Abstract
The progression from metabolic dysfunction-associated steatotic liver disease (MASLD) to metabolic dysfunction-associated steatohepatitis (MASH) is a critical link leading to cirrhosis and hepatocellular carcinoma. Yet the responsible cellular programs remain unclear. We integrated public single-cell, spatial, and bulk transcriptomic datasets to map microenvironmental remodeling and regulatory networks during MASLD-MASH progression. Among the seven major liver cell types identified, monocytes/macrophages and hepatic stellate cells (HSCs) were significantly enriched and demonstrated spatial co-localization within the context of MASH. We identified a DTNA+distinct macrophage subpopulation that was specifically enriched in MASH. This subpopulation exhibited characteristics consistent with M2 polarization, hypoxia, and enhanced inflammatory signaling. Pseudotime trajectory analysis revealed that this state represents a differentiation pathway originating from Kupffer cells to the DTNA+ state. RUNX2 emerged as the key transcriptional regulator. Cell communication analysis demonstrated that DTNA+ macrophages potentially interact with activated HSCs via the RUNX2-PLG-PARD3 axis, contributing to the exacerbation of liver fibrosis. Finally, ensemble machine learning models (mean AUC = 0.839), identified DTNA as the optimal predictive biomarker for distinguishing MASLD from MASH. This study highlight DTNA+ macrophages and the RUNX2–PLG–PARD3 axis as candidate mechanisms and targets for non-invasive diagnosis and therapy in MASH.
Similar content being viewed by others
Introduction
MASLD has emerged as the most prevalent chronic liver disease worldwide, affecting approximately 38% of the adult population globally, with an incidence rate of approximately 50 new cases per 1000 person-years. Its prevalence is notably higher in individuals with obesity and type 2 diabetes1. Driven by changes in lifestyle and dietary patterns, the incidence of MASLD is continuously rising, and this disease is now recognized as an independent risk factor for hepatocellular carcinoma (HCC)2. The pathological spectrum of MASLD ranges from simple steatosis to MASH, the latter being characterized by inflammation, hepatocyte injury, and fibrosis. Approximately 20–30% of MASH patients ultimately progress to cirrhosis and potentially HCC3. Although the “multiple-hit” model, involving lipid accumulation, oxidative stress, inflammatory cell infiltration, and fibrotic activation, is implicated in the transition from MASLD to MASH, the precise underlying molecular mechanisms remain to be determined. This knowledge gap has hindered the development of reliable biomarkers for early prediction and effective intervention strategies4. Conventional diagnostic tools, such as imaging and biochemical markers (e.g., liver enzymes and fibrosis scores), are widely used but fail to capture the cellular-level heterogeneity of disease progression, often resulting in diagnostic delays and missed therapeutic windows5.
While conventional bulk transcriptomics has led to the identification of differentially expressed genes in MASLD/MASH, these methods average signals across all cells, consequently overlooking the cellular heterogeneity and spatial organization within the liver tissue. They are thus unable to resolve the specific cell subpopulations driving disease progression6. The recent rapid development of scRNA-seq and spatial transcriptomics has provided powerful tools for dissecting this complex tissue microenvironment. Specifically, scRNA-seq enables the identification of cell subpopulations, developmental trajectories, and regulatory networks. Spatial transcriptomics complements this by preserving tissue architecture, thereby revealing the spatial proximity and interactions among cells7,8. Furthermore, the combination of multi-omics integration analysis with machine learning algorithms has been successfully applied to biomarker screening and predictive model construction in various diseases9. The application of these technologies in liver disease research has demonstrated that macrophages and HSCs play critical roles in fibrosis progression. For example, it has been shown that inflammatory macrophages can activate HSCs by secreting cytokines, thereby promoting collagen deposition10.
Current research on MASLD/MASH progression is limited by incomplete characterization of cellular heterogeneity in liver diseases and insufficient integration of multi-omics data. For instance, although scRNA-seq has revealed the diversity of HSC subpopulations and macrophage heterogeneity in the human liver, it often lacks spatial context due to tissue dissociation biases and has limited capacity for capturing rare cell types11,12,13. Furthermore, multi-omics integration for MASLD/MASH is constrained by challenges such as the handling of missing data, the interpretation of high-dimensional data, and the selection of optimal omics combinations14,15,16. This has resulted in a limited understanding of the disease’s dynamic progression, as well as a scarcity of dynamic trajectory analyses and translational predictive models targeted at disease advancement. In this study, we sought to address this gap by integrating scRNA-seq, spatial transcriptomics, and bulk RNA-seq data from public datasets. Using this approach, we systematically resolved the key cell subpopulations, molecular networks, and regulatory mechanisms driving the transition from MASLD to MASH. Specifically, we identified a core population of DTNA+ macrophages that drives this progression by interacting with activated HSCs associated with the RUNX2-PLG-PARD3 axis, thereby exacerbating fibrosis. Concurrently, we employed machine learning models to evaluate the predictive value of these key markers. This research not only uncovered a novel role for myeloid cells within the liver microenvironment but also provides a theoretical basis and potential biomarkers for the early diagnosis of, and precision intervention in MASLD/MASH.
Results
The integration of single-cell and spatial transcriptomic atlases reveals the potential role of monocytes/macrophages in MASH progression
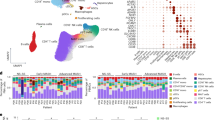
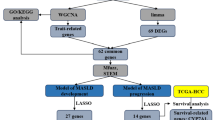
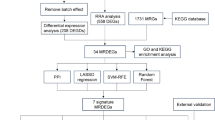
The study design is shown in Fig. 1. To explore the heterogeneity of MASLD, we integrated 115 samples from five scRNA-seq datasets: GSE289173, GSE212837, GSE202379, GSE189600, and GSE189175 (Fig. 2A, B; Supplementary Table 1), together with additional bulk transcriptomic datasets (RNA-seq and microarray), spatial transcriptomic data, and ATAC-seq datasets collected from public repositories (Supplementary Table 2). Among them, the single-cell cohort comprised 14 MASLD samples, 66 MASH samples, and 35 healthy control liver tissue samples (Fig. 2C). The MASH group was further stratified to include 9 cases of eMASH and 10 cases of aMASH. In total, 570,194 cells were retained for subsequent analysis and clustered into seven major cell types (Fig. 2D). These were annotated based on canonical marker gene expression, as follows: hepatocytes (n = 322,546; ASGR1, CYP3A4), endothelial cells (n = 93,372; STAB2), T cells (n = 64,205; CD3E, CD2), monocytes/macrophages (n = 37,907; CD163, MARCO), HSCs (n = 26,073; DCN, COL3A1), cholangiocytes (n = 20,910; KRT7, CFTR), and B cells (n = 5181; MS4A1, CD79A). These annotations were validated via a dot plot, which confirmed the distinct expression levels of marker genes in each cluster (Fig. 2E). The same cellular composition and expression patterns were also identified in the validation cohort (Fig. S1A, B).
Flow chart of the study.
UMAP analysis of the discovery cohort across five datasets before (A) and after (B) batch‑effect removal. C UMAP plots colored by disease status: Healthy (red), MASLD (green), and MASH (blue). D UMAP visualization of the seven major cell types identified in the discovery cohort. E Ratio of observed to expected (Ro/e) for major cell types in Healthy, MASLD, and MASH groups. F Ro/e comparison of major cell types between early MASH (eMASH) and advanced MASH (aMASH). G Dot plot showing the expression of canonical marker genes in major cell types, scaled by expression percentage and average expression level. H Heatmap of enriched metabolic pathways across distinct cell clusters, with significantly upregulated (red) and downregulated (blue) pathways indicated. I Spatial transcriptomics image showing signature scores for monocytes/macrophages, stellate cells, and hepatocytes, highlighting their spatial co‑localization in a MASH sample.
Although all seven major cell types were present across all groups, the infiltration levels varied across populations, potentially reflecting differences in the stages of MASLD progression. Among these major cell types, monocytes/macrophages and HSCs were significantly enriched in MASH (Fig. 2F). We quantified the proportional distribution of various cell types across MASLD, MASH, and Healthy groups. The results showed that the proportions of both hepatic stellate cells and monocytes/macrophages were increased in the NASH group (Supplementary Table 3). This enrichment was further confirmed in the validation cohort, where the numbers of monocytes/macrophages were found to be significantly increased in mice fed a high-fat high-fructose diet (HFHFD) for 30 weeks (Fig. S1C, D). Furthermore, compared with eMASH, the proportion of HSCs was further elevated in aMASH samples (Fig. 2G). We assessed the activity of metabolism-related pathways in each cell type using gene set enrichment analysis (GSEA), and revealing significant metabolic reprogramming in hepatocytes (Fig. 2H). Moreover, spatial deconvolution analysis using RCTD on seven MASH spatial transcriptomics samples revealed that there was a high degree of spatial co-localization between monocytes/macrophages and HSCs (Figs. 2I and S2). These findings suggest that monocytes/macrophages constitute a cell type that extensively interacts with HSCs, emphasizing the importance of exploring the heterogeneity of monocytes/macrophages as well as the characteristics underlying their associations with HSCs in MASH.
