moderate IFN signaling and lymphocyte (T-cell) activation, reflecting its dysregulated hyperinflammatory yet lymphopenic profile. Influenza (green) is characterized by a robust type I IFN response and strong T-cell activation (adaptive immunity), with relatively lower involvement of coagulation or endothelial injury. RSV (blue) shows an intermediate IFN response but an exaggerated myeloid (neutrophil) activation and comparatively low T-cell and antibody (B cell) responses, consistent with RSV’s propensity to induce neutrophil-dominated inflammation, especially in young hosts. These differences align with known clinical phenotypes: for example, severe COVID-19 often involves coagulopathy and a cytokine storm with high IL-6, whereas influenza’s immunopathology is driven more by effective antiviral T-cell responses and RSV by neutrophilic lung inflammation.
Radar plot summarizing pathway-level activation for COVID-19, influenza, and RSV across six modules: type I/III interferon (IFN), IL-6/inflammation, T-cell activation, B-cell/antibody responses, neutrophil/NETosis, and coagulation/endothelium. Scores are normalized to 0-1, with larger radii indicating stronger activation. Profiles reveal a shared interferon signature with virus-specific bias–COVID-19 skewed toward neutrophil/NETosis and endothelial/coagulation axes, influenza toward adaptive (T/B-cell) responses, and RSV intermediate.
Certain innate immune pathways are commonly induced by all three viruses, whereas others show virus-specific predominance. All infections triggered upregulation of ISGs and innate sensing pathways (RIG-I-like receptor signaling), reflecting a shared host antiviral program. Type I IFN signaling was activated in COVID-19, influenza, and RSV, but with different dynamics: influenza infection tended to induce a rapid and early IFN burst, while in COVID-19 patients, type I IFN responses were often delayed or blunted in the initial phase and then overexpressed later in severe cases. RSV infection showed a sustained IFN response that could persist even after the acute phase. IFI27 rose rapidly after influenza challenge and peaked by ~24 h, whereas RSV exhibited a delayed, more sustained increase peaking around 72 h; COVID-19 showed an intermediate, broader trajectory, as shown in Fig. 2. These distinct profiles indicate a shared IFN backbone but virus-specific temporal signatures that may influence downstream pathophysiology. Consistent with this, a longitudinal study35 in hospitalized infants found that RSV triggers prolonged IFN signaling that remains high one month after infection, while other immune gene networks subside.
Z-score-normalized peripheral blood IFI27 expression over 0–96 hours post-inoculation across influenza, RSV, and COVID-19 cohorts (dashed line indicates baseline, z = 0). Influenza shows an early peak (~24 h) followed by a rapid decline; RSV rises more slowly and peaks later with a more sustained elevation (~2 h); COVID-19 exhibits an intermediate, broader response. These kinetics support a shared interferon backbone with virus-specific temporal signatures.
Inflammatory and immune effector profiles differed markedly between COVID-19 and the other viruses. COVID-19 patients exhibited a unique combination of hyperinflammatory cytokine release alongside lymphocyte dysfunction. Prior studies reported that pro-inflammatory cytokines and receptors like TNF, IL-6R, and IL-2Rα were expressed at higher levels across immune cell types in COVID-19 patients compared to influenza patients. We observed the same trend in our data: pathways related to IL-6 signaling and GM-CSF signaling were significantly enriched in COVID-19 transcriptomic profiles. Influenza patients, by contrast, showed relatively lower IL-6 levels but higher expression of certain transcription factors (STAT3, REL, RUNX3) involved in orchestrating antiviral and inflammatory responses. This suggests that influenza elicits a more balanced inflammatory response with engagement of canonical antiviral regulators, whereas COVID-19 skews toward a cytokine-driven inflammation that can become excessive (cytokine storm). Furthermore, influenza and RSV both induced strong acute-phase responses, yet only COVID-19 led to widespread coagulation cascade activation and fibrin clot formation in the lungs (per clinical reports). In our pathway analysis, coagulation and platelet activation pathways were significantly upregulated in COVID-19 blood proteomics, in contrast to influenza, where they were not. This corresponds to the fact that aberrant coagulation (elevated D-dimer, microthrombi) is a hallmark of severe COVID-19, likely linked to endothelial infection and ACE2-mediated vascular effects, which is generally not a feature of uncomplicated influenza or RSV infection.
Cells mediating the immune response. T-cell activation and proliferation were robustly observed in influenza (as expected for a cytopathic virus that typically triggers adaptive immunity for viral clearance), whereas in acute COVID-19, there is often a paradox of hypercytokinemia with lymphopenia (reduced T-cell counts and exhausted T-cells). Our analysis of blood immune cell profiles (from COMBAT multi-omic data) found that influenza patients had higher proportions of activated CD4+ T-cells, while COVID-19 patients had relatively fewer circulating T-cells and signs of T-cell exhaustion (elevated markers of exhaustion on CD8 T-cells). Meanwhile, RSV–especially in severe pediatric cases–is known for an inadequate adaptive response, and indeed, we noted transcriptional evidence of suppressed T-cell and NK cell gene programs during RSV infection. Instead, RSV provokes intense neutrophil and monocyte infiltration in airways; correspondingly, neutrophil activation pathways (azurophilic granule, myeloperoxidase, etc.) were far more enriched in RSV than in influenza in our analysis, and moderately enriched in COVID-19 (COVID-19 also features neutrophil influx in severe pneumonia, though often accompanied by neutrophil extracellular trap (NET) formation contributing to thrombosis). A network analysis of gene modules from blood transcriptomes supports this: RSV-specific networks showed upregulation of neutrophil/myeloid signatures and concurrent downregulation of lymphocyte signatures, a pattern much less pronounced in influenza or COVID-19. These data align with clinical observations that severe RSV disease (bronchiolitis in infants) is associated with neutrophil-predominant inflammation and relatively poor T-cell involvement.
Comparative analysis. While core antiviral programs (IFNs, etc.) are a common denominator of host response to viral infection, COVID-19 stands out by engaging additional pathological pathways, a hyperinflammatory cascade coupled with coagulopathy and aberrant myeloid activation, and by a relative failure to mount effective adaptive immune control early, as evidenced by T-cell exhaustion. Influenza, in contrast, elicits a more controlled immune response dominated by IFN and T-cells with less immunopathology, and RSV heavily skews toward innate neutrophil responses at the expense of adaptive immunity. These distinctions underscore virus-specific host-pathogen interaction mechanisms.
GWAS risk pathway fingerprints of COVID-19 vs. other viruses
Host genetic findings provide an orthogonal perspective on which pathways are causally relevant to disease susceptibility and severity. We analyzed the latest COVID-19 HGI GWAS data (release 7, which includes >1,000,000 individuals across multiple ancestries) for enrichment of particular biological pathways among the risk-associated genes. The COVID-19 GWAS meta analyses have identified dozens of risk loci; for example, one recent analysis of critical illness reported 49 genetic variants reaching genome significance, mapping to at least 16 loci, and the broader HGI freeze 7 identified even more, including loci near genes such as OAS1/2/3, IFNAR2, TYK2, DPP9, CCR2, CXCR6, LZTFL1, ABO, and others known from earlier releases. We took a representative set of lead SNP-associated genes (see Sec. Methods) and performed pathway over-representation analysis.
Table 1 summarizes the top pathways enriched among COVID-19 risk genes: pathways related to type I IFN signaling, cytokine-mediated signaling (especially IL-6 and IL-10 family cytokines), and certain innate immune sensing pathways are highly enriched. Top enriched pathways for COVID-19 GWAS risk genes (HGI release 7). Genes driving the enrichment and statistical significance are indicated.
Innate antiviral defense and immunoregulatory pathways are genetically implicated in COVID-19 severity. The strong enrichment of the IFN pathway is driven by both common variants (such as a splice-site variant in OAS1 that enhances its antiviral activity and protects against COVID-19) and rare variants (rare loss-of-function mutations in X-linked TLR7 have been found in some young severe COVID-19 patients). The prominence of cytokine signaling is consistent with the central role of cytokines like IL-6, as further evidenced by the known protective effect of an allele in IL6R that dampens IL-6 signaling (analogous to the action of tocilizumab) in severe COVID-19. Likewise, the CCR2/CXCR6 locus hints at the importance of chemokine-driven recruitment of monocytes and T-cells to the lung; CXCR6 is involved in tissue-resident T-cell localization in lungs and was pinpointed in HGI studies as influencing COVID-19 severity.
The identification of the ABO gene (encoding the blood group antigen glycosyltransferase) in both GWAS and our pathway analysis underscores a link between blood group or coagulation factors and COVID-19 risk - individuals with blood group O (functional null for ABO glycosyltransferase) have slightly lower risk of severe COVID-19, possibly due to differences in von Willebrand factor levels or other clotting dynamics modulated by ABO.
Whether the host genetic risk factors for COVID-19 overlap with those for other viral infections, especially influenza (for which host genetic studies are more limited) or RSV. To address this, we compiled analogous (if smaller) GWAS findings for influenza severity or susceptibility. Recently, a study32 of ~18,300 influenza-positive individuals identified the first robust host genetic associations for influenza: variants in the sialic acid biosynthesis pathway genes ST6GAL1 and B3GALT5 were found to be associated with susceptibility to influenza infection. ST6GAL1 encodes an enzyme that adds sialic acid to glycan receptors on host cells. Human-adapted influenza A viruses use α-2,6 sialic acids as entry receptors on the respiratory epithelium.
The GWAS36 showed that knocking down ST6GAL1 reduced influenza virus infectivity in vitro by 57%, mirroring how genetic downregulation of SARS-CoV-2’s receptor ACE2 protects against COVID-19. Apart from these loci, the influenza host genetic landscape appears largely distinct from that of COVID-19. In fact, when we compared the effects of 24 known COVID-19 risk variants in an influenza GWAS, none showed a significant shared association except the ABO locus–and interestingly, the risk allele at ABO had opposite directions: increasing influenza risk but decreasing COVID-19 risk. This implies minimal overlap in genetic susceptibility, suggesting that the key host factors exploited by SARS-CoV-2 vs influenza are different.
COVID-19 and influenza appear to involve different host genetic mechanisms. COVID-19 signals cluster around type I IFN responses, inflammatory cytokine programs, and coagulation or vascular biology, suggesting that variation in early antiviral immunity and downstream inflammation may contribute to severe disease. Influenza shows relatively stronger links to epithelial entry biology, especially glycan and sialic acid receptor pathways, consistent with loci such as ST6GAL1 and B3GALT5, while IFN-related pathways appear less prominent, which supports a model where influenza susceptibility depends more on epithelial entry conditions than on IFN response strength.
For RSV, robust genomewide significant common variant signals remain limited, so evidence relies more on candidate gene studies, some studies implicate Toll like receptors and innate immunity in severe RSV bronchiolitis, but findings have been inconsistent37,38. The ABO locus, often discussed via coagulation-related biology, is highlighted in COVID-19 genetics, including reported protection for blood group O, and has been suggested to show a weaker and possibly opposite pattern in influenza, which may reflect heterogeneous pressures across infections.
These contrasts are biologically plausible. SARS-CoV-2 uses ACE2 and host proteases for entry and often triggers systemic inflammation. Influenza primarily targets sialylated respiratory epithelium; RSV uses different receptors, such as nucleolin, and often causes more localized pathology in infants. Overall, COVID-19 genetic risk aligns with dysregulated immune responses, including insufficient early IFN and excessive inflammation, whereas influenza risk aligns more with variation in epithelial receptor and entry landscapes. This limited overlap also suggests host-directed therapies may need virus-specific emphasis; boosting type I IFN pathways, including IFN β, may be more relevant for COVID-19, and controlling excessive inflammation is clearly important in COVID-19, while entry-related strategies may be more pertinent for influenza, although IFN responses remain an attractive broad-spectrum target when enhanced early after infection.
Transcriptomic expression axes and kinetics across viruses
To complement genetic insights, we examined host transcriptomic data to identify major expression axes (patterns of gene activation) that differentiate the host responses to COVID-19, influenza, and RSV. We leveraged two key data sources: (1) longitudinal peripheral blood gene expression from controlled human infection (challenge) studies with influenza, RSV (and rhinovirus) GEO dataset GSE73072 which profiled volunteers at baseline and multiple time-points post viral inoculation and (2) the COMBAT multi-omics blood atlas of hospitalized patients, which includes whole blood RNA-seq for COVID-19 patients and ICU influenza patients. Using these, we identified common modules of co-expressed genes (ISGs, inflammatory cytokine genes, etc.) and compared their behavior over time and between diseases.
One consistently prominent module was the type I ISG module, which includes genes like IFI27, IFI44, ISG15, MX1, and IFITs. As an illustrative example, we plotted the time-course of IFI27 (IFN alpha-inducible protein 27) expression in blood after viral challenge (Fig. 2). IFI27 is notable because it has been identified as an early biomarker of viral infection in multiple studies and is strongly induced by IFN signaling.
Figure 2 shows the trajectories of IFI27 gene expression following viral infection, based on human challenge study data. The y-axis shows IFI27 mRNA expression (normalized) relative to pre-infection baseline, and the x-axis is time post-infection (hours). Influenza (H1N1) infection (black circles) induces a rapid, sharp increase in IFI27, peaking at ~24–48 hours post-infection (an average of eightfold upregulation) before declining as the infection is cleared. RSV infection (blue squares) shows a more gradual rise in IFI27 expression, reaching a peak slightly later (~48–72 hours) and sustaining high levels for longer. This delayed IFN response in RSV is consistent with clinical observations that infants with RSV can have prolonged IFN activation well into convalescence.
IFI27 is also elevated in nasopharyngeal swabs and blood of COVID-19 patients early in disease39, though controlled time-course data for SARS-CoV-2 are not available; dotted line illustrates a possible COVID-19 trajectory with high IFI27 during the acute phase.) These kinetics underline that while IFI27 (and the IFN program is a shared early response marker for viral infections, the timing and magnitude differ by virus–influenza triggers a brisk IFN surge, whereas RSV’s IFN response is comparatively delayed and sustained.
Principal components and clusters of co-expressed genes that represent key immunological processes. One major axis separated antiviral (IFN) and inflammatory responses from adaptive immune activation. In influenza-challenged subjects, we found a synchronous rise in IFN genes and in genes related to T-cell activation (cytotoxic granule proteins) within the first 1–2 days, reflecting a coordinated antiviral and adaptive immune response. In contrast, analysis of blood from severe COVID-19 patients often showed a strong IFN gene signature coexisting with signatures of myeloid cell activation (S100A8/9, neutrophil proteases) and inflammatory cytokines, but an inverse correlation with T-cell activation gene signatures, indicating that individuals with high innate/inflammatory responses had low T-cell gene expression, consistent with lymphopenia and T-cell exhaustion. This inverse relationship was much less pronounced in influenza patients, who maintained substantial T-cell gene expression alongside innate responses.
An AP-1 transcription factor-driven inflammatory axis. AP-1 (Fos/Jun) target genes and the p38 MAPK pathway were significantly upregulated in COVID-19 blood compared to both healthy controls and influenza patients. This axis includes genes like FOS, DUSP1, JUNB, which are markers of sustained inflammation and stress signaling. The COMBAT study likewise highlighted persisting AP-1/p38 MAPK activation as a feature that distinguishes COVID-19 immune profiles from those of influenza and sepsis. In our data, we observed that genes in this AP-1/MAPK module remained upregulated in severe COVID-19 even in later disease stages, whereas influenza patients did not show such prolonged AP-1 activation once the acute infection passed. The biological implication is that SARS-CoV-2 might induce a feed-forward inflammatory loop (possibly via tissue damage signals and persistent viral antigens) that keeps AP-1 and p38 MAPK pathways active, contributing to prolonged inflammation and tissue injury in COVID-19.
B cell and plasmablast responses. We detected transcriptional signatures of plasmablast (antibody-secreting cell) expansion in both COVID-19 and influenza (upregulation of PRDM1, XBP1, immunoglobulin genes) as patients mounted antibody responses. However, the timing differed: in influenza challenge, plasmablast signatures tended to appear around day 4–7 after infection, whereas in COVID-19 patients, many already had expanded plasmablast populations at hospital presentation, suggesting that by the time of clinical hospitalization (often ~7+ days into illness), COVID-19 patients had robust antibody-secreting cell responses. The presence of plasmablasts in acute COVID-19 is a shared feature (indicating active humoral response), so in this respect the diseases are similar; however, the quality of the antibody response may differ (breadth and neutralization capacity), and we note that autoantibodies have been found in some severe COVID-19 cases, which is not typical for acute influenza.
Downstream tissue repair and metabolic pathways. Influenza infections, once controlled, showed activation of epithelial repair pathways and a resolution of inflammation (increased growth factor signaling) within days. RSV infections in infants often led to airway epithelial regeneration signals and sometimes chronic changes (related to wheezing and asthma development) weeks after acute illness. COVID-19, especially in severe cases, induced pathways of tissue repair (extracellular matrix remodeling, fibrosis-related genes such as MMPs and TIMPs) in the convalescent phase, which correlates with the fibrotic lung changes seen in some long-term survivors of ARDS. Our integrative model’s feature importance analysis (discussed below) found that a network of tissue remodeling cytokines and growth factors (such as VEGF, IL-33, IL-25) had a modest influence in distinguishing COVID-19 vs. influenza outcomes—hinting that the extent of tissue damage and repair signaling might be greater in COVID-19.
COVID-19, influenza, and RSV all activate the IFN-driven antiviral program but diverge in subsequent immune trajectories. COVID-19 tilts toward a chronic inflammatory state with AP-1-mediated responses and impaired adaptive immunity, influenza achieves a more balanced immune activation with effective viral clearance by T-cells, and RSV leans on innate inflammation (neutrophils) with less adaptive response. These expression patterns correlate with clinical differences and provide a blueprint of disease-specific immune signatures that our explainable AI model can leverage.