Identification of five HSC subtypes in liver tissue
To identify the five HSC-related subtypes in liver tissue (quiescent HSCs [qHSCs], intermediate HSCs [inHSCs], activated HSCs [aHSCs], hepatocyte-like HSCs [hep-like HSCs], and smooth muscle cells [SMCs]), we first performed UMAP-based dimensional reduction and clustering analysis (Fig. 3A). This achieved a clear separation of the five subtypes in two-dimensional space, with each subtype forming a distinct cluster with clear boundaries. This representation illustrated the population differentiation characteristics of the different subtypes, thereby providing a visual typing basis for subsequent identification. Further analysis of the gene expression markers for each subtype using a bubble plot (Fig. 3B and Supplementary Table 4) showed that resting-state-related genes, such as RELN, were highly expressed in qHSCs. In aHSCs, the expression of inflammation and activation markers such as KALRN and ITGBL1 was significantly upregulated. Smooth muscle differentiation-related genes, such as MYH11, were specifically highly expressed in SMCs. Hep-like HSCs and inHSCs also showed unique gene expression patterns, thus validating the identity of each subtype based on differential marker gene expression.
A UMAP visualization of five HSC subtypes in liver tissue: quiescent HSC (qHSC), inflammatory HSC (inHSC), activated HSC (aHSC), hepatocyte-like HSC (hep-like HSC), and smooth-muscle-cell-like HSC (SMC). B Dot plot displaying marker-gene expression for each HSC subtype, scaled by percentage of expressing cells and mean expression level. C Ro/e analysis quantifying relative abundance trends of each HSC subtype across MASLD and MASH. D Ro/e analysis quantifying relative abundance trends of each HSC subtype across eMASH and aMASH. E Volcano plot of differentially expressed genes (DEGs) between MASLD- and MASH-derived HSCs. F Bar chart showing KEGG pathways enriched in up-regulated DEGs of MASLD versus MASH HSCs. G CytoTRACE analysis evaluating differentiation-potential distribution across HSC subtypes. H, I Pseudotemporal trajectory analysis revealing differentiation paths: qHSC → inHSC → aHSC / hep-like HSC / SMC. J Smoothed heat map illustrating dynamic gene-expression patterns along HSC differentiation.
Quantitative analysis using the Ro/e ratio (the ratio of observed to expected cell numbers, used to highlight dynamic changes in cell subpopulations during disease progression) (Fig. 3C) showed that, with disease progression (from MASLD to MASH), the Ro/e value of aHSCs increased, whereas that of qHSCs decreased. A comparison specifically between eMASH and aMASH demonstrated that the Ro/e value of aHSCs was higher in aMASH than in eMASH (Fig. 3D). Collectively, these findings precisely illustrate the dynamic changes in the characteristics of the five HSC subtypes throughout the pathological process.
To further elucidate the differences in HSCs between MASLD and MASH, we performed a differential analysis of the HSCs from the two groups (Fig. 3E, F and Supplementary Table 5). The results indicated that HSCs in the MASLD group were primarily enriched in pathways such as Axon Guidance, Relaxin Signaling Pathway, and Dopaminergic Synapse. In contrast, HSCs in the MASH group were mainly enriched in pathways related to Cholesterol Metabolism and Bile Secretion.
CytoTRACE analysis (Fig. 3G) was used to evaluate the differentiation potential of each HSC subtype. This analysis revealed that qHSCs were concentrated in the high CytoTRACE value region, while aHSCs and SMCs were distributed in the low CytoTRACE value region. This observation strongly suggests that qHSCs may represent the progenitor state subtype. In combination with cell trajectory analysis (Fig. 3H, I), it was further clarified that the five subtypes follow the differentiation path: “qHSC→inHSC→aHSC/hep-like HSC/SMC”. States 4 and 5 correspond to the qHSC progenitor state, and State 1 corresponds to the aHSC or SMC terminal differentiation state, clearly presenting the developmental hierarchical relationship among the subtypes. Finally, gene expression patterns across these state transitions were analyzed and are presented in a smoothed heatmap in Fig. 3J.
DTNA+ macrophages are key cells in the progression of MASLD to MASH
UMAP analysis revealed seven distinct clusters of hepatic myeloid cells, including VCAM1+ Kupffer cells (KCs), SAA1+ KCs, DTNA+ macrophages, VCAN+ monocytes, myeloid dendritic cells (mDCs), plasmacytoid dendritic cells (pDCs), and MKI67+ macrophages (Fig. 4A). This clustering highlights the spatial distribution of the cells based on transcriptional profiles. We generated a dot plot of key marker genes (Fig. 4B and Supplementary Table 6), which demonstrated the genes that were highly expressed in each cell type: CD163 and MARCO in resident macrophages; VCAM1 and SAA1 in KC subtypes; DTNA and ITGAX in DTNA+ macrophages; VCAN and FCN1 in monocytes; and proliferation markers, such as MKI67 and TOP2A, in MKI67+ macrophages. These markers are scaled by expression percentage and average expression level. We further quantified the cell proportions (Ro/e value) between MASLD and MASH, and found that the number of DTNA+ macrophages was significantly increased in MASH, concurrent with a decrease in other clusters, such as VCAM1+ KCs (Fig. 4C).
A UMAP projection of hepatic myeloid compartments revealing clusters including VCAM1⁺ KCs, SAA1⁺ KCs, and DTNA⁺ macrophages. B Dot plot of key marker genes across myeloid clusters, scaled by percentage of expressing cells and mean expression level. C Ro/e analysis showing relative abundance changes of each myeloid cluster between MASLD and MASH; DTNA⁺ macrophages are significantly expanded in MASH. D Heat map of z-scored metabolic and inflammatory pathways specifically activated in MASH-derived DTNA⁺ macrophages. E Box plots comparing glycolysis, hypoxia, and other pathway scores in DTNA⁺ macrophages between MASLD and MASH (*P < 0.05, ***P < 0.001). F Heat map of Hallmark gene-set enrichment across the seven identified myeloid populations. G irGSEA highlighting differentially enriched pathways between MASLD and MASH myeloid cells; angiogenesis and coagulation are up-regulated in MASH (p <0.05 to <0.0001).
We generated a heatmap of metabolic and inflammatory pathway z-scores, which demonstrated enhanced M2 macrophage polarization and activation of hypoxia/HIF regulatory pathways in DTNA+ macrophages during MASH (Fig. 4D). Significant increases (p < 0.05 to < 0.001) in glycolysis, hypoxia, lipid mediator activity, M1/M2 polarization, pro-inflammatory responses, TCA cycle activity, and interferon scores were also observed in MASH compared with MASLD (Fig. 4E). A heatmap of Hallmark gene set enrichment (Fig. 4F) also revealed elevated inflammatory signaling, including TNFA signaling via NFKB and IL6-JAK-STAT3, alongside increased coagulation and cholesterol homeostasis in DTNA+ macrophages in MASH. Finally, GSEA was performed to visualize the differentially enriched pathways between MASLD and MASH. The results showed significant upregulation of angiogenesis, coagulation, hypoxia, and inflammatory signaling within MASH-oriented clusters, particularly those enriched for DTNA+ macrophages (p < 0.05 to p < 0.0001) (Fig. 4G). Collectively, these findings imply that DTNA+ macrophages comprise a key cell population driving the transition from MASLD to MASH, mediating disease progression through enhanced M2 polarization and pro-inflammatory adaptive mechanisms.
Pseudotime and transcriptional regulatory analysis of DTNA+ macrophages
CytoTRACE analysis was used to establish the stemness score distribution across hepatic myeloid cells, spanning a normalized range from 0.00 (highly differentiated) to 1.00 (highly stem-like). The resulting gradient demonstrated that specific populations, namely VCAM1+ KCs and SAA1+ KCs, exhibit greater stemness potential (Fig. 5A). A subsequent projection plot, delineating cell states (States 1–5), illustrated a defined evolutionary progression from States 1 to 5, indicating a potential differentiation trajectory (Fig. 5B). Furthermore, a projection plot color-coded by cell type distinguished clusters—including VCAM1+ KCs, SAA1+ KCs, DTNA+ macrophages, VCAN+ monocytes, mDCs, pDCs, and MKI67+ macrophages—and specifically highlighted the spatial localization of the DTNA+ macrophages within the plot (Fig. 5C). The accompanying gene expression patterns throughout the state transitions are presented in a smoothed heatmap in Fig. 5D.