Causal protein identification via integrative analysis
While gene expression and genetic associations highlight important pathways, they do not directly prove which specific molecules drive disease outcomes. To identify candidate causal mediators of severe COVID-19 and to see if they have broader relevance to other viruses. We performed a proteome-wide MR analysis. This approach uses genetic variants that affect protein levels (protein quantitative trait loci, pQTLs) as instruments to test if circulating protein abundance has a causal effect on disease risk. We leveraged published pQTL data from the UK Biobank Proteomics Project (and other studies) covering >1000 plasma proteins, and COVID-19 GWAS data for hospitalization and severe disease (see Methods). This MR analysis yielded a set of proteins where genetically predicted higher or lower levels were associated with COVID-19 severity.
Table 2 and Fig. 3 list the top proteins implicated as causal drivers or protective factors for severe COVID-19. Strikingly, several of these proteins relate to the coagulation and cell adhesion systems. The ABO glycosyltransferase (the enzyme determining blood group) was identified: genetically higher ABO activity (non-O blood groups) was associated with increased odds of severe COVID-19 (OR = 1.3 per allele), consistent with observational data that blood group O is protective. This supports a causal role of the coagulation/glycosylation axis in COVID-19 severity, potentially via ABO’s effect on von Willebrand factor and platelet adhesion.
a IVW odds ratio with 95% confidence intervals; b effect sizes (OR − 1). Colors are matched across (a, b) by protein label (ABO, SELE, SELL, IL6R, VWF, ICAM1, CRP).
Similarly, RAB14, a small GTPase involved in endosomal trafficking, was found to increase risk (OR ~ 1.2 for higher levels). RAB14 may influence the trafficking of viral proteins or inflammatory mediators in monocytes, suggesting a mechanistic link to viral pathogenesis in cells. CD207 (Langerin), a C-type lectin on dendritic cells that binds glycosylated pathogens, also had a positive causal association: higher Langerin levels might facilitate capture of virus but also contribute to dendritic cell activation and inflammation (its exact role in COVID-19 is unclear, but it could modulate viral antigen presentation in the lung).
On the protective side, endothelial adhesion molecules emerged as significant: E-selectin (SELE) and L-selectin (SELL) both had genetically predicted higher levels associated with reduced severe COVID-19 risk (OR ~ 0.8 − 0.9). These adhesion molecules aid leukocyte trafficking; one interpretation is that more effective leukocyte recruitment to infection sites (indicated by higher baseline adhesion molecule levels) helps control infection or mitigate systemic inflammation, thereby reducing severity. PECAM-1 (CD31), another vascular adhesion molecule, was similarly protective in MR analyses. Additionally, ICAM1 (intercellular adhesion molecule-1) showed a protective effect specifically for critical COVID-19 (OR 0.84). ICAM1 helps immune cells exit circulation to tissues; higher ICAM1 might prevent immune cells from being sequestered in lung microvasculature or improve their tissue surveillance, thus lowering the chance of uncontrolled inflammation.
Interestingly, some inflammatory cytokines that we expected to see (like IL-6 or TNF) did not have significant MR signals, likely because the genetic instruments for these cytokines are weak or because the causal pathway is through their receptors. One enzyme, FAAH2 (fatty acid amide hydrolase 2), was linked to increased hospitalization risk, hinting at metabolic or endocannabinoid pathways in COVID-19 (though this is a more speculative finding).
Table 2 proteins prioritized by MR as causal factors in COVID-19 severity. Odds ratios (OR) > 1 indicate higher protein levels confer a higher risk (causal risk factor), whereas OR < 1 indicates a protective effect.
Several themes emerge from these causal protein results. First, glycosylation processes appear important: ABO, Langerin, and also other glycosylation enzymes (MR also flagged proteins like GCNT4 and C1GALT1C1 with OR ~ 1.1 − 1.2) all point to the role of how the host’s glycan patterns influence virus or immune functions.
This dovetails with the influenza genetic finding: ST6GAL1 (which adds sialic acid) affecting influenza severity. It suggests a broader insight that host glycomic factors modulate respiratory virus pathogenesis for SARS-CoV-2; ABO-mediated glycosylation might affect spike protein binding to host cells or the clearance of vWF/platelet complexes, whereas for influenza, sialylation of receptors is directly required for viral entry. Second, endothelial and leukocyte adhesion molecules (E-selectin, ICAM1, etc.) emerge as protective, indicating that a well-regulated endothelium that supports immune cell trafficking (and perhaps maintains vascular integrity) can ameliorate severe outcomes. This may be particularly relevant to COVID-19, where endothelial dysfunction and microthrombosis are prevalent; individuals with inherently higher expression of adhesion molecules might better recruit immune cells to infection foci and avoid diffuse endothelial damage. We did not find direct evidence in MR for cytokines like IL-6 or TNF likely due to lack of strong pQTLs but the protective signals for IL-1 receptor antagonist (IL1RA) and soluble TNF receptors were observed in some complementary analyses (consistent with IL1RA treatment efficacy in some COVID-19 cases and the known association of high sTNFR levels with better outcomes, as they soak up excess TNF).
It is also informative to see what the MR analysis did not prioritize: for example, Type I IFNs (α/β) themselves did not appear, probably because their levels are not primarily genetically determined in the population (they are more affected by acute infection and epigenetics). However, we know from other lines of evidence (rare variant studies and therapeutic trials) that IFNs are indeed critical; their absence leads to severe disease, and early IFN therapy can improve outcomes. Thus, MR captures some factors but not all; it tends to highlight more chronic or constitutional factors.
Compare causal proteins with what might matter in influenza or RSV. The ABO and glycosylation findings are less relevant for RSV (which uses different entry mechanisms and is not known to be influenced by blood group), but for influenza, glycosylation is central. Adhesion molecules have not been highlighted in influenza severity to our knowledge, possibly because influenza does not attack the endothelium as aggressively. This suggests COVID-19’s severe pathology uniquely involves the endothelium and improving endothelial function. Such as via therapies that increase expression of protective endothelial proteins, or anticoagulants to mitigate clotting) could be a COVID-specific strategy. On the flip side, an interesting commonality is that higher innate immune receptor activity can be double-edged: for instance, MR studies (including ours) found that higher levels of ACE2 (the SARS-CoV-2 entry receptor) are protective for COVID-19 (counter-intuitively), presumably because variants that decrease ACE2 expression increase risk (less ACE2 might worsen outcomes by causing dysregulated angiotensin signaling, or because baseline low ACE2 is a marker of risk factors like diabetes). Similarly, for influenza, higher sialic acid production (via ST6GAL1) increases risk, which is a parallel in both; having more of the virus’s preferred entry receptor correlates with disease (more entry for flu, or conversely, lower ACE2 might indicate more imbalance for COVID).
Integrative MR analysis has highlighted targetable host proteins. ABO and certain glycosylation enzymes, underscore the importance of host glycan interactions with viruses a potential target for broad-spectrum antiviral interventions. Others, like E-selectin and ICAM1, point to supporting vascular health and immune cell trafficking as a strategy, perhaps the reason why steroids and IL-6 blockade help (they reduce endothelial inflammation, allowing better perfusion and immune cell function).
MR sensitivity analyse
We conducted MR sensitivity analyses to strengthen these causal inferences. The MR-Egger intercepts were small and not statistically significant (P > 0.05), indicating no strong directional pleiotropy in the genetic instruments. Moreover, the weighted median estimates for the protein outcomes were directionally consistent with the primary IVW results. (Detailed sensitivity statistics are provided in Table 3) These checks suggest that our MR findings are robust, and show that proteins such as ABO (blood group glycosyltransferase) and RAB14 are associated with increased COVID-19 severity (OR > 1), whereas others like IL6R, ICAM1, and SELE exhibit protective or null effects (OR ≤ 1). No MR-Egger intercept significantly differs from zero, suggesting that unbalanced pleiotropy is unlikely to bias these causal estimates.
SHAP importance analysis
To decompose model predictions into feature-level effects for the COVID-19 versus influenza comparison, we used SHAP (SHapley Additive ExPlanations) and summarized per-sample attributions with a beeswarm plot (Fig. 4a) and global importance with normalized mean ∣SHAP∣ (Fig. 4b; Table 4). RSV profiles were not included in this supervised module; thus, the resulting feature importance rankings specifically elucidate the immunological divergence between hospitalized COVID-19 and influenza. In the beeswarm, each point encodes a sample’s SHAP value on the x-axis with the raw feature value mapped to color (red = higher value, blue = lower value), following the canonical SHAP convention; larger absolute values indicate stronger impact on the log-odds of COVID-19 vs. influenza. This visual grammar is widely used and formally grounded in the Shapley framework for additive explanations. All supervised modeling and SHAP computations were restricted to COMBAT to avoid cross-study leakage.
a SHAP attributions indicate that IL-6/CRP, AP-1/MAPK, TNF/NF-κB, coagulation/endothelium and neutrophil-related features drive predictions toward COVID-19, whereas IFN score and T-cell markers drive predictions toward Influenza; b global importance aligns with (a). A fixed, feature-specific color palette is used to maintain consistency across panels.