A CytoTRACE projection showing stemness scores (0–1 gradient) across hepatic myeloid cells. B State-map visualization tracing the evolution of cellular states (States 1–5). C State map color-coded by cell-type annotation, highlighting the position of DTNA⁺ macrophages. D Smoothed heat map of gene-expression dynamics along the inferred myeloid differentiation trajectory. E Heat map depicting transcriptional activity of M1–M17 regulon modules. F Top 10 transcription factors (TFs) ranked by regulon specificity score (RSS) in each myeloid subset, with DTNA⁺ macrophage-associated TFs highlighted. G Venn diagram intersecting two TF sets, identifying BHLHE41, RUNX2, BACH2, and MEF2C as shared regulators.
Next, we performed SCENIC analysis to identify key transcription factors (TFs) responsible for driving cell state transitions during disease development. This analysis resolved distinct gene regulatory networks (GRNs), or regulons, across different cell populations. A GRN is defined as a collection of TFs and cofactors that collectively regulate the expression of downstream genes, leading to specific cellular states. We identified a total of 360 significant regulons. These regulons were subsequently categorized and re-clustered using the Connection Specificity Index (CSI). Transcriptional activity across the resulting M1–M17 modules was characterized using a heatmap (Fig. 5E). Notably, DTNA+ macrophages demonstrated significant enrichment in modules M2, M3, M10, and M12, which together comprised 74 TFs (Fig. S3A and Supplementary Table 7). We also ranked the top 10 TFs in various cell types based on the Regulon Specificity Score (RSS), and noted that these TFs were strongly associated with DTNA+ macrophages (Figs. 5F and S3B). A Venn diagram was used to illustrate the intersection of two distinct regulon sets (Fig. 5G), identifying four specific TFs—BHLHE41(+), RUNX2(+), BACH2(+), and MEF2C(+)—that were highly specific to DTNA+ macrophages. Among them, RUNX2 has been reported to promote HSC activation and liver fibrosis by regulating ITGAV expression17. Furthermore, RUNX2 upregulation promotes MCP-1 expression, enhances hepatic macrophage infiltration18, and drives hepatocyte death, leading to fibrosis in MASH19. These findings suggest that RUNX2 may play a crucial role in the development and progression of liver fibrosis through myeloid-mediated mechanisms. Collectively, these data reveal the differentiation trajectory, gene modules, and regulatory networks of DTNA+ macrophages.
DTNA+ macrophages potentially interact with aHSCs via the RUNX2-PLG-PARD3 axis, which may contribute to the exacerbation of liver fibrosis and progression to MASH
The network diagram shown in Fig. 6A illustrates the number of cell-cell interactions in MASLD and MASH. It highlights a significant increase in interactions involving DTNA+ macrophages and other cells—specifically aHSCs, hep-like HSCs, and inHSCs—in MASH, indicating that communication is enhanced during disease progression. A heatmap (Fig. 6B) of differential interaction counts and strength further shows relative changes (from −20 to 20) between senders and receivers in MASH versus MASLD. DTNA+ macrophages, acting as the sender, showed a positive differential interaction enhancement with cells such as aHSCs, meaning that this specific interaction was significantly enhanced in MASH. Intercellular communication mediated by the PLG-PARD3 pair was also observed to be specifically enhanced in MASH, particularly in DTNA+ macrophages (Fig. 6C). Violin plots (Fig. 6D) comparing PLG and PARD3 expression in different cell clusters across MASLD (pink) and MASH (blue) show PLG activation in DTNA+ macrophages in MASH, while PARD3 is expressed in HSCs under both conditions. Finally, correlation scatter plots from the GSE126848 dataset (Fig. 6E) show a correlation between the fibrosis signature score and PLG expression (R = 0.639, p = 0.00011), and between RUNX2 and PLG expression (R = 0.655, p = 6.33 × 10−5), suggesting that RUNX2 influences fibrosis via PLG. These findings were corroborated in other datasets (Fig. S4).
A Network of intercellular interaction numbers showing increased DTNA⁺ macrophage contacts with aHSCs and other cells in MASH. B Differential-interaction heat map (MASH vs. MASLD); red indicates strengthened, blue weakened interactions. C Circle plot displaying communication probability of ligand–receptor pairs originating from DTNA⁺ macrophages. D Violin plots comparing PLG and PARD3 expression in MASLD (pink) versus MASH (blue). E Scatter plots illustrating correlation between fibrosis-signature score and PLG expression (R = 0.639, p = 0.00011), and between RUNX2 and PLG (R = 0.655, p = 6.33 × 10⁻⁵). F Correlation matrix across nine MASH bulk RNA-seq datasets revealing positive association (Pos) between DTNA⁺ macrophages and aHSCs.
We profiled genome-wide chromatin accessibility landscapes using ATAC-seq on liver biopsies from patients with MASH and MASLD. Our specific focus was on the regulatory circuitry of the PLG locus and the annotation of RUNX2 transcription factor binding sites (TFBS). In the MASH versus MASLD comparison, we identified 145,812 upregulated and 10,567 downregulated differential accessibility regions (DARs) (Fig. S5A and Supplementary Table 8). Importantly, all seven PLG-associated cis-regulatory elements were mapped to the up-cluster, emphasizing their functional role in MASH pathogenesis (Supplementary Table 9). Motif enrichment within these seven elements yielded 27 specific RUNX2 TFBS (Supplementary Table 10). Functional annotation of the TFBS (Fig. S5B and Supplementary Table 11) revealed that RUNX2 predominantly binds to PLG promoters and introns, with a smaller number of sites located in distal intergenic regions. This finding experimentally validates the RUNX2-PLG regulatory axis. Distribution analysis relative to the PLG transcription start site (TSS) further showed that most TFBS resided 10–100 kb downstream or upstream of the TSS (Fig. S5C). Collectively, these data uncover MASH-specific epigenetic reprogramming and offer novel mechanistic insights for therapeutic strategies targeting the RUNX2-PLG axis. Additionally, considering the importance of lactate accumulation in liver fibrosis20,21, we supplemented our analysis with a correlation study between key targets and lactate metabolism-related genes (Fig. S6). The results showed that three key genes (RUNX2, PLG, and PARD3) were significantly correlated with multiple lactate metabolism-related genes, suggesting a potential metabolic regulatory mechanism. In the scRNA-seq validation cohort, we found that RUNX2 expression was upregulated in mice fed a HFHFD for 30 weeks(Fig. S7A, B). Finally, a correlation matrix of cell types across nine MASH bulk datasets showed that there was a significant positive correlation between DTNA+ macrophages and aHSCs in six of the datasets (p < 0.05, R > 0.8; Fig. 6F).
To accelerate the translation from mechanistic discovery to therapeutic strategy, we performed a joint drug-repurposing screen targeting the three core genes RUNX2, PLG, and PARD3 via the CTD database. This yielded 18 candidate compounds that simultaneously modulate all three targets. Each of these drugs has been reported for the treatment of fatty-liver disease, liver injury, cirrhosis, or non-alcoholic fatty liver disease, as summarized in Fig. S8. Overall, these data suggest that DTNA+ macrophages play a critical role in MASLD, potentially interacting with aHSCs via the RUNX2-PLG-PARD3 axis, thereby exacerbating liver fibrosis and driving the progression to MASH.
PLG secreted by DTNA+ macrophages may contribute to liver fibrosis via complement and coagulation cascades
MASH samples were stratified into high-PLG and low-PLG expression groups based on the 75th percentile PLG level derived from DTNA+ macrophages. Differentially expressed genes in DTNA+ macrophages were compared between these two groups to characterize transcriptional differences associated with varying levels of macrophage-derived PLG (Fig. 7A and Supplementary Table 12). GSEA (Gene Set Enrichment Analysis) of macrophages in the high-PLG group revealed significant enrichment of the “Complement and coagulation cascades” pathway (Fig. 7B, C). In parallel, spatial transcriptomics data from MASH demonstrated that there was strong spatial co-localization between DTNA+ macrophages and aHSCs. Quantification of this relationship using scatter plots showed the DTNA+ macrophage signature score was significantly and positively correlated with the aHSC signature score across patient samples (Figs. 7D, E and S9). Consistent with these findings, the PLG–PARD3 axis exhibited pronounced spatial co-expression (Fig. S10).
A Volcano plot of differentially expressed genes in DTNA⁺ macrophages between high-PLG and low-PLG groups. B, C GSEA revealing significant enrichment of the “Complement and coagulation cascades” pathway in the high-PLG group. D, E Spatial transcriptomics showing pronounced co-localization of DTNA⁺ macrophages and aHSCs in MASH sections; correlation scatter plot confirms a significant positive association (p < 0.001).
DTNA+ macrophages can accurately predict MASLD and MASH
The predictive power of DTNA+ macrophages for MASLD and MASH was assessed using machine learning. We generated a heatmap illustrating the AUC performance of 113 machine learning models across the training set (GSE135251) and five external validation sets. This analysis highlighted the ElasticNet model (Enet, α = 0.9) as having the highest predictive accuracy (mean AUC = 0.839) (Fig. 8A). This result indicates that ElasticNet is superior for classifying MASLD and MASH. We plotted the Receiver Operating Characteristic (ROC) curves for the model genes derived from the optimal Enet [α = 0.9] model across all cohorts, thereby validating the generalizability of these model genes. Critically, DTNA itself exhibited the highest predictive power across all datasets (Fig. 8B–G). Furthermore, box plots demonstrated that the expression of ARHGAP22, DHRS9, DTNA, ITGAX, RUNX2, and TRPM2 was significantly elevated in the MASH group compared to the MASLD group, whereas that of TESK2 showed no significant difference (Fig. 8H). Furthermore, IGV visualization of ATAC-seq data revealed that chromatin accessibility within ±2 kbp of the promoter regions of eight model genes was significantly higher in MASH samples compared to MASLD samples (Fig. S11). Collectively, these data demonstrate that the signature genes of DTNA+ macrophages, particularly DTNA, achieve high AUC values across multiple independent cohorts and can accurately predict MASLD and MASH.
A Heat map of AUC performance for 113 machine-learning models in training and validation sets; Enet (α = 0.9) achieved the highest mean AUC = 0.839. B–G ROC curves depicting predictive performance of the optimal gene signature across independent cohorts; DTNA consistently yielded the highest AUC in all datasets. H Box plots comparing expression of top signature genes between MASLD and MASH (***p < 0.001, *p < 0.05, ns not significant).
SHAP analysis was subsequently conducted on the Enet [α = 0.9] model to quantify the impact of features on the output for MASLD and MASH classification. The mean absolute SHAP values, visualized in a bar plot (Fig. 9A), indicated that DTNA (0.100) was the most important feature, followed by DHRS9 (0.011), TRPM2 (0.009), TESK2 (0.007), ARHGAP22 (0.006), ITGAX (0.004), RUNX2 (0.003), and AUTS2 (0.002). This hierarchy suggests that DTNA is the primary contributor to the model’s prediction. A bee swarm plot was generated to display the SHAP value distribution. As shown in Fig. 9B, DTNA exhibited the widest horizontal spread in both high (red) and low (blue) values, indicating that its expression level, regardless of direction, exerted a significant effect on the model output. A waterfall plot showed the cumulative feature contribution to the final prediction [f(x) = 0.158] from the base value E[f(x)] = 0. SHAP values indicate the contribution of each feature to the model’s prediction: a positive value means the feature pushes the prediction upward, whereas a negative value pulls it downward. Starting from the baseline E[f(x)] = 0, these contributions are summed to reach the final output, thereby quantifying each feature’s importance. DTNA (+1.01), RUNX2 (+0.643), and TRPM2 (+0.755) functioned as positive drivers, while TESK2 (−0.852) and DHRS9 (−0.748) exerted negative effects from the base value E[f(x)] = 0, showing DTNA and RUNX2 as key features driving the prediction (Fig. 9C). DTNA (+1.01) and RUNX2 (+0.643) were the main positive contributors, with TRPM2 (+0.755) jointly enhancing the prediction, while TESK2 (−0.852) and DHRS9 (−0.748) had a negative impact, collectively supporting the dominant role of DTNA in the model (Fig. 9D). These results consistently demonstrate the high predictive ability of DTNA+ macrophages, via DTNA expression, to accurately distinguish and predict MASLD and its progression to MASH.
A Bar plot of mean absolute SHAP values; DTNA ranks highest (0.100) as the most influential feature. B Bee-swarm plot of SHAP value distribution, revealing DTNA as the dominant driver. C Waterfall plot illustrating cumulative feature contributions to the prediction output; DTNA (+1.01) and RUNX2 (+0.643) are the primary positive contributors. D Force plot confirming the positive impact of DTNA and RUNX2, alongside the negative influence of DHRS9.
Discussion
By integrating multi-omics data from scRNA-seq, spatial transcriptomics, and bulk transcriptomics, this study systematically delineated the key cellular and molecular mechanisms driving the progression from MASLD to MASH. We identified seven major liver cell types, with monocytes/macrophages and HSCs showing marked enrichment and spatial co-localization in MASH. Further subdivision revealed five HSC subtypes, highlighting a marked transition from qHSCs to aHSCs with increasing disease severity. Within the monocyte/macrophage compartment, DTNA+ macrophages emerged as a critical subpopulation, showing an increased proportion in MASH, accompanied by M2 polarization and enhanced hypoxia signaling. Pseudotime analysis suggested a differentiation trajectory from KCs toward the DTNA+ state. Transcriptional regulatory network analysis identified distinct regulon modules in DTNA+ macrophages, including TFs such as RUNX2, that may contribute to pro-inflammatory and fibrotic programs. Cell-cell communication analysis suggested the RUNX2-PLG-PARD3 axis linking DTNA+ macrophages and aHSCs, which may contribute to liver fibrosis. Moreover, an ensemble machine learning model identified markers, including DTNA, with strong predictive performance (mean AUC = 0.839) in distinguishing MASLD from MASH. Collectively, these findings clarify how cellular heterogeneity, spatial organization, and regulatory networks contribute to disease progression, offering a mechanistic foundation for both biological insight and clinical translation.
An integrated multi-omics analysis delineated the central function of the monocyte/macrophage lineage in the progression from MASLD to MASH, specifically emphasizing the enrichment of DTNA+ macrophages and their direct interaction with HSCs. This finding corroborates established literature regarding the pivotal role of the inflammatory microenvironment in liver fibrosis17. Notably, however, our study provides enhanced mechanistic resolution through single-cell and spatial transcriptomic dimensions. Traditional bulk transcriptomics studies, while identifying the differential expression of genes related to lipid metabolism and inflammation in MASLD22, are fundamentally constrained by cellular heterogeneity and cannot resolve the contributions of distinct cellular subpopulations. Our investigation integrated 115 scRNA-seq and 7 spatial transcriptomics samples, classifying them into seven principal cell types: hepatocytes, endothelial cells, T cells, monocytes/macrophages, HSCs, cholangiocytes, and B cells. A significant increase in the number of monocytes/macrophages was detected in MASH, exhibiting spatial co-localization with activated aHSCs, thereby extending prior single-cell liver atlases that reported inflammatory activation of KCs in MASH23 Multiple studies have shown that M1-polarized macrophages serve as the principal mediators of early acute inflammation in MASH, contributing to hepatocyte injury and the consequential release of pro-inflammatory cytokines, such as TNF-α and IL-1β24,25. Conversely, the present study highlights the significant contribution of M2-polarized DTNA+ macrophages to the trajectory of fibrosis progression. This finding is concordant with existing evidence that M2 macrophages are actively involved in the chronic inflammation and tissue remodeling characteristic of late-stage MASH26,27,28. This discrepancy is potentially attributable to disease stage-specific immune response kinetics: M1 macrophages predominantly drive the initial inflammatory phase, while distinct M2-like subpopulations progressively manifest in the later fibrotic phase, thereby stimulating HSC activation and extracellular matrix (ECM) deposition26,27,28. Furthermore, differences arising from experimental models (e.g., human vs. rodent data), combined with the high resolution afforded by single-cell sequencing in this study, may reveal the existence of M1/M2 hybrid phenotypes. In this context, DTNA+ macrophages not only express M2 markers but also exhibit pro-inflammatory signaling, as evidenced by the upregulation of Hallmark gene sets such as TNFA-NFKB and IL6-JAK-STAT3. This observation, derived from our Hallmark GSEA, suggests that DTNA+ macrophages in MASH may simultaneously exacerbate chronic inflammation and accelerate fibrosis26,29. This insight both complements findings from animal models regarding macrophage-driven MASH inflammation19,22 and specifically emphasizes the metabolic-inflammatory coupling mechanisms observed in human samples.