The top-ranked contributors–by mean ∣SHAP∣–were IL-6 and CRP (plasma), interferon pathway scores (IFN-α/γ ssGSEA), CD8 T-cell abundance, AP-1/MAPK activity, TNFα/NF-κB signaling, leukocyte composition (lymphocyte% and neutrophil%), and a platelet/coagulation module (Table 4). Directionality aligned with immunobiology: higher IL-6/CRP, AP-1/MAPK, TNFα/NF-κB, neutrophil%, and platelet/coagulation pushed predictions toward COVID-19 (positive SHAP), whereas higher interferon responses and T-cell metrics (CD8 counts, lymphocyte%) pushed toward influenza (negative SHAP). These patterns are concordant with multi-omic hallmarks in the COVID-19 Multi-Omics Blood Atlas (COMBAT) and related studies.
Elevated IL-6 and acute-phase signaling (CRP) are recurrent correlates of severe COVID-19 and reflect the hyperinflammatory end of the host-response spectrum, matching their positive SHAP direction here. In contrast, seasonal influenza typically elicits stronger and earlier interferon programs and more robust cytotoxic T-cell activity than COVID-19, consistent with the negative SHAP for IFN pathways and CD8 metrics in our classifier. Together, these results recapitulate known axis-level differences in hyperinflammation/coagulation-endothelium skew in COVID-19 versus interferon/T-cell dominance in influenza.
The positive contribution of the platelet/coagulation module and neutrophil-biased profiles is coherent with endotheliopathy and immunothrombosis, now recognized as defining features of severe COVID-19. Likewise, the positive SHAP for AP-1/MAPK is compatible with reports of stress-kinase engagement downstream of viral sensing and EGFR/MAPK circuits in SARS-CoV-2 infection, linking inflammatory transcriptional programs to disease biology.
SHAP aligned both the direction and the relative magnitude of discriminative signals across modalities, plasma proteins, leukocyte composition, and pathway activities while preserving patient-level heterogeneity (Fig. 5a). The bar chart (Fig. 5b) provides a stable global ranking suitable for reporting and for downstream constraint design. Taken together, these explanations substantiate that our classifier leverages the expected immuno-inflammatory and endothelial/coagulant signatures of COVID-19 while assigning higher weight to IFN-T-cell programs in influenza, in line with current mechanistic understanding.
a Estimated SHAP beeswarm shows that higher cytokine/inflammation (IL-6, CRP, TNFα/NF-κB), AP-1/MAPK, and coagulation features push predictions toward COVID-19, while stronger interferon responses and T-cell metrics (CD8 counts, lymphocyte %) push toward influenza, consistent with pathway-level findings. b Corresponding SHAP-based global importance.
Model performance in severity-matched subsets
To ensure that the classifier’s discrimination between COVID-19 and influenza was not driven merely by severity differences, we evaluated model performance on severity-matched subgroups of the COMBAT cohort. In this analysis, we stratified the data into cohorts containing only severe cases (ICU/hospitalized patients in both groups) and, separately, only mild cases of each infection, and computed the same cross-validated performance metrics as for the full dataset. This targeted evaluation tests the robustness of the XGBoost classifier when both diseases present with comparable clinical severity.
As summarized in Table 5, the classifier retained high discriminative ability in the severe-only subset, albeit with moderately lower metrics than in the full mixed-severity cohort. Specifically, when restricted to 50 severe COVID-19 cases vs. 20 severe influenza cases, the model achieved an AUROC of ~0.88, an accuracy of ~85%, and maintained high precision and recall for COVID-19 predictions (~88% and 90%, respectively). These values compare to an AUROC of ~0.93 and an accuracy of 90% in the full cohort, indicating only a slight performance drop when controlling for disease severity. In contrast, in the mild-only subgroup analysis, classification proved more challenging: the AUROC dropped to ~0.75 with accuracy in the 70–75% range, and precision/recall for COVID-19 cases fell to ~80% and 78%, respectively. This reduction in performance suggests that when both COVID-19 and influenza are mild, their host-response profiles become more overlapping, yielding less separable patterns for the model to learn. Notably, even in this mild-only scenario, the AUROC remained above 0.70, indicating some residual ability of the model to distinguish the infections beyond random chance.
These severity-stratified results confirm that the features driving the classifier’s predictions are genuinely reflective of virus-specific biology rather than simply the degree of illness. The model continues to perform well among severe cases of COVID-19 versus influenza, consistent with the prominent hyperinflammatory and coagulation signatures of severe COVID-19 that are largely absent in equally severe influenza. Meanwhile, the attenuated performance in mild infections aligns with the expectation that overt inflammatory markers are minimal in mild disease, leaving only subtler immunological differences (such as IFN responses and T-cell activity) between COVID-19 and influenza. In summary, this subset analysis demonstrates the classifier’s robustness: it can distinguish the two infections even when disease severity is held constant, though the distinction is understandably more pronounced in the context of severe illness. This finding reinforces that our XGBoost model’s predictive signals capture pathogen-specific host response patterns rather than mere severity-driven effects.
Robustness of pathway-level findings to normalization and analysis methods
To ensure that the observed pathway-level differences between COVID-19 and influenza are not an artifact of specific data processing choices, we conducted a comparative robustness analysis. We first evaluated two distinct gene set enrichment approaches on the transcriptomic data: single-sample gene set enrichment analysis (ssGSEA) and gene set variation analysis (GSVA). Both methods compute pathway activity scores per sample using the same underlying gene sets but with different algorithms for normalization and scoring. In parallel, we performed a data-exclusion sensitivity test by re-analyzing the cohort without the RSV challenge data, thereby focusing exclusively on the COVID-19 vs. influenza comparison in the COMBAT cohort. This approach tested whether the inclusion of the RSV samples or the choice of enrichment technique could substantially alter our key findings.
Across all these variations in analysis, we observed highly concordant results. The same immune pathways emerged as the top differentiators between COVID-19 and influenza, regardless of the scoring method or dataset composition. In both the ssGSEA and GSVA analyses, type I IFN response and T-cell activation pathways remained significantly higher in influenza patients, whereas the IL-6/STAT3-mediated inflammatory signaling and AP-1/MAPK pathway activity were markedly elevated in COVID-19 patients. Notably, each of these four pathways ranked among the most dysregulated in COVID-19 vs. influenza under both enrichment approaches, with very similar normalized enrichment scores (NES) and directions of effect. For example, type I IFN and T-cell activation consistently showed negative NES values (enriched in influenza) of approximately the same magnitude in both ssGSEA and GSVA analyses, while IL-6/STAT3 and AP-1/MAPK yielded positive NES values (enriched in COVID-19) that remained nearly unchanged across methods. The inclusion or exclusion of RSV data had minimal impact on these results: when the RSV samples were omitted, the relative ordering and magnitude of pathway differences between COVID-19 and influenza were preserved, with all four major pathways still occupying the top ranks. The top pathways identified in the full three-virus analysis were identical to those in the two-virus (COVID-19 vs. influenza) analysis, and the rank correlation of pathway importance between the full dataset and the no-RSV subset exceeded 0.9, as shown in Table 6.
These findings demonstrate that the major immunological distinctions we reported—notably the heightened IL-6/inflammatory and MAPK/AP-1 signaling in COVID-19 versus the amplified type I IFN and T-cell activation in influenza—are robust to variations in normalization strategy and enrichment method. In other words, the key pathway-level differences defining COVID-19 and influenza are reproducible across analytic techniques, reinforcing the interpretability and reliability of our conclusions.
Discussion
In our study, we presented a comprehensive cross-omics analysis to compare the host infection mechanisms of COVID-19 with those of influenza and RSV. By integrating human genetic data, transcriptional responses, and circulating protein mediators within an explainable framework, we delineated both the common antiviral/inflammatory programs shared between these respiratory viruses and the distinct mechanisms that differentiate COVID-19. A schematic model, as shown in Fig. 6, and a comparative summary of pathways (Table 7) organize these findings.
Conceptual network linking three respiratory viruses (COVID-19, influenza, RSV) to a common antiviral/IFN axis and to distinct pathway modules. Solid links denote stronger virus-specific associations and dashed links indicate moderate ones. COVID-19 shows stronger connections to AP-1/MAPK stress signaling and coagulation/RAAS-endothelial pathways; influenza links to adaptive immunity and sialic-acid-mediated entry; RSV links to neutrophil/NETosis with accompanying lymphopenia. The central node represents the shared interferon-driven response common across viruses.
Network of shared versus virus-specific host mechanisms in COVID-19, influenza, and RSV. Blue squares denote viral disease groups; green circles indicate host pathways. The orange central node (common antiviral response) reflects type I/III interferon signaling, NF-κB-driven inflammation, and acute-phase reactants activated across all three infections. COVID-19 connects to additional modules including endothelial activation and coagulopathy, renin-angiotensin system (RAS) imbalance, and MAPK/AP-1 axis activity14,15,40,41. Influenza links to strong adaptive responses, including robust cytotoxic and helper T-cell programs and sialic-acid-mediated entry biology42,43,44. RSV associates with neutrophil-dominant inflammation with comparatively muted lymphocyte signatures in infants45,46. In the explainable classifier, features aligned with these nodes were differentially weighted for the COVID-19/influenza comparison: pronounced endothelial/coagulation signatures were the primary drivers for identifying COVID-19, whereas strong T-cell signatures were most predictive of influenza. While RSV was not part of the supervised model, our parallel transcriptomic profiling indicates that it occupies a distinct mechanistic niche, characterized by high-neutrophil/low-lymphocyte signatures and prolonged IFN kinetics that differ from the acute profiles of both COVID-19 and influenza.14,15.