Regarding the identification and dynamic alterations of HSC subtypes, this study successfully delineated five distinct populations: qHSC, inHSC, aHSC, hep-like HSC, and SMC. Marker gene expression for these subtypes, such as the prominent expression of KALRN and ITGBL1 in aHSCs, was validated using UMAP and DotPlot visualizations. Consistent with the core pathological process of hepatic fibrosis, which involves the transition of HSCs from a quiescent to an activated state, we observed that the proportion of aHSCs increased while that of qHSCs decreased as the disease progressed30,31. CytoTRACE and pseudotime trajectory analyses inferred a differentiation path that initiates from qHSCs, posited as the progenitor state, continuing toward either aHSCs or SMCs, a transformation accompanied by systematic changes in gene expression patterns during the state transitions. While studies by Zhang et al.32 and Dobie et al.33 provided substantial insights into the diversity and function of HSCs in fibrosis—the former focusing on activation/differentiation and the latter on spatial functional differences—our findings complement this work by supporting the view that HSCs constitute a complex population with multiple differentiation and functional states, rather than being products of a single differentiation trajectory. Crucially, our integration of human MASLD/MASH data unveiled dynamic pathology-specific changes, contrasting with previous work primarily centered on animal models32,33, thus underscoring the clinical relevance of human HSC heterogeneity. Spatial deconvolution analysis further confirmed the physical proximity and interaction between HSCs and monocytes/macrophages, thereby extending the understanding of microenvironment remodeling in MASH7,15.
The identification of DTNA+ macrophages and their role in MASH is a key finding of this study. UMAP clustering analysis demonstrated that DTNA+ macrophages (which co-express DTNA and ITGAX) are significantly distinct from other myeloid clusters, notably VCAM1+ and SAA1+ KCs, and their unique identity was confirmed by marker gene dot plot analysis. The proportion of this specific subpopulation was markedly elevated in MASH samples, concurrent with the activation of both inflammatory and metabolic pathways. Previous research suggested that CD11c+ (encoded by ITGAX) macrophages drive liver fibrosis in MASH mouse models by promoting hepatic crown-like structure (hCLS) formation triggered by hepatocyte death19. Furthermore, related reviews indicate that the composition of macrophage subpopulations exhibits dynamic alterations during MASH fibrosis and is intimately linked to microenvironment remodeling34,35. Building upon this background, our finding that the DTNA+ macrophage subpopulation is substantially enriched in MASH and is associated with the activation of inflammatory and metabolic pathways further substantiates the critical involvement of macrophages in liver disease microenvironment remodeling. However, our gene enrichment analysis provides a nuanced perspective by highlighting the upregulation of signaling pathways associated with angiogenesis, coagulation, and hypoxia, thereby challenging the traditional view that macrophages function solely as pro-inflammatory entities36. Moreover, CytoTRACE analysis indicated that DTNA+ macrophages possess low stemness, and pseudotime trajectory analysis revealed an evolutionary path from VCAM1+ KCs to the DTNA+ state, accompanied by specific changes in gene module expression. This differentiation sequence complements existing research on myeloid cell plasticity within the context of liver disease37.
Transcriptional regulatory network analysis provided a deeper mechanistic understanding of the DTNA+ macrophage population. Employing SCENIC, we identified 360 regulons, which were subsequently clustered into M1–M17 modules based on the CSI. DTNA+ macrophages exhibited enrichment within modules M2, M3, M10, and M12, encompassing a total of 74 TFs. According to our Rank Specificity Score (RSS) analysis, TFs such as BHLHE41, RUNX2, BACH2, and MEF2C demonstrated high specificity within this distinct subpopulation. RUNX2 is known to be upregulated in the fibrotic liver and in activated HSCs, where it promotes a pro-fibrotic HSC phenotype through the transcriptional activation of ITGAV. In MASLD models, HSC-specific RUNX2 can also upregulate MCP-1, thereby driving macrophage infiltration17. Building on this existing evidence, our multi-dataset analysis revealed that RUNX2 exhibits a significant positive correlation with the ligand PLG, and PLG is, in turn, significantly positively correlated with the degree of liver fibrosis. Furthermore, we observed that RUNX2 expression was upregulated in myeloid cells within a MASH mouse model. Accordingly, we propose that the RUNX2-PLG-PARD3 axis serves to enhance the interaction between DTNA+ macrophages and aHSCs. Although most relevant studies indicate that PLG mediates a reparative effect by degrading and remodeling the ECM, consequently inhibiting fibrosis formation38,39,40, our data revealed a significant positive correlation between PLG expression/activity and the severity of hepatic fibrosis. We hypothesize that, within the context of chronic liver injury, PLG function may become dysregulated, culminating in abnormal ECM deposition and subsequent progression of fibrosis. This phenomenon could be closely associated with the excessive activation of PLG, an imbalance of its related factors, or sustained activation of HSCs. Furthermore, PARD3 is an established regulator of cell polarity, known to influence tumor cell proliferation and migration through the modulation of cell-cell and cell-matrix interactions41,42. Given the high expression of PARD3 observed in HCC and its fundamental role in regulating cellular polarity, we propose that PARD3 may contribute to the progression of hepatic fibrosis by actively remodeling both cell polarity and the local microenvironment. The mechanism involving the proposed RUNX2-PLG-PARD3 axis significantly complements existing research on ligand-receptor interactions in MASH. Crucially, however, our study is the first to integrate this mechanism with transcriptional regulation, thereby establishing RUNX2 as an upstream driver of MASH pathogenesis. In addition to this axis, the FN1-(ITGAV + ITGB1) and C3-ITGAX/ITGB2 ligand-receptor pairs also exert significant influence on the MASH fibrosis process. Specifically, ITGB1 has been widely reported to promote the activation of HSCs, drive ECM deposition, and facilitate EMT, all of which exacerbate the progression of fibrosis43,44. This pathway further enhances fibroblast proliferation and collagen production, consequently increasing tissue stiffness and accelerating the overall fibrotic process43,44. Similarly, the complement component C3 promotes myeloid cell adhesion, phagocytosis, and the activation of pro-inflammatory signaling pathways, thereby sustaining chronic inflammation and HSC activation, which collectively drive fibrosis progression45,46,47. The aggregated data underscore the inherent complexity of the cross-cellular signaling networks involved. Consistent with this, the correlation matrix derived from bulk datasets confirmed the existence of a positive correlation between DTNA+ macrophages and aHSCs (R = 0.36, p < 2.2 × 10−16).
The results obtained from the machine learning ensemble model provide critical validation of the translational potential for the identified mechanisms. We constructed 113 distinct model combinations, with the Enet[α = 0.9] model achieving a mean AUC of 0.839 across the training and validation cohorts. Within this model, DTNA demonstrated the highest predictive power. SHAP analyses further quantified the substantial contribution of DTNA (mean absolute SHAP = 0.100), with RUNX2 and TRPM2 identified as significant positive drivers. This method offers a superior predictive capability compared to the application of single algorithms for MASH prediction, such as Random Forest (RF) models, which typically report AUCs of approximately 0.7548,49. Our study’s multi-paradigm ensemble approach ensured robustness and consistently identified markers—including DTNA, ITGAX, and RUNX2—that are significantly upregulated in MASH across multiple cohorts. In comparison to existing biomarkers, such as CK-18, a common MASH diagnostic marker with a reported AUC of approximately 0.80 in multiple studies and meta-analyses50,51, our single-cell-derived markers exhibit higher specificity and demonstrate effective generalizability across diverse GEO datasets. This outcome not only echoes recent multicenter studies employing machine learning for risk stratification of MASH/at-risk MASH33 but also aligns with the concept of leveraging transfer learning and reference atlas mapping to integrate multi-source single-cell data, thereby strengthening the connection between mechanistic insight and clinical prediction52.
The findings of this study have substantial implications for clinical practice in liver disease management. First, the DTNA+ macrophage markers (including DTNA, ITGAX, and RUNX2), which were rigorously screened at single-cell resolution, exhibit considerable non-invasive predictive potential. When integrated with imaging assistance, these markers could facilitate the early stratification of high-risk MASH populations and consequently reduce the current reliance on invasive liver biopsies53. Second, the detailed elucidation of the RUNX2-PLG-PARD3 axis provides a robust mechanistic basis for the development of targeted therapies. For instance, the design of RUNX2 inhibitors to specifically block the detrimental macrophage-HSC interactions could potentially serve as a strategy to delay fibrosis progression17. Furthermore, at the population level, global epidemiological data consistently emphasize the close association between MASLD/MASH and metabolic syndrome. Consequently, enhanced intervention strategies and stratified management approaches for this high-risk population must be prioritized as a public health imperative54]. Although the identification of DTNA as a predictive biomarker is exciting, the challenges and limitations of translating it into clinical application deserve attention. First, these biomarkers lack large-scale, multicenter prospective validation, which may lead to insufficient accuracy and reproducibility across diverse populations. Second, they are not liver-specific and can be influenced by comorbidities such as obesity or metabolic syndrome, potentially reducing diagnostic specificity. Additionally, while the computational pipeline is comprehensive, current evidence for the RUNX2-PLG-PARD3 axis lacks orthogonal validation at the protein or functional level. Future work should confirm causality through these validation approaches.