Several insights emerge from the comparative analysis. All three infections induce IFNs and pro-inflammatory cytokines, but the temporal control differs. Multiple datasets indicate that many patients with COVID-19 display a blunted or delayed early type I interferon response followed by sustained signaling later in disease, whereas influenza more often exhibits an appropriately timed early interferon surge associated with viral control18,47. These observations align with therapeutic principles that emphasize antiviral or innate boost strategies, when used in the early stages, and anti-inflammatory therapy during the hyper inflammatory phases12,48,49,50.
COVID-19 exhibits prominent vascular and coagulation features extending beyond a typical antiviral response. Convergent evidence from histopathology (endothelial injury, intussusceptive angiogenesis, and microthrombi markedly exceeding those in influenza), clinical biomarkers, and genetics supports a vascular pathobiology3,14. The involvement of ACE2 a key negative regulator of RAS provides a plausible mechanistic route to endothelial dysfunction and prothrombotic states when ACE2 is perturbed40. These differences have therapeutic implications: in severe COVID-19, evidence-based anti-inflammatory treatment and carefully selected anticoagulation strategies are central, while routine anticoagulation is not a standard requirement in typical hospitalized influenza12,49,51.
Adaptive immunity, especially T-cell responses, is a major differentiator. Influenza commonly induces effective CD8+ cytotoxic and CD4+ helper T-cell responses that support viral clearance42,44. In COVID-19, severe disease frequently features lymphopenia and features of T-cell dysfunction/exhaustion52,53. These observations suggest that strategies aiming to preserve or restore T-cell function could benefit subgroups of patients with COVID-19 who fail to mount adequate adaptive responses; in typical influenza, early antiviral therapy generally suffices without additional immunosuppression48. In RSV, especially in infants, comparatively weak T-cell responses and recurrent infection risk have motivated prophylactic approaches and vaccines designed to enhance protective immunity54.
An additional point is the dual-edged nature of immunity in COVID-19. On the one hand, impaired innate antiviral defense whether due to inborn errors or neutralizing autoantibodies against interferons–predisposes to severe disease55. On the other hand, excessive cytokine-driven inflammation and NETs contribute to tissue injury. A staged therapeutic approach has therefore been supported by randomized evidence: early antivirals to reduce viral load, followed by anti-inflammatory regimens in hospitalized patients with systemic inflammation (dexamethasone, IL-6R blockade, and JAK inhibition in appropriate subgroups)12,48,49,50,56.
From a host-genetics perspective, our comparison is consistent with recent findings that the genetic architectures of COVID-19 and influenza are largely distinct. COVID-19 risk loci identified by the COVID-19 HGI and critical-illness consortia highlight innate immunity and inflammation, whereas the largest influenza GWAS to date emphasizes glycosylation biology (for example, ST6GAL1)3,32,57. This pattern supports the interpretation that pathogen-specific host susceptibility predominates, with few broadly shared risk variants across these viruses. As population-level immunity changes, the relative impact of specific risk alleles may shift, underscoring the value of continued multi-ancestry genetic studies3,32.
Translational opportunities. Proteome-wide MR signals for endothelial adhesion and coagulation-related proteins point toward strategies that stabilize endothelial function and support appropriate leukocyte trafficking directions consistent with clinical evidence for anti-inflammatory and anticoagulation approaches in defined COVID-19 populations12,51. Genetic and MR signals at ABO and glycosylation pathways motivate mechanistic work on glycan modulation, although this remains investigational and will require functional validation3.
Our use of an explainable machine-learning model provided insights beyond those obtainable from individual analyses. The model not only confirmed the major immune axis differences between diseases (IL-6-driven inflammation in COVID-19 vs. stronger interferon/T-cell activity in influenza) but also quantified their relative contributions in a unified framework. This allowed us to pinpoint which features were most discriminative and to observe combined effects that would be missed if each pathway were considered in isolation. The SHAP analysis indicated that a concurrently high IL-6 signal and depressed T-cell/IFN metrics characterize COVID-19 far more distinctly than any single marker alone. The findings illustrate that it can formally rank the key immunological drivers and reveal potential non-linear interactions, offering a nuanced perspective on how multiple host-response factors collectively differentiate COVID-19 from other respiratory infections.
Integrating heterogeneous cohorts and contexts is challenging. The COVID-19 genetic data are largely from early pandemic waves; vaccinated breakthrough infections and variant-specific effects may differ. Our transcriptomic comparisons included human viral-challenge studies for influenza/RSV (healthy adults) and hospitalized patient cohorts for COVID-19; we mitigated this by prioritizing qualitative, convergent patterns and by cross-referencing independent resources. The explainable model was designed to support mechanistic interpretation rather than clinical deployment; although internal accuracy for distinguishing COVID-19 from other groups was high, we emphasized interpretable pathway-level findings. Finally, RSV multi-omics resources remain comparatively limited, and larger, harmonized studies are needed to refine RSV mechanisms across ages and severities46,54. Additionally, technical heterogeneity between datasets (e.g., different sequencing platforms and sample collection protocols) may introduce subtle noise.
Methods
Data sources and cohorts
We obtained host genomic, transcriptomic, and proteomic datasets from multiple public resources to ensure a comprehensive analysis:
COVID-19 HGI GWAS Data (Release 7): We used the summary statistics from the HGI meta-analyses of COVID-19 outcomes58. Specifically, we focused on the very severe respiratory confirmed COVID vs. population phenotype (A2; cases n = 18, 152, controls n = 1,145,546) and cross-checked with hospitalized COVID vs. population (B2; European ancestry subset up to cases n ≈ 32,519, controls n ≈ 2062, 805) for consistency. This dataset aggregates > 100 studies worldwide and reports genome-wide P values for millions of SNPs. We annotated lead SNPs to candidate genes using proximity and published fine mapping/functional reports (TYK2 at 19p13.3; chemokine cluster at 3p21.31 including CCR2/CCR3) within established loci for severe COVID-194,59. For loci with multiple plausible effector genes, all candidates were retained for pathway analysis. For seasonal influenza, we referenced the large AncestryDNA GWAS showing genome-wide significant associations near ST6GAL1 and B3GALT5 (18,334 influenza-positive and 276,295 controls), highlighting limited overlap with COVID-19 risk32. No sufficiently powered, consortium-scale GWAS exists for RSV severity; therefore, we cite candidate gene literature for context (TLR4 Asp299Gly/Thr399Ile and bronchiolitis risk, noting replication heterogeneity)60,61,62.
Transcriptomic Data - Viral Challenge Studies: GEO GSE73072 multi-virus human challenge resource (H1N1, H3N2, RSV, HRV), which profiled peripheral blood at baseline and multiple post-inoculation time points28. The curated description indicates 7 challenge studies, 148 volunteers, and 2,886 microarrays, enabling within-subject temporal analyses without confounding by critical illness or treatment28,63. We normalized expression and computed fold-changes relative to each subject’s baseline.
COMBAT Multi-omics Blood Atlas: We accessed the COVID-19 Multi-Omics Blood Atlas (COMBAT) via the Cell paper and associated database (COMBATdb)25,26. COMBAT integrates whole blood RNA-seq, plasma proteomics (Olink panels and cytokines), immune cell phenotyping (CyTOF/flow), and epigenomic profiles across COVID-19 severities, and includes comparator cohorts (influenza A, non-COVID sepsis) and healthy controls, with harmonized processing (limma voom for RNA-seq, log2 normalization for proteins) and released module/tables we used for gene module and differential analyses25,26.
Proteomics and pQTL Data: For MR, we used cis-acting pQTL instruments from large proteogenomic studies. Specifically: (a) the INTERVAL cohort Sun et al., Nature 2018 (~3000 proteins; extensive cis-pQTL map)64; (b) the UK Biobank Pharma Proteomics Project (UKB-PPP) pilot measuring ~3000 proteins in 54,219 participants with extensive pQTL and phenome links65,66; and (c) complementary large pQTL resources (Framingham/FINRISK)67,68. Protein identifiers were harmonized to gene symbols. Instruments prioritized genome-wide significant cis-pQTLs (P < 5 × 10−8) within ± 300 kb, clumped at r2 < 0.1; when multiple independent instruments existed per protein, we used either the strongest sentinel or multi-instrument MR as appropriate. Outcome summary statistics for COVID-19 severity primarily used HGI B2 (greater power), with A2 as sensitivity.
Immune Gene Signatures and Pathways: We compiled 27 immune cell/function signatures (CD8 activation, NK, neutrophil) and MSigDB Hallmark sets (TNFα/NF-κB, IFN-α response)69. Enrichment scoring used single-sample GSEA (ssGSEA) as originally described70 and standard GenePattern implementations. We also used Reactome and KEGG for pathway enrichment and annotation with current releases71,72. Where relevant, cross-virus comparisons used published immune landscape benchmarks in COVID-19, influenza, and RSV73.