In this study, through multi-omics integration, we revealed the core mechanisms and regulatory networks underlying the role of DTNA+ macrophages in the progression of MASLD to MASH, providing a new paradigm for liver disease research. These findings not only advance theoretical understanding but also provide a foundation for developing diagnostic applications and therapeutic strategies for MASH.
Methods
Data acquisition and preprocessing
Bulk RNA-seq and Affymetrix microarray datasets from patients with MASLD were downloaded from the Gene Expression Omnibus (GEO) database, including GSE126848, GSE135251, GSE167523, GSE48452, GSE66676, GSE89632, GSE164760, GSE162694, and GSE24807. A dataset comprising 115 scRNA-seq samples assembled from the GEO database (GSE289173, GSE212837, GSE202379, GSE189600, GSE189175) served as the discovery cohort. In the single-cell dataset GSE289173, MASH is classified by fibrosis severity into eMASH (F0-2, representing no to mild fibrosis) and aMASH (F3-4, representing advanced fibrosis) according to the liver fibrosis staging system. Thirty-eight scRNA-seq samples from GSE166504 were employed as the validation cohort. Spatial transcriptomics data included 7 MASH samples from GSE248077. Additionally, ATAC-seq data comprising 44 samples were downloaded from the Sequence Read Archive under accession PRJNA725028.
Single-cell RNA-seq data processing and quality control
The expression profiles were loaded using the Seurat package (v4.3.0)55, following which the cells were filtered based on total UMI counts, the number of expressed genes, and the percentage of mitochondrial gene expression. The mitochondrial gene expression percentage represents the proportion of transcripts derived from mitochondrial genes relative to all transcripts in a cell. Cells with high mitochondrial transcript levels generally exhibit reduced RNA abundance, indicating that these cells are entering apoptosis. Quality control was performed using the Median Absolute Deviation (MAD) method, whereby values exceeding 3 MADs from the median were considered outliers and removed. Data scaling was conducted using the FindVariableFeatures function in Seurat to identify the top 3000 highly variable genes (HVGs), which were subsequently used for principal component analysis (PCA). The FindNeighbors function was then applied to determine nearest neighbors for PC-based graph clustering, and cell subtypes were defined using FindCluster. Visualization was achieved through Uniform Manifold Approximation and Projection (UMAP). To correct for batch effects, the Harmony algorithm was applied prior to clustering. Harmony was applied using the RunHarmony function, which leverages a soft k-means clustering approach to integrate cells across multiple datasets by removing dataset-specific technical variation while preserving genuine biological heterogeneity. Subsequently, we applied FindNeighbors and FindClusters functions in Seurat to perform graph-based clustering and identify distinct cell subtypes.
Cell type annotation and subclustering
Major cell populations were annotated based on canonical marker genes: hepatocytes (ASGR1, CYP2C19, CYP3A4), endothelial cells (PTH1R, PTPRB, STAB2), T cells (CD2, CD3D, CD3E), monocytes/macrophages (CD163, LYZ, MARCO), HSCs (COL3A1, DCN), cholangiocytes (KRT7, CFTR), and B cells (CD79A, MS4A1, IGHM). To explore intra-population heterogeneity, independent subclustering was performed on monocytes/macrophages and HSCs, followed by fine-grained subtype identification using the respective marker genes.
Developmental trajectory inference
A trajectory analysis was conducted on the monocyte/macrophage and HSC subclusters using the Monocle2 and CytoTRACE algorithms. Monocle2, a trajectory analysis tool designed for scRNA-seq data56, reconstructs cellular developmental paths by ordering cells based on their gene expression profiles. Monocle2 can identify key genes driving cell fate decisions and visualize cell progression between different states, thereby inferring cell differentiation trajectories. CytoTRACE evaluates the differentiation potential of individual cells by analyzing gene expression diversity and complexity57. The algorithm predicts cellular developmental potential, where higher scores indicate greater differentiation potential and gene expression diversity. CytoTRACE can assist Monocle2 in determining the starting point of cellular development. First, a CellDataSet object was built using RNA-seq data. Dimensionality reduction was then performed with the DDRTree method, and cells were pseudotemporally ordered along the reconstructed trajectory. In parallel, CytoTRACE was employed to assess the differentiation potential of individual cells based on gene expression diversity. Finally, the obtained CytoTRACE scores were integrated into the Monocle2 object to compare developmental potential across the pseudotemporal trajectory.
Cell-cell communication analysis
Intercellular communication networks were inferred using the CellChat R package58. A CellChat object was constructed using the normalized expression matrix as input. Expressed ligand-receptor pairs and signaling pathways were then identified using default parameters. Communication probabilities and pathway-specific interactions were calculated, and the aggregated network was visualized to reveal alterations in signaling pathways across different cell hierarchies.
Transcription factor regulatory network analysis
Single-cell regulatory network inference and clustering (SCENIC) consists of three main steps: co-expression analysis, target gene motif enrichment analysis, and regulon activity assessment. Briefly, in the first step, GENIE3 is used to infer co-expression modules between TFs and candidate target genes. Each module consists of a TF and its predicted targets, based solely on co-expression. In the second step, RcisTarget is used to analyze each co-expression module for enriched motifs; only modules and targets with TF motif enrichment are retained. Each TF and its potential direct targets constitute a regulon. In the third step, the activity of each regulon in every cell is assessed using AUCell, which calculates the area under the recovery curve (AUC). The AUCell scores are used to generate a regulon activity matrix. This matrix can be binarized by setting an AUC threshold for each regulon, determining in which cells the regulon is “active.” The regulon activity matrix can be used to cluster cells, thereby identifying cell types and states based on shared regulon network activity. pySCENIC, a high-speed Python implementation of the SCENIC workflow59, designed to infer transcriptional regulatory networks from scRNA-seq data, assess regulon activity in individual cells, and perform cell clustering based on shared regulatory network activity. We used pySCENIC to analyze activated regulons in each cell type, with the raw count matrix serving as input. Co-expression networks were computed by GRNboost, and regulons were identified by RcisTarget. Regulon activity for each cell was scored using AUCell.
Regulon module identification
The identification of regulon modules was based on the Connection Specificity Index (CSI), a context-based metric for identifying specific association partners60. The assessment of the CSI involves two steps. First, the Pearson Correlation Coefficient (PCC) is evaluated for each pair of regulon activity scores. Second, for a fixed pair of regulons A and B, the corresponding CSI is defined as the proportion of all other regulons whose PCC with A and B is lower than that between A and B. Based on the resulting CSI matrix, hierarchical clustering was performed using Euclidean distance to identify distinct regulon modules. Furthermore, a regulon association network was constructed using a cutoff of CSI > 0.7 to explore the relationships between different regulons. For each regulon module, its activity score associated with a specific cell type was defined as the average activity score of its member regulons across all cells within that cell type. The top-ranking cell types were then determined for each module.
Quantification of the fibrosis score
The fibrosis-related gene set was retrieved from the Molecular Signatures Database (MsigDB), specifically corresponding to the gene set ID HP_HEPATIC_FIBROSIS (database link: https://www.gsea-msigdb.org/gsea/msigdb/cards/HP_HEPATIC_FIBROSIS). The single-sample Gene Set Enrichment Analysis (ssGSEA) algorithm was used to quantify the fibrosis score. This method calculates an independent enrichment score for each sample-gene set pair, assessing whether the genes in a specific gene set are coordinately upregulated or downregulated in the sample. This scoring mechanism provides a quantitative measure of the degree of fibrosis.
Robust cell type decomposition
The spacexr R package61 was employed for cell type deconvolution, leveraging scRNA-seq data from the validation cohort (GSE166504) as a reference atlas. This dataset comprises transcriptomic profiles from 38 high-fat, high-fructose diet (HFHFD)-fed liver samples alongside matched controls, reflecting the cellular heterogeneity of hepatocytes and non-parenchymal cells in metabolic liver disease contexts. The Robust Cell Type Decomposition (RCTD) algorithm, a core component of spacexr, was then applied to all Visium spatial transcriptomics datasets. This facilitated the precise assignment of cell type annotations to individual spatial spots, while separately enabling deconvolution at the single-cell resolution to map fine-grained cellular distributions.
Feature selection through machine learning-based ensembles
In this study, we developed an integrated ensemble learning framework incorporating 12 diverse machine learning algorithms, encompassing regularization techniques (Lasso, Ridge, and Elastic Net), generalized linear models (Stepwise GLM [Stepglm], boosted GLM [glmBoost], and partial least squares GLM [plsRglm]), ensemble methods (Random Forest [RF], Gradient Boosting Machine [GBM], and eXtreme Gradient Boosting [XGBoost]), and pattern recognition approaches (Support Vector Machine [SVM], Linear Discriminant Analysis [LDA], and Naive Bayes). This heterogeneous ensemble integrates linear regression paradigms with sophisticated non-linear architectures to enhance model robustness and generalizability. The analytical pipeline comprised two principal stages: feature selection via Recursive Feature Elimination (RFE) to pinpoint the most predictive variables, and predictive modeling through stacked generalization. All models underwent rigorous evaluation using stratified 10-fold cross-validation (CV). Following comprehensive hyperparameter tuning, 113 unique model configurations were derived, comprising 22 single-algorithm variants (optimized independently for each algorithm) and 91 ensemble models (formed by pairwise combinations of algorithms). Performance metrics focused on the Area Under the Receiver Operating Characteristic Curve (AUC) for both training and validation sets. The configuration yielding the highest mean AUC across CV folds was selected as the optimal predictor, with its constituent genes designated as pivotal biomarkers for forecasting the progression from MASLD to MASH.
Statistical analysis
All statistical analyses and data visualizations were conducted using R software (version 4.3.0) and Python (version 3.10.11). For continuous variables, Pearson correlation coefficients were calculated when the data followed a normal distribution; otherwise, Spearman correlation was applied. Differences between two groups were assessed using the Wilcoxon rank-sum test. Statistical significance was considered at p < 0.05 (*p < 0.05, **p < 0.01, ***p < 0.001).
Data availability
The datasets used in this study are publicly available. Bulk RNA-sequencing and Affymetrix data can be accessed from GEO under the following accession numbers: GSE126848, GSE135251, GSE167523, GSE48452, GSE66676, GSE89632, GSE164760, GSE162694, and GSE24807. Single-cell RNA-seq data were obtained from GEO (accession numbers: GSE289173, GSE212837, GSE202379, GSE189600, GSE189175, GSE166504). Spatial transcriptomics data are available in GEO under accession number GSE248077. Additionally, ATAC-seq data comprising 44 samples were downloaded from the Sequence Read Archive under accession PRJNA725028. All datasets supporting the findings of this study are fully accessible as described in this article.
Code availability
The code supporting this analysis can be provided upon available request.
References
Younossi, Z. M., Kalligeros, M. & Henry, L. Epidemiology of metabolic dysfunction-associated steatotic liver disease. Clin. Mol. Hepatol. 31, S32–s50, https://doi.org/10.3350/cmh.2024.0431 (2025).
Daher, D., Dahan, K. S. E. & Singal, A. G. Non-alcoholic fatty liver disease-related hepatocellular carcinoma. J. Liver Cancer 23, 127–142, https://doi.org/10.17998/jlc.2022.12.30 (2023).
Lee, K. C., Wu, P. S. & Lin, H. C. Pathogenesis and treatment of non-alcoholic steatohepatitis and its fibrosis. Clin. Mol. Hepatol. 29, 77–98, https://doi.org/10.3350/cmh.2022.0237 (2023).
Xu, X. et al. Targeted therapeutics and novel signaling pathways in non-alcohol-associated fatty liver/steatohepatitis (NAFL/NASH). Signal Transduct. Target Ther. 7, 287, https://doi.org/10.1038/s41392-022-01119-3 (2022).
Castera, L., Friedrich-Rust, M. & Loomba, R. Noninvasive assessment of liver disease in patients with nonalcoholic fatty liver disease. Gastroenterology 156, 1264–1281 e1264, https://doi.org/10.1053/j.gastro.2018.12.036 (2019).
Govaere, O. et al. Transcriptomic profiling across the nonalcoholic fatty liver disease spectrum reveals gene signatures for steatohepatitis and fibrosis. Sci. Transl. Med. 12, eaba4448, https://doi.org/10.1126/scitranslmed.aba4448 (2020).
Wallace, S. J., Tacke, F., Schwabe, R. F. & Henderson, N. C. Understanding the cellular interactome of non-alcoholic fatty liver disease. JHEP Rep. 4, 100524. https://doi.org/10.1016/j.jhepr.2022.100524 (2022).
Watson, B. R. et al. Spatial transcriptomics of healthy and fibrotic human liver at single-cell resolution. Nat. Commun. 16, 319. https://doi.org/10.1038/s41467-024-55325-4 (2025).
Ghosh, S., Zhao, X., Alim, M., Brudno, M. & Bhat, M. Artificial intelligence applied to ‘omics data in liver disease: towards a personalised approach for diagnosis, prognosis and treatment. Gut 74, 295–311, https://doi.org/10.1136/gutjnl-2023-331740 (2025).
Krenkel, O. & Tacke, F. Liver macrophages in tissue homeostasis and disease. Nat. Rev. Immunol. 17, 306–321, https://doi.org/10.1038/nri.2017.11 (2017).
Payen, V. L. et al. Single-cell RNA sequencing of human liver reveals hepatic stellate cell heterogeneity. JHEP Rep. 3, 100278. https://doi.org/10.1016/j.jhepr.2021.100278 (2021).
Andrews, T. S. et al. Single-cell, single-nucleus, and spatial RNA sequencing of the human liver identifies cholangiocyte and mesenchymal heterogeneity. Hepatol. Commun. 6, 821–840, https://doi.org/10.1002/hep4.1854 (2022).
Atif, J., Thoeni, C., Bader, G. D., McGilvray, I. D. & MacParland, S. A. Unraveling the complexity of liver disease one cell at a time. Semin. Liver Dis. 42, 250–270, https://doi.org/10.1055/s-0042-1755272 (2022).
Zhao, C. et al. Single-cell data analysis reveals critical hepatic cells subpopulations in the progression of non-alcoholic fatty liver disease to non-alcoholic steatohepatitis. Comb. Chem. High Throughput Screen. 28, 1251–1263, https://doi.org/10.2174/0113862073303213240523095742 (2025).
Cheng, S. et al. Single-cell RNA sequencing reveals the heterogeneity and intercellular communication of hepatic stellate cells and macrophages during liver fibrosis. MedComm (2020) 4, e378. https://doi.org/10.1002/mco2.378 (2023).
He, W. et al. Identifying a distinct fibrosis subset of NAFLD via molecular profiling and the involvement of profibrotic macrophages. J. Transl. Med. 21, 448. https://doi.org/10.1186/s12967-023-04300-6 (2023).
Zhong, L. et al. Runx2 activates hepatic stellate cells to promote liver fibrosis via transcriptionally regulating Itgav expression. Clin. Transl. Med. 13, e1316. https://doi.org/10.1002/ctm2.1316 (2023).
Zhong, L. et al. Cell-specific elevation of Runx2 promotes hepatic infiltration of macrophages by upregulating MCP-1 in high-fat diet-induced mice NAFLD. J. Cell Biochem. 120, 11761–11774, https://doi.org/10.1002/jcb.28456 (2019).
Itoh, M. et al. CD11c+ resident macrophages drive hepatocyte death-triggered liver fibrosis in a murine model of nonalcoholic steatohepatitis. JCI Insight 2, e92902, https://doi.org/10.1172/jci.insight.92902 (2017).
Yao, S. et al. Role of lactate and lactate metabolism in liver diseases (Review). Int. J. Mol. Med. 54, 59, https://doi.org/10.3892/ijmm.2024.5383 (2024).
Liang, H., Xu, L. & Yang, Y. Lactate and lactylation: novel perspectives on fibrosis pathogenesis and therapeutic directions. J. Transl. Med. 23, 705. https://doi.org/10.1186/s12967-025-06748-0 (2025).
Dam, T. V., Toft, N. I. & Grøntved, L. Cell-type resolved insights into the Cis-regulatory genome of NAFLD. Cells 11, 870 (2022).
Li, J. Z. et al. Spatial and single-cell transcriptomics reveals the regional division of the spatial structure of MASH fibrosis. Liver Int. 45, e16125 https://doi.org/10.1111/liv.16125 (2025).
Tosello-Trampont, A. C., Landes, S. G., Nguyen, V., Novobrantseva, T. I. & Hahn, Y. S. Kuppfer cells trigger nonalcoholic steatohepatitis development in diet-induced mouse model through tumor necrosis factor-α production. J. Biol. Chem. 287, 40161–40172 https://doi.org/10.1074/jbc.M112.417014 (2012).
Zhang, X. et al. Macrophage p38α promotes nutritional steatohepatitis through M1 polarization. J. Hepatol. 71, 163–174 https://doi.org/10.1016/j.jhep.2019.03.014 (2019).
Vonderlin, J., Chavakis, T., Sieweke, M. & Tacke, F. The multifaceted roles of macrophages in NAFLD pathogenesis. Cell Mol. Gastroenterol. Hepatol. 15, 1311–1324 https://doi.org/10.1016/j.jcmgh.2023.03.002 (2023).