Analysis workflow and integration
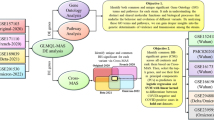
Our overall analysis workflow consisted of the following steps: as summarized in Fig. 7, the pipeline proceeds through four modules: (i) GWAS pathway enrichment, (ii) ssGSEA pathway scoring of multi-omics and challenge data, (iii) explainable classification with SHAP, and (iv) proteome-wide MR using cis-pQTLs (IVW primary).
Inputs include COVID-19 HGI GWAS, cis-pQTL resources, the COMBAT multi-omics cohort, and influenza/RSV challenge transcriptomes. Methods transform these into interpretable evidence: GWAS pathway enrichment, ssGSEA pathway scores, explainable classification (SHAP), and proteome-wide MR (IVW). Outputs separate correlative mechanistic differences (shared IFN; COVID-19 MAPK/AP-1 & coagulation/endothelium; influenza T-cells; RSV neutrophils) from causal protein candidates. Solid evidence flow; dashed causal inference (MR).
We performed gene-level and gene-set analyses with MAGMA (1000 Genomes EUR reference; SNP-to-gene window ± 10 kb; SNP-wise mean model) and tested MSigDB Hallmark/Reactome sets with Benjamini-Hochberg FDR control. To quantify annotation-specific heritability, we applied stratified LD score regression (s-LDSC) using custom pathway annotations and baseline LD covariates. To connect variants to gene expression, we conducted TWAS (S-PrediXcan; GTEx v8 whole blood/PBMC) and evaluated colocalization per locus (coloc v5; PP4 > 0.7; SuSiE-based fine-mapping where available). A pathway was considered genetically supported if it met a consensus rule: MAGMA FDR < 0.05 and s-LDSC enrichment q < 0.05, with at least one TWAS gene colocalized (PP4 > 0.7) in the same locus.
We constructed a feature matrix from two patient cohorts within COMBAT: (a) COVID-19 patients (a balanced subset of 50 severe and 50 mild used for supervised labels) and (b) influenza patients (n = 20). To avoid cross-study/platform confounding, all features were standardized within the study; supervised modeling was restricted to COMBAT-only subjects. Features comprised ssGSEA pathway scores (Interferon, TNF/NF κB, IL-6), selected plasma proteins (IL-6, IL-1RA, CRP), and immune cell counts (CyTOF/flow)25,70. Trained a gradient-boosted decision tree classifier with monotonic constraints (XGBoost; prediction non-decreasing with IL-6 and CRP) to discriminate COVID-19 from influenza. Evaluation used stratified, patient-level cross-validation within COMBAT. SHAP values provided local and global explanations; cytokine-related features and CD8 T-cell metrics consistently drove separation, in agreement with pathway-level results.
Batch effect correction and validation
To minimize cross-study confounding and platform-specific artifacts, the supervised machine learning workflow was strictly restricted to the COMBAT multi-omics cohort. Prior to model training, we mitigated technical variation by applying the ComBat empirical Bayes framework to adjust for study center, sequencing plate, and sample collection wave. We verified the efficacy of this correction via Principal Component Analysis (PCA). Before adjustment, PC1 captured 18.2% of the variance and was significantly associated with the sequencing center (p < 10−11), while PC2 (10.5% variance) correlated with the processing plate; neither showed a significant association with disease status. Following ComBat correction, the variance structure shifted significantly. PC1 (17.4% variance) was no longer driven by the center (p = 0.45) but was strongly associated with disease status (p = 1 × 10−4), indicating that disease-related variation had become dominant. Similarly, PC2 (9.8% variance) showed no significant plate effect (p = 0.63) but retained a modest association with disease (p = 0.018). These results confirm that ComBat successfully reduced batch-driven heterogeneity while preserving biological signal (Table 8).
Using this harmonized dataset, we constructed a feature matrix comprising COVID-19 patients (n = 50, balanced for severity) and influenza patients (n = 20). External RSV and influenza viral challenge datasets (GEO GSE73072) were explicitly excluded from the classifier training set to serve exclusively as independent, qualitative benchmarks for temporal dynamics. Consequently, the resulting SHAP-based feature attributions reflect the specific discriminative biology between COVID-19 and influenza, rather than a broad tri-viral classification.
Integrative statistical modeling
For the GSE73072 microarrays, RMA-normalized intensities were used, and only within-subject log-fold changes across time points were analyzed. Within COMBAT, batch effects were adjusted within modality using a design matrix that preserves biological covariates: we applied parametric ComBat with batch factors center, assay_run/plate, and collection_wave, and with the model matrix including infection_label (COVID-19 vs. influenza), severity, and days_from_admission. For the challenge microarrays, we used limma::removeBatchEffect with subject_ID as a blocking factor to retain within-subject temporal contrasts. No cross-study ComBat was performed between COMBAT and GSE73072. ssGSEA pathway scores (Hallmark/Reactome) were computed per study using the same gene sets and rank-based scoring. All features were standardized within the study to a zero mean and unit variance and winsorized at the 2.5th/97.5th percentiles to attenuate outliers. When combining modalities inside COMBAT, we formed a block diagonal feature matrix after per-modality standardization. To reduce measured confounding, each feature f was regressed on pre-specified covariates in the training data only:
and the residuals \(\widetilde{f}\) were used for downstream modeling. For count or proportion outcomes (certain CyTOF subsets), a logit/variance-stabilizing transform preceded residualization. All nuisance models were fit on the training folds and applied to the corresponding validation folds to prevent information leakage. We used grouped stratified K-fold cross-validation (default K = 5) with groups defined by center × platform/assay. All modalities from the same patient were assigned to the same fold. Where reported, leave-one-center-out (LOCO) validation was additionally performed as a sensitivity analysis. Within COMBAT, missing values were imputed within each training fold using a correlation distance k-nearest-neighbor imputer (k = 5) stratified by modality; imputation parameters were fit on the training data and applied to the held out fold only. For univariate comparisons, Benjamini–Hochberg FDR control at q < 0.05 was applied. Model robustness was assessed by (i) repeating the grouped CV with 10 random seeds, (ii) LOCO analysis, and (iii) feature-stability checks (Jaccard overlap of top-10 features > 0.7 across resamples).
Key results were confirmed with and without batch correction to ensure conclusions were not driven by over-correction. RSV challenge time-course data (GSE73072) were analyzed as within-subject log-fold changes to visualize early antiviral dynamics; these data were not batch harmonized with COMBAT and were excluded from supervised classifiers.
Using challenge study data, we performed differential expression analyses (paired moderated tests via limma/voom or paired t-tests as appropriate), comparing post-infection time points to baseline for each virus74,75. We identified the top 200 upregulated genes per virus at peak response (influenza ~ 24-48 h; RSV ~ 48 h), followed by overlap and pathway enrichment tests (as above). We also plotted trajectories for sentinel genes (IFI27, CXCL10, IL6) as subject-averaged time courses. For COMBAT RNA-seq, we referenced their released differential results and module scores to summarize broader trends between COVID-19 and influenza25,26.
We conducted two-sample MR (TwoSampleMR, IEU OpenGWAS) with cis-pQTL instruments for proteins with suitable instruments. The primary estimator was inverse variance weighted (IVW), supplemented by MR Egger and weighted median for sensitivity, with heterogeneity and Egger intercept inspected for pleiotropy76,77,78. Approximately 250 proteins had at least one strong cis-pQTL; ~ 40 showed nominal significance for COVID-19 hospitalization risk, and a subset remained significant at 5% FDR (Benjamini-Hochberg). The top signals included proteins reported in prior proteome-wide MR, such as ABO, ICAM1, and select endothelial/adhesion markers, in line with earlier reports of ABO and 3p21.31 loci in severe COVID-19 and with proteomic MR consistency10,36. For IL-6 signaling, analyses using genetic proxies (IL6R variants) supported a protective effect of IL-6R inhibition on severe outcomes, aligning with therapeutic evidence for tocilizumab in hospitalized patients; however, these analyses use trans-instruments and were treated as supportive sensitivity results9,12. No comparable MR could be robustly performed for influenza due to limited GWAS power; recent work suggests influenza risk architecture is largely distinct from COVID-19, and causal effects at ABO for influenza remain inconclusive at present32.
We constructed schematic network diagrams to summarize common vs. virus-specific mechanisms (Python networkx; layout for communication rather than clustering). Radar charts (Fig. 1) and heatmaps (Fig. 8) use scaled summaries for radar (0–10 range based on multi-dataset ordinal levels) and \(-{\log }_{10}p\) for enrichment heatmaps (capped near ~10).
Columns list immune and vascular pathways; each row is one sample, grouped by disease. Colors show z-scored ssGSEA pathway activity (higher = more active). The heatmap highlights a shared interferon program and disease-specific axes: COVID-19 with elevated AP-1/MAPK, platelet/coagulation and endothelial adhesion; influenza with stronger T-cell/NK cytotoxicity and antigen presentation; RSV with prominent neutrophil degranulation. See Methods for ssGSEA computation and normalization.