Cao, Y. et al. Macrophages evoke autophagy of hepatic stellate cells to promote liver fibrosis in NAFLD mice via the PGE2/EP4 pathway. Cell Mol. Life Sci. 79, 303 https://doi.org/10.1007/s00018-022-04319-w (2022).
Xi, S. et al. Activated hepatic stellate cells induce infiltration and formation of cd163(+) macrophages via CCL2/CCR2 pathway. Front. Med. 8, 627927 https://doi.org/10.3389/fmed.2021.627927 (2021).
Spite, M. Resolving inflammation in nonalcoholic steatohepatitis. J. Clin. Investig. 129, 1524–1526 https://doi.org/10.1172/JCI127583 (2019).
Tsuchida, T. & Friedman, S. L. Mechanisms of hepatic stellate cell activation. Nat. Rev. Gastroenterol. Hepatol. 14, 397–411 https://doi.org/10.1038/nrgastro.2017.38 (2017).
Merens, V. et al. Hepatic stellate cell single cell atlas reveals a highly similar activation process across liver disease aetiologies. JHEP Rep. 7, 101223 https://doi.org/10.1016/j.jhepr.2024.101223 (2025).
Zhang, W. et al. Heterogeneity of hepatic stellate cells in fibrogenesis of the liver: insights from single-cell transcriptomic analysis in liver injury. Cells 10, 2129 https://doi.org/10.3390/cells10082129 (2021).
Dobie, R. et al. Single-cell transcriptomics uncovers zonation of function in the mesenchyme during liver fibrosis. Cell Rep. 29, 1832–1847 e1838 https://doi.org/10.1016/j.celrep.2019.10.024 (2019).
Tada, Y. et al. Roles of macrophages in advanced liver fibrosis, identified using a newly established mouse model of diet-induced non-alcoholic steatohepatitis. Int. J. Mol. Sci. 23, 13251 https://doi.org/10.3390/ijms232113251 (2022).
Daemen, S. et al. Dynamic shifts in the composition of resident and recruited macrophages influence tissue remodeling in NASH. Cell Rep. 34, 108626 https://doi.org/10.1016/j.celrep.2022.111660 (2021).
Guilliams, M. & Scott, C. L. Does niche competition determine the origin of tissue-resident macrophages? Nat. Rev. Immunol. 17, 451–460, https://doi.org/10.1038/nri.2017.42 (2017).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383. https://doi.org/10.1038/s41467-018-06318-7 (2018).
Banerjee, D., Wei, Z. & Luyendyk, J. P. Role of the plasminogen activation system in liver injury and repair: knowns and known unknowns. Arterioscler. Thromb. Vasc. Biol. 46, 27–36 (2025).
Wang, H., Zhang, Y. & Heuckeroth, R. O. PAI-1 deficiency reduces liver fibrosis after bile duct ligation in mice through activation of tPA. FEBS Lett. 581, 3098–3104, https://doi.org/10.1016/j.febslet.2007.05.049 (2007).
Pohl, J. F., Melin-Aldana, H., Sabla, G., Degen, J. L. & Bezerra, J. A. Plasminogen deficiency leads to impaired lobular reorganization and matrix accumulation after chronic liver injury. Am. J. Pathol. 159, 2179–2186, https://doi.org/10.1016/S0002-9440(10)63069-6 (2001).
Atashrazm, F. & Ellis, S. The polarity protein PARD3 and cancer. Oncogene 40, 4245–4262, https://doi.org/10.1038/s41388-021-01813-6 (2021).
Wu, J. et al. PARD3 drives tumorigenesis through activating Sonic Hedgehog signalling in tumour-initiating cells in liver cancer. J. Exp. Clin. Cancer Res. 43, 42, https://doi.org/10.1186/s13046-024-02967-3 (2024).
Masuzaki, R. et al. Integrin β1 establishes liver microstructure and modulates transforming growth factor β during liver development and regeneration. Am. J. Pathol. 191, 309–319, https://doi.org/10.1016/j.ajpath.2020.10.011 (2021).
Sharip, A. & Kunz, J. Mechanosignaling via integrins: pivotal players in liver fibrosis progression and therapy. Cells 14, 266 (2025).
Prado, L. G. & Nagy, L. E. Role of complement in liver diseases. Semin. Liver Dis.44, 510–522, https://doi.org/10.1055/s-0044-1795143 (2024).
Zhao, J. et al. Association of complement components with the risk and severity of NAFLD: a systematic review and meta-analysis. Front. Immunol. 13, 1054159 https://doi.org/10.3389/fimmu.2022.1054159 (2022).
Feng, L., Zhao, Y. & Wang, W. L. Association between complement C3 and the prevalence of metabolic-associated fatty liver disease in a Chinese population: a cross-sectional study. BMJ Open 11, e051218, https://doi.org/10.1136/bmjopen-2021-051218 (2021).
Lee, J. et al. Machine learning algorithm improves the detection of NASH (NAS-based) and at-risk NASH: A development and validation study. Hepatology 78, 258–271, https://doi.org/10.1097/HEP.0000000000000364 (2023).
Fialoke, S., Malarstig, A., Miller, M. R. & Dumitriu, A. Application of machine learning methods to predict non-alcoholic steatohepatitis (NASH) in non-alcoholic fatty liver (NAFL) patients. AMIA Annu. Symp. Proc. 2018, 430–439 (2018).
Chen, J., Zhu, Y., Zheng, Q. & Jiang, J. Serum cytokeratin-18 in the diagnosis of non-alcoholic steatohepatitis: a meta-analysis. Hepatol. Res.44, 854–862, https://doi.org/10.1111/hepr.12197 (2014).
Feldstein, A. E. et al. Cytokeratin-18 fragment levels as noninvasive biomarkers for nonalcoholic steatohepatitis: a multicenter validation study. Hepatology 50, 1072–1078, https://doi.org/10.1002/hep.23050 (2009).
Lotfollahi, M. et al. Mapping single-cell data to reference atlases by transfer learning. Nat. Biotechnol. 40, 121–130, https://doi.org/10.1038/s41587-021-01001-7 (2022).
Rinella, M. E. et al. AASLD practice guidance on the clinical assessment and management of nonalcoholic fatty liver disease. Hepatology 77, 1797–1835, https://doi.org/10.1097/HEP.0000000000000323 (2023).
Younossi, Z. M. et al. The global epidemiology of nonalcoholic fatty liver disease (NAFLD) and nonalcoholic steatohepatitis (NASH): a systematic review. Hepatology 77, 1335–1347 https://doi.org/10.1097/HEP.0000000000000004 (2023).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 e3529 https://doi.org/10.1016/j.cell.2021.04.048 (2021).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 https://doi.org/10.1038/nmeth.4402 (2017).
Kang, M. et al. Improved reconstruction of single-cell developmental potential with CytoTRACE 2. Nat. Methods 22, 2258–2263 https://doi.org/10.1038/s41592-025-02857-2 (2025).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat. Protoc. 20, 180–219 https://doi.org/10.1038/s41596-024-01045-4 (2025).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 https://doi.org/10.1038/nmeth.4463 (2017).
Suo, S. et al. Revealing the critical regulators of cell identity in the mouse cell Atlas. Cell Rep. 25, 1436–1445 e1433 https://doi.org/10.1016/j.celrep.2018.10.045 (2018).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 https://doi.org/10.1038/s41587-021-00830-w (2022).
Acknowledgements
We sincerely thank all members of the Tumor ImmunoMetabolism Institute (TIMI) of Zhuzhou Hospital for their valuable discussions and technical support. This work was supported by the National Natural Science Foundation of China (Grant Nos. 82300955 and 82303246),the Natural Science Foundation of Hunan Province (Grant Nos. 2023JJ41066, 2025JJ40100, 2024JJ6728 and 2025JJ50493), and the Regional Joint Fundation of Guangdong Province (Grant No. 2022A1515111099), and Health Research Project of Hunan Provincial Health Commission (Grant Nos. Z2023100 and W20243173).
Author information
Authors and Affiliations
Contributions
W.W., Z.L., W.T., Y.T. and W.L. performed data analysis and interpretation. J.W., H.H., Z.J., X.Tang, J.Y., J.X., and X.Tan contributed to data acquisition and figure preparation. Y.L., P.W., and X.C. designed and reviewed the study, supervised the project, and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wen, W., Liu, Z., Tan, W. et al. Integrating multi-omics and machine learning systematically deciphers cellular heterogeneity and fibrotic regulatory networks in the progression from MASLD to MASH. npj Digit. Med. 9, 167 (2026). https://doi.org/10.1038/s41746-026-02352-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-026-02352-8