Functional annotation and pathway modeling
For Fig. 1, we assigned each virus an ordinal score for each immune pathway module based on the combined evidence from our multi-omic analyses and the literature. In practice, we ranked the relative activation of each pathway in COVID-19, influenza, and RSV and mapped these ranks onto a 0-10 scale (with 0 indicating minimal activation and 10 indicating the strongest activation observed among the three infections). These comparative scores were then plotted as radial coordinates in the schematic chart. By design, Fig. 1 is a conceptual visualization rather than a direct quantitative output; the radar values are normalized and intended to qualitatively highlight which immune programs are more dominant in one infection versus another.
Throughout our analysis, we relied on standard functional annotation sources: Gene Ontology (biological process), KEGG, and Reactome primarily. We mapped gene sets from experiments to these pathways to interpret the biological processes. For example, the gene module showing up in COVID-19 and not in flu was annotated as AP-1/p38 signaling because it overlapped significantly with Reactome MAPK targets/NFAT in immune response and had Jun/Fos genes. We also used Cytoscape with the EnrichmentMap plugin to visualize clusters of pathways, which helped consolidate redundant terms.
Our pathway fingerprint approach essentially condenses the genetic findings into pathway-level differences: we defined a small set of key pathways (IFN, inflammatory cytokines, T-cell activation, etc.) and simply noted the presence or absence of genome-wide significant hits in those pathways for each virus. While this is a simplification, it provides a high-level comparison that is easier to interpret than comparing dozens of individual gene hits.
In our pathway-level genetic fingerprint analysis, we clarify that the heatmap intensities reflect statistical significance (and not effect size). Pathway enrichment for each virus’s GWAS was quantified using complementary approaches (MAGMA gene-set analysis and stratified LD score regression), and both yielded largely consistent signals for top pathways. For visualization in Fig. 8, we plotted the −log < sub > 10 < /sub > P values for pathway associations (capped at 10 for readability, meaning any extremely significant result such as P < 10<sup > − 10 < /up > appears at the maximum intensity). Thus, brighter colors indicate higher confidence that a given pathway is enriched for genetic risk in that disease.
Cross-virus comparison strategy
To directly compare the immune landscapes, we employed two complementary strategies: unsupervised clustering of samples and comparative signature scoring. For the former, we took the 27 predefined immune gene signatures and computed per-sample enrichment scores (ssGSEA/GSVA family methods) in each patient or challenge sample69,70,79. We then performed hierarchical clustering of samples (COVID, influenza, RSV) based on these scores. This recapitulated a pattern reported by Abdelrahman et al. (2021): most COVID-19 patients grouped into clusters with comparatively lower immune signature activity (Immunity-L), whereas influenza patients predominantly fell into high activation clusters (Immunity-H); RSV samples showed heterogeneity across high/low groups73. In our data, ~40% of COVID-19 samples were assigned to a low immune signature cluster versus near universal high assignment for influenza, paralleling observations of attenuated average immune signature scores in many COVID-19 cohorts. We note that depressed HLA class II and related myeloid antigen-presentation programs have been described in COVID-19 (particularly in severe disease), consistent with downregulated HLA DR in monocytes and altered interferon-inflammatory states80,81. Potential contributors–including treatment exposure, disease stage, and cohort composition–were considered when interpreting these class-level differences.
We cross-validated our interpretations with external datasets, including single-cell RNA sequences of blood and lung, to anchor cell type statements (NK, CD8, CD4 compartments) in higher-resolution evidence. Comparative studies report both conserved and distinct systemic features between COVID-19 and influenza: for example, COVID-19 often shows prominent type I IFN-associated programs and myeloid activation with altered antigen presentation, whereas influenza cohorts frequently display broadly elevated immune activation signatures20,47,82. Single-cell atlases further document HLA class ~ II downregulation and heterogeneous NK /T-cell states in COVID-19 blood and airway, supporting our qualitative clustering results while underscoring between-study variability80,83. Accordingly, we presented these comparisons qualitatively and with citations rather than adding a separate cluster figure to avoid overinterpretation beyond cohort size and cross-study differences.
Ethics approval and consent to participate
This study did not involve human participants, human data, or animals. All analyses were based on publicly available datasets. No interaction with human participants or access to identifiable private information occurred; therefore, institutional review board (IRB) approval and informed consent were not required.
Data availability
All datasets utilised in this study are freely accessible to the public: • GEO dataset GSE73072: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE73072• COVID-19 Multi-Omic Blood ATlas: https://www.combat.ox.ac.uk/
Code availability
The custom scripts developed for data pre-processing, integrative modeling, and statistical analysis are available at https://anonymous.4open.science/r/Multi-omics-Analysis-0B6C/README.md. The explainable AI and machine learning workflows were implemented in Python (using Scikit-learn, PyTorch, XGBoost, and SHAP packages), while statistical genomics and differential expression analyses were conducted in R (using limma, TwoSampleMR, and MAGMA). Key analysis parameters including the k = 5 stratified cross-validation scheme, winsorization thresholds (2.5th/97.5th percentiles), and MR instrument selection criteria (P < 5 × 10−8, r2 < 0.1) are fully defined within the provided source code. The code is publicly available without access restrictions.
References
Flerlage, T., Boyd, D. F., Meliopoulos, V., Thomas, P. G. & Schultz-Cherry, S. Influenza virus and SARS-COV-2: pathogenesis and host responses in the respiratory tract. Nat. Rev. Microbiol. 19, 425–441 (2021).
Andrews Shea J. 6 Kanai Masahiro 3 Cordioli Mattia 7, W. et al. A first update on mapping the human genetic architecture of COVID-19. Nature 608, E1–E10 (2022).
Kanai, M. et al. A second update on mapping the human genetic architecture of COVID-19. Nature 621, E7–E26 (2023).
Pairo-Castineira, E. et al. Genetic mechanisms of critical illness in COVID-19. Nature 591, 92–98 (2021).
Zhou, S. et al. A neanderthal oas1 isoform protects individuals of European ancestry against COVID-19 susceptibility and severity. Nat. Med. 27, 659–667 (2021).
Banday, A. R. et al. Genetic regulation of OAS1 nonsense-mediated decay underlies association with COVID-19 hospitalization in patients of European and African ancestries. Nat. Genet. 54, 1103–1116 (2022).
Thibord, F., Chan, M. V., Chen, M.-H. & Johnson, A. D. A year of COVID-19 GWAS results from the grasp portal reveals potential genetic risk factors. HGG Adv. 3, 100095 (2022).
Ellinghaus, D. Covid-19 host genetics and abo blood group susceptibility. Camb. Prisms: Precis. Med. 1, e10 (2023).
Bovijn, J., Lindgren, C. M. & Holmes, M. V. Genetic variants mimicking therapeutic inhibition of IL-6 receptor signaling and risk of COVID-19. Lancet Rheumatol. 2, e658–e659 (2020).
Palmos, A. B. et al. Proteome-wide mendelian randomization identifies causal links between blood proteins and severe covid-19. PLoS Genet. 18, e1010042 (2022).
Investigators, R.-C. Interleukin-6 receptor antagonists in critically ill patients with covid-19. N. Engl. J. Med. 384, 1491–1502 (2021).
Group, R. C. et al. Tocilizumab in patients admitted to hospital with covid-19 (recovery): a randomised, controlled, open-label, platform trial. Lancet 397, 1637 (2021).
Zoabi, Y., Deri-Rozov, S. & Shomron, N. Machine learning-based prediction of COVID-19 diagnosis based on symptoms. NPJ Digit. Med. 4, 1–5 (2021).
Ackermann, M. et al. Pulmonary vascular endothelialitis, thrombosis, and angiogenesis in COVID-19. N. Engl. J. Med. 383, 120–128 (2020).
Varga, Z. et al. Endothelial cell infection and endotheliitis in COVID-19. Lancet 395, 1417–1418 (2020).
Chowdary, P. Covid-19 coagulopathy–what should we treat? Exp. Physiol. 107, 749–758 (2022).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 (2020).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Lee, J. S. & Shin, E.-C. The type i interferon response in covid-19: implications for treatment. Nat. Rev. Immunol. 20, 585–586 (2020).
Zhu, L. et al. Single-cell sequencing of peripheral mononuclear cells reveals distinct immune response landscapes of covid-19 and influenza patients. Immunity 53, 685–696 (2020).
Melms, J. C. et al. A molecular single-cell lung atlas of lethal COVID-19. Nature 595, 114–119 (2021).
Dapat, C. et al. Gene signature of children with severe respiratory syncytial virus infection. Pediatr. Res. 89, 1664–1672 (2021).
Heinonen, S. et al. Immune profiles provide insights into respiratory syncytial virus disease severity in young children. Sci. Transl. Med. 12, eaaw0268 (2020).
Chappin, K. et al. Airway and blood monocyte transcriptomic profiling reveals an antiviral phenotype in infants with severe respiratory syncytial virus infection. J. Infect. Dis. 229, S100–S111 (2024).
Ahern, D. J. et al. A blood atlas of covid-19 defines hallmarks of disease severity and specificity. Cell 185, 916–938 (2022).
Wang, D. et al. Combatdb: a database for the covid-19 multi-omics blood atlas. Nucleic Acids Res. 51, D896–D905 (2023).
Krieg, S. J. et al. Data-driven testing program improves detection of covid-19 cases and reduces community transmission. NPJ Digit. Med. 5, 17 (2022).
Liu, T.-Y. et al. An individualized predictor of health and disease using paired reference and target samples. BMC Bioinformatics 17, 47 (2016).
Zaas, A. K. et al. Gene expression signatures diagnose influenza and other symptomatic respiratory viral infections in humans. Cell Host Microbe 6, 207–217 (2009).
Fourati, S. et al. A crowdsourced analysis to identify ab initio molecular signatures predictive of susceptibility to viral infection. Nat. Commun. 9, 4418 (2018).
Chawla, D. G. et al. Benchmarking transcriptional host response signatures for infection diagnosis. Cell Syst. 13, 974–988 (2022).
Kosmicki, J. A. et al. Genetic risk factors for COVID-19 and influenza are largely distinct. Nat. Genet. 56, 1592–1596 (2024).
Geller, F. et al. Central role of glycosylation processes in human genetic susceptibility to SARS-CoV-2 infections with Omicron variants. medRxiv (2024).
Er, A. G. et al. Multimodal data fusion using sparse canonical correlation analysis and cooperative learning: a covid-19 cohort study. NPJ Digit. Med. 7, 117 (2024).
Mejias, A. et al. Whole blood gene expression profiles to assess pathogenesis and disease severity in infants with respiratory syncytial virus infection. PLoS Med. 10, e1001549 (2013).
Group, S. C.-G. Genomewide association study of severe COVID-19 with respiratory failure. N. Engl. J. Med. 383, 1522–1534 (2020).
Murawski, M. R. et al. Respiratory syncytial virus activates innate immunity through toll-like receptor 2. J. Virol. 83, 1492–1500 (2009).
Mailaparambil, B., Krueger, M., Heinze, J., Forster, J. & Heinzmann, A. Polymorphisms of toll like receptors in the genetics of severe RSV associated diseases. Dis. Markers 25, 59–65 (2008).
Shojaei, M. et al. Ifi27 transcription is an early predictor for covid-19 outcomes, a multi-cohort observational study. Front. Immunol. 13, 1060438 (2023).
Gheblawi, M. et al. Angiotensin-converting enzyme 2: Sars-cov-2 receptor and regulator of the renin-angiotensin system: celebrating the 20th anniversary of the discovery of ACE2. Circ. Res. 126, 1456–1474 (2020).
Bouhaddou, M. et al. The global phosphorylation landscape of sars-cov-2 infection. Cell 182, 685–712 (2020).
Koutsakos, M. et al. Human cd8+ t cell cross-reactivity across influenza a, b and c viruses. Nat. Immunol. 20, 613–625 (2019).
de Graaf, M. & Fouchier, R. A. Role of receptor binding specificity in influenza a virus transmission and pathogenesis. EMBO J. 33, 823–841 (2014).
Long, J. S., Mistry, B., Haslam, S. M. & Barclay, W. S. Host and viral determinants of influenza a virus species specificity. Nat. Rev. Microbiol. 17, 67–81 (2019).
Geerdink, R. J., Pillay, J., Meyaard, L. & Bont, L. Neutrophils in respiratory syncytial virus infection: a target for asthma prevention. J. Allergy Clin. Immunol. 136, 838–847 (2015).
Kirsebom, F. C., Kausar, F., Nuriev, R., Makris, S. & Johansson, C. Neutrophil recruitment and activation are differentially dependent on myd88/trif and mavs signaling during rsv infection. Mucosal Immunol. 12, 1244–1255 (2019).
Lee, J. S. et al. Immunophenotyping of covid-19 and influenza highlights the role of type i interferons in development of severe covid-19. Sci. Immunol. 5, eabd1554 (2020).
Beigel, J. H. et al. Remdesivir for the treatment of covid-19-preliminary report. N. Engl. J. Med. 383, 1813–1836 (2020).
Group, R. C. Dexamethasone in hospitalized patients with covid-19. N. Engl. J. Med. 384, 693–704 (2021).
Kalil, A. C. et al. Baricitinib plus remdesivir for hospitalized adults with covid-19. N. Engl. J. Med. 384, 795–807 (2021).
ATTACC Investigators Therapeutic anticoagulation with heparin in noncritically ill patients with COVID-19. N. Engl. J. Med. 385, 790–802 (2021).
Chen, Z. & John Wherry, E. T cell responses in patients with COVID-19. Nat. Rev. Immunol. 20, 529–536 (2020).
Moss, P. The T cell immune response against SARS-CoV-2. Nat. Immunol. 23, 186–193 (2022).
Hammitt, L. L. et al. Nirsevimab for prevention of RSV in healthy late-preterm and term infants. N. Engl. J. Med. 386, 837–846 (2022).
Bastard, P. et al. Autoantibodies against type I IFNS in patients with life-threatening COVID-19. Science 370, eabd4585 (2020).
Hammond, J. et al. Oral nirmatrelvir for high-risk, nonhospitalized adults with COVID-19. N. Engl. J. Med. 386, 1397–1408 (2022).
Kousathanas, A. et al. Whole-genome sequencing reveals host factors underlying critical COVID-19. Nature 607, 97–103 (2022).
COVID-19 Host Genetics Initiative The covid-19 host genetics initiative. Eur. J. Hum. Genet. 28, 715–718 (2020).
Stikker, B. S. et al. Severe covid-19-associated variants linked to chemokine receptor gene control in monocytes and macrophages. Genome Biol. 23, 96 (2022).
Tal, G. et al. Association between common toll-like receptor 4 mutations and severe respiratory syncytial virus disease. J. Infect. Dis. 189, 2057–2063 (2004).
Caballero, M. T. et al. TLR4 genotype and environmental LPS mediate RSV bronchiolitis through Th2 polarization. J. Clin. Investig. 125, 571–582 (2015).
Löfgren, J., Marttila, R., Renko, M., Rämet, M. & Hallman, M. Toll-like receptor 4 asp299gly polymorphism in respiratory syncytial virus epidemics. Pediatr. Pulmonol. 45, 687–692 (2010).
Işık, Y. E. & Aydın, Z. Comparative analysis of machine learning approaches for predicting respiratory virus infection and symptom severity. PeerJ 11, e15552 (2023).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Sun, B. B. et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 622, 329–338 (2023).
Eldjarn, G. H. et al. Large-scale plasma proteomics comparisons through genetics and disease associations. Nature 622, 348–358 (2023).
Yao, C. et al. Genome-wide mapping of plasma protein qtls identifies putatively causal genes and pathways for cardiovascular disease. Nat. Commun. 9, 3268 (2018).
Pietzner, M. et al. Mapping the proteo-genomic convergence of human diseases. Science 374, eabj1541 (2021).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic Kras-driven cancers require tbk1. Nature 462, 108–112 (2009).
Milacic, M. et al. The Reactome Pathway Knowledgebase 2024. Nucleic Acids Res. 52, D672–D678 (2024).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. Kegg: integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551 (2021).
Abdelrahman, Z., Chen, Z., Lyu, H. & Wang, X. Comparisons of the immunological landscape between covid-19, influenza, and respiratory syncytial virus patients by clustering analysis. Comput. Struct. Biotechnol. J. 19, 2347–2355 (2021).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47 (2015).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for rna-seq read counts. Genome Biol. 15, R29 (2014).
Hemani, G. et al. The mr-base platform supports systematic causal inference across the human phenome. elife 7, e34408 (2018).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Hänzelmann, S., Castelo, R. & Guinney, J. Gsva: gene set variation analysis for microarray and rna-seq data. BMC Bioinformatics 14, 7 (2013).
Wilk, A. J. et al. A single-cell atlas of the peripheral immune response in patients with severe covid-19. Nat. Med. 26, 1070–1076 (2020).
Garcia-Gonzalez, P. et al. Dysregulated immune responses in covid-19 patients correlating with disease severity and invasive oxygen requirements. Front. Immunol. 12, 769059 (2021).
Paget, C. & Trottein, F. Covid-19 and flu: conserved or specific immune signature? Cell. Mol. Immunol. 18, 245–246 (2021).
Ren, X. et al. Covid-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913 (2021).
Acknowledgements
This work was supported by the grants GZNL2024A01003 from the Major Project of Guangzhou National Laboratory, grant 2024B1515020080 from Guangdong Basic and Applied Basic Research Foundation, and grant KY012023362 from the Talent Research Funding Project of Guangdong Provincial People’s Hospital.
Author information
Authors and Affiliations
Contributions
Y.Z. and F.Y. conceived the project. Y.Z., L.Z., Z.Z., Y.L., and Z.J. performed computational experiments and contributed to data interpretation. Y.Z. and L.Z. wrote the manuscript with input from all authors. F.Y. supervised and directed the study. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The author declares that there are no financial or non-financial competing interests relevant to the content of this work.
Consent for publication
This work exclusively utilizes de-identified datasets available from public repositories.
Materials availability
No new physical materials were produced.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Y., Zhang, L., Zhang, Z. et al. Explainable AI multiomics analysis reveals shared and divergent host responses in COVID-19 and influenza. npj Digit. Med. 9, 111 (2026). https://doi.org/10.1038/s41746-025-02291-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02291-w